INTRODUCTION
MEF2 transcription factor plays vital role in both physiological and pathological processes. The members of MEF2 family have been mainly involved in neural development, muscle formation, heart development, and carcinogenesis. Emerging evidences indicate that MEF2 family contributes to the development of various human diseases due to its complex function by interacting with numerous signaling pathways and small non-coding RNA such as microRNAs [1–4]. This review summarizes the relationship between MEF2 family and their molecular mechanisms of development and progression of several death causing human diseases. We also described signaling pathways and microRNAs directly involved in regulation of expression and activity of MEF2. In addition, we have summarized some of the new drugs that targeting MEF2 family.
BASIC KNOWLEDGE OF MEF2 FAMILY
The MEF2 family and gene mapping
MEF2 proteins belong to MADS-box family of transcription factors. MADS-box is named from the four proteins minichromosome maintenance genes (MCM1), agamous (AG), deficiens (DEFA) and serum response factor (SRF), which share a contiguous conserved motif in eukaryotic organisms [5, 6]. MEF2 proteins were initially identified as a vital factor for skeletal muscle development that have the ability to bind A/T-rich sequences within gene promoter of muscle creatine kinase (MCK) [7, 8]. MEF2 is a single gene in Drosophila, Caenorhabditis elegans and Saccharomyces cerevisiae, while vertebrates have four distinct numbers—namely, MEF2A, B, C and D [3]. The evolutionary history of MEF2 family is inferred using the Neighbor-Joining method (Figure 1). The sole MEF2 gene in Drosophila melanogaster is located on chromosome 2. Murine MEF2A, C and D are located in chromosome 7, 13 and 3, respectively. A study deduced that MEF2B may be located on chromosome 8. In human, MEF2A, B, C and D are confirmed to be located on 15q26, 19q12, 5q14 and 1q12-q23, respectively [9].
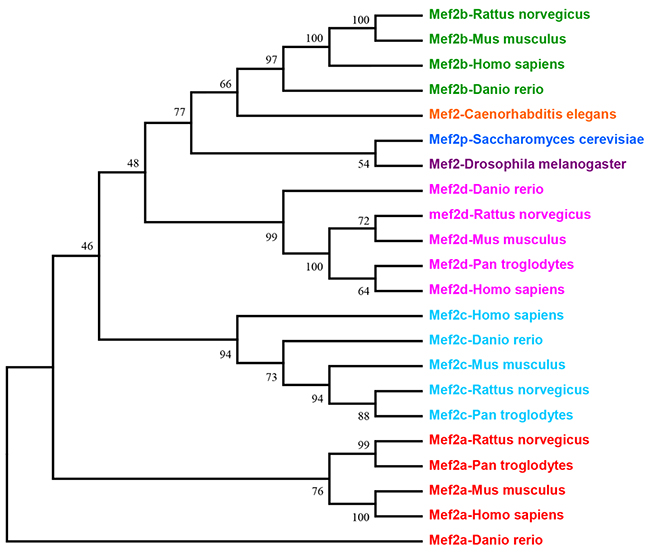

Figure 1: This is the phylogenetic tree of MEF2 family. Evolutionary analyses tree of MEF2 family was conducted in software MEGA7 by using Neighbor-Joining method.
The structure of MEF2 proteins
MEF2 family proteins from different species share a very similar N-termini that contains a highly conserved MADS-box domain and an immediate adjacent MEF2 domain, however, there is a diversity in the structure of C-terminal transactivation domain [4] (Figure 2). The MADS-box domain which is a DNA-binding region of MEF2 protein consists of 55 amino acids with some conserved residues shared by other transcription factors of MADS-box family, and this region plays a key role in the recognition of their target sequences. The main role of these invariable residues are combined with abundant A/T DNA sequences and they mediate dimerization of MADS-box proteins and provide places for interaction of MEF2 with other cofactors. Due to the conserved sequence within MEF2 domain, MEF2A, B, C and D can form homo-dimerization structures with themselves, or hetero-dimerization with other molecules [10]. In C-terminal part of MEF2 proteins, conserved regions contain potential phosphorylation sites recognized by some specific kinases, but other amino acid sequences are highly diverse. Intriguingly, alternative splicing of MEF2 mRNAs take place in the sequences coding their C-terminal regions, which contribute to the regulation of MEF2 proteins’ activity [11].
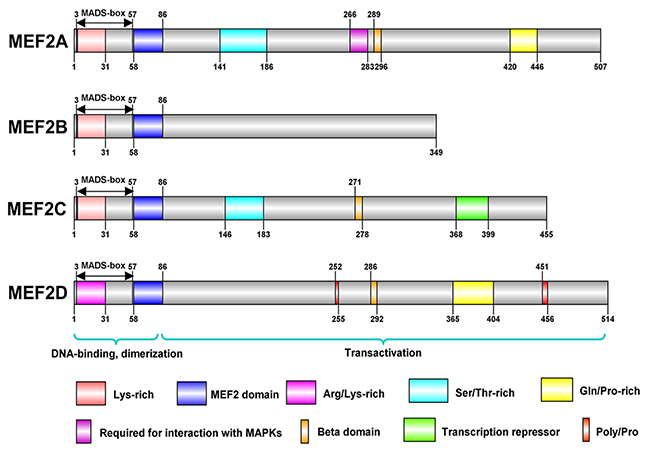
Figure 2: These are structures of MEF2 proteins in human. The structures of MEF2 proteins were conducted in IBS(illustrator for Biological Sequences). MEF2 family proteins, including MEF2A, B, C, and D, share a similar N-terminal that contains a highly conserved MADS-box domain and an immediate adjacent MEF2 domain. MADS-box and MEF2 domain have 55 and 29 amino acids, respectively, and both contribute to DNA binding and dimerization. There is a diversity in the structure of C-terminal transactivation domain of MEF2 proteins. MEF2B is the simplest protein among MEF2 proteins. MEF2A, C, and D contain several amino acids with the potential of phosphorylation.
Transcriptional targets of MEF2 family
In the neural development, MEF2 has directly engaged in many genetic programs. The previous publications had identified that the (C/T)TA(T/A)4TA(G/A),a ten-nucleotide motif, could be MEF2 binging site. In addition, the activity-dependent MEF2 genetic program may use polyA site switch mechanisms to inference the functions of proteins. The activity-regulated neuronal target genes of MEF2 and Poly A switch are JSAP1, Odz2, Arhgef9, Csnk1e, Vesl-1, Omp25, Pkib, Septin 11, Rinzf, Klf6 and Tcf4 [12]. In the cardiovascular development, especially during mouse embryogenesis and adulthood, myocardin had been proved to be a direct transcriptional target of MEF2,Tead and Foxo proteins [13]. Consistently, myomaxin also had been identified as the direct downstream target gene of MEF2A [14]. During the Drosophila embryonic development, researchers had constructed a temporal map of MEF2 activity by immune-precipitation analysis and gene expression profiles. The results indicated that Mhc, mbl, nau and meso18E were directly regulated by MEF2 [15]. Recent researches have determined the putative recognition motif of MEF2 transcription factors within the promoter or enhancer of their downstream target genes, which are enriched in the constitution of adenine and thymine. Recently, researchers used a powerful approach to illuminate the genomic functions of MEF2A and MEF2C and to identify their transcriptional targets by ChIP-seq in mouse cortical neurons. MEF2A and MEF2C may encode similar epigenetic programs by binding enhancer regulatory elements close to the target genes involved in neuronal plasticity and calcium signaling. The results indicated SRF could bind to MEF2,AP1and NeuroD1,CPBE/ATF could bind to MEF2A enhancers and CEBP/A and forkhead box factors could bind to MEF2C. And the LRP8-Reelin-Regulated Neuronal enhancers (LRN) could be recognized by both MEF2A and MEF2C in synapse-to-nucleus pathway. According to KEGG pathway analysis, the MEF2 factors could regulate signaling pathways including glutamatergic synaptic transmission, drug addiction, axon guidance and MAPK signaling pathways [6]. During the evolution of abthropoid primates, a new enhancer element emerged due to 5-10 nucleotide changes in osteocrin (OSTN). The element was an 85 bp-long sequence and located about 600 bp upstream of OSTN transcription initiation site. Intriguingly, MEF2 can bind to this sequence, indicating that OSTN is a potential target of MEF2s. Further study on the mechanism between MEF2 and OSTN may contribute to our better understanding of human cognition and brain function. [16].
MEF2 PHYSIOLOGICAL FUNCTION AND SIGNAL PATHWAYS
MEF2 family is mainly associated with a variety of physiological processes, including muscle formation [17], nervous system development [18], heart development [19], etc. They are also implicated in human diseases, such as liver fibrosis [20], cancers [21] and neurodegenerative diseases [22]. Atkins et al. established a Ras-driven tumorigenesis model in Drosophila epithelial tissues to study gene regulatory networks in cancer cells. They discovered that Mef2 is involved in a cross-regulation with other factors such as Sat, AP-1, Myc, AP-4, Ftz-f1, and Taiman/SRC3 [23]. MEF2 family can be regulated by multiple signaling pathways during normal physiological processes as well as in pathological conditions (Figure 3). Accumulating evidence show that MEF2 proteins promote the differentiation of skeletal, cardiac and smooth muscle myocytes [24, 25]. MEF2 proteins are widely expressed in muscle and nervous tissues, and they can integrate extracellular signaling into promoters or enhancers of tissue-specific genes. Some researchers revealed a link between MEF2A and SRF in the activation of selective muscle-specific promoters in different muscle cell lines. Similarly, MEF2C and SRF can cooperatively regulate the expression of miR-133a [26]. MEF2 activity is controlled by the myogenic regulatory factor MRF4 in the adult skeletal muscle [27]. A recent study showed a significant up-regulation of MEF2C and MEF2D mRNA levels in both sporadic and SOD1+ amyotrophic lateral sclerosis (ALS) patients detected by gene expression analysis, and a down-regulation of their targets, BDNF, KLF6, and RUFY3 [28].
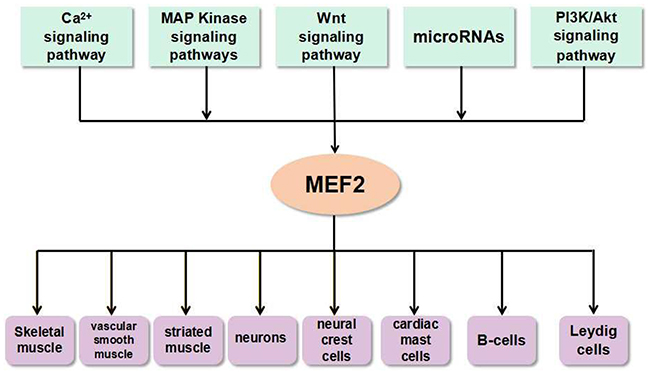
Figure 3: The link between MEF2 proteins and signals in various types of cells and tissues. The microRNAs and signaling pathways such as Ca2+ signaling pathway, MAP Kinase signaling pathways, Wnt signaling pathway, PI3K/Akt signaling pathway, etc. can activate MEF2. This interaction between MEF2 proteins and signals ubiquitously exist in many types of cells and tissues.
The family members of MEF2 transcription factor are closely associated with calcium-dependent signaling pathway [8], which play an important role in the development of nervous system and neuronal differentiation [29]. All MEF2 family members, especially MEF2C, are highly expressed in the neuron of central nervous system [30]. MEF2A and MEF2D are found to exhibit transcriptional activity in post-mitotic cerebellar granule neurons [31]. Surprisingly, MEF2A, C, and D play a distinct role in cell- and non-cell-autonomous control of adult hippocampal neurogenesis [32]. MEF2B acts as a valuable marker of normal germinal center (GC) B cells, and it is potentially useful for differential diagnosis of small B cell lymphomas [33]. As an Epstein-Barr virus (EBV) nuclear antigen 1 (EBNA1)-bound gene, MEF2B is important for the survival of B cells infected with EBV [34].
As a cardiac-specific marker gene, MEF2 can be regulated by focal adhesion kinase to enhance the expression of Jun in cardiomyocytes [35]. MEF2C is also associated with congenital human heart defects due to its regulatory function on teratocarcinoma-derived growth factor 1 expression (Tdgf1), which is essential for the early embryonic heart development. The abnormalities in the expression of MEF2C and its associated Tdgf1 leads to developmental defects and congenital heart problems [36]. Mef2c can physically interact with Suv39h1 and their interaction contributes to the regulation of myogenic expression, histone methylation modification, and myoblast differentiation. Furthermore, MEF2 transcription activity could be inhibited by Suv39h1 in a dose-dependent manner [37].
MEF2 family members also have close connections with biological characteristics (e.g., uncontrolled proliferation and enhancement of invasion) and clinical outcomes of cancer. MEF2D is overexpressed in colorectal cancer tissues and it promotes cancer cell invasion and metastasis [38]. A study in our laboratory also found that MEF2D acts as an oncogene in HCC. The expression of MEF2D is up-regulated in hepatocellular carcinoma (HCC) and this increase can accelerate cell proliferation in HCC [39]. MEF2D co-operationally promotes HCC invasion with other oncogene, such as Pokemon [40]. Molecular studies revealed that MEF2 family members promote invasion properties of HCC by enhancing the effect of TGF-β1 on EMT [41]. Increasing evidences indicate that MEF2C and MEF2D both act as tumor-promoting or -suppressing proteins dependent on the type of cancer. In leukemic patients, the ectopic expression of MEF2s caused by chromosomal rearrangements is responsible for the development of leukemia. A high level expression of MEF2C is observed in T-cell acute lymphoblastic leukemia (T-ALL) [42, 43]. The chromosomal translocation generates reciprocal DAZAP1/MEF2D and MEF2D/DAZAP1 fusion genes that promotes oncogenic properties in NIH 3T3 cells [44]. In contrast, the depression of MEF2C and MEF2D can promote cell proliferation and anchorage independent growth in lipo- and leiomyosarcoma by upregulating HDAC4 and PI3K/Akt signaling [45].
The involvement of MEF2 family in Ca2+ signaling pathway
MEF2 family often acts as an effector of many extracellular and intracellular signals. Ca2+ signaling pathway is the major regulator of MEF2 activity by multiple mechanisms. MEF2 is required for calcineurin signaling in developing skeletal muscle [46]. MEF2 mediates calcium/calmodulin-dependent signaling responsible for the terminal differentiation of skeletal muscle progenitor cells. In specifically, the activation of transcriptional activities of MEF2 proteins by calcineurin, a calcium/calmodulin-dependent protein phosphatase, is required for myogenic differentiation [47]. A further study showed that scaffolding protein mAKAP organizes a calcineurin/MEF2 signaling complex in myocytes, and which regulates gene transcription promoted by MEF2 family members [48].
Apart from this, several cofactors and kinases also regulate MEF2 activity in calcium signaling. Hu et al. (2015) found that endothelin signaling activates various gene expression in neural crest by enhancing MEF2C activity through Calmodulin-CamKII-histone deacetylase signaling cascade [49]. In addition, salt inducible kinase 2 (SIK2) tightly regulates CaMKI dependent transcriptional activities of MEF2C and disruption of SIK2 promotes activation of CaMKI depdendent activation of MEF2C mediated gene expression during cell proliferation and stress response [50]. Interestingly, a novel MHCI signaling regulates MEF2 transcription factors. MHCI pathway requires calcineurin-mediated activation of MEF2 to control synapse density in young rat cortical neurons [51]. In addition, Tsai1 et al. found that the activation of MEF2 proteins could induce synapse elimination and degradation of postsynaptic density protein 95 (PSD-95) through protein phosphatase 2A (PP2A)-mediated dephosphorylation of murine double-2 (MDM2) in wild-type neurons. However, eukaryotic elongation factor 1α (EEF1a) could inhibit MEF2-induced accumulation of MDM2 at the synapses in Fmr1 KO mouse neurons [52].
Surprisingly, MEF2 is not only expressed in myocytes and neurons, but also detected in Sertoli and Leydig cells of fetal and adult testis. In MA-10 Leydig cells, steroid hormone biosynthesis and steroidogenic gene expression are regulated by luteinizing hormone (LH), which activates Ca2+ signaling pathways and MEF2 transcription factors [53]. Another study showed that MEF2 itself can bind and activate Gsta1 promoter as well as cooperate with Ca2+/calmodulin-CamKI to further enhance the initiation of Gsta1 transcription in Sertoli and Leydig cells [54].
The function of MEF2 proteins in MAP kinase signaling pathways
The mitogen-activated protein kinase (MAPK) signaling pathways are ubiquitous type of serine/threonine protein kinase, which plays an important role in many biological processes (cell proliferation, cell differentiation, cell apoptosis, and etc.) [55, 56]. There are three independent MAPKs signaling pathways such as extracellular signal-regulated kinase (ERK) pathway, c-Jun N-terminal kinase/stress-activated protein kinase (JNK/SAPK) pathway and p38 MAPK pathway are found in mammalian cells [57], and MEF2 factors have been frequently found to be activated by these signaling pathways.
ERK protein kinase comprises five members namley ERK1, ERK2, ERK3, ERK4 and ERK5. Among them, ERK5 is closely associated with the activation of MEF2 transcription factors. In myeloid leukemia cells, ERK5 promotes activation of its direct downstream target, MEF2C transcription factor, and this activation required for the monocytic differentiation of leukemic cells [58]. In sensory neurons, the ERK5/MEF2D pathway strictly regulates expression of Bcl-w, an anti-apoptopic bcl-2 family member, which promotes sensory neuron survival [59]. Recently, a groundbreaking finding revealed that the activation of MEF2C/D transcription factors promote the development of early B-cell, which depends on their phosphorylation by ERK5. These researchers also reported that B-cell development is blocked at the pre-B-cell stage in MEF2C/D-knockout mice [60]. The ERK5-MEF2C signaling also plays a critical role in anti-apoptotic and neuroprotective actions of ischemic preconditioning in the hippocampus CA1 region [61]. In Hela cells, ERK5/MEF2B signaling can upregulate apoptosis suppressor induced by DNA damage and promotes cancer cell invasion by activating β-catenin target genes [62].
p38 is an important member of MAPK family, which can be activated by physiological stress, lipopolysaccharide, osmotic stress and ultraviolet irradiation [63]. The relationship between p38 MAPK and MEF2s was first identified in myocardial cells. p38 MAPK can activate MEF2C by phosphorylation of three amino acids located in C-terminal active region [64]. p38 MAPK-mediated phosphorylation-dependent activation of MEF2s increases its interaction with β-catenin, which stimulates cell proliferation in multiple cell types including primary VSMCs [65]. More intriguingly, MEF2 proteins are found to negatively regulate p38 MAPK pathway in a feedback fashion [66]. In recent years, more evidence have been shown that p38 MAPK pathway activates the expression of MEF2 genes under a variety of cellular circumstances. For instances, p38 MAPK signaling activates MEF2C to control B cell differentiation [67] and this signaling axis also promotes osteogenic differentiation [68]. In myocardial hypertrophy, MEF2 family is associated with increased heart mass in response to pressure overload, and its activation in hypertrophic heart requires adiponectin signaling mediated upregulation of p38 MAPK pathway [69]. Some researchers identified that microRNAs are critically regulating MEF2 and p38 MAPK axis. For example, miR-140 is a suppressor of p38 MAPK signal transduction pathway, and overexpression of this microRNA can reduce MEF2C expression [70]. Glycogen Synthase Kinase 3β (GSK3β) is a well-known regulator of striated muscle-related gene expression, which suppresses both myogenesis and cardiomyocyte hypertrophy. It is well known that MEF2 transcription factors are essential for the regulation of skeletal and cardiac muscle gene expression. GSK3β can regulate MEF2 activity indirectly through regulation of p38 MAPK [71].
The link between MEF2 and Wnt signaling
The Wnt signaling is a kind of evolutionarily conservative signaling pathway that exists widely in invertebrates and vertebrates. The Wnt signaling plays a vital role in developmental and physiological processes such as animal embryonic development [72], organ formation and tissue regeneration [73]. The Canonical Wnt/β-catenin pathway can activate expression of many nuclear target genes, and the aberrant activation of the Wnt signaling is closely implicated in malignant transformation and progression [74, 75] and blocking of Wnt/β-catenin signaling inhibits cancer cell proliferation. While a very high expression of MEF2C leads to risk of VEGF-induced malignancy [76].
The proteins of MEF2 family also serve as downstream targets of Wnt/β-cateinin pathway genes, and mediate its physiological functions. MEF2A and Wnt signaling both contribute to skeletal muscle regeneration in adult mice. A molecular study found that MEF2A promotes this process via directly regulating two microRNAs, miR-410 and miR-433. In MEF2A knockout mice, these two microRNAs are downregulated during skeletal muscle regeneration, which results in upregulation of SFRP2 expression and reduction of Wnt activity [77].
microRNAs associated regulation of MEF2 proteins
Many recent studies observed that microRNAs, a type of small non-coding RNAs, participate in the regulation of MEF2s, especially in the progression of malignant diseases. miR-218 regulates MEF2D expression by targeting 3’UTR of its mRNA and down-regulation of miR-218 in cardiac myxoma condition can increase MEF2D expression and promote cell cycle progression [78]. In glioma cells, miR-18a binds with 3’UTR of MEF2D mRNA and negatively regulates its expression [79]. Similarly, miR-122 and miR-1244 negatively regulate MEF2D expression in hepatocellular carcinoma and lung cancer cells, respectively [39, 80]. Furthermore, MEF2D regulate miR-1244 by directly binding to its promoter and this molecular regulatory loop could be a target for lung carcinoma treatment [80]. Apart from their role in cancer, the regulatory functions of MEF2s are also well established in other human diseases. MEF2C is the cardiac-specific markers and its expression is upregulated in congenital heart disease (CHD) due to downregulation of the expression of miR-29C [81]. Li et al. (2015) found that MEF2D is an authentic target of miR-103, which promotes cell differentiation by targeting the AKT/mTOR signaling mediated activation of MEF2D [82]. In hyperhomocysteinemia induced cardiac hypertrophy, the downregulation and inactivation of MEF2C leads to attenuation of expression of miR-133a, an anti-hypertrophic factor in cardiomyocytes. Hydrogen sulfide (H2S) can mitigate hypertrophy via reversal of expression of miR-133a through activation of MEF2C in hyperhomocysteinemia cardiomyocytes [83]. Similarly, MEF2A and MEF2C regulate miR-143 and miR-23a activities via their interaction with 3’UTR region of microRNA in vascular smooth muscle cell (VSMC) and cardiomyocytes from mice with myotonic dystrophy, respectively [84, 85]. These studies reveal that microRNA/MEF2 pathway might play a key role in various types of cells and its dysregulation always contribute to various diseases (Figure 4). Therefore, targeting this molecular pathway could be a promising therapeutic strategy.
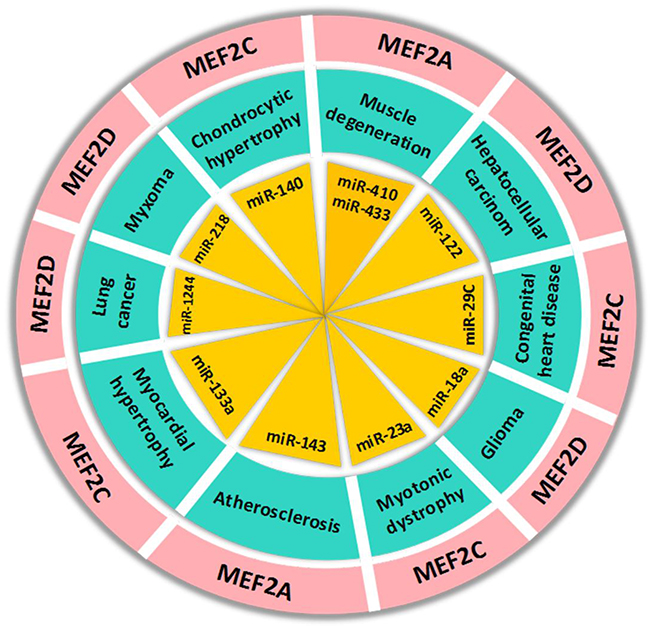
Figure 4: The regulation of MEF2 proteins by microRNAs in human diseases. The microRNAs in the figure can regulate MEF2 expression by targeting 3’UTR of its mRNA in human diseases.
Besides, some studies have revealed that long noncoding RNAs (lncRNAs) play important roles in regulating some critical transcription factors, including MEF2A-D. Linc-MD1 is a kind of competing endogenous lncRNAs that its reduced expression level in Duchenne muscular dystrophy (DMD) promotes differentiation of myoblast. Notably, linc-MD1 detaches miR-135 away from MEF2C mRNA, and thus, depresses the expression of MEF2C in skeletal muscle development and disease [86]. Song and his/her colleagues found that the expression level of uc.167 was robustly elevated in ventricular septum defect (VSD) heart tissues and uc.167 had an effect on cell proliferation, apoptosis and differentiation of P19 cell by regulating MEF2C. In addition, the overexpression of uc.167 induced the reduction of MEF2C expression. Altogether, the concrete relationship between MEF2C and uc.167 still needs further investigation [87].
Other signaling pathways associated with MEF2s
The PI3K/Akt pathway is a newly confirmed MEF2-associated pathway, which is involved in the proliferation, differentiation, apoptosis and the regulation of glucose transport. The PI3K/Akt pathway is closely implicated in the occurrence of many human diseases [88, 89]. The PI3K/Akt signaling can enhance MEF2 transcriptional activity in muscle differentiation [90]. Interestingly, the depletion of MEF2D promotes neonatal cardiomyocyte proliferation by suppressing the expression of PTEN, which is the primary negative regulator of PI3K/Akt signaling. However, prolonged depletion of MEF2D in neonatal cardiomyocytes resulted in significant programmed cell death. This study suggest that MEF2D mediated regulation of PI3K/AKT signaling contributes to post-mitotic state of cardiomyocytes [91]. In contrast to this, a molecular study in sarcoma cells found that PI3K/Akt pathway influences the expression of target genes of MEF2-HDAC axis and increased activity of PI3K/Akt results in decreased expression of MEF2 [45].
Cyclic-AMP dependent protein kinase A (PKA) is a kind of protein kinase with simple structure, which is also connected to the transcriptional activities of MEF2. MEF2 can be a target of cAMP-PKA pathway in neuron [92]. Given the fact that MEF2 proteins are key regulators myogenesis, cAMP-PKA-pathway-mediated myogenic repression occurs through downregulation of MEF2D in myoblasts and this loss leads to inhibition of the skeletal muscle differentiation program. Given the fact that MEF2 proteins are key regulators of myogenesis, cAMP-PKA-pathway-mediated myogenic repression occurs through downregulation of MEF2D in myoblasts and this loss leads to inhibition of the skeletal muscle differentiation program [93].
In iron/sphingolipid/PDK1/Mef2 pathway, loss of FXN in vertebrates can upregulate 3-phosphoinositide dependent protein kinase-1 (Pdk1) and myocyte enhancer factor-2 (Mef2), and induce sphingolipid synthesis. Furthermore, the activation of Mef2 triggers the aberrant transcription of downstream targets. Similarly, loss of frataxin homolog (fh) in Drosophila nervous system causes the accumulation of iron, and in turn, enhances spingolipid synthesis. In addition, PDK1 and Mef2 were both activated to trigger neuro-degeneration of adult photoreceptors. Since the above evidence has well documented that the mechanism of iron/sphingolipid/PDK1/Mef2 pathway is conserved among different species, targeting this pathway may benefit on the hearts of Friedreich ataxia (FRDA) patients [94].
As mentioned above, MEF2 is a single gene in Drosophila, and it is a vital gene in myogenesis of indirect flight muscles (IFMs). MEF2 can inhibit Notch pathway in non-myogenic cells [95]. Like other molecular regulator, MEF2s can also cooperate with key proteins in other signaling to modulate its final biological effect. For example, the Notch-MEF2 synergy acts upon JNK signaling and its downstream events to induce proliferation and metastasis in Drosophila [96].
DISCUSSION
Researchers are engaged in developing new drugs that target MEF2 family in order to treat MEF2 associated human diseases. For instances, oleanolic acid is a new type of anti-tumor drugs that suppresses proliferation of lung cancer cells via inhibition of MEF2D expression [97]. Similarly, estrogen alone or its co-treatment with parathyroid hormone can increase vertebral bone mass by suppressing the expression of MEF2s in ovariectomized rats [98]. Atorvastatin is a drug used to treat angiocardiopathy, which can reverse cardiac remodeling by suppressing protein kinase D/MEF2D activation in spontaneously hypertensive rats [99].
In this review, we summarized that MEF2 family plays significant roles in nervous system diseases, heart diseases, muscular disorders, and cancers (Table 1). The four members in MEF2 family are actually different from each other in their biochemical and physiological features, and these distinctions cause their different roles in human diseases. Although many signaling have been identified as a partner of MEF2 proteins, more new molecular pathways may be related to these transcription factors. MEF2s may participate in the regulation of YAP/Hippo signaling and EGFR-related pathway. Similarly, NF-kappa B pathway may also be regulated by MEF2 proteins during inflammatory reactions. Taken together, MEF2 proteins and their associated signaling pathways play important roles in both physiological and pathological process in human (Figure 5).
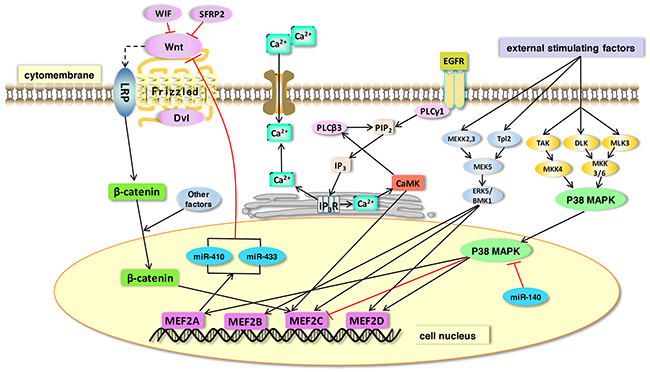
Figure 5: The function of MEF2 proteins in signaling pathways. MEF2 proteins act as effectors of many signaling pathways. In this figure, we show some representative pathways. 1) Wnt protein binds frizzled receptors to activate its downstream reactions. A stablized β-catenin is then translocated into the nucleus where it binds with MEF2 transcription factors. MEF2A can directly regulate microRNAs (miR-410 and miR-433) and thus inhibit the Wnt signaling pathway. 2) MAPK signaling (ERK and p38 MAPK) can be stimulated by external factors. The ERK signaling increases expression of its direct downstream targets, MEF2B, C and D. MEF2A and MEF2C can be activated by p38 MAPK. miR-140 is a suppressor of p38 MAPK signaling and its overexpression can decrease MEF2C expression in a p38MAPK-dependent pathway. 3) The expression of MEF2C can be enhanced by the activation of CaMK in Ca2+ signaling pathway.
Table 1: The MEF2-related human diseases, signaling pathways, targets and drugs
Diseases |
Mainly corresponding MEF2 subtype |
Signaling Pathways |
Targeted Proteins |
Drugs |
|---|---|---|---|---|
Nervous system diseases |
MEF2A |
MAP Kinase signaling pathway |
SIK2,PSD95, |
― |
Heart diseases |
MEF2C |
Ca2+signaling pathway |
Tdgf1,Suv39h1 |
Atorvastatin |
Muscle disorders |
MEF2A |
PI3K/AKT pathway |
BDNF,KLF6 |
― |
Cancer |
MEF2B |
Wnt signaling pathway |
VEGF,SFRP2 |
Oleanolic acid |
It is noteworthy that the MEF2 proteins could be potential targets during the treatment of many diseases. Although previous studies have reported various functions of MEF2 family proteins, further works are still needed to clarify the pathological and pharmacological mechanism by which MEF2 genes is connected to specific signaling pathway. The detailed elucidation of underlying mechanisms of MEF2 activation and its downstream targets could facilitate the development of efficient MEF2-targeted therapeutic strategies.
CONFLICTS OF INTEREST
There is no conflict of interest.
GRANT SUPPORT
This work was supported by funds from National Natural Sciences Foundation of China (81502065), China Postdoctoral Science Foundation Funded Project (2016T90613 and 2015M580574), Natural Sciences Foundation of Shandong Province (ZR2014HQ009), Postdoctoral Innovation Project of Shandong Province (201602037), Qingdao Postdoctoral Research Project (2015167) and Qingdao Innovation Project (16-5-1-56-JCH).
REFERENCES
1. Nakayama H, Fujio Y, Yamaguchi O. [Calcium dependent signaling in cardiac hypertrophy and cell death]. [Article in Japanese]. Clin Calcium. 2013; 23:505–15.
2. Tamir Y, Bengal E. Phosphoinositide 3-kinase induces the transcriptional activity of MEF2 proteins during muscle differentiation. J Biol Chem. 2000; 275:34424–32. https://doi.org/10.1074/jbc.M005815200.
3. Keren A, Tamir Y, Bengal E. The p38 MAPK signaling pathway: a major regulator of skeletal muscle development. Mol Cell Endocrinol. 2006; 252:224–30. https://doi.org/10.1016/j.mce.2006.03.017.
4. McKinsey TA, Zhang CL, Olson EN. MEF2: a calcium-dependent regulator of cell division, differentiation and death. Trends Biochem Sci. 2002; 27:40–7.
5. Shore P, Sharrocks AD. The MADS-box family of transcription factors. Eur J Biochem. 1995; 229:1–13.
6. Ma Q, Telese F. Genome-wide epigenetic analysis of MEF2A and MEF2C transcription factors in mouse cortical neurons. Commun Integr Biol. 2015; 8:e1087624. https://doi.org/10.1080/19420889.2015.1087624.
7. Brand NJ. Myocyte enhancer factor 2 (MEF2). Int J Biochem Cell Biol. 1997; 29:1467–70.
8. Black BL, Olson EN. Transcriptional control of muscle development by myocyte enhancer factor-2 (MEF2) proteins. Annu Rev Cell Dev Biol. 1998; 14:167–96. https://doi.org/10.1146/annurev.cellbio.14.1.167.
9. Hobson GM, Krahe R, Garcia E, Siciliano MJ, Funanage VL. Regional chromosomal assignments for four members of the mads domain transcription enhancer factor 2 (MEF2) gene family to human chromosomes 15q26, 19p12, 5q14, and 1q12-q23. Genomics. 1995; 29:704–11. https://doi.org/10.1006/geno.1995.9007.
10. Molkentin JD, Black BL, Martin JF, Olson EN. Mutational analysis of the DNA binding, dimerization, and transcriptional activation domains of MEF2C. Mol Cell Biol. 1996; 16:2627–36.
11. Martin JF, Miano JM, Hustad CM, Copeland NG, Jenkins NA, Olson EN. A MEF2 gene that generates a muscle-specific isoform via alternative mRNA splicing. Mol Cell Biol. 1994; 14:1647–56.
12. Flavell SW, Kim TK, Gray JM, Harmin DA, Hemberg M, Hong EJ, Markenscoff-Papadimitriou E, Bear DM, Greenberg ME. Genome-wide analysis of MEF2 transcriptional program reveals synaptic target genes and neuronal activity-dependent polyadenylation site selection. Neuron. 2008; 60:1022–38. https://doi.org/10.1016/j.neuron.2008.11.029.
13. Creemers EE, Sutherland LB, McAnally J, Richardson JA, Olson EN. Myocardin is a direct transcriptional target of MEF2, tead and Foxo proteins during cardiovascular development. Development. 2006; 133:4245–56. https://doi.org/10.1242/dev.02610.
14. Huang HT, Brand OM, Mathew M, Ignatiou C, Ewen EP, McCalmon SA, Naya FJ. Myomaxin is a novel transcriptional target of MEF2A that encodes a xin-related alpha-actinin-interacting protein. J Biol Chem. 2006; 281:39370–9. https://doi.org/10.1074/jbc.M603244200.
15. Sandmann T, Jensen LJ, Jakobsen JS, Karzynski MM, Eichenlaub MP, Bork P, Furlong EE. A temporal map of transcription factor activity: MEF2 directly regulates target genes at all stages of muscle development. Dev Cell. 2006; 10:797–807. https://doi.org/10.1016/j.devcel.2006.04.009.
16. Ataman B, Boulting GL, Harmin DA, Yang MG, Baker-Salisbury M, Yap EL, Malik AN, Mei K, Rubin AA, Spiegel I, Durresi E, Sharma N, Hu LS, et al. Evolution of osteocrin as an activity-regulated factor in the primate brain. Nature. 2016; 539:242–7. https://doi.org/10.1038/nature20111.
17. McGee SL, Sparling D, Olson AL, Hargreaves M. Exercise increases MEF2- and GEF DNA-binding activity in human skeletal muscle. FASEB J. 2006; 20:348–9. https://doi.org/10.1096/fj.05-4671fje.
18. Shalizi AK, Bonni A. Brawn for brains: the role of MEF2 proteins in the developing nervous system. Curr Top Dev Biol. 2005; 69:239–66. https://doi.org/10.1016/s0070-2153(05)69009-6.
19. Phan D, Rasmussen TL, Nakagawa O, McAnally J, Gottlieb PD, Tucker PW, Richardson JA, Bassel-Duby R, Olson EN. BOP, a regulator of right ventricular heart development, is a direct transcriptional target of MEF2C in the developing heart. Development. 2005; 132:2669–78. https://doi.org/10.1242/dev.01849.
20. Wang X, Tang X, Gong X, Albanis E, Friedman SL, Mao Z. Regulation of hepatic stellate cell activation and growth by transcription factor myocyte enhancer factor 2. Gastroenterology. 2004; 127:1174–88.
21. Pon JR, Marra MA. MEF2 transcription factors: Developmental regulators and emerging cancer genes. Oncotarget. 2016; 7:2297–312. https://doi.org/10.18632/oncotarget.6223.
22. Verdaguer E, Alvira D, Jimenez A, Rimbau V, Camins A, Pallas M. Inhibition of the cdk5/MEF2 pathway is involved in the antiapoptotic properties of calpain inhibitors in cerebellar neurons. Br J Pharmacol. 2005; 145:1103–11. https://doi.org/10.1038/sj.bjp.0706280.
23. Atkins M, Potier D, Romanelli L, Jacobs J, Mach J, Hamaratoglu F, Aerts S, Halder G. An ectopic network of transcription factors regulated by hippo signaling drives growth and invasion of a malignant tumor model. Curr Biol. 2016; 26:2101–13. https://doi.org/10.1016/j.cub.2016.06.035.
24. Al-Khalili L, Kramer D, Wretenberg P, Krook A. Human skeletal muscle cell differentiation is associated with changes in myogenic markers and enhanced insulin-mediated MAPK and PKB phosphorylation. Acta Physiol Scand. 2004; 180:395–403. https://doi.org/10.1111/j.1365-201X.2004.01259.x.
25. Ma K, Chan JK, Zhu G, Wu Z. Myocyte enhancer factor 2 acetylation by p300 enhances its DNA binding activity, transcriptional activity, and myogenic differentiation. Mol Cell Biol. 2005; 25:3575–82. https://doi.org/10.1128/mcb.25.9.3575-3582.2005.
26. Mughal W, Nguyen L, Pustylnik S, da Silva Rosa SC, Piotrowski S, Chapman D, Du M, Alli NS, Grigull J, Halayko AJ, Aliani M, Topham MK, Epand RM, et al. A conserved MADS-box phosphorylation motif regulates differentiation and mitochondrial function in skeletal, cardiac, and smooth muscle cells. Cell Death Dis. 2015; 6:e1944. https://doi.org/10.1038/cddis.2015.306.
27. Moretti I, Ciciliot S. MRF4 negatively regulates adult skeletal muscle growth by repressing MEF2 activity. 2016; 7:12397. https://doi.org/10.1038/ncomms12397.
28. Arosio A, Sala G, Rodriguez-Menendez V, Grana D, Gerardi F, Lunetta C, Ferrarese C, Tremolizzo L. MEF2D and MEF2C pathways disruption in sporadic and familial als patients. Mol Cell Neurosci. 2016; 74:10–7. https://doi.org/10.1016/j.mcn.2016.02.002.
29. Infantino V, Convertini P, Menga A, Iacobazzi V. MEF2C exon alpha: Role in gene activation and differentiation. Gene. 2013; 531:355–62. https://doi.org/10.1016/j.gene.2013.08.044.
30. Leifer D, Krainc D, Yu YT, McDermott J, Breitbart RE, Heng J, Neve RL, Kosofsky B, Nadal-Ginard B, Lipton SA. MEF2C, a MADS/MEF2-family transcription factor expressed in a laminar distribution in cerebral cortex. Proc Natl Acad Sci U S A. 1993; 90:1546–50.
31. Li M, Linseman DA, Allen MP, Meintzer MK, Wang X, Laessig T, Wierman ME, Heidenreich KA. Myocyte enhancer factor 2A and 2D undergo phosphorylation and caspase-mediated degradation during apoptosis of rat cerebellar granule neurons. J Neurosci. 2001; 21:6544–52.
32. Latchney SE, Jiang Y, Petrik DP, Eisch AJ, Hsieh J. Inducible knockout of MEF2a, -c, and -d from nestin-expressing stem/progenitor cells and their progeny unexpectedly uncouples neurogenesis and dendritogenesis in vivo. FASEB J. 2015; 29:5059–71. https://doi.org/10.1096/fj.15-275651.
33. Krenacs D, Borbenyi Z, Bedekovics J, Mehes G, Bagdi E, Krenacs L. Pattern of MEF2B expression in lymphoid tissues and in malignant lymphomas. Virchows Arch. 2015; 467:345–55. https://doi.org/10.1007/s00428-015-1796-6.
34. Tempera I, De Leo A, Kossenkov AV, Cesaroni M, Song H, Dawany N, Showe L, Lu F, Wikramasinghe P, Lieberman PM. Identification of MEF2B, EBF1, and Il6r as direct gene targets of epstein-barr virus (EBV) nuclear antigen 1 critical for EBV-infected b-lymphocyte survival. J Virol. 2015; 90:345–55. https://doi.org/10.1128/jvi.02318-15.
35. Cardoso AC, Pereira AH, Ambrosio AL, Consonni SR, Rocha de Oliveira R, Bajgelman MC, Dias SM, Franchini KG. FAK forms a complex with MEF2 to Couple Biomechanical Signaling to Transcription in Cardiomyocytes. Structure. 2016; 24:1301–10. https://doi.org/10.1016/j.str.2016.06.003.
36. Barnes RM, Harris IS, Jaehnig EJ, Sauls K, Sinha T, Rojas A, Schachterle W, McCulley DJ, Norris RA, Black BL. MEF2C regulates outflow tract alignment and transcriptional control of Tdgf1. Development. 2016; 143:774–9. https://doi.org/10.1242/dev.126383.
37. Jin W, Shang Y, Peng J, Jiang S. Histone h3 methyltransferase Suv39h1 prevents myogenic terminal differentiation by repressing MEF2 activity in muscle cells. Int J Mol Sci. 2016. https://doi.org/10.3390/ijms17121908.
38. Suryo Rahmanto A, Savov V, Brunner A, Bolin S, Weishaupt H, Malyukova A, Rosen G, Cancer M, Hutter S, Sundstrom A, Kawauchi D, Jones DT, Spruck C, et al. FBW7 suppression leads to SOX9 stabilization and increased malignancy in medulloblastoma. EMBO J. 2016; 35:2192–212. https://doi.org/10.15252/embj.201693889.
39. Ma L, Liu J, Liu L, Duan G, Wang Q, Xu Y, Xia F, Shan J, Shen J, Yang Z, Bie P, Cui Y, Bian XW, et al. Overexpression of the transcription factor MEF2D in hepatocellular carcinoma sustains malignant character by suppressing G2-m transition genes. Cancer Res. 2014; 74:1452–62. https://doi.org/10.1158/0008-5472.can-13-2171.
40. Hong X, Hong XY, Li T, He CY. Pokemon and MEF2D co-operationally promote invasion of hepatocellular carcinoma. Tumour Biol. 2015; 36:9885–93. https://doi.org/10.1007/s13277-015-3744-0.
41. Yu W, Huang C, Wang Q, Huang T, Ding Y, Ma C, Ma H, Chen W. MEF2 transcription factors promotes emt and invasiveness of hepatocellular carcinoma through TGF-beta1 autoregulation circuitry. Tumour Biol. 2014; 35:10943–51. https://doi.org/10.1007/s13277-014-2403-1.
42. Kawashima-Goto S, Imamura T, Tomoyasu C, Yano M, Yoshida H, Fujiki A, Tamura S, Osone S, Ishida H, Morimoto A, Kuroda H, Hosoi H. BCL2 inhibitor (ABT-737): a restorer of prednisolone sensitivity in early t-cell precursor-acute lymphoblastic leukemia with high MEF2C expression? PLoS One. 2015; 10:e0132926. https://doi.org/10.1371/journal.pone.0132926.
43. Homminga I, Pieters R, Langerak AW, de Rooi JJ, Stubbs A, Verstegen M, Vuerhard M, Buijs-Gladdines J, Kooi C, Klous P, van Vlierberghe P, Ferrando AA, Cayuela JM, et al. Integrated transcript and genome analyses reveal NKX2-1 and MEF2C as potential oncogenes in T cell acute lymphoblastic leukemia. Cancer Cell. 2011; 19:484–97. https://doi.org/10.1016/j.ccr.2011.02.008.
44. Prima V, Hunger SP. Cooperative transformation by MEF2D/DAZAP1 and DAZAP1/MEF2D fusion proteins generated by the variant t(1;19) in acute lymphoblastic leukemia. Leukemia. 2007; 21:2470–5. https://doi.org/10.1038/sj.leu.2404962.
45. Di Giorgio E, Clocchiatti A, Piccinin S, Sgorbissa A, Viviani G, Peruzzo P, Romeo S, Rossi S, Dei Tos AP, Maestro R, Brancolini C. MEF2 is a converging hub for histone deacetylase 4 and phosphatidylinositol 3-kinase/Akt-induced transformation. Mol Cell Biol. 2013; 33:4473–91. https://doi.org/10.1128/mcb.01050-13.
46. Friday BB, Mitchell PO, Kegley KM, Pavlath GK. Calcineurin initiates skeletal muscle differentiation by activating MEF2 and MyoD. Differentiation. 2003; 71:217–27. https://doi.org/10.1046/j.1432-0436.2003.710303.x.
47. Boubali S, Liopeta K, Virgilio L, Thyphronitis G, Mavrothalassitis G, Dimitracopoulos G, Paliogianni F. Calcium/calmodulin-dependent protein kinase ii regulates il-10 production by human t lymphocytes: a distinct target in the calcium dependent pathway. Mol Immunol. 2012; 52:51–60. https://doi.org/10.1016/j.molimm.2012.04.008.
48. Li J, Vargas MA, Kapiloff MS, Dodge-Kafka KL. Regulation of MEF2 transcriptional activity by calcineurin/mAKAP complexes. Exp Cell Res. 2013; 319:447–54. https://doi.org/10.1016/j.yexcr.2012.12.016.
49. Dai X, Liu H, Shen S, Guo X, Yan H, Ji X, Li L, Huang J, Feng XH, Zhao B. YAP activates the hippo pathway in a negative feedback loop. Cell Res. 2015; 25:1175–8. https://doi.org/10.1038/cr.2015.101.
50. Lee CW, Yang FC, Chang HY, Chou H, Tan BC, Lee SC. Interaction between salt-inducible kinase 2 and protein phosphatase 2A regulates the activity of calcium/calmodulin-dependent protein kinase i and protein phosphatase methylesterase-1. J Biol Chem. 2014; 289:21108–19. https://doi.org/10.1074/jbc.M113.540229.
51. Elmer BM, Estes ML, Barrow SL, McAllister AK. MHCI requires MEF2 transcription factors to negatively regulate synapse density during development and in disease. J Neurosci. 2013; 33:13791–804. https://doi.org/10.1523/jneurosci.2366-13.2013.
52. Tsai NP, Wilkerson JR, Guo W, Huber KM. FMRP-dependent Mdm2 dephosphorylation is required for MEF2-induced synapse elimination. Hum Mol Genet. 2017; 26:293–304. https://doi.org/10.1093/hmg/ddw386.
53. Daems C, Di-Luoffo M, Paradis E, Tremblay JJ. MEF2 cooperates with forskolin/cAMP and GATA4 to regulate star gene expression in mouse MA-10 leydig cells. Endocrinology. 2015; 156:2693–703. https://doi.org/10.1210/en.2014-1964.
54. Di-Luoffo M, Brousseau C, Bergeron F, Tremblay JJ. The transcription factor MEF2 is a novel regulator of gsta gene class in mouse MA-10 leydig cells. Endocrinology. 2015; 156:4695–706. https://doi.org/10.1210/en.2015-1500.
55. Guo JR, Li W, Wu Y, Wu LQ, Li X, Guo YF, Zheng XH, Lian XL, Huang HF, Chen YZ. Hepatocyte growth factor promotes proliferation, invasion, and metastasis of myeloid leukemia cells through PI3K-AKT and MAPK/ERK signaling pathway. Am J Transl Res. 2016; 8:3630–44.
56. Yang HJ, Ma SP, Ju F, Zhang YP, Li ZC, Zhang BB, Lian JJ, Wang L, Cheng BF, Wang M, Feng ZW. Thrombospondin-4 promotes neuronal differentiation of NG2 cells via the ERK/MAPK pathway. J Mol Neurosci. 2016. https://doi.org/10.1007/s12031-016-0845-1.
57. Seger R, Krebs EG. The MAPK signaling cascade. FASEB J. 1995; 9:726–35.
58. Wang X, Pesakhov S, Harrison JS, Danilenko M, Studzinski GP. ERK5 pathway regulates transcription factors important for monocytic differentiation of human myeloid leukemia cells. J Cell Physiol. 2014; 229:856–67. https://doi.org/10.1002/jcp.24513.
59. Pazyra-Murphy MF, Hans A, Courchesne SL, Karch C, Cosker KE, Heerssen HM, Watson FL, Kim T, Greenberg ME, Segal RA. A retrograde neuronal survival response: Target-derived neurotrophins regulate MEF2D and bcl-w. J Neurosci. 2009; 29:6700–9. https://doi.org/10.1523/jneurosci.0233-09.2009.
60. Herglotz J, Unrau L, Hauschildt F, Fischer M, Kriebitzsch N, Alawi M, Indenbirken D, Spohn M, Muller U, Ziegler M, Schuh W, Jack HM, Stocking C. Essential control of early B-cell development by MEF2 transcription factors. Blood. 2016; 127:572–81. https://doi.org/10.1182/blood-2015-04-643270.
61. Wang RM, Zhang QG, Li J, Yang LC, Yang F, Brann DW. The ERK5-MEF2C transcription factor pathway contributes to anti-apoptotic effect of cerebral ischemia preconditioning in the hippocampal CA1 region of rats. Brain Res. 2009; 1255:32–41. https://doi.org/10.1016/j.brainres.2008.12.011.
62. Im JY, Yoon SH, Kim BK, Ban HS, Won KJ, Chung KS, Jung KE, Won M. DNA damage induced apoptosis suppressor (ddias) is upregulated via ERK5/MEF2B signaling and promotes beta-catenin-mediated invasion. Biochim Biophys Acta. 2016. https://doi.org/10.1016/j.bbagrm.2016.07.003.
63. Gupta J, Nebreda AR. Roles of p38alpha mitogen-activated protein kinase in mouse models of inflammatory diseases and cancer. Febs j. 2015; 282:1841–57. https://doi.org/10.1111/febs.13250.
64. Han J, Jiang Y, Li Z, Kravchenko VV, Ulevitch RJ. Activation of the transcription factor MEF2C by the MAP kinase p38 in inflammation. Nature. 1997; 386:296–9. https://doi.org/10.1038/386296a0.
65. Ehyai S, Dionyssiou MG, Gordon JW, Williams D, Siu KW, McDermott JC. A p38 mitogen-activated protein kinase-regulated myocyte enhancer factor 2-beta-catenin interaction enhances canonical wnt signaling. Mol Cell Biol. 2015; 36:330–46. https://doi.org/10.1128/mcb.00832-15.
66. Wales S, Hashemi S, Blais A, McDermott JC. Global MEF2 target gene analysis in cardiac and skeletal muscle reveals novel regulation of dusp6 by p38mapk-MEF2 signaling. Nucleic Acids Res. 2014; 42:11349–62. https://doi.org/10.1093/nar/gku813.
67. Kong NR, Davis M, Chai L, Winoto A, Tjian R. MEF2C and EBF1 co-regulate B cell-specific transcription. PLoS Genet. 2016; 12:e1005845. https://doi.org/10.1371/journal.pgen.1005845.
68. Shen S, Huang D, Feng G, Zhu L, Zhang Y, Cao P, Zheng K, Zhang D, Feng X. MEF2 transcription factor regulates osteogenic differentiation of dental pulp stem cells. Cell Reprogram. 2016; 18:237–45. https://doi.org/10.1089/cell.2016.0016.
69. Dadson K, Turdi S, Hashemi S, Zhao J, Polidovitch N, Beca S, Backx PH, McDermott JC, Sweeney G. Adiponectin is required for cardiac MEF2 activation during pressure overload induced hypertrophy. J Mol Cell Cardiol. 2015; 86:102–9. https://doi.org/10.1016/j.yjmcc.2015.06.020.
70. Papaioannou G, Mirzamohammadi F, Lisse TS, Nishimori S, Wein MN, Kobayashi T. MicroRNA-140 provides robustness to the regulation of hypertrophic chondrocyte differentiation by the PTHrP-HDAC4 pathway. J Bone Miner Res. 2015; 30:1044–52. https://doi.org/10.1002/jbmr.2438.
71. Dionyssiou MG, Nowacki NB, Hashemi S, Zhao J, Kerr A, Tsushima RG, McDermott JC. Cross-talk between glycogen synthase kinase 3beta (GSK3beta) and p38mapk regulates myocyte enhancer factor 2 (MEF2) activity in skeletal and cardiac muscle. J Mol Cell Cardiol. 2013; 54:35–44. https://doi.org/10.1016/j.yjmcc.2012.10.013.
72. Yi H, Wang Z, Li X, Yin M, Wang L, Aldalbahi A, El-Sayed NN, Wang H, Chen N, Fan C, Song H. Silica nanoparticles target a wnt signal transducer for degradation and impair embryonic development in zebrafish. Theranostics. 2016; 6:1810–20. https://doi.org/10.7150/thno.16127.
73. Nayak L, Bhattacharyya NP, De RK. Wnt signal transduction pathways: modules, development and evolution. BMC Syst Biol. 2016; 10 Suppl 2:44. https://doi.org/10.1186/s12918-016-0299-7.
74. Ota C, Baarsma HA, Wagner DE, Hilgendorff A, Konigshoff M. Linking bronchopulmonary dysplasia to adult chronic lung diseases: role of wnt signaling. Mol Cell Pediatr. 2016; 3:34. https://doi.org/10.1186/s40348-016-0062-6.
75. Sedgwick AE, D’Souza-Schorey C. Wnt signaling in cell motility and invasion: drawing parallels between development and cancer. Cancers (Basel). 2016; 8. https://doi.org/10.3390/cancers8090080.
76. Bai XL, Zhang Q, Ye LY, Liang F, Sun X, Chen Y, Hu QD, Fu QH, Su W, Chen Z, Zhuang ZP, Liang TB. Myocyte enhancer factor 2C regulation of hepatocellular carcinoma via vascular endothelial growth factor and wnt/beta-catenin signaling. Oncogene. 2015; 34:4089–97. https://doi.org/10.1038/onc.2014.337.
77. Snyder CM, Rice AL, Estrella NL, Held A, Kandarian SC, Naya FJ. MEF2A regulates the Gtl2-Dio3 microRNA mega-cluster to modulate WNT signaling in skeletal muscle regeneration. Development. 2013; 140:31–42. https://doi.org/10.1242/dev.081851.
78. Cao Q, Dong P, Wang Y, Zhang J, Shi X, Wang Y. miR-218 suppresses cardiac myxoma proliferation by targeting myocyte enhancer factor 2D. Oncol Rep. 2015; 33:2606–12. https://doi.org/10.3892/or.2015.3861.
79. Zhao YY, Zhao LN, Wang P, Miao YS, Liu YH, Wang ZH, Ma J, Li Z, Li ZQ, Xue YX. Overexpression of miR-18a negatively regulates myocyte enhancer factor 2D to increase the permeability of the blood-tumor barrier via kruppel-like factor 4-mediated downregulation of zonula occluden-1, claudin-5, and occludin. J Neurosci Res. 2015; 93:1891–902. https://doi.org/10.1002/jnr.23628.
80. Zhang R, Zhang Y, Li H. miR-1244/Myocyte enhancer factor 2D regulatory loop contributes to the growth of lung carcinoma. DNA Cell Biol. 2015; 34:692–700. https://doi.org/10.1089/dna.2015.2915.
81. Su L, Luo Y, Yang Z, Yang J, Yao C, Cheng F, Shan J, Chen J, Li F, Liu L, Liu C, Xu Y, Jiang L, et al. MEF2D transduces microenvironment stimuli to ZEB1 to promote epithelial-mesenchymal transition and metastasis in colorectal cancer. Cancer Res. 2016; 76:5054–67. https://doi.org/10.1158/0008-5472.can-16-0246.
82. Li M, Liu Z, Zhang Z, Liu G, Sun S, Sun C. miR-103 promotes 3T3-L1 cell adipogenesis through AKT/mTOR signal pathway with its target being Mef2d. Biol Chem. 2015; 396:235–44. https://doi.org/10.1515/hsz-2014-0241.
83. Kesherwani V, Nandi SS, Sharawat SK, Shahshahan HR, Mishra PK. Hydrogen sulfide mitigates homocysteine-mediated pathological remodeling by inducing miR-133a in cardiomyocytes. Mol Cell Biochem. 2015; 404:241–50. https://doi.org/10.1007/s11010-015-2383-5.
84. Zhao W, Zheng XL, Peng DQ, Zhao SP. Myocyte enhancer factor 2A regulates hydrogen peroxide-induced senescence of vascular smooth muscle cells via microRNA-143. J Cell Physiol. 2015; 230:2202–11. https://doi.org/10.1002/jcp.24948.
85. Kalsotra A, Singh RK, Gurha P, Ward AJ, Creighton CJ, Cooper TA. The MEF2 transcription network is disrupted in myotonic dystrophy heart tissue, dramatically altering miRNA and mRNA expression. Cell Rep. 2014; 6:336–45. https://doi.org/10.1016/j.celrep.2013.12.025.
86. Dey BK, Mueller AC, Dutta A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription. 2014; 5:e944014. https://doi.org/10.4161/21541272.2014.944014.
87. Song G, Shen Y, Ruan Z, Li X, Chen Y, Yuan W, Ding X, Zhu L, Qian L. LncRNA-uc.167 influences cell proliferation, apoptosis and differentiation of p19 cells by regulating Mef2c. Gene. 2016; 590:97–108. https://doi.org/10.1016/j.gene.2016.06.006.
88. Chen H, Zhou L, Wu X, Li R, Wen J, Sha J, Wen X. The PI3K/AKT pathway in the pathogenesis of prostate cancer. Front Biosci (Landmark Ed). 2016; 21:1084–91.
89. Guo H, German P, Bai S, Barnes S, Guo W, Qi X, Lou H, Liang J, Jonasch E, Mills GB, Ding Z. The PI3K/AKT pathway and renal cell carcinoma. J Genet Genomics. 2015; 42:343–53. https://doi.org/10.1016/j.jgg.2015.03.003.
90. Xu Q, Wu Z. The insulin-like growth factor-phosphatidylinositol 3-kinase-Akt signaling pathway regulates myogenin expression in normal myogenic cells but not in rhabdomyosarcoma-derived rd cells. J Biol Chem. 2000; 275:36750–7. https://doi.org/10.1074/jbc.M005030200.
91. Estrella NL, Clark AL, Desjardins CA, Nocco SE, Naya FJ. MEF2D deficiency in neonatal cardiomyocytes triggers cell cycle re-entry and programmed cell death in vitro. J Biol Chem. 2015; 290:24367–80. https://doi.org/10.1074/jbc.M115.666461.
92. Wang X, Tang X, Li M, Marshall J, Mao Z. Regulation of neuroprotective activity of myocyte-enhancer factor 2 by cAMP-protein kinase a signaling pathway in neuronal survival. J Biol Chem. 2005; 280:16705–13. https://doi.org/10.1074/jbc.M501819200.
93. Du M, Perry RL, Nowacki NB, Gordon JW, Salma J, Zhao J, Aziz A, Chan J, Siu KW, McDermott JC. Protein kinase a represses skeletal myogenesis by targeting myocyte enhancer factor 2D. Mol Cell Biol. 2008; 28:2952–70. https://doi.org/10.1128/mcb.00248-08.
94. Chen K, Ho TS, Lin G, Tan KL, Rasband MN, Bellen HJ. Loss of frataxin activates the iron/sphingolipid/pdk1/mef2 pathway in mammals. Elife. 2016; 5:e20732. https://doi.org/10.7554/eLife.20732.001.
95. Caine C, Kasherov P, Silber J, Lalouette A. MEF2 interacts with the notch pathway during adult muscle development in drosophila melanogaster. PLoS One. 2014; 9:e108149. https://doi.org/10.1371/journal.pone.0108149.
96. Pallavi SK, Ho DM, Hicks C, Miele L, Artavanis-Tsakonas S. Notch and MEF2 synergize to promote proliferation and metastasis through JNK signal activation in drosophila. Embo j. 2012; 31:2895–907. https://doi.org/10.1038/emboj.2012.129.
97. Zhao X, Liu M, Li D. Oleanolic acid suppresses the proliferation of lung carcinoma cells by miR-122/cyclin G1/MEF2D axis. Mol Cell Biochem. 2015; 400:1–7. https://doi.org/10.1007/s11010-014-2228-7.
98. Jia HB, Ma JX, Ma XL, Yu JT, Feng R, Xu LY, Wang J, Xing D, Zhu SW, Wang Y. Estrogen alone or in combination with parathyroid hormone can decrease vertebral MEF2 and sclerostin expression and increase vertebral bone mass in ovariectomized rats. Osteoporos Int. 2014; 25:2743–54. https://doi.org/10.1007/s00198-014-2818-y.
99. Geng J, Zhao Z, Kang W, Wang W, Zhang Y, Zhiming GE. Atorvastatin reverses cardiac remodeling possibly through regulation of protein kinase d/myocyte enhancer factor 2D activation in spontaneously hypertensive rats. Pharmacol Res. 2010; 61:40–7. https://doi.org/10.1016/j.phrs.2009.08.004.

