INTRODUCTION
Normal cells produce energy mostly through the oxidation of pyruvate in the mitochondria; however, cancer cells are known to produce energy via increased glycolysis in the cytosol. This effect known as “Warburg Effect” [1] requires a metabolic shift from oxidative phosphorylation to glycolysis or lactate fermentation [2]. Glutamine, one of the most abundant intracellular amino acids, plays an important role in satisfying the biosynthetic needs of proliferating cancer cells by providing carbons to produce tricarboxylic acid (TCA) cycle intermediates, glutathione, fatty acids, and nucleotides [3–6]. As a result of glutamine being a major source of carbon molecules in tumor growth-facilitating metabolic pathways, many cancer cells often become “addicted” to glutaminolysis (a rate limiting step in the TCA cycle) [5]. Glutaminolysis occurs via two steps, first step is catalyzed by glutaminase (GLS) and converts glutamine to glutamate. The second step converts glutamate to α-ketoglutarate (α-KG) and is catalyzed by glutamate dehydrogenase (GDH) [7]. Cancer cells addicted to glutaminolysis often rely on glutamine as the carbon source for the TCA cycle [8].
In humans, GLS exists in two forms: kidney-type glutaminase (GLS1) and liver-type glutaminase (GLS2). While GLS1 is expressed ubiquitously, GLS2 is expressed primarily in the liver [9]. Recent efforts have focused on targeting glutaminolysis by inhibiting the GLS activity in cancer cells [10]. Drugs targeting GLS activity such as BPTES (bis-2-(5-phenylacetamido-1,2,4-thiadiazol-2-yl)ethyl sulfide 3) or CB-839 have gained attention owing to their potent inhibition of GLS1 activity and anti-proliferative effect in multiple tumor subtypes including leukemia and triple negative breast cancer [11].
Neurofibromatosis type 1 (NF1) is an autosomal dominant genetic disorder caused due to the loss and/or mutation of NF1 tumor suppressor gene [12]. The NF1 gene codes for a Ras GTPase activating protein called Neurofibromin (NF) and mutational inactivation and/or loss of NF1 can lead to altered Ras-MAPK signaling [13]. Many patients with NF1 are often at risk of developing cancers such as gliomas, neurofibromas and malignant peripheral nerve sheath tumors (MPNSTs) among others [14, 15]. MPNSTs are soft-tissue tumors that are highly aggressive with a very poor prognosis [16]. NF1 associated MPNSTs are often fatal and there are not many treatment options available to treat these therapeutically resistant tumors.
Although glutamine metabolism has been shown to play a crucial role in tumorigenesis both in vitro and in vivo [17], its role in NF1 disease setting has not been studied before. In this study, we report for the first time that NF1 associated soft-tissue sarcoma cell lines (MPNST, ST8814, S462) are highly dependent on glutamine for proliferation compared to wild-type NF1 cell lines (LS141, CHP100, STS26T). Targeted inhibition of glutaminase (GLS) using inhibitors BPTES and CB-839 results in significant inhibition of cell proliferation and mTORC1 activity. Association between glutamine metabolism and NF1 was also confirmed using siRNA and NF1 over-expression studies in vitro. Furthermore, treatment of MPNST xenografts with CB-839 resulted in significant suppression of tumor volume and inhibition of downstream signaling pathways. Results obtained in this study strongly suggest that glutaminase inhibition in NF1 associated tumors needs to be explored for a potentially novel therapeutic approach in this disease setting.
RESULTS
NF1 mutant/null cell lines show decreased cell viability and mTORC1 activity in response to glutamine deprivation
Although NF1 is known to play a role in the development of malignant peripheral nerve sheath tumors (MPNSTs), its role in modulating glutamine dependency has not been studied before. MPNST, ST8814 and S462 cell lines used in this study have been shown previously to carry a mutation/deletion in NF1, whereas, STS26T cell line carries wild-type NF1 [18–20]. LS141 (Liposarcoma) and CHP100 (Ewing Sarcoma) cell lines, on the other hand, have been used extensively and both these cell lines have not been reported to harbor any NF1 mutation/loss [19, 21–24] (also, personal communication with Kanojia D, Cancer Science Institute, Singapore). Figure 1A shows the expression levels of NF1 in the six soft-tissue sarcoma cell lines that were used in this study. MPNST cell line shows detectable levels of NF1 expression since it is NF1 mutant, whereas, ST8814 and S462 cell lines do not show any detectable levels of NF1 on the western blot (Figure 1A).
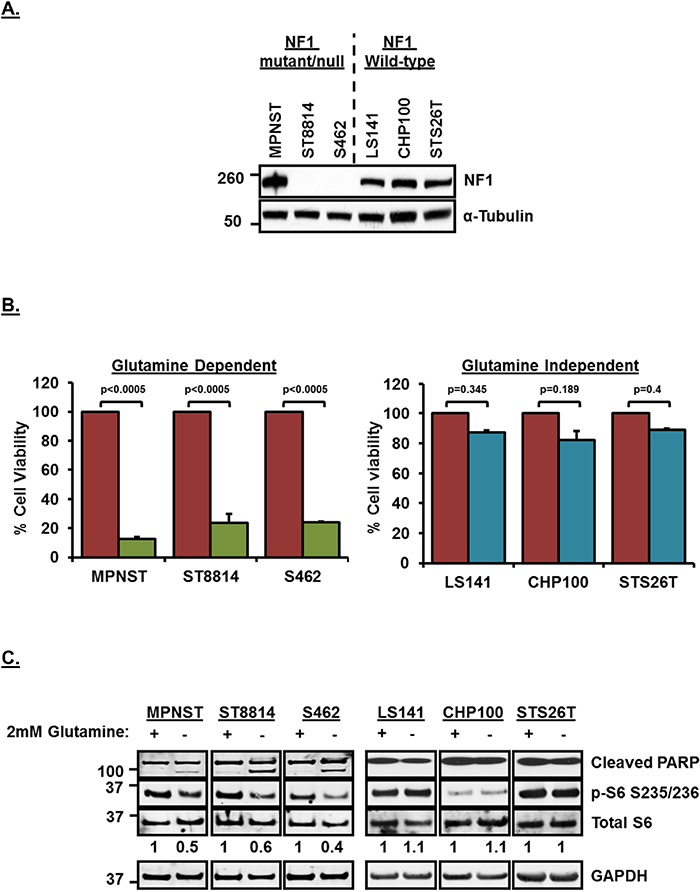
Figure 1: (A) NF1 expression levels in NF1 mutant/null and NF1 wild-type soft-tissue sarcoma cell lines. Cells from a confluent 60mm plate were washed twice with ice-cold PBS and cell pellet was obtained by scraping in PBS and centrifuging. Pellet was lysed with RIPA lysis buffer. 30μg of lysates were loaded on SDS/PAGE and proteins were detected on western blot using indicated antibodies. Numbers on the left indicate molecular weight in kilo Daltons (kDa). (B) Glutamine dependency of NF1 mutant/null cell lines for cell proliferation.1500 cells per well were plated in 96 well plates in triplicate in RPMI+10%FBS without Glutamine for 24 hours. Next day, media was replaced with RPMI+10%FBS with or without 2mM Glutamine. After 72 hours, cell viability was measured using Dojindo CCK-8 kit using manufacturer’s instructions. Cell viability was calculated as percentage of growth in 2mM Glutamine containing media. Combined data from two independent experiments is shown. Error bars represent standard error mean. (C) Induction of apoptosis and downregulation of mTORC1 after glutamine deprivation in NF1 mutant/null sarcoma cell lines. Cells were plated in RPMI+10%FBS without Glutamine for 24 hours. Next day, media was replaced with fresh RPMI+10%FBS without Glutamine or RPMI+10%FBS containing 2mM Glutamine. Cells were incubated for another 48 hours, harvested, cell pellets were lysed in RIPA lysis buffer and 30μg of lysates were loaded on SDS/PAGE. Proteins were detected on western blot using indicated antibodies. Numbers at the bottom of blot indicate densitometric quantitation of p-S6 signal normalized to total S6 levels. Numbers on the left indicate molecular weight in kilo Daltons (kDa). Representative blot from at least 3 independent experiments are shown.
To test the dependency of these cell lines on extracellular glutamine for cell proliferation, we carried out a cell viability assay in the presence or absence of glutamine (2mM) in the growth media. Removal of glutamine from the media significantly (p<0.0005) decreased cell viability (Figure 1B) only in the NF1 mutant/null cell lines, MPNST, ST8814 and S462 compared to wild-type NF1 sarcoma cell lines, LS141, CHP100 and STS26T. A similar decrease in cell viability was observed in an NF1-null metastatic melanoma cell line, MeWo [25] when glutamine was removed from the media, whereas, another melanoma cell line, 92.1 [26], that does not carry any NF1 mutation/deletion did not show any decrease in cell viability after glutamine removal from the media (Supplementary Figure 1, left panel). This clearly suggested that cancer cell lines other than soft-tissue sarcoma carrying NF1 mutation/deletion may also show sensitivity to glutamine deprivation. Western blot analysis shows that removal of glutamine from the media induced apoptosis (shown as induction of cleaved poly ADP-ribose polymerase, PARP) only in NF1 mutant/null but not in wild-type NF1 cell lines (Figure 1C and Supplementary Figure 1, right panel). Since glutaminolysis is known to play an important role in mTORC1 activation [27], removal of glutamine from media downregulated mTORC1 activity (shown as decreased phosphorylation of S6 ribosomal protein, p-S6 S235/236 and also quantitated in arbitrary densitometric units) in NF1 mutant/null but not wild-type NF1 cell lines (Figure 1C and Supplementary Figure 1).
Glutaminase inhibition results in decreased cell proliferation, mTORC1 signaling and glutamine utilization in NF1 mutant/null cell lines
Inhibition of glutaminase, a key enzyme in the conversion of glutamine to glutamate, by BPTES [bis-2-(5-phenylacetamido-1,2,4-thiadiazol-2-yl)ethyl sulfide] and CB-839 has been shown to have anti-proliferative effect in triple negative breast cancer, leukemia and glioma [28–30]. To test the efficacy of glutaminase inhibition by BPTES and CB-839 in these cell lines, we carried out cell viability assays and western blot analysis using increasing concentrations of both the drugs. As shown in Figure 2A and 2B, both BPTES and CB-839 treatment resulted in a significant decrease (p<0.001) in cell viability only in NF1 mutant/null cell lines (also Supplementary Figure 1, left panel). Notably, CB-839 treatment resulted in decreased cell viability at low nanomolar concentrations compared to BPTES (Figure 2A versus 2B). Western blot analysis (Figure 2C) after 48 hours of CB-839 treatment revealed induction of cleaved PARP as well as downregulation of mTORC1 targets such as p-S6 and p-4EBP1 in NF1 mutant/null but not in wild-type NF1 cell lines. Similar decrease in cell viability, induction of PARP and downregulation of downstream signaling pathways was observed for the NF1-null melanoma cell line, MeWo but not for the wild-type NF1 carrying melanoma cell line, 92.1 (Supplementary Figure 1, right panel). CB-839 treatment at time points as early as 3 and 6 hours also showed downregulation of mTORC1 targets such as p-S6 only in the NF1-null cell line, ST8814 (Supplementary Figure 2).
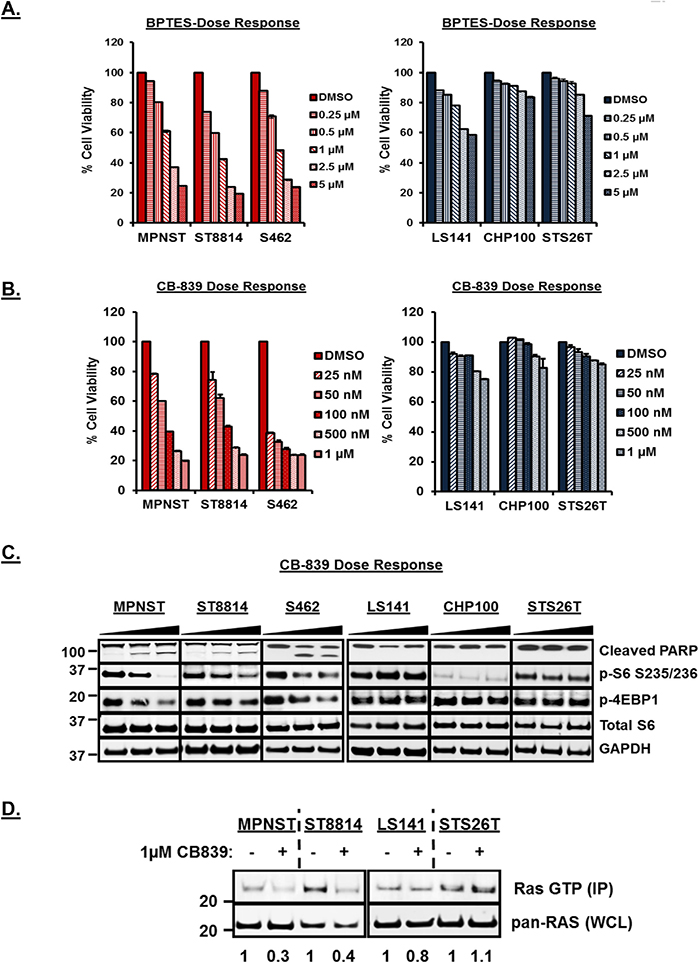
Figure 2: (A and B) Anti-proliferative efficacy of BPTES and CB-839 in NF1 mutant/null (MPNST, ST88 and S462) versus NF1 wild-type (LS141, CHP100 and STS26T) cell lines. 1500 cells per well were plated in 96 well plates in triplicate in RPMI+10%FBS without Glutamine for 24 hours. Next day, cells were treated with increasing dose of CB-839 or BPTES in RPMI+10%FBS with 2mM Glutamine. After 72 hours, cell viability was measured using Dojindo CCK-8 kit using manufacturer’s instructions. Cell viability was calculated as percentage of growth compared to DMSO control. Combined data from two independent experiments is shown. (C) Induction of apoptosis and downregulation of mTORC1 with the treatment of CB-839 in NF1 mutant/null versus wild-type NF1 cell lines. Cells were grown to 60% confluency in 60-mm plates for 24 hours and treated for another 48 hours with DMSO or increasing concentrations of CB-839 (0.1 and 1μM). 30μg of RIPA lysates were loaded on SDS/PAGE and immunoblotted using indicated antibodies. Representative blot from at least 3 independent experiments are shown. Numbers next to the blot indicate molecular weight in kilo Daltons (kDa). (D) Detection of active Ras in NF1 mutant/null and NF1 wild-type cell lines. In vitro Ras activity from cell lysates was determined using active Ras pull down and detection kit as described in materials and methods. Immunoprecipitated samples (IP) along with 25μg of total cell lysates (TCL) were analyzed by Western blotting using an anti-Ras mouse monoclonal antibody. Representative blot from at least 3 independent experiments are shown. Numbers next to the blot indicate molecular weight in kilo Daltons (kDa).
To test whether the inhibitory effects observed after CB-839 treatment were solely a result of inhibition of mTORC1 activity by CB-839; we compared the effects of CB-839 treatment against a known mTORC1 inhibitor, rapamycin, in a cell proliferation and western blot analysis assay. Unlike CB-839, rapamycin treatment at 10nM [24] induced a modest inhibition of cell proliferation in the NF1-null (ST8814) cell line (Supplementary Figure 2). Complete inhibition of mTORC1 targets such as p-S6 was observed in both the cell lines (ST8814 and STS26T) irrespective of their NF1- status (Supplementary Figure 2). Only CB-839 but not rapamycin treatment induced cleaved PARP in the NF1-null ST8814 cell line (Supplementary Figure 2). No changes in the expression levels of GLS1 were observed after CB-839 or rapamycin treatment suggesting that the effects of CB-839 treatment are at the functional rather than expression levels of GLS1.
Neurofibromin (NF), a product of the NF1 gene, is a RAS GTPase-activating protein and is well known to negatively regulate the RAS signaling pathway [31]. Since only the NF1- associated but not wild-type NF1 cell lines were sensitive to glutaminase inhibition, we tested whether CB-839 treatment has any effect on Ras activity in these cell lines. To test this, we carried out an in vitro pulldown assay for active Ras (Ras GTP immunoprecipitated using Ras Binding Domain of Raf, RBD) using cell lysates from two NF1 associated (MPNST, ST8814) and two wild-type NF1 cell lines (LS141, STS26T) treated with 1μM CB-839 for 48 hours. Western blots in Figure 2D (top panel) show a significant decrease in immunoprecipitated Ras-GTP levels (active Ras) only in the NF1-associated but not wild-type NF1 cell lines (also shown as arbitrary densitometric units). No decrease in total Ras was observed in any of the cell lines (Figure 2D, bottom panel). NF1-null melanoma cell line, MeWo, but not wild-type NF1 carrying cell line 92.1, showed similar decrease in active Ras in response to CB-839 treatment (Supplementary Figure 3).
In order to test the metabolic consequences of glutaminase inhibition, we carried out intracellular metabolite analysis in response to CB-839 treatment in two NF1 associated (MPNST and ST8814) and two wild-type NF1 cell lines (LS141 and STS26T). Treatment with 1μM CB-839 for 48 hours resulted in a significant decrease in glutamine utilization (p<0.01) shown as a steep increase in intracellular glutamine levels and a significant decrease in glutamate levels (p<0.005) (Figure 3A), thus, confirming reduced conversion of glutamine to glutamate. The increase in glutamine levels or the decrease in glutamate levels was not as significant in wild-type NF1 cell lines (Figure 3A versus 3B). It must be noted that the basal metabolite levels for glutamine in the NF1 associated cell lines were higher than the wild-type NF1 cell lines (Figure 3A and 3B). In addition to glutamate levels, a decrease in other TCA intermediary metabolites such as succinate, fumarate and α-keto-Glutarate (α-KG) were also more pronounced in NF1 associated compared to wild-type NF1 cell lines (Figure 3A and 3B).
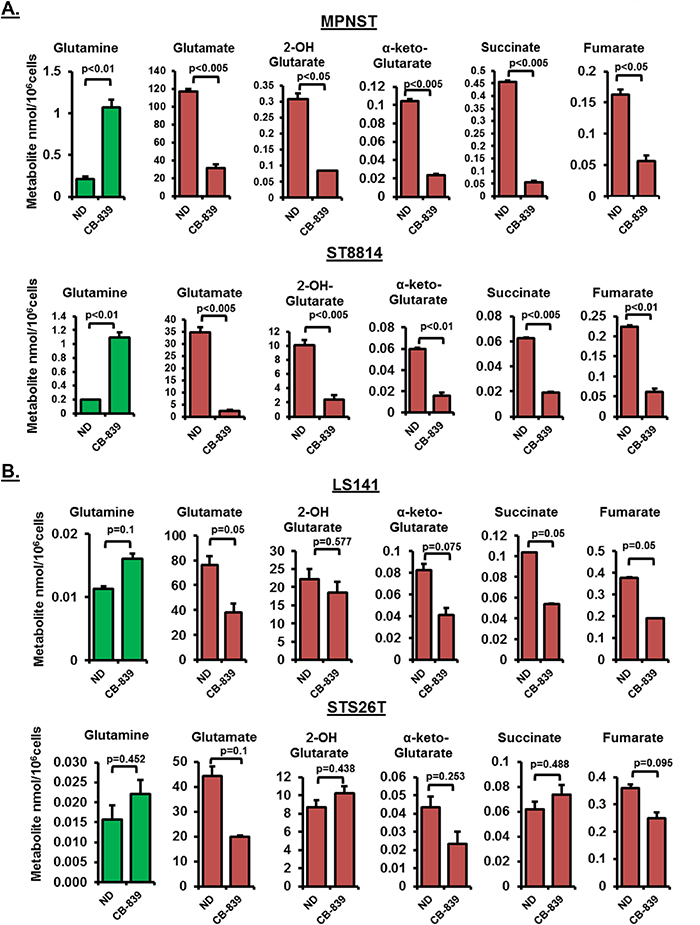
Figure 3: (A and B) Intracellular metabolite profiling after CB-839 treatment. Changes in intracellular metabolite levels in NF1 mutant/null (MPNST, ST8814) and NF1 wild-type (LS141, STS26T) cells treated with DMSO or 1μM CB-839 for 48 hours are shown. The levels of intermediary metabolites (nmol/106cells) were determined by Gas chromatography-mass spectrometry (GC-MS) metabolomics as described in materials and methods. Results are representative of at least two independent experiments. Error bars represent standard error mean. ND=No Drug control.
Effects of glutamine deprivation can be restored by re-addition of glutamate to the media
In order to test whether the decreased cell viability is a result of reduced conversion of glutamine to glutamate, we carried out a rescue experiment by adding 5mM glutamate back to the media. Figure 4A shows that adding glutamate back to the media rescued cell viability in both NF1 associated cell lines (more significantly in the NF1 null ST8814 cell line among the two), thereby, strongly supporting the hypothesis that NF1 associated cell lines are highly dependent on glutaminase activity for cell viability. Re-addition of either 2mM glutamine or 5mM glutamate to the growth media also resulted in restoring downregulated mTROC1 activity as indicated by increased p-S6 signal in NF1 associated cell lines after re-addition (Figure 4B). As seen earlier (Figure 1C), wild-type NF1 cell lines (LS141 and STS26T) did not show any decrease in mTORC1 signaling as a result of glutamine removal and therefore, p-S6 levels stayed the same and did not increase further despite re-addition of either glutamine or glutamate to the growth media (Figure 4B).
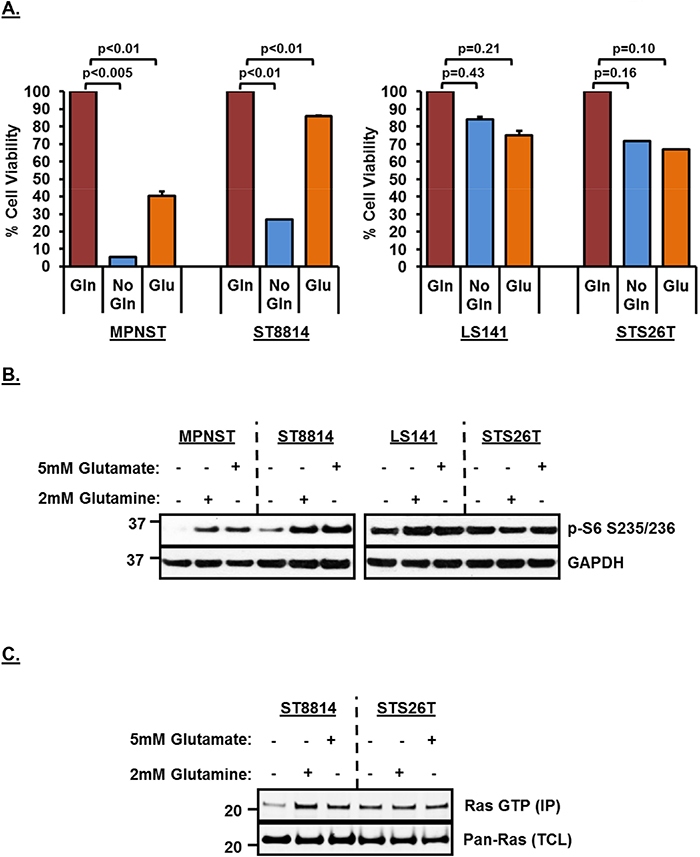
Figure 4: (A) Rescue of growth inhibition in NF1 mutant/null cell lines by Glutamate supplementation in media. 1500 cells per well were plated in 96 well plates in triplicate in RPMI+10%FBS without Glutamine for 24 hours. Next day, media was replaced with fresh RPMI+10%FBS without Glutamine or RPMI+10%FBS containing 2mM Glutamine or 5mM Glutamate. After 72 hours, cell viability was measured using Dojindo CCK-8 kit using manufacturer’s instructions. Cell viability was calculated as percentage of growth in 2mM Glutamine containing media. Combined data from two independent experiments is shown. Error bars represent standard error mean. (B and C) Rescue of mTORC1 and Ras activity after re-addition of Glutamine or Glutamate in the media. Cells were plated in RPMI+10%FBS without Glutamine 24 hours. Next day, media was replaced with fresh RPMI+10%FBS without Glutamine or RPMI+10%FBS containing 2mM Glutamine or 5mM Glutamate. Cells were incubated for another 48 hours, harvested, cell pellets were lysed in RIPA lysis buffer and 30μg of lysates were loaded on SDS/PAGE. Proteins were detected on western blot using indicated antibodies. In vitro Ras activity from cell lysates was determined using active Ras pull down and detection kit as described in materials and methods. Representative blot from at least 2 independent experiments are shown. Numbers next to the blot indicate molecular weight in kilo Daltons (kDa).
To test whether the decreased Ras activity in response to removal of glutamine from the media could also be restored, we carried our Ras pull-down assay after adding 2mM glutamine or 5mM glutamate back to the media. As shown in Figure 4C, re-addition of either glutamine or glutamate to the media restored active Ras (top panel) in the NF1-null cell line, ST8814. No change in active Ras was noted in the wild-type NF1 cell line, STS26T, irrespective of the addition of glutamine or glutamate to the media. No decrease in total Ras was observed in any of the cell lines.
Changes in NF1 expression levels modulates sensitivity and Ras activity in response to glutaminase inhibition
In order to confirm the role of NF1 in conferring sensitivity to glutaminase inhibition, we next carried out NF1 overexpression as well as siRNA mediated NF1 knockdown experiments. Overexpression of NF1-GRD (GAP Related Domain) has been shown previously to restore normal cell growth in NF1-/- cells [32] as well as downregulate high levels of Ras-GTP in leukemia cells [33]. To test this, we overexpressed wild-type NF1-GRD in the NF1 null cell line, ST8814, and then treated the cells with 1μM CB-839 for 48 hours. As shown earlier, CB-839 treatment in empty vector transfected cells resulted in decreased cell viability and mTORC1 signaling (p-S6) as well as reduced Ras-GTP levels (Figure 5A). However, overexpression of NF1-GRD was able to rescue the effects of CB-839 treatment on cell viability as well as mTORC1 and active Ras (Figure 5A), thus, restoring p-S6 as well as Ras-GTP levels in NF1-overexpressing cells. Similar results were obtained in NF1 null melanoma cell line, MeWo, (Supplementary Figure 3) indicating a strong correlation between NF1 status and glutamine dependency.
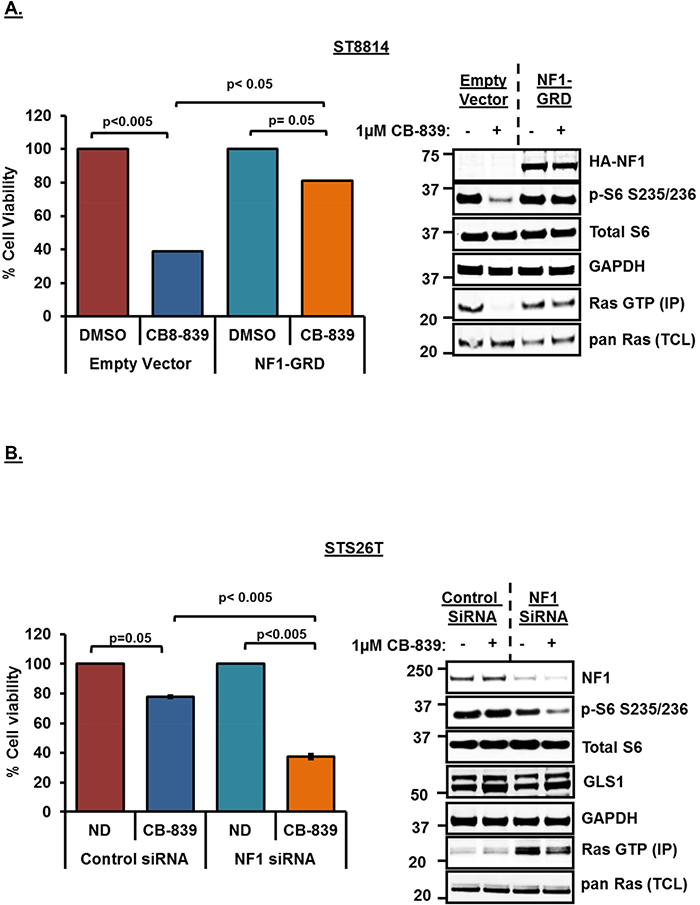
Figure 5: (A) Rescue of cell proliferation and mTORC1 activity by wild-type NF1-GRD overexpression. NF1 null (ST8814) cells were plated overnight and transfected transiently with HA-tagged wild-type NF1 GAP related domain (NF1-GRD) or an empty vector next day and incubated for another 48 hours. Cells were then trypsinized and 1500 cells per well were plated in 96 well plates in triplicate and 100 mm plates ( 106 cells / plate) for cell viability and western blot analysis respectively. Cells were then treated next day with 1μM CB-839 or DMSO for another 72 hours. Cell viability was measured after using Dojindo CCK-8 kit using manufacturer’s instructions as described earlier. Cell viability was calculated as percentage of growth in DMSO control. Western blot analysis and active Ras pull down from the lysates was carried out as described in materials and methods. (B) siRNA mediated knockdown of wild-type NF1 sensitizes STS26T cell line to glutaminase inhibition by CB-839. 100 nM siRNAs (pooled siRNAs, Dharmacon) specific for NF1 or scrambled (non-targeting pool) siRNA as a control were transfected using Lipofectamine RNAiMAX reagent (Invitrogen). 48 hours after transfections, cells were trypsinized, counted using a Nexcelom cell counter and plated in 96-well plates in triplicate for cell viability assays or 60mm plates for western blotting analysis. Cells were also collected, lysed and analyzed by western blot to check for knockdown of protein expression. For western blotting, cells in 60mm plates were treated for 24 hours, whereas, for cell viability assays, cells in 96 well plates were treated for 72 hours with 1μM CB-839. Combined data from two independent experiments is shown. Numbers next to the blot indicate molecular weight in kilo Daltons (kDa).
We also carried out siRNA mediated knockdown of NF1 in the wild-type NF1 cell line, STS26T. As a result of NF1 knockdown, STS26T cell line exhibited decreased cell viability, and reduced mTORC1 activity (p-S6) (Figure 5B). SiRNA mediated loss of NF1 resulted in increased Ras-GTP levels (Figure 5B, western blot, lane 3). CB-839 treatment post NF1 knockdown was able to reduce increased Ras-GTP levels (Figure 5B, western blot, lane 4) clearly suggesting glutamine dependency in these cells when NF1 is knocked down. No changes in the expression levels of GLS1 were observed after SiRNA mediated knockdown of NF1 or post CB-839 treatment (Figure 5B).
Glutaminase inhibition by CB-839 in MPNST xenografts results in suppression of tumor volume and reduced glutamine utilization in tumors
In order to test whether glutaminase inhibition by CB-839 can result in reduced tumor volume in vivo, we carried out mouse xenograft studies using MPNST serial transplant tumors [34]. As shown in Figure 6A, treatment with 200mg/kg CB-839 resulted in significant suppression of tumor volume compared to vehicle control (n=6). Western blot analysis (Figure 6B) revealed inhibition of downstream signaling pathways including p-ERK1/2 as well as mTORC1 targets such as p-S6 and p-4EBP1, thus, confirming the inhibition observed in vitro. Similar to the results obtained in vitro, metabolite profiling of tumor samples (Figure 6C) showed a significant decrease in glutamine utilization (shown as increased accumulation of glutamine in tumors) and reduced levels of TCA cycle intermediary metabolites such as α-keto-Glutarate, succinate and fumarate.
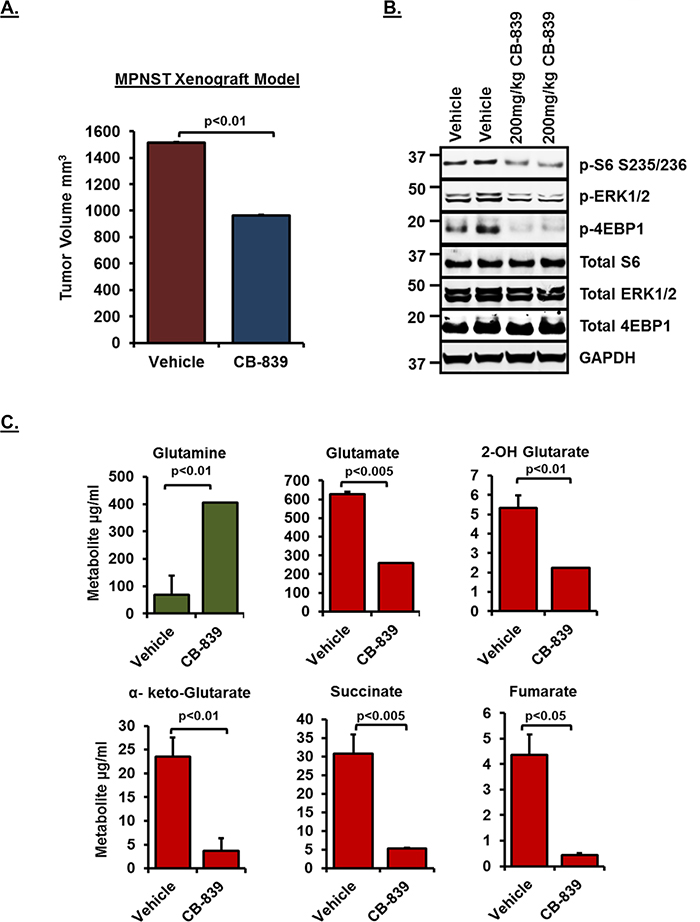
Figure 6: Effect of CB-839 treatment in MPNST xenografts. (A) Tumor growth of MPNST xenografts treated with CB-839 or Vehicle control for a period of 2 weeks is shown. See methods for drug dosing schedule. (B) 30μg of RIPA lysates obtained using sample grinding kit (GE healthcare) from xenograft tissues at the end of 2-week treatment were loaded on SDS/PAGE and immunoblotted using indicated antibodies. Blots from tumor tissues obtained from two independent animals per treatment condition are shown. Numbers next to the blot indicate molecular weight in kilo Daltons (kDa). (C) Relative changes in intermediary metabolite levels (μg/ml) in MPNST xenografts treated with CB-839 or Vehicle controls (n=6) were measured by Gas chromatography-mass spectrometry (GC/MS) based metabolomics as described in materials and methods. Error bars represent standard error mean.
DISCUSSION
The present study is the first report evaluating glutamine dependency and glutaminase inhibition as a potential therapeutic approach in NF1 associated tumors. Though previous studies have evaluated efficacy of glutaminase inhibitors in cancers such as triple negative breast cancer [28], acute myeloid leukemia [35] and pancreatic cancer [36], glutamine dependency in NF1 associated malignancies has not been studied before. Malignant peripheral nerve sheath tumors or MPNSTs arise in patients with type 1 neurofibromatosis (NF1) and are often associated with activation of the Ras pathway due to loss of function mutations in NF1, a gene which encodes the Ras-GTPase activating protein (GAP), Neurofibromin (NF) [37]. MPNSTs are often difficult to treat and the only definitive therapy despite recent advances relies on surgical resection of tumor. Though many molecularly targeted agents are currently being tested in clinical trials [38], the prognosis still remains poor and more effective clinical approaches are needed to treat MPNSTs. In addition to NF1 associated MPNSTs, role of NF1 mutations has also been elucidated in other cancer types such as cutaneous melanoma [25]. Although research aimed at understanding molecular mechanisms that drive NF1 associated malignancies has been in focus for some time [13], not many treatment options are available to treat this disease and novel therapeutic approaches are urgently needed [39].
Cancer cells are known to utilize large amounts of glucose and glutamine in order to fuel the tricarboxylic acid (TCA) cycle. Glutamine, a critical nutrient for driving cancer cell proliferation, acts as a source of fatty acid, lipid as well as glutathione synthesis especially under hypoxic conditions [40]. Glutaminase, a key enzyme that converts glutamine to glutamate, is crucial for generation of intermediary metabolites such as α-keto-Glutarate that drive fatty acid production and glutathione synthesis [41]. Glutamine deprivation and glutaminase inhibition, therefore, has emerged as a potential therapeutic approach in recent years [11, 42–45]. Selective glutaminase inhibitors, such as CB-839 (N-(5-(4-(6-((2-(3-(Trifluoromethoxy)phenyl)acetyl)amino)-3-pyridazinyl)butyl)-1,3,4-thiadiazol-2-yl)-2-pyridineacetamide) have been the subject of clinical trials in solid tumors and hematological malignancies (ClinicalTrials.gov Identifier: NCT02071862, ClinicalTrials.gov Identifier: NCT02071888, and ClinicalTrials.gov Identifier: NCT02071927). Based on preclinical data, combination trials of CB-839 with EGFR inhibitors in EGFR mutant lung cancer and with proteasome inhibitors in multiple myeloma are now being planned [46, 47]. When we tested the effect of glutamine deprivation on cell viability in six soft-tissue sarcoma cell lines, we observed a significant decrease in cell viability and induction of apoptosis after glutamine removal from the media only in the NF1 associated cell lines. This was surprising since no correlation has been reported previously between the NF1 status and glutamine dependency. To explore further, we carried out cell viability and western blot analysis after glutaminase inhibition using either BPTES or CB-839 and observed that glutaminase inhibition not only decreased cell viability and induced apoptosis but also resulted in significant downregulation of p-S6, a known mTORC1 target. When we deprived the NF1 mutant/null cells of glutamine in the media or treated the cells with glutaminase inhibitor, CB-839 and carried out in vitro detection of Ras activity, we observed a significant decrease in active Ras (Ras-GTP), thus, supporting our hypothesis that Ras activity in these cell lines is dependent on glutamine utilization. Moreover, this decrease in Ras activity could be recapitulated when NF1 was knocked down using siRNA in the wild-type NF1 cell line, STS26T. Conversely, when wild-type NF1-GRD (GAP Related Domain) was overexpressed in NF1 null cell line (ST8814), the decrease in Ras activity in response to glutaminase inhibition could be restored. Taken together, our data strongly suggests that Ras activity in the NF1 mutant/null cell lines is highly dependent on the ability of cells to utilize glutamine.
Interestingly, we observed that sensitivity to glutaminase inhibition is modulated depending on the presence or absence of wild-type NF1. Decreased cell viability in response to CB-839 treatment in NF1 null cell line (ST8814) could be reversed when wild-type NF1 was overexpressed. On the other hand, siRNA mediated knockdown of NF1 in wild-type NF1 cell line (STS26T) resulted in increased sensitivity to glutaminase inhibition.
In accordance with the previously reported studies [7, 27], we observed a reduction in mTORC1 signaling (shown as decreased p-S6 and p-4EBP1) in response to glutaminase inhibition and glutamine deprivation confirming the role played by glutaminolysis in mTORC1 activation. Similar to cell viability assays, knockdown or overexpression of NF1 modulated mTORC1 activity in response to glutaminase inhibition. Thus, decreased mTORC1 (p-S6) signaling after CB-839 treatment was rescued when NF1 was overexpressed, whereas, mTORC1 activity was reduced when NF1 was knocked down.
Western blot analysis using tissue lysates obtained from tumor xenograft samples also showed a decrease in p-ERK as well as p-S6 and p-4EBP1, thus, validating the results obtained in vitro. Loss of NF1 is known to induce Ras activation [31] and our data showing blockade of p-ERK (a critical component of Ras/Raf/ERK pathway) in response to glutaminase inhibition suggested that glutamine metabolism in the NF1 associated tumors is important for Ras activity.
Decreased glutamine utilization after CB-839 treatment has been reported earlier in triple negative breast cancer [28]. Intermediary metabolite profiling showed a similar decrease in utilization of glutamine and a significant reduction in key TCA metabolites such as α-keto-Glutarate, succinate and fumarate in NF1 associated cell lines (MPNST and ST8814) compared to wild-type NF1 cell lines (LS141 and STS26T) as well as in MPNST tumor xenograft samples in vivo. It was noteworthy that cell viability as well as mTORC1 signaling and Ras activity were rescued when one of the key metabolites, glutamate, was provided in the media suggesting that cells were able to take up externally added glutamate, thus, overcoming the blockade of intracellular glutamine utilization. It must be noted though that the glutamate levels were similar in both glutamine dependent and independent cell lines despite intracellular levels of glutamine being low in glutamine independent cell lines. We postulate that these cell lines are able to generate intermediary metabolites like glutamate via glucose consumption since no glutamine dependency was observed. On the other hand, cell lines highly dependent on glutamine (MPNST and ST8814) are able to generate glutamate mostly via glutaminolysis. Therefore, the decrease in glutamate levels post CB-839 treatment is more pronounced in glutamine dependent cell lines compared to glutamine independent cell lines. More studies need to be undertaken to understand the intricacies of glutamine versus glucose consumption in NF1 mutant/null and NF1 wild-type cell lines. The other notable observation from the metabolite profiling analysis was the high-levels of the intermediary metabolite 2-hydorxy-glutarate (2-HG), a known oncometabolite, in the NF1-null ST8814 as well as wild-type NF1 carrying LS141 and STS26T cell lines. Cancer associated mutations in isocitrate dehydrogenase (IDH) often lead to high levels of 2-HG [48]. While the D-isomer of 2-HG is often associated with mutant IDH, L-isomer is produced under hypoxic conditions [49]. The metabolites quantitated by the mass spectrometry approach used in this study are L-enantiomers. Moreover, previous mutational genomic analysis of the NF1-null ST8814 [20] and wild-type NF1 carrying LS141 [22] cell lines have not identified any mutations in IDH, therefore, we do not anticipate that the high levels of 2-HG observed in these cell lines are a result of possible IDH mutations.
Glutamine has been shown to play a major role not only as a carbon and nitrogen source but also contributes to the anti-oxidative pathways and chromatin organization [50]. Other studies have shown that pancreatic cancer cells with oncogenic KRas are dependent on glutamine metabolism for proliferation [51]. As mentioned earlier, studies have reported that mTORC1 activation is dependent on glutaminolysis and glutamine metabolism is required to promote cell growth and proliferation [27]. We believe that in our study, NF1 mutant/null cells are highly dependent on glutamine metabolism for proliferation and withdrawal of glutamine from media or inhibition of GLS1 by CB-839 results in decreased glutaminolysis and thus, reduced mTORC1 activity (shown as reduced p-S6 and p-4EBP1). Additionally, we observe that the Ras activity in our NF1 mutant/null cells is highly dependent on glutamine metabolism. Therefore, we believe that the decreased cell proliferation in the NF1 mutant/null cells is a combined result of decreased mTORC1 and Ras activity and this can be rescued either by overexpression of NF1-GRD or exogenous addition of glutamate to the media.
In summary, our data shows a strong correlation between NF1 status and glutamine dependency in tumors. In addition to MPNSTs, a known NF1 associated malignancy, such a correlation was also observed only in NF1 null metastatic melanoma cell line, MeWo but not wild-type NF1 carrying 92.1 cell line. This observation certainly warrants further studies to validate whether observed glutamine dependency is a universal phenomenon in other NF1 associated malignancies such as leukemia and gliomas. Nonetheless, data from our study provides a strong rationale to explore targeted glutaminase inhibition as a potential therapeutic approach in the NF1 disease setting. In fact based on these results a single agent CB-839 study for patients with NF1 mutant MPNST, as well as a “basket” study for all NF1 mutant patients, are now planned.
MATERIALS AND METHODS
Chemicals and drugs
CB-839 was provided by Calithera Biosciences. BPTES was purchased from Sigma Aldrich. Both the drugs were dissolved in dimethyl sulfoxide (DMSO) for in vitro studies and stored at -20°C.
Cell culture and reagents
NF1 null ST8814 and S462 cell lines were a kind gift by Dr. Karen Cichowski (BWH Biomedical Research Institute, Boston, MA). NF1 mutant MPNST cell line has been described elsewhere [18]. STS26T, a wild-type NF1 harboring MPNST cell line, was a generous gift by Dr. Steven. A Porcelli (Albert Einstein College of Medicine, Bronx, NY). De-differentiated Liposarcoma cell line (LS141) was obtained from Dr. Samuel Singer (Memorial Sloan Kettering Cancer Center (MSKCC), New York, NY). Ewing Sarcoma (CHP100) cell line was obtained from Dr. Melinda S. Merchant (Center for Cancer Research, NCI/NIH, Bethesda, MD). MeWo cell line was purchased from American Type Culture Collection All the cell lines were cultured in RPMI media with 10% FBS, 100 U/mL penicillin, and 100 mg/mL streptomycin, maintained at 37°C in 5% CO2, and passaged for no more than 4 months. Initial stocks of all cell lines were received from their sources within the past 3 years. Cell lines were determined to be mycoplasma free using MycoAlert Mycoplasma detection Kit (Lonza).
Cell viability assays
For Glutamine depletion experiments, cells were plated in 96-well plates at 1,500-2,000 cells per well density in 0.1 ml Glutamine free RPMI media, and then treated with the indicated drugs in RPMI containing 2mMol/L Glutamine the next day for an additional 72 hours. Cell viability assays were carried out using the Dojindo Molecular Technologies (CCK-8) kit as per manufacturer’s instructions. The optical density was read at 450nm using a Spectra Max 340 PC (Molecular Devices Corporation). Cell viability is expressed as a percentage of untreated cells. Half maximal inhibitory concentrations (IC50) were extrapolated from cell viability data using CompuSyn software according to the manufacturer’s instructions.
Western immunoblotting
Cells were lysed with 1X radioimmunoprecipitation assay (RIPA) buffer (Cell Signaling Technologies) supplemented with protease inhibitor cocktail tablets (Roche Diagnostics) and 1mM Na3VO4 (Sodium orthovanadate). Equal amounts (20-30μg) of protein were electrophoresed onto 4% to 12% gradient gels (Life Technologies) and transferred onto polyvinylidene difluoride (PVDF) membrane (Immobilon-FL, catalog # IPFL00010). Membranes were blocked in 1X Tris-buffered saline (TBS) containing 5% non-fat dry milk and 0.1% Tween 20 (TBS-T), and probed with primary antibodies. A complete list of antibodies used in the study is included in the supplementary methods. Bound antibodies were detected with horseradish peroxidase secondary antibodies (GE Healthcare) and visualized by Enhanced Chemiluminescence Reagent (GE Healthcare).
Tissues from xenograft experiment were homogenized using a mortar and pestle. Lysates were prepared using sample grinding kit form GE-healthcare (catalog # 80-6483-37), lysed with radioimmunoprecipitation assay (RIPA) buffer and western blotting was carried out as described above.
Quantitation of organic acids by GC-MS
Standards for organic acids (Succinic acid, fumaric acid, alpha keto glutaric acid, citric acid, 2 hydroxyglutaric acid and amino acids (asparagine, aspartic acid, glutamic acid and glutamine) were purchased from Sigma (St. Louis, MO). Internal standard, succinate d4 was obtained from CDN Isotopes, Quebec, Canada. N, O Bis (trimethylsilyl) trifluoroacetamide (BSTFA) + 1% Trimethylsilyl chloride (TMCS) were purchased from Regis Technologies (Morton Grove, IL). Mox reagent (methoxamine in pyridine) was purchased form Supelco (St. Louis, MO). All other chemicals were of analytical reagent grade.
Cells were plated at a density of 3000 cells per plate in 60mm plates (Corning) in Glutamine free RPMI 1640 media supplemented with 10% FBS. 18 h after seeding, cells were treated with indicated drug in RPMI containing 2mM Glutamine. After 48 hours, culture medium was aspirated and attached cells were washed twice with cold saline water (9 gm/L NaCl) and quenched with 1.5ml ice-cold methanol. After 5 minutes of incubation on ice, cells were collected by scraping with disposable cell scraper (Fisher Scientific) and collected in glass tubes with Teflon sealed caps (Corning Inc. catalog # 9826-16X). One volume of chloroform was added to the cell suspension and vortexed at 4°C for 10 seconds. 1.5 ml water was added and vortexed for another 1 minute. Samples were centrifuged at 2000 rpm at 4ºC for 20 min and the aqueous phase was collected in a new tube and evaporated under airflow at 37ºC with nitrogen using an evaporator (Turbovap).
The extracted samples were evaporated under nitrogen stream prior to derivatization. The derivatization involves the addition of the internal standard, succinate d4 followed by 50uL of methoxyamine-HCl in pyridine and incubating for 90 min at 37°C. The final derivatization step involves the addition of 80uL of BSTFA+1% TMCS and incubating at 60°C for 30min.
GC-MS analysis
GCMS analysis was performed using an Agilent 7000 triple quadrupole mass spectrometer coupled to an Agilent 7693 GC with an autosampler (Agilent Technologies, Palo Alto, CA).
Two microliters of the sample was injected onto a GC column (30mx0.2mmx0.25um) at a flow rate of 1mL/min. The initial temperature of the oven was set to 70°C kept for 1min followed by a ramp of 5°C/min to 200°C, held for 1 min and increasing to 300°C at 40°C/min with a total run time of 33.5min. The compounds were scanned in EI and spilt less injection mode by selected ion monitoring (SIM) using the following diagnostic ions: succinate, m/z 247; fumarate, m/z 245; alpha ketoglutarate, m/z 288; 2 hydroxyglutaric acid, m/z 247; citrate, m/z 273; glutamate, m/z 246; aspartic acid, m/z 232; asparagine m/z 231; glutamine, m/z 245; and succinate d4, m/z 251. Data acquisition, peak integration and quantitation were done by Mass Hunter software (version B. 07.04). The intra-assay precision was in the range of 1.5-6.0% and the accuracy was between 92-106%. The inter-assay precision and accuracy was in the range of 4.0-9.1% and 92-110% respectively.
Gene silencing
Cells were plated at 50–60% confluency in 60 mm plates (Corning) and incubated for 24 h. Cells were then transiently transfected using Lipofectamine RNAiMAX (Invitrogen) using pooled siRNAs (GE Dharmacon). Nonspecific scrambled siRNA was used as control. After 48 hours of transfection, cells were lysed for western immunoblotting analysis.
Detection of Ras activity
To determine the activity of Ras, EZ-Detect Ras Activation Kit (catalog # 89855, Thermo Scientific Pierce, Rockford, IL) was used according to manufacturer’s protocol. Briefly, cells were plated in RPMI+10%FBS without Glutamine and grown to approximately 60% confluency. Next day, media was replaced with fresh RPMI+10%FBS either without Glutamine or RPMI+10%FBS containing indicated reagents and incubated for another 48 hours. Cell lysates were prepared using the lysis buffer provided in the kit and then treated with GTPγS or GDP to activate or inactivate Ras, respectively. The nucleotide exchange reaction was terminated within 15 minutes by placing the samples on ice. The lysates were then incubated with a GST-fusion protein containing the Ras Binding Domain (RBD) of Raf1 to pull down active Ras. Detection of Ras was done by immunoblotting using pan-Ras antibody provided with the kit
Xenograft studies
Briefly, MPNST xenografts were transplanted subcutaneously in the flank of ICR/SCID mice. Once tumors reached a volume of 80–100mm3, the mice were randomized into two groups of 7–10 animals each and treated with vehicle control or CB-839 (200mg/kg/day) orally for 2 weeks. Vehicle and CB-839 were prepared as described previously [28]. Tumor size was measured twice weekly by caliper. The average tumor volume in each group was expressed in cubic millimeter and calculated using the formula p/6 × (large diameter) × (small diameter) [2]. Animals were sacrificed after 3 weeks of drug treatment and the resected tumor tissues were snap frozen for western blot analysis. Experiments were carried out under an Institutional Animal Care and Use Committee-approved protocol, and institutional guidelines for the proper and humane use of animals were followed.
Statistical analysis
All in vitro experiments were carried out at least three times unless otherwise indicated. P-values were calculated using Student’s T-test, with values of ≤0.05 determined to be statistically significant. Standard error was calculated as the standard deviation divided by the square root of the number of samples tested.
ACKNOWLEDGMENTS
The authors would like to thank Calithera Biosciences for providing CB-839 and helpful scientific discussions. The authors would also like to thank Dr. Elisa de Stanchina (MSKCC) for help with tumor xenograft experiments.
CONFLICTS OF INTEREST
The authors have no conflicts to disclose in relation to this work.
FINANCIAL SUPPORT
Funding for this work was provided by the Herbert Irving Comprehensive Cancer Center.
REFERENCES
1. Warburg O. On the origin of cancer cells. Science. 1956; 123:309-14.
2. Koppenol WH, Bounds PL, Dang CV. Otto Warburg’s contributions to current concepts of cancer metabolism. Nat Rev Cancer. 2011; 11:325-37.
3. Hensley CT, Wasti AT, DeBerardinis RJ. Glutamine and cancer: cell biology, physiology, and clinical opportunities. J Clin Invest. 2013; 123:3678-84.
4. DeBerardinis RJ, Cheng T. Q’s next: the diverse functions of glutamine in metabolism, cell biology and cancer. Oncogene. 2010; 29:313-24.
5. DeBerardinis RJ, Mancuso A, Daikhin E, Nissim I, Yudkoff M, Wehrli S, Thompson CB. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc Natl Acad Sci U S A. 2007; 104:19345-50.
6. Le A, Lane AN, Hamaker M, Bose S, Gouw A, Barbi J, Tsukamoto T, Rojas CJ, Slusher BS, Zhang H, Zimmerman LJ, Liebler DC, Slebos RJ, et al. Glucose-independent glutamine metabolism via TCA cycling for proliferation and survival in B cells. Cell Metab. 2012; 15:110-21.
7. Duran RV, Oppliger W, Robitaille AM, Heiserich L, Skendaj R, Gottlieb E, Hall MN. Glutaminolysis activates Rag-mTORC1 signaling. Mol Cell. 2012; 47:349-58.
8. Deberardinis RJ, Sayed N, Ditsworth D, Thompson CB. Brick by brick: metabolism and tumor cell growth. Curr Opin Genet Dev. 2008; 18:54-61.
9. Xiang Y, Stine ZE, Xia J, Lu Y, O'Connor RS, Altman BJ, Hsieh AL, Gouw AM, Thomas AG, Gao P, Sun L, Song L, Yan B, et al. Targeted inhibition of tumor-specific glutaminase diminishes cell-autonomous tumorigenesis. J Clin Invest. 2015; 125:2293-306.
10. Jin L, Alesi GN, Kang S. Glutaminolysis as a target for cancer therapy. Oncogene. 2016; 35:3619-25.
11. Chen L, Cui H. Targeting glutamine induces apoptosis: a cancer therapy approach. Int J Mol Sci. 2015; 16:22830-55.
12. Ferner RE. Neurofibromatosis 1. Eur J Hum Genet. 2007; 15:131-8.
13. Rad E, Tee AR. Neurofibromatosis type 1: fundamental insights into cell signalling and cancer. Semin Cell Dev Biol. 2016; 52:39-46.
14. Tonsgard JH. Clinical manifestations and management of neurofibromatosis type 1. Semin Pediatr Neurol. 2006; 13:2-7.
15. McCaughan JA, Holloway SM, Davidson R, Lam WW. Further evidence of the increased risk for malignant peripheral nerve sheath tumour from a Scottish cohort of patients with neurofibromatosis type 1. J Med Genet. 2007; 44:463-6.
16. Durbin AD, Ki DH, He S, Look AT. Malignant peripheral nerve sheath tumors. Adv Exp Med Biol. 2016; 916:495-530.
17. Altman BJ, Stine ZE, Dang CV. From Krebs to clinic: glutamine metabolism to cancer therapy. Nat Rev Cancer. 2016; 16:619-34.
18. Ambrosini G, Cheema HS, Seelman S, Teed A, Sambol EB, Singer S, Schwartz GK. Sorafenib inhibits growth and mitogen-activated protein kinase signaling in malignant peripheral nerve sheath cells. Mol Cancer Ther. 2008; 7:890-6.
19. Sun D, Tainsky MA, Haddad R. Oncogene mutation survey in MPNST cell lines enhances the dominant role of hyperactive Ras in NF1 associated pro-survival and malignancy. Transl Oncogenomics. 2012; 5:1-7.
20. Barkan B, Starinsky S, Friedman E, Stein R, Kloog Y. The Ras inhibitor farnesylthiosalicylic acid as a potential therapy for neurofibromatosis type 1. Clin Cancer Res. 2006; 12:5533-42.
21. Barretina J, Taylor BS, Banerji S, Ramos AH, Lagos-Quintana M, Decarolis PL, Shah K, Socci ND, Weir BA, Ho A, Chiang DY, Reva B, Mermel CH, et al. Subtype-specific genomic alterations define new targets for soft-tissue sarcoma therapy. Nat Genet. 2010; 42:715-21.
22. Kanojia D, Nagata Y, Garg M, Lee DH, Sato A, Yoshida K, Sato Y, Sanada M, Mayakonda A, Bartenhagen C, Klein HU, Doan NB, Said JW, et al. Genomic landscape of liposarcoma. Oncotarget. 2015; 6:42429-44. https://doi.org/10.18632/oncotarget.6464.
23. Reuss DE, Mucha J, Hagenlocher C, Ehemann V, Kluwe L, Mautner V, von Deimling A. Sensitivity of malignant peripheral nerve sheath tumor cells to TRAIL is augmented by loss of NF1 through modulation of MYC/MAD and is potentiated by curcumin through induction of ROS. PLoS One. 2013; 8:e57152.
24. Slotkin EK, Patwardhan PP, Vasudeva SD, de Stanchina E, Tap WD, Schwartz GK. MLN0128, an ATP-competitive mTOR kinase inhibitor with potent in vitro and in vivo antitumor activity, as potential therapy for bone and soft-tissue sarcoma. Mol Cancer Ther. 2015; 14:395-406.
25. Nissan MH, Pratilas CA, Jones AM, Ramirez R, Won H, Liu C, Tiwari S, Kong L, Hanrahan AJ, Yao Z, Merghoub T, Ribas A, Chapman PB, et al. Loss of NF1 in cutaneous melanoma is associated with RAS activation and MEK dependence. Cancer Res. 2014; 74:2340-50.
26. Griewank KG, Yu X, Khalili J, Sozen MM, Stempke-Hale K, Bernatchez C, Wardell S, Bastian BC, Woodman SE. Genetic and molecular characterization of uveal melanoma cell lines. Pigment Cell Melanoma Res. 2012; 25:182-7.
27. Duran RV, Hall MN. Glutaminolysis feeds mTORC1. Cell Cycle. 2012; 11:4107-8.
28. Gross MI, Demo SD, Dennison JB, Chen L, Chernov-Rogan T, Goyal B, Janes JR, Laidig GJ, Lewis ER, Li J, Mackinnon AL, Parlati F, Rodriguez ML, et al. Antitumor activity of the glutaminase inhibitor CB-839 in triple-negative breast cancer. Mol Cancer Ther. 2014; 13:890-901.
29. Jacque N, Ronchetti AM, Larrue C, Meunier G, Birsen R, Willems L, Saland E, Decroocq J, Maciel TT, Lambert M, Poulain L, Hospital MA, Sujobert P, et al. Targeting glutaminolysis has antileukemic activity in acute myeloid leukemia and synergizes with BCL-2 inhibition. Blood. 2015; 126:1346-56.
30. Seltzer MJ, Bennett BD, Joshi AD, Gao P, Thomas AG, Ferraris DV, Tsukamoto T, Rojas CJ, Slusher BS, Rabinowitz JD, Dang CV, Riggins GJ. Inhibition of glutaminase preferentially slows growth of glioma cells with mutant IDH1. Cancer Res. 2010; 70:8981-7.
31. Bollag G, Clapp DW, Shih S, Adler F, Zhang YY, Thompson P, Lange BJ, Freedman MH, McCormick F, Jacks T, Shannon K. Loss of NF1 results in activation of the Ras signaling pathway and leads to aberrant growth in haematopoietic cells. Nat Genet. 1996; 12:144-8.
32. Hiatt KK, Ingram DA, Zhang Y, Bollag G, Clapp DW. Neurofibromin GTPase-activating protein-related domains restore normal growth in Nf1-/- cells. J Biol Chem. 2001; 276:7240-5.
33. Morgan KJ, Rowley MA, Wiesner SM, Hasz DE, Van Ness B, Largaespada DA. The GAP-related domain of neurofibromin attenuates proliferation and downregulates N- and K-Ras activation in Nf1-negative AML cells. Leuk Res. 2007; 31:1107-13.
34. Patwardhan PP, Surriga O, Beckman MJ, de Stanchina E, Dematteo RP, Tap WD, Schwartz GK. Sustained inhibition of receptor tyrosine kinases and macrophage depletion by PLX3397 and rapamycin as a potential new approach for the treatment of MPNSTs. Clin Cancer Res. 2014; 20:3146-58.
35. Matre P, Velez J, Jacamo R, Qi Y, Su X, Cai T, Chan SM, Lodi A, Sweeney SR, Ma H, Davis RE, Baran N, Haferlach T, et al. Inhibiting glutaminase in acute myeloid leukemia: metabolic dependency of selected AML subtypes. Oncotarget. 2016; 7:79722-35. https://doi.org/10.18632/oncotarget.12944.
36. Elgogary A, Xu Q, Poore B, Alt J, Zimmermann SC, Zhao L, Fu J, Chen B, Xia S, Liu Y, Neisser M, Nguyen C, Lee R, et al. Combination therapy with BPTES nanoparticles and metformin targets the metabolic heterogeneity of pancreatic cancer. Proc Natl Acad Sci U S A. 2016; 113:E5328-36.
37. Kraniak JM, Sun D, Mattingly RR, Reiners JJ Jr, Tainsky MA. The role of neurofibromin in N-Ras mediated AP-1 regulation in malignant peripheral nerve sheath tumors. Mol Cell Biochem. 2010; 344:267-76.
38. Kim A, Stewart DR, Reilly KM, Viskochil D, Miettinen MM, Widemann BC. Malignant peripheral nerve sheath tumors state of the science: leveraging clinical and biological insights into effective therapies. Sarcoma. 2017; 2017:7429697.
39. Bakker AC, La Rosa S, Sherman LS, Knight P, Lee H, Pancza P, Nievo M. Neurofibromatosis as a gateway to better treatment for a variety of malignancies. Prog Neurobiol. 2017; 152:149-65.
40. De Vitto H, Perez-Valencia J, Radosevich JA. Glutamine at focus: versatile roles in cancer. Tumour Biol. 2016; 37:1541-58.
41. Martinez-Outschoorn UE, Peiris-Pages M, Pestell RG, Sotgia F, Lisanti MP. Cancer metabolism: a therapeutic perspective. Nat Rev Clin Oncol. 2017; 14:11-31.
42. Erickson JW, Cerione RA. Glutaminase: a hot spot for regulation of cancer cell metabolism? Oncotarget. 2010; 1:734-40. https://doi.org/10.18632/oncotarget.208.
43. Katt WP, Cerione RA. Glutaminase regulation in cancer cells: a druggable chain of events. Drug Discov Today. 2014; 19:450-7.
44. McDermott LA, Iyer P, Vernetti L, Rimer S, Sun J, Boby M, Yang T, Fioravanti M, O'Neill J, Wang L, Drakes D, Katt W, Huang Q, Cerione R. Design and evaluation of novel glutaminase inhibitors. Bioorg Med Chem. 2016; 24:1819-39.
45. Wang JB, Erickson JW, Fuji R, Ramachandran S, Gao P, Dinavahi R, Wilson KF, Ambrosio AL, Dias SM, Dang CV, Cerione RA. Targeting mitochondrial glutaminase activity inhibits oncogenic transformation. Cancer Cell. 2010; 18:207-19.
46. Momcilovic M, Bailey ST, Lee JT, Fishbein MC, Magyar C, Braas D, Graeber T, Jackson NJ, Czernin J, Emberley E, Gross M, Janes J, Mackinnon A, et al. Targeted inhibition of EGFR and glutaminase induces metabolic crisis in EGFR mutant lung cancer. Cell Rep. 2017; 18:601-10.
47. Thompson RM, Dytfeld D, Reyes L, Robinson RM, Smith B, Manevich Y, Jakubowiak A, Komarnicki M, Przybylowicz-Chalecka A, Szczepaniak T, Mitra AK, Van Ness BG, Luczak M, Dolloff NG. Glutaminase inhibitor CB-839 synergizes with carfilzomib in resistant multiple myeloma cells. Oncotarget. 2017; 8:35863-76. https://doi.org/10.18632/oncotarget.16262.
48. Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, Marks KM, Prins RM, Ward PS, et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature. 2009; 462:739-44.
49. Intlekofer AM, Dematteo RG, Venneti S, Finley LW, Lu C, Judkins AR, Rustenburg AS, Grinaway PB, Chodera JD, Cross JR, Thompson CB. Hypoxia induces production of L-2-hydroxyglutarate. Cell Metab. 2015; 22:304-11.
50. Zhang J, Pavlova NN, Thompson CB. Cancer cell metabolism: the essential role of the nonessential amino acid, glutamine. EMBO J. 2017; 36:1302-15.
51. Son J, Lyssiotis CA, Ying H, Wang X, Hua S, Ligorio M, Perera RM, Ferrone CR, Mullarky E, Shyh-Chang N, Kang Y, Fleming JB, Bardeesy N, et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature. 2013; 496:101-5.

