INTRODUCTION
According to a recent report, carcinoma is a major cause of mortality, accounting for the death of 8.2 million people and 14.1 million newly diagnosed cancer cases worldwide in the year 2102 [1]. The mechanism of tumor formation varies in different cancers, but carcinoma with metastasis is an important prognostic factor, and the presence or absence of metastasis determines treatment strategies [2, 3].
A majority of malignant tumor cases can ultimately develop metastases, which include lymph node metastases (LNM) and distant metastases (DM). Most cancers are diagnosed at an advanced stage, and the rate of lymph node metastasis or distant metastasis remains high. Until now, the precise metastasis mechanism of cancer cells is still unclear. In recent years, microRNAs (miRNAs), which act as gene expression regulators by regulating messenger RNA translation and degradation, have been found to have an effect on the occurrence and development of cancer. Accordingly, they have become a hot research topic, and their role in cancer is being recognized by more and more scientists. A better understanding of the molecular mechanism underlying carcinoma development, invasion and metastasis is of great clinical value to find reliable diagnostic markers and develop novel therapeutic strategies.
MicroRNA-10b (miR-10b) has been reported to be dysregulated in certain types of cancer and to play an important role in invasion and metastasis [4–6]. Previous studies show that miR-10b is up-regulated in breast cancer with distant metastasis, liver, pancreatic and esophageal cancers, and in glioma tissues [7–9]. Its expression was higher in cancerous tissues compared with paired noncancerous tissues or adjacent tissues. Also, some studies have demonstrated that the expression level of miR-10b was up-regulated in cancerous tissues compared to non-neoplastic tissue or adjacent tissue, and was associated with some clinical features, including LNM and DM. Moreover, there were associations between miR-10b expression and certain clinical characteristics of cancer patients. However, even though miR-10b may act as a common molecular marker for both LNM and DM, currently, we still know nothing about its role in predicting metastases of various cancers. Therefore, in the present study, the relevant literature was retrieved and evaluated according to the inclusion and exclusion criteria. The studies that met the criteria were included in this meta-analysis. The correlation between the expression level of miR-10b and metastasis status was analyzed and to further determine whether miR-10b could be a promising noninvasive biomarker to predict the prognosis of cancer patients.
RESULTS
Studies’ characteristics
The literature retrieval process is depicted in detail in Figure 1. A total of 9 studies [10–18] were ultimately found to be eligible according to the selection criteria and included in this meta-analysis. A total of 962 patients were included in the present meta-analysis, and the mean sample size of patients per study was 106.9 (range:34–436). Among the 9 studies, 8 were from the People’s Republic of China and one was from Japan. Five different cancer types were covered in this meta-analysis, including one breast cancer, five non-small cell lung cancer, one gastric cancer, one colorectal cancer and one hepatocellular carcinoma. All cancerous specimens were well preserved before RNA extraction. The diagnoses of LNM and DM were all based on pathological examination.
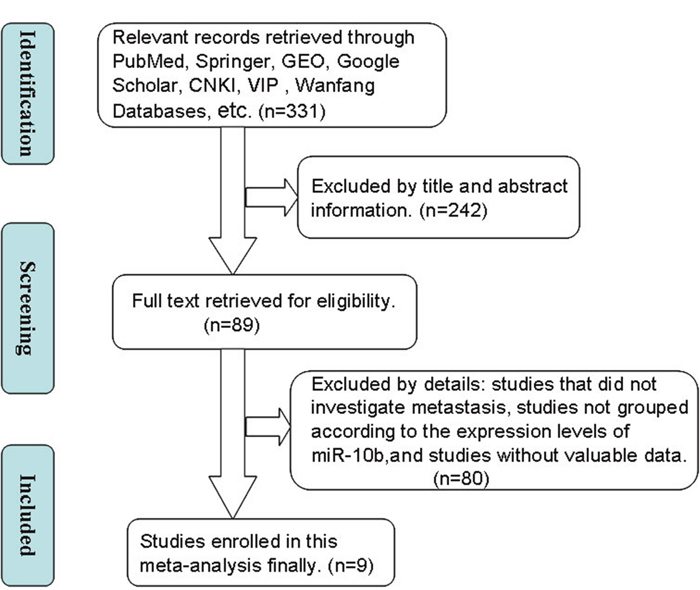
Figure 1: A flowchart presenting the steps of literature retrieval and selection.
CKNI, China National Knowledge Infrastructure; VIP, Chongqing VIP Information Network; GEO, Gene expression Omnibus.
There were eight studies that reported on the association between the expression of miR-10b and LNM occurrence [10, 11, 13–18], and six articles focused on the association of miR-10b expression and distant metastases [10, 12–15, 17]. The detection methods, namely reverse transcription polymerase chain reaction was used to determine the expression levels of miR-10b in cancerous tissues. The criteria of high miR-10b expression in all included studies are shown in Table 1. All studies were divided into two groups (high miR-10b expression group and low miR-10b expression group).
Table 1: The basic information and data of all included studies in the meta-analysis
Author(year) |
Country |
Race |
Cancer type |
Total number |
miR-10b expression |
Detection method |
Criterion of high expression |
|||
|---|---|---|---|---|---|---|---|---|---|---|
High LNM/DM |
High |
Low LNM/DM |
Low |
|||||||
Nishida et al (2012) [13] |
JAPAN |
Asian |
CRC |
88 |
21/10 |
44 |
15/7 |
44 |
qRT PCR |
above median |
Luan et al (2014) [18] |
CHINA |
Asian |
NSCLC |
50 |
15/- |
25 |
6/- |
25 |
qRT PCR |
RQ>1.18 |
Wang et al (2013) [14] |
CHINA |
Asian |
GC |
436 |
151/40 |
183 |
119/21 |
253 |
qRT PCR |
Staining index score≥4 |
Zhang et al (2014) [16] |
CHINA |
Asian |
NSCLC |
73 |
21/- |
37 |
5/- |
36 |
qRT PCR |
HOTAIR/GAPDH ≥1.0 |
Li et al (2012) [12] |
CHINA |
Asian |
HCC |
34 |
-/13 |
17 |
-/7 |
17 |
qRT PCR |
above median |
Li YW et al (2016) [11] |
CHINA |
Asian |
NSCLC |
58 |
23/- |
25 |
21/- |
33 |
qRT PCR |
2-ΔCt≥0.003 |
Huang et al (2015) [10] |
CHINA |
Asian |
NSCLC |
114 |
45/7 |
55 |
34/6 |
59 |
qRT PCR |
ΔCt>range/2 |
Yang et al (2015) [15] |
CHINA |
Asian |
NSCLC |
74 |
30/6 |
64 |
1/0 |
10 |
qRT PCR |
RQ≥1.15 |
Cheng et al (2012) [17] |
CHINA |
Asian |
BC |
35 |
17/14 |
18 |
9/6 |
17 |
qRT PCR |
Fold control >23.73 |
The dashes represent no data.
BC, breast cancer; CRC, colorectal cancer; HCC, hepatocellular carcinoma; NSCLC, non-small cell lung cancer; GAPDH, glyceraldehyde 3-phosphate dehydrogenase; GC, gastric cancer; DM, distant metastases; LNM, lymph node metastases; RT-PCR, reverse transcription polymerase chain reaction; RQ=2-ΔΔCT; Staining index score:staining intensity score and the proportion of positive tumor cells.
Association between miR-10b expression level and LNM
Eight studies reported the number of patients with LNM based on different miR-10b expression levels, which comprised 928 cases. The fixed-effects model was adopted because there was limited heterogeneity across studies (I2=15%, PQ=0.31). The odds ratio (OR), expressed as the high miR-10b expression group versus low miR-10b expression group, was 4.65 (95%CI: 3.40–6.37, P<0.00001) is shown in Figure 2. When the LNM incidence in the cancer patients was compared between the two groups, the results of the meta-analysis revealed that the difference in the LNM incidence between the high and low expression groups was statistically significant. This result demonstrated that cancer patients with high miR-10b expression in cancerous tissues were more prone to develop LNM.
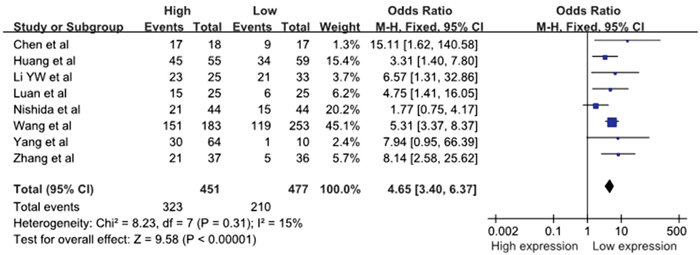
Figure 2: A forest plot for the association between the miR-10b expression levels with LNM.
LNM, lymph node metastases; M–H, Mantel–haenszel test; df, degrees of freedom.
Association between miR-10b expression level and DM
Six studies reported the number of patients with DM based on different miR-10b expression levels, which comprised 781 patients. There was no significant heterogeneity among the studies (I2=0%, PQ=0.46); thus, the fixed-effects model was adopted. The analysis showed a pooled OR =2.70 (95% CI: 1.79–4.08, P<0.00001), as shown in Figure 3. Compared with the low miR-10b expression group, the DM rate was significantly increased in the high miR-10b expression group. The result revealed that the high miR-10b expression level in the patients’ tumor tissues may be an increased risk of developing DM. It must be mentioned that, due to the relatively small heterogeneity across studies on distant metastasis and the limited number of included studies, the sensitivity analysis and assessment of publication bias were not performed.
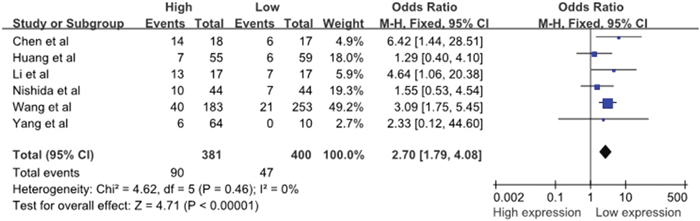
Figure 3: A forest plot for the association between the miR-10b expression levels with DM.
DM, distant metastases; M–H, Mantel–haenszel test; df, degrees of freedom.
Publication bias
For the meta-analysis of the association between the miR-10b expression level and LNM occurrence, the funnel plot was slightly asymmetrical, as shown in Figure 4. Additionally, the statistical assessment of publication bias using the Begg’s test (Pr>|z|=0.266) and the Egger’s test (P>|t|=0.609, 95% CI: -1.602 to 2.509), as shown in Figure 5, revealed no publication bias (P>0.05). The trim and fill method was also adopted to test for publication bias, and the results of this test suggested that the meta-analysis results were identical.
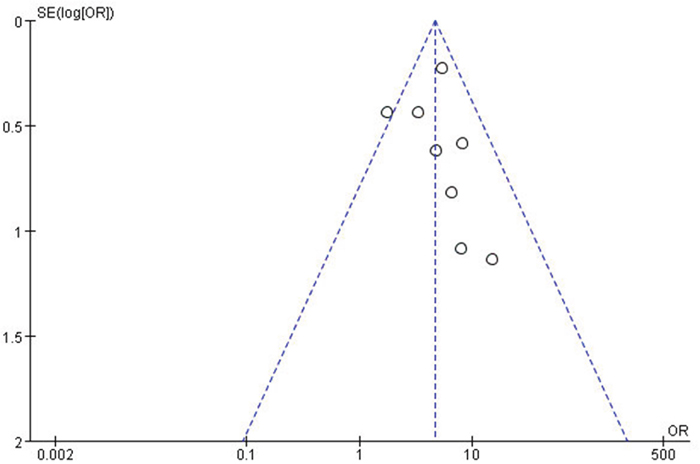
Figure 4: A funnel plot analysis of potential publication bias.
SE, standard error; OR, odds ratio.
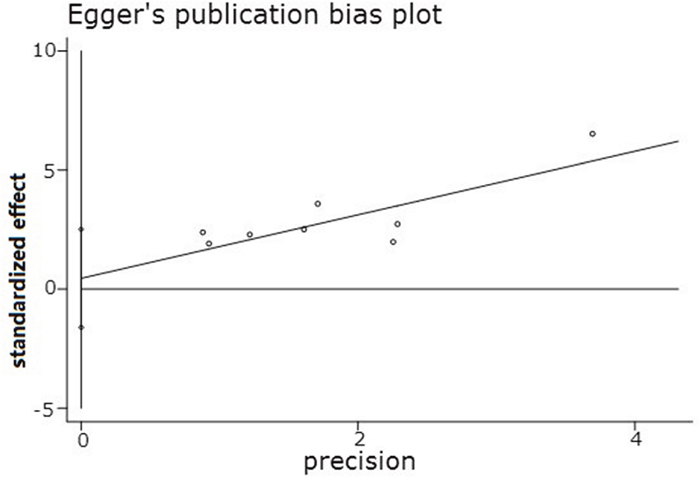
Figure 5: Egger’s publication bias plot.
Sensitivity analysis
For the meta-analysis of the association between the miR-10b expression level and LNM occurrence, a sensitivity analysis was performed by deleting each study in turn from the pooled analysis. It aimed to test the influence of the removed data set on the overall ORs. The result was not significantly influenced by the exclusion of any of the studies, suggesting that the result of the synthetic analysis was robust.
DISCUSSION
The LNM is the most common metastasis pathway in most cancers, and distant metastasis often occurs in the advanced stages of cancer. LNM and DM are positively important for diagnosis in tumor–node–metastasis staging and the treatment strategy for cancer patients, as well as indicators for predicting prognosis. Accordingly, there is a need to identify new, prognostic biomarkers for cancer to establish prognostic sets needing different therapeutic intensity.
MiRNAs are members of a family of endogenous, highly conserved, small non-coding RNA molecules (18–24 nucleotides) that regulate the expression of a wide variety of genes and have similar physiological functions [19]. Through base pairing with the 3’-untranslated region (3’-UTR) of target genes, miRNAs enhance mRNA degradation or inhibit posttranscriptional translation [20]. miRNAs can direct a wide variety of normal biological mechanisms, such as embryonic development, cell growth, apoptosis and differentiation. MiRNAs have been found to regulate both oncogenes, such as ras, and tumor suppressor genes, such as PTEN [21, 22]. Although miRNAs have been widely investigated in recent years, the molecular regulatory mechanisms of miRNAs and their roles in the development of cancers remain largely unknown and need further investigation.
MiR-10b has been shown to be dysregulated in certain types of cancer and to play an important role in invasion and metastasis [7–9, 23, 24]. However, the exact mechanism of miR-10b in cancer is not completely understood. Overexpression of miR-10b was shown to exert multiple effects on various cancers, through different mechanisms. In addition, miR-10b was shown to be highly expressed in metastatic breast cancer cell lines as well as in patients’ metastatic breast tumors. Upregulation of miR-10b, which is induced by Twist, inhibits the translation of HOXD10, a transcription factor known for its roles in cell motility, resulting in increased expression of the prometastatic gene RHOC [24]. Silencing of miR-10b, both in vitro and in vivo, with antagomirs significantly decreases miR-10b levels and increases the levels of HOXD10, thereby inhibiting metastasis [25].
MiR-10b also plays an important role in the invasion and metastasis of nasopharyngeal carcinoma cells. For instance, after transfection of the miR-10b inhibitor, the ability of nasopharyngeal carcinoma cell lines to migrate and invade was reduced [26]. Also, through regulation of Kruppel-like factor 4 (KLF4) expression in human esophageal cancer cell lines, miR-10b was found to induce tumor migration and invasion [27]. Additionally, the loss of CADM1, a tumor suppressor gene involved in cell–cell interactions and epithelial-like phenotype, can contribute to cancer invasion and metastasis in epithelial-derived cancers by inducing epithelial mesenchymal transition (EMT) [4, 28, 29]. Indeed, downregulation or loss of CADM1 expression is frequently observed in various cancers, such as NSCLC, HCC, cervix, prostate and pancreas cancer [30–35], especially in those with LNM and DM [36]. Multiple studies showed that miR-10b regulates CADM1 expression at the post transcriptional level. Loss of Cadherin 1 (CDH1), a glycoprotein responsible for calcium-dependent cell-to-cell adhesion [37], can also induced EMT, and its downregulation is associated with tumor grade and stage and contributes to the transition of adenoma to carcinoma in animal models [38]. The research by Zhang et al. revealed that the CDH1mRNA and protein were overexpressed in miR-10b-suppressed NSCLC cells compared with controls, suggesting that miR-10b might be necessary to drive the expression of CDH1 [39]. In addition, target genes, such as MMP-9, vimentin (VIM), and CDH1, which were found to be differently expressed in the nasopharyngeal carcinoma cells transfected with miR-10b mimics and with miR-10b inhibitors, were also involved in migration and invasion. Moreover, miR-10b was also confirmed to be overexpressed in malignant glioma, hepatocellular carcinoma, and pancreatic cancer [40, 41].
This study demonstrated that the high expression of miR-10b was significantly correlated with DM and regional lymph node involvement. This indicated that overexpression of miR-10b was closely related to the aggressive behavior of tumor cells, thus suggesting miR-10b as a promising biomarker to predict the prognosis of cancer patients. In future experiments, the exact mechanism whereby miR-10b exerts its effects on promoting tumor cell invasion and metastasis will be further investigated.
In assessing the relationship between miR-10b expression and LNM, the heterogeneity tests revealed mild heterogeneity. The heterogeneity may originate from different tumor types and different definitions of high expression of miR-10b. At the same time, some small sample studies may also be the cause of heterogeneity. Additionally, the proportion of advanced tumors in different research centers may vary, and can also be a source of heterogeneity.
Since most of the studies enrolled lacked miR-10b expression data based on different ages and tumor stages, no meaningful subgroup analysis was conducted.
Almost all relevant studies were browsed in full text and carefully evaluated. Unfortunately, in many studies, patients (especially those of the European and African populations) were not grouped according to the level of miR-10b expression to described their clinicopathological parameters. Many researchers detected tissue levels of miR-10b, based on LNM and DM. Thus, there is a lack of pathological parameters related to metastasis based on different levels of miR-10b expression, and more data are needed to verify the results of this study in the future.
MATERIALS AND METHODS
Literature retrieval strategy
To obtain the potentially eligible studies, systematic online literature search was conducted against multiple databases, including PubMed, Springer, Google Scholar, Gene expression Omnibus, China National Knowledge Infrastructure (CNKI), Chongqing VIP Information Network, and Wanfang. The keywords for the search were as follows: “miR-10b”, “microRNA-10b”, “Prognosis”, “Clinicopathology” and “metastasis”. The relevant documents found were retrieved, using April 1, 2017, as the retrieval period deadline. In addition, other relevant articles were also obtained by manually viewing the reference list.
Inclusion and exclusion criteria
The inclusion criteria for the articles were as follows: 1) the role of miR-10b in the development of human carcinoma was investigated; 2) the expression level of miR-10b in primary cancerous tissue was measured; 3) patients were grouped according to the expression level of miR-10b; 4) related clinicopathologic parameters were reported in the study. The exclusion criteria for the articles were as follows: 1) duplicate study publications; 2) studies without valuable data or data acquired through animal experiments; 3) reviews, letters, case reports, and expert opinions; 4) the expression level of miR-10b is detected in serum.
Date extraction and quality assessment
The data and information from all eligible studies were independently extracted by two investigators (RB Liao and BX Tong) through cross-check. The following data and information were collected from each study: author, publication year, country, race, cancer type, total number of patients, number of patients with high miR-10b expression in the group and with low miR-10b expression in the group, number of patients with LNM and DM in each group, and the criteria for high miR-10b expression. If there were disagreements, a consensus was reached by a third investigator (HJ Wu). The Newcastle-Ottawa Scale (NOS) was applied to assess the quality of all included studies. The NOS scores ranged from 0 to 9, and the study with an NOS score ≥ 6 was considered to be of high quality. The quality of all studies included in this meta-analysis was varied from 5 to 9, with a mean value of 6.7.
Statistical methods
The present meta-analysis was conducted using the RevMan5.2 software and Stata SE12.0 software. The heterogeneity among the enrolled studies was determined by the chi-square-based Q-test and I2 statistics; a P-value for the Q-test <0.05 and a I2-value>50% were considered as indicators of severe heterogeneity. The random-effects model was applied to the studies with a significant heterogeneity (PQ≤0.05, I2≥50%). Otherwise, the fixed-effects model was adopted (PQ>0.05, I2<50%). Potential publication bias was assessed using a funnel plot, and the sensitivity analysis was also performed to ensure the reliability of the results. The P-value <0.05 was considered statistically significant.
CONCLUSION
This meta-analysis has analyzed the association between the miR-10b expression levels with LNM and DM occurrence in patients with carcinoma. Compared with their incidence in those with low miR-10b expression, the incidence of LNM and DM was higher in cancer patients with high miR-10b expression. However, there were also some limitations in this meta-analysis. First, the number of included cancer patients was still relatively small, thus larger and better designed studies would be necessary to confirm the results obtained in this meta-analysis. Also, all patients included in this meta-analysis were Asian, and no patients of other races could be included. Additionally, potential publication bias may exist, which may originate from unpublished negative results or the small sample studies included, although no significant publication bias was observed based on the trim and fill method and the sensitivity analysis also showed the results were robust. Furthermore, the criterion for high miR-10b expression was varied in these studies. Therefore, larger size, multicenter, and higher quality studies are needed for further research, based on a unique criterion to classify the miR-10b expression groups.
Author contributions
HJ Wu and LL Hu were involved in the design of this meta-analysis, revised this manuscript and gave final approval for submission. TF Hao and BX Tong were involved in the collection of publications. Y Zhang and RB Liao conducted the data analysis of the study and manuscript writing. All authors contributed toward data analysis, drafted, critically revised the manuscript and agree to be accountable for all aspects of the work.
CONFLICTS OF INTEREST
The authors declared that there are no conflicts of interest regarding the publication of this paper.
FUNDING
We thank for the fnancial support of Graduate Innovation Fund of Nanchang University (No. cx2015168).
REFERENCES
1. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015; 136:E359-E386.
2. Cho JH, Lee YS, Sun DI, Kim MS, Cho KJ, Nam IC, Kim CS, Kim SY, Park YH, Joo YH. Prognostic impact of lymph node micrometastasis in oral and oropharyngeal squamous cell carcinomas. Head Neck. 2016; 38:E1777-E1782.
3. Li P, Wu F, Zhao H, Dou L, Wang Y, Guo C, Wang G, Zhao D. Analysis of the factors affecting lymph node metastasis and the prognosis of rectal neuroendocrine tumors. Int J Clin Exp Pathol. 2015; 8:13331-13338.
4. Ladeiro Y, Couchy G, Balabaud C, Bioulac-Sage P, Pelletier L, Rebouissou S, Zucman-Rossi J. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology. 2008; 47:1955-1963.
5. Bloomston M, Frankel WL, Petrocca F, Volinia S, Alder H, Hagan JP, Liu CG, Bhatt D, Taccioli C, Croce CM. MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA. 2007; 297:1901-1908.
6. Tian Y, Luo A, Cai Y, Su Q, Ding F, Chen H, Liu Z. MicroRNA-10b promotes migration and invasion through KLF4 in human esophageal cancer cell lines. J Biol Chem. 2010; 285:7986-7994.
7. Fritz HK, Lindgren D, Ljungberg B, Axelson H, Dahlback B. The miR(21/10b) ratio as a prognostic marker in clear cell renal cell carcinoma. Eur J Cancer. 2014; 50:1758-1765.
8. Xiao H, Li H, Yu G, Xiao W, Hu J, Tang K, Zeng J, He W, Zeng G, Ye Z, Xu H. MicroRNA-10b promotes migration and invasion through KLF4 and HOXD10 in human bladder cancer. Oncol Rep. 2014; 31:1832-1838.
9. Han X, Yan S, Weijie Z, Feng W, Liuxing W, Mengquan L, Qingxia F. Critical role of miR-10b in transforming growth factor-beta1-induced epithelial-mesenchymal transition in breast cancer. Cancer Gene Ther. 2014; 21:60-67.
10. Huang J, Sun C, Wang S, He Q, Li D. microRNA miR-10b inhibition reduces cell proliferation and promotes apoptosis in non-small cell lung cancer (NSCLC) cells. Mol Biosyst. 2015; 11:2051-2059.
11. Li Y, Li Y, Liu J, Fan Y, Li X, Dong M, Liu H, Chen J. Expression levels of microRNA-145 and microRNA-10b are associated with metastasis in non-small cell lung cancer. Cancer Biol Ther. 2016; 17:272-279.
12. Li QJ, Zhou L, Yang F, Wang GX, Zheng H, Wang DS, He Y, Dou KF. MicroRNA-10b promotes migration and invasion through CADM1 in human hepatocellular carcinoma cells. Tumour Biol. 2012; 33:1455-1465.
13. Nishida N, Yamashita S, Mimori K, Sudo T, Tanaka F, Shibata K, Yamamoto H, Ishii H, Doki Y, Mori M. MicroRNA-10b is a prognostic indicator in colorectal cancer and confers resistance to the chemotherapeutic agent 5-fluorouracil in colorectal cancer cells. Ann Surg Oncol. 2012; 19:3065-3071.
14. Wang YY, Ye ZY, Zhao ZS, Li L, Wang YX, Tao HQ, Wang HJ, He XJ. Clinicopathologic significance of miR-10b expression in gastric carcinoma. Hum Pathol. 2013; 44:1278-1285.
15. Yang YL, Xu LP, Zhuo FL, Wang TY. Prognostic value of microRNA-10b overexpression in peripheral blood mononuclear cells of nonsmall-cell lung cancer patients. Tumour Biol. 2015; 36:7069-7075.
16. Zhang J, Xu L, Yang Z, Lu H, Hu D, Li W, Zhang Z, Liu B, Ma S. MicroRNA-10b indicates a poor prognosis of non-small cell lung cancer and targets E-cadherin. Clin Transl Oncol. 2015; 17:209-214.
17. Chen XL. The relationship of miR-10b expression differences and clinicopathological characteristics and prognosis of breast cancer [D]. Guangdong Pharmaceutical University, 2012. (in Chinese)
18. Luan L, Han B, Bai Y, Teng M, Wang CF. Analysis of clinical and pathological significance of expression of miR-10b in the non-small cell lung cancer. [Article in Chinese]. Chin J Cancer Prev Treat. 2014; 21:1419-1422.
19. Chen PY, Meister G. microRNA-guided posttranscriptional gene regulation. Biol Chem. 2005; 386:1205-1218.
20. Brodersen P, Voinnet O. Revisiting the principles of microRNA target recognition and mode of action. Nat Rev Mol Cell Biol. 2009; 10:141-148.
21. Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST, Patel T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology. 2007; 133:647-658.
22. Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A, Labourier E, Reinert KL, Brown D, Slack FJ. RAS is regulated by the let-7 microRNA family. Cell. 2005; 120:635-647.
23. Sun XJ, Liu H, Zhang P, Zhang XD, Jiang ZW, Jiang CC. miR-10b promotes migration and invasion in nasopharyngeal carcinoma cells. Asian Pac J Cancer Prev. 2013; 14:5533-5537.
24. Ma L, Teruya-Feldstein J, Weinberg RA. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature. 2007; 449:682-688.
25. Ma L, Reinhardt F, Pan E, Soutschek J, Bhat B, Marcusson EG, Teruya-Feldstein J, Bell GW, Weinberg RA. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat Biotechnol. 2010; 28:341-347.
26. Li G, Wu Z, Peng Y, Liu X, Lu J, Wang L, Pan Q, He ML, Li XP. MicroRNA-10b induced by Epstein-Barr virus-encoded latent membrane protein-1 promotes the metastasis of human nasopharyngeal carcinoma cells. Cancer Lett. 2010; 299:29-36.
27. Nakata K, Ohuchida K, Mizumoto K, Kayashima T, Ikenaga N, Sakai H, Lin C, Fujita H, Otsuka T, Aishima S, Nagai E, Oda Y, Tanaka M. MicroRNA-10b is overexpressed in pancreatic cancer, promotes its invasiveness, and correlates with a poor prognosis. Surgery. 2011; 150:916-922.
28. Sakurai-Yageta M, Masuda M, Tsuboi Y, Ito A, Murakami Y. Tumor suppressor CADM1 is involved in epithelial cell structure. Biochem Biophys Res Commun. 2009; 390:977-982.
29. Kalluri R, Weinberg RA. The basics of epithelial-mesenchymal transition. J Clin Invest. 2009; 119:1420-1428.
30. Soltermann A, Tischler V, Arbogast S, Braun J, Probst-Hensch N, Weder W, Moch H, Kristiansen G. Prognostic significance of epithelial-mesenchymal and mesenchymal-epithelial transition protein expression in non-small cell lung cancer. Clin Cancer Res. 2008; 14:7430-7437.
31. Heller G, Fong KM, Girard L, Seidl S, End-Pfutzenreuter A, Lang G, Gazdar AF, Minna JD, Zielinski CC, Zochbauer-Muller S. Expression and methylation pattern of TSLC1 cascade genes in lung carcinomas. Oncogene. 2006; 25:959-968.
32. Fukuhara H, Kuramochi M, Fukami T, Kasahara K, Furuhata M, Nobukuni T, Maruyama T, Isogai K, Sekiya T, Shuin T, Kitamura T, Reeves RH, Murakami Y. Promoter methylation of TSLC1 and tumor suppression by its gene product in human prostate cancer. Jpn J Cancer Res. 2002; 93:605-609.
33. Jansen M, Fukushima N, Rosty C, Walter K, Altink R, Heek TV, Hruban R, Offerhaus JG, Goggins M. Aberrant methylation of the 5′ CpG island of TSLC1 is common in pancreatic ductal adenocarcinoma and is first manifest in high-grade PanlNs. Cancer Biol Ther. 2002; 1:293-296.
34. Steenbergen RD, Kramer D, Braakhuis BJ, Stern PL, Verheijen RH, Meijer CJ, Snijders PJ. TSLC1 gene silencing in cervical cancer cell lines and cervical neoplasia. J Natl Cancer Inst. 2004; 96:294-305.
35. Zhang W, Zhou L, Ding SM, Xie HY, Xu X, Wu J, Chen QX, Zhang F, Wei BJ, Eldin AT, Zheng SS. Aberrant methylation of the CADM1 promoter is associated with poor prognosis in hepatocellular carcinoma treated with liver transplantation. Oncol Rep. 2011; 25:1053-1062.
36. Uchino K, Ito A, Wakayama T, Koma Y, Okada T, Ohbayashi C, Iseki S, Kitamura Y, Tsubota N, Okita Y, Okada M. Clinical implication and prognostic significance of the tumor suppressor TSLC1 gene detected in adenocarcinoma of the lung. Cancer. 2003; 98:1002-1007.
37. Wang G, Dong W, Shen H, Mu X, Li Z, Lin X, Liu Y, Du J. A comparison of Twist and E-cadherin protein expression in primary non-small-cell lung carcinoma and corresponding metastases. Eur J Cardiothorac Surg. 2011; 39:1028-1032.
38. Thiery JP. Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer. 2002; 2:442-454.
39. Zhang J, Xu L, Yang Z, Lu H, Hu D, Li W, Zhang Z, Liu B, Ma S. MicroRNA-10b indicates a poor prognosis of non-small cell lung cancer and targets E-cadherin. Clin Transl Oncol. 2015; 17:209-214.
40. Ouyang H, Gore J, Deitz S, Korc M. microRNA-10b enhances pancreatic cancer cell invasion by suppressing TIP30 expression and promoting EGF and TGF-beta actions. Oncogene. 2014; 33:4664-4674.
41. Lin J, Teo S, Lam DH, Jeyaseelan K, Wang S. MicroRNA-10b pleiotropically regulates invasion, angiogenicity and apoptosis of tumor cells resembling mesenchymal subtype of glioblastoma multiforme. Cell Death Dis. 2012; 3:e398.

