INTRODUCTION
Breast cancer is the most frequently diagnosed female tumor type, and its incidence continues to rise [1, 2]. Despite the development in early clinical diagnosis and treatment strategies, breast cancer remains a significant global burden [3, 4]. Detection of breast cancer at an early stage is a critical step for successful treatment and improvement of prognosis. Therefore, to identify sensitive and specific biomarkers for prognosis of breast cancer patients has important clinical implications.
Syndecan family is a new found evolutionarily conserved transmembrane proteoglycan which can regulate cell proliferation, migration, adhesion, angiogenesis, and modulate cell-cell and cell-matrix interactions during wound reparation [5, 6]. Syndecan-1 (SDC1) is an important member of syndecan family, which expression has been associated with prognosis and treatment response in a various tumor types, including solid tumors and hematolymphoid malignancies [7–14]. SDC1 has also been demonstrated to be enhance endometrial cancer invasion by modulating matrix metalloproteinase-9 (MMP-9) expression through nuclear factor kappaB (NF-κb) [15]. In addition, the proliferation and invasion of glioma cell has been attenuated through knockdown SDC1 [5]. Moreover, syndecan-1 has been identified at high levels in a significant percentage of breast carcinomas and related to an aggressive phenotype and poor clinical behavior [16]. Together, SDC1 may not only act as a potential therapeutic target, but also as a novel prognostic biomarker in cancer. However, the role of SDC1 in breast cancer and the underlying molecular mechanism responsible for its involvement in the generation and development of tumor are still unclear. The clinicopathological and prognostic value of SDC1 expression patterns in breast patients remains controversial [17, 18].
In this study, we evaluated the significance of SDC1 mRNA in human breast cancer using Oncomine microarray datasets. We further analyzed the relationship between SDC1 expression and clinicopathologic parameters including prognostic significance, and explored the biological function and molecular mechanism of SDC1 in breast cancer patients by pooling all currently available data.
RESULTS
SDC1 transcript expression and methylation status in human breast cancer.
As a cell surface heparin sulfate proteoglycan (HSPG), the expression of profile of SDC1 was analyzed though using Oncomine database. SDC1 has been identified in human cancers, including hematological malignancies and solid tumors (Figure 1). Oncomine analysis of tumor versus. normal samples revealed that SDC1 mRNA was significantly elevated in invasive breast carcinoma, invasive ductal breast carcinoma, mixed lobular and ductal breast carcinoma, invasive lobular breast carcinoma (Table 1, Figure 2) [19–26]. Through comparing SDC1 expression heat map and its DNA methylation status, we observed that its expression might be negatively related with some CpG sites (blank frame) (Figure 3)
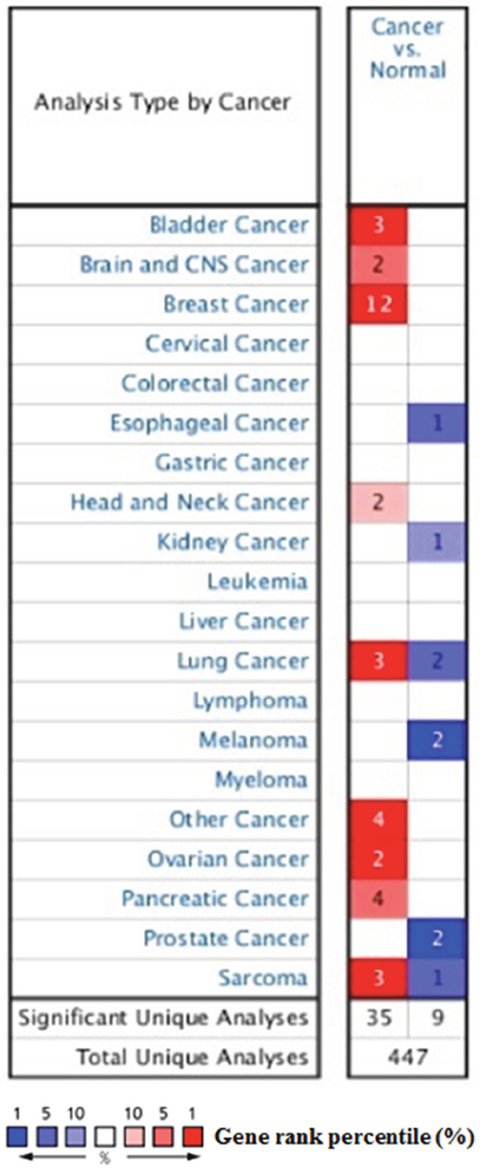
Figure 1: The transcription levels of SDC1 in different types of cancers. This graphic was generated from Oncomine, indicating the numbers of datasets with statistically significant (p<0.01) mRNA over-expression (Red) or down-expression (Blue) of SDC1 (different types of cancer vs. corresponding normal tissue). The threshold was designed with following parameters: p-value of 1E-4, fold change of 2, and gene ranking of 10%.
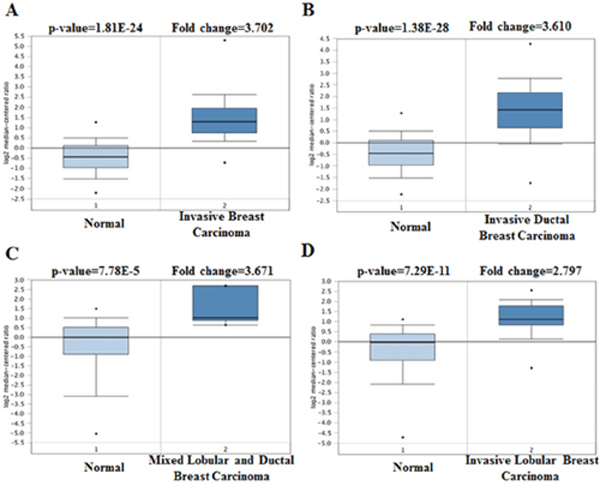
Figure 2: SDC1 analysis in breast cancer (Oncomine database). The box plot comparing specific SDC1 expression in normal (left plot) and cancer tissue (right plot) was derived from Oncomine database. The analysis was shown in invasive breast carcinoma, invasive ductal breast carcinoma, mixed lobular and ductal breast carcinoma, and invasive lobular breast carcinoma relative to normal breast.
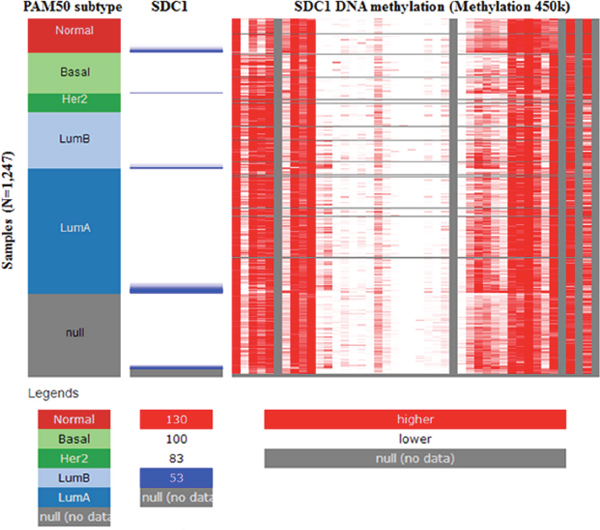
Figure 3: The heat map of SDC1 expression and its DNA methylation status.
Table 1: The significant changes of SDC1 expression in transcription level between different types of breast cancer and normal tissues (ONCOMINE database)
Subtype of breast cancer |
p-value |
Fold change |
Rank (%) |
Sample |
Reference |
|---|---|---|---|---|---|
Ductal Breast Carcinoma |
7.15E-7 |
2.567 |
1 |
69 |
[29] |
Ductal Breast Carcinoma |
3.62E-6 |
2.317 |
2 |
39 |
[30] |
Ductal Breast Carcinoma |
1.17E-5 |
2.754 |
2 |
94 |
[31] |
Invasive Ductal Breast Carcinoma |
4.86E-5 |
5.223 |
1 |
22 |
[32] |
Invasive Ductal Breast Carcinoma |
4.58E-5 |
3.165 |
1 |
23 |
[33] |
Invasive Breast Carcinoma |
1.81E-24 |
3.702 |
2 |
137 |
TCGA |
Invasive Ductal Breast Carcinoma |
1.38E-28 |
3.610 |
4 |
450 |
TCGA |
Mixed Lobular and Ductal Breast Carcinoma |
7.78E-5 |
3.671 |
4 |
68 |
TCGA |
Invasive Lobular Breast Carcinoma |
7.29E-11 |
2.797 |
5 |
97 |
TCGA |
Tubular Breast Carcinoma |
1.25E-26 |
3.407 |
2 |
211 |
[34] |
Ductal Breast Carcinoma |
4.38E-7 |
2.549 |
3 |
47 |
[35] |
Invasive Breast Carcinoma |
1.65E-17 |
2.245 |
3 |
59 |
[36] |
SDC1 mutation in human breast cancer
The pie chart that included the information of mutations of substitution nonsense, missense, synonymous, insertion frame shift, and inframe deletion was generated by performing COSMIC. Substitution missense rate is 51.85% and substitution synonymous rate is 37.04% of mutant samples of breast cancer (Figure 4A). Breast cancer has 35.62% C > T and 20.55% G > A mutation in SDC1 coding strand (Figure 4A). Alteration frequency of SDC1 mutation in breast cancer was identified by performing cBioPortal. Less than 1.6% mutation in the patients with breast cancer was observed (Figure 4B)
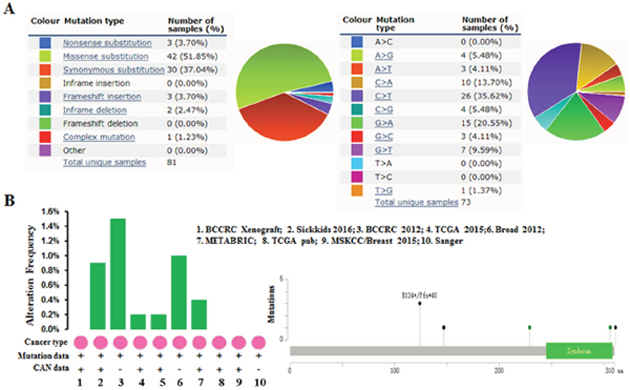
Figure 4: SDC1 mutation in human breast cancer. (A) Pie-chart showed the percentage of the mutation type of SDC1 in breast cancer according to COSMIC database. (B) Alteration frequency of SDC1 mutation in breast cancer was analyzed by using BioPortal.
Genetic alterations of SDC1 and clinicopathological parameters in breast cancer patients
As displayed in Table 2, we performed a welch’s test to compare the transcription levels of SDC1 between groups of patients by bc-GenExMiner 4.0, basing on different clinicopathological parameters. For age criterion, there was no significant difference between ≤ 51 y and > 51 y groups (Table 2). Breast cancer patients with positive nodal status displayed elevated SDC1 mRNA than negative-nodal patients (Table 2, Figure 5). Estrogen receptor (ER) status was confirmed to be negatively related with SDC1 expression. Conversely, in breast cancer patients with higher epidermal growth factor receptor 2 (HER2), the mRNA levels of SDC1 was significantly increased compared with HER2-negative groups. However, there was no significant difference between positive progesterone receptor (PR) and negative PR groups. Meanwhile, the basal-like and triple-negative breast cancer (TNBC) characteristics also has no difference between positive and negative status (Table 2). In Scarff Bloom & Richardson grade status (SBR) criterion, more advanced SBR grade was associated with the higher mRNA level of SDC1 (Figure 5).
Table 2: The relationship between mRNA expression of SDC1 and clinicopathological parameters of breast carcinoma
Variables |
SDC1 |
||
|---|---|---|---|
No.* |
mRNA |
p-value |
|
Age |
|||
≤ 51 |
1392 |
- |
0.0135 |
> 51 |
2210 |
↑ |
|
Nodal Statas |
|||
- |
2493 |
- |
0.0045 |
+ |
1562 |
↑ |
|
ER |
|||
- |
1559 |
- |
<0.0001 |
+ |
3988 |
↓ |
|
PR |
|||
- |
946 |
- |
0.8436 |
+ |
1439 |
- |
|
HER2 |
|||
- |
1409 |
- |
<0.0001 |
+ |
201 |
↑ |
|
Basal-like Status |
|||
Not |
4205 |
- |
0.6699 |
Basal-like |
1144 |
- |
|
Triple-negative Status |
|||
Not |
4099 |
- |
0.0681 |
TNBC |
374 |
- |
|
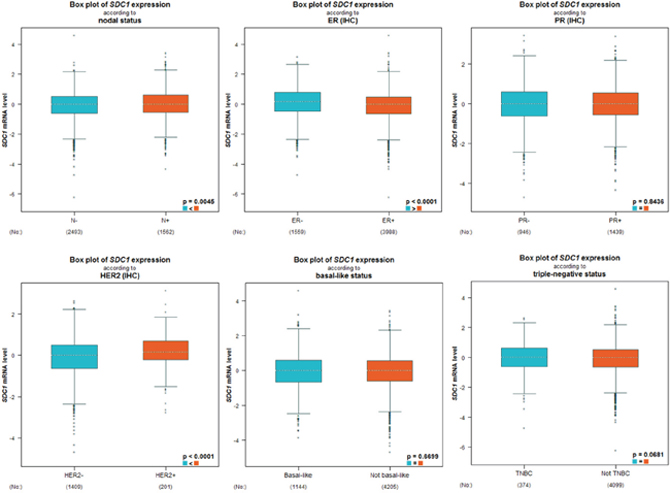
Figure 5: The relationship between mRNA expression of SDC1 and clinicopathological parameters in breast cancer patients. Global signifcant different between groups was assessed by Welch’s test to generate p value, along with Dunnett-Tukey-Kramer’s.
Association of SDC1 expression and prognosis in breast cancer patients
Kaplan-Meir analysis revealed that high SDC1 mRNA was correlated with a lower overall survival (OS), relapse-free survival (RFS), distance metastasis free survival (DMFS) in breast cancer (Figure 6). By performing data mining in bc-GenExMiner, we pooled previous available annotated genomic data to analysis the relationship between SDC1 expression and prognosis metastatic relapse-free survival (MRFS) in breast cancer. High CD24 expression was associated with higher risk of metastatic relapse (HR =1.15, 95% CI 1.08 - 1.23, p < 0.0001) (Figure 7A). Similarly, high SDC1 expression was also associated with worse MRFS (HR =1.20, 95% CI 1.06–1.36, p =0.0038) (Figure 7B).
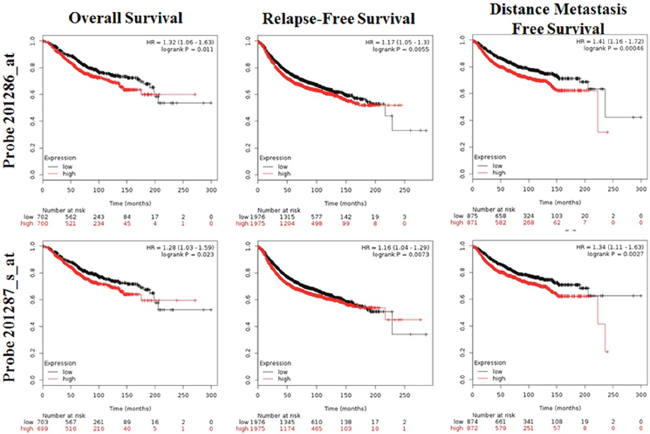
Figure 6: The prognostic value of mRNA level of SDC1 in breast cancer patients (OS, RFS, DMFS in Kaplan-Meier plotter).
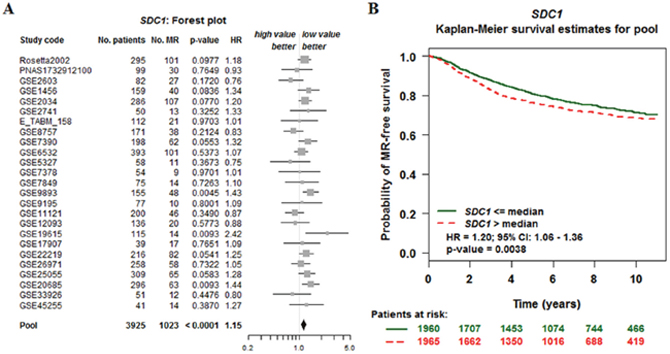
Figure 7: The relationship between SDC1 expression and prognosis metastatic relapse-free survival (MRFS) in breast cancer (bc-GenExMiner v4. 0). (A) Forest plots displaying univariate Cox’s analysis of SDC1 expression and the risk of metastatic relapse (MR). (B) Kaplan–Meier survival analysis showing the relationship between SDC1 expression and MR-free survival.
Co-expression of SDC1 mRNA
To further explore the potential regulation of SDC1 in breast cancer, we performed data mining in breast cancer cohort using cBioPortal. PLAU (urokinase plasminogen activator gene, uPA gene) is a top correlated gene (Figure 8A), which can enhance multiple oncogene expression in breast cancer [27, 28]. Regression analysis showed that SDC1 and PLAU had high correlation coefficients (Pearson’s correlation = 0.59; Spearman’s correlation = 0.58) (Figure 8B). Data mining in bc-GenExMiner 4.0 confirmed the positive correlation between SDC1 and PLAU mRNA (Figure 8C). By analyzing breast cancer patient data in TCGA database using UCSC Xena (http://xena.ucsc.edu/), we also confirmed the positive correlation (Figure 8D-8E). These findings suggest that SDC1 might be closely related to the PLAU signaling pathways in breast cancer.
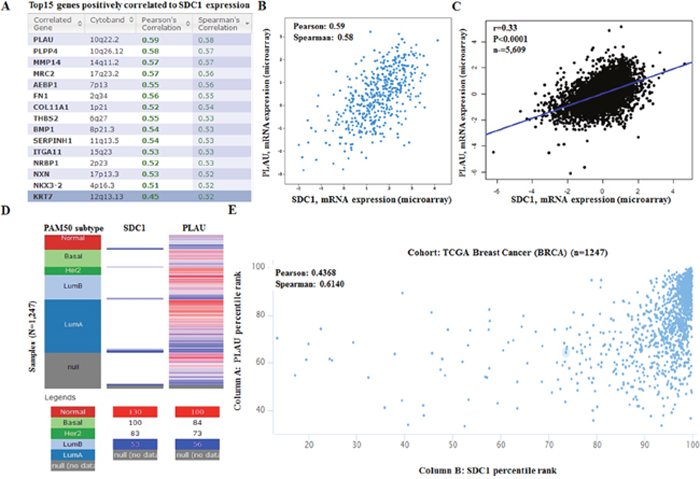
Figure 8: SDC1 mRNA expression is correlated to PLAU mRNA expression in breast cancer. (A) Top 20 genes positively correlated with SDC1 mRNA expression based on 482 breast cancer patients in TCGA. Analysis was performed using cBioportal (http://cbioportal.org). (B) Regression analysis showed that SDC1 and PLAU had high correlation coefficients. (C) Data mining in bc-GenExMiner 4.0 confirmed the positive correlation between SDC1 and PLAU mRNA. (D) The heat map of SDC1 mRNA and PLAU mRNA expression across PAM50 breast cancer subtypes in TCGA database. Data was analyzed using UCSC Xena (http://xena.ucsc.edu/). (E) The correlation between SOX10 and NES mRNA expression in TCGA database.
DISCUSSION
SDC1 is a heparin cell surface proteoglycan acting as a co-receptor for growth factors and chemokines, which strongly associated with the tumor aggressiveness and clinical outcomes [5, 8, 29, 30]. Previous studies have revealed that SDC1 has prognostic value in various types of cancers such as hepatocellular cancer [31], gastric cancer [32], lung cancer [33], and colorectal cancer [7]. In particular, higher expression of SDC1 has been correlated with poorer prognosis for breast cancer patients and correlated with more malignant and higher grade breast cancer tissues [34]. Meanwhile, knockdown of SDC1 expression can inhibits glioma proliferation and invasion through deregulating the c-src/FAK-associated signaling pathway [5]. However, the distinct role of SDC1 expression in the development and metastasis of breast cancer is still unclear.
In the present study, we analyzed the data based on amounts of gene expression with clearly defined parameters between cancer and normal tissues. Oncomine analysis of cancer vs. normal tissue revealed that SDC1 mRNA was significantly over-expressed in invasive breast carcinoma, invasive ductal breast carcinoma, mixed lobular and ductal breast carcinoma, invasive lobular breast carcinoma. Meanwhile, high SDC1 expression is associated with increased risked of age, nodal, HER2 and higher SBR grade status, which predicted fast-growing, spreading tumors. We further analyzed the mutations and alteration frequency of SDC1 using COSMIC and cBioPortal database. Our analysis suggested that the major proportion of mutation in breast cancer is synonymous mutations. Meanwhile, the low alteration frequency was observed in breast cancer. Then, we tried to investigate the mechanisms of SDC1 dysregulation. Through examining its DNA methylation status in TCGA-BRCA, we observed a negative correlation between methylation status of some CpG sites and SDC1 expression. These findings suggest that DNA methylation might be an important mechanism of SDC1 dysregulation in breast cancer.
Survival analysis indicated SDC1 high expression was significantly associated with poor OS, RFS and DMFS. In addition, the following data mining and meta-analysis suggested that high SDC1 expression also associated with increased risk of MR and indicates significantly worse MR-free survival among patients with breast cancer. Depend on these findings, we demonstrated that the expression level of SDC1 might be a useful marker of the prognosis of breast cancer.
Through co-expression and correlation analysis, we observed that PLAU was positively correlated with the expression of SDC1. In fact, as a component of complement and coagulation cascades, PLAU (uPA) can also play a variety of functions in a non-proteolytic fashion, such as cellular adhesion, differentiation, proliferation and migration, via signals triggered by its receptor PLAUR (uPAR) [35, 36]. Dawei Li et al. found that aberrant FOXM1-PLAUR signaling plays an critical role in progression and metastasis of colon cancer [35]. Furthermore, Overexpression of PLAU can largely rescue self-renewal, migration, invasion and vascular formation defects of repressed glioma cells [36]. These results indicated that SDC1 expression might regulated tumor migration and invasion through promoting PLAU expression.
MATERIALS AND METHODS
Oncomine database analysis
Oncomine gene expression array datasets (https://www.oncomine.org/resource/login.html), a publicly accessible online cancer microarray database, was performed to determine the mRNA levels of distinct SDC1 in breast cancer [37]. In this study, we compared the clinical specimens of cancer vs. normal control datasets, using the Student’s t–test to generate a p-value. The fold change was defined as 2 and p value was set up at 1E-4, whereas the data type was restricted to mRNA.
COSMIC analysis for SDC1 mutations
The Catalog of Somatic Mutations in Cancer (COSMIC) database (http://www.sange r.ac.uk/cosmic/), the world’s largest and most comprehensive resource for exploring the impact of somatic mutations in human cancer, was performed to analysis the mutations of SDC1. Pie charts were generated for a distribution overview and substitutions on the coding strand in breast cacner.
UCSC cancer genomics browser analysis
The heat map of SDC1 expression and methylation status and the correlation between SDC1 and PLAU expression were identified using data in TCGA breast cancer cohort (TCGA-BRCA) by performing the UCSC (University of California at Santa Cruz) Cancer Genomics Browser (http://xena.ucsc.edu/) [38–40].
cBioPortal database analysis
The cBioPortal for Cancer genomics (http://www.cbioportal.org/) is affiliated with Memorial Sloan Kettering Cancer Center and provides integrative analysis of complex cancer genomics and clinical profiles from 105 cancer studies in TCGA pipeline [41, 42]. The term “SDC1” was searched in cBioPortal database and a cross-cancer summary was obtained for it. The search parameters included alterations (amplification, deep deletion, missense mutations), copy-number variance (CNV) from GISTIC and RNA seq data with the default setting. In addition, cBioPortal were performed to identified the positive correlative genes with SDC1 expression in breast cancer.
Breast cancer gene-expression miner v4.0
Breast Cancer Gene-Expression Miner v4.0 (bc-GenExMiner v4.0), a statistical mining tool of published annotated genomic data including 36 annotated genomic datasets and 5861 breast cancer patients, was used to assess the expression of SDC1 in breast cancer, as well as the association between SDC1 mRNA expression and prognosis in breast cancer patients [43, 44]. Furthermore, the correlation of SDC1 and PLAU were assessed by correlation module of bc-GenExMiner v4.0.
Kaplan-Meier plotter database analysis
Kaplan-Meier Plotter (http://kmplot.com/analysis/), an online database including gene expression data and clinical data, was performed to analysis the prognostic significance of the mRNA expression of SDC1 in breast cancer [45 46]. Log rank p-value and hazard ratio (HR) with 95% confidence intervals (CI) were calculated and displayed on the webpage.
CONCLUSIONS
In summary, overexpressed SDC1 was identified in breast cancer than in matched normal tissues and is associated with methylation status of SDC1 promoter. Additionally, SDC1 is positively associated with PLAU and might act as a potential prognostic indicator for breast cancer. However, future research is required to validate our findings and thus promote the clinical utility of SDC1 in breast cancer prognosis evaluation.
Author contributions
XRC and XJ carried out the data gathering, data analyses, and figure/table preparations. QY, CLL, BT, XL, XC, YH, ZPX, and JT provided material input and helped revising the manuscript. JZ supervised the experimental design, data analyses, and manuscript writing. All authors read and approved the final manuscript.
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
Availability of data and materials
All data generated or analyzed during this study are included in this published article (and its additional information files).
Consent for publication
All the authors in this paper consent to publication of the work.
Ethics approval and consent to participate
Not applicable.
FUNDING
This work is partially supported a grant from the National Natural Science Foundation of China (No.81670270) to J. Zhu and the Scientific Research Project of Shanxi Provincial Department of health [201601070] to X.R. Cui.
REFERENCES
1. Xu H, Yu S, Liu Q, Yuan X, Mani S, Pestell RG, Wu K. Recent advances of highly selective CDK4/6 inhibitors in breast cancer. J Hematol Oncol. 2017; 10:97.
2. Stellrecht CM, Vangapandu HV, Le XF, Mao W, Shentu S. ATP directed agent, 8-chloro-adenosine, induces AMP activated protein kinase activity, leading to autophagic cell death in breast cancer cells. J Hematol Oncol. 2014; 7:23.
3. Shan M, Yin H, Li J, Li X, Wang D, Su Y, Niu M, Zhong Z, Wang J, Zhang X, Kang W, Pang D. Detection of aberrant methylation of a six-gene panel in serum DNA for diagnosis of breast cancer. Oncotarget. 2016; 7:18485–18494. https://doi.org/10.18632/oncotarget.7608.
4. Baldwin RM, Haghandish N, Daneshmand M, Amin S, Paris G, Falls TJ, Bell JC, Islam S, Cote J. Protein arginine methyltransferase 7 promotes breast cancer cell invasion through the induction of MMP9 expression. Oncotarget. 2015; 6:3013–3032. https://doi.org/10.18632/oncotarget.3072.
5. Shi S, Zhong D, Xiao Y, Wang B, Wang W, Zhang F, Huang H. Syndecan-1 knockdown inhibits glioma cell proliferation and invasion by deregulating a c-src/FAK-associated signaling pathway. Oncotarget. 2017; 8:40922–40934. https://doi.org/10.18632/oncotarget.16733.
6. Binder Gallimidi A, Nussbaum G, Hermano E, Weizman B, Meirovitz A, Vlodavsky I, Gotte M, Elkin M. Syndecan-1 deficiency promotes tumor growth in a murine model of colitis-induced colon carcinoma. PLoS One. 2017; 12:e0174343.
7. Jary M, Lecomte T, Bouche O, Kim S, Dobi E, Queiroz L, Ghiringhelli F, Etienne H, Leger J, Godet Y, Balland J, Lakkis Z, Adotevi O, et al. Prognostic value of baseline seric Syndecan-1 in initially unresectable metastatic colorectal cancer patients: a simple biological score. Int J Cancer. 2016; 139:2325–2335.
8. Beauvais DM, Jung O, Yang Y, Sanderson RD, Rapraeger AC. Syndecan-1 (CD138) suppresses apoptosis in multiple myeloma by activating IGF1 receptor: prevention by synstatinIGF1R inhibits tumor growth. Cancer Res. 2016; 76:4981–4993.
9. Goto K. CD138 expression is observed in the urothelial epithelium and in various urothelial carcinomas, and cannot be evidence for plasmacytoid urothelial carcinoma. Int J Surg Pathol. 2016; 24:614–619.
10. Guo T, Yu W, Lv S, Zhang C, Tian Y. MiR-302a inhibits the tumorigenicity of ovarian cancer cells by suppression of SDC1. Int J Clin Exp Pathol. 2015; 8:4869–4880.
11. Gharbaran R. Advances in the molecular functions of syndecan-1 (SDC1/CD138) in the pathogenesis of malignancies. Crit Rev Oncol Hematol. 2015; 94:1–17.
12. Szarvas T, Reis H, Kramer G, Shariat SF, Vom Dorp F, Tschirdewahn S, Schmid KW, Kovalszky I, Rubben H. Enhanced stromal syndecan-1 expression is an independent risk factor for poor survival in bladder cancer. Hum Pathol. 2014; 45:674–682.
13. Shimada K, Anai S, Fujii T, Tanaka N, Fujimoto K, Konishi N. Syndecan-1 (CD138) contributes to prostate cancer progression by stabilizing tumour-initiating cells. J Pathol. 2013; 231:495–504.
14. Seftalioglu A, Karakus S. Syndecan-1/CD138 expression in normal myeloid, acute lymphoblastic and myeloblastic leukemia cells. Acta Histochem. 2003; 105:213–221.
15. Yang N, Mosher R, Seo S, Beebe D, Friedl A. Syndecan-1 in breast cancer stroma fibroblasts regulates extracellular matrix fiber organization and carcinoma cell motility. Am J Pathol. 2011; 178:325–335.
16. Barbareschi M, Maisonneuve P, Aldovini D, Cangi MG, Pecciarini L, Angelo Mauri F, Veronese S, Caffo O, Lucenti A, Palma PD, Galligioni E, Doglioni C. High syndecan-1 expression in breast carcinoma is related to an aggressive phenotype and to poorer prognosis. Cancer. 2003; 98:474–483.
17. Dhodapkar MV, Sanderson RD. Syndecan-1 (CD 138) in myeloma and lymphoid malignancies: a multifunctional regulator of cell behavior within the tumor microenvironment. Leuk Lymphoma. 1999; 34:35–43.
18. Akl MR, Nagpal P, Ayoub NM, Prabhu SA, Gliksman M, Tai B, Hatipoglu A, Goy A, Suh KS. Molecular and clinical profiles of syndecan-1 in solid and hematological cancer for prognosis and precision medicine. Oncotarget. 2015; 6:28693–28715. https://doi.org/10.18632/oncotarget.4981.
19. Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, Barrette T, Pandey A, Chinnaiyan AM. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004; 6:1–6.
20. Zhu J, Sanborn JZ, Benz S, Szeto C, Hsu F, Kuhn RM, Karolchik D, Archie J, Lenburg ME, Esserman LJ, Kent WJ, Haussler D, Wang T. The UCSC cancer genomics browser. Nat Methods. 2009; 6:239–240.
21. Sanborn JZ, Benz SC, Craft B, Szeto C, Kober KM, Meyer L, Vaske CJ, Goldman M, Smith KE, Kuhn RM, Karolchik D, Kent WJ, Stuart JM, et al. The UCSC cancer genomics browser: update 2011. Nucleic Acids Res. 2011; 39:D951–D959.
22. Goldman M, Craft B, Swatloski T, Ellrott K, Cline M, Diekhans M, Ma S, Wilks C, Stuart J, Haussler D, Zhu J. The UCSC cancer genomics browser: update 2013. Nucleic Acids Res. 2013; 41:D949–D954.
23. Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N. Integrative analysis of complex cancer genomics and clinical profiles using the cBioportal. Sci Signal. 2013; 6:pl1.
24. Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012; 2:401–404.
25. Jezequel P, Campone M, Gouraud W, Guerin-Charbonnel C, Leux C, Ricolleau G, Campion L. Bc-GenExMiner: an easy-to-use online platform for gene prognostic analyses in breast cancer. Breast Cancer Res Treat. 2012; 131:765–775.
26. Jezequel P, Frenel JS, Campion L, Guerin-Charbonnel C, Gouraud W, Ricolleau G, Campone M. bc-GenExMiner 3.0: new mining module computes breast cancer gene expression correlation analyses. Database (Oxford). 2013; 2013:bas060.
27. Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010; 123:725–731.
28. Gyorffy B, Surowiak P, Budczies J, Lanczky A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS One. 2013; 8:e82241.
29. Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A. 2001; 98:10869–10874.
30. Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, et al. Molecular portraits of human breast tumours. Nature. 2000; 406:747–752.
31. Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, Demeter J, Perou CM, Lonning PE, et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci U S A. 2003; 100:8418–8423.
32. Karnoub AE, Dash AB, Vo AP, Sullivan A, Brooks MW, Bell GW, Richardson AL, Polyak K, Tubo R, Weinberg RA. Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature. 2007; 449:557–563.
33. Ma XJ, Dahiya S, Richardson E, Erlander M, Sgroi DC. Gene expression profiling of the tumor microenvironment during breast cancer progression. Breast Cancer Res. 2009; 11:R7.
34. Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Graf S, Ha G, Haffari G, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012; 486:346–352.
35. Richardson AL, Wang ZC, De Nicolo A, Lu X, Brown M, Miron A, Liao X, Iglehart JD, Livingston DM, Ganesan S. X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell. 2006; 9:121–132.
36. Finak G, Bertos N, Pepin F, Sadekova S, Souleimanova M, Zhao H, Chen H, Omeroglu G, Meterissian S, Omeroglu A, Hallett M, Park M. Stromal gene expression predicts clinical outcome in breast cancer. Nature Med. 2008; 14:518–527.
37. Viala M, Alexandre M, Thezenas S, Lamy PJ, Maran-Gonzalez A, Gutowski M, Colombo PE, Romieu G, Jacot W, Guiu S. Prognostic impact of the inclusion of uPA/PAI-1 for adjuvant treatment decision-making in ER+/Her2- pN0 early breast cancers. Breast Cancer Res Treat. 2017; 165:611–621.
38. Li H, Chen C. Quercetin has antimetastatic effects on gastric cancer cells via the interruption of uPA/uPAR function by modulating NF-κb, PKC-δ, ERK1/2, and AMPKα. Integr Cancer Ther. 2017.
39. Chen D, Zou J, Zong Y, Meng H, An G, Yang L. Anti-human CD138 monoclonal antibodies and their bispecific formats: generation and characterization. Immunopharmacol Immunotoxicol. 2016; 38:175–183.
40. Liakou E, Mavrogonatou E, Pratsinis H, Rizou S, Evangelou K, Panagiotou PN, Karamanos NK, Gorgoulis VG, Kletsas D. Ionizing radiation-mediated premature senescence and paracrine interactions with cancer cells enhance the expression of syndecan 1 in human breast stromal fibroblasts: the role of TGF-beta. Aging. 2016; 8:1650–1669.
41. Matsumoto A, Ono M, Fujimoto Y, Gallo RL, Bernfield M, Kohgo Y. Reduced expression of syndecan-1 in human hepatocellular carcinoma with high metastatic potential. Int J Cancer. 1997; 74:482–491.
42. Hu XF, Yao J, Gao SG, Yang YT, Peng XQ, Feng XS. Midkine and syndecan1 levels correlate with the progression of malignant gastric cardiac adenocarcinoma. Mol Med Rep. 2014; 10:1409–1415.
43. Al-Shibli K, Al-Saad S, Andersen S, Donnem T, Bremnes RM, Busund LT. The prognostic value of intraepithelial and stromal CD3-, CD117- and CD138-positive cells in non-small cell lung carcinoma. APMIS. 2010; 118:371–382.
44. Lendorf ME, Manon-Jensen T, Kronqvist P, Multhaupt HA, Couchman JR. Syndecan-1 and syndecan-4 are independent indicators in breast carcinoma. J Histochem Cytochem. 2011; 59:615–629.
45. Li D, Wei P, Peng Z, Huang C, Tang H, Jia Z, Cui J, Le X, Huang S, Xie K. The critical role of dysregulated FOXM1-PLAUR signaling in human colon cancer progression and metastasis. Clin Cancer Res. 2013; 19:62–72.
46. Zhong X, Liu X, Li Y, Cheng M, Wang W, Tian K, Mu L, Zeng T, Liu Y, Jiang X, Yu L, Gao L, Zhou Y. HMGA2 sustains self-renewal and invasiveness of glioma-initiating cells. Oncotarget. 2016; 7:44365–44380. https://doi.org/10.18632/oncotarget.9744.

