INTRODUCTION
Among pancreatic tumors, solid pseudopapillary neoplasm (SPN) is very rare and comprises only 1–3% of primary pancreatic tumors. In spite of its clinical rarity, the clinical and pathologic findings are well recognized. SPN is composed of poorly cohesive monomorphic epithelial cells that form solid, pseudopapillary structures, and it occurs predominantly in young women [1].
SPNs are genetically characterized by somatic mutations in exon 3 of CTNNB1, which encodes β-catenin [2, 3]. In addition, the molecular regulatory networks of SPN have recently been well-characterized [4-7]. Through an integrative analysis of mRNA, microRNA, and proteome expression profiles, we have identified many (> 1,000) SPN-specific upregulated genes, specifically activated Wnt/β-catenin, Hedgehog, and androgen receptor (AR) signaling pathways [6, 8].
However, there are several unresolved issues in SPNs. First, putative diagnostic immunohistochemical markers are necessary. The pathologic diagnosis of SPNs is usually made using a combination of histologic and immunohistochemical markers, including beta-catenin, CD10, chromogranin, and E-cadherin [1, 9, 10]. Diffuse cytoplasmic and nuclear localization of β-catenin is almost always found in SPN, and these findings are helpful in the diagnosis of SPNs. Nevertheless, some SPNs are histologically and immunohistochemically similar to pancreatic neuroendocrine tumors (NETs), and nuclear localization of β-catenin can also be found in some pancreatic ductal adenocarcinomas (PDCs) and NETs. Second, there are still no definite prognostic and diagnostic criteria for predicting the malignant behavior of SPNs. Although most SPNs have low malignant potential and can be cured using surgical resection [11-13], it has been reported that up to 20% of SPNs showed recurrence or distant metastasis [11-16].
In this study, we selected several immunohistochemical markers from the SPN-specific up-regulated genes and validated their diagnostic usability in SPNs. Additionally, we investigated clinicopathologic parameters associated with the malignant behavior of SPNs.
RESULTS
Clinicopathologic characteristics of SPN
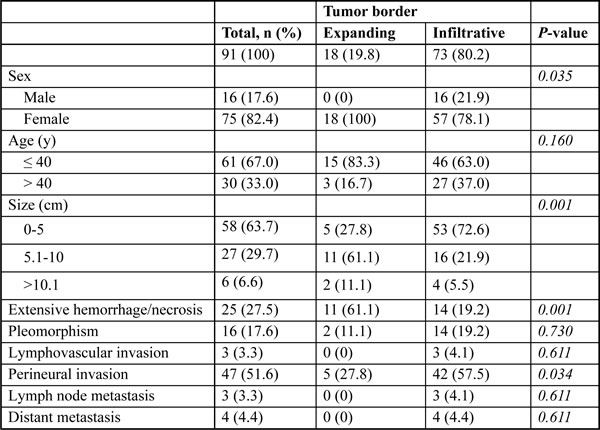
The basic clinicopathologic findings are listed in Table 1 and Figure 1. Sixty-one patients (67.0%) were ≤ 40 years old (median, 34 years old; range, 8–67 years), with a predominance of women (75, 82.4%). The tumor size varied from 1 to 16 cm (average, 4.7 cm). Six (6.6%) tumors measured more than 10 cm. More than 80% of the tumors had at least a focally infiltrative border (73; 80.2%, Figure 1A) and 40 (44%) showed peripancreatic adipose tissue invasion. Cellular pleomorphism (Figure 1C) was observed in 6 (6.6%) cases and mitoses were found in up to 2/10 high-power fields (HPFs) in 4 (4.3%) cases. Calcification and extensive hemorrhage/necrosis were present in 38 (43.6%) and 25 (27.5%) of tumors, respectively. Small cytoplasmic hyaline globules, foamy histiocytes, cholesterol clefts, and myxoid stromal changes were also observed. Three cases (3.3%) showed lymphovascular invasion (Figure 1D) and lymph nodes metastasis, respectively. Perineural invasion (Figure 1E) was a relatively frequent finding in 47 (51.6%) cases. Distant metastasis occurred in 4 (4.4%, Figure 1F) patients and recurrence developed in 3 (3.3%) patients.
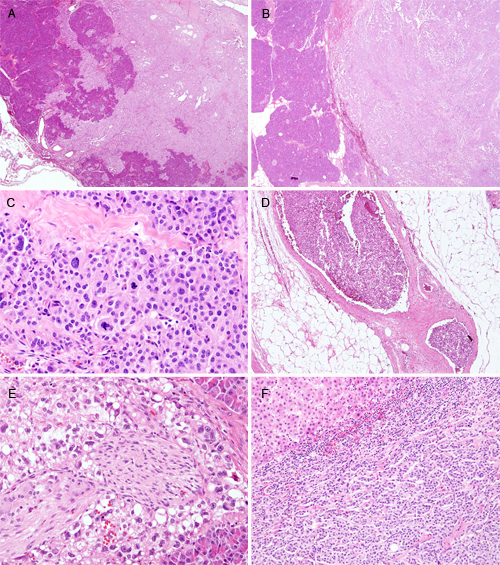
Figure 1: Histologic features (H & E stain) of SPNs. A. Infiltrative and B. expanding border of tumor (A and B, each original magnification x12.5). C. Pleomorphism and mitosis (original magnification x100). D. Lymphovascular invasion (original magnification x40). E. Perineural invasion (original magnification x100). F. Liver metastasis (original magnification x40).
The rim of most SPNs is known to be infiltrative. However, since some SPNs show totally expanding borders, we examined whether clinicopathologic parameters differ depending on whether the margin of tumor invades surrounding parenchyma. When the border of SPNs was divided into at least focally infiltrative pattern and totally expanding pattern, male sex (p = 0.035), smaller tumor size (p < 0.001), non-extensive hemorrhage and/or necrosis (p = 0.001), and frequent perineural invasion (p = 0.034) were found in the infiltrative SPNs (Table 1). We also found that SPNs in male patients had a lower incidence of hemorrhage and/or necrosis, more frequent infiltrative tumor borders, and peripancreatic invasion (Supplementary Table 2). They also showed a tendency toward more frequent lymph node metastasis (p = 0.079).
Table 1: Clinicopathologic characteristics of 91 SPNs
Immunohistochemical expression in SPN, PDC, and NET
The results of the immunohistochemical analysis of 91 SPNs, 51 PDCs, and 48 NETs are shown in Table 2. Beta-catenin showed diffuse membranous expression in normal pancreatic tissue and AR was expressed in some of the normal pancreatic ductal epithelium. FUS and WIF-1 showed occasional staining in the cytoplasm of pancreatic ductal cells and acinar cells. LEF1 and TFE3 were not expressed in normal pancreatic tissue.
Table 2: Expression pattern of 6 markers in SPN
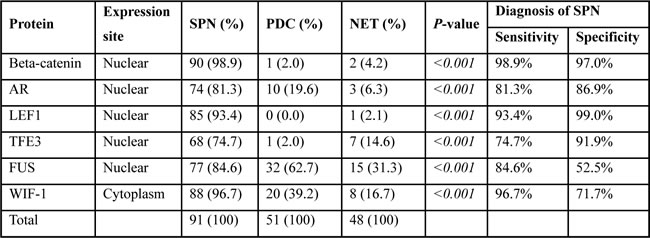
In the SPNs, nuclear expressions of β-catenin (98.9%), AR (81.3%), LEF1 (93.4%), TFE3 (74.7%), and FUS (84.6%), and cytoplasmic expression of WIF-1 (96.7%) were noted (Figure 2, p < 0.001). In contrast, PDCs and NETs showed no or occasional expression. Although some markers were positive in both SPNs and/or PDCs and NETs, there was no case showing strong expression of these markers in PDCs and NETs. Two of the 3 cases that had undergone decalcification during processing showed no expression of any of the markers, except for WIF-1 (weak) and β-catenin (moderate).
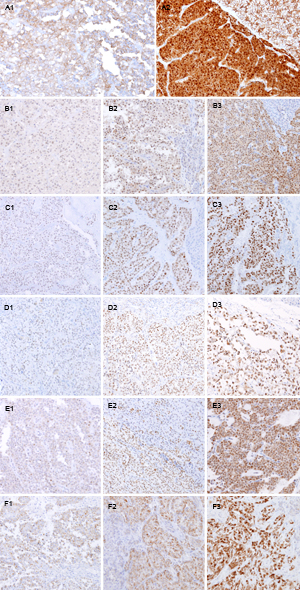
Figure 2: Representative immunohistochemical results of SPNs. Beta-catenin (A1, negative and A2, strong positive), AR (B1-B3), LEF1 (C1-C3), TFE3 (D1-D3), and FUS (E1-E3) show nuclear expression. WIF-1 (F1-F3) show cytoplasmic expression. Sub-number indicates weak (1), moderate (2), and strong (3), respectively. Original magnification x100.
After evaluating the diagnostic significance of these markers in pancreatic tumors, we described the sensitivity and specificity in Table 2 and Figure 3. Based on these findings, we concluded that LEF1, TFE3, and AR could be used as putative diagnostic markers of SPN. When β-catenin, LEF1, and TFE3 staining results were combined, the sensitivity and specificity were 100% and 91.9%, respectively.
We also evaluated the relationship between the clinicopathologic findings and the expression of these 6 markers. Among them, AR was more frequently expressed in male patients (p = 0.035). However, there were no other significant correlations between the expression of other markers and clinicopathologic parameters.
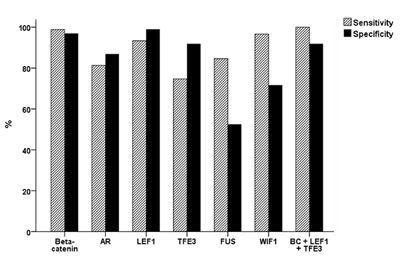
Figure 3: Sensitivity and specificity of six immunohistochemical markers for the diagnosis of SPN. When beta-catenin, LEF1, and TFE3 are combined, the sensitivity and specificity are 100% and 91.9%, respectively.
A high Ki-67 proliferative index is associated with metastatic SPN
In our SPN cohort, distant metastasis was noted in 4 (4.4%) cases. The sites of metastasis were the liver (3, 3.3%; Figure 4B) and peritoneum (1, 1.1%). The Ki-67 proliferative index was correlated with metastasis (p = 0.013, Table 3 and Figure 4A). Univariate logistic regression analysis revealed that lymphovascular invasion (OR = 21.75; p = 0.030), lymph node metastasis (OR = 22.50; p = 0.028), and a Ki-67 proliferative index of more than 5% (OR = 18.75; p = 0.014) were statistically significant risk factors for distant metastasis. The three recurred SPNs were associated with a larger tumor size (> 5cm, p = 0.043), as well as a Ki-67 proliferative index of more than 5% (p = 0.004). In univariate logistic regression analysis, lymphovascular invasion (OR = 44.00; p = 0.017) and lymph node metastasis (OR = 45.50; p = 0.011) were statistically significant risk factors for recurrence. However, nuclear pleomorphism (Figure 4C), mitotic count, perineural invasion, peripancreatic extension, and infiltrative tumor borders were not associated with distant metastasis or recurrence.
Table 3: Relationship between Ki-67 labeling index and metastasis/recurrence
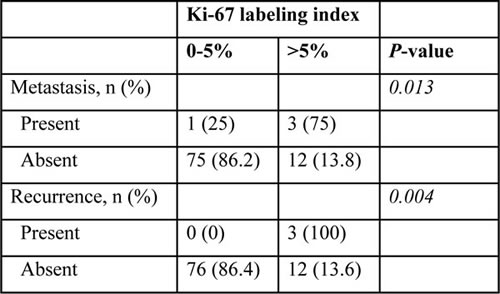
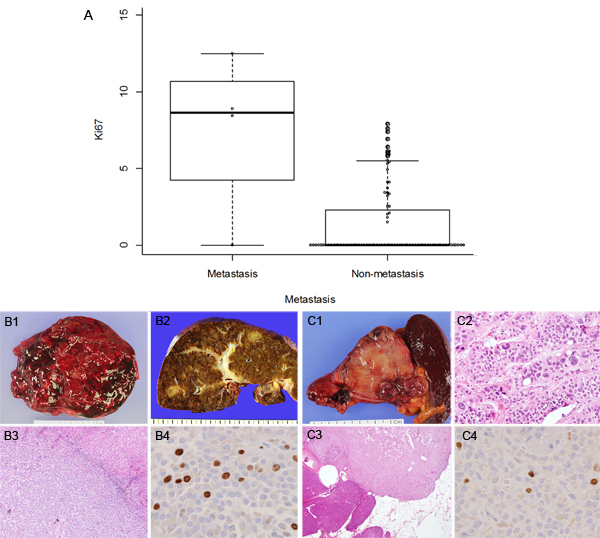
Figure 4: Ki-67 proliferative index and representative cases of SPNs with metastasis. Ki-67 proliferative index is related to distant metastasis (A, p = 0.013). Gross features of primary SPN (B1) and metastatic SPN in the liver (B2). H & E appearances metastatic SPN (B3, original magnification x40) and relatively high Ki-67 proliferative index (8.9%; B4, original magnification x400). Gross features of non-metastatic SPN (C1) may have pleomorphism (C2, original magnification x100) or peripancreatic extension (C3, original magnification x40), but show relatively low Ki-67 proliferative index (1.5%; C4, original magnification x400).
DISCUSSION
At present, the pathologic diagnosis of SPN is based on histologic and immunohistochemical characteristics. SPN is composed of small monomorphic cells with clear or eosinophilic cytoplasm that form solid or pseudopapillary structures with poor cohesion. Mutations in exon 3 of the β-catenin gene have been identified in the majority of SPNs [2, 3] and diffuse nuclear expression of β-catenin is known to be a characteristic feature used in the diagnosis of SPN. Other diagnostic markers include E-cadherin (loss) [9, 17, 18], CD10 [19], alpha-1-antitrypsin [19], chromogranin (negative) [9, 16], CD99 (dot-like pattern) [20], beta form of estrogen receptors [21], galectin 3 [21], KIT [22], DOG1 [23], FLI-1 [24], and alpha-Methylacyl-CoAracemase (AMACR, P504S) [25]. However, a confirmatory diagnosis of SPN is still challenging because SPN shares many morphological and immunophenotypical features with other pancreatic tumors, especially PDCs and NETs. In this study, we aimed to identify additional and putative diagnostic markers of SPN based on upregulated genes analyzed using mRNA expression profiling [6] and availability of antibodies.
We demonstrated that the combination of several immunohistochemical markers enhanced the diagnostic accuracy of SPN. We initially selected 6 markers and found frequent and strong nuclear expression of beta-catenin, AR, LEF1, TFE3, and FUS, and cytoplasmic expression of WIF-1 in SPNs. All 6 markers showed expression differences between SPNs and other pancreatic tumors. Although some markers were positive in both SPNs and/or PDCs and NETs, strong expression of these markers was not found in any PDCs or NETs. Our results confirmed that beta-catenin is the most sensitive (98.9%) and specific (97%) marker for SPN. Additionally, we demonstrated that 3 putative markers (LEF1, TFE3, and AR) showed high rates of sensitivity and specificity. When beta-catenin, LEF1, and TFE3 were combined, the sensitivity and specificity were 100% and 91.9%, respectively. Based on these findings, we suggest that LEF1, TFE3, and AR are useful for the diagnosis of SPN, and the combination of these markers with beta-catenin is helpful to improve their sensitivity and specificity in the diagnosis of SPNs.
LEF1 is a member of the lymphoid enhancer-binding factor 1/T-cell factor (LEF1/TCF) complex [26]. It interacts with mutant CTNNB1 and act as a regulator of the Wnt/CTNNB1 signaling pathway [26, 27]. It has been reported to be upregulated in SPNs compared with normal pancreatic tissue [6], PDCs, and NETs [7]. Singhi et al. [28] have demonstrated diffuse nuclear LEF1 overexpression in 27 SPNs with high sensitivity and specificity. We observed similar results in a larger cohort.
TFE3 is a member of the microphthalmia (MiT) family of transcription factors, which is composed of MITF, TFE3, TFEB, and TFEC. MiT transcription factors form homodimers or heterodimers that bind target promoters and act as regulators of melanocyte development, cellular proliferation, survival, motility, and metabolism, which are deregulated during oncogenic processes [29, 30]. The MITF/TFE3 family has also been shown to bind and enhance the activity of the LEF1 protein, and may promote propagation of Wnt signals in many cell types [31]. TFE3 also had been reported to interact with AR and FUS in SPNs based on gene regulatory network analysis, and it has also been suggested as a novel biomarker of SPNs [7].
AR is associated with cell cycle regulation and with the AR signaling pathway, which is activated in SPN [6, 7]. Moreover, codon changes and codon deletion mutations in AR have been found using whole exome sequencing, which also reveals its close relationship with CTNNB1 signaling [32]. Recent studies have aimed to identify gene regulatory networks in SPN and have found that CTNNB1, LEF1, AR, and TFE3 interact with each other by diverse pathways [6, 7]. These findings indicate that CTNNB1, LEF1, AR, and TFE3 are closely interrelated with each other.
In this study, we aimed to identify useful biomarkers for predicting the malignant behavior of SPN. There remains a lack of consensus of the criteria for “malignant” SPN. The concerning features, including perineural invasion, vascular invasion, or deep infiltrative growth, that are commonly found in other malignant neoplasms do not indicate malignant behavior in SPNs [1]. Consequently, all SPNs are presently classified as low-grade malignant neoplasms. For these reasons, we considered distant metastasis and recurrence as decisive malignant features of SPN. Several studies have also failed to demonstrate significant impacts of certain features that can predict metastasis or recurrence; however, an undifferentiated carcinoma component, a larger tumor size (> 8cm), microscopic malignant features (such as cellular atypia), pleomorphism, capsule invasion, peripancreatic tissue invasion, perineural invasion, lymphovascular invasion, and a high mitotic rate have all been suggested as risk factors for recurrence [12, 14-16]. Despite this, we could not find any correlation between metastasis and clinicopathologic parameters, including nuclear pleomorphism, mitotic count, perineural invasion, peripancreatic extension, an infiltrative tumor border, and our putative biomarkers. For recurrence, a larger tumor size (> 5cm) was the only correlated factor. We evaluated the association between our immunohistochemical results and the malignant behavior of SPN, and found no relationship. We also performed Ki-67 immunohistochemistry and showed that SPNs with a KI-67 proliferative index of more than 5% were significantly associated with malignant behavior. A high proliferative index using Ki-67 immunohistochemistry has been associated with aggressive biological behavior of SPNs [14, 15]. We therefore suggest that close follow-up is needed in patients with a Ki-67 proliferative index of more than 5%.
In summary, we found that LEF1, TFE3, and AR were putative diagnostic markers of SPN auxiliary to β-catenin. Incorporated into an immunohistochemical panel, they could be beneficial to distinguish SPN from PDC and NET. In addition, we suggest that the Ki-67 proliferative index could be a predictive marker of metastasis in SPN.
MATERIALS AND METHODS
Case selection
Ninety-one formalin-fixed, paraffin-embedded (FFPE) specimens of SPN were obtained from the archives of the Department of Pathology, Yonsei University, Seoul, Korea and from the Liver Cancer Specimen Bank of the National Research Resource Bank Program of the Korean Science and Engineering Foundation of the Ministry of Science and Technology; the cases were encountered between 1999 and 2016. From consecutive cases of surgically resected pancreases, 51 PDCs were retrieved from 2009 to 2011 and 48 NETs were obtained from 2009 to 2015. Hematoxylin and eosin (H&E)-stained slides of all cases were reviewed. Histologic diagnosis of SPN was made based on microscopic features and one or more immunohistochemical markers including beta-catenin, CD10, E-cadherin, and chromogranin [1].
Tissue microarray construction
Ninety-one cases of SPN, 51 of PDC, and 48 of NET were used for tissue microarray (TMA) construction using FFPE samples. A representative area of each case was selected under the microscope. Core tissues (3 mm in diameter) were taken from the individual FFPE blocks (donor blocks) and arranged in recipient paraffin blocks (tissue array blocks) using a trephine apparatus. Thirty-eight SPNs were selected and examined using conventional whole sections to rule out intra-tumoral heterogeneity.
Antibody selection and immunohistochemistry
In our previous study, we found 2,026 up-regulated genes in SPNs compared with non-neoplastic pancreas. Among them, 1,119 genes were exclusively up-regulated in SPNs compared to PDCs and NETs. Beta-catenin, AR and WNT inhibitory factor 1 (WIF-1) have been identified by western blotting and immunohistochemical analysis [6]. Beta-catenin and fused in sarcoma (FUS) have also been identified as overexpressed proteins in our previous proteome expression profiling study of SPN [8]. We chose lymphoid enhancer-binding factor 1 (LEF1) and transcription factor for immunoglobulin heavy-chain enhancer 3 (TFE3), which were associated with Wnt signaling pathway. Therefore, immunohistochemistry was performed with six commercially available antibodies to these proteins (Beta-catenin, AR, WIF-1, FUS, LEF1, and TFE3).
Immunohistochemistry was conducted on 4-μm TMA tissue sections by a Ventana Bench Mark XT Autostainer (Ventana Medical Systems, Tucson, AZ, USA). Details of the tested primary antibodies are shown in Supplementary Table 1. The pattern of immunohistochemical staining was examined by intensity and area (%) as follows; negative (no staining or staining of less than 5% of tumor cells), weak positive (faint protein expression in 5%–30% of tumor cells), moderate positive (faint protein expression in > 30% of tumor cells or definite protein expression in less than 30% of tumor cells), and strong positive (definite protein expression in > 30% of tumor cells) [9, 20, 24].
We also evaluated Ki-67 proliferative index using whole section slides to investigate its associations with prognosis. For calculating Ki-67 proliferative index (Ki-67 positive tumor cells/total counted tumor cells, %), each tumor slide was manually scanned with a microscope at × 20 objective, and the area of greatest Ki67 positivity (hot spot) was selected for photography. Colored image of the hot spot was captured, and Ki67-negative and -positive tumor cells were marked in different colors. At least 500 tumor cells were counted. Pale staining nuclei were ignored during counting [33].
Statistical analyses
Comparison of qualitative variables between groups was performed using the chi-square or Fisher’s exact test. The diagnostic sensitivity and the specificity for each of the markers individually, as well as in different combinations, were calculated. Using a logistic regression model, univariate analysis was conducted and odds ratios were estimated to assess predictive markers for distant metastasis. Two-sided P-values of less than 0.05 were considered statistically significant. Statistical analyses were performed using IBM SPSS 22 software for Windows (IBM Corp, New York, USA).
Abbreviations
AR: Androgen receptor; FFPE: formalin-fixed, paraffin-embedded; FUS: fused in sarcoma; H&E: Hematoxylin and eosin; LEF1: Lymphoid enhancer-binding factor 1; NET: neuroendocrine tumor; OR: odds ratio; PDC: pancreatic ductal carcinoma; SPN: Solid pseudopapillary neoplasm; TFE3: Transcription factor for immunoglobulin heavy-chain enhancer 3; TMA: tissue microarray; WHO: World Health Organization; WIF-1: WNT inhibitory factor 1.
CONFLICTS OF INTEREST
The authors of this article have no conflict of interest.
FUNDING
This work was supported by grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: HI14C1324) and Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: HI16C0257).
REFERENCES
1. Bosman FT, World Health Organization, International Agency for Research on Cancer. WHO classification of tumours of the digestive system. Lyon: International Agency for Research on Cancer; 2010.
2. Abraham SC, Klimstra DS, Wilentz RE, Yeo CJ, Conlon K, Brennan M, Cameron JL, Wu TT, Hruban RH. Solid-pseudopapillary tumors of the pancreas are genetically distinct from pancreatic ductal adenocarcinomas and almost always harbor beta-catenin mutations. Am J Pathol. 2002; 160:1361-9.
3. Tanaka Y, Kato K, Notohara K, Hojo H, Ijiri R, Miyake T, Nagahara N, Sasaki F, Kitagawa N, Nakatani Y, Kobayashi Y. Frequent beta-catenin mutation and cytoplasmic/nuclear accumulation in pancreatic solid-pseudopapillary neoplasm. Cancer Res. 2001; 61:8401-4.
4. Tiemann K, Heitling U, Kosmahl M, Kloppel G. Solid pseudopapillary neoplasms of the pancreas show an interruption of the Wnt-signaling pathway and express gene products of 11q. Mod Pathol. 2007; 20:955-60. doi: 10.1038/modpathol.3800902.
5. Cavard C, Audebourg A, Letourneur F, Audard V, Beuvon F, Cagnard N, Radenen B, Varlet P, Vacher-Lavenu MC, Perret C, Terris B. Gene expression profiling provides insights into the pathways involved in solid pseudopapillary neoplasm of the pancreas. J Pathol. 2009; 218:201-9. doi: 10.1002/path.2524.
6. Park M, Kim M, Hwang D, Park M, Kim WK, Kim SK, Shin J, Park ES, Kang CM, Paik YK, Kim H. Characterization of gene expression and activated signaling pathways in solid-pseudopapillary neoplasm of pancreas. Mod Pathol. 2014; 27:580-93. doi: 10.1038/modpathol.2013.154.
7. Li P, Hu Y, Yi J, Li J, Yang J, Wang J. Identification of potential biomarkers to differentially diagnose solid pseudopapillary tumors and pancreatic malignancies via a gene regulatory network. J Transl Med. 2015; 13:361. doi: 10.1186/s12967-015-0718-3.
8. Park M, Lim JS, Lee HJ, Na K, Lee MJ, Kang CM, Paik YK, Kim H. Distinct Protein Expression Profiles of Solid-Pseudopapillary Neoplasms of the Pancreas. J Proteome Res. 2015; 14:3007-14. doi: 10.1021/acs.jproteome.5b00423.
9. Kim MJ, Jang SJ, Yu E. Loss of E-cadherin and cytoplasmic-nuclear expression of beta-catenin are the most useful immunoprofiles in the diagnosis of solid-pseudopapillary neoplasm of the pancreas. Hum Pathol. 2008; 39:251-8. doi: 10.1016/j.humpath.2007.06.014.
10. Burford H, Baloch Z, Liu X, Jhala D, Siegal GP, Jhala N. E-cadherin/beta-catenin and CD10: a limited immunohistochemical panel to distinguish pancreatic endocrine neoplasm from solid pseudopapillary neoplasm of the pancreas on endoscopic ultrasound-guided fine-needle aspirates of the pancreas. Am J Clin Pathol. 2009; 132:831-9. doi: 10.1309/ajcpvt8fclfdtzwi.
11. Law JK, Ahmed A, Singh VK, Akshintala VS, Olson MT, Raman SP, Ali SZ, Fishman EK, Kamel I, Canto MI, Dal Molin M, Moran RA, Khashab MA, et al. A systematic review of solid-pseudopapillary neoplasms: are these rare lesions? Pancreas. 2014; 43:331-7. doi: 10.1097/mpa.0000000000000061.
12. Kang CM, Choi SH, Kim SC, Lee WJ, Choi DW, Kim SW. Predicting recurrence of pancreatic solid pseudopapillary tumors after surgical resection: a multicenter analysis in Korea. Ann Surg. 2014; 260:348-55. doi: 10.1097/sla.0000000000000583.
13. Papavramidis T, Papavramidis S. Solid pseudopapillary tumors of the pancreas: review of 718 patients reported in English literature. J Am Coll Surg. 2005; 200:965-72. doi: 10.1016/j.jamcollsurg.2005.02.011.
14. Zhang H, Wang W, Yu S, Xiao Y, Chen J. The prognosis and clinical characteristics of advanced (malignant) solid pseudopapillary neoplasm of the pancreas. Tumour Biol. 2016; 37:5347-53. doi: 10.1007/s13277-015-4371-5.
15. Tang LH, Aydin H, Brennan MF, Klimstra DS. Clinically aggressive solid pseudopapillary tumors of the pancreas: a report of two cases with components of undifferentiated carcinoma and a comparative clinicopathologic analysis of 34 conventional cases. Am J Surg Pathol. 2005; 29:512-9.
16. Estrella JS, Li L, Rashid A, Wang H, Katz MH, Fleming JB, Abbruzzese JL, Wang H. Solid pseudopapillary neoplasm of the pancreas: clinicopathologic and survival analyses of 64 cases from a single institution. Am J Surg Pathol. 2014; 38:147-57. doi: 10.1097/pas.0000000000000141.
17. El-Bahrawy MA, Rowan A, Horncastle D, Tomlinson I, Theis BA, Russell RC, Stamp G. E-cadherin/catenin complex status in solid pseudopapillary tumor of the pancreas. Am J Surg Pathol. 2008; 32:1-7. doi: 10.1097/PAS.0b013e31813e0676.
18. Chetty R, Jain D, Serra S. p120 catenin reduction and cytoplasmic relocalization leads to dysregulation of E-cadherin in solid pseudopapillary tumors of the pancreas. Am J Clin Pathol. 2008; 130:71-6. doi: 10.1309/feyd99txc4lmyva5.
19. Notohara K, Hamazaki S, Tsukayama C, Nakamoto S, Kawabata K, Mizobuchi K, Sakamoto K, Okada S. Solid-pseudopapillary tumor of the pancreas: immunohistochemical localization of neuroendocrine markers and CD10. Am J Surg Pathol. 2000; 24:1361-71.
20. Guo Y, Yuan F, Deng H, Wang HF, Jin XL, Xiao JC. Paranuclear dot-like immunostaining for CD99: a unique staining pattern for diagnosing solid-pseudopapillary neoplasm of the pancreas. Am J Surg Pathol. 2011; 35:799-806. doi: 10.1097/PAS.0b013e318219c036.
21. Geers C, Moulin P, Gigot JF, Weynand B, Deprez P, Rahier J, Sempoux C. Solid and pseudopapillary tumor of the pancreas--review and new insights into pathogenesis. Am J Surg Pathol. 2006; 30:1243-9. doi: 10.1097/01.pas.0000213311.28682.b2.
22. Cao D, Antonescu C, Wong G, Winter J, Maitra A, Adsay NV, Klimstra DS, Hruban RH. Positive immunohistochemical staining of KIT in solid-pseudopapillary neoplasms of the pancreas is not associated with KIT/PDGFRA mutations. Mod Pathol. 2006; 19:1157-63. doi: 10.1038/modpathol.3800647.
23. Bergmann F, Andrulis M, Hartwig W, Penzel R, Gaida MM, Herpel E, Schirmacher P, Mechtersheimer G. Discovered on gastrointestinal stromal tumor 1 (DOG1) is expressed in pancreatic centroacinar cells and in solid-pseudopapillary neoplasms--novel evidence for a histogenetic relationship. Hum Pathol. 2011; 42:817-23. doi: 10.1016/j.humpath.2010.10.005.
24. Tiemann K, Kosmahl M, Ohlendorf J, Krams M, Kloppel G. Solid pseudopapillary neoplasms of the pancreas are associated with FLI-1 expression, but not with EWS/FLI-1 translocation. Mod Pathol. 2006; 19:1409-13. doi: 10.1038/modpathol.3800664.
25. Shen Y, Wang Z, Zhu J, Chen Y, Gu W, Liu Q. alpha-Methylacyl-CoA racemase (P504S) is a useful marker for the differential diagnosis of solid pseudopapillary neoplasm of the pancreas. Ann Diagn Pathol. 2014; 18:146-50. doi: 10.1016/j.anndiagpath.2014.02.006.
26. Eastman Q, Grosschedl R. Regulation of LEF-1/TCF transcription factors by Wnt and other signals. Curr Opin Cell Biol. 1999; 11:233-40.
27. Aoki M, Hecht A, Kruse U, Kemler R, Vogt PK. Nuclear endpoint of Wnt signaling: neoplastic transformation induced by transactivating lymphoid-enhancing factor 1. Proc Natl Acad Sci U S A. 1999; 96:139-44.
28. Singhi AD, Lilo M, Hruban RH, Cressman KL, Fuhrer K, Seethala RR. Overexpression of lymphoid enhancer-binding factor 1 (LEF1) in solid-pseudopapillary neoplasms of the pancreas. Mod Pathol. 2014; 27:1355-63. doi: 10.1038/modpathol.2014.40.
29. Hemesath TJ, Steingrimsson E, McGill G, Hansen MJ, Vaught J, Hodgkinson CA, Arnheiter H, Copeland NG, Jenkins NA, Fisher DE. microphthalmia, a critical factor in melanocyte development, defines a discrete transcription factor family. Genes Dev. 1994; 8:2770-80.
30. Steingrimsson E, Copeland NG, Jenkins NA. Melanocytes and the microphthalmia transcription factor network. Annu Rev Genet. 2004; 38:365-411. doi: 10.1146/annurev.genet.38.072902.092717.
31. Yasumoto K, Takeda K, Saito H, Watanabe K, Takahashi K, Shibahara S. Microphthalmia-associated transcription factor interacts with LEF-1, a mediator of Wnt signaling. Embo j. 2002; 21:2703-14. doi: 10.1093/emboj/21.11.2703.
32. Guo M, Luo G, Jin K, Long J, Cheng H, Lu Y, Wang Z, Yang C, Xu J, Ni Q, Yu X, Liu C. Somatic Genetic Variation in Solid Pseudopapillary Tumor of the Pancreas by Whole Exome Sequencing. Int J Mol Sci. 2017; 18. doi: 10.3390/ijms18010081.
33. Reid MD, Bagci P, Ohike N, Saka B, Erbarut Seven I, Dursun N, Balci S, Gucer H, Jang KT, Tajiri T, Basturk O, Kong SY, Goodman M, et al. Calculation of the Ki67 index in pancreatic neuroendocrine tumors: a comparative analysis of four counting methodologies. Mod Pathol. 2015; 28:686-94. doi: 10.1038/modpathol.2014.156.

