INTRODUCTION
Ubiquitination modification is a ubiquitous modifying method of post-translation of proteins to induce protein degradation, which can be regulated and reversed by deubiquitinating enzymes [1, 2]. UCHL1 also known as PGP9.5, is a deubiquitinating enzyme that hydrolyzes C-terminal esters and amides for generating monomeric ubiquitin [3]. UCHL1 appears to be a general marker of nerves and neuroendocrine cells across the vertebrate kingdom [3], i.e., mammals, birds, reptiles, amphibians, fish, and even marsupials. UCHL1/PGP 9.5 has also been reported in various locations in numerous species, including rainbow trout, Oncorhychus mykiss [4]; barfin flounder, Verasper moseri [5]; penguins, Spheniscidae Family [6]; Surinam caiman, Caiman crocodilus crocodilus [7]; flying fox, Pteropus vampyrus [8]. This pattern suggests that UCHL1 is a conservative protein enzyme involved in special biological functions.
Different localization of UCHL1 plays an important role in regulating cellular proliferation and differentiation by ubiquitination modification of post-translational proteins. Three existence forms of UCHL1 appear in animals, including cytoplasmic localizing, nuclear localizing, and secretory UCHL1. Cytoplasmic localization of UCHL1 is the main localization in cells that serve as markers for the non-proliferating spermatogonia in primates [9], and neurons in mice and humans. Nuclear localization of UCHL1 occurs in approximately 5% of neurons in the central and peripheral nervous systems [10], but the function is unknown. Extracellular UCHL1 in the body fluid and spaces within tissues has also been reported. UCHL1/PGP 9.5 in serum is measured at approximately 100 ng/ml [10]. There have been several reports regarding UCHL1/PGP 9.5 localization in the renal tubules of mice and its distribution in the epididymis of rat [11], human [12], and mouse [13]. It was thought that the expression of UCHL1/PGP 9.5 in any part of the renal tubule or the epididymal cauda was possibly caused by re-absorption from plasma [11] or circulatory system, because there was no expression of UCHL1 mRNA in the renal tubule or in the epididymal cauda [10].
The ubiquitin pathway plays a critical role in the progression of spermatogenesis through the mitotic, meiotic, and post-meiotic phases [14]. Generally, conserved expression of UCHL1 regulates spermatogensis in mammalian testes [15]. UCHL1 is expressed in spermatogonia and Sertoli cells in rodents [16], whereas in other mammals, including humans, non-human primates [14], and domestic animals, such as bovines [17, 18], pigs [19], goats [20, 21], it is only expressed in gonocytes and spermatogonia in seminiferous tubules. UCHL1 is considered an optimal marker for spermatogonia and for pre-meiotic male germ cells [16]. For example, in fish, UCHL1, via the ubiquitin-proteasome system, may play an important role in the gonadal transformation process in the rice field eel (Monopterus albus) [22]. In amphibians, UCHL1 is possibly involved in the process of progesterone-induced toad oocyte maturation [23].
The Chinese giant salamander (CGS, Andrias davidianus) is the largest extant amphibian in the world [24, 25], belonging to the Cryptobranchidae family, which only contains three species: (Cryptobranchus alleganiensisin in North America, Andrias japonicus in Japan, and Andrias davidianus in China). As an endangered species, the CGS has received increasing attention in evolutionary, comparative biology, and other studies [26-29]. Although there are morphological differences between urodele testis and mammal testis, the spermatogenetic process in urodele amphibians is similar to that of mammals. Many genes take part in regulating the process of spermatogenesis. To date, the molecular characterization and cellular localization of UCHL1 has not been reported during development of the CGS gonad. In addition, the potential function of UCHL1 in spermatogenesis of CGS is not clear. In this study, the cDNA sequence of UCHL1 was identified and characterized for the Chinese giant salamander and the potential function of UCHL1 was analyzed during spermatogenesis and the development of CGS gonads.
RESULTS
Sequence characterization of UCHL1
A fragment of UCHL1 was obtained from the transcriptome database for the Chinese giant salamander. It was confirmed by specific amplification with primers UCHL1-F and UCHL1-R. The total sequence of UCHL1 mRNA has been obtained by 3′ and 5′ RACE and deposited in the GenBank database under accession number (MF418646). The cDNA sequence consisted of 1087 bp, including a 195 bp 5′ UTR, 223 bp 3′ UTR, and a 669 bp ORF with content of G + C (48.6 %) (Figure 1). The CDS of UCHL1 encoded a putative polypeptide of 222 amino acids with a calculated molecular mass of 24.6 kDa and a theoretical pI of 5.19. SignalP analysis showed the typical signal peptide was not found. The number of the potential phosphorylation sites and O-linked glycosylation sites were 14 and one, respectively (Figure 1).
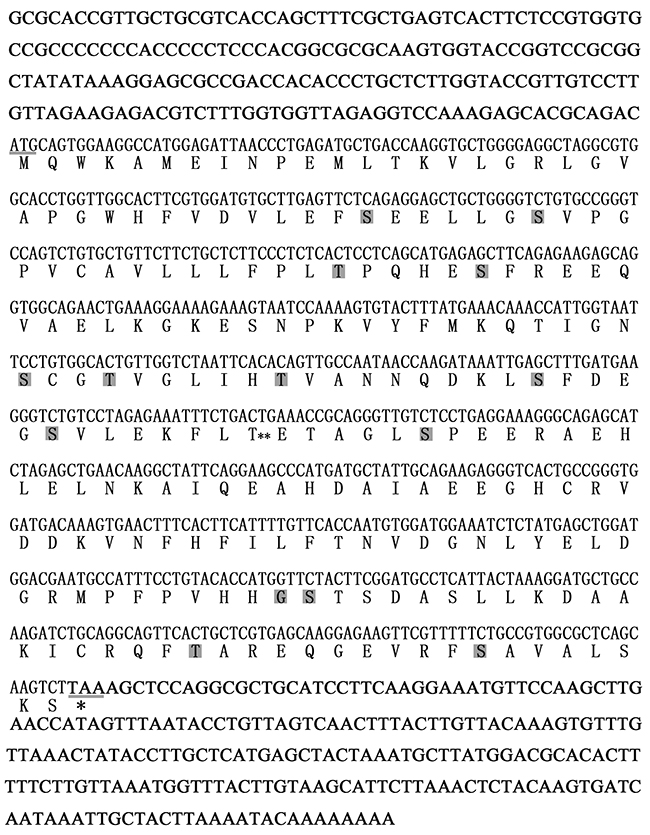
Figure 1: Characterization of amino acid sequences and cDNA nucleotide sequences of CGS UCHL1. The start codon and stop codon are underlined. The gray boxes show the phosphorylation sites. The double asterisks (**) display the O-linked glycosylation site.
The results of Blast showed that amino acid sequences of CGS UCHL1 was most homologous with human UCHL1 (up to 73.99 %), secondly that of mice (up to 73.09 %) and then Xenopus. laevis (up to 70.4 %) (Figure 2). The phylogenetic analysis demonstrated that the phylogeny of CGS UCHL1 and human UCHL1 was more closely related than that of any other vertebrate (Figure 3).
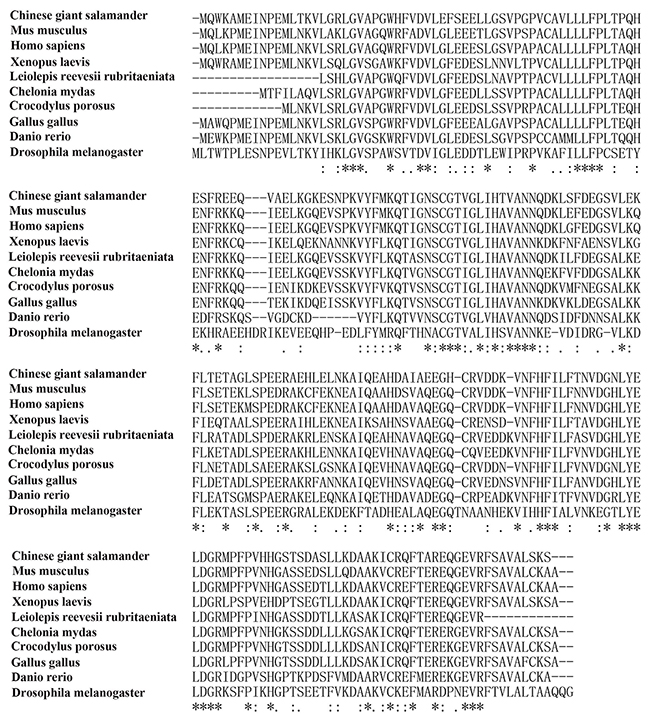
Figure 2: Multiple alignments of UCHL1 amino acid sequences of 10 species.
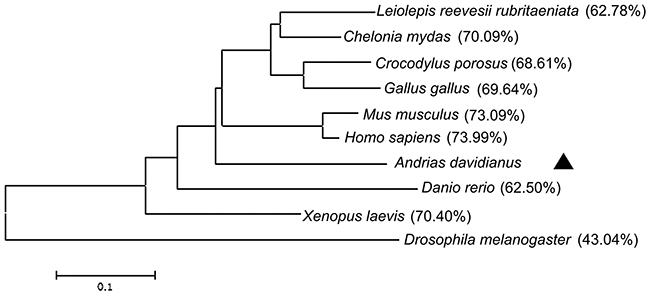
Figure 3: Phylogenetic tree of UCHL1 and other homologs. The phylogenetic tree was constructed based on the full-length amino acid sequences by the neighbor-joining method and the bootstrap value was set at 1000. The CGS UCHL1 is labeled with triangle (▲). Percentage of bootstrap replications is shown in the figure. The scale bar (0.01) represents the genetic distance.
Expression level of UCHL1 mRNA in testes of CGSs at different developmental stages
The RT-qPCR was employed to examine the level of UCHL1 in testes of CGSs at different developmental stages (Figure 4). We found that UCHL1 mRNA was expressed in all different developmental testis tissues. The peak level of UCHL1 mRNA appeared in testes of 2-year-old CGSs. Further, the expression level of UCHL1 in adult testes (at least 4 years old) was lower during the breeding period (August) than in the post-breeding period (November).
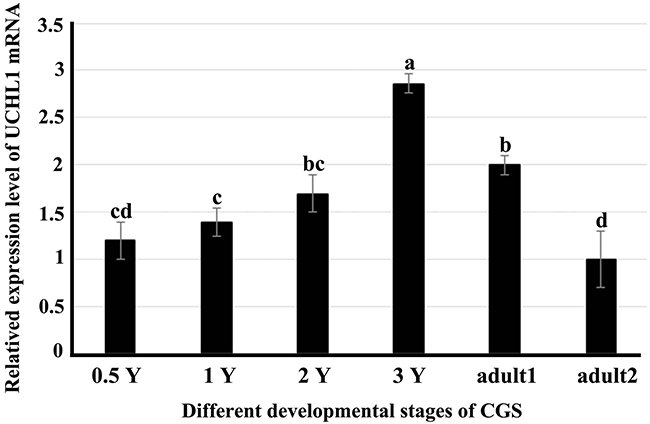
Figure 4: Expression profiles of UCHL1 mRNA in male gonads at different developmental stages by real-time PCR with β-actin as an internal reference. 0.5 Y, 1 Y, 2 Y, and 3 Y represented samples of testes or gonads from 0.5-year-old, 1-year-old, 2-year-old, 3-year-old and adult CGSs. Adult1 denoted the tissue samples of adult CGSs in November (post-breeding period); adult2 denoted the tissue samples of adult CGSs in August (breeding period). Similar letters denote non-significant differences (p > 0.05). Different letters indicate a significant difference (p < 0.05).
Localization of UCHL1 in testis of CGSs at different developmental stages by IHC
As stated above, the amino acids sequences of UCHL1 protein were highly conservative. An antibody of human UCHL1 was used to localize UCHL1 in the testes of CGSs at different developmental stages by IHC.
In the gonad of the 0.5-year-old CGSs, UCHL1 was located in the nucleus of large cells with loose chromatin, which was distributed in the peripheral region of the gonads (Figure 5A).
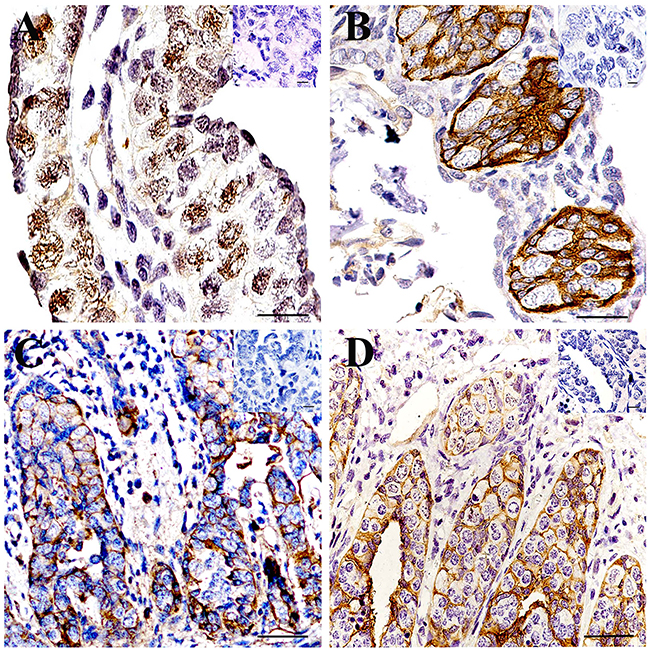
Figure 5: Localization of UCHL1 in gonads at different developmental CGS by IHC. (A) 0.5-year-old CGSs; (B) 1-year-old CGS; (C) 2-year-old CGS; (D) 3-year-old CGS. The inset represents the negative controls. Scale bar, 80 μm.
In the testes of the 1-year-old CGSs, these UCHL1 positive cells aggregated together to form the primary seminiferous lobules without a lumen. UCHL1 was only distributed on the membrane of the cells in seminiferous lobules (Figure 5B).
In histologically, the lumen of seminiferous lobules of CGSs had formed after 2 years of age (Figure 5C) and the lumen was smaller than those of 3-year-old CGSs (Figure 5D). The UCHL1 positive cells displayed a tight junction state to form the lobular lumen wall at this stage (Figure 5C, 5D). In the testes of the 3-year-old CGSs, the UCHL1 positive cells were regularly arranged in the seminiferous lobules wall (Figure 5D).
In testes of adult CGSs, the expression of UCHL1 significantly decreased in the different segments of the seminiferous lobules. From the proximal to the distal end of the seminiferous lobules, the expression level of UCHL1 gradually decreased, until it was no longer detected (Figure 6A). Interestingly, UCHL1 was distributed in the extracellular matrix, and little of the UCHL1 positive signal appeared in the cells of the seminiferous lobules (Figure 6). The seminiferous lobule was divided into different cysts by the extracellular UCHL1. High levels of UCHL1 expression occurred in the proximal seminiferous lobule where there were germ cells at different developmental stages. Meanwhile, no UCHL1 expression occurred in the distal end of the seminiferous lobule, which was full of mature sperm (Figure 6).
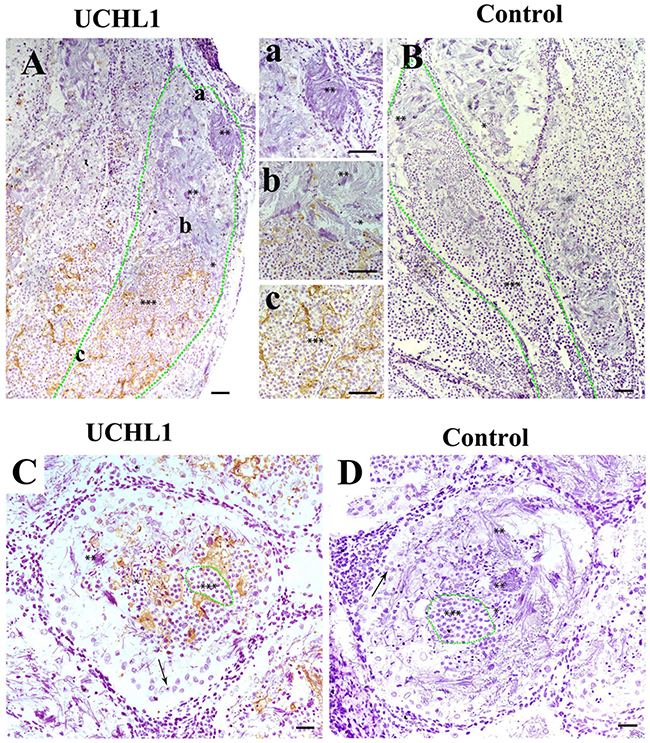
Figure 6: Localization of UCHL1 in male gonads of adult CGS by IHC. The different morphological sperm was in seminiferous lobules, i.e., the round spermatid (***), sperm with condensed nucleus (**); sperm with non-condensing nucleus (*). (A, B) Vertical section. The zone of green dots denotes a seminiferous lobule. a, b, and c represent the enlarged corresponding regions of Figure A, respectively. (C, D) Transverse section. The zone of green dots denote a speculative cyst. Arrows denote a type of large cell near the well of seminiferous lobules. Scale bar, 200 μm.
Developmental expression of UCHL1 protein in testes of CGSs at different stages
Western blot was performed to detect UCHL1 protein level of testes at different developmental stages (Figure 7). The testis tissue was divided into the pellets and the supernatant fractions. The levels of UCHL1 in testicular tissue pellets removed and tissue fluid are shown in Figure 7A; furthermore, the expression level of UCHL1 protein in testicular leaching liquid is displayed in Figure 7B (see materials and methods). The peak level of UCHL1 protein in testis pellets appeared for the 3-year-old CGSs, however, the level of UCHL1 protein was lowest in testes of the 2-year-old CGSs. In adult testes, the expression level of UCHL1 protein was lower in the breeding season (August) than in the post-reproduction period (November) (Figure 7A). Furthermore, the level of UCHL1 protein in testicular leaching liquid was gradually increased with male gonad development. Interestingly, the expression level of UCHL1 protein in testicular leaching liquid was higher in breeding period than after reproduction in adult testes (Figure 7B). Furthermore, UCHL1 protein was detected in the exosomes from testicular leaching liquid (Figure 8).
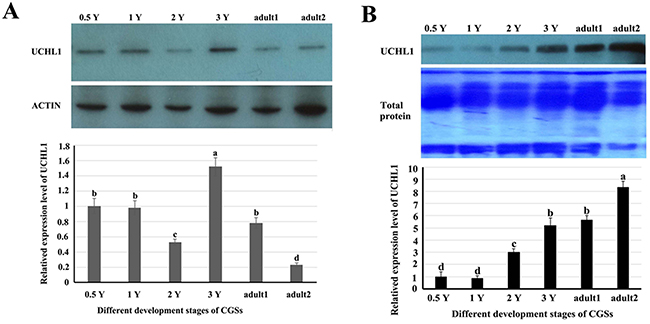
Figure 7: The related expression level of UCHL1 protein in the tissues or in the leaking liquid of male gonads at different developmental stages. (A) The level of UCHL1 protein in the tissues of male gonads at different developmental stages. 0.5 Y, 1 Y, 2 Y, 3 Y, adult1, and adult2 represented samples of gonads from 0.5-year-old, 1-year-old, 2-year-old, 3-year-old, adult CGSs in November, and adult CGSs in August, respectively. (B) The level of UCHL1 protein in the tissues or in the leaking liquid of male gonads at different development stages. 0.5 Y, 1 Y, 2 Y, 3 Y, adult 1, and adult2 represent the samples of the gonads leaking liquid from 0.5-year-old, 1-year-old, 2-year-old, 3-year-old, adult CGSs in November, and adult CGSs in August, respectively. Same letters denote non-significant difference (p > 0.05). Different letters indicate a significant difference (p < 0.05).
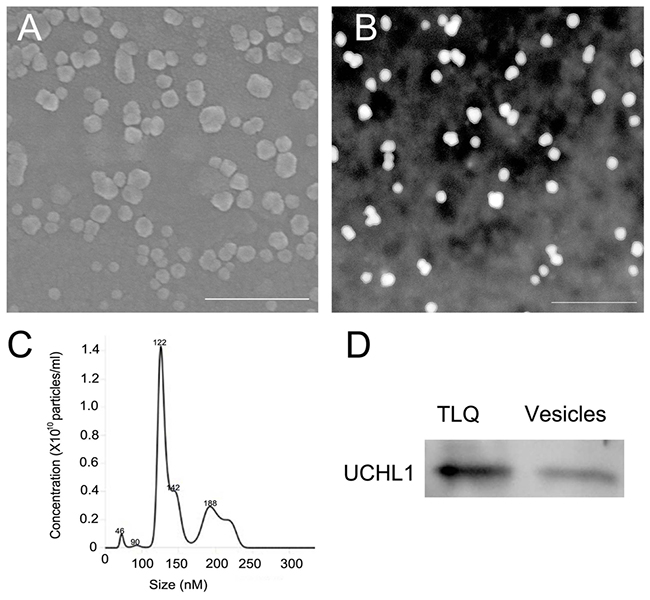
Figure 8: Detection of CGS UCHL1 in extracellular vesicles from the leaching liquid of the adult testes. (A) Detection of the vesicles by scanning electron microscopy. (B) Detection of the vesicles by transmission electron microscopy. (C) Size distribution and quantification of the extracellular vesicles analyzed by NanoSight NS300. (D) Detection of UCHL1 protein by western blot. TLQ represented the protein samples from tissue leaching liquid of adult testes; vesicles represent the protein samples from extracellular vesicles of the tissue leaching liquid. Scale bar: 500 nm.
DISCUSSION
UCHL1 (or PGP 9.5) is a conservative protein extensively presents in animals, including nematodes, Drosophila, fish, amphibians, birds, and mammals [3]. In this study, the sequence of CGS UCHL1 cDNA was obtained and the molecular characteristics were revealed by bioinformation analysis. CGS UCHL1 was most homologous with human UCHL1 (up to 73.99 %) among 10 different species compared. In the phylogenetic tree, CGS UCHL1 was clustered together with the UCHL1 of other higher vertebrates. This illustrated that the evolutional characterization of CGS UCHL1 is conservative. The homology of CGS UCHL1 and X. laevis UCHL1 was also very high (up to 70.04 %), whereas X. laevis UCHL1 formed a clade in the phylogenetic tree with CGS and other vertebrates (Figure 3). It was inferred that CGS UCHL1 could play a special function in CGSs.
UCHL1 was initially found to be expressed specifically in the human brain [33]. The mRNA for UCHL1/PGP 9.5 is known to be expressed extremely early in the differentiation of the nervous system [34]. Subsequent research identified UCHL1 was prominently present in the kidneys [35], ovaries [36], and testes [19, 21]. In mammals, UCHL1/PGP 9.5, expression in spermatogonia cells, was found to play an important role in spermatogenesis [9, 19]. In this study, we observed regular expression of UCHL1 during the gonad development of the Chinese giant salamander. The peak level of UCHL1 mRNA appeared in testes of 3-year-old CGSs, as well as the expression pattern of UCHL1 protein in testis tissues (Figure 4, 6A). Specially, the expression level of UCHL1 mRNA and protein in adult testes was lower during the breeding season than during the post-reproduction period (Figure 7A). However, the expression level of UCHL1 protein in the testicular leaking liquid of adult CGSs was higher during the breeding season than during the post-reproduction period (Figure 7B). The results suggested that UCHL1 played an important role in regulating the gonad development of CGSs and the testicular annual cycles.
Three localizations of UCHL1 in tissue have been previously reported [10, 11, 35]. The intracellular and extracellular localization of UCHL1 were also observed in testes during the developmental stages of CGSs. In the seminiferous lobules of CGS testes before sexual maturation, the UCHL1 was only localized on primordial germ cells and Sertoli cells (Figure 5). This proved that UCHL1 could play an important role in regulating the gonadal development of CGSs.
CGS UCHL1 should be thought of as a non-secretary protein because the typical signal peptide in the UCHL1 protein molecular was not found. Although our results showed that the UCHL1 was in the extracellular matrix and loaded in exosomes (Figure 8). The UCHL1 could be partly released into the extracellular matrix through the exosome pathway. In mammals, UCHL1 protein can be transferred by loading on exosomes [37, 38]. The UCHL1 loading on exosomes could have important biological functions in the male gonads of adult CGSs.
In fish and amphibians, there are cysts that could be related with homogeneous development of germ cells in seminiferous lobules after sexual maturity [39, 40]. The exosomes loading UCHL1 could be involved in forming the cysts (or niches) in which the homogeneous germ cells aggregated. The cysts separated germ cells at different stages from each other to ensure the synchronous maturation of sperm in fish and amphibians [41, 42]. The synchronous maturation of sperm allow for the ejection of a copious amount of sperm to fertilize eggs in vitro in water during the breeding season of amphibians and fish [43, 44]. It was inferred that the extracellular UCHL1 could construct a barrier to protect normal development of the germ cells by reversing the protein ubiquitination pathway. UCHL1 could also regulate spermatogenesis of adult CGS during the annual testicular cycle.
CONCLUSION
In conclusion, we have identified UCHL1 from CGS and investigated the expression level and localization pattern of UCHL1 in seminiferous lobules during development of testes. Moreover, during the annual cycle of testicular development in adult CGS, the extracellular UCHL1 in exosomes is an important component of cysts (or niches), which could be related to the synchronous development of germ cells. The present study enriched our knowledge of UCHL1 and provided valuable information for better understanding of the spermatogenesis characteristics of CGS. In the future, the mechanism of UCHL1 regulation in spermatogenesis will be further studied to reveal the reproductive characteristics of CGS.
MATERIALS AND METHODS
Collection of tissues samples of Chinese giant salamanders
Chinese giant salamanders were collected from eight farms in Hanzhong County (Shaanxi Province, China) in August and October, 2016. CGSs were anesthetized using 0.6 mg/L MS-222 and sacrificed by destroying the spinal cord of the CGS with a needle. These CGSs were the second or third generation of individuals that were permitted to be used in this study by the Wildlife Conservation Bureau of Shaanxi Province, China. The tissue samples were collected immediately after dissection. Some samples were fixed in Bouin’s liquid for section, and other aliquots were stored in liquid nitrogen for protein and RNA extraction. This study was approved by the Animal Care and Use Committee of Northwest A&F University in China. All protocols were carried out in accordance with the approved guidelines and regulations.
Total RNA extraction and cDNA synthesis
Total RNA was extracted using RNAiso Reagent following the manufacturer’s protocol. The concentration, purity, and integrity of RNA were detected by the NanoDrop 1000 Spectrophotometer or electrophoresis on 1 % agarose gel. Genomic DNA was removed from the total RNA using Rnase-free DNase before cDNA synthesis. All cDNA samples were synthesized from 1 mg total RNA with the PrimeScript RT Reagent Kit. The first-strand cDNA was synthesized by the PrimeScript RT Enzyme following the manufacturer’s instructions with random primers.
cDNA cloning of UCHL1
A partial cDNA sequence of UCHL1 was acquired from the transcriptome database for the Chinese giant salamander in our lab. The primer for amplifying UCHL1 cDNAs fragments were designed (UCHL1-F and UCHL1-R) and is shown in Table 1. PCR amplification was performed as follows: 1 cycle of 94°C for 5 min, 35 cycles, including denaturation, at 94°C for 40 s, annealing at 56°C for 40 s, and extension at 72°C for 1 min, followed by 1 cycle at 72°C for 10 min. The specific products were assessed by electrophoresis on 1 % agarose gel and cloned into the plasmid vector pMD19-T and then transformed into E. coli TOP10 for sequencing. The nucleotide sequences were determined for three sense and antisense strands of plasmid inserts for three independent clones.
Table 1: Primer pairs in the study
Primer |
Sequence (5′ to 3′) |
Nucleotide position (nt) |
Remark |
|---|---|---|---|
UCHL1-F |
CCAAAGAGCACGCAGACATGC |
179-299 |
UCHL1 fragment |
UCHL1-R |
TTAAGACTTGCTGAGCGCCAC |
844-864 |
|
AP |
GGCCACGCGTCGACTAGTACT18 |
3′ Race |
|
AAP |
GGCCACGCGTCGACTACGGGIIGGGIIGGGIIG |
5′Race |
|
AUAP |
GGCCACGCGTCGACTAGTAC |
3′, 5′Race |
|
5′RTP |
GGCATTCGTCCATCCAGCTC |
715-734 |
5′Race |
5′GSP1 |
GGCATTCGTCCATCCAGCTC |
715-734 |
5′Race |
5′GSP2 |
GTTCACTTTGTCATCCACCC |
653-672 |
5′Race |
5′GSP3 |
CTTTGTCATCCACCCGGCAG |
658-677 |
5′Race |
3′GSP1 |
CCCTCTCACTCCTCAGCATG |
354-373 |
3′Race |
3′GSP2 |
CTGTGCTGTTCTTCTGCTCT |
333-352 |
3′Race |
3′GSP3 |
GGATGTGCTTGAGTTCTCAG |
282-302 |
3′Race |
F-UCHL1 |
GTGGTTAGAGGTCCAAAGAG |
167-186 |
qPCR |
R-UCHL1 |
ATCCACGAAGTGCCAACCAG |
266-285 |
|
F-β-actin |
AAAAGCCAACCGAGAAAAG |
qPCR |
|
R-β-actin |
TACGACCAGAGGCATACAG |
To obtain the 5′ and 3′ ends of each cDNA, a number of gene-specific sense or antisense primers (Table 1) were designed for the three UCHL1 transcripts to replicate sequence regions for their core fragment sequenced above. 5′- and 3′-RACE were performed with the RACE cDNA Amplification Kit using ovarian or liver total RNA according to manufacturer’s protocols, respectively. Nested 5′- and 3′-RACE products of the expected size were sub-cloned and sequenced as described above.
Bioinformatic analysis of UCHL1
The complete ORF regions and amino acid sequences of UCHL1 were deduced using ORFfinder at NCBI (http://www.ncbi.nlm.nih.gov/gorf/gorf.html). The cleavage site for the signal peptide was predicted with SignalP (http://www.cbs.dtu.dk/services/SignalP/). O-linked and N-linked glycosylation sites were detected with the NetOGlyc Server (http://www.cbs.dtu.dk/services/NetOGlyc/) and with the NetNGlyc Server (http://www.cbs.dtu.dk/services/NetNGlyc/), respectively. To evaluate the evolutionary relationship of CGS UCHL1 with 13 other species, nucleic acid and amino acid sequences of the UCHL1 protein from different species were used to perform multiple alignments using ClustalX2 and a phylogenic tree was constructed by the neighbor-joining (NJ) method within MEGA version 5.0 [30]. GenBank accession numbers of the UCHL1 amino acid sequences included in the tree are: BC005117.1 (H. sapiens), NM_011670.2 (M. musculus), NM_001080212.1 (G. gallus), XM_019553393.1 (C. porosus), AY147851.1 (D. rerio), NM_001096657.2 (X. laevis), XM_007069383.1 (C. mydas), AB490372.1 (L. reevesii rubritaeniata), NM_057592.4 (D. melanogaster).
Developmental expression of UCHL1 mRNA in testes of CGSs at different stages
The expression patterns of UCHL1 in different developmental stages of gonads were detected by RT-qPCR. The qRT-PCR mixture consisted of 1 μL of the diluted cDNA samples (50 ng/ μL), 6 μL Power 2×SYBR Real-time PCR premixture (BioTeKe), 1 μL of each primer (0.4 μM), and 11 μL H2O. The qRT-PCR cycle profile included 1 cycle at 95°C for 5 min, then 32 cycles at 95°C for 15 s, 58°C for 15 s, and 72°C for 20 s. To determine the relative mRNA expression level of UCHL1, QRT-PCR was performed via the 2−ΔΔct method with the β-actin gene used as an internal control gene for cDNA normalization. The special primer pairs for UCHL1 (F-UCHL1, R-UCHL1) and β-actin (F-β-actin, R-β-actin) were designed and are listed in Table 1.
Histological analysis
For histological analysis, tissues from the normal CGSs were dissected and fixed in Bouin’s fixative overnight. After successive dehydration, tissues were imbedded in paraffin. Sections (5 μm) were mounted on slides with polylysine pretreatment, dewaxed in a xylene bath, and then rehydrated through gradient ethanol solutions. The sections were used for immunohistochemistry (IHC). IHC was performed for locating the UCHL1 in the tissues. After rinsing in PBS, heat-mediated antigen retrieval was performed in boiling citrate for 10 min. Slides were cooled for 40 min at room temperature and then rinsed three times with PBS before treating with 3 % H2O2. Nonspecific protein binding was blocked for 1 h by normal goat serum before incubating with the UCHL1 antibodies (1:200; Cat: ab916, Abcam) overnight in a damp box. Slides were washed (3 × 5 min) in PBS, followed by incubation for 1 h with biotinylated anti-mouse. Next biotinylated horseradish peroxidase was used for 1 h. Slides were stained by DAB according to the manufacturer’s instructions before counterstaining with hematoxylin. Images were obtained using a microscope. As controls, the primary antibodies (rabbit anti-UCHL1, Millipore, 1:200) were replaced with PBS.
Testes sample collection and protein extraction
The testes were harvested at different developmental stages. Firstly, the testes were split using small forceps on ice and 10 mg tissue fragments were washed using 200 μL PBS containing 1 % protease inhibitors. Subsequently, the samples were centrifuged at 800 g for 10 min at 4°C. The pellets and the supernatant fractions (tissue liquid) were collected for protein extraction. Thirdly, the pellets were lysed using RIPA buffer containing 1 % protease inhibitors. The lysates and the tissue liquid were cleared by centrifugation at 14,000 g for 20 min. Then the concentration of protein was analyzed using the BCA method.
Separation and identification of exosomes from the adult testes
Separation of exosomes
The extracellular vesicles (or exosomes) was separated from the adult testes. Firstly, the testes were split by small forceps on ice and 10 mg tissue fragments were washed using 200 μL PBS containing 1 % protease inhibitors. Subsequently, the samples were centrifuged at 800 g for 10 min at 4°C. The supernatant was centrifuged at 12,000 g for 20 min and the supernatant was filtered through 0.22-μm membranes. Subsequently, ultracentrifugation at 120,000 g for 2 h was performed to collect exosomes. Finally, exosomes pellets were resuspended in 200 μL cold PBS.
Transmission electron microscopy (TEM)
Fresh isolated exosomes were diluted in cold PBS through serial dilutions. After mounting on copper grids, the samples were fixed in 1 % glutaraldehyde for 10 min, washed in sterile distilled water, and then incubated with phosphotungstic acid for 1–2 min. Excess reagent was removed and samples were dried at room temperature. Finally, the HT7700 TEM was used to image the exosome samples at 80–100 kV.
Scanning electron microscopy (SEM)
The method was as previously described, with minor modifications [31]. Exosomes were fixed in 2.5 % glutaraldehyde for 1 h at 4°C. The samples were diluted in distilled water through serial dilutions, and then 5 μL were added to a cleaned silicon slice. After air drying under a ventilation hood, exosomes were post-fixed in 1 % osmic acid for 1 h and then dehydrated in increasing concentrations of ethyl alcohol (35, 50, 70, 80, 90, and 100 %) every 20 min. Next, isoamyl acetate was used to treat samples for 10 min. Finally, the specimens were sputter-coated with gold palladium and viewed under a scanning electron microscope.
Nanoparticle tracking analysis
Size distribution and quantification of the extracellular vesicles were analyzed using a NanoSight NS300 instrument as described previously [32]. Briefly, exosomes from testes were diluted in 1mL of PBS and disaggregated by using a syringe and needle. Then, the sample was injected into the chamber and three individual samples were tested.
Western blot analysis
Protein samples were boiled in 5× loading buffer for 10 min, and separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Subsequently, proteins were transferred to a polyvinylidene difluoride membrane. Then, the membranes were blocked with 5 % (w/v) skim milk and 0.05 % (v/v) Tween 20 in Tris buffered saline (TBS; 20 mmol/L Tris, 500 mmol/L NaCl; pH 7.0). The mouse anti-UCHL1 (1:200, Cat:ab916, Abcam) was used as the primary antibodies and incubated with the membrane at a dilution of 1:500 overnight at 4°C. The membranes were washed with TBS containing 0.05 % (v/v) Tween 20 and subsequently incubated with HRP-conjugated anti-mouse IgG antibody at a dilution of 1:1000. Finally, after the membrane was incubated with ECL for chemiluminescence developing by BIO-RAD ChemiDoc XRS. β-actin antibody was used to detect β-actin protein as an internal control in the examination.
Statistical analyses
All experiments were performed with at least three replicates. Data were expressed as mean ± standard deviation (S.D.) and were analyzed with an analyses of variance (ANOVA) and Duncan’s multiple comparisons. The statistical analyses were conducted with SPSS software. The significance level was defined as p < 0.05.
Abbreviations
UCHL1: Ubiquitin carboxyl-terminal hydrolase L1.
CDS: Coding sequences.
CGS: Chinese giant salamander.
ORF: Open reader form.
IHC: Immunohistochemistry.
PBS: Phosphate buffer saline.
DAB: Diaminobenzidine.
TEM: Transmission electron microscopy.
SEM: Scanning electron microscopy.
TBS: Tris buffered saline.
Author contributions
Y.W. and L.W. performed IHC, TEM, Western blot.
H.G. and Y.G. cloned the UCHL1 gene and bioinformatics analysis.
Y. M. and Y. G. prepared the samples of CGSs.
H. J. wrote manuscript and corrected the paper.
W. D. designed all of the experiments and wrote manuscript.
ACKNOWLEDGMENTS
We thank Prof. Wen-xian Zeng at College of Animal Science and Technology, Northwest A&F University for commenting on our manuscript.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
This work was supported by the Fund of the Agriculture Sci-Tech Project of Shaanxi Province (No.2014K01-20-01) and the National Natural Science Foundation China (NSFC) (No. C170104-31172205).
REFERENCES
1. Amerik AY, Hochstrasser M. Mechanism and function of deubiquitinating enzymes. Biochimica et biophysica acta. 2004; 1695:189-207.
2. Reyes-Turcu FE, Ventii KH, Wilkinson KD. Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annual review of biochemistry. 2009; 78:363-397.
3. Day IN, Thompson RJ. UCHL1 (PGP 9.5): neuronal biomarker and ubiquitin system protein. Progress in neurobiology. 2010; 90:327-362.
4. Esteban FJ, Jimenez A, Barroso JB, Pedrosa JA, del Moral ML, Rodrigo J, Peinado MA. The innervation of rainbow trout (Oncorhynchus mykiss) liver: protein gene product 9.5 and neuronal nitric oxide synthase immunoreactivities. J Anat. 1998; 193:241-249.
5. Saito S, Yamamoto Y, Mori M, Amano M, Yamanome T, Taniguchi K, Yamamori K. Variety in histochemical characteristics of the olfactory receptor cells in a flatfish, barfin flounder (Verasper moseri). J Vet Med Sci. 2004; 66:1409-1412.
6. Kobayashi K, Kumakura M, Yoshimura K, Inatomi M, Asami T. Fine structure of the tongue and lingual papillae of the penguin. Arch Histol Cytol. 1998; 61:37-46.
7. Yoshie S, Yokosuka H, Kanazawa H, Fujita T. The existence of Merkel cells in the lingual connective tissue of the Surinam caiman, Caiman crocodilus crocodilus (order Crocodilia). Arch Histol Cytol. 1999; 62:97-106.
8. Kusindarta DL, Wijayanto H, Atoji Y. Intrinsic innervation in the tracheal smooth muscle of the large flying fox (Pteropus vampyrus): an immunohistochemical study. European journal of morphology. 2003; 41:111-116.
9. Tokunaga Y, Imai S, Torii R, Maeda T. Cytoplasmic liberation of protein gene product 9.5 during the seasonal regulation of spermatogenesis in the monkey (Macaca fuscata). Endocrinology. 1999; 140:1875-1883.
10. Wilson PO, Barber PC, Hamid QA, Power BF, Dhillon AP, Rode J, Day IN, Thompson RJ, Polak JM. The immunolocalization of protein gene product 9.5 using rabbit polyclonal and mouse monoclonal antibodies. British journal of experimental pathology. 1988; 69:91-104.
11. Martin R, Santamaria L, Fraile B, Paniagua R, Polak JM. Ultrastructural localization of PGP 9.5 and ubiquitin immunoreactivities in rat ductus epididymidis epithelium. The Histochemical journal. 1995; 27:431-439.
12. Fraile B, Martin R, De Miguel MP, Arenas MI, Bethencourt FR, Peinado F, Paniagua R, Santamaria L. Light and electron microscopic immunohistochemical localization of protein gene product 9.5 and ubiquitin immunoreactivities in the human epididymis and vas deferens. Biol Reprod. 1996; 55:291-297.
13. Kwon J, Sekiguchi S, Wang YL, Setsuie R, Yoshikawa Y, Wada K. The region-specific functions of two ubiquitin C-terminal hydrolase isozymes along the epididymis. Exp Anim. 2006; 55:35-43.
14. Sheng K, Liang X, Huang S, Xu W. The role of histone ubiquitination during spermatogenesis. BioMed research international. 2014; 2014:870695.
15. Devi L, Pawar RM, Makala H, Goel S. Conserved expression of ubiquitin carboxyl-terminal esterase L1 (UCHL1) in mammalian testes. Indian journal of experimental biology. 2015; 53:305-312.
16. Kon Y, Endoh D, Iwanaga T. Expression of protein gene product 9.5, a neuronal ubiquitin C-terminal hydrolase, and its developing change in sertoli cells of mouse testis. Molecular reproduction and development. 1999; 54:333-341.
17. Fujihara M, Kim SM, Minami N, Yamada M, Imai H. Characterization and in vitro culture of male germ cells from developing bovine testis. The Journal of reproduction and development. 2011; 57:355-364.
18. Oatley JM, de Avila DM, Reeves JJ, McLean DJ. Testis tissue explant culture supports survival and proliferation of bovine spermatogonial stem cells. Biology of reproduction. 2004; 70:625-631.
19. Luo J, Megee S, Rathi R, Dobrinski I. Protein gene product 9.5 is a spermatogonia-specific marker in the pig testis: application to enrichment and culture of porcine spermatogonia. Molecular reproduction and development. 2006; 73:1531-1540.
20. Rodriguez-Sosa JR, Dobson H, Hahnel A. Isolation and transplantation of spermatogonia in sheep. Theriogenology. 2006; 66:2091-2103.
21. Heidari B, Rahmati-Ahmadabadi M, Akhondi MM, Zarnani AH, Jeddi-Tehrani M, Shirazi A, Naderi MM, Behzadi B. Isolation, identification, and culture of goat spermatogonial stem cells using c-kit and PGP9.5 markers. Journal of assisted reproduction and genetics. 2012; 29:1029-1038.
22. Sun J, Shang X, Tian Y, Zhao W, He Y, Chen K, Cheng H, Zhou R. Ubiquitin C-terminal hydrolase-L1 (Uch-L1) correlates with gonadal transformation in the rice field eel. The FEBS journal. 2008; 275:242-249.
23. Sun ZG, Kong WH, Zhang YJ, Yan S, Lu JN, Gu Z, Lin F, Tso JK. A novel ubiquitin carboxyl terminal hydrolase is involved in toad oocyte maturation. Cell research. 2002; 12:199-206.
24. Dong W, Zhang X, Yang C, An J, Qin J, Song F, Zeng W. Iridovirus infection in Chinese giant salamanders, China, 2010. Emerging infectious diseases. 2011; 17:2388-2389.
25. Du J, Wang L, Wang Y, Shen J, Pan C, Meng Y, Yang C, Ji H, Dong W. Autophagy and apoptosis induced by Chinese giant salamander (Andrias davidianus) iridovirus (CGSIV). Veterinary microbiology. 2016; 195:87-95.
26. Fortuny J, Marce-Nogue J, Heiss E, Sanchez M, Gil L, Galobart A. 3D bite modeling and feeding mechanics of the largest living amphibian, the Chinese giant salamander Andrias davidianus (Amphibia:Urodela). PloS one. 2015; 10:e0121885.
27. Huang Y, Gao XC, Xiong JL, Ren HT, Sun XH. Sequencing and de novo transcriptome assembly of the Chinese giant salamander (Andrias davidianus). Genomics data. 2017; 12:109-110.
28. Huang Y, Ren HT, Xiong JL, Gao XC, Sun XH. Identification and characterization of known and novel microRNAs in three tissues of Chinese giant salamander base on deep sequencing approach. Genomics. 2017; 109:258-264.
29. Sun J, Geng X, Guo J, Zang X, Li P, Li D, Xu C. Proteomic analysis of the skin from Chinese fire-bellied newt and comparison to Chinese giant salamander. Comparative biochemistry and physiology Part D, Genomics and proteomics. 2016; 19:71-77.
30. Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular biology and evolution. 2007; 24:1596-1599.
31. Du J, Shen J, Wang Y, Pan C, Pang W, Diao H, Dong W. Boar seminal plasma exosomes maintain sperm function by infiltrating into the sperm membrane. Oncotarget. 2016; 7:58832-58847. https://doi.org/10.18632/oncotarget.11315.
32. Mazzeo C, Canas JA, Zafra MP, Rojas Marco A, Fernandez-Nieto M, Sanz V, Mittelbrunn M, Izquierdo M, Baixaulli F, Sastre J, Del Pozo V. Exosome secretion by eosinophils: A possible role in asthma pathogenesis. The Journal of allergy and clinical immunology. 2015; 135:1603-1613.
33. Jackson P, Thompson RJ. The demonstration of new human brain-specific proteins by high-resolution two-dimensional polyacrylamide gel electrophoresis. Journal of the neurological sciences. 1981; 49:429-438.
34. Bilguvar K, Tyagi NK, Ozkara C, Tuysuz B, Bakircioglu M, Choi M, Delil S, Caglayan AO, Baranoski JF, Erturk O, Yalcinkaya C, Karacorlu M, Dincer A, et al. Recessive loss of function of the neuronal ubiquitin hydrolase UCHL1 leads to early-onset progressive neurodegeneration. Proceedings of the National Academy of Sciences of the United States of America. 2013; 110:3489-3494.
35. Shirato I, Asanuma K, Takeda Y, Hayashi K, Tomino Y. Protein gene product 9.5 is selectively localized in parietal epithelial cells of Bowman’s capsule in the rat kidney. Journal of the American Society of Nephrology. 2000; 11:2381-2386.
36. Sekiguchi S, Kwon J, Yoshida E, Hamasaki H, Ichinose S, Hideshima M, Kuraoka M, Takahashi A, Ishii Y, Kyuwa S, Wada K, Yoshikawa Y. Localization of ubiquitin C-terminal hydrolase L1 in mouse ova and its function in the plasma membrane to block polyspermy. The American journal of pathology. 2006; 169:1722-1729.
37. Ning K, Wang T, Sun X, Zhang P, Chen Y, Jin J, Hua D. UCH-L1-containing exosomes mediate chemotherapeutic resistance transfer in breast cancer. Journal of surgical oncology. 2017; 115:932-940.
38. Kumar D, Manek R, Raghavan V, Wang KK. Protein Characterization of Extracellular Microvesicles/Exosomes Released from Cytotoxin-Challenged Rat Cerebrocortical Mixed Culture and Mouse N2a Cells. Molecular neurobiology. 2017. https://doi.org/10.1007/s12035-017-0474-x.
39. Uribe MC, Mejia-Roa V. Testicular structure and germ cells morphology in salamanders. Spermatogenesis. 2014; 4:e988090.
40. Uribe MC, Grier HJ, Mejia-Roa V. Comparative testicular structure and spermatogenesis in bony fishes. Spermatogenesis. 2014; 4:e983400.
41. Almeida FF, Kristoffersen C, Taranger GL, Schulz RW. Spermatogenesis in Atlantic cod (Gadus morhua): a novel model of cystic germ cell development. Biology of reproduction. 2008; 78:27-34.
42. Yoshida S. From cyst to tubule. innovations in vertebrate spermatogenesis. Wiley interdisciplinary reviews Developmental biology. 2016; 5:119-131.
43. Kihlstrom JE, Lakomaa E, Hall H. A probably nongonadotropic sperm-releasing activity in the pituitary gland from mammals, amphibians, and fishes. General and comparative endocrinology. 1971; 17:573-575.
44. Sasso-Cerri E, Freymuller E, Miraglia SM. Testosterone-immunopositive primordial germ cells in the testis of the bullfrog, Rana catesbeiana. Journal of anatomy. 2005; 206:519-523.

