INTRODUCTION
The identity of a cell is determined by the genes it expresses. Protein-coding genes had been the focus of research in the past, in recent years the importance of non-coding genes has also been reported to differentially express and play a role in processes such as differentiation, normal cell state maintenance and malignancy [1-5]. The hematopoietic system consists of well-characterized cell types from the multipotent hematopoietic stem cells (HSC) through defined progenitors and down to the various effector cells. Studying non-coding genes in normal and perturbed hematopoiesis is of interest in order to reveal new insights into this essential process of blood cell generation, and possibly related malignancies. Specific studies of non-coding genes of interest are needed to further characterize their expression along hematopoiesis.
Nuclear Enriched Abundant Transcript 1 (Neat1) is a long non-coding RNA (lncRNA) that was cloned as polyadenylated nuclear enriched RNA [6]. Neat1 has two isoforms: Neat1_1 and Neat1_2 also known as Multiple Endocrine Neoplasia ε and β (Menε and Menβ, respectively) [1]. These two isoforms are transcribed from a single locus of chromosome 19 in mice or 11 in humans, with a difference in their 3’ ends [6, 7]. Neat1_1 is a 3170 bp polyadenylated transcript, whereas Neat1_2 is a 20,177 bp transcript containing a genomic poly(A)-rich tract on its 3’ end [1, 8-10]. Both isoforms were reported to play an essential role as the core of the sub-nuclear structures called paraspeckles [11]. Intriguingly, it is not yet clear which of the isoforms is more significant in maintaining paraspeckle integrity, due to the fact that they are produced from a single locus. Moreover, it is difficult to stop the transcription of Neat1_1 without affecting Neat1_2, although vice versa might be possible. Thus, both Neat1_1 and Neat1_2 are considered as core-components of the paraspeckles, and are of interest for specific studies on different cell types [1, 11].
Paraspeckles are sub-nuclear structures composed of functionally distinct proteins including PSPC1, SFPQ and NONO (also known as p54nrb), that are in a complex together with Neat1 [8, 12, 13]. Paraspeckles play a role in the regulation of certain genes in differentiated cells by nuclear retention of RNA, controlling gene expression by trapping adenosine to inosine (A to I) hyper-edited RNA within the nucleus [14, 15]. This mechanism of mRNA retention can be used to coordinate gene expression by release upon need, such as stress [12, 15]. Furthermore, a recent study discovered that NONO, PSF and NEAT1 in HeLa cells mediate pri-miRNA processing, with a structural role for NEAT1_2 isoform in recruitment of miRNA microprocessors, highlighting potential role in broad regulation of gene expression [16]. The physiological role of Neat1 is not yet known, as viability under normal conditions was not severely affected in knockout mice [11], while the paraspeckle proteins were distributed across the nucleoplasm and the number of paraspeckle foci was decreased in Neat1 deficiency [10]. Interestingly, it was shown that upon infliction of stress such as Polyinosinic:polycytidylic acid induction of type-I interferon response, there was an increase in the transcription levels of Neat1 and in the formation of paraspeckle foci in the nucleus [4, 9, 10, 17]. Neither Neat1 nor the paraspeckles proteins were studied so far in normal or perturbed hematopoiesis.
Paraspeckles are found in almost every cell type, including primary cells and cell lines, except for human embryonic stem cells (hESC) [11]. Interestingly, when hESC were differentiated in culture, Neat1 was upregulated and paraspeckle foci were demonstrated to form in non-pluripotent cells [11]. Furthermore, recent studies suggested that the lack of expression of Neat1 and paraspeckles might indicate a loss of pluripotency in hESC [18]. Therefore, if a cell does not express Neat1 and/or paraspeckles this might serve as a marker for loss of pluripotency [18]. Induction of Neat1 and paraspeckles was also shown on the differentiation of myoblasts into myotubes, with a three-fold up-regulation of Neat1 and an increase in paraspeckle number and size [1]. This is most significant with the recent discovery of NEAT1 isoforms role in pri-miRNA processing [16]. Intriguingly, the role of Neat1 and paraspeckles has not yet been studied in hematopoiesis before, although findings in other cell types make it an interesting topic for research in the context of adult stem cell and differentiation.
HSCs research is leading both basic research and clinical applications of adult stem cells [19]. Therefore, the finding of paraspeckles which hold the potential to influence proliferation and differentiation of HSCs is of great interest. Characterization of Neat1 expression and its isoforms in primary HSCs and in defined hematopoietic progenitors is needed in order to establish its possible role in early hematopoiesis. Through the study of HSCs and the factors that influence their potential to proliferate and differentiate, we may achieve new innovations in the study of bone marrow transplantation and blood cancers [19]. Paraspeckles were reported to correlate with the cell cycle and are associated with the appearance of RNA polymerase II and changes in the metabolic activity of cells [8, 12]. These properties are of major interest in hematopoiesis, making Neat1 and paraspeckles a focal point of interest for research in HSCs. Recently, it was reported that Neat1 is induced by p53 and plays a role in suppressing transformation and cancer initiation [20], thus strongly suggesting Neat1 as a new factor on early malignancy.
In this study, we analyzed the expression of Neat1 lncRNA and of paraspeckle proteins in mouse HSCs and in their direct progeny. RNA-seq data and RT-qPCR show that Neat1_1 is highly expressed in HSCs and in early progenitors, while Neat1_2 is expressed at lower levels. FISH identified bright Neat1 foci in the nucleus, and paraspeckle proteins NONO and PSPC1 show distinguished nuclear expression. As for differentiated hematopoietic cells, granulocytes have the lower expression levels of Neat1, NONO and PSPC1, while B-cells maintain relatively higher expression levels of all three. Furthermore, ex-vivo cultured cells presented increased expression of Neat1 when differentiation was induced. Taken together, our data present multiple evidence for Neat1 and paraspeckles in different stages of hematopoiesis for the first time.
RESULTS
Neat1 is expressed throughout hematopoiesis
To begin the study of Neat1 in HSCs, we first evaluated the specific expression levels of Neat1_1 and Neat1_2 by analyzing RNA-seq data [21]. The aligned-reads clearly showed that the Neat1_1 isoform is highly expressed in HSCs (Figure 1A). RNA-seq data suggested that Neat1_2 isoform was expressed to a much lesser extent, and there was no complete transcript of Neat1_2 profoundly detected (Figure 1A, see longer-transcript). Since Neat1 has a structural role, and the longer-isoform might be less well detected by the RNA-seq, we performed independent RT-qPCR for Neat1_1 and Neat 1_2 on mouse HSCs, progenitors and effector immune cells. HSCs, as well as other hematopoietic progenitors and differentiated cell types, showed a significant difference between Neat1_1 to Neat1_2 expression in primary cells (Figure 1B with HSC: p<0.01; MPP: p<0.01; CMP: p<0.01; GMP: p<0.01; MEP: p<0.01; Gran: p<0.01; B-Cells: p=0.018). Moreover, there was a significant difference in the levels of Neat1_1 expression between HSCs to GMPs (p<0.01), MEP (p=0.019), and granulocytes (p=0.012) in that HSCs had higher expression by 2.45, 3.12, and 2.52-fold, respectively. The rest of the cells examined (MPP, CMP and B-Cells) showed Neat1_1 expression levels which did not significantly differ of HSCs (Figure 1B). Concerning the relative Neat1_2 expression, HSCs showed higher expression levels compared to GMPs, by 3.09-fold (p=0.032). Notably, the RT-qPCR data suggests relatively high levels of Neat1 in HSCs, at about 0.1-fold of the expression of β-Actin which is an extremely abundant transcript (Figure 1B).
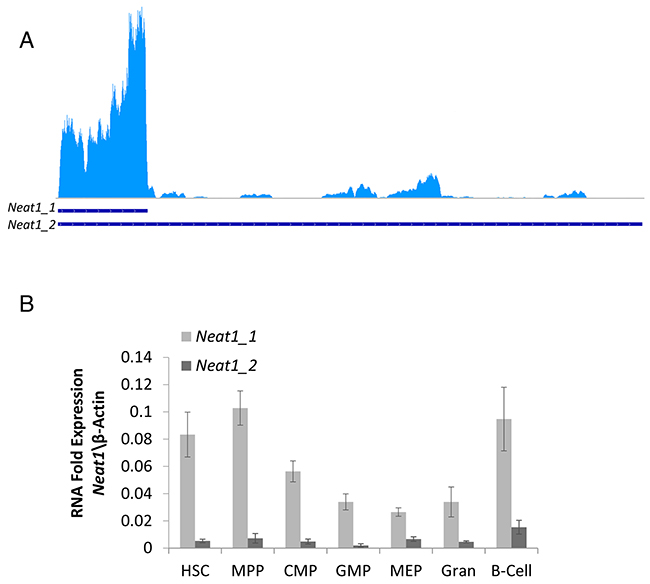
Figure 1: Neat1_1 and Neat1_2 RNA expression in hematopoietic stem cells. (A) Neat1_1 and Neat1_2 lncRNA transcripts of 3170bp and 20177bp, respectively, presented by blue lines on bottom. RNA-seq data show that Neat1_1 Is highly expressed in HSCs, whereas Neat1_2 is expressed to a much lesser extent. (B) RT-qPCR detection of Neat1_1 and Neat1_2, shown as relative to β-actin, from HSC, MPP, CMP, GMP, MEP, granulocyte, B-cells. Histograms show averages ±SD.
Neat1 RNA forms sharp foci in HSCs and progenitors
After we managed to show RNA expression of Neat1_1 and Neat1_2, we wanted to visualize it in HSCs, progenitors and differentiated cells. We used FISH technique aimed for Neat1 isoforms, using commercial Stellaris® FISH Probes. The representative panels show that each primary cell type has usually 1 or 2 foci of Neat1_1/2 which are all localized in the nucleus, and may suggest the formation of paraspeckles in these cells (Figure 2). Interestingly, when both Neat1_1 and Neat1_2 are detected they seem to colocalize as shown for example in HSCs and GMPs (Figure 2). This reinforces the existence of paraspeckles in mouse hematopoietic stem- and progenitor cells.
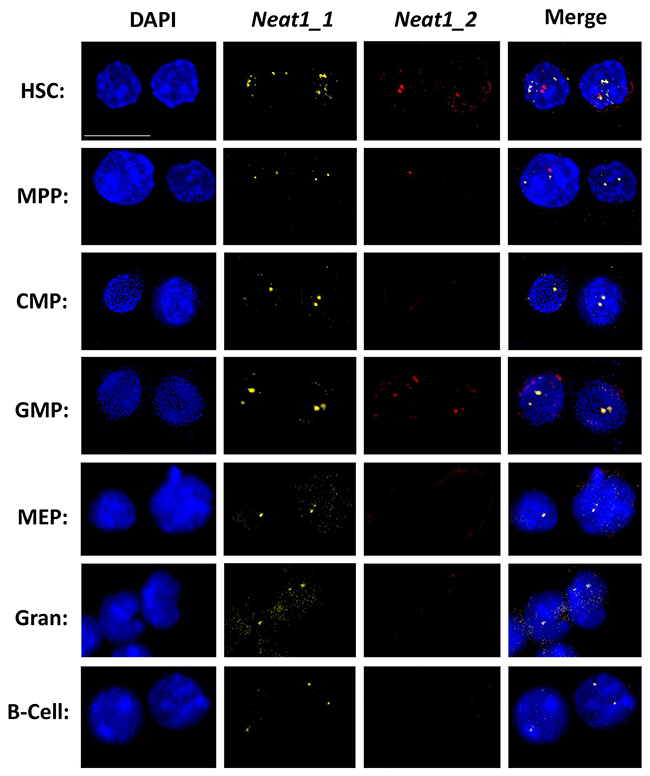
Figure 2: Primary NEAT RNA forms sharp foci in HSCs and progenitors. Confocal microscopy analysis of Neat1_1 and Neat1_2 using FISH, in representative primary cell types through hematopoiesis, HSC, MPP, CMP, GMP, MEP, granulocyte, B cell. Neat1_1 (yellow) and Neat1_2 (red) are superimposed over nuclei stained with DAPI (blue). Scale bar denotes 10 μ.m.
Regarding expression levels, Neat1_1 was more vivid in all the cell types that we examined, in agreement with RNA-seq and RT-qPCR data (Figure 1). Neat1_2 was bright only in some cells such as HSCs, but not easily detected in granulocytes (Figure 2). It seems that there is a slight agreement between the RT-qPCR data (Figure 1B) and the images acquired using FISH, possibly due to inherent differences between the techniques. Taken together, our data demonstrate the expression and the nuclear localization of Neat1 in hematopoietic stem-cells, progenitors, and effector immune cells.
Paraspeckle protein NONO displays aggregated clusters in hematopoietic cells
In addition to Neat1 RNA, we wanted to visualize known paraspeckle proteins [12]. We visualized NONO and PSPC1 using immunofluorescence. Representative images show that each primary cell type tested expresses nuclear NONO foci with variance in amount, fluorescence intensity and size (Figure 3A). These foci may present paraspeckles which according to the literature can be in different size, number and intensity depending on cell type, and its cell cycle state [1, 7, 22]. Furthermore, the fact that they are seen in foci and not stained in a diffuse manner suggests that they might be attached to a paraspeckle. Interestingly, the visualized NONO-foci seem to localize mostly in the sub-areas of the nucleus that are more euchromatin, and there were no foci in dense DNA areas which are heterochromatin rich (Figure 3A and Supplementary Figure 1).
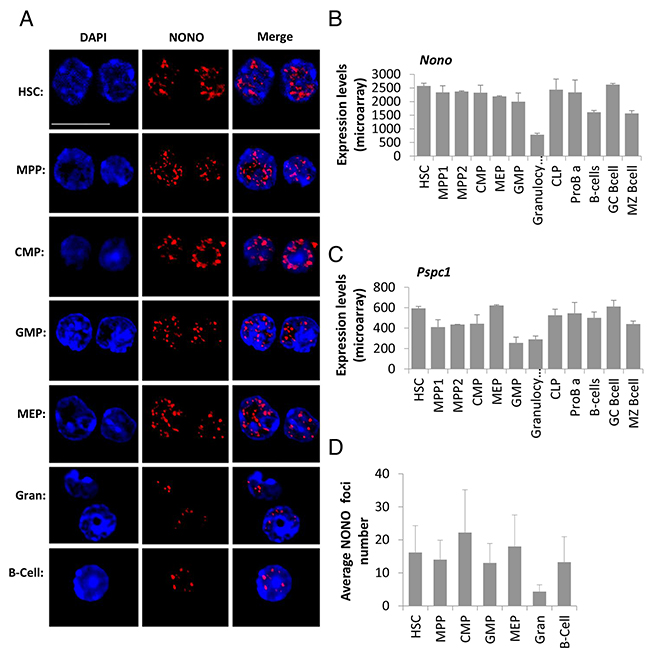
Figure 3: Paraspeckle protein NONO presents aggregated clusters in hematopoietic cells’ nuclei and reduced expression in granulocytes. (A) Confocal microscopy analysis of NONO, a paraspeckle protein, using an immunofluorescent staining technique, in representative primary cell types through hematopoiesis, HSC, MPP, CMP, GMP, MEP, granulocyte, B cell. NONO (red) is superimposed over nuclei stained with DAPI (blue). Scale bar denotes 10 μ.m. (B and C) Expression levels of mRNA of the paraspeckles proteins NONO and PSPC1, respectively. Data is from ImmGen’s microarray database, showing averages of at least triplicates per cell type except for MPP and MEP that are duplicates; and SD per cell type. (D) Quantification of the foci number per cell type from immunofluorescent stained cells for NONO. Histograms show averages ±SD.
Paraspeckles proteins show reduced expression in granulocytes
In order to quantify the paraspeckle proteins we first used published expression data from the ImmGen consortium [23] to determine the mRNA expression levels of Nono and Pspc1. It shows that while both are expressed in all cell types, Nono expression levels throughout hematopoiesis are higher than Pspc1 (Figure 3B and 3C, about 4-fold). This may explain why the PSPC1 immunostaining was less vivid compared to NONO (Supplementary Figure 2). Furthermore, this data suggests no substantial differences among the examined cell types, with the exception of granulocytes. HSCs have a higher expression of Nono and Pspc1 as compared to granulocytes by 3.28 and 2.04-fold respectively (p<0.01). On the other hand, only Pspc1 showed higher expression in HSCs compared to GMPs, by 2.32-fold (p<0.01). Also, HSCs have a slightly higher mRNA expression for Nono and Pspc1 compared with naïve B-cells by 1.59 and 1.18-fold, respectively (p<0.01 and p=0.018), and for marginal-zone B-cells by 1.64 and 1.34-fold for Nono and Pspc1, respectively (p<0.01, both). This RNA expression data is in concordance with the protein staining quantification we performed on the cells stained for NONO (Figure 3A). Nevertheless, the enumeration for the NONO foci done by CellProfiler found no significant differences between the number of foci in the cells that were stained, except for granulocytes which showed reduced number of foci compared to HSCs (Figure 3D, average foci per cell 4.3 to 16.1 respectively, p=0.018). Thus, we can see a good agreement between RNA and protein levels. The high standard deviation of CellProfiler analysis might derive at least in part from inherent technical inability to enumerate the foci in the highest microscopic resolution. Nevertheless, microscopic images present a substantial variability of paraspeckle formation, suggesting that along with the significant difference between the population-averages of HSCs and granulocytes, there is truly a substantial variability between cells of the same population with respect to NONO foci numbers.
Neat1 expression is enhanced by differentiation in vitro
Subsequent to the findings in primary fresh cells, we tested cells in culture. To induce differentiation, cultured primary cells were split into separate wells and while some remained on minimal-media the others were supplemented with GM-CSF and Serum to induce further differentiation. After 96 hours of induction the cells were examined by RNA quantification, FISH, and Immunofluorescence staining. RT-qPCR for Neat1 isoforms expression was normalized to the housekeeping gene β-Actin. The addition of GM-CSF and Serum to the culture medium increased the RNA expression of Neat1_1 by 1.56-fold (p<0.01) and Neat1_2 by 1.59-fold (p=0.03, Figure 4A and 4B). Similar results were seen in independent experiments (with the total of 5 biological replicates; Supplementary Figure 3A and 3B). FISH detection of Neat1_1 and Neat1_2 RNA in the cultured cells yielded bright expression. Microscopic images show that as a consequence of adding GM-CSF and serum to the cultured cells there was an increase in the staining of Neat1_1 and relatively higher increase of Neat1_2 (Figure 4C). This comes in concordance with the higher expression levels shown using RT-qPCR. Moreover, there is a better colocalization of Neat1_1 and Neat1_2, suggesting the enhancement of paraspeckles in cultured hematopoietic cells that were pushed towards differentiation. Similar results were visualized for cultured primary MPPs (Supplementary Figure 3), further implying that the induction of differentiation in-vitro upregulates Neat1 expression, and possibly the paraspeckle formation in hematopoietic cells.
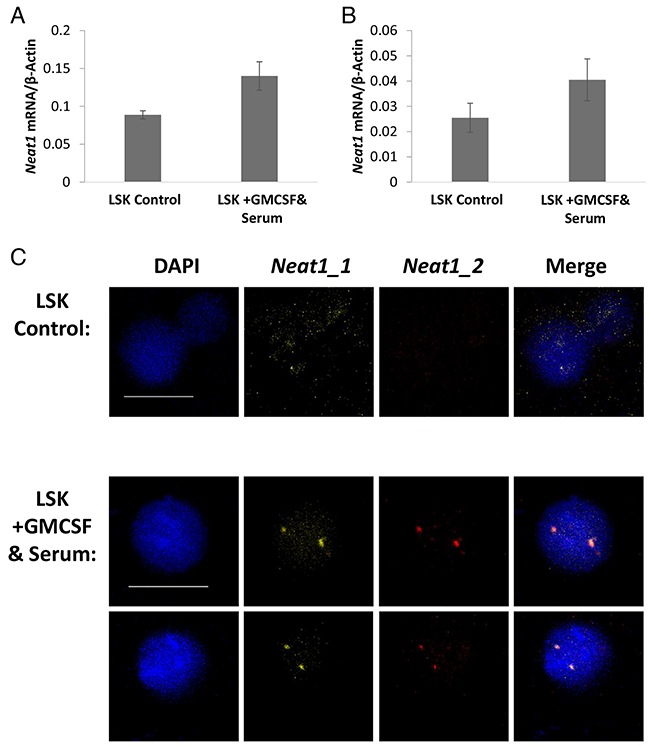
Figure 4: Neat1 expression is enhanced by differentiation in vitro. (A and B) Expression levels of Neat1_1 and Neat1_2 measured by RT-qPCR and shown relative to β-Actin. GMCSF and Serum were added to the medium for the last 96 hours of the culture. Histograms show averages ±SD. (C) Confocal microscopy FISH of Neat1_1 and Neat1_2. Scale bar denotes 10 μ.m.
NONO sustains aggregated nuclear expression in cultured cells
To further visualize paraspeckles in the cultured cells we stained for the NONO protein. The cultured cells did not show any significant difference in the amount, intensity or size of NONO foci (Figure 5 and Supplementary Figure 4). Although there was no difference between cells that were induced into differentiation, cells of both groups sustained nuclear foci integrity, suggesting that the culture conditions are not dismantling paraspeckles. This finding strengthens the claim of presence of these sub-nuclear structures in hematopoietic cells, not only in vivo but also in vitro. In addition, it further suggests that there is no strict limiting correlation between the paraspeckle proteins and the Neat1 RNA.
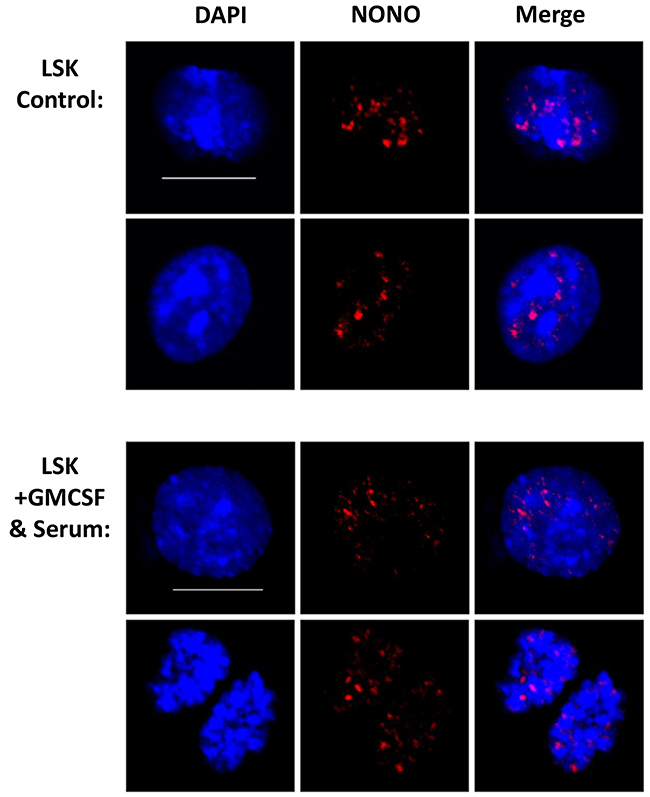
Figure 5: NONO sustains aggregated nuclear expression in cultured cells. Confocal microscopy analysis of NONO using Immunofluorescent staining technique for cultured cells. Representative images are shown from independent experiments and total of 5 biological replicates. Scale bar denotes 10 μ.m.
DISCUSSION
Learning of the expression of Neat1 and formation of paraspeckles in hematopoietic cells will advance the research in the field of normal hematopoiesis and blood malignancy, considering the data on Neat1 in other cancer types and the lack of data in the hematopoietic system [24-26]. In this study, we have laid the foundation to elucidating the manner in which Neat1 and paraspeckles are expressed in hematopoietic stem-cells, progenitors, and differentiated immune cells both in vivo and in vitro. Our data demonstrate the expression of Neat1 and visualize paraspeckle formation in HSCs and progenitors. We do identify pronounced downregulation of Neat1 and paraspeckles proteins in granulocytes, but not in B-cells. These findings will lead for innovative research in the fields of hematopoiesis, stem cell differentiation and in blood cancers.
Neat1 and its isoforms in hematopoietic cells
Our data demonstrate that Neat1 is expressed in HSCs, progenitors and immune cells, by analysis of RNA-seq, independent RT-qPCR and direct FISH visualization. Furthermore, Neat1 seems to form sharp nuclear foci in these cells, both in vivo and in vitro. Neat1_1 is highly expressed, quantified at 0.1-fold of β-Actin which is known to be an extremely-highly expressed gene, and is visualized most clearly in all cells. Although Neat1_1 is very profound, Neat1_2 is not expressed as much, and when attempting to visualize it appeared stronger in HSCs and MPPs than in the differentiated cells such as granulocytes or B-cells. This may suggest that when a cell has the potential to proliferate and differentiate it expresses higher levels of Neat1, especially Neat1_2. This might be of special interest with recent discovery of NEAT1_2 in pri-miRNA processing [16]. Cells which are terminally differentiated, and post-mitotic (like granulocytes) do not express Neat1 as much. This is in concordance with the data in the literature concerning the role of Neat1 and paraspeckles in cell cycle of other cell types [1, 7, 8, 12, 22, 26]. This point is of interest in malignant hematopoietic cells, which may consist of highly-proliferative along with more quiescent cells, and would specifically be of interest regarding Leukemia-Initiating-Cells (LICs, sometimes called “cancer stem cells”). Our data indicate that HSCs, which are deeply quiescent, express Neat1 at their dormant primary multipotent state.
Paraspeckles are formed in hematopoietic cells
Paraspeckles are formed in most cell types, with the exception of hESC [11], yet up until now paraspeckles were not visualized in hematopoietic cells. In this study, we show for the first time the foci formed by paraspeckles proteins NONO and PSPC1 in primary hematopoietic cells. We found that Nono is highly expressed compared to Pspc1 at the mRNA level, and the microscopic images suggested similar order with substantially Higher NONO than PSPC1 protein staining (Figure 3A and Supplementary Figure 2). PSPC1 images were generally less-sharp, possibly due to its lower expression levels. Interestingly, enumeration of NONO foci, as a proxy for paraspeckles, found substantial variability among cells (Figure 3D). This is in concordance with the literature as NONO is expressed in foci of various sizes, intensities and number per nucleus in various cells such as Myoblasts, HeLa, NIH 3T3, and other cell lines [1, 7, 22]. Finding paraspeckles protein’s foci is not obvious, because in some cases it was reported that when Neat1 expression is downregulated they are disrupted, showing diffused staining, suggesting dismantle of the sub-nuclear structure [10]. All the hematopoietic cells tested presented with aggregated NONO and PSPC1 proteins, suggesting no dismantling of paraspeckle foci in these normal primary cells. It was reported that paraspeckles and Neat1 may change with the cell cycle of other cells [8, 12, 26]. However, finding high levels of Neat1, NONO and PSPC1 in HSCs is in contrast with their non-proliferative dormant state. Our current data suggest no correlation of Neat1 or paraspeckle formation with HSC quiescence. Moreover, we visualized paraspeckles in sub-nuclear areas that are euchromatic, thus their DNA is in a transcription-ready state. These two findings imply that Neat1 and paraspeckles are fairly abundant in primary quiescent HSCs and active progenitor cells, thus they may have regulatory role on the proliferative potential rather than the acute stage within the cell cycle.
Paraspeckle foci are sharpened with induced-differentiation ex-vivo
After previous publications regarding Neat1’s role in other cell types, and its overexpression during excessive transcription and DNA replication [8-10, 12, 17, 26], we hypothesized that hematopoietic cells pushed towards differentiation would upregulate their expression of Neat1 and paraspeckle formation. Our RT-qPCR and the images acquired using FISH to visualize Neat1 were in line with this hypothesis. Thus, it is suggested that Neat1 may play a role in differentiation of effector cells. Interestingly, the aggregation of paraspeckle proteins at euchromatic regions in the nucleus and their reported role in DNA-breaks in Mouse Embryonic Fibroblasts (MEFs) [27] may further suggest a possible similar function in hematopoietic cells. It would therefore be interesting to further study Neat1 and paraspeckles in HSCs throughout aging, which is reported to involve the accumulation of DNA-damage and double-strand breaks [28].
MATERIALS AND METHODS
RT-qPCR
Experiments were performed according to the local and state ethics committee approval. Cells were extracted from the femur, pelvis, and tibia of wildtype C57BL/6J mice that are housed at the SPF unit of BGU; mice were euthanized using isoflurane and bone marrow cells were obtained by crushing; mononuclear cells were enriched by Histopaque (H1083, Sigma-Aldrich). Cells were stained for Lineage (Ter119, CD11b, GR-1, CD3e, CD4, CD8, and B220), Sca1, cKit, CD34, CD150 and CD48, or for B220, CD19, GR-1 and MAC1; all antibodies from Biolegend, see Supplementary Figure 5 for FACS-gating. Cells were sorted by FACSAriaIII (BD biosciences) and stored in TRIzol (Invitrogen) for RNA, or fixed and stained for FISH and immunohistochemistry. Total of 6 mice in 3 independent experiments were used for RT-qPCRs; triplicates were routinely used for statistical analysis of RT-qPCR. RNA was extracted according to the manufacturer’s protocol, followed by reverse transcription using PrimeScriptTM RT-PCR Kit (Takara). qPCR reactions were set with Kappa SYBR Fast qPCR master mix, and run on LightCycler 480 (Roche), using following Primers; Neat1_1 forward: GATCGGGACCCCAGTGACCT Reverse: AGCTTTCCCCAACACCCACA, Neat1_2 forward: GCTCTGGGACCTTCGTGACTCT Reverse: CTGCCTTGGCTTGGAAATGTAA; Actin-beta forward: CTCTGGCTCCTAGCACCATGAAGA Reverse: GTAAAACGCAGCTCAGTAACAGTCCG. PCR program was: Pre-incubation heat to 95°C 5min sec; amplification 45 cycles of 95°C 5sec, 60°C 15sec, 72°C 20sec; melting heat to 95°C 5sec, 65°C 1min and then heat to 97°C by 0.11C/sec.
Immunohistochemistry
Bone-marrow cells were extracted from femur, tibia and pelvis as above; for primary fresh cells, a total of 9 mice in 3 independent experiments were used, and for cultured cells a total of 5 mice in 2 independent experiments were used. Sorted cells were taken for immunostaining in 96U well Falcon plate. Fixation and permeabilization of cells using “True-NuclearTM Transcription Factor Buffer Set [CAT:424401, BioLegend]. The anti-mouse H-2Dd clone: 34-2-12 [CAT: 110602 BioLegend] was used for isotype control. Monoclonal Anti-PSPC1 cat#SAB4200503 and monoclonal Anti-NONO cat#SAB4200502 (Sigma). Cells were stained overnight at 4°C, washed twice and fluorescently labeled with secondary DylightTM 594-conjugated AffiniPure Rat Anti-Mouse IgG [cat# 415-515-166, Jackson ImmunoResearch]. DNA was stained with DAPI (4’,6-Diamidino-2-Phenylindole, Dihydrochloride; 1μg/ml). Cells were mounted on slides and images taken with Olympus Confocal Microscope using Olympus Fluoview FV1000. Quantification of foci was done using the CellProfiler software by Broad Institute [32]. For fresh or for cultured cells about 10 or 30 cells were used for CellProfiler analysis, respectively.
Fluorescence in situ hybridization (FISH)
Sorted cells (as above) were obtained from a total of 5 mice in 2 independent experiments. FISH was performed in 96U well Falcon plates after Fixation for 10 minutes with 4%PFA in PBS [cat:#15710 Electron Microscopy Science]; Permeabilization for two hours in 70% Ethanol; Hybridization using the two probes overnight at 30°C. Probes were Stellaris® FISH Probe Mouse Neat1 5’ Segment with Quasar® 570 Dye [SMF-3009-1], and Stellaris® FISH Probe Mouse Neat1 Middle Segment with Quasar® 670 Dye [SMF-3010-1, custom-conjugated to enable dual-color detection]. Nuclei were counter-stained using DAPI 1μg/ml. Cells were mounted on slides using GLOX mounting media [contains: Tris pH 8.0, SSC, glucose and nuclease free water] supplemented with 1μL Catalase and 1μL Glucose Oxidase. Cells were observed using Olympus Confocal Microscope and images were taken using Olympus Fluoview FV1000.
Primary culture
Sorted cells were cultured in BioTraget media (Biological Industries, Israel) supplemented with 10ng/ml of SCF, TPO, IL-3 and Flt3L (Peprotech) as a minimal media that support cell viability in vitro. Cells were grown in 96U plates with the media changed twice a week and split according to their growth. Differentiation was further induced by adding GM-CSF (10ng/ml) and Fetal Bovine Serum (10%). Viability was tested microscopically to ensure that cells were in good condition. Cells from individual mice were cultured in parallel to obtain biological replicates, and independent experiments were performed from independent primary sorts, using total of 5 mice in 2 independent experiments.
Statistical analysis
Results are shown as the mean ±standard deviation. Two-tailed and one-tailed paired Student’s t-tests were used to compare different groups. Statistical p < 0.05 was considered significant.
CONCLUSIONS
The data we present hereby provide the foundation for studying the effect(s) of lncRNA Neat1 and paraspeckles on blood cancer, including Leukemia, Myeloma or Lymphoma. Other studies showed that Neat1 may increase proliferation, invasion, Endothelial-to-Mesenchymal-Transition (EMT), and chemo-resistance in multiple cancer types including lung cancer, breast cancer and in renal-cell carcinoma [24-26]. Other lncRNAs were reported to genetically associate and functionally regulate blood cancers [2, 5]. Yet, there was no study that spotlighted Neat1 and focused on its role in paraspeckle formation in hematologic cancers. Most interestingly, a recent study revealed Neat1 dysregulation in Multiple Myeloma [29], pioneering its clinical hematological relevance. Strikingly, Neat1 was recently reported to be directly induced by p53, and to suppress early malignant neoplasm [4, 20]. Neat1 had been reported to relate with breast cancer, suggesting it as a biomarker [30]. We had searched for hematological cancers survival correlation through SurvExpress [31], finding significant Hazard Risk with some of the Lymphoma and in Multiple Myeloma (data not shown), suggesting further focused interest. Our study offers a base for future research, with solid findings of the normal state and the ability to quantify and visualize Neat1 and paraspeckle proteins in primary hematopoietic cells. This study demonstrates, for the first time, the consistent expression of Neat1_1 and the formation of paraspeckles in HSCs, progenitors and effector immune cells.
Abbreviations
Neat1: Nuclear Enriched Abundant Transcript 1; HSCs: hematopoietic stem cells; lncRNAs: long non-coding RNAs; Menε: Multiple Endocrine Neoplasia ε; Menβ: Multiple Endocrine Neoplasia β; hESC: human embryonic stem cell; MPP: multipotent progenitors; CMP: common myeloid progenitor; GMP: granulocyte-macrophage progenitor; MEP: megakaryocyte-erythroid progenitor; Gran: granulocyte; CLP: common lymphoid progenitors; FISH: Fluorescence in Situ Hybridization; MEFs: Mouse Embryonic Fibroblasts; GM-CSF: granulocyte-macrophage colony-stimulating factor; EMT: Endothelial-to-Mesenchymal-Transition; RNA-seq: RNA sequencing; RT-qPCR: reverse transcription quantitative polymerase chain reaction; PFA: paraformaldehyde; PBS: phosphate-buffered saline.
ACKNOWLEDGMENTS
Authors are thankful to Shalev Itskovitch for help with FISH and for discussions and advice; to Debbie Toiber and Daniel Stein for technical help with CellProfiler; to Alon Zilcha, Alex Braiman and Uzi Hadad for technical help with microscopy.
CONFLICTS OF INTEREST
Authors declare no conflicts of interest on this study.
FUNDING
This study was supported by ISF grant 1690/13, PCIG13-GA-2013-618647, and ICRF grant 16-1232-RCDA.
REFERENCES
1. Sunwoo H, Dinger ME, Wilusz JE, Amaral PP, Mattick JS, Spector DL. MEN epsilon/beta nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res. 2009; 19: 347-59. https://doi.org/10.1101/gr.087775.108.
2. Trimarchi T, Bilal E, Ntziachristos P, Fabbri G, Dalla-Favera R, Tsirigos A, Aifantis I. Genome-wide mapping and characterization of Notch-regulated long noncoding RNAs in acute leukemia. Cell. 2014; 158: 593-606. https://doi.org/10.1016/j.cell.2014.05.049.
3. Luo M, Jeong M, Sun D, Park HJ, Rodriguez BA, Xia Z, Yang L, Zhang X, Sheng K, Darlington GJ, Li W, Goodell MA. Long non-coding RNAs control hematopoietic stem cell function. Cell Stem Cell. 2015; 16: 426-38. https://doi.org/10.1016/j.stem.2015.02.002.
4. Adriaens C, Standaert L, Barra J, Latil M, Verfaillie A, Kalev P, Boeckx B, Wijnhoven PW, Radaelli E, Vermi W, Leucci E, Lapouge G, Beck B, et al. p53 induces formation of NEAT1 lncRNA-containing paraspeckles that modulate replication stress response and chemosensitivity. Nat Med. 2016; 22: 861-8. https://doi.org/10.1038/nm.4135.
5. Schwarzer A, Emmrich S, Schmidt F, Beck D, Ng M, Reimer C, Adams FF, Grasedieck S, Witte D, Kabler S, Wong JW, Shah A, Huang Y, et al. The non-coding RNA landscape of human hematopoiesis and leukemia. Nat Commun. 2017; 8: 218. https://doi.org/10.1038/s41467-017-00212-4.
6. Hutchinson JN, Ensminger AW, Clemson CM, Lynch CR, Lawrence JB, Chess A. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics. 2007; 8: 39. https://doi.org/10.1186/1471-2164-8-39.
7. Clemson CM, Hutchinson JN, Sara SA, Ensminger AW, Fox AH, Chess A, Lawrence JB. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell. 2009; 33: 717-26. https://doi.org/10.1016/j.molcel.2009.01.026.
8. Fox AH, Lam YW, Leung AK, Lyon CE, Andersen J, Mann M, Lamond AI. Paraspeckles: a novel nuclear domain. Curr Biol. 2002; 12: 13-25.
9. Hirose T, Virnicchi G, Tanigawa A, Naganuma T, Li RH, Kimura H, Yokoi T, Nakagawa S, Benard M, Fox AH, Pierron G. NEAT1 long noncoding RNA regulates transcription via protein sequestration within subnuclear bodies. Mol Biol Cell. 2014; 25: 169-83. https://doi.org/10.1091/mbc.E13-09-0558.
10. Imamura K, Imamachi N, Akizuki G, Kumakura M, Kawaguchi A, Nagata K, Kato A, Kawaguchi Y, Sato H, Yoneda M, Kai C, Yada T, Suzuki Y, et al. Long noncoding RNA NEAT1-dependent SFPQ relocation from promoter region to paraspeckle mediates IL8 expression upon immune stimuli. Mol Cell. 2014; 53: 393-406. https://doi.org/10.1016/j.molcel.2014.01.009.
11. Nakagawa S, Naganuma T, Shioi G, Hirose T. Paraspeckles are subpopulation-specific nuclear bodies that are not essential in mice. J Cell Biol. 2011; 193: 31-9. https://doi.org/10.1083/jcb.201011110.
12. Fox AH, Bond CS, Lamond AI. P54nrb forms a heterodimer with PSP1 that localizes to paraspeckles in an RNA-dependent manner. Mol Biol Cell. 2005; 16: 5304-15. https://doi.org/10.1091/mbc.E05-06-0587.
13. Bond CS, Fox AH. Paraspeckles: nuclear bodies built on long noncoding RNA. Journal of Cell Biology. 2009; 186: 637-44. https://doi.org/10.1083/jcb.200906113.
14. Kawaguchi T, Hirose T. Architectural roles of long noncoding RNAs in the intranuclear formation of functional paraspeckles. Frontiers in Bioscience-Landmark. 2012; 17: 1729-46. https://doi.org/10.2741/4015.
15. Zhang Z, Carmichael GG. The fate of dsRNA in the nucleus: a p54(nrb)-containing complex mediates the nuclear retention of promiscuously A-to-I edited RNAs. Cell. 2001; 106: 465-75. https://doi.org/10.1016/S0092-8674(01)00466-4
16. Jiang L, Shao C, Wu QJ, Chen G, Zhou J, Yang B, Li H, Gou LT, Zhang Y, Wang Y, Yeo GW, Zhou Y, Fu XD. NEAT1 scaffolds RNA-binding proteins and the Microprocessor to globally enhance pri-miRNA processing. Nat Struct Mol Biol. 2017. https://doi.org/10.1038/nsmb.3455.
17. Naganuma T, Hirose T. Paraspeckle formation during the biogenesis of long non-coding RNAs. RNA Biology. 2013; 10: 456-61. https://doi.org/10.4161/rna.23547.
18. Chen LL, Carmichael GG. Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: functional role of a nuclear noncoding RNA. Mol Cell. 2009; 35: 467-78. https://doi.org/10.1016/j.molcel.2009.06.027.
19. Orkin SH, Zon LI. Hematopoiesis: an evolving paradigm for stem cell biology. Cell. 2008; 132: 631-44. https://doi.org/10.1016/j.cell.2008.01.025.
20. Mello SS, Sinow C, Raj N, Mazur PK, Bieging-Rolett K, Broz DK, Imam JFC, Vogel H, Wood LD, Sage J, Hirose T, Nakagawa S, Rinn J, et al. Neat1 is a p53-inducible lincRNA essential for transformation suppression. Genes Dev. 2017; 31: 1095-108. https://doi.org/10.1101/gad.284661.116.
21. Wilson NK, Kent DG, Buettner F, Shehata M, Macaulay IC, Calero-Nieto FJ, Sanchez Castillo M, Oedekoven CA, Diamanti E, Schulte R, Ponting CP, Voet T, Caldas C, et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell. 2015; 16: 712-24. https://doi.org/10.1016/j.stem.2015.04.004.
22. Souquere S, Beauclair G, Harper F, Fox A, Pierron G. Highly ordered spatial organization of the structural long noncoding NEAT1 RNAs within paraspeckle nuclear bodies. Mol Biol Cell. 2010; 21: 4020-7. https://doi.org/10.1091/mbc.E10-08-0690.
23. Gazit R, Garrison BS, Rao TN, Shay T, Costello J, Ericson J, Kim F, Collins JJ, Regev A, Wagers AJ, Rossi DJ; Immunological Genome Project Consortium. Transcriptome analysis identifies regulators of hematopoietic stem and progenitor cells. Stem Cell Rep. 2013; 1: 266-80. https://doi.org/10.1016/j.stemcr.2013.07.004.
24. Li S, Yang J, Xia Y, Fan Q, Yang KP. LncRNA NEAT1 promotes proliferation and invasion via targeting MiR-181a-5p in non-small cell lung cancer. Oncol Res. 2017. https://doi.org/10.3727/096504017X15009404458675.
25. Li X, Wang S, Li Z, Long X, Guo Z, Zhang G, Zu J, Chen Y, Wen L. The lncRNA NEAT1 facilitates cell growth and invasion via the miR-211/HMGA2 axis in breast cancer. Int J Biol Macromol. 2017. https://doi.org/10.1016/j.ijbiomac.2017.07.053.
26. Liu F, Chen N, Gong Y, Xiao R, Wang W, Pan Z. The long non-coding RNA NEAT1 enhances epithelial-to-mesenchymal transition and chemoresistance via the miR-34a/c-Met axis in renal cell carcinoma. Oncotarget. 2017; 8: 62927-38. https://doi.org/10.18632/oncotarget.17757.
27. Li S, Li Z, Shu FJ, Xiong H, Phillips AC, Dynan WS. Double-strand break repair deficiency in NONO knockout murine embryonic fibroblasts and compensation by spontaneous upregulation of the PSPC1 paralog. Nucleic Acids Res. 2014; 42: 9771-80. https://doi.org/10.1093/nar/gku650.
28. Rossi DJ, Seita J, Czechowicz A, Bhattacharya D, Bryder D, Weissman IL. Hematopoietic stem cell quiescence attenuates DNA damage response and permits DNA damage accumulation during aging. Cell Cycle. 2007; 6: 2371-6. https://doi.org/10.4161/cc.6.19.4759.
29. Sedlarikova L, Gromesova B, Kubaczkova V, Radova L, Filipova J, Jarkovsky J, Brozova L, Velichova R, Almasi M, Penka M, Bezdekova R, Stork M, Adam Z, et al. Deregulated expression of long non-coding RNA UCA1 in multiple myeloma. Eur J Haematol. 2017; 99: 223-33. https://doi.org/10.1111/ejh.12908.
30. Ding X, Zhang Y, Yang H, Mao W, Chen B, Yang S, Ding X, Zou D, Mo W, He X, Zhang X. Long non-coding RNAs may serve as biomarkers in breast cancer combined with primary lung cancer. Oncotarget. 2017; 8: 58210-21. https://doi.org/10.18632/oncotarget.17356.
31. Aguirre-Gamboa R, Gomez-Rueda H, Martinez-Ledesma E, Martinez-Torteya A, Chacolla-Huaringa R, Rodriguez-Barrientos A, Tamez-Pena JG, Trevino V. SurvExpress: an online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS One. 2013; 8: e74250. https://doi.org/10.1371/journal.pone.0074250.
32. Carpenter AE, Jones TR, Lamprecht MR, Clarke C, Kang IH, Friman O, Guertin DA, Chang JH, Lindquist RA, Moffat J, Golland P, Sabatini DM. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 2006; 7: R100. https://doi.org/10.1186/gb-2006-7-10-r100.

