INTRODUCTION
Breast carcinoma is the leading cause of cancer-related death among female patients with malignant tumors worldwide [1]. The morbidity and mortality rates of breast carcinoma have steadily increased since 1980s [2]. In the past few decades, great progress has been made in the treatment of breast cancer, particularly in the field of chemotherapy. However, treatment efficacy varies greatly due to the inherent genetic heterogeneity of individuals.
Efflux of agents from oncocyte by ATP-binding cassette (ABC) transporter is the most predominant and common mechanism of multiple drug resistance (MDR) among carcinoma [3]. In humans, 49 ABC genes have been reported and classified into seven different families [4, 5]. However, ABC gene subtypes involved in drug efflux from human cells do not belong to any particular family. For example, 12 transporters have been reported to regulate drug efflux; however, permeability glycoprotein (P-gp/ABCB1), multidrug resistance protein (MRP1/ABCC1), and breast cancer resistance protein (BCRP/ABCG2) are important for the efflux of a variety of drugs [3].
Human ABCB1 gene is located in chromosome region 7q21; this gene encodes a transmembrane transporters of 170 kDa that acts as an efflux pump for a variety of environmental carcinogens and antineoplastic drugs and plays an important role in resulting MDR of tumors [6-9]. Previous studies have shown that 66 coding single-nucleotide polymorphisms (SNPs) in ABCB1 gene have been identified, including 22 synonymous mutations and 44 non-synonymous mutations [10]. Several studies suggested that the expression level of ABCB1 gene was influenced by its polymorphism status [11, 12]. One of the most critical ABCB1 gene polymorphism is C3435T (rs1045642). Although it is a synonymous mutation, C3435T can alter the mRNA expression level of ABCB1, protein activity, and substrate specificity [13-15]. The variant allele frequency of C3435T is significantly different among various populations and races [16].
Human ABCG2 gene is located in chromosomal region 4q22 and encodes a 72-kDa membrane protein [17]. ABCG2 can transport numerous substrates, ranging from chemotherapeutics to carcinogenic xenobiotics [18-20]. Thus far, several SNPs in ABCG2 gene have been identified that can alter its expression and functionality [21]. One of the most important ABCG2 gene polymorphisms is C421A (rs2231142). The C421A SNP can lead to a glutamine-to-lysine amino acid substitution, resulting in decreased expression and activity of the ABCG2 protein [22-24]. Similar to C3435T, the mutation rate of C421A in ABCG2 gene is significantly different among different populations [25].
This study investigated the distribution of rs1045642 and rs2231142 polymorphisms in a Chinese Han breast cancer population who had been treated with post-operative chemotherapy. Correlation between the genetic polymorphism and breast cancer incidence, clinical features, and prognosis were explored.
RESULTS
Baseline characteristics of study subjects
The distributions of characteristics of the 100 breast cancer cases and 100 healthy controls are presented in Table 1. There were no significant differences in the distributions of age and menopausal state between cases and controls (p=0.571 and p= 0.48, respectively), and the average age was matched for breast cancer cases (range, 23–77 years; median, 50 years) and controls (range, 20–75 years; median, 50 years). Of the 100 breast cancer cases, 75 had invasive ductal carcinoma, 20 had invasive lobular carcinoma, and 5 had medullary carcinoma. Furthermore, 63 patients were diagnosed with stage II and 37 patients were diagnosed with stage III. Age at menarche of the patients was 12-18 years old. None of them had family history of breast cancer. Of the patients in our cohort, 70% / 60% / 40% were estrogen receptor (ER) /progesterone receptor (PR) / Her2 positive, respectively. IHC results showed 34% / 38% / 28% of patients with low / intermediate / high Ki67 expression. Lymph node metastasis was detected in 52% of the patients.
Table 1: Demographic and clinicopathological parameters of patients (n = 100)
Characteristic |
Case number |
Controls number |
P |
|---|---|---|---|
Age (years) |
|||
Median |
50 |
50 |
|
Range |
23-77 |
20-75 |
|
<50 |
44 |
50 |
0.479 |
≥50 |
56 |
50 |
|
Age at menarche (years) |
|||
≤14 |
68 |
||
>14 |
32 |
||
Menopausal state |
|||
Premenopausal |
48 |
54 |
0.48 |
Postmenopausal |
52 |
46 |
|
Reproductive history |
|||
One child |
35 |
||
Two children |
53 |
||
Three or more children |
12 |
||
Onset age (year,¯x±s) |
51.29±9.48 |
||
Family history |
|||
Yes |
0 |
||
No |
100 |
||
Five year survival rate |
65 |
||
Pathological location |
|||
Left breast |
53 |
||
Right breast |
47 |
||
Pathological type |
|||
Invasive ductal carcinoma |
75 |
||
Invasive lobular carcinoma |
20 |
||
Medullary carcinoma |
5 |
||
Clinical stage |
|||
II |
63 |
||
III |
37 |
||
Estrogen receptor |
|||
+ |
70 |
||
- |
30 |
||
Progesterone receptor |
|||
+ |
60 |
||
- |
40 |
||
Her2 |
|||
+ |
40 |
||
- |
60 |
||
Ki67 |
|||
Low (<14%) |
34 |
||
Intermediate (14%-30%) |
38 |
||
High (>30%) |
28 |
||
Lymph node metastasis |
|||
Node-positive |
52 |
||
Node-negative |
48 |
||
Surgery |
|||
Yes |
100 |
||
No |
0 |
||
Postoperative chemotherapy |
|||
Yes |
96 |
||
No |
4 |
Comparison of genotype distribution between patients with breast carcinoma and healthy controls
Overall genotype and allele frequencies for ABCB1 C3435T and ABCG2 C421A polymorphisms in cases and controls are listed in Table 2 and Supplementary Figure 1. The observed genotype frequency among individuals in the control group was in agreement with Hardy–Weinberg equilibrium. There was no significant difference in the distribution of three genotypes (CC, CT, and TT) of ABCB1 C3435T between the breast cancer cases and healthy controls (p>0.05). No significant difference was detected in the distribution of C and T alleles between the breast cancer patients and healthy controls (p>0.05). Moreover, we did not find significant difference in the distribution of three genotypes (CC, CA, and AA) of ABCG1 C421A between the breast cancer cases and healthy controls (p>0.05). There were no significant difference in the distribution of C and A alleles between the breast cancer patients and healthy controls (p>0.05).
Table 2: Genotype and allele frequencies of ABCB1 C3435T and ABCG2 C421A polymorphisms in normal tissues of breast cancer patients and controls
Variable |
No. of cases |
No. of controlsa |
P-valueb |
OR (95% CI)c |
|
|---|---|---|---|---|---|
ABCB1 |
Allele |
||||
C |
122 |
120 |
- |
- |
|
T |
78 |
80 |
0.919 |
1.043(0.698-1.557) |
|
Genotype |
|||||
CC |
40 |
35 |
- |
||
CT |
42 |
50 |
0.353 |
1.361(0.738-2.508) |
|
TT |
18 |
15 |
1 |
0.952(0.419-2.166) |
|
CT+TT |
60 |
65 |
0.559 |
1.238(0.698-2.197) |
|
ABCG2 |
Allele |
||||
C |
135 |
133 |
|||
A |
65 |
67 |
0.915 |
1.046 (0.69-1.587) |
|
Genotype |
|||||
CC |
47 |
46 |
- |
||
CA |
41 |
41 |
1 |
1.022 (0.564-1.85) |
|
AA |
12 |
13 |
1 |
1.107 (0.457-2.678) |
|
CA + AA |
53 |
54 |
1 |
1.041 (0.597-1.815) |
OR: odds ratio, CI: confidence interval
a The observed genotype frequency among individuals in the control group was in agreement with Hardy–Weinberg equilibrium. (p2 + 2pq + q2)= 1; χ2 = 0.174, p = 0.677 for ABCB1 (C3435T); χ2 = 0.636, p = 0.425 for ABCG2 (C421A).
b P values were calculated from two-sided chi-square tests for either genotype distribution or allele frequency
c OR and 95 % CI values were calculated by unconditional logistic regression adjusted for age and menopausal state.
Comparison of genotype distribution between breast cancer tissues and adjacent tissues
Overall genotype and allele frequencies for ABCB1 C3435T and ABCG2 C421A polymorphisms in cancer tissues and adjacent tissues are presented in Table 3. The observed genotype frequency among the para-carcinoma tissues was in agreement with Hardy–Weinberg equilibrium. There were no significant difference in the distribution of three genotypes (CC, CT, and TT) of ABCB1 C3435T between the carcinoma and para-carcinoma adjacent tissues (p>0.05). However, we found that 16% of carcinoma tissues had genetic mutations (Table 4). The mutation rates of CC, CT, and TT genotypes were 15.8%, 11.6%, and 26.3%, respectively. Although no significant difference was found in the distribution of three genotypes (CC, CA, and AA) of ABCG2 C421A between the cancerous tissues and adjacent tissues (p>0.05), 17% mutation rate was detected in the cancerous tissues. The mutation rates of CC, CA, and AA genotypes were 1.9%, 43.3%, and 22.2%, respectively.
Table 3: Genotype and allele frequencies of ABCB1 C3435T and ABCG2 C421A polymorphisms in cancer tissues and adjacent tissues
Variable |
No. of cancer tissue |
No. of adjacent tissuea |
P-valueb |
OR (95% CI)c |
|
|---|---|---|---|---|---|
ABCB1 |
Allele |
||||
C |
119 |
122 |
- |
- |
|
T |
81 |
78 |
0.838 |
0.939(0.629-1.402) |
|
Genotype |
|||||
CC |
38 |
40 |
- |
- |
|
CT |
43 |
42 |
0.876 |
0.928(0.502-1.716) |
|
TT |
19 |
18 |
0.843 |
0.9(0.411-1.969) |
|
CT+TT |
62 |
60 |
0.885 |
0.919(0.521-1.623) |
|
ABCG2 |
Allele |
||||
C |
134 |
135 |
- |
- |
|
A |
66 |
65 |
0.915 |
1.023 (0.674-1.553) |
|
Genotype |
|||||
CC |
52 |
47 |
|||
CA |
30 |
41 |
0.214 |
1.512 (0.818-2.795) |
|
AA |
18 |
12 |
0.534 |
0.738 (0.322-1.692) |
|
CA+AA |
48 |
53 |
0.572 |
1.222 (0.701-2.128) |
OR: odds ratio, CI: confidence interval
a The observed genotype frequency among cases in the adjacent tissues was in agreement with Hardy–Weinberg equilibrium. (p2 + 2pq + q2)= 1; χ2 = 1.375, p = 0.241 for ABCB1 (C3435T); χ2 = 0.429, p = 0.512 for ABCG2 (C421A).
b P values were calculated from two-sided chi-square tests for either genotype distribution or allele frequency
c OR and 95 % CI values were calculated by unconditional logistic regression adjusted for age and menopausal state.
Table 4: The mutation rates of ABCB1 C3435T and ABCG2 C421A in breast cancer patients
Adjacent tissue |
Cancer tissue |
Mutation rate |
||
|---|---|---|---|---|
ABCB1 |
CC |
CT |
4 |
15.8% |
TT |
2 |
|||
CT |
CC |
1 |
11.6% |
|
TT |
4 |
|||
TT |
CC |
2 |
26.3% |
|
CT |
3 |
|||
Total |
16 |
16% |
||
ABCG2 |
||||
CC |
AA |
1 |
1.9% |
|
CA |
AA |
9 |
43.3% |
|
CC |
4 |
|||
AA |
CA |
2 |
22.2% |
|
CC |
2 |
|||
Total |
17 |
17% |
Relation between genotype distribution and clinicopathological characteristics
Clinicopathological features of the patients were distinguished according to ABCB1 C3435T and ABCG2 C421A genotypes and are shown in Table 5. Chi-square tests or Fisher’s exact test was used to assess the effect of the SNPs on the clinicopathological characteristics of the 100 breast cancer patients. We found that there was no significant correlation between genotype distribution of ABCG2 and age at diagnosis, menopausal state, age at menarche, histology, clinical stage, lymph node metastasis, Ki67 expression level, ER status, PR status, or HER2 status. We also investigated the effect of ABCB1 C3435T genotype distribution on the above clinicopathological features, and only clinical stage and Ki67 expression level were significantly associated with ABCB1 C3435T genotype. Moreover, we detected that the distribution frequency of the ABCB1 C3435T CC genotype was lower among the cases diagnosed with stage II than stage III (p= 0.018 and OR (95 % CI) 0.34 (0.146–0.793)). Based on our research, compared to patients with intermediate/high expression of Ki67, patients with low expression of Ki67 showed a significant reduction in mutation rate of ABCB1 C3435T (p< 0.001 and OR (95 % CI) 4.656 (1.922–11.279)).
Table 5: Correlation of clinical characteristics of ABCB1 C3435T and ABCG2 C421A polymorphisms in patients with breast cancer
Characteristics |
ABCB1 C3435T |
ABCG2 C421A |
||||||
|---|---|---|---|---|---|---|---|---|
CC |
CT+TT |
Pa |
ORb |
CC |
CA+AA |
Pa |
ORb |
|
Age (year) |
||||||||
<50 |
16 |
28 |
0.765 |
0.883 |
23 |
21 |
0.961 |
1.02 |
≥50 |
22 |
34 |
29 |
27 |
||||
Menopausal state |
||||||||
Premenopausal |
18 |
30 |
0.921 |
0.96 |
25 |
23 |
0.987 |
1.006 |
Postmenopausal |
20 |
32 |
27 |
25 |
||||
Pathological type |
||||||||
IDC |
32 |
43 |
0.125 |
2.357 |
39 |
36 |
1 |
1 |
Others |
6 |
19 |
13 |
12 |
||||
Clinical stage |
||||||||
II |
18 |
45 |
0.018 |
0.34* |
35 |
28 |
0.41 |
1.471 |
III |
20 |
17 |
17 |
20 |
||||
Lymph node metastssis |
||||||||
Node-negative |
16 |
32 |
0.412 |
0.682 |
23 |
25 |
0.548 |
0.73 |
Node-positive |
22 |
30 |
29 |
23 |
||||
ER |
||||||||
- |
13 |
17 |
0.506 |
1.376 |
12 |
18 |
0.131 |
0.5 |
+ |
25 |
45 |
40 |
30 |
||||
PR |
||||||||
- |
15 |
25 |
1 |
0.965 |
18 |
22 |
0.309 |
0.626 |
+ |
23 |
37 |
34 |
26 |
||||
Her2 |
||||||||
- |
25 |
35 |
0.405 |
1.484 |
30 |
30 |
0.685 |
0.818 |
+ |
13 |
27 |
22 |
18 |
||||
Ki67 |
||||||||
Low (<14%) |
21 |
13 |
<0.001 |
4.656 |
14 |
20 |
0.12 |
0.516 |
Intermediate (14%-30%) and High (>30%) |
17 |
49 |
(1.922-11.279) |
38 |
28 |
(0.223-1.194) |
||
OR: odds ratio, CI: confidence interval
*: p<0.05
a P values were calculated from two-sided chi-square tests or Fisher’s exact test
b OR and 95 % CI values were calculated by unconditional logistic regression adjusted for age and menopausal state.
Association between ABCB1 and ABCG2 gene variants and patient survival
To investigate the association between ABCB1 C3435T and ABCG2 C421A gene polymorphisms and progression-free survival (PFS) and 5-year survival rate in breast carcinoma patients, the cases were followed up for 5 years. We observed that patients with TT genotype showed significantly longer PFS than patients harboring CC genotype in ABCB1 C3435T (p=0.03). There was no statistically significant difference in PFS between the patients with the CC genotype and those with the CT genotype in ABCB1 C3435T (p>0.05). Moreover, there was no significant association between the genotype of ABCB1 C3435T and the 5-year survival rate of breast cancer patients (Figure 1, p>0.05). We also detected that patients with CC genotype had significantly longer PFS than patients with 421AA genotype in ABCG2 (p=0.012). However, there was no significant difference in PFS between the patients with CC genotype and those harboring the CA genotype in ABCG2 C421A (p>0.05). There was no significant association between the polymorphism of ABCG2 C421A and the 5-year survival rate of breast cancer patients (Figure 1, p>0.05).
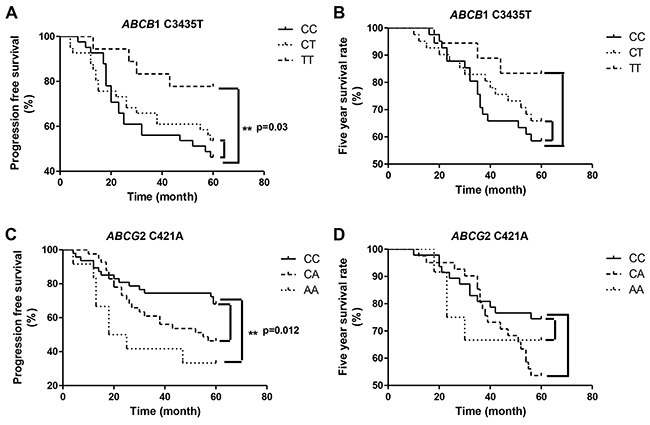
Figure 1: Association between ABCB1 and ABCG2 polymorphism and breast cancer patient survival (A) Association between ABCB1 C3435T polymorphism and progression-free survival. (B) Association between ABCB1 C3435T polymorphism and five year survival rate of the patients. (C) Association between ABCG2 C421A polymorphism and progression-free survival. (D) Association between ABCG2 C421A polymorphism and five year survival rate of the patients.
DISCUSSION
Gene polymorphism plays a vital role in human phenotypic variability including cancer susceptibility and patient response to therapy. In recent decades, considerable progress has been made in the study of the polymorphism of tumor-resistance genes. ABCB1 and ABCG2 have been extensively studied as important resistance genes. Previous studies have shown that the polymorphisms of ABCB1 and ABCG2 can alter the mRNA expression levels and protein activity [11, 12, 21]. The C3435T polymorphism in ABCB1 is a synonymous mutation that can alter its mRNA expression level, protein activity, and substrate specificity [13-15]. The C421A polymorphism in ABCG2 can lead to a glutamine-to-lysine amino acid substitution, resulting in the decrease of its expression level and protein activity [22-24]. In this study, we examined the genotype frequencies of ABCB1 C3435T and ABCG2 C421A distribution in Chinese female breast cancer patients and healthy controls.
The frequencies of the CC, CT, TT genotype of C3435T in the breast cancer cases were 40%, 42%, and 18%, and in the healthy controls, the frequencies were 35%, 50%, and 15%, respectively. Our results suggested that there was no significant association between ABCB1 C3435T polymorphism and the breast cancer susceptibility in Chinese women (p>0.05). Tatari et al. also reported that ABCB1 C3435T polymorphism was not associated with breast cancer susceptibility in Iran [26]. Gervasini et al. found no association between ABCB1 C3435T polymorphism and risk of lung cancer [27]. In contrast, Siegsmund et al. reported that in renal cell carcinoma patients, the frequency of the exon 26 C3435T allele was significantly higher than the normal population [28]. The study of Jamroziak et al. in acute lymphoblastic leukemia showed that the ABCB1 3435TT genotype was associated with the occurrence of ALL. Wu et al. found that the frequency of the homozygous variant TT genotype of C3435T in breast carcinoma patients was significantly higher than in controls [29]. The difference in tumor type and patient ethnicity may contribute to these inconsistent findings. The frequencies of the CC, CA, and AA genotype of C421A in the breast carcinoma patients were 47%, 41%, and 12%, while 46%, 41%, and 13% in healthy controls, respectively. However, our findings suggested that ABCG2 C421A polymorphism was not associated with the susceptibility to breast cancer in Chinese women. A number of studies indicated that ABCG2 C421A polymorphism was not associated with the susceptibility to prostate cancer [30], colorectal cancer [31, 32]. In contrast, some studies suggested that ABCG2 C421A polymorphism may be useful as a biomarker for the prediction of susceptibility to diffuse large B-cell lymphoma [33], lymphoma [34], and nonpapillary renal cell carcinoma [35]. Previous studies have shown that the ABCG2 C421A polymorphism varies widely among different tumor types and populations. Nonetheless, research on the correlation of ABCG2 gene polymorphism and breast cancer susceptibility is lacking, which this study attempts to address in a Chinese Han population.
We also detected ABCB1 C3435T and ABCG2 C421A genotypes in the cancerous and normal tissues from breast cancer patients. Interestingly, we found that the cancer tissue of 16% breast carcinoma patients harbored gene mutations in the ABCB1 C3435T loci. The mutation rate of the CC, CT, and TT genotype was 15.8%, 11.6%, and 26.3% respectively. We also found that 17% of carcinoma cases had gene mutations in ABCG2 C421A loci. The mutation rate of the CC, CA, and AA genotype was 1.9%, 43.3%, and 22.2% respectively. Therefore, genotype analysis of cancer tissue cannot be replaced by detecting the peripheral blood or normal tissue of patients.
This study analyzed the correlation between the clinicopathological features of breast cancer patients and ABCB1 C3435T and ABCG2 C421A gene polymorphisms. The frequency of 3435CC genotype in patients with stage III was significantly higher than patients with stage II. Compared to patients with intermediate/high expression of Ki67, patients with low expression of Ki67 got a significant higher frequency of 3435CC genotype. Other clinicopathological features in this study were not significantly related to ABCB1 C3435T polymorphisms. Turgut et al. [36], Wu et al.[9] and Macías-Gómezdid et al. [37] reported similar results. The correlation between the clinicopathological features of breast carcinoma in our study, according to ABCG2 C421A polymorphism revealed no significant association at this level. Our results showed that no significant association between ABCG2 C421A polymorphisms and clinicopathological features (age at diagnosis, menopausal state, age at menarche histology, clinical stage, lymph node metastasis, Ki67 status, ER status, PR status, HER2 status) of breast cancer patients. Our results are in agreements with those from Korenaga et al. [35]. We speculate that other factors may obscure the relationship between ABCG2 C421A polymorphisms and clinicopathological characteristics.
All the patients in this study received surgical treatment and postoperative adjuvant chemotherapy. We followed up patients for five years, and assessed their progression-free survival and 5-year survival rates. Our results showed that breast carcinoma patients with ABCB1 3435TT genotype had significantly longer PFS than those with CC genotype. Our results were consistent with the studies of Madrid-Paredes et al[38] and Wu et al[9]. However, the studies of Cizmarikova et al[39] and Ji et al[40] are inconsistent with our findings. Potentially, the difference in pathological stage and treatment regimen may explain the inconsistent results. We also analyzed the 5-year survival rate of the patients, and found that there was no significant correlation with the polymorphisms of ABCG1 C34535T and ABCG2 C421A.
In conclusion, this study found that ABCB1 C3435T and ABCG2 C421A genotypes were not significantly correlation with the susceptibility to breast carcinoma in a Chinese Han population. However, these two loci had a higher rate of mutation in breast cancer tissue. ABCB1 3435TT and ABCG2 421CC genotypes were significantly correlated with longer PFS of the breast cancer patients. But the result of multivariate Cox regression analysis suggested that they cannot be used as predictors for the PFS of breast cancer patients (Table 6). Here, we suggest that in detecting breast cancer resistance-related genes, samples should be selected from cancer tissue, and not peripheral blood or normal tissue. Meanwhile, a multi-gene joint analysis will be better.
Table 6: Univariate and multivariate analysis of PFS in breast cancer patients
Variables |
Univariate analyses |
Multivariate analyses |
||
|---|---|---|---|---|
Hazard ratio (95%CI) |
p-valuea |
Hazard ratiob (95%CI) |
p-valuea |
|
Age |
1.064(0.587-1.928) |
0.838 |
- |
- |
Lymph node metastssis |
0.852(0.473-1.535) |
0.594 |
- |
- |
Pathological type |
1.435(0.74-2.84) |
0.286 |
- |
- |
Clinical stage |
1.073(0.584-1.973) |
0.281 |
- |
- |
Menopausal state |
0.657 (0.342–1.324) |
0.246 |
- |
- |
ER |
2.136(1.103-4.136) |
0.024* |
1.787 (0.983-3.248) |
0.057 |
PR |
1.836(0.998-3.379) |
0.051 |
- |
- |
HER2 |
0.666(0.362-1.226) |
0.192 |
- |
- |
Ki67 |
0.999(0.54-1.847) |
0.997 |
||
ABCB1 C3435T |
2.124 (1.035-4.358) |
0.04* |
1.785 (0.753-4.231) |
0.188 |
ABCG2 C421A |
2.039 (1.129-3.681) |
0.018* |
0.577 (0.312-1.067) |
0.08 |
OR: odds ratio, CI: conWdence interval
*: p<0.05
a P values were calculated from two-sided chi-square tests or Fisher’s exact test
b OR and 95 % CI values were calculated by unconditional logistic regression adjusted for age and menopausal state.
MATERIALS AND METHODS
Study subjects
In this study, 100 breast cancer patients (female, median age: 50 years, range: 23–77 years) with incident breast carcinoma who were admitted to the 3201 hospital affiliated to Xi’an Jiao Tong University between 2010 and 2016 were enrolled. In addition 100 healthy control subjects (female, median age: 50 years, range: 20–75 years) were enrolled. All breast cancer patients underwent surgical treatment. Post-operative chemotherapy was based on a docetaxel and epirubicin regimen. Carcinoma and para-carcinoma tissues were collected from the patients and blood samples were collected from the healthy donors. Malignancy of the carcinoma tissues was confirmed by pathological analysis. The local ethics committee approved the research protocol for this study and all volunteers signed the study informed consent form.
Date collection
Two clinicians collected clinical features and treatment outcomes from medical records and followed patients on a regular basis. Complete information about the treatment was obtained from all 100 breast carcinoma patients.
DNA extraction
Paraffin-embedded tissue DNA extraction kit (TIANGEN, DP331, China) was used to extract DNA from carcinoma and para-carcinoma tissues. Blood genomic DNA extraction kit (TIANGEN, DP318, China) was used to extract DNA from blood samples. All protocols were in strict accordance with the manufacturers’ instructions.
Single nucleotide polymorphism analysis
The SNPs in ABCB1 C3435T and ABCG2 C421A were detected by Taqman method using the primer sequences and probes shown in Table 7. Each PCR reaction mixture contained 2x Hotstart Fluo-PCR mix 10 μl, sense primer 0.5 μl (10 μM), anti- sense primer 0.5μl (10 μM), probe 0.8 μl (10 μM), template DNA 2 μl (20 ng/μl), PCR-grade water 6.2 μl. The amplification consisted of an initial denaturation step for 4 min at 95°C followed by 40 cycles of melting 95°C for 15 s, and annealing/extension at 60°C for 60 s. PCR reactions were carried out in a Roche, LightCycler480 Real-time PCR System.
Table 7: Primers and probes used for taqman assays
SNP |
Primer |
Probe |
|---|---|---|
rs2231142 |
F:5’-ATGTTGTGATGGGCACTCTG-3’ |
P-A:TGCTGAGAACTTTAAGT |
R:5’-GTCATAGTTGTTGCAAGCCG-3’ |
P-C:TGCTGAGAACTGTAAGT |
|
rs1045642 |
F:5’-CCTATGGAGACAACAGCCG-3’ |
P-T:CCTCACAATCTCTTC |
R:5’-ACTCGATGAAGGCATGTATGTT-3’ |
P-C:CTCACGATCTCTTC |
Immunohistochemical analysis
A standard protocol was used for the immunohistochemistry (IHC) of the samples that were detected as breast cancer by hematoxylin and eosin staining. Briefly, formalin fixed, paraffin embedding, paraffin-embedded specimens, dewaxing to water, antigen repair, serum blocking, primary antibody incubation (ER antibody, abcam, ab27595; PR antibody, abcam, ab32063; HER2 antibody, abcam, ab16901; Ki67 antibody, abcam, ab8191), secondary antibody incubation, coloration.
Statistical analyses
All statistical analyses were carried out using Statistical Program for Social Sciences (SPSS) software 17.0 (SPSS Inc., USA). Hardy–Weinberg equilibrium and pairwise haplotype frequencies were estimated using the Hardy–Weinberg calculator and CubeX tools respectively, both provided by the Online Encyclopedia for Genetic Epidemiology studies. Statistical significance was set at p < 0.05 for all tests, and all tests were two-sided. Chi-square (Pearson’s χ2 test) or Fisher’s exact test was used to determine the differences in distributions of demographic, epidemiologic, and clinical variables between the two groups. Survival analysis was performed by Kaplan–Meier method and compared by log-rank test. Factors with significant influence on univariate analysis were further analyzed by multivariate Cox regression analysis. The minimum level of significance was established at p<0.05.
Compliance with ethical standards
The study was approved by the ethics committee of 3201 hospital affiliated to Xi’an Jiaotong University.
ACKNOWLEDGMENTS
This study was funded by State Key Laboratory of Tumor Biology (CBSKL201601) and Hanzhong 3201 Hospital Research Foundation (3201yk201550, 3201yk201607).
CONFLICTS OF INTEREST
All of the authors declared that they have no conflicts interest in relation to this study.
REFERENCES
1. Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011; 61:69–90.
2. Xing P, Dong H, Liu Q, Yao F, Xu Y, Chen B, Zheng X, Wu Y, Jin F, Li J. Impact of persistence on survival of patients with breast cancer treated with endocrine therapy in Northeast China: A prospective study. Oncotarget. 2017; 8:102499-102510. https://doi.org/10.18632/oncotarget.18454.
3. El-Awady R, Saleh E, Hashim A, Soliman N, Dallah A, Elrasheed A, Elakraa G. The Role of Eukaryotic and Prokaryotic ABC Transporter Family in Failure of Chemotherapy. Front Pharmacol. 2017; 7:535.
4. Dean M, Hamon Y, Chimini G. The human ATP-binding cassette (ABC) transporter superfamily. J Lipid Res. 2001; 42:1007–17.
5. Auner V, Sehouli J, Oskay-Oezcelik G, Horvat R, Speiser P, Zeillinger R. ABC transporter gene expression in benign and malignant ovarian tissue. Gynecol Oncol. 2010; 117:198–201.
6. Kerb R. Implications of genetic polymorphisms in drug transporters for pharmacotherapy. Cancer Lett. 2006; 234:4–33.
7. Ueda K, Clark DP, Chen CJ, Roninson IB, Gottesman MM, Pastan I. The human multidrug resistance (mdr1) gene. cDNA cloning and transcription initiation. J Biol Chem. 1987; 262:505–08.
8. Zhou SF. Structure, function and regulation of P-glycoprotein and its clinical relevance in drug disposition. Xenobiotica. 2008; 38:802–32.
9. Wu H, Kang H, Liu Y, Tong W, Liu D, Yang X, Lian M, Yao W, Zhao H, Huang D, Sha X, Wang E, Wei M. Roles of ABCB1 gene polymorphisms and haplotype in susceptibility to breast carcinoma risk and clinical outcomes. J Cancer Res Clin Oncol. 2012; 138:1449–62.
10. Wolf SJ, Bachtiar M, Wang J, Sim TS, Chong SS, Lee CG. An update on ABCB1 pharmacogenetics: insights from a 3D model into the location and evolutionary conservation of residues corresponding to SNPs associated with drug pharmacokinetics. Pharmacogenomics J. 2011; 11:315–25.
11. Ieiri I, Takane H, Hirota T, Otsubo K, Higuchi S. Genetic polymorphisms of drug transporters: pharmacokinetic and pharmacodynamic consequences in pharmacotherapy. Expert Opin Drug Metab Toxicol. 2006; 2:651–74.
12. Hoffmeyer S, Burk O, von Richter O, Arnold HP, Brockmöller J, Johne A, Cascorbi I, Gerloff T, Roots I, Eichelbaum M, Brinkmann U. Functional polymorphisms of the human multidrug-resistance gene: multiple sequence variations and correlation of one allele with P-glycoprotein expression and activity in vivo. Proc Natl Acad Sci USA. 2000; 97:3473–78.
13. Hemauer SJ, Nanovskaya TN, Abdel-Rahman SZ, Patrikeeva SL, Hankins GD, Ahmed MS. Modulation of human placental P-glycoprotein expression and activity by MDR1 gene polymorphisms. Biochem Pharmacol. 2010; 79:921–25.
14. Hutson JR, Koren G, Matthews SG. Placental P-glycoprotein and breast cancer resistance protein: influence of polymorphisms on fetal drug exposure and physiology. Placenta. 2010; 31:351–57.
15. Kimchi-Sarfaty C, Oh JM, Kim IW, Sauna ZE, Calcagno AM, Ambudkar SV, Gottesman MM. A "silent" polymorphism in the MDR1 gene changes substrate specificity. Science. 2007; 315:525–28.
16. Tulsyan S, Mittal RD, Mittal B. The effect of ABCB1 polymorphisms on the outcome of breast cancer treatment. Pharmgenomics Pers Med. 2016; 9:47–58.
17. Bailey-Dell KJ, Hassel B, Doyle LA, Ross DD. Promoter characterization and genomic organization of the human breast cancer resistance protein (ATP-binding cassette transporter G2) gene. Biochim Biophys Acta. 2001; 1520:234–41.
18. Mao Q, Unadkat JD. Role of the breast cancer resistance protein (ABCG2) in drug transport. AAPS J. 2005; 7:E118–33.
19. Krishnamurthy P, Schuetz JD. Role of ABCG2/BCRP in biology and medicine. Annu Rev Pharmacol Toxicol. 2006; 46:381–410.
20. Huss WJ, Gray DR, Greenberg NM, Mohler JL, Smith GJ. Breast cancer resistance protein-mediated efflux of androgen in putative benign and malignant prostate stem cells. Cancer Res. 2005; 65:6640–50.
21. Wang C, Xie L, Li H, Li Y, Mu D, Zhou R, Liu R, Zhou K, Hua Y. Associations between ABCG2 gene polymorphisms and isolated septal defects in a Han Chinese population. DNA Cell Biol. 2014; 33:689–98.
22. Kobayashi D, Ieiri I, Hirota T, Takane H, Maegawa S, Kigawa J, Suzuki H, Nanba E, Oshimura M, Terakawa N, Otsubo K, Mine K, Sugiyama Y. Functional assessment of ABCG2 (BCRP) gene polymorphisms to protein expression in human placenta. Drug Metab Dispos. 2005; 33:94–101.
23. Imai Y, Nakane M, Kage K, Tsukahara S, Ishikawa E, Tsuruo T, Miki Y, Sugimoto Y. C421A polymorphism in the human breast cancer resistance protein gene is associated with low expression of Q141K protein and low-level drug resistance. Mol Cancer Ther. 2002; 1:611–16.
24. Mizuarai S, Aozasa N, Kotani H. Single nucleotide polymorphisms result in impaired membrane localization and reduced atpase activity in multidrug transporter ABCG2. Int J Cancer. 2004; 109:238–46.
25. Chen P, Zhao L, Zou P, Xu H, Lu A, Zhao P. The contribution of the ABCG2 C421A polymorphism to cancer susceptibility: a meta-analysis of the current literature. BMC Cancer. 2012; 12:383.
26. Tatari F, Salek R, Mosaffa F, Khedri A, Behravan J. Association of C3435T single-nucleotide polymorphism of MDR1 gene with breast cancer in an Iranian population. DNA Cell Biol. 2009; 28:259–63.
27. Gervasini G, Carrillo JA, Garcia M, San Jose C, Cabanillas A, Benitez J. Adenosine triphosphate-binding cassette B1 (ABCB1) (multidrug resistance 1) G2677T/A gene polymorphism is associated with high risk of lung cancer. Cancer. 2006; 107:2850–57.
28. Siegsmund M, Brinkmann U, Scháffeler E, Weirich G, Schwab M, Eichelbaum M, Fritz P, Burk O, Decker J, Alken P, Rothenpieler U, Kerb R, Hoffmeyer S, Brauch H. Association of the P-glycoprotein transporter MDR1(C3435T) polymorphism with the susceptibility to renal epithelial tumors. J Am Soc Nephrol. 2002; 13:1847–54.
29. Jamroziak K, Młynarski W, Balcerczak E, Mistygacz M, Trelinska J, Mirowski M, Bodalski J, Robak T. Functional C3435T polymorphism of MDR1 gene: an impact on genetic susceptibility and clinical outcome of childhood acute lymphoblastic leukemia. Eur J Haematol. 2004; 72:314–21.
30. Gardner ER, Ahlers CM, Shukla S, Sissung TM, Ockers SB, Price DK, Hamada A, Robey RW, Steinberg SM, Ambudkar SV, Dahut WL, Figg WD. Association of the ABCG2 C421A polymorphism with prostate cancer risk and survival. BJU Int. 2008; 102:1694–99.
31. Andersen V, Ostergaard M, Christensen J, Overvad K, Tjønneland A, Vogel U. Polymorphisms in the xenobiotic transporter Multidrug Resistance 1 (MDR1) and interaction with meat intake in relation to risk of colorectal cancer in a Danish prospective case-cohort study. BMC Cancer. 2009; 9:407.
32. Campa D, Pardini B, Naccarati A, Vodickova L, Novotny J, Försti A, Hemminki K, Barale R, Vodicka P, Canzian F. A gene-wide investigation on polymorphisms in the ABCG2/BRCP transporter and susceptibility to colorectal cancer. Mutat Res. 2008; 645:56–60.
33. Hu LL, Wang XX, Chen X, Chang J, Li C, Zhang Y, Yang J, Jiang W, Zhuang SM. BCRP gene polymorphisms are associated with susceptibility and survival of diffuse large B-cell lymphoma. Carcinogenesis. 2007; 28:1740–44.
34. Campa D, Butterbach K, Slager SL, Skibola CF, de Sanjosé S, Benavente Y, Becker N, Foretova L, Maynadie M, Cocco P, Staines A, Kaaks R, Boffetta P, et al, and GEC Consortium. A comprehensive study of polymorphisms in the ABCB1, ABCC2, ABCG2, NR1I2 genes and lymphoma risk. Int J Cancer. 2012; 131:803–12.
35. Korenaga Y, Naito K, Okayama N, Hirata H, Suehiro Y, Hamanaka Y, Matsuyama H, Hinoda Y. Association of the BCRP C421A polymorphism with nonpapillary renal cell carcinoma. Int J Cancer. 2005; 117:431–34.
36. Turgut S, Yaren A, Kursunluoglu R, Turgut G. MDR1 C3435T polymorphism in patients with breast cancer. Arch Med Res. 2007; 38:539–44.
37. Macías-Gómez NM, Gutiérrez-Angulo M, Leal-Ugarte E, Ramírez-Reyes L, Peregrina-Sandoval J, Meza-Espinoza JP, Ramos Solano F, de la Luz Ayala-Madrigal M, Santoyo Telles F. MDR1 C3435T polymorphism in Mexican patients with breast cancer. Genet Mol Res. 2014; 13:5018–24.
38. Madrid-Paredes A, Cañadas-Garre M, Sánchez-Pozo A, Segura-Pérez AM, Chamorro-Santos C, Vergara-Alcaide E, Castillo-Portellano L, Calleja-Hernández MÁ. ABCB1 C3435T gene polymorphism as a potential biomarker of clinical outcomes in HER2-positive breast cancer patients. Pharmacol Res. 2016; 108:111–18.
39. Cizmarikova M, Wagnerova M, Schonova L, Habalova V, Kohut A, Linkova A, Sarissky M, Mojzis J, Mirossay L, Mirossay A. MDR1 (C3435T) polymorphism: relation to the risk of breast cancer and therapeutic outcome. Pharmacogenomics J. 2010; 10:62–69.
40. Ji M, Tang J, Zhao J, Xu B, Qin J, Lu J. Polymorphisms in genes involved in drug detoxification and clinical outcomes of anthracycline-based neoadjuvant chemotherapy in Chinese Han breast cancer patients. Cancer Biol Ther. 2012; 13:264–71.

