Introduction
Head and neck squamous cell carcinoma (HNSCC) constitutes the sixth most common malignancy worldwide [1]. In spite of considerable advances in multimodality therapy including surgery, radiotherapy, and chemotherapy, the overall five year survival rate for patients with this type of cancer is among the lowest of all major cancer types and has not improved during the last decade [2]. Local tumor recurrence and distant metastasis after conventional therapy appear to be major contributing factors for restricted survival of HNSCC patients. Therefore, understanding the molecular oncogenic pathways underlying HNSCC would help to improve diagnosis, approaches to therapy, and prevention of the disease.
MicroRNAs (miRNAs) are endogenous, short, non-coding RNA molecules which regulate gene expression by translational repression or degradation of mRNA in a sequence-specific manner [3]. Bioinformatic prediction indicates that miRNAs regulate more than 30% of the protein coding genes [4]. It is estimated that approximately 1,000 miRNAs exist in the vertebrate genome. At this time, 1,048 human miRNAs are registered at miRBase release 16.0 (http://microrna.sanger.ac.uk/). A growing body of evidence suggests that miRNAs are aberrantly expressed in many human cancers, and that they play significant roles in carcinogenesis and cancer progression [5]. miRNAs can be divided into two classes: those which are oncogenic miRNAs and those which are tumor suppressive miRNAs. Up-regulated miRNAs could act as oncogenes by negatively regulating tumor suppressor genes, while down-regulated miRNAs could function as tumor suppressors by repressing oncogenes [6,7].
Studies of tumor suppressive miRNAs and searches for their target genes are important for our understanding of miRNA-regulated cancer pathways, including those specific miRNAs altered in HNSCC [8-10]. Recently, our miRNA profiles showed that miR-133a was down-regulated in cancer cells and that miR-133a had tumor suppressive functions [11-13]. Interestingly, miR-1 and miR-133a are located on the same chromosomal locus, forming a so called cluster. miR-1 and miR-133a are expressed in muscle and might be the most studied miRNAs in skeletal and cardiac muscle development [14,15].
miR-1 was also down-regulated in our miRNA screening of hypopharyngeal and esophageal squamous cell carcinoma [13,16]. However, the functional significance of miR-1 has not been clarified in HNSCC. The aim of this study was to investigate the function of miR-1 in HNSCC cell lines and to identify miR-1-regulated cancer pathways. For target genes searches of miR-1 in HNSCC cells, we performed genome-wide gene expression analysis. We focused on transgelin 2 (TAGLN2) as a candidate target of miR-1, as it was among the most down-regulated genes. Insight into the association between tumor suppressive miRNA and their target oncogene networks could enhance our understanding of the molecular mechanism of HNSCC carcinogenesis.
Results
Effect of miR-1 transfection on cell proliferation, migration, and invasion in HNSCC cell lines
To determine the function of miR-1, we performed gain-of-function analysis using miR-1 transfectants. The XTT assay showed statistically significant inhibition of cell proliferation in miR-1 transfectants in comparison with miRNA controls after 72 hr and 96 hr. For example, after 72 hr, both miR-1 transfected HSC3 and FaDu cultures grew only ~42% as much as control cultures (both P < 0.05, Figure 1A, top), while after 96 hr, proliferation fell to 23% and 9% of controls, respectively, (both P < 0.05, Figure 1A, bottom). Wound healing assays of miR-1-transfected HSC3 demonstrated that cell migration was significantly inhibited to 5% that of control (P < 0.05; Figure 1B).The Matrigel invasion assay showed that the number of invading cells was significantly decreased in miR-1 transfectants. Relative to control values (100%), the percentages of invading HSC3 and FaDu cells were only 45% and 27.1%, respectively (both P < 0.05; Figure 1C).
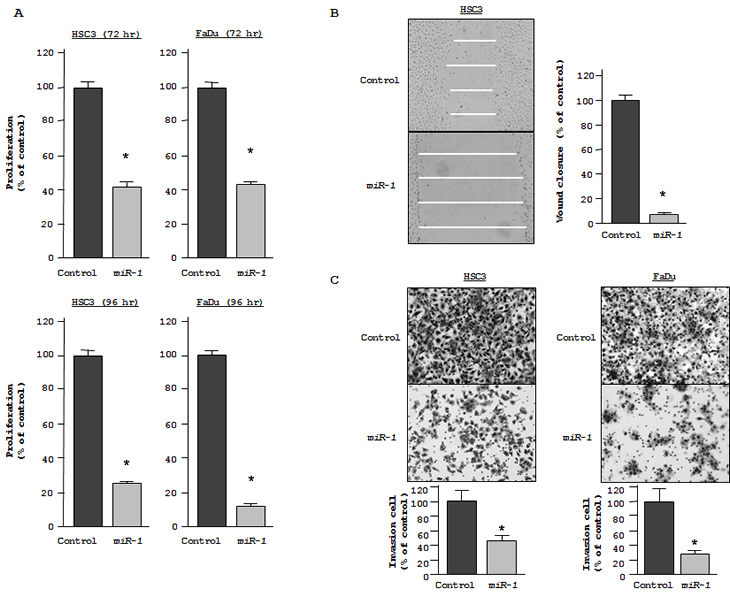
Figure 1: Gain-of-function studies using miR-1 transfected HNSCC cell lines (HSC3 and FaDu). (A) Cell growth as revealed by the XTT assay after 72 hr [above] and 96 hr [below]. (B) HSC3 cell migration activity (wound healing assay).(D) Cell invasion activity ( Matrigel invasion assay) in HSC3 and FaDu transfected with miR-1. *P < 0.05
Effect of miR-1 transfection on cell apoptosis and cell cycle in HNSCC cell lines
Cell apoptosis in miR-1 transfected cells was assessed by flow cytometry. The fraction of early apoptotic cells significantly increased in miR-1 transfectants approximately 3-fold in HSC3 and 12-fold in FaDu compared with controls (both P < 0.05, Figure 2A). We also confirmed induction of apoptosis approximately 4-fold in FaDu by ectopic miR-1 performing TUNEL assay (data not shown). As for cell cycle distribution, cells in G0/G1 phase were significantly greater in miR-1 transfectant than those in the control (Figure 2B). These results suggest that ectopic miR-1 expression induces G0/G1 arrest in both HNSCC cell lines
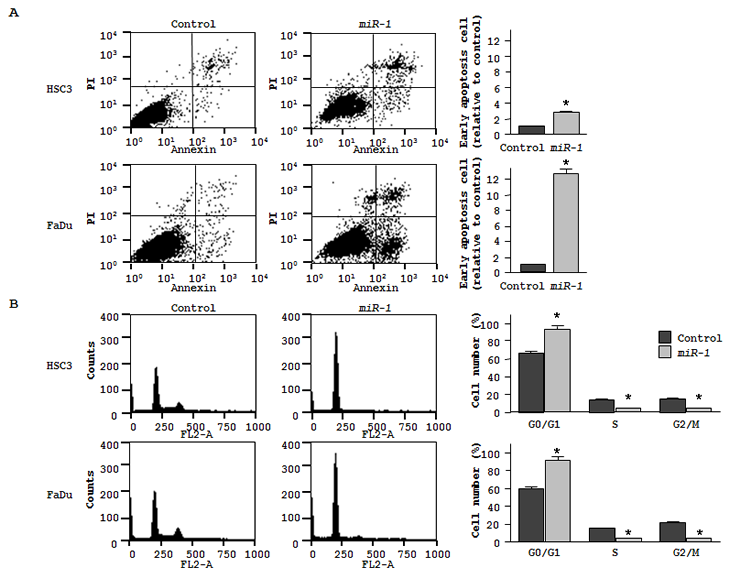
Figure 2: Effect of miR-1 transfection on cell apoptosis and cell cycle. (A) The representative quadrant figures of miRNA-control, or miR-1 transfectants in HSC3 [above] and FaDu [below] cells are shown. The bar chart shown on the right of each quadrant indicates the ratio of the early apoptotic cell fraction in miR-1 transfectants in comparison with miRNA-control transfectant. The data for the early apoptotic cell fraction is expressed as the relative value of the average expression of the miRNA-control transfectant. * P < 0.05. (B) The typical figures of cell cycle analysis of miRNA-control, or miR-1 transfectants are shown. The bar chart shown on the right of each figures represents the percentage of the cells in G0/G1, S, or G2/M phase as indicated. * P < 0.05.
Gene expression profiling identifies down-regulated genes in miR-1 transfectants
To investigate candidate molecular targets of miR-1 in HNSCC cells, we examined the effect of miR-1 on protein coding genes. Mature miR-1 was transiently transfected into HSC3 and FaDu cells, with negative miRNA transfection used as a control. Comprehensive gene expression analysis (see Methods) clearly showed changes in gene expression patterns between miR-1 and negative-control transfectants. To identify candidate miR-1 target genes, a cut-off of value less than -2.00-fold was applied to the array data. This filter resulted in the identification of 59 genes that were significantly down-regulated upon miR-1 transfection in both HSC3 and FaDu cells (top 20 genes are shown in Table 1). Entries from the microarray data were approved by the Gene Expression Omnibus (GEO), and were assigned GEO accession number GSE24782. The 3’ UTR of these down-regulated genes were examined for miR-1 target sites using the TargetScan database. Of the top 20 putative gene targets, 17 genes contained miR-1 target sites.
Table 1: Down-regulated genes in miR-1 transfectants
| No. | Entrez Gene ID | ||||||
’’’’
<
<
±±±±±±±±±±
<
<<<<<<
’’°
’
’®™
μ’°°°®®
μβ
–’™’–
μ™μ™
°
<<
’

