INTRODUCTION
Following long non-coding RNAs (lncRNA) and microRNAs (miRNAs), circular RNAs (circRNAs) are a novel class of non-coding RNAs (ncRNAs), representing a new paradigm of gene regulation in many biological processes, such as proliferation, senescence and apoptosis [1, 2]. As early as the 1980s, circRNAs were initially discovered in RNA viruses, such as plant viroids. Unfortunately, reflecting their low abundance, circRNAs were originally considered as splicing byproducts or background noise [3, 4]. Currently, thousands of circRNAs have been identified and characterized with the development of high throughput RNA sequencing (RNA-seq) technologies and powerful computational analyses, and the expression level of some circRNAs is significantly higher than that of their corresponding linear isoforms from the same genes [5, 6].
To date, emerging studies have confirmed that circRNAs are endogenous, abundant and conserved in mammalian cells [7, 8]. The modulation of circRNAs on gene expression plays a significant role in various physiological and pathological procedures, particularly carcinogenesis [9, 10]. Moreover, subsequent reports have established the circRNAs-miRNAs-mRNA signaling axis, which participates in the occurrence and development of various cancers, such as oral carcinomas [11], hypopharyngeal cell carcinoma [12], osteosarcoma [13], etc. Thus, circRNAs show great potential as promising clinical biomarkers and therapeutic targets for cancers.
In this review, we primarily focus on the functions and potential roles of circRNAs in cancer research and therapy. In addition, we discuss some inspiring areas for future research, which may influence potential clinical implications, specifically serving as therapeutic targets. Finally, some algorithms detecting circRNAs from RNA-seq data are also preliminarily summarized.
CHARACTERISTICS OF circRNAs
Unlike linear RNAs, circRNAs form a covalently closed continuous loop without free caps and poly(A) tails [14-16] (Figure 1). Reflecting their unique structure, circRNAs have several noteworthy properties. First, the lack of exposed 3’ and 5’ terminals makes circRNAs more stable and less susceptible to degradation by ribonuclease R (RNase R) or other exonucleases. Studies have indicated that RNase R could degrade any linear RNA, resulting in the enrichment of pure circRNAs in RNA samples. After treatment with RNase R, numerous differentially expressed circRNAs could be precisely identified in tumor tissues relative to adjacent nonneoplastic tissues [17, 18]. Second, based on differences in backsplicing mechanisms, circRNAs could be divided into three types: intronic circRNAs, exonic circRNAs and exonic-intron circRNAs [19]. Quaking (QKI), an RNA-binding protein regulated in cancer cells during the epithelial-mesenchymal transition (EMT), promotes RNA backsplicing, producing mature circRNA [20]. Third, the majority of circRNAs are evolutionarily conserved. Thomas et al. confirmed an decreased SNP (single nucleotide polymorphism) density in circRNA sequences, particularly the miRNA-binding sites [21]. Fourth, most circRNAs are located in the cytoplasm, whereas the circRNAs from introns are abundant in the nucleus [22]. Nuclear circRNAs regulate RNA transcription by binding the cognate DNA locus [23] or RNA polymerase II [24]. Cytoplasmic circRNAs primarily function as miRNA sponges that inhibit miRNA expression and weaken the translation suppression on the corresponding target molecules [25] (Figure 2). Through paired-end RNA-seq on samples obtained from three ovarian cancer patients, Ahmed et al. showed that thousands of circRNA isoforms have predicted miRNA response elements (MREs). Over-expression of these circRNAs could effectively regulate the downstream target genes by sponging the miRNAs involved in carcinogenesis [26]. However, another study showed that cytoplasmic circHIAT1 could also serve as a miRNA reservoir, further stabilizing the miRNA expression in renal cell carcinoma (RCC) [27] (Figure 2). These contradictory findings indicate that further studies are needed to explore circRNA-miRNA networks and obtain a comprehensive understanding of the properties of circRNAs in cancer biology. Fifth, the expression profiles showed that these molecules are expressed in a tissue- or cancer-specific manner [28]. The database CircNet was used to successfully illustrate this profile across human transcriptome samples [29]. Last, it is clear that as potential tumor markers, circRNAs are sensitive and specific. Chen et al. reported that the sensitivity and specificity of has_circ_0000190, a downregulated circRNA in gastric cancer, is much better than that of carcino-embryonic antigen (CEA) and CA19-9, two commonly used biomarkers in cancer diagnoses [30]. Taken together, the abovementioned characteristics of circRNAs suggest that these molecules are promising molecular biomarkers and novel therapeutic targets for future drugs.
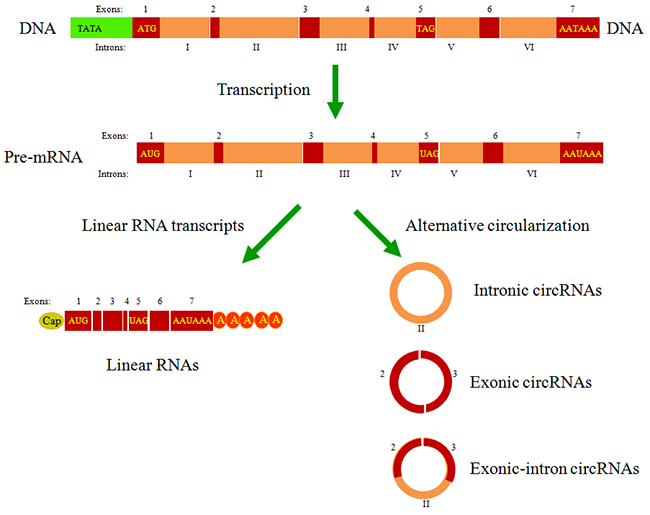
Figure 1: The types of circRNAs generated from the pre-mRNA by backsplicing. Three main types of circRNAs were recently identified: intronic circRNAs, exonic circRNAs and exonic-intron circRNAs. Studies have demonstrated that circRNAs are most likely generated co-transcriptionally from the same gene locus in competition with the formation of canonical linear RNA transcripts, a phenomenon called alternative circularization.
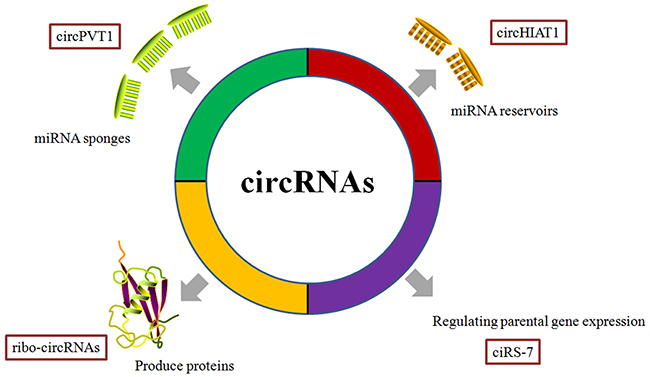
Figure 2: Functions of circRNAs determined through cancer research. CircRNAs serve as miRNA sponges/reservoirs or parental gene expression regulators involved in the onset and progression of cancers. Although the complete function of circRNAs in cancer biology remains unidentified clearly, several probable mechanisms have been reported so far. In general, a well-known function of circRNAs is of miRNA regulation through sponging miRNAs, if circRNA contains the corresponding miRNA binding sites. For example, circPVT1 promotes gastric cancers progression via acting as miR-125 sponge. Surprisingly, a specific circRNAs with corresponding miRNA binding sites, like cytoplasmic circHIAT1, serves as a miRNA reservoir, further stabilizing the miR-195-5p/29a-3p/29c-3p in renal cell carcinoma. In addition, some circRNAs, like ciRS-7/CDR1-AS, regulate parental genes through direct interactions. CiRS-7 is reported to enhance the CDR1 expression by interacting and stabilizing the sense CDR1 mRNA.
Additionally, although circRNAs were originally defined as a class of ncRNAs, newly published studies revealed that several circRNAs could function as mRNAs that produce functional proteins, indicating cap-dependent translation [31-33] (Figure 2). By screening the open reading frames (ORFs) using ribosome footprinting (RFP) datasets, Pamudurti et al. showed ribosomes associated circRNAs (ribo-circRNAs) have coding potential in Drosophila S2 cells. Further mass spectrometry analysis demonstrated that the encoded proteins from ribo-circRNA isoforms could be modulated by the transcription factor forkhead box protein O (FOXO) and starvation [34]. Similarly, through polysome profiling analysis, computational prediction and mass spectrometry in human embryonic kidney 293 cells and cervical cancer HeLa cells, Yang et al. identified hundreds of endogenous circRNAs with translation abilities, which were dependent on the motifs of N6-methyladenosine (m6A) in the sequence of the molecules, and the initiation factor eIF4G2 (eukaryotic translation initiation factor 4 gamma 2) and m6A reader YTHDF3 (YTH m6A RNA-binding protein 3) were shown to drive this m6A-mediated translation [35]. Legnini’s group provided another example of protein-coding circRNA, Circ-ZNF609. These studies showed that Circ-ZNF609 contains an ORF, similar to its parent linear transcript, and could be translated into protein in a cap-independent manner [36]. Further studies are needed to determine whether circRNA translation responds to different stress stimuli or cancer-related signals, eventually leading to tumorigenesis.
circRNAs ACT AS PROMISING TARGETS FOR CANCER BIOLOGY
In tumor biology, it is important to identify new potential biomarkers, which are differentially expressed between cancers and normal tissues. A number of studies have demonstrated that circRNAs are closely associated with cancer initiation and development [37-39] (Table 1). In addition, circRNAs could be secreted into human body fluids, such as saliva [40] and blood [41], or enriched in exosomes [42], emerging as effective biomarkers for the detection of cancers. The aberrant circRNAs discovered in cancers are discussed in detail below.
Table 1: CircRNA expression profiles in cancer tissues and cell lines
Cancers |
CircRNAs |
Expression level |
Samples |
Mechanisms |
Refs |
|---|---|---|---|---|---|
Oral carcinoma |
circRNA_100290 |
Up |
Tissues |
MiR-29 sponge |
[11] |
Osteosarcoma |
circUBAP2 |
Up |
Tissues |
MiR-143 sponge |
[13] |
Thyroid carcinoma |
hsa_circRNA_100395 |
Down |
Tissues |
Interaction with miR-141-3p/200a-3p |
[17] |
Renal carcinoma |
circHIAT1 |
Down |
Tissues |
MiR-195-5p/29a-3p/29c-3p sponge |
[27] |
Glioma |
cZNF292 |
Up |
Cells |
Promoting Wnt/β-catenin signaling |
[47] |
ciRS-7 |
Down |
Tissues/Cells |
MiR-671-5p targets |
[49] |
|
Lung cancer |
circRNA_100876 |
Up |
Tissues |
– |
[50] |
hsa_circ_0013958 |
Up |
tissues/Cells/Plasma |
MiR-134 sponge |
[51] |
|
cir-ITCH |
Down |
Tissues |
Inhibiting Wnt/β-catenin signaling |
[52] |
|
Hepatic carcinoma |
circZKSCAN1 |
Down |
Tissues |
Regulating cancer-related pathways |
[53] |
hsa_circ_0005075 |
Up |
Tissues |
MiR-23b-5p sponge |
[56] |
|
hsa_circ_0001649 |
Down |
Tissues |
– |
[57] |
|
circRNA_100338 |
Up |
Tissues/Cells |
MiR-141-3p sponge |
[58] |
|
hsa_circ_0004018 |
Down |
Tissues |
Interaction with miR-30e-5p/626 |
[59] |
|
ciRS-7 |
Up |
Targeting miR-7 |
[60] |
||
ciRS-7 |
No significance |
Tissues |
– |
[61] |
|
Colorectal cancer |
circ-BANP |
Up |
Tissues |
Promoting Akt phosphorylation |
[65] |
hsa_circ_0000069 |
Up |
Tissues |
– |
[66] |
|
circCCDC66 |
Up |
Tissues |
Regulating a subset of oncogenes |
[67] |
|
circRNA_001569 |
Up |
Tissues |
MiR-145 sponge |
[68] |
|
has_circ_0020397 |
Up |
Cells |
Promoting the miR-138 target genes |
[69] |
|
hsa_circ_001988 |
Down |
Tissues |
– |
[70] |
|
Gastric cancer |
hsa_circ_0000190 |
Down |
Tissues |
– |
[30] |
circRNA-0026 |
Down |
Tissues |
Regulating cancer-associated miRNAs |
[76] |
|
circPVT1 |
Up |
Tissues |
MiR-125 family sponge |
[77] |
|
hsa_circ_0000096 |
Down |
Tissues |
Inhibiting cyclin D1, CDK6 and MMP2/9 |
[78] |
|
circRNA_100269 |
Down |
Tissues/Cells |
Targeting miR-630 |
[79] |
|
hsa_circ_0001895 |
Down |
Tissues/Cells |
– |
[80] |
|
has_circ_0003159 |
Down |
Tissues |
– |
[81] |
|
Leukemia |
f-circRNAs |
Up |
Cells |
– |
[86] |
hsa_circ_0004277 |
Down |
Tissues |
– |
[88] |
|
Bladder carcinoma |
circRNA MYLK |
Up |
Tissues |
Binding competitively miRNA-29a family |
[90] |
circTCF25 |
Up |
Tissues |
miR-103a-3p/107 sponge |
[91] |
|
Esophageal carcinoma |
has_circ_0067934 |
Up |
Tissues/Cells |
– |
[93] |
Breast cancer |
has_circ_100219 |
Down |
Tissues |
Sequestering miRNAs |
[96] |
circDENND4C |
Up |
Cells |
HIF1α dependent |
[97] |
|
circ-ABCB10 |
Up |
Tissues |
MiR-1271 sponge |
[98] |
|
circ-Amotl1 |
Up |
Tissues/Cells |
inducing c-Myc nuclear translocation |
[99] |
circRNAs AND GLIOMAS
Glioblastoma or glioblastoma multiform (GBM) is a highly aggressive and invasive astrocytic cell neoplasm, with malignancy grades and histological subtypes. After initial diagnosis, the median survival time of glioma patients is approximately 12-15 months [43]. To inhibit disease progression and improve the therapeutic effect of patients with GBM, it is imperative to screen for more efficient biomarkers involved in glioma biological behaviors. Studies on ncRNAs, including lncRNAs [44] and circRNAs [45], have recently become a hotspot topic that could provide insight into gliomagenesis. Using the UROBORUS tool to analyze the RNA-seq data from 27 gliomas and 19 normal brain samples, studies have revealed more than 476 circRNAs differentially expressed in gliomas and control samples [46]. Further studies have demonstrated that cZNF292, an oncogenic circRNA, is significantly upregulated in human glioma cell lines U87 and U251. Targeted silencing of cZNF292 using short interfering RNA (siRNA) could induce cell cycle arrest via the Wnt/β-catenin signaling pathway, resulting in proliferation inhibition in glioma cells [47]. In addition, some circRNAs, such as ciRS-7/CDR1-AS (cerebellar degeneration-related protein 1 antisense), could stabilize and promote the expression of parental genes through direct interactions [48] (Figure 2). In gliomas, the upregulation of oncomiR miR-671-5p induced Argonaute-mediated ciRS-7 degradation. The overexpression of ciRS-7 positively regulated the expression of CDR1, abrogating the oncogenic function of miR-671-5p in glioma biopsies and cell lines [49] (Figure 3). Based on these findings, circRNAs are proposed to play key roles in the modification of glioma pathological profiles.
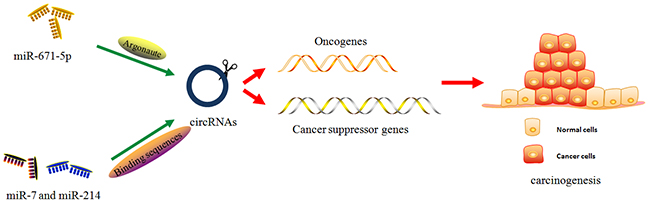
Figure 3: The miRNA-mediated cirRNA degradation. Emerging studies have demonstrated miRNA could mediate the cirRNA degradation via Argonaute-mediated signaling or circRNA-binding sequences, further to regulating the downstream cancer-associated genes, oncogenes or cancer suppressor genes. The detailed mechanisms are well explained in the relevant sections.
circRNAs AND LUNG CANCERS
The clinical application of circRNAs as diagnostic and prognostic indicators for lung cancers is increasingly investigated. Through the analysis of the circRNA expression features in lung cancer and adjacent normal tissues using qPCR, Yao et al. showed that circRNA_100876, encoded by RNF121 (ring finger protein 121) gene, is significantly upregulated in lung cancer samples, and Kaplan-Meier survival analysis revealed that patients with high expression levels of circRNA_100876 showed shorter overall survival times [50]. Similarly, using microarrays to screen the tumour-specific circRNA candidates in lung adenocarcinoma patients, Zhu et al. identified a significantly upregulated circRNA, hsa_circ_0013958, and found that hsa_circ_0013958 overexpression could promote tumor progression and apoptosis inhibition in lung adenocarcinoma via sponging miR-134 [51]. Wan et al. recently evaluated the roles of cir-ITCH (itchy E3 ubiquitin protein ligase) in the progression of lung cancer cells using a Taqman-based qPCR method, and the results showed that the expression of cir-ITCH was decreased in two different cancer cell lines, A549 and NIC-H460. The results of mechanism studies showed that ectopic expression of cir-ITCH could facilitate ITCH expression via sponging miR-214 or miR-7, further leading to the inhibition lung cancer cell proliferation. Interestingly, in turn, both miR-214 and miR-7 could degrade cir-ITCH through binding to the ITCH sequences of cir-ITCH [52], supporting miRNA-mediated circRNA degradation (Figure 3). Results above indicated that, by interfering with cancers-associated-miRNAs, circRNAs are differentially expressed in lung cancer and involved in tumor formation.
circRNAs AND HEPATOCELLULAR CARCINOMAS
The advent of RNA-seq or microarray initially suggested that circRNAs could be novel targets for clinical treatment in hepatocellular carcinomas (HCC). Furthermore, circRNA and its relative mRNA or miRNAs could co-exist in the same HCC patient and cooperate closely to promote carcinogenesis [53-55]. Currently, gene expression microarray has been regarded as a feasible and helpful method to recognize the molecular signatures, including circRNAs. Shang et al. used microarray analysis to investigate the circRNA expression in HCC and showed that the global expression profile of circRNAs between HCC tissue and normal liver tissue is significantly different. Among these alternative circRNAs, hsa_circ_0005075, located on chromosome 1, exhibits significantly downregulated expression and shows good potential for the diagnosis of HCC. Mechanistically, hsa_circ_0005075 acts as the sponge of miR-23b-5p to regulate the circRNA-miRNA-mRNA networks involved in liver cancer development [56]. Another low expressed circRNA, hsa_circ_0001649, derived from exons 26-29 of SHPRH (SNF2 histone linker PHD RING helicase) gene, is negatively associated with tumor size and occurrence. Knockdown of hsa_circ_0001649 could promote the expression of matrix metalloproteinases, further enhancing cancer cell metastasis [57]. Through a circRNA microarray analysis, one upregulated circRNAs, circRNA_100338, was recently identified as another potentially valuable biomarker for hepatitis B-related HCC. Functioning as an endogenous sponge for miR-141-3p, circRNA_100338 expression could enhance metastatic progression and shorten survival time [58]. Moreover, these differentially expressed circRNAs exhibit HCC-stage specific properties [59], further suggesting an independent role for the prognosis prediction of HCC. Through analyzing the expression profile of human circRNAs in HCC tissues, Han et al. identified circMTO1 as the significantly downregulated circRNA in HCC tissues. The results from this group suggested that circMTO1 could enhance the HCC cell proliferation and invasion by sponging miR-9 and increasing p21 level. Further Kaplan-Meier survival study revealed that the HCC patients with low circMTO1 expression have a shorter survival time [54]. Yu’s group executed qPCR analysis of circRNAs in 66 HCC tissues and reported a carcinogenic circRNA, ciRS-7, which promotes HCC cell proliferation by targeting miR-7 and elevating the expression of the direct target gene PIK3CD (phosphatidylinositol 3-kinase catalytic subunit delta) [60]. However, as measured by qPCR,, Xu et al. have showed no obvious difference in ciRS-7 expression in 108 pairs of HCC and matched non-tumor tissues [61]. These contradictory findings may reflect the inconsistent sample capacity. What–s more, as ciRS-7 contains more than 70 binding sites for miR-7 [62], these discrepant findings may also be due to the different down-stream factors of miR-7 influenced by ciRS-7 in different group of characters. Therefore, it is of importance to explore the detailed function and mechnism of ciRS-7 in pathophysiology and treatment of different HCC patients.
circRNAs AND COLORECTAL CANCERS
As potential predictive or diagnostic biomarkers for colorectal cancers (CRC), circRNAs are attracting increasing research attention. Moreover, the activities of these molecules as gene expression regulators or miRNA sponges have been associated with CRC progression and chemoradiation senstitivity, with high sensitivity and specificity [63-65]. The differential expression profiles of circRNAs in 3 paired CRC cancerous and normal tissues indicated the significant upregulation (n=136) or downregulation (n=243) of 2608 circRNAs in CRC samples (fold changes > 2). Moreover, circ-BANP, generated from exons 5-11 of the BANP gene, is the most upregulated circRNA, and knockout of circ-BANP obviously reduced the proliferation rates of HT29 and HCT116 CRC cells by attenuating Akt/PKB (protein kinase B) phosphorylation [66]. Using unsupervised hierarchical clustering analysis (UHCA) technology, Guo et al. identified the most upregulated circRNA, has_circ_0000069, in 6 paired CRC cancerous and normal tissues and further verified its expression level in cancer cells using qPCR. Function analysis suggested that has_circ_0000069 could promote cell proliferation and metastasis in HT29 CRC cells [67]. Similarly, Hsiao’s group showed that elevated circCCDC66, derived from exons 6-11 of CCDC66, is associated with the poor prognosis of CRC. Mechanistically, after hypoxic stress, accumulated circCCDC66 could protect a specific group of oncogenes, such as the MYC gene, from interactions with miR-33b, miR-93, and miR-185, leading to the proliferation and metastasis of HT29 and HCT116 cells [68]. The overexpression of has_circ_001569 in CRC cells also functions as a miR-145 sponge and promotes the expression of its targets genes E2F5 (E2F transcription factor 5) and BAG4 (BCL2 associated athanogene 4) [69]. High expression of has_circ_0020397 could positively influence the cell viability and invasion via promoting the expression of miR-138 target genes, telomerase reverse transcriptase (TERT) and programmed death-ligand 1 (PD-L1) [70]. Apart from these markedly upregulated circRNAs, the global reduction of the abundance of circRNAs, including hsa_circ_001988, has been negatively associated with the proliferation index, and these molecules may also represent potential novel targets for CRC therapy [71, 72]. These studies clearly suggest the clinical relevance of circRNAs as promising biomakers, since their expression is able to distinguish CRC tissues from the adjacent non-tumor tissues. Moreover, these differentially-expressed circRNAs afore-mentioned may also be recognized as molecular biomakers in assessing the curative effect of chemoradiotherapy on CRC patients. In addition, deep RNA-seq analysis of the exosomes from CRC cells showed that hundreds of candidate circRNAs could be transferred into exosomes [73], suggesting that extracellular-vesicle derived circRNAs in CRC patients may also represent potential targets for therapeutic interventions in the future.
circRNAs AND GASTRIC CANCERS
CircRNA profiles and circRNA-miRNA-mRNA interactions in gastric cancers (GC) have recently been discussed, and studies have demonstrated that differentially expressed circRNAs could serve as novel and stable makers for the diagnosis and progression of GC patients [74, 75]. In a recent study, based on microarray chip technology, Sui et al. screened 467 significant differences (>2-fold changes, p-values<0.05) in circRNA expression, with 214 upregulated and 253 downregulated molecules identified in 8 GC tissues and adjacent tissue samples. In-house miRNA target prediction software revealed miRNA-binding sites in 69 differentially expressed circRNAs [76]. Using a circRNA microarray, Huang et al. confirmed only one significantly differently down-expressed circRNA, circRNA-0026, in GC tissues. Biomathematical analyses indicated that circRNA-0026 regulates RNA transcription, RNA metabolism, and other biological functions by adsorbing cancer-associated miRNAs [77]. Thereby, these differentially expressed circRNAs could sponge corresponding miRNAs to modulate the expression of their target genes. For example, the upregulated circPVT1, derived from the PVT1 locus on chromosome 8q24, promotes GC progression via acting as a sponge for the miR-125 family [78]. Additionally, the low expressed hsa_circ_0000096 could regulate the expression levels of cell cycle- or metastasis-correlated proteins, affecting the growth and migration of GC cells [79]. Another downregulated circRNA_100269 could target miR-630, resulting in GC cell growth suppression [80]. These data laid the foundation to investigate the attractive clinical significance of distinct circRNA candidates for GC pathobiology. According to the data obtained from CircBase database, Shao et al. showed that has_circ_0001895 is downregulated in GC cells and significantly associated with histological type and CEA expression [81]. Tian et al. demonstrated the negatively association between has_circ_0003159 expression and clinicopathological factors of GC patients [82]. Similarly, another two typically downregulated circRNAs, hsa_circ_002059 and hsa_circ_00001649, have been associated with clinicopathological characteristics, such as differentiation, TNM stage, gender, etc. [83, 84]. In addition, Zhang et al. generated a circRNA-based classifier to assess the recurrence risk after radical surgery using four significantly differentially expressed circRNAs in GC samples. These studies showed that this predictive tool could effectively distinguish the high-risk patients with recurrence [85].
circRNAs AND HEMATOLOGICAL MALIGNANCIES
Increasing evidence has demonstrated that fusion genes, encoded by aberrant chromosomal translocations, are undoubtedly involved in malignant hematological disorders. In addition, chromosomal rearrangements affect ncRNAs levels in tumor cells through complicated mechanisms, such as the direct generation of fusion circRNAs (f-circRNAs) [86] (Figure 4). Further studies have suggested that these cancer-associated fusion genes could give rise to multiple f-circRNAs, such as f-circPR and f-circM9_1, which show pro-proliferative and proto-oncogenic properties [87]. Using ACFS software to analysis the data from single- and paired-ended RNA-Seq, You et al. identified a several differentially expressed circRNAs between AML and acute promyelocytic leukemia (APL) patients. The ACFS database could further verify the f-circRNAs in leukemia tissues with high accuracy and a low false discovery rate [88]. These data support the notion that f-circRNA plays an active function in favoring the progression of hematological malignancies. Apart from the f-circRNAs, Li et al. identified another remarkably downregulated circRNA, has_circ_0004277, in a cohort of 115 human acute myelocytic leukemia (AML) samples. The findings from this group indicated that standard chemotherapy could recover the expression of has_circ_0004277 and its corresponding linear isoform WDR37 (WD repeat domain 37), suggesting that elevated has_circ_0004277 may affect the chemotherapy response [89].
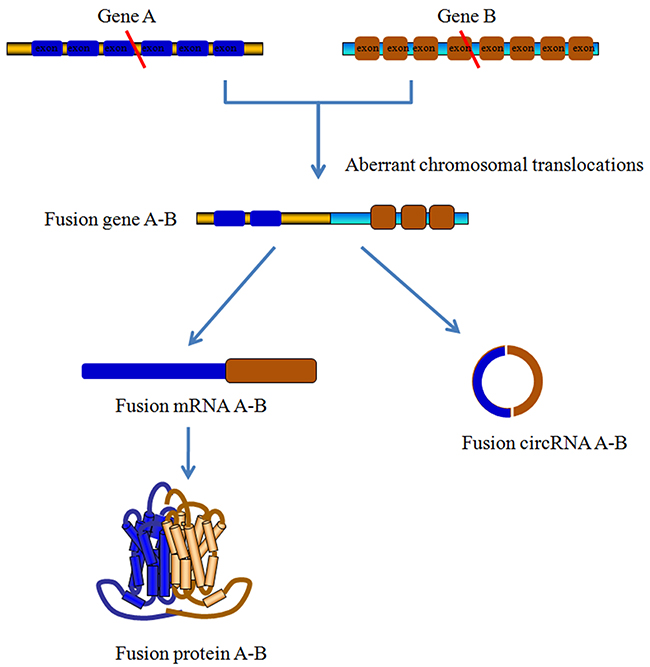
Figure 4: Fusion-circRNAs derived from aberrant chromosomal translocations. Cancer-related chromosomal translocations could produce linear fusion mRNA and fusion circRNAs (f-circRNAs). The formation of f-circRNAs is shown here. Briefly, after the chromosomal translocation of Gene A and Gene B, the fusion and backsplice junctions of one exon from Gene A and another exon from Gene B result in the formation of f-circRNA. Meanwhile, corresponding linear fusion mRNA can be translated into functional fusion proteins. Both fusion proteins and f-circRNAs involve in the cancer development and progression, especially the hematological malignancies.
circRNAs AND BLADDER CARCINOMAS
Competing endogenous RNAs (ceRNA) have recently been identified specifically upon cancer onset and development, reflecting the pivotal effect of these molecules on the expression of cancer-associated miRNAs (Figure 2). A recent study analyzing the ceRNA network from the microarray data of bladder cancer (BC) and para-cancer tissues showed that uniform MREs were observed in both circRNAs and lncRNAs. For example, lncRNA H19 and circRNA MYLK competitively bind miR-29a-3p via its corresponding MREs, resulting in an increase of miR-29a-3p target genes. Moreover, through regulating the expression of miR-29a, circRNA MYLK plays a key role in accelerating cancer progression of BC patients [90, 91]. In addition, Zhong et al. identified 469 dysregulated cercRNAs (285 upregulated and 184 downregulated) in BC compared with normal samples. Bioinformatics approaches preliminarily showed that these abnormally expressed circRNAs, particularly circTCF25, participate in multiple cancer-associated signaling pathways, functioning as valuable biomarkers for the diagnosis and therapy of BC [92]. However, the detailed mechanisms and functions of these differentially expressed circRNAs in BC development require further discussion.
circRNAs AND ESOPHAGEAL CELL CARCINOMAS
Function analysis of circRNA in esophageal cell carcinomas (ESCC) has provided much evidence supporting the exploitation of RNA-based treatment strategy. Expression profile analysis and bioinformatics methods revealed that clusters of circRNAs are aberrantly expressed in radioresistant KYSE-150R esophageal cancer cells compared with parental KYSE-150 cells (fold-change ≥2.0 and P < 0.05). Gene ontology (GO) analysis further demonstrated that the genes in the Wnt signaling pathway are the main targets of these candidate circRNAs [93]. In addition, a highly expressed circRNA, has_circ_0067934, has been associated with poor differentiation and TNM stage in ESCC patients. Silencing of hsa_circ_0067934 using siRNA could alleviate the proliferation and migration of ECA-109 and TE-13 ESCC cells [94]. Taken together, these findings indicated key roles for circRNAs in ESCC development and treatment.
circRNAs AND BREAST CANCER
Latest studies have shown that circRNAs could be the important regulatory molecules for the development of breast cancer [95], one of the most frequently occurring cancers worldwide [96]. Using the Arraystar Human circRNA Array to construct a genome-wide circRNA profile, researchers have found about 1155 differently expressed circRNAs (715 up-regulated and 440 down-regulated) in breast cancer tissues. Then they choose three elevated circRNAs (has_circ_103110, has_circ_104689 and has_circ_104821) and three down-regulated circRNA (has_circ_006054, has_circ_100219 and has_circ_406697) as the promising candidates for the diagnosis of breast cancer. The areas under the curve (AUC) of has_circ_100219 was 0.78, indicating the highest diagnostic accuracy [97]. Liang et al. identified the hypoxic-inducible factor 1 alpha (HIF1α) associated cricRNA, circDENND4C, and found that knockdown of circDENND4C by siRNAs resulted in decreased proliferation of breast cancer cells MCF7 and MDA-MB-231 in hypoxic environment. The clinical relevance analysis indicated a positive correlation between circDENND4C level and the tumor size [98]. Through investigating the differently expressed circRNAs in human breast cancers, another group has found that circ-ABCB10, the most significantly up-regulated circRNA, could facilitate the cancer proliferation and progression by sponging miR-1271 [99]. Meanwhile, Yang et al. demonstrated the tumorigenic capacity of circ-Amotl1, an important circRNA highly expressed in breast cancer tissues and cells. Mechanistically, ectopic expression of circ-Amotl1 mainly triggered the nuclear translocation of proto-oncogene c-Myc, enhancing the c-Myc stability and furtherly increaseing the c-myc targets [100]. Taken together, all these studies reveal the important regulatory role of circRNAs for breast cancer pathogenesis.
circRNAs AND PANCREATIC DUCTAL ADENOCARCINOMAS
Accumulating studies have reported a cluster of circRNAs in the malignant behavior of pancreatic ductal adenocarcinomas (PDAC). The abundance of aberrantly expressed circRNAs, identified using microarray technology, have been implicated in carcinogenesis, providing novel insight into PDAC biology [101, 102].
DATABASES FOR circRNA DETECTING IN CANCER CELLS
In recent years, several algorithms have been developed to annotate and identify circRNAs from RNA-seq data obtained from cancer samples (Table 2). The CircNet database provides comprehensive information on circRNAs, including genome sequences and loci, and more importantly, this database also provides information for integrated ciRNA-miRNA-mRNA networks [29]. The UROBORUS database, developed by Song et al., contains the low expressed circRNAs from the total RNA-seq without RNase R treatment [46]. CIRCexplorer tools can detect candidate circRNAs in breast cancer tissues with high accuracy and good sensitivity, but it requires their gene annotation [41]. ACFS software proposes a fast and quantitative detection method to identify circRNAs from single- and paired-ended RNA-seq datasets of leukemia. In addition, compared with other databases, ACFS shows the highest accuracy and lowest false discovery rate [88]. Circ-Seq is a comprehensive algorithm used to define coincident circRNAs from public breast cancer samples provided by The Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression Project (GTEx) [103]. The website circBase has offered a method to freely access and download the merged and unified data sets of cancer related circRNAs [83, 104]. The study by Ghosal et al. introduced another detection tool, Circ2Traits, for circRNAs potentially associated with human diseases, like cancers. Using this algorithm, researchers could also get the cancer associated single nucleotide polymorphisms (SNPs) on circRNA loci [105]. StarBase v2.0, designed by Li’s group, systematically provides the circRNA–miRNA-mRNA and protein–circRNA interaction networks from large-scale data sets of normal tissues and cancer cells [106]. Other databases, such as PcircRNA_finder [107], CircInteractome [108], deepBase v2.0 [109], CIRI2 [110], et al. have not currently been used to screen circRNAs in cancerous samples, but these algorithms would offer more choices to compare and obtain more reliable results. Although these algorithms have their own advantages, problems, such as little overlap in prediction and no clear gold standard, should be addressed in the future [111]. Therefore, an important step for using these bioinformatics approaches is to adjust the search strategy appropriately to elevate the confidence with an appropriate threshold.
Table 2: Databases used for circRNAs detection in cancers tissues and cells
Databases |
De novo? |
Characteristics |
URL |
Refs |
|---|---|---|---|---|
CircNet |
No |
Tissue-specific circRNA expression profiles; |
http://circnet.mbc.nctu.edu.tw/ |
[29] |
CIRCexplorer |
No |
Depending on the mapping strategy of aligners; |
[41] |
|
UROBORUS |
No |
Total RNA-seq data with no poly(A) depletion or RNase R treatment; |
http://uroborus.openbioinformatics.org/en/latest/ |
[46] |
Acfs |
Yes |
Quantification and fast; |
[87] |
|
Circ-Seq |
No |
Genomic annotation; |
[102] |
|
circBase |
No |
Downloading freely the merged and unified data sets of circRNAs |
[103] |
|
Circ2Traits |
No |
A disease-circRNA association database |
http://gyanxet-beta.com/circdb/ |
[104] |
starBase v2.0 |
No |
Providing comprehensive circRNA–miRNA-mRNA and protein–circRNA interaction networks |
[105] |
DISCUSSION AND CONCLUSION
The recent revelation of the widespread existence of circRNAs sheds an attractive light on the research and therapy of human diseases. Mechanistic and functional studies have revealed that circRNAs are no longer considered useless junk sequences in genomes. Although current knowledge of the biological effect of circRNAs remains limited, the exploration of the roles of circRNA in cancer biology could bring oncologist closer to the tumor cure. Particularly, in recent years, advancements in molecular medicine have helped to predict the potential for circRNAs serving as profound markers for cancer diagnosis and prognosis [112]. Currently, the abundance of natural compounds has emphasized the anti-tumor effects of these molecules via different signaling pathways [113]. Exploring the therapeutic agents that interfere with circRNAs or their effectors could reveal beneficial strategies to eradicate cancer cells.
The clinical potential for the use of circRNAs as diagnostic biomarkers and predictive targets for cancers continues to expand. Recent systemic deep RNA-seq studies have verified that the differential expression of circRNAs is well correlated with some clinicopathological features, such as tumor TNM stages, recurrence, metastasis, etc [61, 85]. For example, the upregulated level of circPVT1 in GC tissues could be served as an independent prognostic marker for overall survival and disease-free survival of cancer patients [78]. Another candidate circRNA, hsa_circ_0004018, has been proved to harbor the tumor stage-specific expression features in diverse chronic liver diseases [59]. Meanwhile, with the AUC of 0.75 in tissues and 0.60 in plasma, respectively, hsa_circ_0000190 showed better sensitivity and specificity than commonly used biomarkers such as CEA and CA19-9 for GC patients [30]. In addition, the specimens for the circRNAs detection should be easily acquired from the cancer patients, including the blood, urine, saliva, etc. CircRNAs have been proved to be detected in the cell-free components of the body fluids, with benign extracellular stability [114]. Reports on circRNAs in exosomes might represent a novel aspect for life sciences [42]. Extracellular vesicles, including exosomes, isolated from the blood and other biofluids of cancerous patients, contain a specific cargo of RNA and protein species, which regulate various cell bio-behaviors in human diseases with both diagnostic and therapeutic implications [115]. Recent studies have demonstrated that cancer-associated circRNAs could be identified in tumor-derived exosomes in patients with colon cancer or leukemia [73, 87]. Thus, the circRNAs in body fluids could be used as promising pathognomonic markers for different types of cancers, particularly the cancers for which biopsies are difficult to obtain. Though these above-mentioned studies have showed preliminary findings about the clinical significances of circRNAs, much clinical relevance still remains to be clarified about the roles of circRNAs before coming full article.
In addition, other cancer-related biological pathways, such as immunocyte aging within the immune system, could combat or sustain cancer development and progression. Recently, Wang et al. analyzed the comprehensive circRNA profile in individuals undergoing immunosenescence, and showed that circular RNA100783 plays an important role in immune T cell aging [116]. Further examination revealed that the expression of circRNAs could be regulated by several Toll-like receptors [117], which induce the appropriate immune responses in cancer patients [118]. Although the detailed mechanisms of circRNAs in immune regulation remain largely unclear, understanding the potential functions of circRNAs in immune responses is of significant relevance to cancer therapy. Thus, extensive studies should focus on the role of circRNAs in certain aspects of cancer biology, such as immunity regulation signaling, which may represent another useful therapeutic strategy for human cancers.
PERSPECTIVE
As a kind of newly discovered endogenous ncRNAs, circRNAs have been proved to involve in multiple cellular processes of tumorigenesis by regulating gene expression at the transcriptional or post-transcriptional level. Thousands of well-expressed circRNAs are abundant and stable in cancer cells, suggesting that they might be the key factors for cancer occurrence and development. Especially, numerous reports have shown that the circRNAs-miRNA-mRNA axis plays a pivotal role in several cancer-associated biological behaviors, such as proliferation, angiogenesis and metastasis [51, 119, 120]. Meanwhile, by interacting with miRNAs, circRNAs also have the potential of impacting a variety of signaling pathways in cancers, such as vascular endothelial growth factor A (VEGFA) signaling pathway [91], large tumor suppressor kinase 1 (LATS1) [121], Src family tyrosine kinases (SFKs) [122] and so on. More importantly, the specificity and sensitivity of circRNAs are helpful to make them be possible biomakers, like the hsa_circ_0004018 in hepatocellular carcinoma [59]. With the progress of molecular biological technique and bioinformatics, other novel circRNAs that were identified recently might also serve as promising biomakers.
However, in the clinical setting, the detailed functions of circRNAs in human cancers need to be further explored. And several outstanding issues of circRNA study to focus on in the future are as follows: (1) Although identified in the 1980s [123], the information of circRNAs in human diseases, especially cancers, is still limited. There is a need to screen and recognize new candidate circRNAs that can serve as promising therapeutic targets, not just the diagnostic and prognosis biomarkers. (2) To data, rapid development of technology have made RNA-seq to be the wide method to identify circRNAs in cancer samples and cells accurately. However, as the next generation sequencing is expensive and time-consuming, the use of circRNAs for the detection of cancers needs to be detected in much easier and cheaper methods. For example, two other methods, qPCR and in-situ hybridization, have been proved to be more sensitive and specific for detecting circRNAs [124]. (3) The potential of circRNAs as valuable makers for cancer diagnose and prognosis is needs to be further well-explored in the future. Because most studies above-mentioned lack the tangible proofs and well-established standard supporting the reasonable clinical application.
ACKNOWLEDGMENTS
We thank Elsevier–s English Language Editing Service for assistance with language editing. Dr. Zhijie Xu is right now a Postdoctoral Fellow in Department of Pharmacy, Xiangya Hospital, Central South University, and thanks all members in Institute of Hospital Pharmacy for their critical comments.
COMPETING INTEREST
The authors declare that they have no potential conflicts in relation to publishing this paper.
FUNDING
This work is supported by the National Natural Science Foundation of China (No. 81572946, 81703036), the China Postdoctoral Science Foundation (No. 2017M610510), Changsha Science and Technology Project (No. k1508024-31), The Postdoctoral Science Foundation of Central South University (185702), and Clinical and Rehabilitation Research Foundation of Xiangya hospital - Beidaweiming.
REFERENCES
1. Maiese K. Disease onset and aging in the world of circular RNAs. J Transl Sci. 2016; 2:327–9.
2. Beermann J, Piccoli MT, Viereck J, Thum T. Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev. 2016; 96:1297–325. https://doi.org/10.1152/physrev.00041.2015.
3. Hou LD, Zhang J. Circular RNAs: an emerging type of RNA in cancer. Int J Immunopathol Pharmacol. 2017; 30:1–6. https://doi.org/10.1177/0394632016686985.
4. Liu J, Liu T, Wang X, He A. Circles reshaping the RNA world: from waste to treasure. Mol Cancer. 2017; 16:58. https://doi.org/10.1186/s12943-017-0630-y.
5. Qu S, Yang X, Li X, Wang J, Gao Y, Shang R, Sun W, Dou K, Li H. Circular RNA: a new star of noncoding RNAs. Cancer Lett. 2015; 365:141–8. https://doi.org/10.1016/j.canlet.2015.06.003.
6. He J, Xie Q, Xu H, Li J, Li Y. Circular RNAs and cancer. Cancer Lett. 2017; 396:138–44. https://doi.org/10.1016/j.canlet.2017.03.027.
7. Rong D, Tang W, Li Z, Zhou J, Shi J, Wang H, Cao H. Novel insights into circular RNAs in clinical application of carcinomas. Onco Targets Ther. 2017; 10:2183–8. https://doi.org/10.2147/OTT.S134403.
8. Chen LL. The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol. 2016; 17:205–11. https://doi.org/10.1038/nrm.2015.32.
9. Chen Y, Li C, Tan C, Liu X. Circular RNAs: a new frontier in the study of human diseases. J Med Genet. 2016; 53:359–65. https://doi.org/10.1136/jmedgenet-2016-103758.
10. Zhao ZJ, Shen J. Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA Biol. 2015:1–8. https://doi.org/10.1080/15476286.2015.1122162.
11. Chen L, Zhang S, Wu J, Cui J, Zhong L, Zeng L, Ge S. circRNA_100290 plays a role in oral cancer by functioning as a sponge of the miR-29 family. Oncogene. 2017. https://doi.org/10.1038/onc.2017.89.
12. Cao S, Wei D, Li X, Zhou J, Li W, Qian Y, Wang Z, Li G, Pan X, Lei D. Novel circular RNA expression profiles reflect progression of patients with hypopharyngeal squamous cell carcinoma. Oncotarget. 2017; 8:45367–79. https://doi.org/10.18632/oncotarget.17488.
13. Zhang H, Wang G, Ding C, Liu P, Wang R, Ding W, Tong D, Wu D, Li C, Wei Q, Zhang X, Li D, Liu P, et al. Increased circular RNA UBAP2 acts as a sponge of miR-143 to promote osteosarcoma progression. Oncotarget. 2017; 8:61687–61697. https://doi.org/10.18632/oncotarget.18671.
14. Wang F, Nazarali AJ, Ji S. Circular RNAs as potential biomarkers for cancer diagnosis and therapy. Am J Cancer Res. 2016; 6:1167–76.
15. Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014; 56:55–66. https://doi.org/10.1016/j.molcel.2014.08.019.
16. Vicens Q, Westhof E. Biogenesis of circular RNAs. Cell. 2014; 159:13–4. https://doi.org/10.1016/j.cell.2014.09.005.
17. Peng N, Shi L, Zhang Q, Hu Y, Wang N, Ye H. Microarray profiling of circular RNAs in human papillary thyroid carcinoma. PLoS One. 2017; 12:e0170287. https://doi.org/10.1371/journal.pone.0170287.
18. Xuan L, Qu L, Zhou H, Wang P, Yu H, Wu T, Wang X, Li Q, Tian L, Liu M, Sun Y. Circular RNA: a novel biomarker for progressive laryngeal cancer. Am J Transl Res. 2016; 8:932–9.
19. Wang Y, Mo Y, Gong Z, Yang X, Yang M, Zhang S, Xiong F, Xiang B, Zhou M, Liao Q, Zhang W, Li X, Li X, et al. Circular RNAs in human cancer. Mol Cancer. 2017; 16:25. https://doi.org/10.1186/s12943-017-0598-7.
20. Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA, Roslan S, Schreiber AW, Gregory PA, Goodall GJ. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015; 160:1125–34. https://doi.org/10.1016/j.cell.2015.02.014.
21. Thomas LF, Saetrom P. Circular RNAs are depleted of polymorphisms at microRNA binding sites. Bioinformatics. 2014; 30:2243–6. https://doi.org/10.1093/bioinformatics/btu257.
22. Ebbesen KK, Kjems J, Hansen TB. Circular RNAs: identification, biogenesis and function. Biochim Biophys Acta. 2016; 1859:163–8. https://doi.org/10.1016/j.bbagrm.2015.07.007.
23. Conn VM, Hugouvieux V, Nayak A, Conos SA, Capovilla G, Cildir G, Jourdain A, Tergaonkar V, Schmid M, Zubieta C, Conn SJ. A circRNA from SEPALLATA3 regulates splicing of its cognate mRNA through R-loop formation. Nat Plants. 2017; 3:17053. https://doi.org/10.1038/nplants.2017.53.
24. Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L, Zhu P, Chang Z, Wu Q, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015; 22:256–64. https://doi.org/10.1038/nsmb.2959.
25. Cortes-Lopez M, Miura P. Emerging functions of circular RNAs. Yale J Biol Med. 2016; 89:527–37.
26. Ahmed I, Karedath T, Andrews SS, Al-Azwani IK, Mohamoud YA, Querleu D, Rafii A, Malek JA. Altered expression pattern of circular RNAs in primary and metastatic sites of epithelial ovarian carcinoma. Oncotarget. 2016; 7:36366–81. https://doi.org/10.18632/oncotarget.8917.
27. Wang K, Sun Y, Tao W, Fei X, Chang C. Androgen receptor (AR) promotes clear cell renal cell carcinoma (ccRCC) migration and invasion via altering the circHIAT1/miR-195-5p/29a-3p/29c-3p/CDC42 signals. Cancer Lett. 2017; 394:1–12. https://doi.org/10.1016/j.canlet.2016.12.036.
28. Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G, Liang L, Gu J, He X, et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun. 2016; 7:11215. https://doi.org/10.1038/ncomms11215.
29. Liu YC, Li JR, Sun CH, Andrews E, Chao RF, Lin FM, Weng SL, Hsu SD, Huang CC, Cheng C, Liu CC, Huang HD. CircNet: a database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res. 2016; 44:D209–15. https://doi.org/10.1093/nar/gkv940.
30. Chen S, Li T, Zhao Q, Xiao B, Guo J. Using circular RNA hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin Chim Acta. 2017; 466:167–71. https://doi.org/10.1016/j.cca.2017.01.025.
31. Wang Y, Wang Z. Efficient backsplicing produces translatable circular mRNAs. RNA. 2015; 21:172–9. https://doi.org/10.1261/rna.048272.114.
32. Tatomer DC, Wilusz JE. An unchartered journey for ribosomes: circumnavigating circular RNAs to produce proteins. Mol Cell. 2017; 66:1–2. https://doi.org/10.1016/j.molcel.2017.03.011.
33. Schneider T, Bindereif A. Circular RNAs: coding or noncoding? Cell Res. 2017. https://doi.org/10.1038/cr.2017.70.
34. Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E, Shenzis S, Samson M, Dittmar G, et al. Translation of CircRNAs. Mol Cell. 2017; 66:9-21 e7. https://doi.org/10.1016/j.molcel.2017.02.021.
35. Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, Jin Y, Yang Y, Chen LL, Wang Y, Wong CC, Xiao X, Wang Z. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res. 2017; 27:626–41. https://doi.org/10.1038/cr.2017.31.
36. Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, Fatica A, Santini T, Andronache A, Wade M, Laneve P, Rajewsky N, Bozzoni I. Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol Cell. 2017; 66:22-37 e9. https://doi.org/10.1016/j.molcel.2017.02.017.
37. Qu S, Zhong Y, Shang R, Zhang X, Song W, Kjems J, Li H. The emerging landscape of circular RNA in life processes. RNA Biol. 2016:1–8. https://doi.org/10.1080/15476286.2016.1220473.
38. Dong Y, He D, Peng Z, Peng W, Shi W, Wang J, Li B, Zhang C, Duan C. Circular RNAs in cancer: an emerging key player. J Hematol Oncol. 2017; 10:2. https://doi.org/10.1186/s13045-016-0370-2.
39. Xin Z, Ma Q, Ren S, Wang G, Li F. The understanding of circular RNAs as special triggers in carcinogenesis. Brief Funct Genomics. 2017; 16:80–6. https://doi.org/10.1093/bfgp/elw001.
40. Bahn JH, Zhang Q, Li F, Chan TM, Lin X, Kim Y, Wong DT, Xiao X. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem. 2015; 61:221–30. https://doi.org/10.1373/clinchem.2014.230433.
41. Galasso M, Costantino G, Pasquali L, Minotti L, Baldassari F, Corra F, Agnoletto C, Volinia S. Profiling of the predicted circular RNAs in ductal insitu and invasive breast cancer: a pilot study. Int J Genomics. 2016; 2016:4503840. https://doi.org/10.1155/2016/4503840.
42. Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J, Chen D, Gu J, He X, Huang S. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 2015; 25:981–4. https://doi.org/10.1038/cr.2015.82.
43. Yan Y, Xu Z, Dai S, Qian L, Sun L, Gong Z. Targeting autophagy to sensitive glioma to temozolomide treatment. J Exp Clin Cancer Res. 2016; 35:23. https://doi.org/10.1186/s13046-016-0303-5.
44. Yan Y, Xu Z, Li Z, Sun L, Gong Z. An Insight into the Increasing Role of LncRNAs in the Pathogenesis of Gliomas. Front Mol Neurosci. 2017; 10:53. https://doi.org/10.3389/fnmol.2017.00053.
45. Kumar L, Shamsuzzama, Haque R, Baghel T, Nazir A. Circular RNAs: the emerging class of non-coding RNAs and their potential role in human neurodegenerative diseases. Mol Neurobiol. 2016. https://doi.org/10.1007/s12035-016-0213-8.
46. Song X, Zhang N, Han P, Moon BS, Lai RK, Wang K, Lu W. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res. 2016; 44:e87. https://doi.org/10.1093/nar/gkw075.
47. Yang P, Qiu Z, Jiang Y, Dong L, Yang W, Gu C, Li G, Zhu Y. Silencing of cZNF292 circular RNA suppresses human glioma tube formation via the Wnt/beta-catenin signaling pathway. Oncotarget. 2016; 7:63449–55. https://doi.org/10.18632/oncotarget.11523.
48. Peng L, Yuan XQ, Li GC. The emerging landscape of circular RNA ciRS-7 in cancer (Review). Oncol Rep. 2015; 33:2669–74. https://doi.org/10.3892/or.2015.3904.
49. Barbagallo D, Condorelli A, Ragusa M, Salito L, Sammito M, Banelli B, Caltabiano R, Barbagallo G, Zappala A, Battaglia R, Cirnigliaro M, Lanzafame S, Vasquez E, et al. Dysregulated miR-671-5p / CDR1-AS / CDR1 / VSNL1 axis is involved in glioblastoma multiforme. Oncotarget. 2016; 7:4746–59. https://doi.org/10.18632/oncotarget.6621.
50. Yao JT, Zhao SH, Liu QP, Lv MQ, Zhou DX, Liao ZJ, Nan KJ. Over-expression of CircRNA_100876 in non-small cell lung cancer and its prognostic value. Pathol Res Pract. 2017; 213:453–6. https://doi.org/10.1016/j.prp.2017.02.011.
51. Zhu X, Wang X, Wei S, Chen Y, Chen Y, Fan X, Han S, Wu G. hsa_circ_0013958: a circular RNA and potential novel biomarker for lung adenocarcinoma. FEBS J. 2017; 284:2170–82. https://doi.org/10.1111/febs.14132.
52. Wan L, Zhang L, Fan K, Cheng ZX, Sun QC, Wang JJ. Circular RNA-ITCH suppresses lung cancer proliferation via inhibiting the Wnt/beta-catenin pathway. Biomed Res Int. 2016; 2016:1579490. https://doi.org/10.1155/2016/1579490.
53. Yao Z, Luo J, Hu K, Lin J, Huang H, Wang Q, Zhang P, Xiong Z, He C, Huang Z, Liu B, Yang Y. ZKSCAN1 gene and its related circular RNA (circZKSCAN1) both inhibit hepatocellular carcinoma cell growth, migration, and invasion but through different signaling pathways. Mol Oncol. 2017; 11:422–37. https://doi.org/10.1002/1878-0261.12045.
54. Han D, Li J, Wang H, Su X, Hou J, Gu Y, Qian C, Lin Y, Liu X, Huang M, Li N, Zhou W, Yu Y, et al. Circular RNA MTO1 acts as the sponge of miR-9 to suppress hepatocellular carcinoma progression. Hepatology. 2017. https://doi.org/10.1002/hep.29270.
55. Ren S, Xin Z, Xu Y, Xu J, Wang G. Construction and analysis of circular RNA molecular regulatory networks In liver cancer. Cell Cycle. 2017:0. https://doi.org/10.1080/15384101.2017.1346754.
56. Shang X, Li G, Liu H, Li T, Liu J, Zhao Q, Wang C. Comprehensive circular RNA profiling reveals that hsa_circ_0005075, a new circular RNA biomarker, is involved in hepatocellular crcinoma development. Medicine (Baltimore). 2016; 95:e3811. https://doi.org/10.1097/MD.0000000000003811.
57. Qin M, Liu G, Huo X, Tao X, Sun X, Ge Z, Yang J, Fan J, Liu L, Qin W. Hsa_circ_0001649: a circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016; 16:161–9. https://doi.org/10.3233/CBM-150552.
58. Huang XY, Huang ZL, Xu YH, Zheng Q, Chen Z, Song W, Zhou J, Tang ZY, Huang XY. Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-100338/miR-141-3p pathway in hepatitis B-related hepatocellular carcinoma. Sci Rep. 2017; 7:5428. https://doi.org/10.1038/s41598-017-05432-8.
59. Fu L, Yao T, Chen Q, Mo X, Hu Y, Guo J. Screening differential circular RNA expression profiles reveals hsa_circ_0004018 is associated with hepatocellular carcinoma. Oncotarget. 2017; 8:58405–58416. https://doi.org/10.18632/oncotarget.16881.
60. Yu L, Gong X, Sun L, Zhou Q, Lu B, Zhu L. The circular RNA Cdr1as act as an oncogene in hepatocellular carcinoma through targeting miR-7 expression. PLoS One. 2016; 11:e0158347. https://doi.org/10.1371/journal.pone.0158347.
61. Xu L, Zhang M, Zheng X, Yi P, Lan C, Xu M. The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J Cancer Res Clin Oncol. 2017; 143:17–27. https://doi.org/10.1007/s00432-016-2256-7.
62. Xu H, Guo S, Li W, Yu P. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci Rep. 2015; 5:12453. https://doi.org/10.1038/srep12453.
63. Taborda MI, Ramirez S, Bernal G. Circular RNAs in colorectal cancer: possible roles in regulation of cancer cells. World J Gastrointest Oncol. 2017; 9:62–9. https://doi.org/10.4251/wjgo.v9.i2.62.
64. Zhang P, Zuo Z, Shang W, Wu A, Bi R, Wu J, Li S, Sun X, Jiang L. Identification of differentially expressed circular RNAs in human colorectal cancer. Tumour Biol. 2017; 39:1010428317694546. https://doi.org/10.1177/1010428317694546.
65. Xiong W, Ai YQ, Li YF, Ye Q, Chen ZT, Qin JY, Liu QY, Wang H, Ju YH, Li WH, Li YF. Microarray analysis of circular RNA expression profile associated with 5-fluorouracil-based chemoradiation resistance in colorectal cancer cells. Biomed Res Int. 2017; 2017:8421614. https://doi.org/10.1155/2017/8421614.
66. Zhu M, Xu Y, Chen Y, Yan F. Circular BANP, an upregulated circular RNA that modulates cell proliferation in colorectal cancer. Biomed Pharmacother. 2017; 88:138–44. https://doi.org/10.1016/j.biopha.2016.12.097.
67. Guo JN, Li J, Zhu CL, Feng WT, Shao JX, Wan L, Huang MD, He JD. Comprehensive profile of differentially expressed circular RNAs reveals that hsa_circ_0000069 is upregulated and promotes cell proliferation, migration, and invasion in colorectal cancer. Onco Targets Ther. 2016; 9:7451–8. https://doi.org/10.2147/OTT.S123220.
68. Hsiao KY, Lin YC, Gupta SK, Chang N, Yen L, Sun HS, Tsai SJ. Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 2017; 77:2339–50. https://doi.org/10.1158/0008-5472.CAN-16-1883.
69. Xie H, Ren X, Xin S, Lan X, Lu G, Lin Y, Yang S, Zeng Z, Liao W, Ding YQ, Liang L. Emerging roles of circRNA_001569 targeting miR-145 in the proliferation and invasion of colorectal cancer. Oncotarget. 2016; 7:26680–91. https://doi.org/10.18632/oncotarget.8589.
70. Zhang XL, Xu LL, Wang F. Hsa_circ_0020397 regulates colorectal cancer cell viability, apoptosis, and invasion by promoting the expression of the miR-138 targets TERT and PD-L1. Cell Biol Int. 2017. https://doi.org/10.1002/cbin.10826.
71. Wang X, Zhang Y, Huang L, Zhang J, Pan F, Li B, Yan Y, Jia B, Liu H, Li S, Zheng W. Decreased expression of hsa_circ_001988 in colorectal cancer and its clinical significances. Int J Clin Exp Pathol. 2015; 8:16020–5.
72. Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, Mesteri I, Grunt TW, Zeillinger R, Pils D. Correlation of circular RNA abundance with proliferation--exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015; 5:8057. https://doi.org/10.1038/srep08057.
73. Dou Y, Cha DJ, Franklin JL, Higginbotham JN, Jeppesen DK, Weaver AM, Prasad N, Levy S, Coffey RJ, Patton JG, Zhang B. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci Rep. 2016; 6:37982. https://doi.org/10.1038/srep37982.
74. Zhang M, Du X. Noncoding RNAs in gastric cancer: research progress and prospects. World J Gastroenterol. 2016; 22:6610–8. https://doi.org/10.3748/wjg.v22.i29.6610.
75. Shao Y, Li J, Lu R, Li T, Yang Y, Xiao B, Guo J. Global circular RNA expression profile of human gastric cancer and its clinical significance. Cancer Med. 2017. https://doi.org/10.1002/cam4.1055.
76. Sui W, Shi Z, Xue W, Ou M, Zhu Y, Chen J, Lin H, Liu F, Dai Y. Circular RNA and gene expression profiles in gastric cancer based on microarray chip technology. Oncol Rep. 2017; 37:1804–14. https://doi.org/10.3892/or.2017.5415.
77. Huang YS, Jie N, Zou KJ, Weng Y. Expression profile of circular RNAs in human gastric cancer tissues. Mol Med Rep. 2017; 16:2469–76. https://doi.org/10.3892/mmr.2017.6916.
78. Chen J, Li Y, Zheng Q, Bao C, He J, Chen B, Lyu D, Zheng B, Xu Y, Long Z, Zhou Y, Zhu H, Wang Y, et al. Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett. 2017; 388:208–19. https://doi.org/10.1016/j.canlet.2016.12.006.
79. Li P, Chen H, Chen S, Mo X, Li T, Xiao B, Yu R, Guo J. Circular RNA 0000096 affects cell growth and migration in gastric cancer. Br J Cancer. 2017; 116:626–33. https://doi.org/10.1038/bjc.2016.451.
80. Zhang Y, Liu H, Li W, Yu J, Li J, Shen Z, Ye G, Qi X, Li G. CircRNA_100269 is downregulated in gastric cancer and suppresses tumor cell growth by targeting miR-630. Aging (Albany NY). 2017; 9:1585–94. https://doi.org/10.18632/aging.101254.
81. Shao Y, Chen L, Lu R, Zhang X, Xiao B, Ye G, Guo J. Decreased expression of hsa_circ_0001895 in human gastric cancer and its clinical significances. Tumour Biol. 2017; 39:1010428317699125. https://doi.org/10.1177/1010428317699125.
82. Tian M, Chen R, Li T, Xiao B. Reduced expression of circRNA hsa_circ_0003159 in gastric cancer and its clinical significance. J Clin Lab Anal. 2017. https://doi.org/10.1002/jcla.22281.
83. Li P, Chen S, Chen H, Mo X, Li T, Shao Y, Xiao B, Guo J. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta. 2015; 444:132–6. https://doi.org/10.1016/j.cca.2015.02.018.
84. Li WH, Song YC, Zhang H, Zhou ZJ, Xie X, Zeng QN, Guo K, Wang T, Xia P, Chang DM. Decreased expression of hsa_circ_00001649 in gastric cancer and its clinical significance. Dis Markers. 2017; 2017:4587698. https://doi.org/10.1155/2017/4587698.
85. Zhang Y, Li J, Yu J, Liu H, Shen Z, Ye G, Mou T, Qi X, Li G. Circular RNAs signature predicts the early recurrence of stage III gastric cancer after radical surgery. Oncotarget. 2017; 8:22936–43. https://doi.org/10.18632/oncotarget.15288.
86. Bonizzato A, Gaffo E, Te Kronnie G, Bortoluzzi S. CircRNAs in hematopoiesis and hematological malignancies. Blood Cancer J. 2016; 6:e483. https://doi.org/10.1038/bcj.2016.81.
87. Guarnerio J, Bezzi M, Jeong JC, Paffenholz SV, Berry K, Naldini MM, Lo-Coco F, Tay Y, Beck AH, Pandolfi PP. Oncogenic role of fusion-circRNAs derived from cancer-associated chromosomal translocations. Cell. 2016; 165:289–302. https://doi.org/10.1016/j.cell.2016.03.020.
88. You X, Conrad TO. Acfs: accurate circRNA identification and quantification from RNA-Seq data. Sci Rep. 2016; 6:38820. https://doi.org/10.1038/srep38820.
89. Li W, Zhong C, Jiao J, Li P, Cui B, Ji C, Ma D. Characterization of hsa_circ_0004277 as a new biomarker for acute myeloid leukemia via circular RNA profile and bioinformatics analysis. Int J Mol Sci. 2017; 18. https://doi.org/10.3390/ijms18030597.
90. Huang M, Zhong Z, Lv M, Shu J, Tian Q, Chen J. Comprehensive analysis of differentially expressed profiles of lncRNAs and circRNAs with associated co-expression and ceRNA networks in bladder carcinoma. Oncotarget. 2016; 7:47186–200. https://doi.org/10.18632/oncotarget.9706.
91. Zhong Z, Huang M, Lv M, He Y, Duan C, Zhang L, Chen J. Circular RNA MYLK as a competing endogenous RNA promotes bladder cancer progression through modulating VEGFA/VEGFR2 signaling pathway. Cancer Lett. 2017; 403:305–17. https://doi.org/10.1016/j.canlet.2017.06.027.
92. Zhong Z, Lv M, Chen J. Screening differential circular RNA expression profiles reveals the regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in bladder carcinoma. Sci Rep. 2016; 6:30919. https://doi.org/10.1038/srep30919.
93. Su H, Lin F, Deng X, Shen L, Fang Y, Fei Z, Zhao L, Zhang X, Pan H, Xie D, Jin X, Xie C. Profiling and bioinformatics analyses reveal differential circular RNA expression in radioresistant esophageal cancer cells. J Transl Med. 2016; 14:225. https://doi.org/10.1186/s12967-016-0977-7.
94. Xia W, Qiu M, Chen R, Wang S, Leng X, Wang J, Xu Y, Hu J, Dong G, Xu PL, Yin R. Circular RNA has_circ_0067934 is upregulated in esophageal squamous cell carcinoma and promoted proliferation. Sci Rep. 2016; 6:35576. https://doi.org/10.1038/srep35576.
95. Zhang C, Wu H, Wang Y, Zhao Y, Fang X, Chen C, Chen H. Expression patterns of circular RNAs from primary kinase transcripts in the mammary glands of lactating rats. J Breast Cancer. 2015; 18:235–41. https://doi.org/10.4048/jbc.2015.18.3.235.
96. Greenlee H, DuPont-Reyes MJ, Balneaves LG, Carlson LE, Cohen MR, Deng G, Johnson JA, Mumber M, Seely D, Zick SM, Boyce LM, Tripathy D. Clinical practice guidelines on the evidence-based use of integrative therapies during and after breast cancer treatment. CA Cancer J Clin. 2017; 67:194–232. https://doi.org/10.3322/caac.21397.
97. Lu L, Sun J, Shi P, Kong W, Xu K, He B, Zhang S, Wang J. Identification of circular RNAs as a promising new class of diagnostic biomarkers for human breast cancer. Oncotarget. 2017; 8:44096–107. https://doi.org/10.18632/oncotarget.17307.
98. Liang G, Liu Z, Tan L, Su AN, Jiang WG, Gong C. HIF1alpha-associated circDENND4C promotes proliferation of breast cancer cells in hypoxic environment. Anticancer Res. 2017; 37:4337–43.
99. Liang HF, Zhang XZ, Liu BG, Jia GT, Li WL. Circular RNA circ-ABCB10 promotes breast cancer proliferation and progression through sponging miR-1271. Am J Cancer Res. 2017; 7:1566–76.
100. Yang Q, Du WW, Wu N, Yang W, Awan FM, Fang L, Ma J, Li X, Zeng Y, Yang Z, Dong J, Khorshidi A, Yang BB. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 2017. https://doi.org/10.1038/cdd.2017.86.
101. Li H, Hao X, Wang H, Liu Z, He Y, Pu M, Zhang H, Yu H, Duan J, Qu S. Circular RNA expression profile of pancreatic ductal adenocarcinoma revealed by microarray. Cell Physiol Biochem. 2016; 40:1334–44. https://doi.org/10.1159/000453186.
102. Qu S, Song W, Yang X, Wang J, Zhang R, Zhang Z, Zhang H, Li H. Microarray expression profile of circular RNAs in human pancreatic ductal adenocarcinoma. Genom Data. 2015; 5:385–7. https://doi.org/10.1016/j.gdata.2015.07.017.
103. Nair AA, Niu N, Tang X, Thompson KJ, Wang L, Kocher JP, Subramanian S, Kalari KR. Circular RNAs and their associations with breast cancer subtypes. Oncotarget. 2016; 7:80967–79. https://doi.org/10.18632/oncotarget.13134.
104. Glazar P, Papavasileiou P, Rajewsky N. circBase: a database for circular RNAs. RNA. 2014; 20:1666–70. https://doi.org/10.1261/rna.043687.113.
105. Ghosal S, Das S, Sen R, Basak P, Chakrabarti J. Circ2Traits: a comprehensive database for circular RNA potentially associated with disease and traits. Front Genet. 2013; 4:283. https://doi.org/10.3389/fgene.2013.00283.
106. Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014; 42:D92–7. https://doi.org/10.1093/nar/gkt1248.
107. Chen L, Yu Y, Zhang X, Liu C, Ye C, Fan L. PcircRNA_finder: a software for circRNA prediction in plants. Bioinformatics. 2016; 32:3528–9. https://doi.org/10.1093/bioinformatics/btw496.
108. Dudekula DB, Panda AC, Grammatikakis I, De S, Abdelmohsen K, Gorospe M. CircInteractome: a web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol. 2016; 13:34–42. https://doi.org/10.1080/15476286.2015.1128065.
109. Zheng LL, Li JH, Wu J, Sun WJ, Liu S, Wang ZL, Zhou H, Yang JH, Qu LH. deepBase v2.0: identification, expression, evolution and function of small RNAs, LncRNAs and circular RNAs from deep-sequencing data. Nucleic Acids Res. 2016; 44:D196–202. https://doi.org/10.1093/nar/gkv1273.
110. Gao Y, Zhang J, Zhao F. Circular RNA identification based on multiple seed matching. Brief Bioinform. 2017. https://doi.org/10.1093/bib/bbx014.
111. Szabo L, Salzman J. Detecting circular RNAs: bioinformatic and experimental challenges. Nat Rev Genet. 2016; 17:679–92. https://doi.org/10.1038/nrg.2016.114.
112. Meng S, Zhou H, Feng Z, Xu Z, Tang Y, Li P, Wu M. CircRNA: functions and properties of a novel potential biomarker for cancer. Mol Cancer. 2017; 16:94. https://doi.org/10.1186/s12943-017-0663-2.
113. Xu Z, Yan Y, Li Z, Qian L, Gong Z. The antibiotic drug tigecycline: a focus on its promising anticancer properties. Front Pharmacol. 2016; 7:473. https://doi.org/10.3389/fphar.2016.00473.
114. Salzman J. Circular RNA expression: its potential regulation and function. Trends Genet. 2016; 32:309–16. https://doi.org/10.1016/j.tig.2016.03.002.
115. Syn NL, Wang L, Chow EK, Lim CT, Goh BC. Exosomes in cancer nanomedicine and immunotherapy: prospects and challenges. Trends Biotechnol. 2017. https://doi.org/10.1016/j.tibtech.2017.03.004.
116. Wang YH, Yu XH, Luo SS, Han H. Comprehensive circular RNA profiling reveals that circular RNA100783 is involved in chronic CD28-associated CD8(+) T cell ageing. Immun Ageing. 2015; 12:17. https://doi.org/10.1186/s12979-015-0042-z.
117. Ng WL, Marinov GK, Liau ES, Lam YL, Lim YY, Ea CK. Inducible RasGEF1B circular RNA is a positive regulator of ICAM-1 in the TLR4/LPS pathway. RNA Biol. 2016; 13:861–71. https://doi.org/10.1080/15476286.2016.1207036.
118. Wang RF, Miyahara Y, Wang HY. Toll-like receptors and immune regulation: implications for cancer therapy. Oncogene. 2008; 27:181–9. https://doi.org/10.1038/sj.onc.1210906.
119. Li Y, Zheng F, Xiao X, Xie F, Tao D, Huang C, Liu D, Wang M, Wang L, Zeng F, Jiang G. CircHIPK3 sponges miR-558 to suppress heparanase expression in bladder cancer cells. EMBO Rep. 2017; 18:1646–59. https://doi.org/10.15252/embr.201643581.
120. Liao JY, Wu J, Wang YJ, He JH, Deng WX, Hu K, Zhang YC, Zhang Y, Yan H, Wang DL, Liu Q, Zeng MS, Phillip Koeffler H, et al. Deep sequencing reveals a global reprogramming of lncRNA transcriptome during EMT. Biochim Biophys Acta. 2017; 1864:1703–13. https://doi.org/10.1016/j.bbamcr.2017.06.003.
121. Zhang J, Liu H, Hou L, Wang G, Zhang R, Huang Y, Chen X, Zhu J. Circular RNA_LARP4 inhibits cell proliferation and invasion of gastric cancer by sponging miR-424-5p and regulating LATS1 expression. Mol Cancer. 2017; 16:151. https://doi.org/10.1186/s12943-017-0719-3.
122. Fang Z, Yin S, Sun R, Zhang S, Fu M, Wu Y, Zhang T, Khaliq J, Li Y. miR-140-5p suppresses the proliferation, migration and invasion of gastric cancer by regulating YES1. Mol Cancer. 2017; 16:139. https://doi.org/10.1186/s12943-017-0708-6.
123. Yang Z, Xie L, Han L, Qu X, Yang Y, Zhang Y, He Z, Wang Y, Li J. Circular RNAs: regulators of cancer-related signaling pathways and potential diagnostic biomarkers for human cancers. Theranostics. 2017; 7:3106–17. https://doi.org/10.7150/thno.19016.
124. Han YN, Xia SQ, Zhang YY, Zheng JH, Li W. Circular RNAs: a novel type of biomarker and genetic tools in cancer. Oncotarget. 2017; 8:64551–63. https://doi.org/10.18632/oncotarget.18350.
