INTRODUCTION
Genetically unstable regions of the human genome frequently harbor tumor-suppressor genes [1, 2]. The second most frequently affected site of human chromosomal fragility is FRA16D located at 16q23 within the WWOX (tryptophan domain containing oxidoreductase) locus [3]. The sequences of both orthologous chromosomal fragile site regions for human FRA16D/WWOX and mouse Fra8E1/Wwox exhibit remarkable conservation [4, 5]. Deletion of WWOX in mice has resulted in the spontaneous development of osteosarcoma (OS) [6]. Likewise, studies have reported that the majority of human OS patients have undetectable WWOX protein levels [7, 8] or its compromised functionality in cases with metastasis [9]. Nevertheless, the lack of correlation between WWOX expression and osteosarcoma patient prognosis suggests that its loss is an early event in cancer pathogenesis with the phenotypic results of its deletion not contributing directly to patient demise [7]. The fact that WWOX deletion does not occur frequently but that loss of its expression is common suggests that factors influencing the epigenetic alterations of this gene might also be involved in the pathogenesis of osteosarcoma [7].
The involvement of non-coding RNA transcripts is proposed due to their reported accumulation at fragile sites, where they are believed to influence the structure of specific chromosomal regions associated with malignancy [10, 11]. Long non-coding RNA (lncRNA, transcripts longer than 200 nucleotides with limited coding potential) can induce epigenetic variations in the chromatin state of cancer cells [12]. Observed alterations in the abundance of many lncRNA transcripts in cancer, implicates their putative role in tumor pathogenesis [13]. Although seen as mainly regulators of gene expression, lncRNAs can also act via post-transcriptional mechanisms that are capable of influencing chromosome structure and stability [14].
The lncRNA PARTICLE (promoter of MAT2A antisense radiation-induced circulating long non-coding RNA [15]) is transcribed in the antisense direction within the promoter of methionine adenosyltransferase 2A (MAT2A), the product of which encodes the catalytic subunit of methionine adenosyltransferase [16]. PARTICLE forms a triplex within the MAT2A promoter and operates an active feedback silencing loop limiting the upregulation of MAT2A in response to low dose radiation [15]. In contrast to earlier models that showed PARTICLEs repressive ability was, typical of most lncRNAs, restricted to a specific gene at a local (usually in cis) level, it has been recently discovered that PARTICLE over-expression enhances trimethylation of histone H3 lysine 27 across the human genome with specific enrichment within WWOX [17, 18]. In contrast, diminished hyper-methylation of a WWOX promoter CpG island was evident upon PARTICLE knockdown [17]. Given the lack of FRA16D breakage and WWOX loss in some osteosarcomas we have investigated alternate mechanisms for suppressing WWOX expression. We hypothesized that the lncRNA PARTICLE might provide an alternative path for inactivation of WWOX in osteosarcoma where the fragile site remained intact.
RESULTS
WWOX and PARTICLE reveal contrasting expression patterns in osteosarcoma tumor samples
Differential WWOX and PARTICLE expression was determined in OS tumor samples and normal tissue controls [19] obtained from individuals of ~ 15 yr median age (n = 40, Table 1). WWOX was significantly down regulated (p = 0.000005) in OS tumor samples compared to controls. In contrast, PARTICLE expression was increased by 19.7 ± 0.8 % (p = 0.011) in OS tumor samples when similarly compared (Figure 1A). These findings revealed the distinct expression of PARTICLE in malignant tissue.
Table 1: Osteosarcoma clinical data from fourty patients partaking in this study
Osteosarcoma Patient Clinical Data |
WWOX FRA16D Non-breakage (OS - WT) |
WWOX FRA16D Breakage (OS - BRK) |
|---|---|---|
Gender |
n = 23 |
n =17 |
Male |
14 |
9 |
Female |
9 |
8 |
Age at Diagnosis (Years) |
n = 23 |
n = 17 |
Average |
22.9 |
16.4 |
Median |
15.3 |
14.9 |
Range |
8 - 85 |
7 - 47 |
Metastases |
n = 21 |
n = 15 |
Yes |
10 |
4 |
No |
11 |
11 |
Observation Period (Months) |
n = 22 |
n = 17 |
Average |
4.9 |
6.4 |
Median |
5.3 |
2.1 |
Range |
0.3 – 12.2 |
0.2 - 17 |
Response to Neo-adjuvants |
n = 21 |
n = 15 |
Good |
13 |
7 |
Poor |
8 |
8 |
Survival |
n = 19 |
n = 12 |
Alive |
15 |
8 |
Dead |
4 |
4 |
Event* |
n = 19 |
n = 12 |
Yes |
8 |
4 |
No |
11 |
8 |
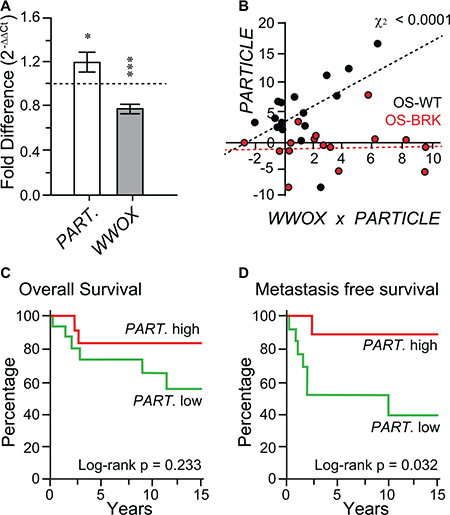
Figure 1: In the absence of FRA16D breakage the WWOX tumor suppressor is accompanied by elevated PARTICLE expression. (A) Histograms depicting PARTICLE and WWOX expression in unselected osteosarcoma specimens relative to paired controls. (B) Scatterplot of Chi-square correlation between PARTICLE and WWOX in OS–WT (black dots) and OS–BRK (red dots). (C and D) Association of PARTICLE with overall survival (C) or metastasis free survival (D) amongst osteosarcoma patients. PARTICLE expression relative to the endogenous control gene encoding the TATA-binding protein in patient tumors was used for comparison with retrospective outcome. Event (defined as metastasis free survival) comparison using log-rank (Mantel-Cox) testing.
Absence of FRA16D breakage in the WWOX tumor suppressor is accompanied by elevated PARTICLE expression
Tumors were classified according to their FRA16D breakage status (non-breakage: OS-WT; breakage: OS-BRK). When these tumor groups were compared, highly significant increased PARTICLE levels were noted when the WWOX FRA16D was intact (Chi-square association p = 0.000059, Figure 1B). This positive correlation suggests that the absence of FRA16D breakage within the WWOX tumor suppressor is accompanied by elevated PARTICLE expression.
PARTICLE expression is associated with metastasis free survival in osteosarcoma patients
No significant difference was found from Kaplan Mayer analysis of PARTICLE expression levels and overall patient survival (log-rank Mantel-Cox text p = 0.23, Figure 1C). Nevertheless, elevated PARTICLE expression was significantly associated with metastasis free survival in osteosarcoma patients within this sample cohort (log-rank Mantel-Cox test p = 0.0324; Figure 1D).
Osteosarcoma cell lines elicit differential PARTICLE levels in response to radiation
PARTICLE transcript levels were examined in osteosarcoma cell lines (MG63, SJSA and U2OS, Figure 2) at 4 or 24 hr following low (0.25 Gy) or medium (2.5 Gy) dose radiation in comparison to sham-irradiated controls (Supplementary Figure 1). In situ hybridization revealed that PARTICLE transcripts were located pre-dominantly in the cytoplasm of MG63 post low dose irradiation (Figure 2A). Quantitative PCR analysis demonstrated a highly significant decrease in PARTICLE at 4 hr post low or medium irradiation (p = 0.002 and p = 0.005, respectively) with similar effects at 24 hr post 0.25 Gy (p = 0.005) (Figure 2B).
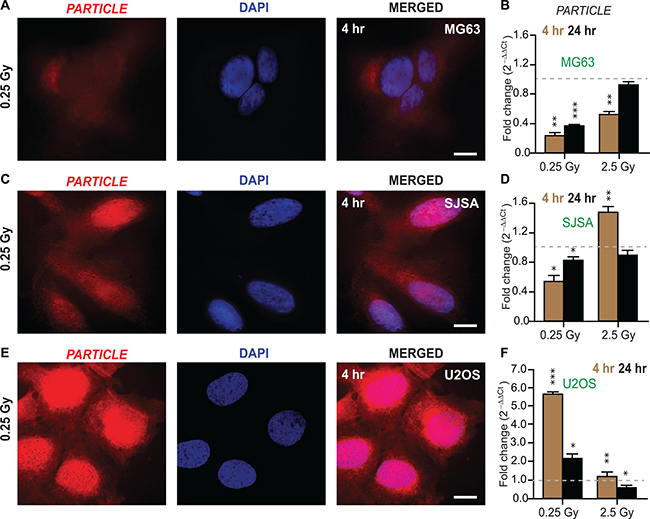
Figure 2: Differential PARTICLE expression in osteosarcoma cell lines in response to radiation. (A, C and E) Representative epifluorescence microscopic images of MG63, SJSA and U2OS 4 hr post 0.25 Gy labelled with in situ hybridization probes (Quasar 570 (red)) specific for PARTICLE (left). Nuclei stained with DAPI (blue, middle). Merged images (right). Scale bar 10 μm. (B, D and F) Histograms of PARTICLE expression in osteosarcoma cell lines MG63 (B), SJSA (D) and U2OS (F) exposed 4 hr (brown bars) or 24 hr (black bars) post 0.25 or 2.5 Gy irradiation exposure.
In SJSA, PARTICLE transcripts were more highly expressed in the nucleus in comparison to the cytoplasm (Figure 2C). In this cell line decreased PARTICLE expression was noted particularly at the 4 hr time point (p = 0.003) post 0.25 Gy (Figure 2D). In contrast, significant elevation in PARTICLE (p = 0.006, Figure 2D) was found 4 hr post exposure to medium dose irradiation (p < 0.0005, Figure 2D).
In situ hybridization, revealed that the intracellular distribution of PARTICLE in U2OS was predominantly nuclear 4 hr post 0.25 Gy exposure (Figure 2E). Ameliorated PARTICLE expression was noted (4.8 ± 0.6 fold increase) compared to sham irradiated cells 4 hr post low dose irradiation (p = 0.0002, Figure 2F). These findings demonstrate variable PARTICLE intracellular distribution and transcript levels post irradiation in representative osteosarcoma cell lines.
WWOX transcript levels and promoter activity in osteosarcoma cell lines
The osteosarcoma cell lines MG63, SJSA and U2OS are characterized by different WWOX genomic backgrounds [21]. MG63 and SJSA cells harbor a deletion in WWOX intron 3 (Figure 3A). In addition, WWOX contains FRA16D breakage in SJSA cells (Figure 3A). U2OS cells represent WWOX allelic imbalance and no FRA16D breakage in WWOX (Figure 3A). Following the revelation that PARTICLE was highly variable in these cell lines, comparison with WWOX expression and its promoter activity was instigated. These cell lines presented very different expression profiles for PARTICLE and WWOX at 4 or 24 hr following low (0.25 Gy) or medium (2.5 Gy) dose radiation exposure in comparison to sham-irradiated controls (Figures 2 and 3). In contrast to PARTICLE in MG63 (Figure 2B), significantly increased WWOX transcription was evident 4 hr post 0.25 or 2.5 Gy exposure (p = 0.0048 and p = 0.00005, respectively) in this cell line (Figure 3A). Luciferase reporter analysis of the WWOX promoter in MG63 also reflected significantly escalated activity at 4 hr post 0.25 or 2.5 Gy exposure after these dosages (p < 0.05) in comparison to sham irradiated controls (Figure 3B). In SJSA, a comparable pattern to MG63 emerged after low dose irradiation whereby increases in WWOX transcript levels (4 hr: p = 0.003; 24 hr: p = 0.01) and promoter activity (4 hr: p = 0.02; 24 hr: p = 0.05) were again accompanied by reduced PARTICLE levels (Figures 2B and 3A). Significant increases in PARTICLE (p = 0.006, Figure 2D) was likewise accompanied by reduction in WWOX expression 4 hr post exposure to medium dose irradiation (p < 0.0005, Figure 3A).
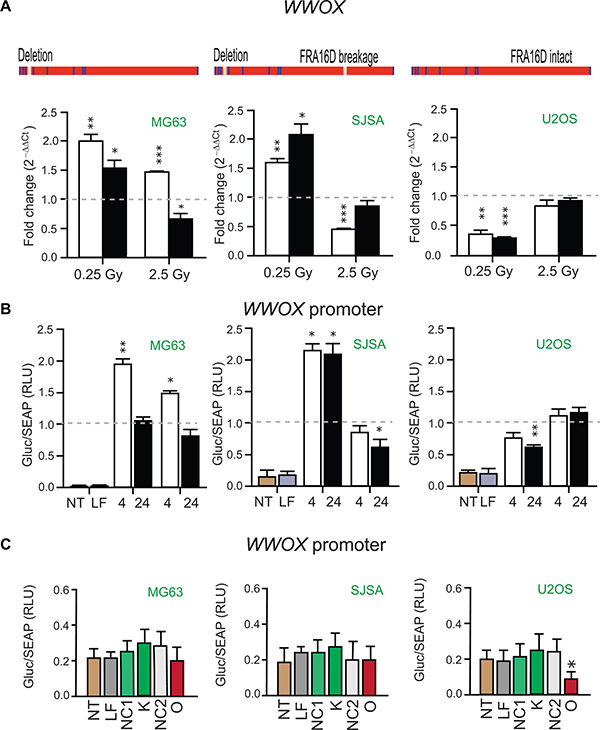
Figure 3: WWOX transcript levels and promoter activity in Osteosarcoma cell lines. (A) Schematics of the WWOX gene in MG63, SJSA and U2OS with presence or absence of intron 3 deletion or FRA16D breakage indicated (above). Histograms of WWOX expression in these cell lines 4 hr (white bars) or 24 hr (black bars) post 0.25 or 2.5 Gy irradiation exposure. Values were normalized with TATA-binding protein (TBP) encoding the endogenous gene with relative expression comparison to sham-irradiated (0 Gy) equivalent cells (dashed lines). Data are represented as mean ± SEM (n = 3) with significance represented by asterisks (p < 0.05) where appropriate. (B) Histograms of Gaussia luciferase (GLuc) activity normalized to alkaline phosphatase activity (SEAP) indicative of WWOX promoter activity in MG63, SJSA and U2OS. Tissue culture media analyzed 4 hr and 24 hr following 0.25 Gy or 2.5 Gy cellular irradiation or sham irradiation (dashed line). Data are represented as mean ± SEM (n = 3). (C) Histograms of GLuc activity normalized to SEAP in non-transfected (NT), lipofectamine transfected (LF), PARTICLE knockdown (K), PARTICLE overexpression (O), siRNA negative control (NC1) or in vitro transcript control (NC2) in MG63, SJSA and U2OS. Data are represented as mean ± SEM (n = 3).
Diminished WWOX levels were detected in U2OS at 4 and 24 hr post low dose irradiation (p = 0.003 and p = 0.0004 respectively, Figure 3A). WWOX promoter activity was likewise reduced in this cell line particularly 24 hr post low dose irradiation exposure (p = 0.006, Figure 3B). Post irradiation the contrasting expression profiles of PARTICLE and WWOX in such cell lines that represented various WWOX genetic backgrounds suggest a relationship may exist between this lncRNA and this tumor suppressor gene.
PARTICLE overexpression compromises WWOX promoter activity
Knockdown of PARTICLE expression did not significantly affect WWOX promoter activity in MG63, SJSA or U2OS (Figure 3C) in comparison to lipofectamine alone transfection or negative control (NC1). Overexpression of PARTICLE elicited different effects on WWOX promoter activity in these cell lines (Figure 3C). No significant decrease in WWOX promoter activity was noted in MG63 or SJSA (p > 0.05) when compared to lipofectamine alone transfection or an over-expressed ‘run-off’ transcript control (NC2). In U2OS, PARTICLE overexpression significantly reduced WWOX promoter activity by 55 ± 2 % (p = 0.013). Evidence is provided here that WWOX promoter activity is curtailed by the lncRNA PARTICLE.
DISCUSSION
Herein, this report shows that higher PARTICLE expression levels are associated with malignant tissue and the absence of WWOX FRA16D breakage. Importantly, elevated levels of this lncRNA are significantly associated with metastasis free survival in osteosarcoma patients. Focus has shifted recently towards finding therapeutic biomarkers associated with metastasis-free survival as an alternative and important predictor of osteosarcoma outcome. Currently, patients with localized osteosarcoma have a 70% chance of 5-year prognosis. However, with the onset of metastasis or recurrent disease, patients have less than 20% chance of long-term survival despite aggressive therapies [22, 23]. A recent report [24] has shown that the provision of increased therapy to high risk patients was ineffective (ie. post 10 weeks operative and chemotherapy intervention), as was therapeutic intensity reduction for low risk patients. These prognostic outcomes for osteosarcoma have not changed significantly over the last 25 years.
It has been shown that osteosarcoma is the dominant phenotype in WWOX gene knockout mice [7]. Numerous studies have reported that WWOX alterations occur in many types of cancer cells and in situations of cellular stress [3]. Nonetheless, studies have failed to correlate the occurrence of genetic modifications in WWOX with cancer development, suggesting non-mutational regulation may occur [25]. Conflicting evidence exists related to the correlation between loss of WWOX protein and cancer prognosis [7, 26, 27]. Evidence has emerged that PARTICLE over-expression enhances the histone repressive modification mark across the human genome and specifically within WWOX in a breast cancer cell line [17]. PARTICLE was reported to increase the methylation status of a WWOX promoter CpG island (annotated CpG105476) of 990 bp located on chromosome 16: 78133076 – 78134066 (NCBI homo sapiens build number 37/hg19). The transcription initiation site for WWOX resides within this region at position chromosome 16: 78133327 orientated in a forward direction (NCBI gene id. 51741) [17]. It has also been demonstrated that PARTICLE overexpression leads to a reduction in WWOX expression [18].
Emerging evidence using bio-informatic tools from the latest miRbase and the ENCODE projects have revealed that chromosomal fragile sites are enriched in various non-coding binding elements, as well as specific variations in histone modifications [28]. This prompted us to investigate the association of elevated PARTICLE and an absence of FRA16D breakage in WWOX in tumor specimens. At the structural level, chromosomal fragile sites have the propensity to form secondary non-B DNA structures that interfere with the movement of the replication fork thus leading to its collapse and associated DNA breaks [29].
Intriguingly, transiently elevated expression of the lncRNA PARTICLE was found to accompany diminished WWOX transcript levels and promoter activity within in vitro models of osteosarcoma. Manipulation of PARTICLE transcript levels via siRNA interference or in vitro transcription and transfection demonstrated reduction of WWOX promoter activity in U2OS. Such an effect was absent from the other tested cell lines perhaps due to an alternative PARTICLE intracellular distribution in MG63 or the presence of FRA16D breakage in SJSA.
It could be envisaged that PARTICLE association within the WWOX gene might avert FRA16D breakage through fork remodeling/scaffolding yet hindering replication and transcription in tumor cells that lead to metastasis (Figure 4). Rather than merely structural domains vulnerable to breakage, chromosomal fragile sites may be highly organized functional stress sensors cooperating with non-coding elements recruited to harness the effects of osteosarcoma progression.
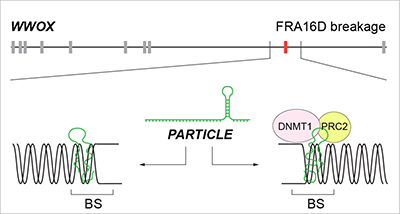
Figure 4: Schematic of the potential mechanisms by which PARTICLE association within the WWOX gene might avert FRA16D breakage yet hindering replication and transcription in tumor cells that lead to metastasis. Replication fork remodeling (left) or scaffold formation (right) for epigenetic silencers. BS: breakage site; PRC2: Polycomb repressive complex 2; DNMT1: DNA methyltransferase 1.
MATERIALS AND METHODS
Ethical approval
This study was carried out in accordance with the recommendations of the medical faculty of the Technical University of Munich (TUM) with written informed consent from all subjects in accordance with the Declaration of Helsinki. The protocol was approved by the research ethics board (Reference 1867/07).
Clinical information and osteosarcoma specimen preparation
Clinicopathological data (n = 40) were obtained from osteosarcoma patients with informed consent. Cases included 23 male and 17 female patients with osteosarcoma; mean age was 19.6 years. Metastases information was available for 90% of cases with 61% classified with local tumor status. The majority of patients (90%) had received neoadjuvant chemotherapy with 44% responding poorly to treatment. Anonymized primary tumors and paired controls [19] were obtained from this patient cohort (n = 40). All tumor samples were evaluated by an experienced bone pathologist who confirmed the diagnosis and the tumor content to be > 70% per sample. Total RNA was isolated from tumor tissue (~ 10 mg) or controls using an RNeasy mini-kit (cat # 74124, Qiagen).
Propagation and maintenance of cell lines
U2OS, SJSA and MG63 human osteosarcoma cell lines (American Type Culture Collection) were cultivated in Roswell Park Memorial Institute (RPMI) 1640 media (GibcoTM cat # 21875-034) and fetal bovine serum (10%) at 37°C in a humidified incubator containing 95% oxygen/5% carbon dioxide atmosphere. The identity of these cell lines was verified by microsatellite analysis (Forensik GmbH, Germany). All cultures were routinely checked for mycoplasma contamination using a MycoAlert mycoplasma detection kit (Lonza, cat. # LT07-218). In general, cells were grown to 80% confluency prior to removal from the dish using trypsin (0.25%)/ EDTA (0.02%) and sub-culturing or harvesting.
RNA interference targeting PARTICLE
PARTICLE knockdown was undertaken with Silencer® Select siRNA interference technology (siRNA id: n307629; Part # 4390771). U2OS, SJSA and MG63 were grown to 60% confluence and transfected with these siRNAs (10 nM) using lipofectamine as per manufacturer instructions. After 72 hr, cells were transferred onto 6 well dishes for WWOX promoter luminescence assay determination (see below). Control conditions included transfection with lipofectamine and/or negative siRNA (NC1; cat # AM4615 no.3, Thermo Scientific).
PARTICLE overexpression
PARTICLE was cloned into the pGEM® - T vector (p.PART) (GenScript) and transformed into Top10 bacteria. A colony was grown in ampicillin (100 mg/ml) overnight and plasmid midiprep (Promega) performed. Plasmid concentration and purity was assessed (NanoDrop 1000, Thermo Scientific) with A260/280 ratio determination with automated sequence validation (GenScript). Plasmid linearization was carried out using 1mg plasmid DNA and SacI overnight digestion at 37°C. PARTICLE (1432bp) was in vitro transcribed from a pGEM®-T vector (GenScript) using the TranscriptAid T7 High Yield transcription kit (Thermo Scientific, cat # K0441). Transcripts were treated with RNase-free DNase 1 (Thermo Scientific) and purified using an RNeasy mini-elute cleanup kit (Qiagen, cat # 74204) and verified by TBE-agarose (1.8%) electrophoresis. Prior (24 hr) to transfection, U2OS, SJSA and MG63 were seeded (105 cells/well in a 6 well dish) in growth media (described above) in the absence of antibiotic/anti-mycotic to ~ 70% confluence at the time of transfection. The control template included in the Transcript T7 High Yield Transcription kit (Thermo Scientific, cat # K0441) as utilised for the production of a 2223bp ‘run off’ transcript serving as a negative control (NC2) for over-expression studies. Cells were transfected with lipofectamine and PARTICLE (4 mg) or negative control (4 mg) as per standard conditions with incubation for 72 hr prior to WWOX promoter luminescence assay.
Irradiation
All irradiation was performed using a closed HWM-D 2000 Cesium137 source (Wälischmiller Engineering GmbH, Markdorf, DE; 10 cm height, 33 cm diameter) at a dose rate of 0.0082Gy/sec. Sham irradiation of controls involved only transport to the irradiation facility.
RNA isolation and cDNA synthesis
Total RNA was isolated from cell lines and purified using TriFast peqGOLD (Peqlab, cat # 30–2010) and a Maxwell® 16 LEV Blood DNA kit (Promega cat # AS1290) with solution substitution (ie. isopropanol replacement by 100 % ethanol in cartridge number 1) and a Maxwell® 16 machine (Promega). The final elution was in ultra-pure water with concentration and purity assessment using O.D. 260/280 ratio determination (NanoDrop 1000, Thermo Scientific). Total RNA was stored at –80°C. Total RNA (1 mg) from sham irradiated or irradiated cells was converted into first strand cDNA using standard protocol procedures (with the inclusion of random hexamers and oligo dT primers) and reagents from Life Technologies, Germany.
Real time PCR quantification
PARTICLE and TBP primers and probes [15] or pre-designed WWOX (Hs00249590_ml: single Taqman gene expression assay spanning the exon 1–2 boundary) (ThermoFisher cat # 4331182) were utilized for gene expression determinations. The reaction conditions for single gene assays were as such: cDNA (50–100 ng), 1× Taqman universal PCR master mix (no AmpErase UNG; Life Technologies, cat. # 4324018), forward and reverse primers (10 pmol), specific fluorescent probe (5 pmol), nuclease-free water up to 25 ml. For pre-designed assays, similar conditions were used except for utilization of an assay mix (1 X) instead of individual primers and probes. When pre-amplification was necessary (following total RNA extraction from human osteosarcoma and control specimens), a pooled assay mix was initially prepared (1 pmol primers and 0.5 pmol probes) for PARTICLE, WWOX and TBP (endogenous control) genes. Pre-amplification conditions were prepared as per manufacturer instructions (Life Technologies, cat # 4384556) and performed as follows: 95°C 10 min (1 cycle), 95°C 15 sec and 60°C 4 min (14 cycles), 4°C hold. Cycle threshold values were extracted and fold changes in gene expression determined by 2(-ΔΔCt) [20]. Negative controls were normalized to a value = 1 and samples relatively compared.
WWOX promoter luminescence assay
MG63, SJSA and U20S were plated on 6 well dishes (2 × 105 cells/well) in growth media (2 ml) (above). After 24 hr cells were transfected as per manufacturer’s instructions with a WWOX GLuc-on promoter reporter clone (GeneCopoeia, Rockville, MD, USA, cat. # HPRM19966-PG04) cloned into pEZX-LvPG04. This vector contains reporter and tracking genes encoding Gaussia luciferase (GLuc) and alkaline phosphatase (SEAP) secreted reporter proteins, respectively. Cells were irradiated 72 hr after transfection using lipofectamine. Culture medium (100 ml) was collected for analysis 4 hr and 24 hr after 0.25 Gy or 2.5 Gy irradiation exposures. Cell culture medium was also collected from sham-irradiated +/− lipofectamine transfection as controls. In addition, a positive control (EF1A-PG04, GeneCopoeia, Rockville, MD, USA) was assessed in parallel for verification purposes. The GLuc assay was conducted using buffer GL-S (GeneCopoeia, Rockville, MD, USA) for stable activity. Duplicate luminescence measurements were obtained with 3 second integration using white opaque 96 well flat bottom plates (Becton Dickinson Labware Europe, cat. # 353296). Signal normalization was done using the SEAP signal as an internal standard control (ratio of GLuc/SEAP activities) to eliminate the impact of variation in transfection efficiency.
PARTICLE in situ hybridization and microscopic analysis
Procedures were followed as previously reported [15] and in accordance with Stellaris fluorescence in situ hybridization (FISH) (Biosearch Technologies) website information (www.biocat.com). Using the online probe designer tool (www.biosearchtech.com/stellarisdesigner/), 40 specific probes were selected from an input sequence (PARTICLE NR_038942.1) for optimal binding properties to the target RNA sequence. Search parameters were selected that included a masking level (3–5), maximum probe coverage = 40 and minimum 2 nucleotides spacing level. The probe fluorophore Quasar 570 (Ex: 552 nm; Em: 570 nm) was used for PARTICLE detection. Cells were mounted in VECTASHIELD™ HardSet™ containing the nuclear counterstain DAPI (Vector lab, cat # H-1500). PARTICLE or nuclear images were acquired using a TexasRed or DAPI (Ex/Em: 350/470 nm) dual filter wheel respectively on an inverted Axiovert 200 (Zeiss) fluorescence microscope with apotome slide module activation.
Statistical analysis
Values are expressed as the mean ± S.E.M., with n = 3 referring to the number of independent biological replicates. Comparison of means or groups were tested using the Student’s t-test or ANOVA with p values < 0.05 taken to indicate statistical significance. Comparisons of frequencies were performed using the Chi square test using GraphPad Prism statistical analysis software. Clinical parameters included metastasis or osteosarcoma cause-specific mortality. The time to occurrence of an event was computed from the date of diagnosis to the date when the event was first recorded. Kaplan-Meier survival analysis was used to examine the relationship between overall survival or metastasis-free survival and PARTICLE expression. High or low expression levels of PARTICLE were considered either above or below a value of 1 respectively following normalization. A log-rank Mantel-Cox test was used (GraphPad Prism) to test the statistical significance of the differences in survival rate estimators.
Author contributions
VBOL performed experimental work, designed and supervised the research and wrote the paper; DM performed experimental work; JS performed bioinformatics analysis and contributed graphics; DB provided histology and patient diagnosis; MN provided patient samples and cohort data; SVO designed the graphics for figures and performed statistical analysis; MJA directed the research.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
This work was supported by the EURATOM Fission, European Commission 7th Framework Programme, Dark.Risk project (contract number 323216). D.M was funded by the BMBF (TranSarNet, FKZ 01GM0870).
REFERENCES
1. Richards RI. Fragile and unstable chromosomes in cancer: causes and consequences. Trends Genet. 2001; 17:339–45.
2. Ruiz-Herrera A, Garcia F, Fronicke L, Ponsa M, Egozcue J, Caldes MG, Stanyon R. Conservation of aphidicolin-induced fragile sites in Papionini (Primates) species and humans. Chromosome Res. 2004; 12:683–90. https://doi.org/10.1023/B:CHRO.0000045753.88789.ea.
3. Smith DI, McAvoy S, Zhu Y, Perez DS. Large common fragile site genes and cancer. Semin Cancer Biol. 2007; 17:31–41. https://doi.org/10.1016/j.semcancer.2006.10.003.
4. Aldaz CM, Ferguson BW, Abba MC. WWOX at the crossroads of cancer, metabolic syndrome related traits and CNS pathologies. Biochim Biophys Acta. 2014; 1846:188–200. https://doi.org/10.1016/j.bbcan.2014.06.001.
5. Krummel KA, Denison SR, Calhoun E, Phillips LA, Smith DI. The common fragile site FRA16D and its associated gene WWOX are highly conserved in the mouse at Fra8E1. Genes Chromosomes Cancer. 2002; 34:154–67. https://doi.org/10.1002/gcc.10047.
6. Aqeilan RI, Trapasso F, Hussain S, Costinean S, Marshall D, Pekarsky Y, Hagan JP, Zanesi N, Kaou M, Stein GS, Lian JB, Croce CM. Targeted deletion of Wwox reveals a tumor suppressor function. Proc Natl Acad Sci U S A. 2007; 104:3949–54. https://doi.org/10.1073/pnas.0609783104.
7. Yang J, Cogdell D, Yang D, Hu L, Li H, Zheng H, Du X, Pang Y, Trent J, Chen K, Zhang W. Deletion of the WWOX gene and frequent loss of its protein expression in human osteosarcoma. Cancer Lett. 2010; 291:31–8. https://doi.org/10.1016/j.canlet.2009.09.018.
8. Mirabello L, Troisi RJ, Savage SA. Osteosarcoma incidence and survival rates from 1973 to 2004: data from the Surveillance, Epidemiology, and End Results Program. Cancer. 2009; 115:1531–43. https://doi.org/10.1002/cncr.24121.
9. Kurek KC, Del Mare S, Salah Z, Abdeen S, Sadiq H, Lee SH, Gaudio E, Zanesi N, Jones KB, DeYoung B, Amir G, Gebhardt M, Warman M, et al. Frequent attenuation of the WWOX tumor suppressor in osteosarcoma is associated with increased tumorigenicity and aberrant RUNX2 expression. Cancer Res. 2010; 70:5577–86. https://doi.org/10.1158/0008–5472.CAN-09–4602.
10. Rossi S, Sevignani C, Nnadi SC, Siracusa LD, Calin GA. Cancer-associated genomic regions (CAGRs) and noncoding RNAs: bioinformatics and therapeutic implications. Mamm Genome. 2008; 19:526–40. https://doi.org/10.1007/s00335-008-9119-8.
11. Brown JD, Mitchell SE, O'Neill RJ. Making a long story short: noncoding RNAs and chromosome change. Heredity (Edinb). 2012; 108:42–9. https://doi.org/10.1038/hdy.2011.104.
12. Ishibashi M, Kogo R, Shibata K, Sawada G, Takahashi Y, Kurashige J, Akiyoshi S, Sasaki S, Iwaya T, Sudo T, Sugimachi K, Mimori K, Wakabayashi G, et al. Clinical significance of the expression of long non-coding RNA HOTAIR in primary hepatocellular carcinoma. Oncol Rep. 2013; 29:946–50. https://doi.org/10.3892/or.2012.2219.
13. van Leeuwen S, Mikkers H. Long non-coding RNAs: Guardians of development. Differentiation. 2010; 80:175–83. https://doi.org/10.1016/j.diff.2010.07.003.
14. Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012; 482:339–46. https://doi.org/10.1038/nature10887.
15. O'Leary VB, Ovsepian SV, Carrascosa LG, Buske FA, Radulovic V, Niyazi M, Moertl S, Trau M, Atkinson MJ, Anastasov N. PARTICLE, a Triplex-Forming Long ncRNA, Regulates Locus-Specific Methylation in Response to Low-Dose Irradiation. Cell Rep. 2015; 11:474–85. https://doi.org/10.1016/j.celrep.2015.03.043.
16. Alvarez L, Sanchez-Gongora E, Mingorance J, Pajares MA, Mato JM. Characterization of rat liver-specific methionine adenosyltransferase gene promoter. Role of distal upstream cis-acting elements in the regulation of the transcriptional activity. J Biol Chem. 1997; 272:22875–83.
17. O'Leary VB, Hain S, Maugg D, Smida J, Azimzadeh O, Tapio S, Ovsepian SV, Atkinson MJ. Long non-coding RNA PARTICLE bridges histone and DNA methylation. Sci Rep. 2017; 7:1790. https://doi.org/10.1038/s41598-017-01875-1.
18. O'Leary VB, Smida J, Buske FA, Carrascosa LG, Azimzadeh O, Maugg D, Hain S, Tapio S, Heidenreich W, Kerr J, Trau M, Ovsepian SV, Atkinson MJ. PARTICLE triplexes cluster in the tumor suppressor WWOX and may extend throughout the human genome. Sci Rep. 2017; 7:7163. https://doi.org/10.1038/s41598-017-07295-5.
19. Kovac M, Blattmann C, Ribi S, Smida J, Mueller NS, Engert F, Castro-Giner F, Weischenfeldt J, Kovacova M, Krieg A, Andreou D, Tunn PU, Durr HR, et al. Exome sequencing of osteosarcoma reveals mutation signatures reminiscent of BRCA deficiency. Nat Commun. 2015; 6:8940. https://doi.org/10.1038/ncomms9940.
20. Ottaviano L, Schaefer KL, Gajewski M, Huckenbeck W, Baldus S, Rogel U, Mackintosh C, de Alava E, Myklebost O, Kresse SH, Meza-Zepeda LA, Serra M, Cleton-Jansen AM, et al. Molecular characterization of commonly used cell lines for bone tumor research: a trans-European EuroBoNet effort. Genes Chromosomes Cancer. 2010; 49:40–51. https://doi.org/10.1002/gcc.20717.
21. Chou AJ, Merola PR, Wexler LH, Gorlick RG, Vyas YM, Healey JH, LaQuaglia MP, Huvos AG, Meyers PA. Treatment of osteosarcoma at first recurrence after contemporary therapy: the Memorial Sloan-Kettering Cancer Center experience. Cancer. 2005; 104:2214–21. https://doi.org/10.1002/cncr.21417.
22. Daw NC, Billups CA, Rodriguez-Galindo C, McCarville MB, Rao BN, Cain AM, Jenkins JJ, Neel MD, Meyer WH. Metastatic osteosarcoma. Cancer. 2006; 106:403–12. https://doi.org/10.1002/cncr.21626.
23. Marina NM, Smeland S, Bielack SS, Bernstein M, Jovic G, Krailo MD, Hook JM, Arndt C, van den Berg H, Brennan B, Brichard B, Brown KL, Butterfass-Bahloul T, et al. Comparison of MAPIE versus MAP in patients with a poor response to preoperative chemotherapy for newly diagnosed high-grade osteosarcoma (EURAMOS-1): an open-label, international, randomised controlled trial. Lancet Oncol. 2016; 17:1396–408. https://doi.org/10.1016/S1470-204530214-5.
24. Chang NS, Hsu LJ, Lin YS, Lai FJ, Sheu HM. WW domain-containing oxidoreductase: a candidate tumor suppressor. Trends Mol Med. 2007; 13:12–22. https://doi.org/10.1016/j.molmed.2006.11.006.
25. Pluciennik E, Kusinska R, Potemski P, Kubiak R, Kordek R, Bednarek AK. WWOX--the FRA16D cancer gene: expression correlation with breast cancer progression and prognosis. Eur J Surg Oncol. 2006; 32:153–7. https://doi.org/10.1016/j.ejso.2005.11.002.
26. Donati V, Fontanini G, Dell'Omodarme M, Prati MC, Nuti S, Lucchi M, Mussi A, Fabbri M, Basolo F, Croce CM, Aqeilan RI. WWOX expression in different histologic types and subtypes of non-small cell lung cancer. Clin Cancer Res. 2007; 13:884–91. https://doi.org/10.1158/1078-0432.CCR-06-2016.
27. Georgakilas AG, Tsantoulis P, Kotsinas A, Michalopoulos I, Townsend P, Gorgoulis VG. Are common fragile sites merely structural domains or highly organized "functional" units susceptible to oncogenic stress? Cell Mol Life Sci. 2014; 71:4519–44. https://doi.org/10.1007/s00018-014-1717-x.
28. Durkin SG, Glover TW. Chromosome fragile sites. Annu Rev Genet. 2007; 41:169–92. https://doi.org/10.1146/annurev.genet.41.042007.165900.
29. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001; 25:402–8. https://doi.org/10.1006/meth.2001.1262.

