INTRODUCTION
The 7th edition of the Union for International Cancer Control–American Joint Committee on Cancer (UICC-AJCC) tumor, node and metastasis (TNM) staging system moved gastroesophageal junction (GEJ) cancers from the gastric staging group to esophageal cancers [1]. This change was based on a review of esophageal cancer staging systems that evaluated a data set of 4,627 patients who were treated by surgical resection without previous or adjuvant therapy; the data set was assembled by the Worldwide Esophageal Cancer Collaboration and analyzed using Random Forest Analysis [2].
Although the application of the 7th UICC-AJCC staging system seems to result in a better prognostic stratification of overall survival compared with the 6th edition for esophageal cancer, according to some reports [3, 4], others have demonstrated that the 7th UICC-AJCC staging system does not demonstrate advantages in the assessment of patient prognosis [5]. The current 8th UICC-AJCC classification kept esophagus and esophagogastric tumors as a same tumor group [6].
According to the current UICC classification, GEJ cancers should more closely resemble esophageal tumors than gastric tumors, especially regarding their clinical and epidemiologic features. In addition, the staging system should favor the best decision regarding medical procedures by applying the rules for esophageal tumor management instead of those for gastric cancers [4]. Furthermore, this modification enhances the burden of esophageal cancer and do not increase the incidence of gastric cancer, thus providing a false perception of better gastric cancer control [7].
Nevertheless, clinical management algorithms and protocols are strongly influenced by the UICC-AJCC staging system, so the current classification has a tremendous impact on medical care. The number of retried lymph nodes necessary for adequate staging differs between gastric and esophageal tumors, and the specific lymph node stations that should be resected vary among these tumors. Thus, if a GEJ tumor is classified as a gastric cancer (by the older classification system), at least 16 lymph nodes are necessary to provide an adequate classification, whereas 12 lymph nodes are sufficient for esophageal cancers, according to the current UICC staging. Moreover, the proportion of positive to negative lymph nodes plays a role as a prognostic factor in gastric cancer. Given that fewer lymph nodes are necessary for staging, a negative impact for this benefit might occur [8].
An intensive debate is required to better define GEJ cancer as a specific entity or both a gastric and esophageal cancer or to keep these tumors classified as gastric or esophageal tumors [8].
The identification of the molecular characteristics of each tumor type can favor its classification and determine molecular targets [9–12].
The mRNAs expression pattern provides the specific tumor’s “molecular fingerprints”, and this information has been thoroughly investigated and reported for the majority of human cancers [13–15]. Moreover, the mRNA expression patterns can also differentiate subtypes or peculiar characteristics of a given tumor, such as aggressiveness or prognosis [15, 16]. It is necessary to consider that only transcribed mRNA will code for the proteins and enzymes that are typically present in tumors [17]. In some cases, these proteins and enzymes are used as tumor markers or even targets for therapy [18, 19]. Thus, it seems reasonable to use the mRNA profile to classify GEJ tumors as either gastric or esophageal cancers.
microRNAs are small non coding RNAs that function as post transcriptional regulators of gene expression. Their main function is to block translation by silencing mRNAs and impeding protein synthesis [20]. Due to been differentially expressed among tissues, microRNAs profiles can provide tissue signatures as previously demonstrated [21–24]. It means that looking at the microRNA is a feasible strategy to identify the origin of a given tissue, as a fingerprint.
There are plenty of reports implicating microRNA expression in human cancers [25–29]. Besides been able to characterize a given cancer, microRNAs profiles also work as markers of specific tumor behaviors as metastasis potential and prognosis [30–33]. As an advantage regarding, microRNAs are more stable and easily recovered from tissues specimens, including fixed samples [34–35]. Due to the specificity of expression in normal and cancer tissues, microRNA emerge as a powerful tool for characterizing GEJ tumors as either gastric or esophagus cancers.
The aims of this study are to enhance decision making regarding the best classification of GEJ cancers among the existing UICC categories. Thus, we provide the molecular fingerprints of these tumors and compared this information with gastric and esophageal molecular profiles to elucidate whether GEJs cluster in the gastric cancer or esophageal cancer groups based on mRNA and microRNA expression patterns because these information might represent a coherent tool for defining tumor identity.
RESULTS
After dividing gastric cancer samples between cardia (SCA) and non-cardia (SNCA) tumors and esophageal cancers between adenocarcinomas (EA) and squamous cell carcinomas (ES), the number of tumors was defined as shown in Table 1.
Table 1: Sample data obtained from TCGA, discriminated by cancer type
Cancer type |
mRNA samples |
microRNA samples |
|---|---|---|
Stomach non-cardia adenocarcinoma |
302 |
299 |
Cardia adenocarcinoma |
93 |
93 |
Esophageal adenocarcinoma |
89 |
87 |
Esophageal squamous cell carcinoma |
95 |
88 |
Colon adenocarcinoma |
521 |
459 |
Total |
1100 |
1026 |
Differential expression (D.E.) analysis was performed by grouping the samples as follows: i) (SNCA+SCA+EA) vs. ES; ii) EA vs. ES; iii) SNCA vs. EA; iv) (SNCA + SCA) vs. EA; and v) (SCA+EA) vs. SNCA. The number of differentially expressed mRNAs was 2366, 2550, 512, 375 and 186 for each analysis respectively. The number of differentially expressed microRNAs was 61, 60, 7, 5 and 2 for each analysis respectively (Table 2; Supplementary Figure 1; Supplementary Table 1). Esophageal squamous cell carcinoma clustered apart of the other tumors, whereas adenocarcinomas clustered together, independent of local origin (Figure 1A). The Heat Map of these analyses also demonstrates that squamous cell carcinoma belongs to a different group of tumors, whereas adenocarcinomas of the esophagus resemble adenocarcinomas of the cardia and non-cardia regions of the stomach (Figure 1B).
Table 2: Number of differentially expressed mRNAs and microRNAs in each analysis
Analysis |
Number of D. E. mRNAs |
Number of D.E. microRNAs |
|---|---|---|
(SNCA+SCA+EA) vs. ES |
2366 |
61 |
EA vs. ES |
2550 |
60 |
SNCA vs. EA |
512 |
7 |
(SNCA + SCA) vs. EA |
375 |
5 |
(SCA+EA) vs. SNCA |
186 |
2 |
Tissue discrimination was given as follows: stomach non-cardia adenocarcinoma (SNCA), stomach cardia adenocarcinoma (SCA), esophageal adenocarcinoma (EA) and esophageal squamous cell carcinoma (ES).
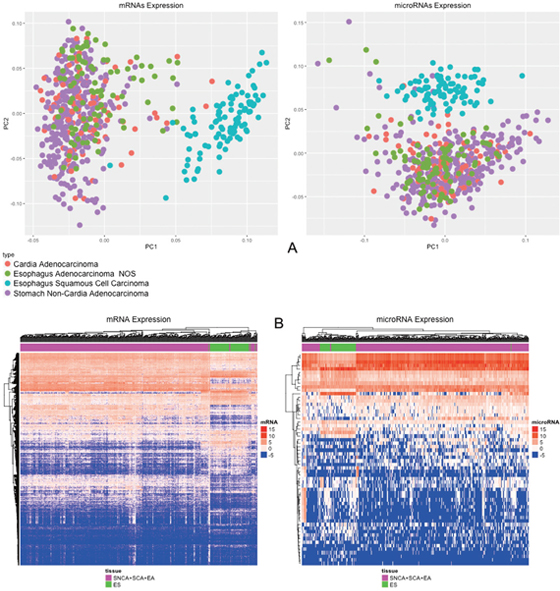
Figure 1: (A) PCA of 20,349 mRNAs and 910 microRNAs; (B) Heat Map of 2,366 mRNAs and 61 microRNAs differentially expressed (p-value < 0.05; |Log2(fold-change)| > 2) comparing non-cardia gastric adenocarcinomas, cardia adenocarcinomas and esophageal adenocarcinomas with esophageal squamous cell carcinomas.
The PCA (Principal Component Analysis) clearly categorized esophageal adenocarcinomas and esophageal squamous cell carcinomas as two different cancers (Figure 2A).
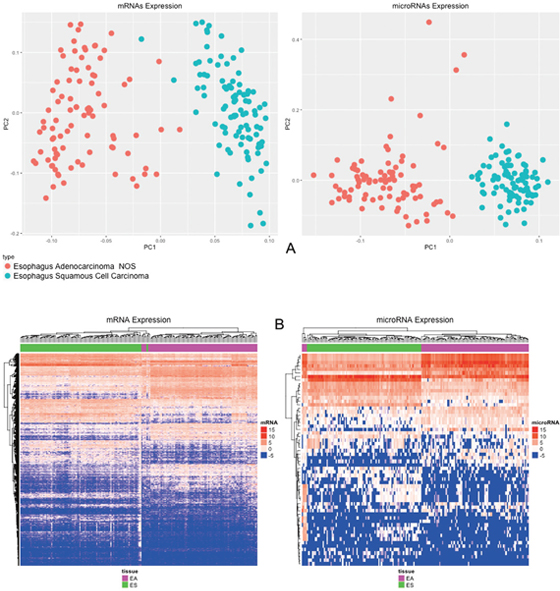
Figure 2: (A) PCA of 20,349 mRNAs and 910 microRNAs; (B) Heat Map of 2550 mRNAs and 60 microRNAs differentially expressed (p-value < 0.05; |Log2(fold-change)| > 2) comparing esophageal adenocarcinomas and esophageal squamous cell carcinomas.
The Heat Map of differentially expressed mRNAs and microRNAs (Figure 2B) strongly demonstrates that esophageal adenocarcinomas and esophageal squamous cell carcinomas are different entities that should not be classified together as a unique tumor, according to the mRNA expression patterns.
According to mRNA and microRNA expression profile, even distal tumors of the stomach are quite similar to adenocarcinomas of the lower esophagus, demonstrating that lower esophageal adenocarcinomas are more similar to gastric cancers than esophageal cancers. The PCA plot robustly represents this observation (Figure 3A).
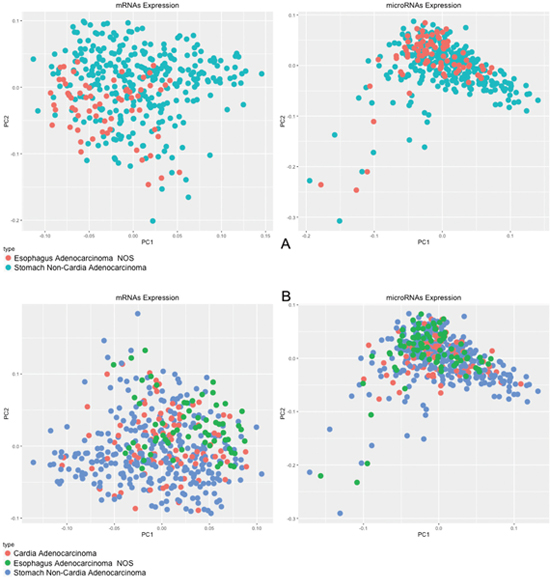
Figure 3: PCA of 20,349 mRNAs and 910 microRNAs. In (A) comparison between esophagus adenocarcinomas and non-cardia gastric adenocarcinomas; in (B) comparison between lower esophagus, gastric cardia region and non-cardia gastric adenocarcinomas.
By analyzing adenocarcinomas from the lower esophagus, adenocarcinomas of the gastric cardia region, and adenocarcinomas of the gastric non-cardia region, it was possible to identify that the mRNA and microRNA expression patterns of these tumors are quite similar (Figure 3B). Briefly, based on this molecular fingerprint, adenocarcinomas of the lower esophagus and adenocarcinomas of the cardia region of stomach could be classified as gastric cancers.
Finally, to determine whether the unique differences among these tumors are related to histological type, i.e., adenocarcinoma or squamous cell carcinoma, we also analyzed samples of colon adenocarcinomas and compared these samples with all of the tumors of esophagus and stomach regions previously analyzed.
The PCA images clearly demonstrated that adenocarcinomas of the colon are different from adenocarcinomas of the stomach and esophagus, thus strengthening the conclusions that GEJ tumors (including cardia cancers and esophageal adenocarcinomas from The Cancer Genome Atlas – TCGA) exhibit mRNA expression patterns similar to that of non-cardia stomach cancers and are extremely different from squamous cell esophagus carcinomas. This finding allows these tumors to be classified as gastric cancers instead of esophageal cancers. In addition, GEJ cancers were distinguished from both esophageal squamous cell carcinomas and colon adenocarcinomas (Figure 4).
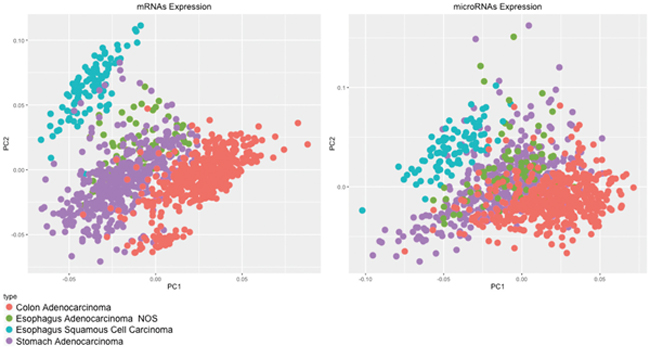
Figure 4: PCA of 20,349 mRNAs and 910 microRNAs from gastric cancer, esophageal squamous cell carcinoma and colon adenocarcinoma samples.
DISCUSSION
Staging GEJ cancer remains a challenge because a special group of tumors integrates this entity, which encompasses two peculiar organs. Nevertheless, the UICC 7th and 8th edition classified GEJ tumors as esophageal cancer. In addition, many clinicians and researchers include these cancers in the group of stomach cancers when reporting clinical and basic research data.
Different approaches have been applied to elucidate the best mode to stage and, consequently, define clinical management in GEJ cancers. Frequently, these approaches take advantage of institutional or even personal case series and compare the 6th and 7th UICC editions to determine which method better reproduces their expected results in terms of surgical and clinical outcomes [3, 7].
Comparing the 6th and 7th UICC staging systems in these case series resulted in conflicting findings due to different approaches and sample peculiarities. Thus, these studies were unable to shed light on this matter [3, 7].
Aiming at removing personal treatment and sample biases, we took advantage of the most important available cancer data bank, named TCGA. We also sought to perform an objective search of information: the genetic features of tumors.
The UICC staging does not differentiate esophageal adenocarcinomas from squamous cell esophageal carcinomas. Thus, for the initial analyses, every esophageal cancer that was available from TCGA, from both histological subtypes, was included.
Since the analyzed data was extracted from TCGA database, and not reviewed case by case by the pathologist author, the topographic accuracy of the tumors might have some misinterpretation, nevertheless the quality control of TCGA registries.
Despite tumor heterogeneity, it was strongly demonstrated that GEJ are much more similar to gastric cancers than they are to esophageal cancers. Using data from every expressed gene (mRNAs from 20,349 genes), the GEJ tumors clustered to gastric cancers instead of esophageal cancers according to the mRNA expression analyses. Regarding microRNAs (910 expressed microRNAs) the results were identical. This preliminary observation suggests that moving GEJ cancers from the stomach to esophageal staging group is not supported by molecular fingerprint analyses.
According to the differential expression analysis, there is a small number of mRNAs and microRNAs differentially expressed when comparing samples from non-cardia gastric adenocarcinomas, cardia adenocarcinomas and esophageal adenocarcinomas, indicating that these cancers have similar expression profile. Otherwise, when comparing these samples with esophageal squamous cell samples, the number of differentially expressed mRNAs and microRNAs increase greatly (Table 2; Supplementary Figure 1). These finds corroborates PCAs and support the suggestion that esophageal and cardia adenocarcinomas expression profiles are more similar to gastric adenocarcinoma than to esophageal cancers.
A possible major inconvenience of this first round of comparison is related to the eventual influence of squamous cell cancers in separating esophageal from cardia cancers. Regarding squamous cell cancers, it was strongly demonstrated, either by mRNAs or microRNAs expression patterns, that this histological type of tumor is significantly different from every group of adenocarcinomas, including non-cardia gastric adenocarcinomas, cardia adenocarcinomas and esophageal adenocarcinomas. Focusing exclusively on esophageal cancers, adenocarcinomas and squamous cell carcinomas of the esophagus are clearly different, thus raising a discussion regarding the traditional staging of these so different tumors as the same entity. In addition to their molecular fingerprints, their histology, epidemiology, risk factors, aggressiveness, and clinical management are completely different [36–38]; thus, this classification should be avoided in clinical practice.
In addition, when comparing esophageal adenocarcinomas to non-cardia gastric adenocarcinomas, no important discriminating differences were observed regarding mRNAs and microRNAs expression patterns. These data reinforce the notion that the placement of GEJ adenocarcinomas in the gastric cancer staging group is much less deleterious than classifying them as esophagus cancers.
Finally, assessing the global data, it could be concluded that the only real difference among these tumors involves the histological type, i.e., adenocarcinoma or squamous cell carcinoma, regardless of the location or organ of origin. To elucidate this notion, we compared the mRNA expression of non-cardia gastric adenocarcinomas, cardia adenocarcinomas and distal esophagus adenocarcinomas with adenocarcinomas of the colon, as well as the microRNAs expression of these tumors. According to the investigated molecular signatures, it was possible to clearly discriminate the organ of origin if colon adenocarcinoma or adenocarcinomas from stomach, esophagus and GEJ adenocarcinomas, thus demonstrating that the issues are much more complex than the tumor histologic type.
The provided data enable the authors to suggest the inclusion of an additional criterion to categorize these tumors – the molecular identity – by UICC. In special, in case of the discussed tumors, it should be reasonable to review the current classification of squamous cell and adenocarcinoma of the esophagus as a unique entity. Adenocarcinoma of GEJ might also be classified in the same gastric cancer staging group, or even as an additional specific tumor category.
In conclusion, by using robust molecular fingerprints of a large series of tumors from the most important available cancer data bank, it was strongly demonstrated that GEJ tumors more closely resemble gastric cancers than esophagus cancers, despite tumor heterogeneity.
MATERIALS AND METHODS
Cancer samples
To better estimate the similarities and differences in mRNA expressions patterns, we took advantage of TCGA and downloaded data (http://gdc.cancer.gov) regarding mRNA and microRNA expression, as well as tumor location and histologic types from gastric, esophagus, GEJ and colorectal cancers. Stomach cancers were categorized as cardia cancers (typically reported as GEJ tumors) and non-cardia cancers according to the available TCGA clinical data. Cases without clear definitions of tumor site or histological information were excluded.
Esophageal cancers were also classified as adenocarcinomas, which corresponds to tumors of the lower esophagus (usually included as GEJ tumors), and squamous cell carcinoma, as defined in TCGA data. Additionally, to determine whether mRNA and microRNA expression patterns differentiate adenocarcinomas of the stomach and GEJ from other organ adenocarcinomas instead of only being able to distinguish squamous cell carcinoma from adenocarcinoma, we included colon adenocarcinomas analyses.
Data analysis
The clinical and expression data used to perform the analysis were downloaded from TCGA. For the mRNA analysis, 395 stomach samples, 184 esophagus samples and 521 colon samples were used (1100 samples in total). For the microRNA analysis, 392 stomach samples, 175 esophagus samples and 459 colon samples were used (1026 samples in total), to avoid outliers, all samples used in microRNAs analysis were also present in mRNA analysis.
For each differential analysis, we excluded samples from non-cancer tissues and then excluded those mRNAs and microRNAs that exhibited no expression in all samples. The main reason for excluding non-cancer tissues adjacent to tumor samples was the possible misinterpretation related to the cancer field effect [39].
Both Principal Component Analysis (PCA) and the Heat Map plots were performed in R platform (http://www.rproject.org) using a Log2 transformation of RPKM (Reads Per Kilobase per Million mapped reads) normalized data. Differential expression analysis was also performed in R, using RAW data and the DESeq2 package [40, 41]. To avoid the multiple test problem, Benjamini-Hochberg False Discovery Rate (FDR) corrections were used. The mRNAs and microRNAs were tagged as differentially expressed if they met the following criteria: i) FDR adjusted p-value < 0.05 and ii) |Log2(fold-change)| > 2 (expressions greater or lower than four-fold change).
Abbreviations
UICC-AJCC: Union for International Cancer Control–American Joint Committee on Cancer; TNM: Tumor, Node and Metastasis staging system; GEJ: Gastroesophageal Junction; D.E.: Differential Expression; SCA: Stomach Cardia; SNCA: Stomach Non-Cardia; EA: Esophageal Adenocarcinoma; ES: Esophageal Squamous cell carcinoma; TCGA: The Cancer Genome Atlas; PCA: Principal Component Analysis; RPKM: Reads Per Kilobase per Million mapped reads; FDR: False Discovery Rate
Author contributions
Study concept and design and critical revision/supervision: Paulo Assumpção, Ândrea Ribeiro dos Santos, Sidney Santos, Gregory Riggins, André Khayat and Rommel Burbano.
Acquisition of data and interpretation: Williams Barra and Fabiano Moreira.
Drafting of the manuscript: Paulo Assumpção, Williams Barra, Fabiano Moreira and Aline Cruz.
Statistical analysis: Fabiano Moreira and Ney Santos.
Technical and material support: Taíssa Araújo, Danielle Calcagno, Rui Mascarenhas Jr, Geraldo Ishak and Samia Demachki.
ACKNOWLEDGMENTS
We acknowledge PROPESP/Universidade Federal do Pará for financial support and CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior) and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) for fellowship support.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
FUNDING
CAPES (Coordenação de Aperfeiçoamento de Pessoal do Nível Superior) and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) were the funding sources of this study that helped in purchasing equipaments.
REFERENCES
1. Sobin LH, Compton CC. TNM seventh edition: what’s new, what's changed. Cancer. 2010; 116: 5336-9. https://doi.org/10.1002/cncr.25537.
2. Rice TW, Rusch VW, Apperson-Hansen C, Allen MS, Chen LQ, Hunter JG, Kesler KA, Law S, Lerut TE, Reed CE, Salo JA, Scott WJ, Swisher SG, et al. Worldwide esophageal cancer collaboration. Dis Esophagus. 2009; 22: 1-8. https://doi.org/10.1111/j.1442-2050.2008.00901.x.
3. Talsma K, van Hagen P, Grotenhuis BA, Steyerberg EW, Tilanus HW, van Lanschot JJ, Wijnhoven BP. Comparison of the 6th and 7th editions of the UICC-AJCC TNM classification for esophageal cancer. Ann Surg Oncol. 2012; 19: 2142-8. https://doi.org/10.1245/s10434-012-2218-5.
4. Lü F, Xue Q, Shao K, Mao YS, Gao SG, Mu JW, Tan FW, Cheng GY, He J. Preliminary experience of clinical applications of the 7th UICC-AJCC TNM staging system of esophageal carcinoma. Zhonghua Zhong Liu Za Zhi. 2012; 34: 461-4. https://doi.org/10. 3760/cma.j.issn.0253-3766.2012.06.013.
5. Suh YS, Han DS, Kong SH, Lee HJ, Kim YT, Kim WH, Lee KU, Yang HK. Should adenocarcinoma of the esophagogastric junction be classified as esophageal cancer? A comparative analysis according to the seventh AJCC TNM classification. Ann Surg. 2012; 255: 908-15. https://doi.org/10.1097/SLA.0b013e31824beb95.
6. Amin MB, Greene FL, Edge SB, Compton CC, Gershenwald JE, Brookland RK, Meyer L, Gress DM, Byrd DR, Winchester DP. The eighth edition AJCC cancer staging manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J Clin. 2017; 67: 93-9. https://doi.org/10.3322/caac.21388.
7. Schuhmacher C, Novotny A, Feith M, Friess H. [The new TNM classification of tumors of the esophagogastric junction. Surgical consequences]. [Article in German]. Chirurg. 2012; 83: 23-30. https://doi.org/10.1007/s00104-011-2146-0.
8. Peyre CG, Hagen JA, DeMeester SR, Altorki NK, Ancona E, Griffin SM, Hölscher A, Lerut T, Law S, Rice TW, Ruol A, van Lanschot JJ, Wong J, et al. The number of lymph nodes removed predicts survival in esophageal cancer: an international study on the impact of extent of surgical resectiresection. Ann Surg. 2008; 248: 549-56. https://doi.org/10.1097/SLA.0b013e318188c474.
9. Paterson AL, Shannon NB, Lao-Sirieix P, Ong CA, Peters CJ, O'Donovan M, Fitzgerald RC. A systematic approach to therapeutic target selection in oesophago-gastric cancer. Gut. 2013; 62: 1415-24. https://doi.org/10.1136/gutjnl-2012-302039.
10. Sonoda A, Mukaisho K, Nakayama T, Diem VT, Hattori T, Andoh A, Fujiyama Y, Sugihara H. Genetic lineages of undifferentiated-type gastric carcinomas analysed by unsupervised clustering of genomic DNA microarray data. BMC Med Genomics. 2013; 6: 25. https://doi.org/10.1186/1755-8794-6-25.
11. Okines AF, Gonzalez de Castro D, Cunningham D, Chau I, Langley RE, Thompson LC, Stenning SP, Saffery C, Barbachano Y, Coxon F, Middleton G, Ferry D, Crosby T, et al. Biomarker analysis in oesophagogastric cancer: results from the REAL3 and TransMAGIC trials. Eur J Cancer. 2013; 49: 2116-25. https://doi.org/10.1016/j.ejca.2013.02.007.
12. Cui J, Yin Y, Ma Q, Wang G, Olman V, Zhang Y, Chou WC, Hong CS, Zhang C, Cao S, Mao X, Li Y, Qin S, et al. Comprehensive characterization of the genomic alterations in human gastric cancer. Int J Cancer. 2015; 137: 86-95. https://doi.org/10.1002/ijc.29352.
13. Ladanyi M, Chan WC, Triche TJ, Gerald WL. Expression profiling of human tumors: the end of surgical pathology? J Mol Diagn. 2001; 3: 92-7. https://doi.org/10.1016/S1525-1578(10)60657-5.
14. Erkan M, Weis N, Pan Z, Schwager C, Samkharadze T, Jiang X, Wirkner U, Giese NA, Ansorge W, Debus J, Huber PE, Friess H, Abdollahi A, et al. Organ-, inflammation- and cancer specific transcriptional fingerprints of pancreatic and hepatic stellate cells. Mol Cancer. 2010; 9: 88. https://doi.org/10.1186/1476-4598-9-88.
15. Olar A, Aldape KD. Using the molecular classification of glioblastoma to inform personalized treatment. J Pathol. 2014; 232: 165-77. https://doi.org/10.1002/path.4282.
16. Chanrion M, Fontaine H, Rodriguez C, Negre V, Bibeau F, Theillet C, Hénaut A, Darbon JM. A new molecular breast cancer subclass defined from a large scale real-time quantitative RT-PCR study. BMC Cancer. 2007; 7: 39. https://doi.org/10.1186/1471-2407-7-39.
17. Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. Molecular biology of the cell. 4th edn. 2002. New York: Garland Science. 1616 pp.
18. Mehta S, Shelling A, Muthukaruppan A, Lasham A, Blenkiron C, Laking G, Print C. Predictive and prognostic molecular markers for cancer medicine. Ther Adv Med Oncol. 2010; 2: 125-48. https://doi.org/10.1177/1758834009360519.
19. Baudino TA. Targeted cancer therapy: the next generation of cancer treatment. Curr Drug Discov Technol. 2015; 12: 3-20.
20. Macfarlane LA, Murphy PR. MicroRNA: biogenesis, function and role in cancer. Curr Genomics. 2010; 11: 537-61. https://doi.org/10.2174/138920210793175895.
21. Darnet S, Moreira FC, Hamoy IG, Burbano R, Khayat A, Cruz A, Magalhães L, Silva A, Santos S, Demachki S, Assumpção M, Assumpção P, Ribeiro-Dos-Santos Â. High-throughput sequencing of microRNAs reveals a tissue signature in gastric cancer and suggests novel potential biomarkers. Bioinform Biol Insights. 2015; 9: 1-8. https://doi.org/10.4137/BBI.S23773.
22. Hur K, Toiyama Y, Schetter AJ, Okugawa Y, Harris CC, Boland CR, Goel A. Identification of a metastasis-specific MicroRNA signature in human colorectal cancer. J Natl Cancer Inst. 2015; 107: dju492. https://doi.org/10.1093/jnci/dju492.
23. Nadal E, Truini A, Nakata A, Lin J, Reddy RM, Chang AC, Ramnath N, Gotoh N, Beer DG, Chen G. A novel serum 4-microRNA signature for lung cancer detection. Sci Rep. 2015; 5: 12464. https://doi.org/10.1038/srep12464.
24. Wotschofsky Z, Gummlich L, Liep J, Stephan C, Kilic E, Jung K, Billaud JN, Meyer HA. Integrated microRNA and mRNA signature associated with the transition from the locally confined to the metastasized clear cell renal cell carcinoma exemplified by miR-146-5p. PLoS One. 2016; 11: e0148746. https://doi.org/10.1371/journal.pone.0148746.
25. Xu C, Zheng Y, Lian D, Ye S, Yang J, Zeng Z. Analysis of microRNA expression profile identifies novel biomarkers for non-small cell lung cancer. Tumori. 2015; 101: 104-10. https://doi.org/10.5301/tj.5000224.
26. Shrestha S, Hsu SD, Huang WY, Huang HY, Chen W, Weng SL, Huang HD. A systematic review of microRNA expression profiling studies in human gastric cancer. Cancer Med. 2014; 3: 878-88. https://doi.org/10.1002/cam4.246.
27. Zhu J, Zheng Z, Wang J, Sun J, Wang P, Cheng X, Fu L, Zhang L, Wang Z, Li Z. Different microRNA expression profiles between human breast cancer tumors and serum. Front Genet. 2014; 5: 149. https://doi.org/10.3389/fgene.2014.00149.
28. Farazi TA, Hoell JI, Morozov P, Tuschl T. MicroRNAs in human cancer. Adv Exp Med Biol. 2013; 774: 1-20. https://doi.org/10.1007/978-94-007-5590-1_1.
29. Ribeiro-dos-Santos Â, Khayat AS, Silva A, Alencar DO, Lobato J, Luz L, Pinheiro DG, Varuzza L, Assumpção M, Assumpção P, Santos S, Zanette DL, Silva WA Jr, et al. Ultra-deep sequencing reveals the microRNA expression pattern of the human stomach. PLoS One. 2010; 5: e13205. https://doi.org/10.1371/journal.pone.0013205.
30. Chen J, Zhang K, Xu Y, Gao Y, Li C, Wang R, Chen L. The role of microRNA-26a in human cancer progression and clinical application. Tumour Biol. 2016; 37: 7095-108. https://doi.org/10.1007/s13277-016-5017-y.
31. Doldi V, Pennati M, Forte B, Gandellini P, Zaffaroni N. Dissecting the role of microRNAs in prostate cancer metastasis: implications for the design of novel therapeutic approaches. Cell Mol Life Sci. 2016; 73: 2531-42. https://doi.org/10.1007/s00018-016-2176-3.
32. Frediani JN, Fabbri M. Essential role of microRNAs in orchestrating the biology of the tumor microenvironment. Mol Cancer. 2016; 15: 42. https://doi.org/10.1186/s12943-016-0525-3.
33. Mutlu M, Raza U, Saatci Ö, Eyüpoğlu E, Yurdusev E, Şahin Ö. miR-200c: a versatile watchdog in cancer progression, EMT, and drug resistance. J Mol Med (Berl). 2016; 94: 629-44. https://doi.org/10.1007/s00109-016-1420-5.
34. Pritchard CC, Cheng HH, Tewari M. MicroRNA profiling: approaches and considerations. Nat Rev Genet. 2012; 13: 358-69. https://doi.org/doi:10.1038/nrg3198.
35. Doleshal M, Magotra AA, Choudhury B, Cannon BD, Labourier E, Szafranska AE. Evaluation and validation of total RNA extraction methods for microRNA expression analyses in formalin-fixed, paraffin-embedded tissues. J Mol Diagn. 2008; 10: 203-11. https://doi.org/10.2353/jmoldx.2008.070153.
36. Tustumi F, Takeda FR, Kimura CM, Sallum RA, Ribeiro U Jr, Cecconello I. Esophageal carcinoma: is squamous cell carcinoma different disease compared to adenocarcinoma? A transversal study in a quaternary high volume hospital in Brazil. Arq Gastroenterol. 2016; 53: 44-8. https://doi.org/10.1590/S0004-28032016000100009.
37. Mariette C, Finzi L, Piessen G, Van Seuningen I, Triboulet JP. Esophageal carcinoma: prognostic differences between squamous cell carcinoma and adenocarcinoma. World J Surg. 2005; 29: 39-45. https://doi.org/10.1007/s00268-004-7542-x.
38. Zhang Y. Epidemiology of esophageal cancer. World J Gastroenterol. 2013; 19: 5598-606. https://doi.org/10.3748/wjg.v19.i34.5598.
39. Assumpção MB, Moreira FC, Hamoy IG, Magalhães L, Vidal A, Pereira A, Burbano R, Khayat A, Silva A, Santos S, Demachki S, Ribeiro-Dos-Santos Â, Assumpção P. High-throughput microRNA sequencing reveals a field effect in gastric cancer and suggests an epigenetic network mechanism. Bioinform Biol Insights. 2015; 6: 111-7. https://doi.org/10.4137/BBI.S24066.
40. Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014; 15: 1-34. https://doi.org/10.1186/s13059-014-0550-8.
41. R Core Team R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2015. https://www.R-project.org/.

