INTRODUCTION
The number of people who will develop and die from cancer is expected to climb rapidly around the world [1]. In 2016, a total of 1,685,210 cases of cancer were newly diagnosed and 595,690 people died from cancer in the United States [2]. Meanwhile, approximately 4,292,000 new cancer cases and 2,814.000 cancer deaths were projected in China in 2015 [3]. It is imperative to deal with the rising cancer burden by taking full advantage of the knowledge embedded in cancer statistics, causes, and mechanisms and apply it to cancer prevention and screening [4]. Genetic-environmental interaction has been considered an essential factor in carcinogenesis [5]. Meanwhile, the roles of multiple genetic changes in influencing the evolution of cancer have been investigated over the years [6].
Recently, a number of investigations have been conducted to study the potential influence of Telomerase reverse transcriptase (TERT) and cleft lip and palate transmembrane 1-like (CLPTM1L) gene variation on cancer susceptibility. The TERT gene encodes the rate-limiting catalytic subunit of the telomerase enzyme that is vital to the maintenance of telomere DNA length, chromosomal stability, and cellular immortality [7]. CLPTM1L, encodes a protein linked to cisplatin resistance and associated with the susceptibility to cleft lip palate. CLPTM1L was found to be overexpressed in cisplatin-resistant ovarian cancer cells and to promote apoptosis in cisplatin-sensitive cells [8]. TERT-CLPTM1L genetic variants, including single nucleotide polymorphisms, have been reported by several genome-wide association studies (GWAS) to associate with the risk of multiple cancer types [9–11]. Among these polymorphisms, rs402710 (C > T) and rs401681 (C > T) have been widely studied for their potential effect on the susceptibility to cancer. Jiang et al. found that rs401681 polymorphism was associated with a decreased risk of lung cancer [12], but other studies found no association was found between rs401681 polymorphism and lung cancer risk [13, 14]. For the rs402710 polymorphism, Ito et al. observed remarkable effects on susceptibility in lung cancer [15], but the results were not repeated in other similar studies [16, 17]. We conducted a comprehensive meta-analysis of all relevant articles to provide a systematic and cumulative assessment of the association of CLPTM1L rs402710 and rs401681 polymorphisms and overall cancer risk.
RESULTS
Study characteristics
As shown in Figure 1, 176 publications were initially identified from PubMed and EMBASE electronic databases that met our inclusion and exclusion criteria. After title and abstract screening, 61 records were excluded. The remaining 115 full text articles were further assessed and 67 were further excluded for the following reasons: 1) 12 were meta-analyses; 2) 43 were irrelevant; and 3) 12 had insufficient information to calculate odds ratios (ORs) and 95% confidence intervals (95% CIs). Additionally, 3 relevant studies were also retrieved manually from the pool. In the end, 51 eligible articles were included in the final meta-analysis.
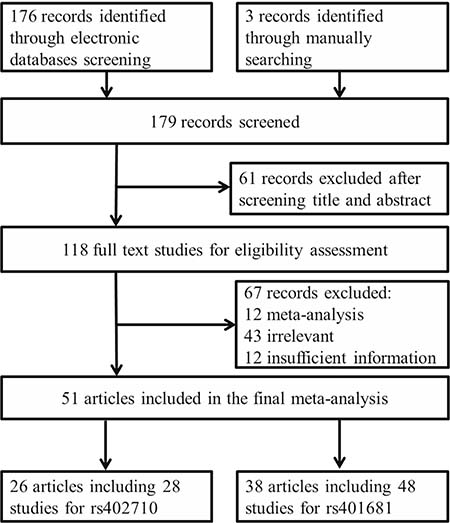
Figure 1: Flow diagram of studies included in our meta-analysis.
The genotypic distribution of rs402710 and rs401681 polymorphisms in the controls in almost all of the studies followed Hardy-Weinberg equilibrium (HWE); four did not (Table 1) [11, 17–19]. Since the genotypic distribution of other polymorphisms was in agreement with HWE in these four studies, the investigators decided to include them in the final analysis. Together, there were 26 articles with 28 studies including 30,770 cases and 34,089 controls for the rs402710 polymorphism [13, 15–39] and 38 articles with 48 studies including 67, 849 cases and 328, 226 controls for the rs401681 polymorphism (Table 1) [9–12, 14, 16–18, 21, 23–25, 28, 29, 32, 34, 35, 37, 38, 40–58]. For the rs402710 polymorphism, 19 studies focused on lung cancer, 2 on bladder cancer, and the rest on other individual types of cancer. There were 21 studies conducted on Asian populations, 5 on Caucasians, and 2 on Africans. 15 were hospital-based, and 12 were population-based. One was nested. As to the rs401681 polymorphism, 20 studies focused on lung cancer, 5 on bladder cancer and pancreatic cancer, 4 on melanoma, 3 on basal cell carcinoma, 2 on esophageal cancer and squamous cell carcinoma, and 7 on other individual cancer types, which we grouped as “the others” for our analyses. 24 were conducted on Asians, 21 on Caucasians, 2 on Africans, and only one on mixed ethnicity. There were 19 studies with a hospital-based design, 21 with a population-based design, and 8 with a nested design.
Table 1: Characteristics of case-control studies included in the current meta-analysis
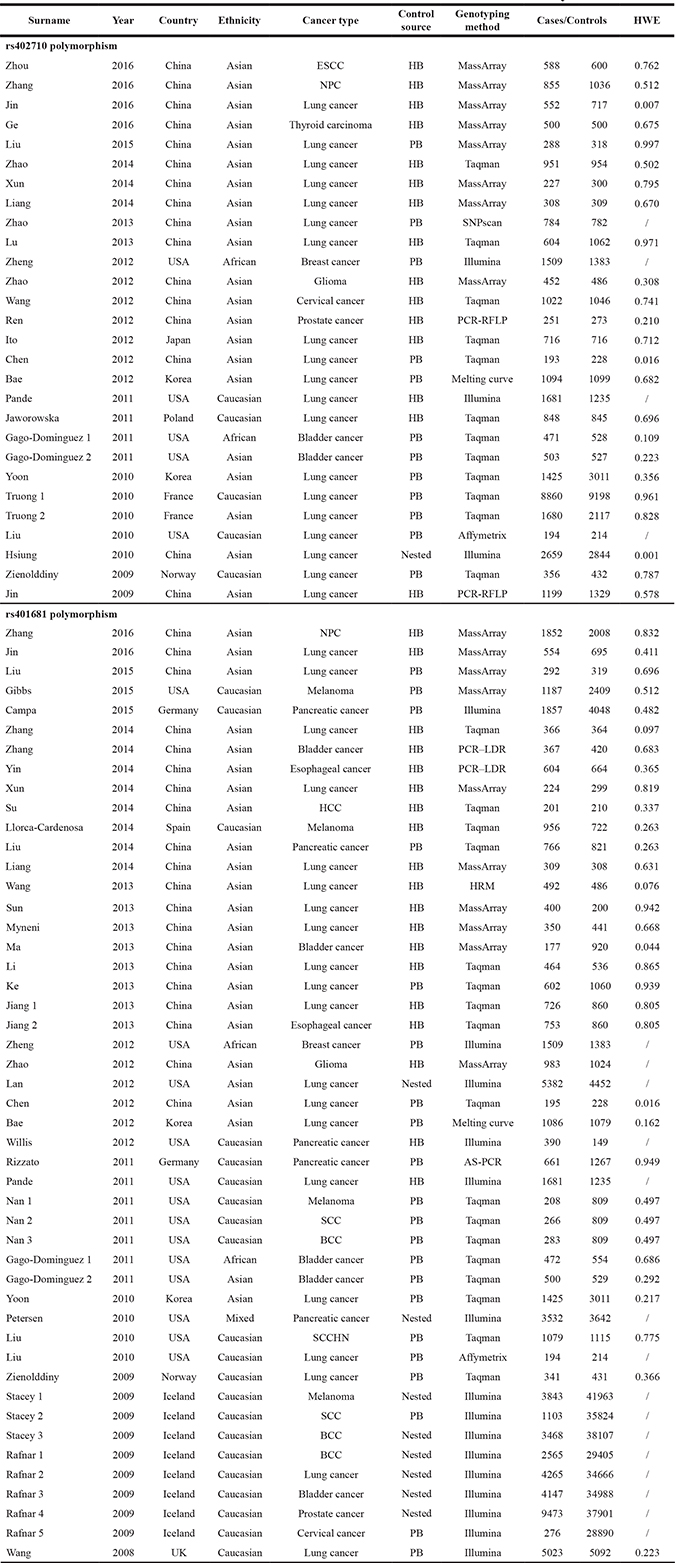
ESCC, esophageal squamous cell carcinoma; NPC, nasopharyngeal carcinoma; HCC, hepatocellular carcinoma; SCC, squamous cell carcinoma; BCC, basal cell carcinoma; SCCHN, squamous cell carcinoma of the head and neck; HB, hospital based; PB, population based; PCR-RFLP, polymerase chain reaction-restriction fragment length polymorphism; PCR–LDR, polymerase chain reaction–ligation detection reaction; HRM, high-resolution melt; AS-PCR, allele-specific polymerase chain reaction; HWE, Hardy-Weinberg equilibrium.
Meta-analysis results of the rs402710 polymorphism
As shown in Table 2 and Figure 2, there was evidence of a significant association between the rs402710 polymorphism and overall cancer risk under all of the five genetic models [homozygous (TT vs. CC): OR = 0.77, 95% CI = 0.70–0.84; heterozygous (CT vs. CC): OR = 0.89, 95% CI = 0.86–0.92; recessive (TT vs. CT+CC): OR = 0.81, 95% CI = 0.74–0.88; and dominant (CT+TT vs. CC): OR = 0.87, 95% CI = 0.83–0.91; as well as the allele comparison model (T vs. C): OR = 0.88, 95% CI = 0.84–0.92]. In the stratification analysis by cancer type, a statistically significant association was identified with lung cancer [homozygous (TT vs. CC): OR = 0.73, 95% CI = 0.67–0.79; heterozygous (CT vs. CC): OR = 0.88, 95% CI = 0.85–0.91; recessive (TT vs. CT+CC): OR = 0.77, 95% CI = 0.71–0.84; and dominant (CT+TT vs. CC): OR = 0.85, 95% CI = 0.82–0.88; as well as allele comparison model (T vs. C): OR = 0.85, 95% CI = 0.81–0.89] and bladder cancer [dominant (CT+TT vs. CC): OR = 0.83, 95% CI = 0.70–0.99; and allele comparison model (T vs. C): OR = 0.85, 95% CI = 0.75–0.98]. In the subgroup analysis by ethnicity and control source, there was a statistically significant association found in the studies of Asians and Caucasians in the hospital-based and population-based studies under all genetic models.
Table 2: Meta-analysis of the association between rs402710 and rs401681 polymorphisms and cancer risk
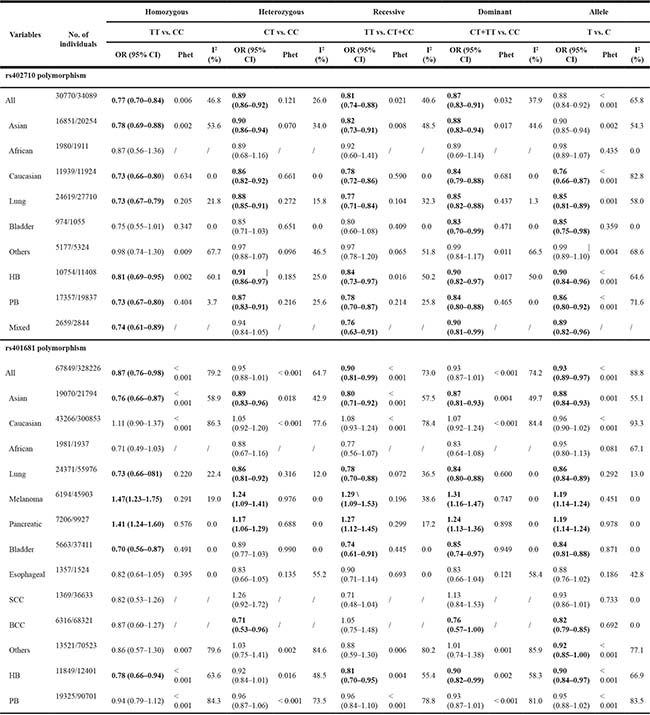
HB, hospital based; PB, population based; SCC, squamous cell carcinoma; BCC, basal cell carcinoma
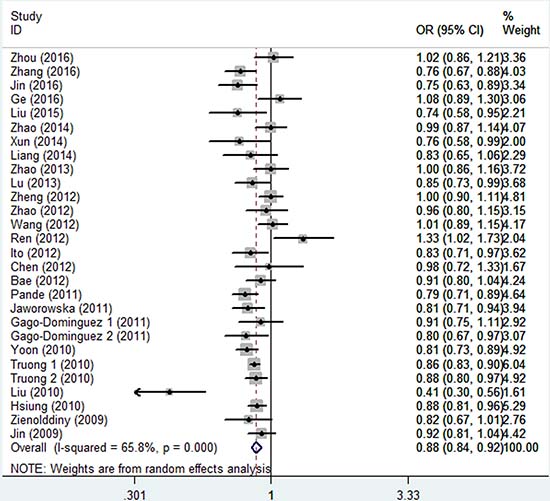
Figure 2: Forest plot of the association between CLPTM1L rs402710 polymorphism and overall cancer risk under the allele contrast model.
Meta-analysis results of rs401681 polymorphism
As shown in Table 2 and Figure 3, similar to the rs402710 polymorphism, a significant association between the rs401681 polymorphism and overall cancer risk was also observed [homozygous (TT vs. CC): OR = 0.87, 95% CI = 0.76–0.98; recessive (TT vs. CT+CC): OR = 0.90, 95% CI = 0.81–0.99; and allele comparison model (T vs. C): OR = 0.93, 95% CI = 0.89–0.97]. In the stratification analysis by cancer type, a decreased association was identified with lung cancer [homozygous (TT vs. CC): OR = 0.73, 95% CI = 0.66–0.81; heterozygous (CT vs. CC): OR = 0.86, 95% CI = 0.81–0.92; recessive (TT vs. CT+CC): OR = 0.78, 95% CI = 0.70–0.88; and dominant (CT+TT vs. CC): OR = 0.84, 95% CI = 0.80–0.88; as well as allele comparison model (T vs. C): OR = 0.86, 95% CI = 0.84–0.89], bladder cancer and basal cell carcinoma. On the contrary, there was an increased risk for melanoma [homozygous (TT vs. CC): OR = 1.47, 95% CI = 1.23–1.75; heterozygous (CT vs. CC): OR = 1.24, 95% CI = 1.09–1.41; recessive (TT vs. CT+CC): OR = 1.29, 95% CI = 1.09–1.53; and dominant (CT+TT vs. CC): OR = 1.31, 95% CI = 1.16–1.47; as well as allele comparison model (T vs. C): OR = 1.19, 95% CI = 1.14–1.24] and pancreatic cancer [homozygous (TT vs. CC): OR = 1.41, 95% CI = 1.24–1.60; heterozygous (CT vs. CC): OR = 1.17, 95% CI = 1.06–1.29; recessive (TT vs. CT+CC): OR = 1.27, 95% CI = 1.12–1.45; and dominant (CT+TT vs. CC): OR = 1.24, 95% CI = 1.13–1.36; as well as allele comparison model (T vs. C): OR = 1.19, 95% CI = 1.14–1.24]. When the analysis was stratified by ethnicity, there was a statistically significant association among Asians, but not in Caucasians or Africans. When the analysis was stratified by control source, there was a statistically significant association in the hospital-based studies, but not in population-based studies.
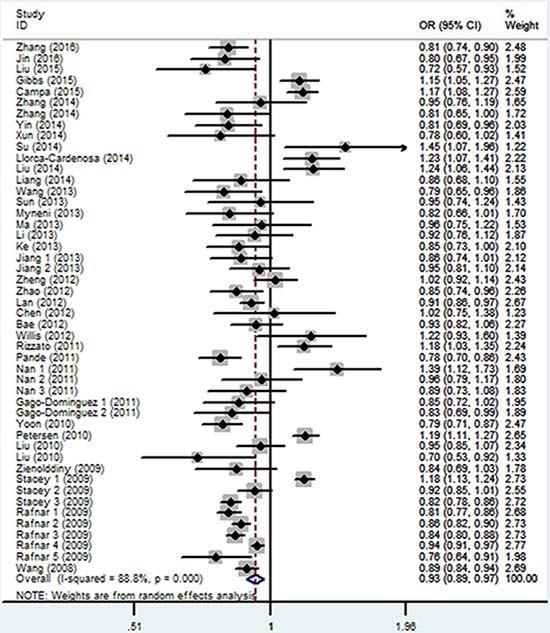
Figure 3: Forest plot of the association between CLPTM1L rs401681 polymorphism and overall cancer risk under the allele contrast model.
Heterogeneity and sensitivity analyses
Substantial heterogeneities were observed among all studies of the rs402710 polymorphism and cancer risk (homozygous: p = 0.006; recessive: p = 0.021; and dominant: p = 0.032; as well as allele comparison model: p < 0.001), except for the heterozygous model (p = 0.121). As to the rs401681 polymorphism, we also observed considerable heterogeneities (homozygous: p < 0.001; heterozygous: p < 0.001; recessive: p < 0.001; and dominant: p < 0.001; as well as allele comparison model: p < 0.001). So, the random-effects model was applied to generate crude ORs and 95% CIs. In addition, the leave-one-out sensitivity analysis indicated that no single study altered the corresponding pooled ORs and 95% CIs.
Publication bias
The shape of our funnel plot did not reveal any evidence of obvious asymmetry for the rs402710 polymorphism (homozygous: p = 0.738; heterozygous: p = 0.291; recessive: p = 0.954; and dominant: p = 0.318; as well as allele comparison model: p = 0.946) and rs401681 polymorphism (homozygous: p = 0.264; heterozygous: p = 0.451; recessive: p = 0.114; and dominant: p = 0.564; as well as allele comparison model: p = 0.967).
DISCUSSION
The CLPTM1L gene encodes the CLPTM1-like protein which is highly expressed in cisplatin-resistant ovarian tumor cell lines and is associated with cisplatin-induced apoptosis [8]. This gene has also been demonstrated to be overexpressed in many other cancers [58–61]. Blockade of CLPTM1-like protein has been shown to inhibit K-Ras-induced lung tumorigenesis [61]. CLPTM1L, near the TERT gene, is located at chromosome 5p15.33 which is widely identified as a susceptibility region associated with several types of cancer [62]. Some common genetic variants of TERT-CLPTM1L are hypothesized to have an important role in initiation and development of many cancers, including lung, bladder, pancreatic, thyroid, and breast cancer.
To the best of our knowledge, the meta-analysis in this study is the largest to study the associations between CLPTM1L rs402710 (C>T) and rs401681 (C>T) polymorphisms and overall cancer risk. In this meta-analysis, based on the database of publications in respect of the rs402710 polymorphism and the rs401681 polymorphism, among which there are 26 articles with 28 studies including 30,770 cases and 34,089 controls for rs402710 polymorphism and 38 articles with 48 studies including 67,849 cases and 328,226 controls for rs401681 polymorphism respectively, we observed both rs402710 and rs401681 polymorphisms were significantly associated with a decreased overall cancer risk. In the stratification analysis, as to the rs402710 polymorphism, a statistically significant association was also identified for lung and bladder cancer. Likewise, the rs401681 polymorphism was significantly associated with a decreased risk for lung cancer, bladder cancer and basal cell carcinoma. Interestingly, an increased risk of melanoma and pancreatic cancer was found for the rs401681 polymorphism.
Up until now, there had been only one meta-analysis (in 2013) focusing on rs402710 and rs401681 polymorphisms and overall cancer risk [62]; which only enrolled seven studies with 4,667 cancer patients and 4,990 controls for the rs402710 polymorphism and eight studies with 6,867 cancer patients and 7,746 controls for the rs401681 polymorphism. The said study found that rs402710 (A>G) and rs401681 (A>G) polymorphisms were associated with an increased cancer risk. In other words, rs402710 (C>T) and rs401681 (C>T) polymorphisms were significantly associated with a decreased overall cancer risk. In the stratification analysis by cancer type, Li et al. [62] included a few studies and did further stratification analysis only for lung and bladder cancer. Such associations were also observed with lung cancer from the meta-analyses before 2014 [63–65]. Since then, at least 8 studies for the rs402710 polymorphism and 13 studies for the rs401681 polymorphism have been published investigating the association with overall cancer risk, most of which focused on lung cancer. As the largest study with the strongest statistical power so far, the current meta-analysis also observed an association between the rs401681 polymorphism and an increased risk for melanoma and pancreatic cancer, which had been found in other meta-analyses [66, 67]. This observation suggested that the rs401681 polymorphism could play different roles in different types of cancer.
Several limitations in our meta-analysis should be acknowledged. First, we only included the studies published in English, which may have missed publications in other languages. Second, in the stratification analysis by cancer type and ethnicity, the sample sizes are relatively small and may be insufficient, which may have diminished the statistical power. Third, due to the lack of raw information such as age, sex, body mass index, smoking habits, and alcohol consumption, this meta-analysis was conducted on the basis of unadjusted estimates. Therefore, larger and well-designed studies with different types of cancer and multiple ethnic with sufficient raw information are warranted to validate the findings of this study.
In conclusion, the meta-analysis in this study demonstrated that CLPTM1L gene rs402710 and rs401681 polymorphisms were associated with a decreased risk of various types of cancer, especially for lung cancer among Asians.
MATERIALS AND METHODS
Search strategy
PubMed and EMBASE electronic databases were screened for publications investigating the association between CLPTM1L polymorphisms and overall cancer risk published prior to May 1, 2017. The following search terms were used: “CLPTM1L”, “5p15”, “polymorphism”, “variant”, “cancer”, “tumor” and “carcinoma”. Additional relevant studies were also retrieved manually from all the relevant publications, including the eligible original publications and reviews. Only the latest or the largest study would be included in the final meta-analysis.
Inclusion and exclusion criteria
All studies included in the current meta-analysis met the following criteria: (1) evaluating the association between CLPTM1L rs402710 and rs401681 polymorphisms and the risk of all types of cancer; (2) case-control or cohort studies; (3) published in English; (4) including sufficient data for genotype frequencies; and (5) providing sufficient data to calculate ORs and 95% CIs. Exclusion criteria include: (1) insufficient raw data; (2) meta-analyses, reviews and comments; (3) duplicates of previous publications; and (4) case-only studies. Studies that were departure from HWE in controls were also excluded, unless further evidence indicated that another polymorphism was in agreement with HWE.
Data extraction
Two investigators extracted the following information from all the eligible studies independently: surname of the first author, publication year, country of origin, ethnicity, cancer type, control source, genotyping method, number of cases and controls, and genotype frequencies of cases and controls. Different ethnicities were stratified into Asian, African, and Caucasian. The source of controls was categorized as hospital-based and population-based studies. Any disagreements were discussed and resolved until the consensus was reached.
Statistical analysis
The strength of association between the CLPTM1L rs402710 and rs401681 polymorphisms and the risk of all types of cancer were assessed by the calculation of crude ORs and 95% CIs. Stratification analyses were further performed according to ethnicity, cancer type and control source. The pooled ORs and 95% CIs were estimated for rs402710 polymorphism under the homozygous (TT vs. CC), heterozygous (CT vs. CC), recessive (TT vs. CT+CC), and dominant (CT+TT vs. CC), as well as allele comparison model (T vs. C) and so was rs401681 polymorphism under the homozygous (TT vs. CC), heterozygous (CT vs. CC), recessive (TT vs. CT+CC), and dominant (CT+TT vs. CC), as well as allele comparison model (T vs. C).
The Chi square-based Q-test was applied to evaluate between-study heterogeneity. If the studies were found to have no heterogeneity (p > 0.10), the fixed-effects model (the Mantel-Haensze method) was used [68]. Otherwise, the random-effects model (the DerSimonian and Laird method) was adopted [69]. Furthermore, heterogeneity was also identified with the I2 test [70]. The higher values suggest the greater degree of heterogeneity. Thus the publication bias was quantified by Begg’s funnel plot and Egger’s linear regression test [71].
All the statistical analyses were conducted using STATA software (version 12.0, STATA Corporation, College Station, TX) and were two-sided, with P values less than 0.05 as statistically significant.
CONFLICTS OF INTEREST
The authors have declared that no competing interests exist.
REFERENCES
1. Bender E. Developing world: Global warning. Nature. 2014; 509:S64–5. https://doi.org/10.1038/509S64a.
2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016; 66:7–30. https://doi.org/10.3322/caac.21332.
3. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66:115–32. https://doi.org/10.3322/caac.21338.
4. McGuire S. World Cancer Report 2014. Geneva, Switzerland: World Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Adv Nutr. 2016; 7:418–9. https://doi.org/10.3945/an.116.012211.
5. Song M, Lee KM, Kang D. Breast cancer prevention based on gene-environment interaction. Mol Carcinog. 2011; 50:280–90. https://doi.org/10.1002/mc.20639.
6. Ponder BA. Cancer genetics. Nature. 2001; 411:336–41. https://doi.org/10.1038/35077207.
7. Cong YS, Wen J, Bacchetti S. The human telomerase catalytic subunit hTERT: organization of the gene and characterization of the promoter. Hum Mol Genet. 1999; 8:137–42.
8. Yamamoto K, Okamoto A, Isonishi S, Ochiai K, Ohtake Y. A novel gene, CRR9, which was up-regulated in CDDP-resistant ovarian tumor cell line, was associated with apoptosis. Biochem Biophys Res Commun. 2001; 280:1148–54. https://doi.org/10.1006/bbrc.2001.4250.
9. Wang Y, Broderick P, Webb E, Wu X, Vijayakrishnan J, Matakidou A, Qureshi M, Dong Q, Gu X, Chen WV, Spitz MR, Eisen T, Amos CI, et al. Common 5p15.33 and 6p21.33 variants influence lung cancer risk. Nat Genet. 2008; 40:1407–9. https://doi.org/10.1038/ng.273.
10. Petersen GM, Amundadottir L, Fuchs CS, Kraft P, Stolzenberg-Solomon RZ, Jacobs KB, Arslan AA, Bueno-de-Mesquita HB, Gallinger S, Gross M, Helzlsouer K, Holly EA, Jacobs EJ, et al. A genome-wide association study identifies pancreatic cancer susceptibility loci on chromosomes 13q22.1, 1q32.1 and 5p15.33. Nat Genet. 2010; 42:224–8. https://doi.org/10.1038/ng.522.
11. Ma Z, Hu Q, Chen Z, Tao S, Macnamara L, Kim ST, Tian L, Xu K, Ding Q, Zheng SL, Sun J, Xia G, Xu J. Systematic evaluation of bladder cancer risk-associated single-nucleotide polymorphisms in a Chinese population. Mol Carcinog. 2013; 52:916–21. https://doi.org/10.1002/mc.21932.
12. Jiang M, Wu H, Qin C. Genetic variant rs401681 at 5p15.33 modifies susceptibility to lung cancer but not esophageal squamous cell carcinoma. PLoS One. 2013; 8: e84277. https://doi.org/10.1371/journal.pone.0084277.
13. Zhao MM, Zhang Y, Shen L, Ren YW, Li XL, Yin ZH, Zhou BS. Genetic variations in TERT-CLPTM1L genes and risk of lung cancer in a Chinese population. Asian Pac J Cancer Prev. 2014; 15:2809–13.
14. Zhang Y, Zhao M, Shen L, Ren Y, Su L, Li X, Yin Z, Zhou B. Genetic polymorphisms of TERT and CLPTM1L and risk of lung cancer: a case-control study in northeast Chinese male population. Med Oncol. 2014; 31: 18. https://doi.org/10.1007/s12032-014-0018-4.
15. Ito H, McKay JD, Hosono S, Hida T, Yatabe Y, Mitsudomi T, Brennan P, Tanaka H, Matsuo K. Association between a genome-wide association study-identified locus and the risk of lung cancer in Japanese population. J Thorac Oncol. 2012; 7:790–8. https://doi.org/10.1097/JTO.0b013e3182475028.
16. Bae EY, Lee SY, Kang BK, Lee EJ, Choi YY, Kang HG, Choi JE, Jeon HS, Lee WK, Kam S, Shin KM, Jin G, Yoo SS, et al. Replication of results of genome-wide association studies on lung cancer susceptibility loci in a Korean population. Respirology. 2012; 17:699–706. https://doi.org/10.1111/j.1440-1843.2012.02165.x.
17. Chen XF, Cai S, Chen QG, Ni ZH, Tang JH, Xu DW, Wang XB. Multiple variants of TERT and CLPTM1L constitute risk factors for lung adenocarcinoma. Genet Mol Res. 2012; 11:370–8. https://doi.org/10.4238/2012.February.16.2.
18. Jin T, Li B, He N, Zhang Y, Xia R, Kang L, Ding Y, Yuan D. CLPTM1L polymorphism as a protective factor for lung cancer: a case-control study in southern Chinese population. Tumour Biol. 2016; 37:10533–8. https://doi.org/10.1007/s13277-016-4938-9.
19. Hsiung CA, Lan Q, Hong YC, Chen CJ, Hosgood HD, Chang IS, Chatterjee N, Brennan P, Wu C, Zheng W, Chang GC, Wu T, Park JY, et al. The 5p15.33 locus is associated with risk of lung adenocarcinoma in never-smoking females in Asia. PLoS Genet. 2010; 6. https://doi.org/10.1371/journal.pgen.1001051.
20. Zhou L, Fu G, Wei J, Shi J, Pan W, Ren Y, Xiong X, Xia J, Shen Y, Li H, Yang M. The identification of two regulatory ESCC susceptibility genetic variants in the TERT-CLPTM1L loci. Oncotarget. 2016; 7:5495–506. https://doi.org/10.18632/oncotarget.6747.
21. Zhang Y, Zhang X, Zhang H, Zhai Y, Wang Z, Li P, Yu L, Xia X, Zhang Y, Zeng Y, He F, Zhou G. Common variations in TERT-CLPTM1L locus are reproducibly associated with the risk of nasopharyngeal carcinoma in Chinese populations. Oncotarget. 2016; 7:759–70. https://doi.org/10.18632/oncotarget.6397.
22. Ge M, Shi M, An C, Yang W, Nie X, Zhang J, Lv Z, Li J, Zhou L, Du Z, Yang M. Functional evaluation of TERT-CLPTM1L genetic variants associated with susceptibility of papillary thyroid carcinoma. Sci Rep. 2016; 6: 26037. https://doi.org/10.1038/srep26037.
23. Liu SG, Ma L, Cen QH, Huang JS, Zhang JX, Zhang JJ. Association of genetic polymorphisms in TERT-CLPTM1L with lung cancer in a Chinese population. Genet Mol Res. 2015; 14:4469–76. https://doi.org/10.4238/2015.May.4.4.
24. Xun X, Wang H, Yang H, Wang B, Kang L, Jin T, Chen C. CLPTM1L genetic polymorphisms and interaction with smoking and alcohol drinking in lung cancer risk: a case-control study in the Han population from northwest China. Medicine (Baltimore). 2014; 93: e289. https://doi.org/10.1097/MD.0000000000000289.
25. Liang Y, Thakur A, Gao L, Wang T, Zhang S, Ren H, Meng J, Geng T, Jin T, Chen M. Correlation of CLPTM1L polymorphisms with lung cancer susceptibility and response to cisplatin-based chemotherapy in a Chinese Han population. Tumour Biol. 2014; 35:12075–82. https://doi.org/10.1007/s13277-014-2508-6.
26. Zhao Z, Li C, Yang L, Zhang X, Zhao X, Song X, Li X, Wang J, Qian J, Yang Y, Jin L, Chen H, Lu D. Significant association of 5p15.33 (TERT-CLPTM1L genes) with lung cancer in Chinese Han population. Exp Lung Res. 2013; 39:91–8. https://doi.org/10.3109/01902148.2012.762436.
27. Lu X, Ke J, Luo X, Zhu Y, Zou L, Li H, Zhu B, Xiong Z, Chen W, Deng L, Lou J, Wang X, Zhang Y, et al. The SNP rs402710 in 5p15.33 is associated with lung cancer risk: a replication study in Chinese population and a meta-analysis. PLoS One. 2013; 8: e76252. https://doi.org/10.1371/journal.pone.0076252.
28. Zheng Y, Ogundiran TO, Adebamowo C, Nathanson KL, Domchek SM, Rebbeck TR, Simon MS, John EM, Hennis A, Nemesure B, Wu SY, Leske MC, Ambs S, et al. Lack of association between common single nucleotide polymorphisms in the TERT-CLPTM1L locus and breast cancer in women of African ancestry. Breast Cancer Res Treat. 2012; 132:341–5. https://doi.org/10.1007/s10549-011-1890-7.
29. Zhao Y, Chen G, Song X, Chen H, Mao Y, Lu D. Fine-mapping of a region of chromosome 5p15.33 (TERT-CLPTM1L) suggests a novel locus in TERT and a CLPTM1L haplotype are associated with glioma susceptibility in a Chinese population. Int J Cancer. 2012; 131:1569–76. https://doi.org/10.1002/ijc.27417.
30. Wang S, Wu J, Hu L, Ding C, Kan Y, Shen Y, Chen X, Shen H, Guo X, Hu Z. Common genetic variants in TERT contribute to risk of cervical cancer in a Chinese population. Mol Carcinog. 2012; 51 Suppl 1: E118-22. https://doi.org/10.1002/mc.21872.
31. Ren Q, Xu B, Chen SQ, Yang Y, Wang CY, Wang YD, Wang XH, Hua LX, Chen M. A common genetic variant of 5p15.33 is associated with risk for prostate cancer in the Chinese population. Genet Mol Res. 2012; 11:1349–56. https://doi.org/10.4238/2012.May.15.5.
32. Pande M, Spitz MR, Wu X, Gorlov IP, Chen WV, Amos CI. Novel genetic variants in the chromosome 5p15.33 region associate with lung cancer risk. Carcinogenesis. 2011; 32:1493–9. https://doi.org/10.1093/carcin/bgr136.
33. Jaworowska E, Trubicka J, Lener MR, Masojc B, Zlowocka-Perlowska E, McKay JD, Renard H, Oszutowska D, Wokolorczyk D, Lubinski J, Grodzki T, Serwatowski P, Nej-Wolosiak K, et al. Smoking related cancers and loci at chromosomes 15q25, 5p15, 6p22.1 and 6p21.33 in the Polish population. PLoS One. 2011; 6: e25057. https://doi.org/10.1371/journal.pone.0025057.
34. Gago-Dominguez M, Jiang X, Conti DV, Castelao JE, Stern MC, Cortessis VK, Pike MC, Xiang YB, Gao YT, Yuan JM, Van Den Berg DJ. Genetic variations on chromosomes 5p15 and 15q25 and bladder cancer risk: findings from the Los Angeles-Shanghai bladder case-control study. Carcinogenesis. 2011; 32:197–202. https://doi.org/10.1093/carcin/bgq233.
35. Yoon KA, Park JH, Han J, Park S, Lee GK, Han JY, Zo JI, Kim J, Lee JE, Takahashi A, Kubo M, Nakamura Y, Lee JS. A genome-wide association study reveals susceptibility variants for non-small cell lung cancer in the Korean population. Hum Mol Genet. 2010; 19:4948–54. https://doi.org/10.1093/hmg/ddq421.
36. Truong T, Hung RJ, Amos CI, Wu X, Bickeboller H, Rosenberger A, Sauter W, Illig T, Wichmann HE, Risch A, Dienemann H, Kaaks R, Yang P, et al. Replication of lung cancer susceptibility loci at chromosomes 15q25, 5p15, and 6p21: a pooled analysis from the International Lung Cancer Consortium. J Natl Cancer Inst. 2010; 102:959–71. https://doi.org/10.1093/jnci/djq178.
37. Liu Z, Li G, Wei S, Niu J, Wang LE, Sturgis EM, Wei Q. Genetic variations in TERT-CLPTM1L genes and risk of squamous cell carcinoma of the head and neck. Carcinogenesis. 2010; 31:1977–81. https://doi.org/10.1093/carcin/bgq179.
38. Zienolddiny S, Skaug V, Landvik NE, Ryberg D, Phillips DH, Houlston R, Haugen A. The TERT-CLPTM1L lung cancer susceptibility variant associates with higher DNA adduct formation in the lung. Carcinogenesis. 2009; 30:1368–71. https://doi.org/10.1093/carcin/bgp131.
39. Jin G, Xu L, Shu Y, Tian T, Liang J, Xu Y, Wang F, Chen J, Dai J, Hu Z, Shen H. Common genetic variants on 5p15.33 contribute to risk of lung adenocarcinoma in a Chinese population. Carcinogenesis. 2009; 30:987–90. https://doi.org/10.1093/carcin/bgp090.
40. Gibbs DC, Orlow I, Kanetsky PA, Luo L, Kricker A, Armstrong BK, Anton-Culver H, Gruber SB, Marrett LD, Gallagher RP, Zanetti R, Rosso S, Dwyer T, et al. Inherited genetic variants associated with occurrence of multiple primary melanoma. Cancer Epidemiol Biomarkers Prev. 2015; 24:992–7. https://doi.org/10.1158/1055-9965.EPI-14-1426.
41. Campa D, Rizzato C, Stolzenberg-Solomon R, Pacetti P, Vodicka P, Cleary SP, Capurso G, Bueno-de-Mesquita HB, Werner J, Gazouli M, Butterbach K, Ivanauskas A, Giese N, et al. TERT gene harbors multiple variants associated with pancreatic cancer susceptibility. Int J Cancer. 2015; 137:2175–83. https://doi.org/10.1002/ijc.29590.
42. Zhang Y, Sun Y, Chen T, Hu H, Xie W, Qiao Z, Ding N, Xie L, Li S, Wang W, Xing C, Wang Y, Qie Y, et al. Genetic variations rs11892031 and rs401681 are associated with bladder cancer risk in a Chinese population. Int J Mol Sci. 2014; 15:19330–41. https://doi.org/10.3390/ijms151119330.
43. Yin J, Wang L, Zheng L, Wang X, Shi Y, Shao A, Ding G, Liu C, Chen S, Tang W, Gu H. TERT-CLPTM1L Rs401681 C>T polymorphism was associated with a decreased risk of esophageal cancer in a Chinese population. PLoS One. 2014; 9: e100667. https://doi.org/10.1371/journal.pone.0100667.
44. Su LY, Li XL, Shen L, Zhang Y, Zhao MM, Yin ZH, Su HY, Zhou BS. Polymorphisms of TERT and CLPTM1L and the risk of hepatocellular carcinoma in Chinese males. Asian Pac J Cancer Prev. 2014; 15:8197–201.
45. Llorca-Cardenosa MJ, Pena-Chilet M, Mayor M, Gomez-Fernandez C, Casado B, Martin-Gonzalez M, Carretero G, Lluch A, Martinez-Cadenas C, Ibarrola-Villava M, Ribas G. Long telomere length and a TERT-CLPTM1 locus polymorphism association with melanoma risk. Eur J Cancer. 2014; 50:3168–77. https://doi.org/10.1016/j.ejca.2014.09.017.
46. Liu C, Wang Y, Huang H, Wang C, Zhang H, Kong Y. Association between CLPTM1L-TERT rs401681 polymorphism and pancreatic cancer risk among Chinese Han population. Tumour Biol. 2014; 35:5453–7. https://doi.org/10.1007/s13277-014-1711-9.
47. Wang H, Zhao Y, Ma J, Zhang G, Mu Y, Qi G, Fang Z, Wang L, Fan Q, Ma Z. The genetic variant rs401681C/T is associated with the risk of non-small cell lung cancer in a Chinese mainland population. Genet Mol Res. 2013; 12:67–73. https://doi.org/10.4238/2013.January.22.5.
48. Sun Y, Zhang YJ, Kong XM. No association of XRCC1 and CLPTM1L polymorphisms with non-small cell lung cancer in a non-smoking Han Chinese population. Asian Pac J Cancer Prev. 2013; 14:5171–4.
49. Myneni AA, Chang SC, Niu R, Liu L, Ochs-Balcom HM, Li Y, Zhang C, Zhao B, Shi J, Han X, Li J, Su J, Cai L, et al. Genetic polymorphisms of TERT and CLPTM1L and risk of lung cancer--a case-control study in a Chinese population. Lung Cancer. 2013; 80:131–7. https://doi.org/10.1016/j.lungcan.2013.01.021.
50. Li C, Yin Z, Wu W, Li X, Ren Y, Zhou B. Genetic variations in TERT-CLPTM1L genes and risk of lung cancer in Chinese women nonsmokers. PLoS One. 2013; 8: e64988. https://doi.org/10.1371/journal.pone.0064988.
51. Ke J, Zhong R, Zhang T, Liu L, Rui R, Shen N, Sun Y, Cheng L, Miao XP. Replication study in Chinese population and meta-analysis supports association of the 5p15.33 locus with lung cancer. PLoS One. 2013; 8: e62485. https://doi.org/10.1371/journal.pone.0062485.
52. Willis JA, Olson SH, Orlow I, Mukherjee S, McWilliams RR, Kurtz RC, Klein RJ. A replication study and genome-wide scan of single-nucleotide polymorphisms associated with pancreatic cancer risk and overall survival. Clin Cancer Res. 2012; 18:3942–51. https://doi.org/10.1158/1078-0432.CCR-11-2856.
53. Lan Q, Hsiung CA, Matsuo K, Hong YC, Seow A, Wang Z, Hosgood HD 3rd, Chen K, Wang JC, Chatterjee N, Hu W, Wong MP, Zheng W, et al. Genome-wide association analysis identifies new lung cancer susceptibility loci in never-smoking women in Asia. Nat Genet. 2012; 44:1330–5. https://doi.org/10.1038/ng.2456.
54. Rizzato C, Campa D, Giese N, Werner J, Rachakonda PS, Kumar R, Schanne M, Greenhalf W, Costello E, Khaw KT, Key TJ, Siddiq A, Lorenzo-Bermejo J, et al. Pancreatic cancer susceptibility loci and their role in survival. PLoS One. 2011; 6: e27921. https://doi.org/10.1371/journal.pone.0027921.
55. Nan H, Qureshi AA, Prescott J, De Vivo I, Han J. Genetic variants in telomere-maintaining genes and skin cancer risk. Hum Genet. 2011; 129:247–53. https://doi.org/10.1007/s00439-010-0921-5.
56. Liu P, Vikis HG, Lu Y, Wang Y, Schwartz AG, Pinney SM, Yang P, de Andrade M, Gazdar A, Gaba C, Mandal D, Lee J, Kupert E, et al. Cumulative effect of multiple loci on genetic susceptibility to familial lung cancer. Cancer Epidemiol Biomarkers Prev. 2010; 19:517–24. https://doi.org/10.1158/1055-9965.EPI-09-0791.
57. Stacey SN, Sulem P, Masson G, Gudjonsson SA, Thorleifsson G, Jakobsdottir M, Sigurdsson A, Gudbjartsson DF, Sigurgeirsson B, Benediktsdottir KR, Thorisdottir K, Ragnarsson R, Scherer D, et al. New common variants affecting susceptibility to basal cell carcinoma. Nat Genet. 2009; 41:909–14. https://doi.org/10.1038/ng.412.
58. Rafnar T, Sulem P, Stacey SN, Geller F, Gudmundsson J, Sigurdsson A, Jakobsdottir M, Helgadottir H, Thorlacius S, Aben KK, Blondal T, Thorgeirsson TE, Thorleifsson G, et al. Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat Genet. 2009; 41:221–7. https://doi.org/10.1038/ng.296.
59. Asakura T, Imai A, Ohkubo-Uraoka N, Kuroda M, Iidaka Y, Uchida K, Shibasaki T, Ohkawa K. Relationship between expression of drug-resistance factors and drug sensitivity in normal human renal proximal tubular epithelial cells in comparison with renal cell carcinoma. Oncol Rep. 2005; 14:601–7.
60. Colombo J, Fachel AA, De Freitas Calmon M, Cury PM, Fukuyama EE, Tajara EH, Cordeiro JA, Verjovski-Almeida S, Reis EM, Rahal P. Gene expression profiling reveals molecular marker candidates of laryngeal squamous cell carcinoma. Oncol Rep. 2009; 21:649–63.
61. James MA, Vikis HG, Tate E, Rymaszewski AL, You M. CRR9/CLPTM1L regulates cell survival signaling and is required for Ras transformation and lung tumorigenesis. Cancer Res. 2014; 74:1116–27. https://doi.org/10.1158/0008-5472.CAN-13-1617.
62. Li C, Yin Z, Wu W, Li X, Zhou B. Genetic variants in TERT-CLPTM1L genetic region associated with several types of cancer: a meta-analysis. Gene. 2013; 526:390–9. https://doi.org/10.1016/j.gene.2013.05.003.
63. Wu H, Zhu R. Quantitative assessment of common genetic variants on chromosome 5p15 and lung cancer risk. Tumour Biol. 2014; 35:6055–63. https://doi.org/10.1007/s13277-014-1802-7.
64. Zhang XL, Zhang XJ, Zhang YM, Zhang Q, Cao CX, Gu DY, Gong YL, Chen JF, Tang CJ. Decreased risk of developing lung cancer in subjects carrying the CLPTM1L rs401681 (G > A) polymorphism: evidence from a meta-analysis. Genet Mol Res. 2014; 13:1373–82. https://doi.org/10.4238/2014.February.28.10.
65. Zhao DP, Yang CL, Zhou X, Ding JA, Jiang GN. Association between CLPTM1L polymorphisms (rs402710 and rs401681) and lung cancer susceptibility: evidence from 27 case-control studies. Mol Genet Genomics. 2014; 289:1001–12. https://doi.org/10.1007/s00438-014-0868-7.
66. Liu CL, Zang XX, Wang C, Kong YL, Zhang H, Zhang HY. Association between CLPTM1L-TERT rs401681 polymorphism and risk of pancreatic cancer: a meta-analysis. Clin Exp Med. 2015; 15:477–82. https://doi.org/10.1007/s10238-014-0316-3.
67. Yang X, Yang B, Li B, Liu Y. Association between TERT-CLPTM1L rs401681[C] allele and NMSC cancer risk: a meta-analysis including 45,184 subjects. Arch Dermatol Res. 2013; 305:49–52. https://doi.org/10.1007/s00403-012-1275-8.
68. Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959; 22:719–48.
69. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7:177–88.
70. Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002; 21:1539–58. https://doi.org/10.1002/sim.1186.
71. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629–34.

