INTRODUCTION
The therapeutic options for castration-resistant prostate cancer (CRPC) have increased dramatically in the past years. Recent clinical trials have led to the FDA approval of cabazitaxel [1], abiraterone [2–4], enzalutamide [5, 6], radium-223 [7], and Sipuleucel-T [8]. Nevertheless, most CRPC patients develop resistance to these new agents, making it of utmost importance to develop newer treatment strategies [9]. The objective responses to immunotherapy in other cancers are driving renewed enthusiasm for cancer immunotherapy, however, the role of immunotherapy in CRPC is yet to be fully elucidated [10].
Programmed Death Receptor 1 (PD-1, or CD279), a member of the extended family of T cell regulators, is expressed on the surface of activated T cells, B cells, and macrophages [12]. Its ligand, Programmed Death Receptor Ligand 1 (PD-L1, or B7-H1 or CD274), is expressed on tumor cells, macrophages, T cells and certain other cell types [12], and the interaction of these two molecules negatively regulates immune responses. Of major interest is that inhibition of the interaction between PD1 and PD-L1 can enhance T-cell responses in vitro, and mediates clinical anti-tumor activity [13–20]. A correlation between PD-L1 expression on tumor or immune cells in the tumor specimen and tumor response to anti-PD1 or anti-PD-L1 immunotherapy has been described in various advanced tumors [13–21]. PD-L1 is expressed in a wide range of tumors, at a frequency of up to 88% in some types of cancer [22]. In the tumor microenvironment, PD-L1 expressed on tumor cells binds to PD-1 on activated T cells that have migrated to the tumor. This delivers an inhibitory signal to those T cells, preventing them from killing target tumor cells, and protecting the tumor from immune elimination [22]. Recently, in a small study with ten CRPC patients treated with pembrolizumab, response in three patients has been reported [23]. Tumor tissue was available for two of these three patients, and showed PD-L1 expression. In addition, primary and metastatic CRPC showed robust synergistic responses when immune checkpoint blockade was combined with myeloid-derived suppressor cells (MDSC)-targeted therapy. Mechanistically, combination therapy efficacy stemmed from the upregulation of interleukin-1 receptor antagonist and suppression of MDSC-promoting cytokines secreted by prostate cancer cells. These recent observations by Lu et al. illuminate a new treatment concept, combining immune checkpoint blockade with MDSC-targeted therapies for mCRPC [11].
The first aim of this study was to systematically describe the expression of PD-L1 in benign prostatic hyperplasia (BPH), localized prostate cancer (PC), and CRPC using two anti-PD-L1 antibodies. The second aim was to observe if PD-L1 status was dependent on previous treatment modalities. The third aim was to develop a new method to describe the relationship between immune cells and tumor-related proteins using pseudo-colored protein expression maps, and to use dot-wise correlation coefficients of superimposed images as an objective co-location measurement.
RESULTS
Expression of PD-L1 in benign and malignant prostate tissue
This study included 248 tissue samples (70 BPH, 96 PC, 82 CRPC) and 3 PC cell lines. There was no expression of PD-L1 observed in either BPH or localized PC samples (0%). Examples of PD-L1 tumor cell (TC) and immune cell (IC) staining in CRCP specimens and cell lines (clone SP263 and E1L3N) are given in Figure 1A–1F. Of the three tested paraffin embedded cell cultures (E1L3N), only PC3 and LnCaP showed membranous PD-L1 expression. A comparison of staining patterns and scoring was performed on serial sections of tissue microarrays with CRPC samples (Figure 2A–2B). Two antibody clones were used for PD-L1 immunohistochemistry assays (E1L3N, SP263). Overall, PD-L1 expression patterns were similar in both assays, and CRPC samples displayed heterogeneous PD-L1 expression. As illustrated in Figure 2B, clone SP263 showed the strongest membranous staining in tumor cells. However, with clone SP263, only three of 82 analyzable cases showed membranous immunoreactivity in more than 1% of tumor cells (3.7%), whereas clone E1L3N revealed 5 positive out of 81 analyzable CRCP samples (6.0%). Figure 3A displays the direct expression values (% positive tumor cells) by using clone E1L3N versus SP263. The comparison of PD-L1 expression (%) with clone SP263 versus E1L3N showed significant correlation coefficients (Figure 3B and 3C). No significant association of PD-L1 immunoreactivity with expression of phospho-ERK1/2 (Figure 4A and 4D), phospho-mTOR (Figure 4B and 4E), phospho-4E-BP1 (Figure 4C and 4F), and Ki-67 proliferation fraction (data not shown) could be observed, after correction for multiple testing (Figure 4A–4F).
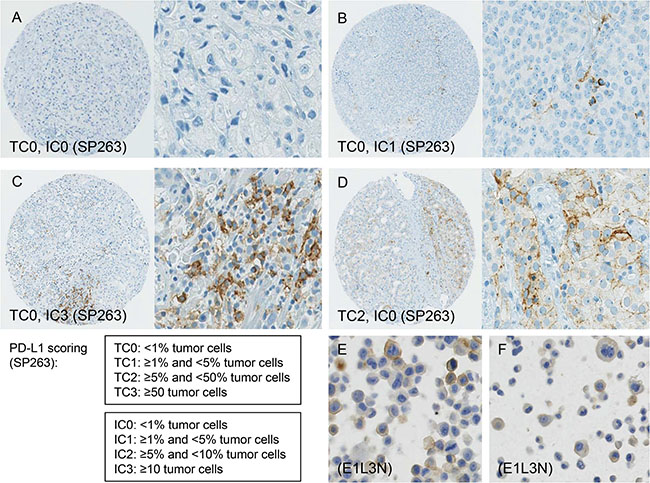
Figure 1: (A–F) Programmed death ligand 1 (PD-L1) scoring criteria and examples of tumor cell (TC) and immune cell (IC) staining in CRCP specimens and cell lines (clone SP263 and E1L3N). A: negative PD-L1 immunoreactivity in a CRCP specimen. B: PD-L1 staining in >1% of immune cells in a CRPC sample. C: PD-L1 expression in >30% of immune cells.. D: PD-L1 expression in > 5% of tumor cells. E, F: staining of PD-L1 in formalin-fixed paraffin-embedded PC3 (E) and LnCaP (F) cell culture pellets. Diameter of tissue microarray spot 0.6 mm. Inserts and cell line images, original magnification 400x. Scoring criteria of PD-L1 immunohistochemistry according to Spira et al. [43].
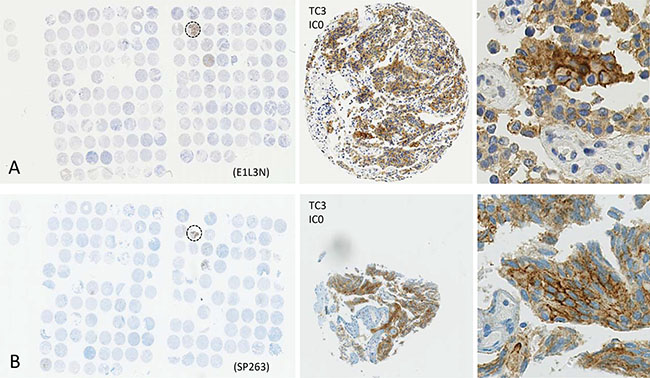
Figure 2: Comparison of PD-L1 staining results for two different monoclonal anti-PD-L1 antibodies using tissue microarrays of CRPC specimens (panel (A) E1L3N, panel (B) SP263). Single tissue spots have a diameter of 0.6 mm. Zoomed versions, showing positive membranous PD-L1 immunoreactivity in a positive CRCP patient (dashed circle).
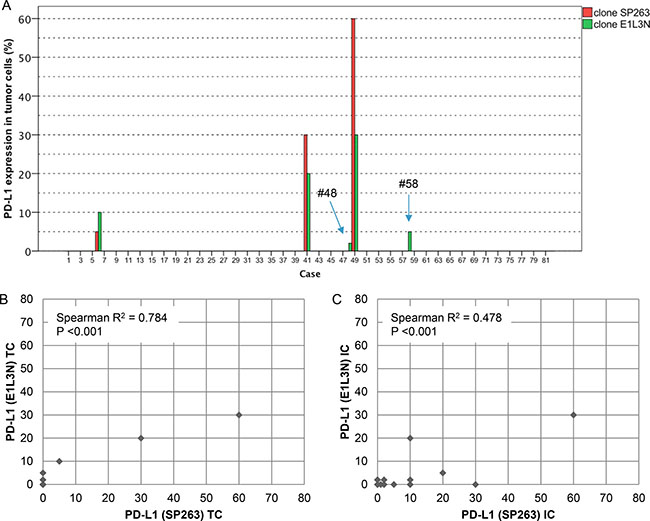
Figure 3: Direct comparison of anti-PD-L1 antibodies E1L3N and SP263. (A) Direct expression values (% positive tumor cells) for clone E1L3N (green) and SP263 (red). (B and C) Scatter plots for the comparison of immunoreactivity in % (clones E1L3N, SP263) regarding tumor cells (3B) and immune cells (3C).
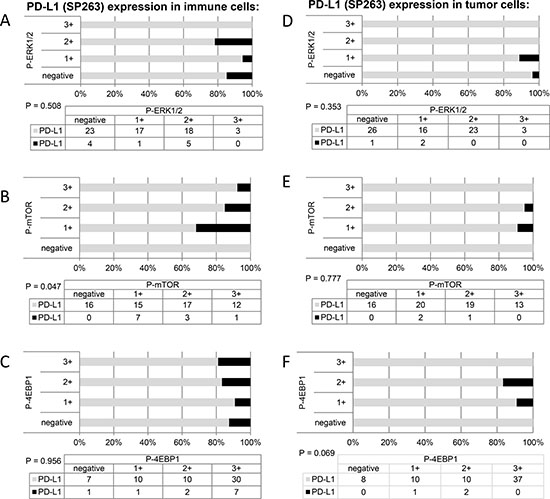
Figures 4: Cumulative bar charts showing the association of PD-L1 immunoreactivity with the expression of phospho-ERK1/2, phospho-mTOR, and phospho-4E-BP1.
PD-L1 status and previous treatment modalities
In CRPC samples, expression of PD-L1 (E1L3N) was present in 5 of 81 (6%) cases. All five positive specimens were retrieved by palliative transurethral resection of the prostate (TURP), and no PD-L1 expression was found in bone (n = 10), brain (n = 1), lung (n = 1), or lymph node (n = 1) metastases (Supplementary Table 1).
Co-localization experiments using clone E1L3N
Immunohistochemistry revealed co-localization of PD-L1 and PanCK in tumor areas, whereas stroma containing CD4 and CD8 positive lymphocytes was predominantly located surrounding the tumor tissue (Figure 5). This was confirmed by dot-wise correlation showing a moderate positive Pearson correlation coefficient for PD-L1 and PanCK expression (r = 0.51). In addition there were weak negative Pearson correlation coefficients for PanCK and CD4 (r =–0.26) and CD8 (r = –0.28), and for PD-L1 and CD4 (r =–0.05) and CD8 (r = –0.20) expression (Figure 6).
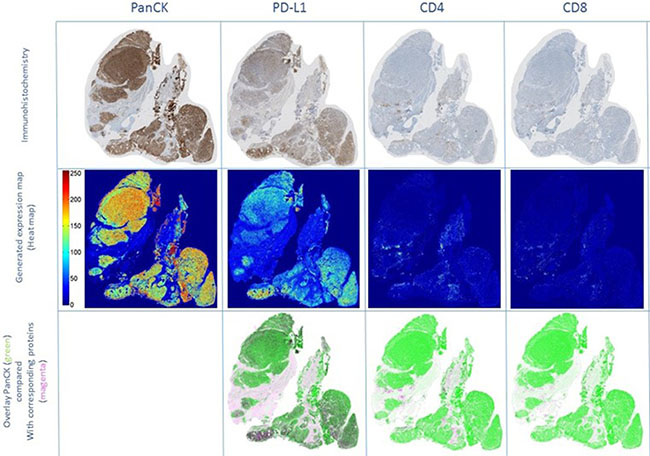
Figure 5: Co-localization experiments using clone E1L3N. Immunohistochemical staining for PanCK, PD-L1, CD4, and CD8 on serial sections of a PD-L1 positive CRPC sample (upper row). Immunohistochemical stainings were transformed into expression maps, using false colors for high (red) to low (blue) protein expression (middle row). Overlay images (lower row) were created displaying PanCK as green and PD-L1, CD4, and CD8 proteins as magenta. PD-L1 was co-localized with PanCK positive tumor cell areas but not with the surrounding stroma containing CD4- or CD8-positive lymphocytes. Abbreviations: CD4: cluster of differentiation 4; CD8: cluster of differentiation 8; CRPC: castration resistant prostate cancer; PanCK: pan cytokeratin; PD-L1: programmed cell death ligand 1.
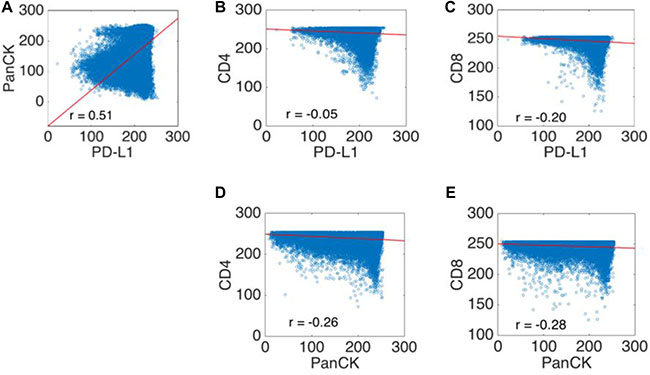
Figure 6: Scatter plots of the co-localization experiments using clone E1L3N. PD-L1 and PanCK expression showed a moderate positive Pearson’s correlation coefficient indicating overlapping expression areas (A). The comparison of PD-L1 and PanCK with CD4 and CD8 immunoreactivity showed negative weak Pearson correlation coefficients, indicating that PD-L1 and PanCK are not expressed in the same location as CD4+ or CD8+ lymphocytes (B–E). The x and y axis are represented by (normalized) intensity, ranging from 0 to 255. Abbreviations: CD4: cluster of differentiation 4; CD8: cluster of differentiation 8; PanCK: pan cytokeratin; PD-L1: programmed cell death ligand 1.
DISCUSSION
In the present study, we observed no PD-L1 expression in BPH or localized PC, and only limited expression on CRPC TCs and ICs, comparing two different PD-L1 antibodies. We did not observe any association between PD-L1 and previous treatment modalities. We demonstrated how protein expression maps and dot-wise comparison could be a useful objective measurement to describe the relationship between immune cells and tumor-related proteins.
Our findings regarding PD-L1 expression are only partially consistent and extend those from prior reports. PD-L1 expression seems to be an important biomarker as it has been associated with biochemical recurrence in PC [24], and overall survival in other cancers [25]. Furthermore, clinical trials have suggested that tissue expression of PD-L1 might be predictive for therapy response to PD-1 and PD-L1 immunotherapy [12, 14–19, 26–28]. The expression of PD-L1 observed in this study is slightly lower compared to previously published advanced PC [29] and CRPC cohorts (15% and 19% PD-L1 expression respectively) [30], and is distinct to a radical prostatectomy cohort (52.5% PD-L1 expression) [24]. These contradictory results in PD-L1 expression can be explained by different antibodies and scoring methods. Our scoring methods and compared antibodies are in line with recently accepted methodologies and guidelines for non-small cell lung cancer [31] and are currently regarded a common standard for PD-L1 testing.
Several PD-L1 immunohistochemistry assays with custom reagents and scoring systems have been developed in parallel, therefore, studies reporting PD-L1 expression and its value as a biomarker cannot be compared accurately. Whereas the heterogeneous methodologies are addressed by first harmonization studies for lung cancer [32], there is no consensus regarding preferred assay or scoring systems for PC. According to our results, we finally implemented the SP263 antibody for PD-L1 staining at our institution due to its satisfactory staining properties. To improve the described interobserver variability of the different PD-L1 scoring methods [32], digital pathology and protein maps using dot-wise comparison might be a useful tool for pathologists.
The scarce expression of PD-L1 in our CPRC cohort is consistent with the results of several clinical trials, which showed no PD-L1 expression in specimens retrieved before starting anti-PD-L1 treatment [14, 18, 33–35]. However, recently a small cohort of ten CRPC patients treated with pembrolizumab has been published, whereby three patients showed response to pembrolizumab [22]. In two of these three patients, tumor tissue was available and showed PD-L1 expression using the same antibody as in the present study (E1L3N). The authors described that both samples showed a pronounced leukocyte infiltration. Of note is the fact that the lymph node metastasis showed PD-L1 expression in cancer cells (cytokeratin positive), whereas in the liver metastases, PD-L1 expression was found in the infiltrating leukocytes. This early phase II trial [22], along with our results, stress the potential influence of the tumor microenvironment regarding treatment response. Ongoing clinical studies with immunotherapeutic agents in men with CRPC [36] are expected to report PD-L1 expression in pre-therapy specimens, and will hopefully allow for the establishment of harmonized methodologies to assess relevant features.
As PD-L1 expression in other tumor types shows variability at various time points [37], a major question remains if there is a therapeutic window for the optimal timing of immunotherapy, and if PD-L1 expression and response to immunotherapy is associated with previous treatment modalities (e.g. androgen deprivation, radio-, or chemotherapy). The observation in our study that no case after radiotherapy expressed PD-L1 is consistent with a study by Bernstein et al., who showed that treating cell lines with radiotherapy resulted in a decrease in the expression of PD-L1 [38]. Even though radiation therapy seems to decrease PD-L1 expression in this study, there might still be a role for radiotherapy. Further studies with optimal fractionation schemes may be able to induce an immune-stimulatory activity. A thorough understanding of the molecular mechanisms of immune checkpoint regulation will help to make use of this biological knowledge, and to design therapies or combinations with established agents.
These data must be interpreted in the context of the study design. First, although we have analyzed a large cohort, this is a retrospective analysis and there is a potential selection bias. Second, it is difficult to compare the results from our study with previous work due to different methodologies used to evaluate PD-L1 expression. Nevertheless, the positive staining in the included cell lines, our previous publication in germ cell tumors [39], and the accordance of our two antibodies, which was also observed in a harmonization study [32], support our protocol. Third, even though we tried to minimize tumor heterogeneity using two different cores for each patient in the tissue micro arrays (TMA), the real PD-L1 expression might be higher because of heterogeneity of PD-L1 expression within the tumor. Fourth, the therapeutic landscape has changed dramatically in the past years and our cohort was treated with only a subset of the current therapeutic options.
MATERIALS AND METHODS
Patients and tissue samples
Clinico-pathological data and follow up information on all patients was retrieved from the Department of Pathology and Molecular Pathology, University Hospital Zurich, and the clinical file of the University Hospital Zurich, Switzerland from 2000 to 2013. The study was approved by the local ethical committee (reference number KEK StV. 25-2008). All patients were treated in different hospitals with the available options: surgical ADT (orchiectomy), chemical ADT (LHRH agonists e.g. goserelin, leuprorelin, anti-androgens (bicalutamide)), chemotherapy (docetaxel) or radiotherapy. Inclusion criteria for this retrospective analysis required tissue collection during transurethral resection of the prostate (TURP), or tumor metastasis resection, and rising PSA or clinical progress under androgen deprivation therapy. The first TMA included 78 formalin-fixed paraffin embedded tissue specimens of patients, which were sampled after the diagnosis of CPRC. Suitable areas for prostate tissue retrieval were marked on routine H&E-stained sections. A section of the paraffin block, 0.6 mm in diameter, was punched out of the suitable area and inserted into the recipient block using a tissue arrayer (Beecher Instruments, Inc., Sun Prairie, WI, USA). The whole tissue microarray preparation was accomplished on the paraffin blocks using cores from two different areas of each tumor or control tissue to account for tissue heterogeneity.
The second TMA consisted of BPH samples (n = 70), localized PC (n = 96), the androgen-resistant human PC cell lines (PC3 and DU145) and the androgen-sensitive PC cell line (LnCaP). The detailed characteristics of this TMA have been reported previously [40].
Immunohistochemistry
Three-micron-thick sections of TMA blocks and formalin-fixed, paraffin-embedded tissues were mounted on glass slides (Super-Frost Plus; Menzel, Braunschweig, Germany), deparaffinized, rehydrated and stained with hematoxylin–eosin using standard histological techniques. For immunohistochemical staining of both TMA and large sections, the Ventana Benchmark automated staining system and Ventana reagents (both Ventana Medical Systems, Tucson, AZ, USA) were used. After deparaffinization in xylene, slides were rehydrated in decreasing concentrations of ethanol. Endogenous peroxidase was blocked using the Ventana endogenous peroxidase blocking kit after a rinse with distilled water. For antigen retrieval, slides were heated with cell conditioning solution (CC1, Ventana) according to the manufacturer’s instructions.
For the detection of the PD-L1 protein in BPH, PC, and cell line specimens, one anti-human PD-L1 rabbit monoclonal antibody was used: E1L3N (Cell Signaling Technology, Inc., Danvers, MA, USA). For the detection of the PD-L1 protein in CRPC, two anti-human PD-L1 rabbit monoclonal antibodies were used: E1L3N (Cell Signaling Technology, Inc., Danvers, MA, USA), SP263 (Ventana Medical Systems, Tucson, AZ, USA). A multi-tumor TMA and a tonsillectomy specimen were used as positive and negative controls to establish the staining protocol for the PD-L1 antibodies. A dilution of 1:1000 resulted in a strong membranous signal without non-specific background staining in positive controls (squamous epithelium of tonsillectomy specimen). PD-L1 negative lung cancer cases were used as negative controls. For the expression maps, primary antibodies against CD4 (clone SP35 pre-diluted, Ventana Medical Systems, Tucson, AZ, USA), CD8 (clone C8/114B, 1:100, Dako, CA, USA), and PanCK (clone AE1/AE3, 1:50, Dako) were applied and adjusted to the Ventana Benchmark system after performing titrations. Immunohistochemistry with antibodies against Phospho-ERK1/2, Phospho-mTOR, Phospho-4E-BP1, and Ki-67 (MIB1) was performed, as previously described [41].
An experienced pathologist (PJW) evaluated all scanned and digitized tissue microarray spots (Spot browser, Alphelys, Plaisir, France). A second independent observer (CDF) re-evaluated the TMAs. In discrepant cases, the two observers achieved consensus after detailed case discussion. Percentages of PD-L1 positive tumor cells and the staining pattern were evaluated for each punch. PD-L1 expression was recorded if a distinct membranous staining signal on the tumor cell surface was observed. In analogy to the scoring guidelines for non-small cell lung cancer [31] , we used a > 1% cutoff for membranous PD-L1 positivity in tumor cells.
Generation of expression maps
The staining images for CD4, CD8, PanCK, and PD-L1 were aligned, superimposed, and normalized for the generation of a pseudo-colored expression map. Whole slide images of one case, and of all performed stainings were digitized (digital slide scanner NanoZoomer 2.0-HT C9600, Hamamatsu Photonics, Solothurn, Switzerland) at 40x magnification (approximately 2k-3k x 2k-3k pixels per image). The images of consecutive tissue slices were pre-processed and aligned for dot-wise correspondence in four steps. First, the original images were white-balanced to remove grayish background. Second, the tissue regions were cropped and rotated, translated and rescaled for rigid alignment of the consecutive slices to the middle slice. The background was then removed to exclude background artifacts that could disturb the following finer alignment process. Third, to refine the alignment and to address for morphological changes of the tissues in consecutive slices, we incorporated a non-linear SIFT flow algorithm [42]. For SIFT flow, the images were scaled down to half edge size, and the standard parameters were used except for a cell size of 5 pixels and a top window size of 20 pixels to obtain optimal alignment results. Finally, the fully dot-wise aligned and downscaled images were then gray-scaled, inverted, and independently normalized for intensities between 0 and 255. The intensity maps were then compared for each case using a dot-wise Pearson correlation between the pixels of the images. All processes have been implemented in MATLAB R2014a. For visualization purposes only, the gray-scaled images have been pseudo-colored with MATLAB’s jet color map function, assigning blue values for low intensities and red values for high intensities.
CONCLUSIONS
In summary, the scarce PD-L1 expression in CRPC and absence in localized PC indicates that advanced PC patients may have to be selected carefully when offered checkpoint inhibition. However, prospective clinical trial data is needed. Our results do not confirm that PD-L1 expression can be induced by orchiectomy, LHRH-agonists, anti-androgens, radiation therapy or chemotherapy. Neither did we find an association with PD-L1 expression and ERK- or mTOR signaling activity. Protein expression maps and dot-wise comparison could be a useful objective method to describe the topographical relationship between immunological and tumor-related proteins, in general.
ACKNOWLEDGMENTS
The authors thank S. Dettwiler, F. Prutek, P. Schraml, M. Storz, N. Wey, M. Bieri and A. Wethmar for excellent technical assistance.
CONFLICTS OF INTEREST
CDF, PJS, CP, NJR, JHR, TH, TS, HM declare no conflicts of interest.
AO: Advisory boards (compensated, institutional): Bayer, Astellas, Janssen, Sanofi, Pfizer. Travel support: Bayer, Astellas, Janssen, Sanofi. Research support: Janssen, Teva
SG: advisory boards (compensated institutional): Active Biotech, Astellas, Bayer, Bristol-Myers Squibb, Curevac, Dendreon, Ferring, Janssen Cilag, Janssen Diagnostics MaxiVAX, Millennium, Novartis, Orion Pharma, Pfizer, Sanofi Aventis. Advisory Boards (uncompensated): ProteoMediX, ESSA Pharmaceuticals Corp, Nectar. Speakers Bureau (without honorarium): Amgen, Astellas, Bayer, Janssen Cilag, Sanofi Aventis. Patent: Pending patent application for a method for biomarker WO 2009138392 A1.
PJW: Participation in advisory boards or speakers bureau (compensated): Roche Diagnostics AG, Sophia Genetics SA, Myriad Genetics, Ventana Medical Systems, Life Technologies, Astellas Pharma AG, Merck AG, Sanofi-Aventis, Janssen-Cilag AG, Astra Zeneca. Research Support: Gilead, Astra Zeneca, Ventana Medical Systems.
GRANT SUPPORT
This work was supported by a SystemsX.ch (Phosphonet-PPM) and a University Hospital Zurich grant (SWF) to PJW, and by a University Hospital of Zurich and medAlumni Zurich grant to CDF.
REFERENCES
1. de Bono JS, Oudard S, Ozguroglu M, Hansen S, Machiels JP, Kocak I, Gravis G, Bodrogi I, Mackenzie MJ, Shen L, Roessner M, Gupta S, Sartor AO. Prednisone plus cabazitaxel or mitoxantrone for metastatic castration-resistant prostate cancer progressing after docetaxel treatment: a randomised open-label trial. Lancet. 2010; 376:1147–54. https://doi.org/10.1016/s0140-6736(10)61389-x.
2. Fizazi K, Scher HI, Molina A, Logothetis CJ, Chi KN, Jones RJ, Staffurth JN, North S, Vogelzang NJ, Saad F, Mainwaring P, Harland S, Goodman OB, et al. Abiraterone acetate for treatment of metastatic castration-resistant prostate cancer: final overall survival analysis of the COU-AA-301 randomised, double-blind, placebo-controlled phase 3 study. Lancet Oncol. 2012; 13:983–92. https://doi.org/10.1016/s1470-2045(12)70379-0.
3. Ryan CJ, Smith MR, de Bono JS, Molina A, Logothetis CJ, de Souza P, Fizazi K, Mainwaring P, Piulats JM, Ng S, Carles J, Mulders PFA, Basch E, et al. Abiraterone in metastatic prostate cancer without previous chemotherapy. N Engl J Med. 2013; 368:138–48. https://doi.org/10.1056/NEJMoa1209096.
4. de Bono JS, Logothetis CJ, Molina A, Fizazi K, North S, Chu L, Chi KN, Jones RJ, Goodman OB Jr, Saad F, Staffurth JN, Mainwaring P, Harland S, et al. Abiraterone and increased survival in metastatic prostate cancer. N Engl J Med. 2011; 364:1995–2005. https://doi.org/10.1056/NEJMoa1014618.
5. Beer TM, Armstrong AJ, Rathkopf DE, Loriot Y, Sternberg CN, Higano CS, Iversen P, Bhattacharya S, Carles J, Chowdhury S, Davis ID, de Bono JS, Evans CP, et al. Enzalutamide in metastatic prostate cancer before chemotherapy. N Engl J Med. 2014; 371:424–33. https://doi.org/10.1056/NEJMoa1405095.
6. Scher HI, Fizazi K, Saad F, Taplin ME, Sternberg CN, Miller K, de Wit R, Mulders P, Chi KN, Shore ND, Armstrong AJ, Flaig TW, Flechon A, et al. Increased survival with enzalutamide in prostate cancer after chemotherapy. N Engl J Med. 2012; 367:1187–97. https://doi.org/10.1056/NEJMoa1207506.
7. Parker C, Nilsson S, Heinrich D, Helle SI, O’Sullivan JM, Fosså SD, Chodacki A, Wiechno P, Logue J, Seke M, Widmark A, Johannessen DC, Hoskin P, et al. Alpha emitter radium-223 and survival in metastatic prostate cancer. N Engl J Med. 2013; 369:213–23. https://doi.org/10.1056/NEJMoa1213755.
8. Kantoff PW, Higano CS, Shore ND, Berger ER, Small EJ, Penson DF, Redfern CH, Ferrari AC, Dreicer R, Sims RB, Xu Y, Frohlich MW, Schellhammer PF, et al. Sipuleucel-T immunotherapy for castration-resistant prostate cancer. N Engl J Med. 2010; 363:411–22. https://doi.org/10.1056/NEJMoa1001294.
9. Perner S, Cronauer MV, Schrader AJ, Klocker H, Culig Z, Baniahmad A. Adaptive responses of androgen receptor signaling in castration-resistant prostate cancer. Oncotarget. 2015; 6:35542–55. https://doi.org/10.18632/oncotarget.4689.
10. Alme AK, Karir BS, Faltas BM, Drake CG. Blocking immune checkpoints in prostate, kidney, and urothelial cancer: An overview. Urol Oncol. 2016; 34:171–81. https://doi.org/10.1016/j.urolonc.2016.01.006.
11. Keir ME, Butte MJ, Freeman GJ, Sharpe AH. PD-1 and its ligands in tolerance and immunity. Annu Rev Immunol. 2008; 26:677–704. https://doi.org/10.1146/annurev.immunol.26.021607.090331.
12. Hamid O, Robert C, Daud A, Hodi FS, Hwu WJ, Kefford R, Wolchok JD, Hersey P, Joseph RW, Weber JS, Dronca R, Gangadhar TC, Patnaik A, et al. Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N Engl J Med. 2013; 369:134–44. https://doi.org/10.1056/NEJMoa1305133.
13. Robert C, Ribas A, Wolchok JD, Hodi FS, Hamid O, Kefford R, Weber JS, Joshua AM, Hwu WJ, Gangadhar TC, Patnaik A, Dronca R, Zarour H, et al. Anti-programmed-death-receptor-1 treatment with pembrolizumab in ipilimumab-refractory advanced melanoma: a randomised dose-comparison cohort of a phase 1 trial. Lancet. 2014; 384:1109–17. https://doi.org/10.1016/s0140-6736(14)60958-2.
14. Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF, Powderly JD, Carvajal RD, Sosman JA, Atkins MB, Leming PD, Spigel DR, Antonia SJ, et al. Safety, Activity, and Immune Correlates of Anti–PD-1 Antibody in Cancer. New England Journal of Medicine. 2012; 366:2443–54. https://doi.org/10.1056/NEJMoa1200690.
15. Powles T, Eder JP, Fine GD, Braiteh FS, Loriot Y, Cruz C, Bellmunt J, Burris HA, Petrylak DP, Teng SL, Shen X, Boyd Z, Hegde PS, et al. MPDL3280A (anti-PD-L1) treatment leads to clinical activity in metastatic bladder cancer. Nature. 2014; 515:558–62. https://doi.org/10.1038/nature13904.
16. Herbst RS, Soria JC, Kowanetz M, Fine GD, Hamid O, Gordon MS, Sosman JA, McDermott DF, Powderly JD, Gettinger SN, Kohrt HE, Horn L, Lawrence DP, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014; 515:563–7. https://doi.org/10.1038/nature14011.
17. Brahmer JR, Tykodi SS, Chow LQ, Hwu WJ, Topalian SL, Hwu P, Drake CG, Camacho LH, Kauh J, Odunsi K, Pitot HC, Hamid O, Bhatia S, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med. 2012; 366:2455–65. https://doi.org/10.1056/NEJMoa1200694.
18. Brahmer JR, Drake CG, Wollner I, Powderly JD, Picus J, Sharfman WH, Stankevich E, Pons A, Salay TM, McMiller TL, Gilson MM, Wang C, Selby M, et al. Phase I study of single-agent anti-programmed death-1 (MDX-1106) in refractory solid tumors: safety, clinical activity, pharmacodynamics, and immunologic correlates. J Clin Oncol. 2010; 28:3167–75. https://doi.org/10.1200/jco.2009.26.7609.
19. Berger R, Rotem-Yehudar R, Slama G, Landes S, Kneller A, Leiba M, Koren-Michowitz M, Shimoni A, Nagler A. Phase I safety and pharmacokinetic study of CT-011, a humanized antibody interacting with PD-1, in patients with advanced hematologic malignancies. Clin Cancer Res. 2008; 14:3044–51. https://doi.org/10.1158/1078-0432.ccr-07-4079.
20. Topalian SL, Sznol M, McDermott DF, Kluger HM, Carvajal RD, Sharfman WH, Brahmer JR, Lawrence DP, Atkins MB, Powderly JD, Leming PD, Lipson EJ, Puzanov I, et al. Survival, durable tumor remission, and long-term safety in patients with advanced melanoma receiving nivolumab. J Clin Oncol. 2014; 32:1020–30. https://doi.org/10.1200/jco.2013.53.0105.
21. Zou W, Chen L. Inhibitory B7-family molecules in the tumour microenvironment. Nat Rev Immunol. 2008; 8:467–77. https://doi.org/10.1038/nri2326.
22. Graff JN, Alumkal JJ, Drake CG, Thomas GV, Redmond WL, Farhad M, Cetnar JP, Ey FS, Bergan RC, Slottke R, Beer TM. Early evidence of anti-PD-1 activity in enzalutamide-resistant prostate cancer. Oncotarget. 2016; 7:52810–52817. https://doi.org/10.18632/oncotarget.10547.
23. Lu X, Horner JW, Paul E, Shang X, Troncoso P, Deng P, Jiang S, Chang Q, Spring DJ, Sharma P, Zebala JA, Maeda DY, Wang YA, et al. Effective combinatorial immunotherapy for castration-resistant prostate cancer. Nature. 2017; 543:728–32. https://doi.org/10.1038/nature21676.
24. Gevensleben H, Dietrich D, Golletz C, Steiner S, Jung M, Thiesler T, Majores M, Stein J, Uhl B, Muller S, Ellinger J, Stephan C, Jung K, et al. The Immune Checkpoint Regulator PD-L1 Is Highly Expressed in Aggressive Primary Prostate Cancer. Clin Cancer Res. 2016; 22:1969–77. https://doi.org/10.1158/1078-0432.ccr-15-2042.
25. Zhang Y, Kang S, Shen J, He J, Jiang L, Wang W, Guo Z, Peng G, Chen G, He J, Liang W. Prognostic Significance of Programmed Cell Death 1 (PD-1) or PD-1 Ligand 1 (PD-L1) Expression in Epithelial-Originated Cancer: A Meta-Analysis. Medicine (Baltimore). 2015; 94:e515. https://doi.org/10.1097/md.0000000000000515.
26. Tumeh PC, Harview CL, Yearley JH, Shintaku IP, Taylor EJM, Robert L, Chmielowski B, Spasic M, Henry G, Ciobanu V, West AN, Carmona M, Kivork C, et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. 2014; 515:568–71. https://doi.org/10.1038/nature13954.
27. Rosenberg JE, Hoffman-Censits J, Powles T, van der Heijden MS, Balar AV, Necchi A, Dawson N, O’Donnell PH, Balmanoukian A, Loriot Y, Srinivas S, Retz MM, Grivas P, et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet. 2016. https://doi.org/10.1016/s0140-6736(16)00561-4.
28. Carbognin L, Pilotto S, Milella M, Vaccaro V, Brunelli M, Caliò A, Cuppone F, Sperduti I, Giannarelli D, Chilosi M, Bronte V, Scarpa A, Bria E, et al. Differential Activity of Nivolumab, Pembrolizumab and MPDL3280A according to the Tumor Expression of Programmed Death-Ligand-1 (PD-L1): Sensitivity Analysis of Trials in Melanoma, Lung and Genitourinary Cancers. PLoS One. 2015; 10:e0130142. https://doi.org/10.1371/journal.pone.0130142.
29. Martin AM, Nirschl TR, Nirschl CJ, Francica BJ, Kochel CM, van Bokhoven A, Meeker AK, Lucia MS, Anders RA, DeMarzo AM, Drake CG. Paucity of PD-L1 expression in prostate cancer: innate and adaptive immune resistance. Prostate Cancer Prostatic Dis. 2015; 18:325–32. https://doi.org/10.1038/pcan.2015.39.
30. Massari F, Ciccarese C, Calio A, Munari E, Cima L, Porcaro AB, Novella G, Artibani W, Sava T, Eccher A, Ghimenton C, Bertoldo F, Scarpa A, et al. Magnitude of PD-1, PD-L1 and T Lymphocyte Expression on Tissue from Castration-Resistant Prostate Adenocarcinoma: An Exploratory Analysis. Target Oncol. 2016; 11:345–51. https://doi.org/10.1007/s11523-015-0396-3.
31. Garon EB, Rizvi NA, Hui R, Leighl N, Balmanoukian AS, Eder JP, Patnaik A, Aggarwal C, Gubens M, Horn L, Carcereny E, Ahn MJ, Felip E, et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med. 2015; 372:2018–28. https://doi.org/10.1056/NEJMoa1501824.
32. Scheel AH, Dietel M, Heukamp LC, Johrens K, Kirchner T, Reu S, Ruschoff J, Schildhaus HU, Schirmacher P, Tiemann M, Warth A, Weichert W, Fischer RN, et al. Harmonized PD-L1 immunohistochemistry for pulmonary squamous-cell and adenocarcinomas. Mod Pathol. 2016. https://doi.org/10.1038/modpathol.2016.117.
33. Bishop JL, Sio A, Angeles A, Roberts ME, Azad AA, Chi KN, Zoubeidi A. PD-L1 is highly expressed in Enzalutamide resistant prostate cancer. Oncotarget. 2015; 6:234–42. https://doi.org/10.18632/oncotarget.2703.
34. Gatalica Z, Snyder C, Maney T, Ghazalpour A, Holterman DA, Xiao N, Overberg P, Rose I, Basu GD, Vranic S, Lynch HT, Von Hoff DD, Hamid O. Programmed cell death 1 (PD-1) and its ligand (PD-L1) in common cancers and their correlation with molecular cancer type. Cancer Epidemiol Biomarkers Prev. 2014; 23:2965–70. https://doi.org/10.1158/1055-9965.EPI-14-0654.
35. Grenga I, Donahue R, Lepone L, Bame J, Schlom J, Farsaci B. PD-L1 and MHC-I expression in 19 human tumor cell lines and modulation by interferon-gamma treatment. Journal for ImmunoTherapy of Cancer. 2014; 2:P102.
36. Modena A, Ciccarese C, Iacovelli R, Brunelli M, Montironi R, Fiorentino M, Tortora G, Massari F. Immune Checkpoint Inhibitors and Prostate Cancer: A New Frontier? Oncol Rev. 2016; 10:293. https://doi.org/10.4081/oncol.2016.293.
37. Chen DS, Mellman I. Oncology meets immunology: the cancer-immunity cycle. Immunity. 2013; 39:1–10. https://doi.org/10.1016/j.immuni.2013.07.012.
38. Bernstein MB, Garnett CT, Zhang H, Velcich A, Wattenberg MM, Gameiro SR, Kalnicki S, Hodge JW, Guha C. Radiation-induced modulation of costimulatory and coinhibitory T-cell signaling molecules on human prostate carcinoma cells promotes productive antitumor immune interactions. Cancer Biother Radiopharm. 2014; 29:153-61. https://doi.org/10.1089/cbr.2013.1578.
39. Fankhauser CD, Curioni-Fontecedro A, Allmann V, Beyer J, Tischler V, Sulser T, Moch H, Bode PK. Frequent PD-L1 expression in testicular germ cell tumors. Br J Cancer. 2015; 113:411–3. https://doi.org/10.1038/bjc.2015.244.
40. Cima I, Schiess R, Wild P, Kaelin M, Schüffler P, Lange V, Picotti P, Ossola R, Templeton A, Schubert O, Fuchs T, Leippold T, Wyler S, et al. Cancer genetics-guided discovery of serum biomarker signatures for diagnosis and prognosis of prostate cancer. Proc Natl Acad Sci U S A. 2011; 108:3342–7. https://doi.org/10.1073/pnas.1013699108.
41. Wild PJ, Ikenberg K, Fuchs TJ, Rechsteiner M, Georgiev S, Fankhauser N, Noske A, Roessle M, Caduff R, Dellas A, Fink D, Moch H, Krek W, et al. p53 suppresses type II endometrial carcinomas in mice and governs endometrial tumour aggressiveness in humans. EMBO Mol Med. 2012; 4:808–24. https://doi.org/10.1002/emmm.201101063.
42. Ce L. SIFT Flow: Dense Correspondence across Scenes and Its Applications. IEEE Transactions on Pattern Analysis and Machine Intelligence. 2011; 33:978–94.
43. Fehrenbacher L, Spira A, Ballinger M, Kowanetz M, Vansteenkiste J, Mazieres J, Park K, Smith D, Artal-Cortes A, Lewanski C, Braiteh F, Waterkamp D, He P, et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet. 2016; 387:1837–46. https://doi.org/10.1016/S0140-6736(16)00587-0.

