INTRODUCTION
Worldwide, lung cancer is one of the most common cancers and the leading cause of cancer-related mortality among men, and the fourth leading cause of cancer morbidity and the second leading cause of cancer mortality among women [1, 2]. In China, lung cancer ranks first among the causes of cancer-related death [3]. Despite improvement in multimodal therapies, the 5-year survival rate for lung cancer is less than 10%. A major reason for this outcome is that a large proportion of lung cancer patients are diagnosed at advanced stages. The definitive mechanism of lung cancer is not fully understood. Evidence suggests that lung cancer is a multifactorial disease caused by genetic and environmental interactions [4–6].
NAD(P)H: quinone oxidoreductase (NQO1), formerly called DT-diaphorase, is a flavoenzyme associated with carcinogen metabolism [7]. As a two-electron reductase, the NQO1 protein can detoxify highly toxic quinones to less toxic hydroquinone analogues. NQO1 also acts as an antioxidant enzyme in vivo. By maintaining antioxidant forms of ubiquinone and reducing the endogenous quinones, NQO1 protects cells from oxidative damage [8]. Research has demonstrated that NQO1 can regulate tumor suppressor gene p53 activity, and thus influence cancer cell life [9]. The human NQO1 gene, consisting of 6 exons and 5 introns, is located on chromosome 16q22. Numerous single nucleotide polymorphisms (SNPs) of the NQO1 gene have been identified, but the most frequently studied SNP is NQO1 C609T [10–12]. NQO1 C609T (rs1800566, Pro187Ser) polymorphism is a C-to-T allele base-pair mutation in exon 6 (position 609) of the NQO1 gene. Such a gene mutation changes the encoded protein from a proline to a serine at position 187. This Pro187Ser mutated protein shows a reduced quinine reductase activity when compared with the wild-type protein [13].
Epidemiological studies have assessed the correlation between the NQO1 C609T polymorphism and lung cancer risk, but conflicting conclusions remain. In addition, several meta-analyses have been performed to reach a consensus but failed. To clarify the current uncertainty, we performed this comprehensive updated meta-analysis by incorporating all available literature.
RESULTS
Study characteristics
In the initial search stage, 81 potentially relevant published records were obtained. Through literature screening and abstracts reading, 38 articles were selected. We excluded 13 of those articles for the following reasons: 3 articles were case only research [14–16]; 1 article was duplicative [17], and 8 articles were meta-analyses [18–25]. However, 3 additional articles were identified from the references of the retrieval articles [26–28]. The workflow of the study selection process is shown in Figure 1. As a result, 37 studies (29 publications [21, 27–54]) with 8493 cases and 10,999 controls were used in the investigation (Table 1). Among the studies, 16 focused on Asians, 14 focused on Caucasians, 4 focused on African-Americans, 1 focused on Hawaiians, 1 focused on Hispanics, and 1 focused on a mixed population; 17 studies were hospital-based designs, 19 were population-based designs, and 1 was a mixed-based design. Of these studies, 23 were considered low quality (quality score ≤ 9), and 14 were considered high quality (quality score > 9). The control genotype frequencies in agreement with the Hardy-Weinberg equilibrium (HWE) were observed in 28 studies, but were not available in 9 studies.
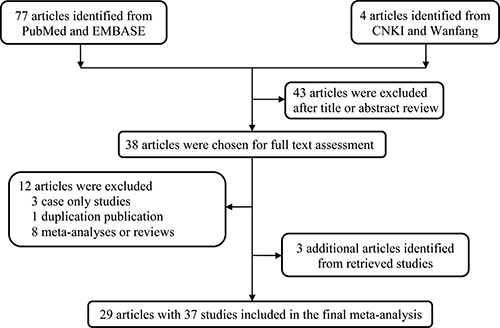
Figure 1: The main flowchart of this work.
Table 1: The baseline characteristics of all qualified studies in this meta-analysis
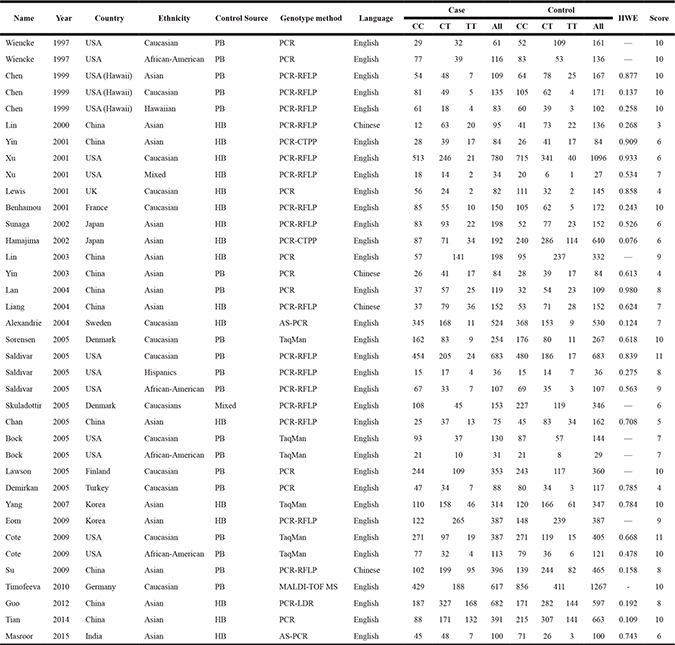
HB, hospital based; PB, population based; PCR, polymerase chain reaction; AS-PCR, Allele specific-PCR; PCR-RFLP, PCR-restriction fragment length polymorphism; MALDI-TOF MS, Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry; HWE, Hardy-Weinberg equilibrium.
Meta-analysis results
The calculated results of the meta-analysis are shown in Table 2 and Figure 2. Overall, no significant correlation between the NQO1 C609T polymorphism and lung cancer risk was observed in any of the genetic models (TT vs. CC: OR = 0.91, 95% CI = 0.74–1.12; CT vs. CC: OR = 1.08, 95% CI = 0.98–1.22; TT vs. CT+CC: OR = 1.13, 95% CI = 0.97–1.33; T vs. C: OR = 1.09, 95% CI = 0.99–1.20; and CT+TT vs. CC: OR = 1.04, 95% CI = 0.94–1.16).
Table 2: Meta-analysis of the correlation between NQO1 C609T polymorphism and lung cancer risk
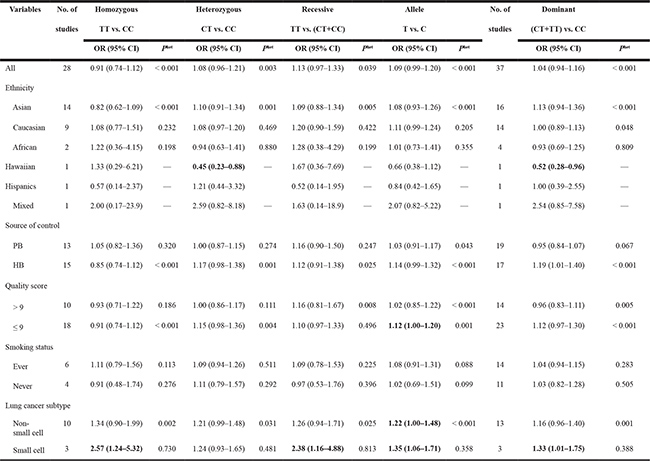
Het, heterogeneity; HB, hospital based; PB, population based; NA, not available.
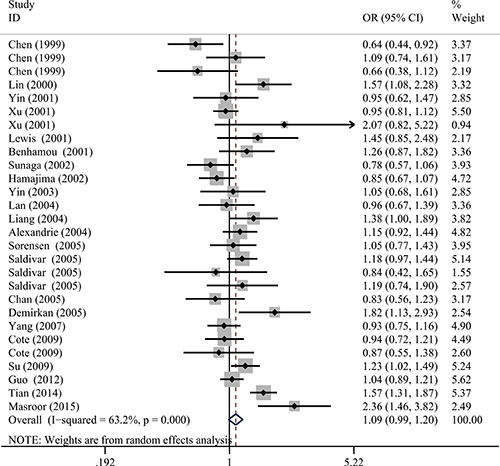
Figure 2: Forest plot for the correlation between the NQO1 C609T polymorphism and lung cancer susceptibility under the allele comparison model. The horizontal lines represent the study-specific ORs and 95% CIs. The diamond represents the pooled results of OR and 95% CI.
Further subgroup analysis by ethnicity, control source, quality score, and smoking status still did not yield a significant association, except for the heterozygous model and dominant model in the Hawaiian subgroup (CT vs. CC: OR = 0.45, 95% CI = 0.23–0.88; CT+TT vs. CC: OR = 0.52, 95% CI = 0.28–0.96) and the allele model in the quality score ≤ 9 subgroup (T vs. C: OR = 1.12, 95% CI = 1.00–1.20). In the subgroup analysis by lung cancer subtype, statistically significant increased risks were found among non-small cell lung cancer for T vs. C (OR = 1.22, 95% CI = 1.00–1.48) and small cell lung cancer for TT vs. CC (OR = 2.57, 95% CI = 1.24–5.32), TT vs. CT+CC (OR = 2.38, 95% CI = 1.16–4.88), T vs. C (OR = 1.35, 95% CI = 1.06–1.71) and CT+TT vs. CC (OR = 1.33, 95% CI = 1.01–1.75).
Heterogeneity and sensitivity analysis
Before calculating the ORs and 95% CIs, we used the Q-test and the I-squared statistics to test between-study heterogeneity. In the pooled analysis, significant heterogeneity exists among all five genetic models (P < 0.001, I2 = 55.2% for Homozygous, P = 0.003, I2 = 47.2% for Heterozygous, P = 0.039, I2 = 34.5% for Recessive, P < 0.001, I2 = 63.2% for Allele and P < 0.001, I2 = 57.0% for Dominant genetic models). Thus, the random-effect model was used to generate wider CIs. Moreover, we adopted a sequential leave-one-out sensitivity analysis to assess the influence of a single study on the combined ORs. After omitting each study, no substantial changes in ORs were observed, which suggested the robust and reliability of this meta-analysis (Figure 3).
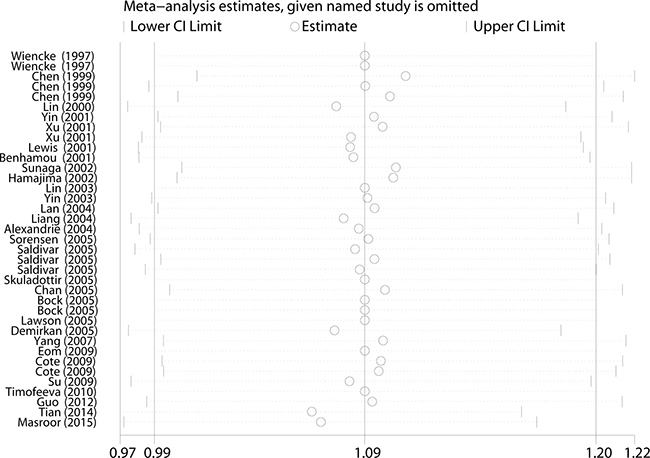
Figure 3: Sensitivity analysis of the association between NQO1 C609T and lung cancer susceptibility under the allele comparison model. Each point represents the recalculated OR after deleting a separate study.
Publication bias
No evidence of obvious asymmetry in Begg’s funnel plots was identified by visual observation (Figure 4). In addition, Egger’s test also indicated that no publication bias was shown among the studies (TT vs. CC: P = 0.48; CT vs. CC: P = 0.45; TT vs. CT+CC: P = 0.76; T vs. C: P = 0.93; and CT+TT vs. CC: P = 0.79).
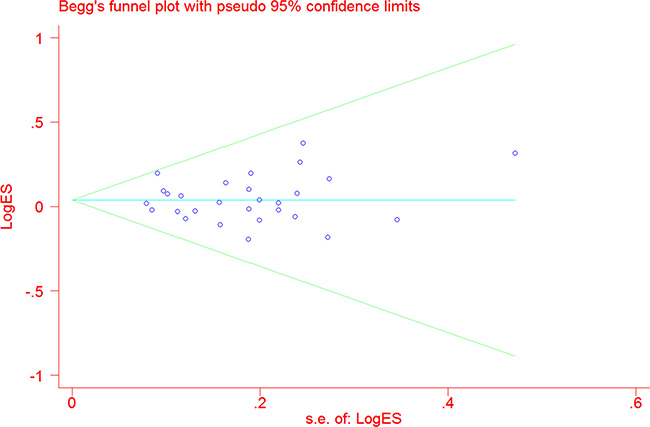
Figure 4: Funnel plot analysis to detect publication bias for NQO1 C609T polymorphism under the allele comparison model. Each point represents a separate study.
Trial sequential analysis
The TSA showed that the cumulative Z-curve failed to cross the trial monitoring boundary before reaching the required information size, suggesting that more trials are needed to further verify the conclusions (Figure 5).
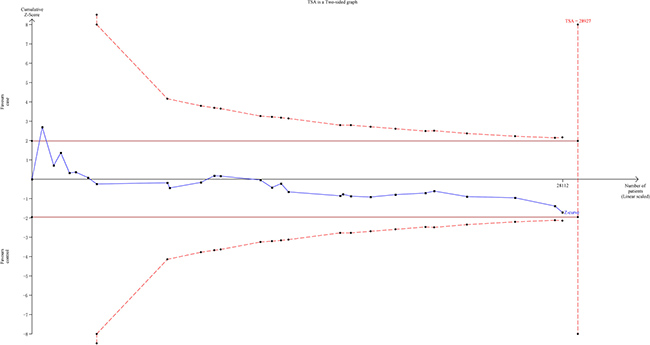
Figure 5: The required information size to demonstrate the correlation between NQO1 C609T polymorphism with lung cancer susceptibility. The solid blue line is the cumulative Z-curve. The dashed inward-sloping line to the left represents the trial sequential monitoring boundaries.
DISCUSSION
In this meta-analysis, we investigated the role of NQO1 C609T polymorphism in lung cancer susceptibility. The obtained results suggest that no significant relation exists. To date, this meta-analysis includes the largest samples used in the investigation of the function of NQO1 C609T in lung cancer susceptibility.
Numerous studies have investigated the activity of the NQO1 gene C609T polymorphism in lung cancer risk. In 1999, Chen et al. [30] claimed that NQO1 C609T polymorphism correlates with decreased lung cancer risk in Japanese. In another study conducted in England that consists of 82 lung cancer patients and 145 controls, Lewis et al. [31] failed to detect any correlation between NQO1 C609T and lung cancer risk. However, Masroor et al. [32] reported that the NQO1 609TT genotype could increase the risk of lung cancer in an Indian population of 100 lung cancer cases and 100 healthy controls. Several meta-analyses have been performed to obtain a clear correlation between NQO1 C609T and lung cancer risk. In a pilot meta-analysis conducted by Kiyohara et al. [22], only 10 studies were included, consisting of 2746 cases and 3902 controls. They found that the Ser allele is a protective factor against lung cancer in Asians, but not in Caucasians. In a meta-analysis that included 25 articles (32 studies) with 7522 cases and 9291 controls, Lou et al. [25] failed to observe any correlation between NQO1 C609T polymorphism and lung cancer risk overall in African-Americans, East Asians, or Hispanics. However, the results suggested that NQO1 C609T polymorphism might correlate with lung cancer risk in Caucasians.
This updated and comprehensive meta-analysis was performed to better elucidate the correlation between NQO1 C609T polymorphism and lung cancer risk. However, we failed to observe any significant relation between NQO1 C609T and lung cancer risk in the pooled analysis in any of the five genetic models. Subgroup analysis by ethnicity suggested that the T allele might be a protective factor in Hawaiians (CT vs. CC: OR = 0.45, 95% CI = 0.23–0.88; CT+TT vs. CC: OR = 0.52, 95% CI = 0.28–0.96). In this Hawaiian subgroup, only one study, with 83 cases and 102 controls was included, which limits the strength of this result. Thus, more case-control studies in Hawaiians are needed to elucidate the role of C609T polymorphism in lung cancer risk. In another subgroup stratified by quality score, we did not detect any correlation between NQO1 C609T and lung cancer risk, yet the T allele acts as a risk factor for lung cancer in low-quality studies under the allele model (T vs. C: OR = 1.12, 95% CI = 1.00–1.20). The insufficient statistical power of the relatively low-quality studies should be also considered. It is possible that such insufficient statistical power might result in false positive results. When we stratified the analysis on the basis of lung cancer subtype, an increased risk of the NQO1 C609T polymorphism on small cell lung cancer was observed. Previous pooled analyses have demonstrated that small cell lung cancer was more frequently observed in patients exposed to tobacco smoke [29]. The NQO1 C609T polymorphism might result in failure of NQO1 protein to detoxify highly toxic quinones, and thus have some consequences for smoking-related small cell lung cancer risk.
In this meta-analysis, we adopted many measurements to be certain of the credibility of our conclusion. First, we incorporated all eligible studies not only written in English but also written in Chinese to expand the included sample numbers. Then the sensitivity analysis and the publication bias were assessed according to the Cochrane protocol. The sensitivity analysis indicated that the results were robust, and Funnel plots suggested that no obvious publication bias was observed. The TSA was also used to test the conclusion reliability of this meta-analysis. The Z-curve failed to cross the trial monitoring boundary before reaching the required information size. Thus, more studies are needed to confirm or refute this finding. However, the limitations must be pointed out when the results of this meta-analysis are interpreted. First, we found that most of the comparisons show significant between-study heterogeneity, which might reduce the validity of the conclusion. Second, the strength of correlation was obtained by use of unadjusted estimates. Adjustment analysis was absent because of the lack of original data, such as environmental factors, age, drinking status, and gene-environment interactions, which restrains further analysis for risk factors. Nearly all the included case-control studies were conducted among Asians and Caucasians. Other ethnicities, such as Africans, were not studied. Additional studies are needed to confirm such conclusions from other ethnicities, especially Africans. Publication bias and language bias still exist, because only published studies and only the studies written in English or Chinese are included.
The current meta-analysis provides evidence that NQO1 C609T polymorphism is not correlated with lung cancer risk, from the perspective of the former case-control studies. Further high-quality investigations with more detailed environmental exposure information and larger sample sizes are warranted to confirm our findings.
MATERIALS AND METHODS
Publication search
We performed a comprehensive literature search of PubMed and EMBASE by adopting the combination of the following phrases: “polymorphism or single nucleotide polymorphism or SNP or variant” and “NQO1 or quinone reductase or quinone oxidoreductase or DT-diaphorase or DTD,” and “lung cancer or lung neoplasm or lung carcinoma.” To enlarge the included studies, we also searched China National Knowledge Infrastructure (CNKI) and Wanfang databases using the above phrases in Chinese. In addition, eligible studies in the references of retrieved articles were also screened. The literature search was performed before April 2017 without any language publication restrictions. If an article contained two or more ethnic subpopulations, they were treated as separate studies. Only the largest study was included if two or more articles contained overlapping data. The designation and writing of this meta-analysis were under the guidelines of Preferred Reporting Items for Systematic Reviews and Meta-analyses.
Eligibility criteria
All the included articles in this meta-analysis met the following criteria: (1) contained unrelated case-control studies, (2) evaluated the correlation of NQO1 C609T polymorphism with lung cancer risk, (3) contained enough genotype distribution information to calculate ORs and 95% CIs, and (4) NQO1 C609T genotype frequency in control subjects were in agreement of the Hardy-Weinberg equilibrium (HWE).
Data extraction
Authors Jiawen Huang and Huiran Lin screened the articles and extracted data from all eligible studies, respectively. The data include first author’s surname, publication year, country, ethnicity, the source of control subjects, genotyping methods, quality score, and numbers of cases and controls with CC, CT, and TT genotypes. Conflicting data were resolved by discussion after consensus was reached
Quality assessment
The quality of each study was assessed by use of the quality assessment criteria [30–32]. The evaluation items were as follows: representativeness of case, representativeness of control, ascertainment of cancer, control selection, genotyping examination, HWE, and total sample size. Each study was evaluated on a scale from 0–15. Quality score of studies ranges from 0 to 15 points. Studies with scores ≤ 9 were of low quality, whereas those with scores > 9 were of high quality. The detail of score of quality assessment was listed in Supplementary Table 1.
Statistical methods
All five genetic models, homozygous model (TT vs. CC), heterozygous model (CT vs. CC), recessive model (TT vs. CT + CC), dominant model (CT + TT vs. CC), and allele comparison (T vs. C) were adopted to investigate the correlation between NQO1 C609T polymorphism and lung cancer risk. The strength of such correlation was assessed by calculating ORs with the corresponding 95% CIs. Stratification analyses were also performed by ethnicity, and source of control subjects, quality score, HWE in control subjects, smoking status, and lung cancer subtype. Between-study heterogeneity was analyzed by chi squared-based Q-test. When the studies were found to be homogeneous (with P > 0.10 for the Q-test), the fixed-effects model (the Mantel-Haenszel method) was used to estimate the pooled OR. Otherwise, the random-effects model (the DerSimonian and Laird method) was adopted. Sensitivity analysis was done by individually removing studies one by one and reanalyzing the pooled risk estimates. Begg’s funnel plot and Egger’s linear regression were used to estimate potential publication bias, with that asymmetric plot and a P-value < 0.05, indicating the presence of publication bias. HWE in the control subjects was tested by use of goodness-of-fit chi-squared test. A value of P < 0.05 was considered as departure from HWE. All statistical analyses were completed by STATA Version 11.0 software (Stata Corporation, College Station, TX). All the statistics were two-sided, with significant findings set at a P-value of less than 0.05.
Trial sequential analysis
To avoid the random errors caused by repeated significance testing and dispersed data, we performed the trial sequential analysis (TSA). First, the required information size was evaluated by considering an overall type I error (a) of 5% and type II error (b) of 20%. Then TSA monitoring boundaries were constructed on the basis of required information size as well as risk for type I and type II errors. If the cumulative Z-curve (blue line) crosses a TSA monitoring boundary (red lines) before reaching the required information size, the robustness of evidence might be confirmed and no further trials are necessary. Otherwise, to get a robust conclusion, more trials are needed.
ACKNOWLEDGMENTS AND FUNDING
This study was supported by grant from Science and Technology Planning Project of Guangdong Province (2016ZC0045).
CONFLICTS OF INTEREST
The authors have declared that no competing interests exist.
REFERENCES
1. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015; 136:E359–386.
2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015; 65:5–29.
3. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66:115–132.
4. Lan Q, Hsiung CA, Matsuo K, Hong YC, Seow A, Wang Z, Hosgood HD 3rd, Chen K, Wang JC, Chatterjee N, Hu W, Wong MP, Zheng W, et al. Genome-wide association analysis identifies new lung cancer susceptibility loci in never-smoking women in Asia. Nat Genet. 2012; 44:1330–1335.
5. Fang X, Yin Z, Li X, Xia L, Quan X, Zhao Y, Zhou B. Multiple functional SNPs in differentially expressed genes modify risk and survival of non-small cell lung cancer in Chinese female non-smokers. Oncotarget. 2017; 8:18924–18934. https://doi.org/10.18632/oncotarget.14836.
6. David SP, Wang A, Kapphahn K, Hedlin H, Desai M, Henderson M, Yang L, Walsh KM, Schwartz AG, Wiencke JK, Spitz MR, Wenzlaff AS, Wrensch MR, et al. Gene by Environment Investigation of Incident Lung Cancer Risk in African-Americans. EBioMedicine. 2016; 4:153–161.
7. Jaiswal AK, McBride OW, Adesnik M, Nebert DW. Human dioxin-inducible cytosolic NAD(P)H:menadione oxidoreductase. cDNA sequence and localization of gene to chromosome 16. J Biol Chem. 1988; 263:13572–13578.
8. Siegel D, Gustafson DL, Dehn DL, Han JY, Boonchoong P, Berliner LJ, Ross D. NAD(P)H:quinone oxidoreductase 1: role as a superoxide scavenger. Mol Pharmacol. 2004; 65:1238–1247.
9. Asher G, Lotem J, Kama R, Sachs L, Shaul Y. NQO1 stabilizes p53 through a distinct pathway. Proc Natl Acad Sci USA. 2002; 99:3099–3104.
10. Bagatir G, Sirma S, Palanduz S, Ozturk S, Cefle K, Ozbek U, Yenerel M, Nalcaci M. The frequency of C609T polymorphism in the NQO1 gene and its relation to cytogenetic abnormalities in patients with myelodysplastic syndrome. Cell Mol Biol (Noisy-le-grand). 2016; 62:61–65.
11. Yang S, Jin T, Su HX, Zhu JH, Wang DW, Zhu SJ, Li S, He J, Chen YH. The association between NQO1 Pro187Ser polymorphism and bladder cancer susceptibility: a meta-analysis of 15 studies. PLoS One. 2015; 10:e0116500.
12. Zhang Y, Yang D, Zhu JH, Chen MB, Shen WX, He J. The association between NQO1 Pro187Ser polymorphism and urinary system cancer susceptibility: a meta-analysis of 22 studies. Cancer Invest. 2015; 33:39–40.
13. Siegel D, Anwar A, Winski SL, Kepa JK, Zolman KL, Ross D. Rapid polyubiquitination and proteasomal degradation of a mutant form of NAD(P)H:quinone oxidoreductase 1. Mol Pharmacol. 2001; 59:263–268.
14. Kolesar JM, Dahlberg SE, Marsh S, McLeod HL, Johnson DH, Keller SM, Schiller JH. The NQO1*2/*2 polymorphism is associated with poor overall survival in patients following resection of stages II and IIIa non-small cell lung cancer. Oncol Rep. 2011; 25:1765–1772.
15. Song SY, Jeong SY, Park HJ, Park SI, Kim DK, Kim YH, Shin SS, Lee SW, Ahn SD, Kim JH, Lee JS, Choi EK. Clinical significance of NQO1 C609T polymorphisms after postoperative radiation therapy in completely resected non-small cell lung cancer. Lung Cancer. 2010; 68:278–282.
16. Yang SY, Yang TY, Li YJ, Chen KC, Liao KM, Hsu KH, Tsai CR, Chen CY, Hsu CP, Hsia JY, Chuang CY, Tsai YH, Chen KY, et al. EGFR exon 19 in-frame deletion and polymorphisms of DNA repair genes in never-smoking female lung adenocarcinoma patients. Int J Cancer. 2013; 132:449–458.
17. Lin P, Wang HJ, Lee H, Lee HS, Wang SL, Hsueh YM, Tsai KJ, Chen CY. NAD(P)H: quinone oxidoreductase polymorphism and lung cancer in Taiwan. J Toxicol Environ Health A. 1999; 58:187–197.
18. Chao C, Zhang ZF, Berthiller J, Boffetta P, Hashibe M. NAD(P)H:quinone oxidoreductase 1 (NQO1) Pro187Ser polymorphism and the risk of lung, bladder, and colorectal cancers: a meta-analysis. Cancer Epidemiol Biomarkers Prev. 2006; 15:979–987.
19. Gresner P, Gromadzinska J, Wasowicz W. Polymorphism of selected enzymes involved in detoxification and biotransformation in relation to lung cancer. Lung Cancer. 2007; 57:1–25.
20. Guo S, Gao M, Li X, Li Y, Chu S, Zhu D, Niu W. Lack of association between NADPH quinone oxidoreductase 1 (NQO1) gene C609T polymorphism and lung cancer: a case-control study and a meta-analysis. PLoS One. 2012; 7:e47939.
21. Huang RHZ, Zhou X, Lin Z, Peng X. The Meta analysis of the NQO1 C609T polymorphism and lung cancer risk. J Fujian Med Univ. 2014; 48:141–146.
22. Kiyohara C, Yoshimasu K, Takayama K, Nakanishi Y. NQO1, MPO, and the risk of lung cancer: a HuGE review. Genet Med. 2005; 7:463–478.
23. Liu Y, Zhang D. The NQO1 C609T polymorphism and risk of lung cancer: a meta-analysis. Asian Pac J Cancer Prev. 2011; 12:3091–3095.
24. Lou Y, Li R, Xiong L, Gu A, Shi C, Chu T, Zhang X, Gu P, Zhong H, Wen S, Han B. NAD(P)H: quinone oxidoreductase 1 (NQO1) C609T polymorphism and lung cancer risk: a meta-analysis. Tumour Biol. 2013; 34:3967–3979.
25. Zhang L, Yan H, Wang S, Hong J, Wang X, Liu Z, Zhang Y, Cao B. A systematic analysis between NQO1 gene polymorphism and susceptibility of lung cancer. J Clin Pathol Res. 2014; 34:701–712.
26. Demirkan A, Vural B, Ustuner Z, Uygun K, Acikalin B, Derin D, Saip P, Ozbek U. CYP1A1, GSTM1 and NQO1 gene polymorphisms: genetic risk factors for small cell lung cancer. Turkish Journal of Cancer. 2005; 35:171–176.
27. Eom SY, Zhang YW, Kim SH, Choe KH, Lee KY, Park JD, Hong YC, Kim YD, Kang JW, Kim H. Influence of NQO1, ALDH2, and CYP2E1 genetic polymorphisms, smoking, and alcohol drinking on the risk of lung cancer in Koreans. Cancer Causes Control. 2009; 20:137–145.
28. Hamajima N, Matsuo K, Iwata H, Shinoda M, Yamamura Y, Kato T, Hatooka S, Mitsudomi T, Suyama M, Kagami Y, Ogura M, Ando M, Sugimura Y, Tajima K. NAD(P)H: quinone oxidoreductase 1 (NQO1) C609T polymorphism and the risk of eight cancers for Japanese. Int J Clin Oncol. 2002; 7:103–108.
29. Pesch B, Kendzia B, Gustavsson P, Jockel KH, Johnen G, Pohlabeln H, Olsson A, Ahrens W, Gross IM, Bruske I, Wichmann HE, Merletti F, Richiardi L, et al. Cigarette smoking and lung cancer--relative risk estimates for the major histological types from a pooled analysis of case-control studies. Int J Cancer. 2012; 131:1210–1219.
30. He J, Liao XY, Zhu JH, Xue WQ, Shen GP, Huang SY, Chen W, Jia WH. Association of MTHFR C677T and A1298C polymorphisms with non-Hodgkin lymphoma susceptibility: evidence from a meta-analysis. Sci Rep. 2014; 4:6159.
31. Fu W, Zhuo ZJ, Chen YC, Zhu J, Zhao Z, Jia W, Hu JH, Fu K, Zhu SB, He J, Liu GC. NFKB1 -94insertion/deletion ATTG polymorphism and cancer risk: Evidence from 50 case-control studies. Oncotarget. 2017; 8:9806–9822. https://doi.org/10.18632/oncotarget.14190.
32. Huang J, Liu X, Tang LL, Long JT, Zhu J, Hua RX, Li J. XPG gene polymorphisms and cancer susceptibility: evidence from 47 studies. Oncotarget. 2017; 8:37263–37277. https://doi.org/10.18632/oncotarget.16146.

