INTRODUCTION
Numerous studies reported expression levels of tissue [1–194] or blood [195–221] miRNAs were related with prognosis of HCC patients. HCC is one of the most common tumors, over 700,000 new cases are reported yearly, and HCC is considered as the third primary etiology of tumor-associated mortality rate globally [222–224]. In spite of enormous process in diagnosis and comprehensive therapy over the last few decades, HCC patients still have poor prognosis, primarily due to its high rate of recurrence [225] and metastasis [226].
miRNAs, a cluster of endogenous short non-coding single strand RNAs, serve as significant post-transcriptional regulatory factor of genetic expression via interacting with the 3′-UTR of the targeted mRNAs [227]. Conspicuously, due to widespread RNAase in the blood environment, circulating miRNAs displayed predominant stability. As a noninvasive detection method, circulating miRNA (blood) demonstrated more potential value as diagnostic and prognostic biomarkers than tissue miRNAs. Studies [228, 229] conducted in preclinical models and cancer patients proved that malignant tumor influences expression levels of miRNAs in the blood and that certain serum miRNAs are correlated with particular cancers. Though the way requires more validation, the finding possibly discloses the avenue to a creative method of detecting cancers via measurement of serum or plasma miRNAs.
Thus far, substantial investigations have discovered that miRNAs are involved and play a crucial role in the carcinogenesis of HCC [230, 231] while some miRNAs are up-regulated and others down-regulated in HCC. For example, Wong et al. [232] gained contrasting results that identifiable difference in miRNA expression pattern could not be discovered between primary HCC and venous metastases. However, comparing venous metastases to primary HCC, a prominent universal decrease of miRNA expression levels was assayed. Their results indicated that miRNA abnormality relatively early occured in liver carcinogenesis and the later universally decreased miRNA aggravated the preexisting miRNA abnormity to further accelerate HCC metastasis.
Nevertheless, there has not been a synthetic meta-analysis to assess precise prognostic value of miRNAs in HCC. As a consequence, it is of vital significance to develop a meta-analysis with an aim to evaluate it.
RESULTS
Study selection
Figure 1 showed a flow chart with details about the study selection process.
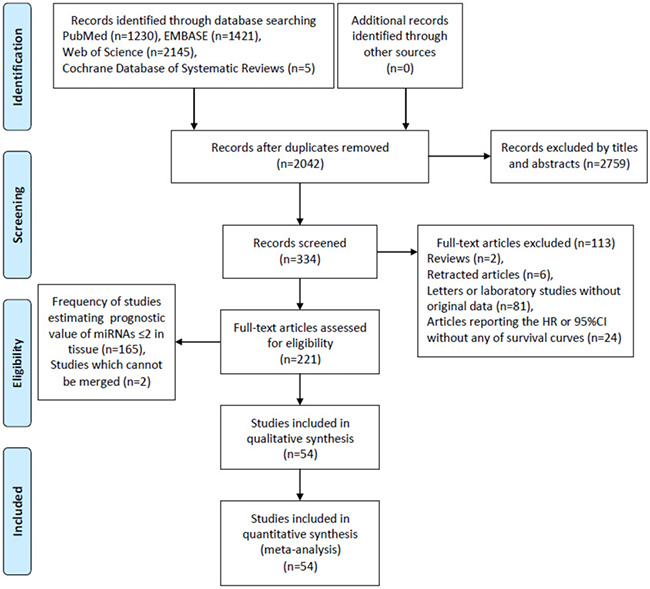
Figure 1: Flow diagram of literature search and selection.
Study frequency
Tables 1 (tissue) and 2 (blood) showed the frequency of researches evaluating prognostic value of miRNAs, including miRNA name, number of investigations assessing prognostic value, and reference.
Table 1: Frequency of studies estimating prognostic value of tissue miRNA expression in hepatocellular carcinoma
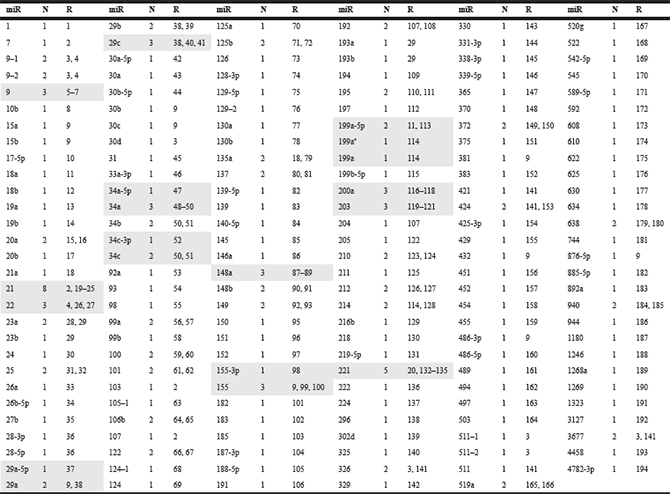
Highlighted studies were included in the present meta-analysis; N: Number of studies estimating prognostic value; R: References.
Table 2: Frequency of studies estimating prognostic value of blood miRNA expression in hepatocellular carcinoma
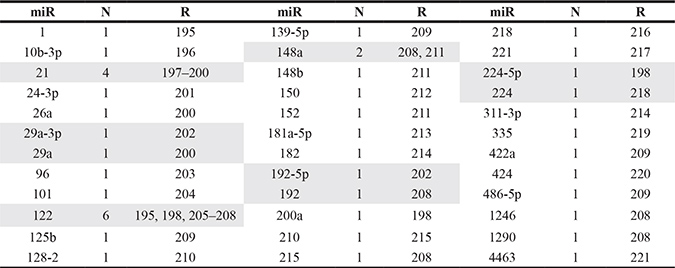
Highlighted studies were included in the present meta-analysis; N: Number of studies estimating prognostic value; R: References.
Study characteristics
Supplementary Table 1 comprehensively presented the characteristics and details (names of miRNAs, information about the included articles, detected samples, sample size, stage, cut-off value, detection methods, follow-up, survival outcome with HR and 95% CI) of studies with Kaplan-Meier survival curves (K-M curves) in HCC. If the survival outcome was not furnished directly and merely as K-M curves, we used the software Engauge Digitizer version 4.1 [233] to extract the data from K-M curves. Additionally, if both the univariate and multivariate outcomes were covered, we just chose the latter in that the confounding factors were corrected.
Meta-analysis
Table 3 presented a summary of the HR estimated from pooled analysis for the included miRNAs. A total of 16 miRNAs were screened by our present study.
Table 3: Summary of the HR for miRNA expression in hepatocellular carcinoma
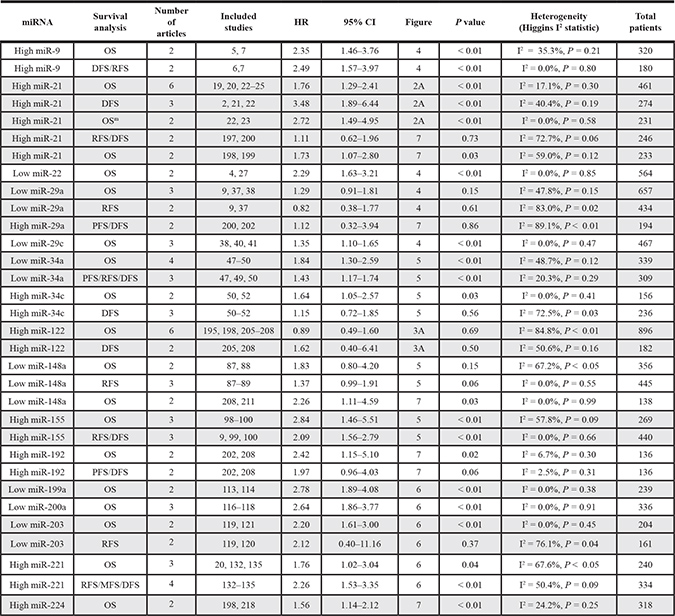
HR: hazard ratios; CI: confidence intervals; OS: overall survival; DFS: disease-free survival; RFS: recurrence-free survival; PFS: progression-free survival; MFS: metastasis-free survival; mMultivariate analysis.
Significantly prognostic value of high tissue miR-21 expression in OS
Six studies [19, 20, 22–25] focused on the correlation between high tissue miR-21 level and OS, suggesting that HCC patients with high tissue miR-21 level demonstrated a significantly worse OS than those with low tissue miR-21 level (HR = 1.76, 95% CI = 1.29–2.41, P < 0.01, Figure 2A).
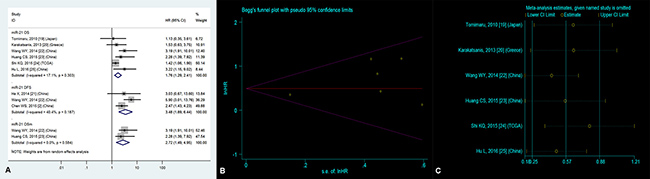
Figure 2: (A) Forest plot of the analyses about high expression of tissue miR-21 and OS, DFS or OS (multivariate analysis); (B) Publication bias of the analysis about high expression of tissue miR-21 and OS and (C) Sensitivity analysis of the study about high expression of tissue miR-21 and OS.
Publication bias
For the purpose of evaluating publication bias on OS of HCC patients with high tissue miR-21 level, we employed the Begg’s funnel plot (Figure 2B). Accordingly, the P value was 0.21, suggesting nonexistent publication bias.
Sensitivity analysis
The sensitivity analysis did not manifest variances among the outcomes in terms of the exclusion of any single research (Figure 2C) in the estimation on OS of HCC patients with high tissue miR-21 level, indicating that no individual investigation significantly affected the merged HR with 95% CI.
No significantly prognostic value of high blood miR-122 expression in OS
Six researches [1, 4, 11–14] concentrated on the relationship between high blood miR-122 level and OS, manifesting that there was no significant correlation between high blood miR-122 level and OS (HR = 0.89, 95% CI = 0.49–1.60, P = 0.69, Figure 3A).
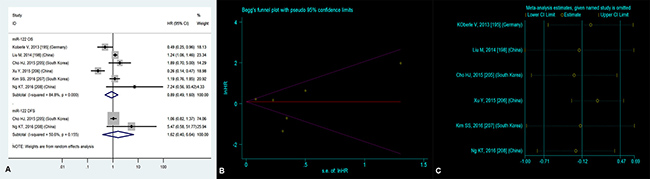
Figure 3: (A) Forest plot of the analyses about high expression of blood miR-122 and OS or DFS; (B) Publication bias of the analysis about high expression of blood miR-122 and OS and (C) Sensitivity analysis of the study about high expression of blood miR-122 and OS.
Publication bias
For the sake of estimating publication bias on OS of HCC patients with high blood miR-122 level, we employed the Begg’s funnel plot (Figure 3B). Consequently, the P value was 0.56, suggesting nonexistent publication bias.
Sensitivity analysis
The sensitivity analysis did not manifest variances among the outcomes in terms of the exclusion of any single research (Figure 3C) in the estimation on OS of HCC patients with high blood miR-122 level, indicating that no individual investigation significantly affected the merged HR with 95% CI.
Meta-regression
We employed the meta-regression to seek source of heterogeneity (I2 = 84.8%) on OS of HCC patients with high blood miR-122 level. The details were shown in Table 4, and source of heterogeneity was significantly caused by maximum months of follow-up (P = 0.01).
Table 4: Results of meta-regression on OS of blood miR-122 expression in hepatocellular carcinoma
Variables |
Details |
tau2 |
I2(%) |
Adj R2(%) |
P value |
|---|---|---|---|---|---|
Year |
2013–2016 |
0.73 |
87.82 |
–29.29 |
0.46 |
Country |
Germany, China, South Korea |
0.50 |
86.34 |
11.27 |
0.34 |
Design |
Prospective, Retrospective |
0.61 |
88.00 |
–21.49 |
0.51 |
Sample |
Serum, Plasma |
0.45 |
87.43 |
20.85 |
0.16 |
Number |
295, 136, 120, 122, 161, 62 |
0.69 |
86.03 |
–21.90 |
0.47 |
Stage |
None, I–IV |
0.52 |
87.59 |
8.14 |
0.39 |
Method |
qRT-PCR, RT-qPCR |
0.51 |
87.01 |
10.36 |
0.22 |
Follow-up |
26, 48, 96, 40, 79, 125 |
0.28 |
84.28 |
50.82 |
0.08 |
Follow-up |
< 48, ≥ 48 |
0.00 |
9.67 |
100.00 |
0.01 |
Tissue miR-9, miR-22, miR-29c, miR-34a, miR-34c, miR-155, miR-199a, miR-200a, miR-203, miR-221 and blood miR-21, miR-148a, miR-192, miR-224 have significantly prognostic values in OS
Table 3 and Figures 4–7 showed the details.
Tissue miR-29a, miR-148a and blood miR-29a do not have significantly prognostic values in OS
Table 3 and Figures 4, 5 and 7 showed the details.
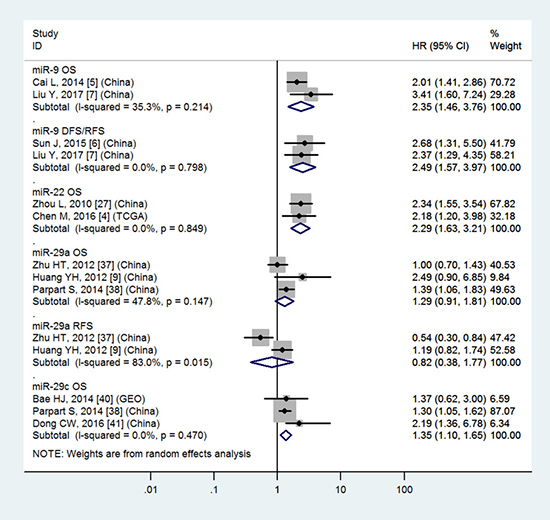
Figure 4: Forest plot of the analyses about high expression of tissue miR-9 or low expression of tissue miR-22, 29a, 29c and OS, DFS/RFS or RFS.
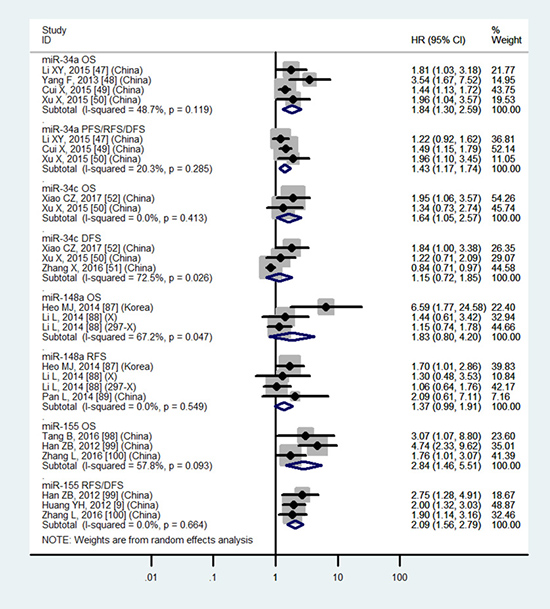
Figure 5: Forest plot of the analyses about high expression of tissue miR-34c, 155 or low expression of tissue miR-34a, 148a and OS, PFS/RFS/DFS, DFS, RFS or RFS/DFS.
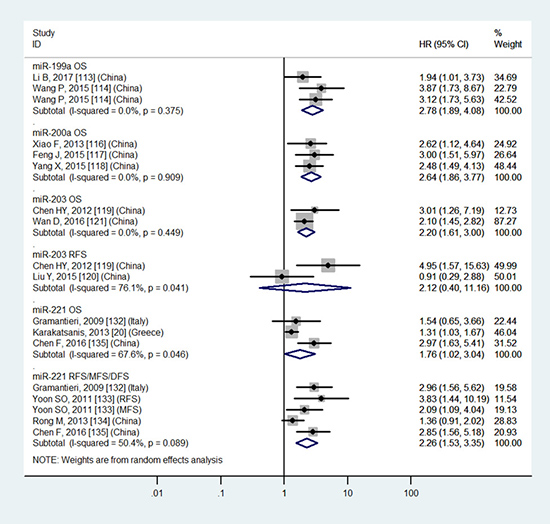
Figure 6: Forest plot of the analyses about high expression of tissue miR-221 or low expression of tissue miR-199a, 200a, 203 and OS, RFS or RFS/MFS/DFS.
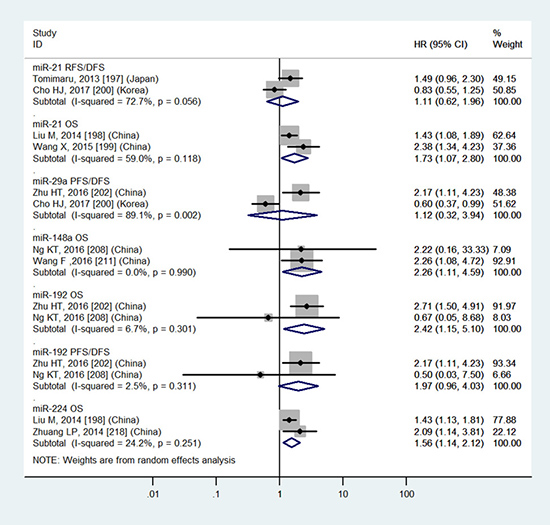
Figure 7: Forest plot of the analyses about high expression of blood miR-21, 29a, 192, 224 or low expression of blood miR-148a and RFS/DFS, OS or PFS/DFS.
DISCUSSION
Status quo
Numerous articles reported that dysregulated expression levels of miRNAs correlated with survival time of HCC patients [1–221]. Nevertheless, there has not been a comprehensive meta-analysis to assess the accurate prognostic value of miRNAs in HCC. Therefore, it was conducted to estimate the relationship between dysregulated miRNA level and survival time of HCC patients.
Main discoveries
For HCC patients, tissue miR-9, miR-21, miR-22, miR-29c, miR-34a, miR-34c, miR-155, miR-199a, miR-200a, miR-203, miR-221 and blood miR-21, miR-148a, miR-192, miR-224 demonstrate significantly prognostic value (P < 0.05). In the light of our reference standard, tissue miR-9, miR-22, miR-155, miR-199a, miR-200a, miR-203 and blood miR-148a, miR-192 potential prognostic candidates for predicting the OS of HCC patients (HR ≥ 2).
Molecular mechanisms for included miRNAs
For included miRNAs in the current study, a summary of miRNAs with changed levels, their possible targets and pathways enrolled in the present study has been presented in Table 5. From the data of the table, these potential targets and pathways may be involved with survival outcome of HCC patients.
Table 5: Summary of miRNAs with altered expression, their potential targets and pathways entered this study
miRNA |
Reference |
Expression |
Potential target |
Pathway |
|---|---|---|---|---|
9 |
5–7 |
Up |
GALNT4 |
None |
21 |
2, 19–25, 197–200 |
Up |
None |
None |
22 |
4, 26, 27 |
Down |
YWHAZ, HDAC4 |
Cell invasion, migration, proliferation, tumourigenicity and YWHAZ/AKT1/foxo3a signaling |
29a |
9, 37, 38, 200, 202 |
Up |
None |
None |
29c |
38, 40, 41 |
Down |
SIRT1 |
None |
34a |
47–50 |
Down |
FOXM1, MYC, BCL2, AXL |
Cell apoptosis, chemoresistance, proliferation, viability and FOXM1/MYC signaling |
34c |
50–52 |
Down |
NCKAP1 |
Cell cycle, growth, invasion and proliferation |
122 |
195, 198, 205–208 |
None |
None |
None |
148a |
87-89, 208, 211 |
Down |
USP4, SIP1 |
Cell invasion, migration and proliferation |
155 |
9, 98–100 |
Up |
ARID2, FBXW7 |
Cell apoptosis, cycle, invasion, proliferation and tumorigenesis |
192 |
202, 208 |
Up |
None |
None |
199a |
11, 113, 114 |
Down |
HIF1A, VEGFA, IGF1, IGF2 |
Cell growth, invasion, proliferation and Warburg effect |
200a |
116-118 |
Down |
MACC1, CDK6, ZEB2 |
Cell cycle, growth, metastasis and proliferation |
203 |
119–121 |
Down |
ADAM9, HULC |
Cell apoptosis, invasion and proliferation |
221 |
20, 132–135 |
Up |
Bmf |
Cell apoptosis and growth |
224 |
198, 218 |
None |
None |
None |
GALNT4: polypeptide N-acetylgalacosaminyltransferase 4; YWHAZ: tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein zeta; HDAC4: histone deacetylase 4; AKT1: AKT serine/threonine kinase 1; foxo3a: forkhead box O3A; SIRT1: sirtuin 1; FOXM1: forkhead box M1; MYC: MYC proto-oncogene, bHLH transcription factor; BCL2: BCL2, apoptosis regulator; AXL: AXL receptor tyrosine kinase; NCKAP1: NCK associated protein 1; USP4: ubiquitin specific peptidase 4; SIP1: Sip1p; ARID2: AT-rich interaction domain 2; FBXW7: F-box and WD repeat domain containing 7; HIF1A: hypoxia inducible factor 1 alpha subunit; VEGFA: vascular endothelial growth factor A; IGF1: insulin like growth factor 1; IGF2: insulin like growth factor 2; MACC1: MACC1, MET transcriptional regulator; CDK6: cyclin dependent kinase 6; ZEB2: zinc finger E-box binding homeobox 2; ADAM9: ADAM metallopeptidase domain 9; HULC: hepatocellular carcinoma up-regulated long non-coding RNA; BMF: Bcl2 modifying factor.
Strengths of the meta-analysis
There are a few strengths in this study, which are as follows: (1) nearly all articles estimating associations between miRNA level and survival result of HCC patients are shown in the current meta-analysis; (2) the number about HCC patients of all researches included in this study are more than or equal to 30, which makes the meta-analyses more convincing; (3) the Begg’s funnel plot and sensitivity analysis were used for miR-21 and miR-122, which excluded publication bias and excessive influence of individual study; (4) we employed meta-regression to seek source of heterogeneity, which indicated that months of follow-up were significantly associated with it; (5) studies merely proposing HR or 95% CI without K-M curves were excluded.
Limitations
Simultaneously, there are also limitations for the current work: (1) only English articles were included by us, which possibly excluded some studies written in other languages; (2) not all the articles assessing associations between miRNA level and survival time were included in the present study, which might neglect some potential miRNAs; (3) a few variables emerged among the included investigations, including different kinds of samples from HCC patients at different stages, cut-off values and detection methods, and only random-effects models were employed for all meta-analyses; (4) although overall studies included 54 relevant articles and 6464 patients in the present study, the number of articles and patients may be not enough for 16 miRNAs focused on.
Implications for future clinical and scientific research
With expression profiles shown in Tables 1 and 2, we can conveniently find relevant article about a single miRNA. Thus, the present study tendency for miRNAs in HCC can be easily obtained by basic researchers. Meanwhile, combined detection of multi-miRNAs can greatly increase the predict level for HCC patients. Besides, for clinical doctors, combined use of tissue and blood from HCC patients can bring about synergistic effect to estimation of prognosis.
MATERIALS AND METHODS
Search strategy, inclusion criteria and exclusion criteria
The details were presented in Table 6. Two authors (Yue Zhang and Chao Wei) independently performed this comprehensive online search.
Table 6: Information of search methods and criteria of inclusion and exclusion
Methods |
Information |
|---|---|
Search strategy |
4 search engines, including PubMed, EMBASE, Web of Science and Cochrane Database of Systematic Reviews |
Search deadline |
15 April, 2017 |
Search terms |
mir and hepatocellular carcinoma |
Inclusion criteria |
(1) Patients with hepatocellular carcinoma; |
(2) Expression of miRNAs and survival outcome in tissue, plasma or serum were measured; |
|
(3) At least, one of survival curves about overall survival (OS), cause-specific survival (CSS), disease-free survival (DFS), recurrence-free survival (RFS), progression-free survival (PFS) and metastasis-free survival (MFS) was measured, with or without the HR or 95% CI; |
|
(4) Full text articles published in English |
|
Exclusion criteria |
(1) Reviews, letters or laboratory studies without original data and retracted articles; |
(2) Frequency of studies estimating prognostic value of miRNAs ≤ 2 in tissue; |
|
(3) Studies which cannot be merged; |
|
(4) If more than one article had been published on the identical study cohort, only the most comprehensive study was selected for the present meta-analysis |
Quality assessment
Yue Zhang and Chao Wei confirmed all eligible investigations that analyzed the prognostic value of miRNAs in HCC, and Yue-Hua Jiang reassessed uncertain data.
Statistical analysis
All analyses were conducted using Stata version 13.0 (StataCorp, College Station, Texas, USA). The relative effect sizes for HR were characterized as moderate (protective [0.51–0.75] or contributory [1.35–1.99]) and large (≤ 0.50 or ≥ 2). The HR was considered significant at the P < 0.05 level if the 95% CI did not include the value 1. If the P values from OS and other survival results about corresponding miRNAs were inconsistent, the HR from OS was considered to the main reference standard. Because different types of samples (tissue, plasma and serum) from HCC patients at different disease stages, cut-off values and miRNA methods were used in individual studies, random-effects models (DerSimonian-Laird method) were more appropriate than fixed-models (Mantel-Haenszel method) for most of the analyses. Consequently, the random-effects models were used in the current meta-analysis. Source of heterogeneity was explored through meta-regression. Publication bias was estimated using the Begg’s funnel plot. A two-tailed P value < 0.05 was considered significant. Sensitivity analysis (influence analysis) was carried out to test how powerful the combined effect size was to removal of individual investigation. If the point assessment was out of the 95% CI of the pooled effect size after it was removed from the analysis, an individual study was doubted to have excessive influence.
CONCLUSIONS
In conclusion, tissue miR-9, miR-21, miR-22, miR-29c, miR-34a, miR-34c, miR-155, miR-199a, miR-200a, miR-203, miR-221 and blood miR-21, miR-148a, miR-192, miR-224 demonstrate significantly prognostic value. Among them, tissue miR-9, miR-22, miR-155, miR-199a, miR-200a, miR-203 and blood miR-148a, miR-192 are potential prognostic candidates for predicting OS in HCC.
Author contributions
Study concept and design: Yue Zhang (e-mail: [email protected]) and Yue-Hua Jiang.
Acquisition of data: Yue Zhang and Chao Wei.
Analysis and interpretation of data: Yue Zhang, Chao Wei and Cong-Cong Guo.
Drafting of the manuscript: Yue Zhang.
Revision of manuscript: Yue Zhang, Chao Wei, Cong-Cong Guo, Rong-Xiu Bi, Jin Xie, Dong-Hui Guan, Chuan-Hua Yang and Yue-Hua Jiang.
Supervision of work: Rong-Xiu Bi, Jin Xie, Dong-Hui Guan, Chuan-Hua Yang and Yue-Hua Jiang.
All authors read and approved the final manuscript.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
FUNDING
This work was supported by the National Natural Science Foundation of China (No. 81673807).
Role of funding source: The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
REFERENCES
1. Wang X, Huang Y, Zhuang H, Qian Y, Zhao Q, Yang L, Gu H, Chen J, Guo R, Liu Y. Downregulation of MicroRNA-1 is Associated with Poor Prognosis in Hepatocellular Carcinoma. Clin Lab. 2015; 61:1331–1336.
2. Chen WS, Yen CJ, Chen YJ, Chen JY, Wang LY, Chiu SJ, Shih WL, Ho CY, Wei TT, Pan HL, Chien PH, Hung MC, Chen CC, et al. miRNA-7/21/107 contribute to HBx-induced hepatocellular carcinoma progression through suppression of maspin. Oncotarget. 2015; 6:25962–25974. https://doi.org/10.18632/oncotarget.4504.
3. Zhang J, Chong CC, Chen GG, Lai PB. A Seven-microRNA Expression Signature Predicts Survival in Hepatocellular Carcinoma. PLoS One. 2015; 10:e0128628.
4. Chen M, Hu W, Xiong CL, Qu Z, Yin CQ, Wang YH, Luo CL, Guan Q, Yuan CH, Wang FB. miR-22 targets YWHAZ to inhibit metastasis of hepatocellular carcinoma and its down-regulation predicts a poor survival. Oncotarget. 2016; 7:80751–80764. https://doi.org/10.18632/oncotarget.13037.
5. Cai L, Cai X. Up-regulation of miR-9 expression predicate advanced clinicopathological features and poor prognosis in patients with hepatocellular carcinoma. Diagn Pathol. 2014;9:1000.
6. Sun J, Fang K, Shen H, Qian Y. MicroRNA-9 is a ponderable index for the prognosis of human hepatocellular carcinoma. Int J Clin Exp Med. 2015; 8:17748–17756.
7. Liu Y, Liu H, Yang L, Wu Q, Liu W, Fu Q, Zhang W, Zhang H, Xu J, Gu J. Loss of N-Acetylgalactosaminyltransferase-4 Orchestrates Oncogenic MicroRNA-9 in Hepatocellular Carcinoma. J Biol Chem. 2017; 292:3186–3200.
8. Li QJ, Zhou L, Yang F, Wang GX, Zheng H, Wang DS, He Y, Dou KF. MicroRNA-10b promotes migration and invasion through CADM1 in human hepatocellular carcinoma cells. Tumour Biol. 2012;33:1455–1465.
9. Huang YH, Lin KH, Chen HC, Chang ML, Hsu CW, Lai MW, Chen TC, Lee WC, Tseng YH, Yeh CT. Identification of postoperative prognostic microRNA predictors in hepatocellular carcinoma. PLoS One. 2012; 7:e37188.
10. Chen L, Jiang M, Yuan W, Tang H. miR-17-5p as a novel prognostic marker for hepatocellular carcinoma. J Invest Surg. 2012; 25:156–161.
11. Morita K, Shirabe K, Taketomi A, Soejima Y, Yoshizumi T, Uchiyama H, Ikegami T, Yamashita Y, Sugimachi K, Harimoto N, Itoh S, Ikeda T, Maehara Y. Relevance of microRNA-18a and microRNA-199a-5p to hepatocellular carcinoma recurrence after living donor liver transplantation. Liver Transpl. 2016; 22:665–676.
12. Murakami Y, Tamori A, Itami S, Tanahashi T, Toyoda H, Tanaka M, Wu W, Brojigin N, Kaneoka Y, Maeda A, Kumada T, Kawada N, Kubo S, et al. The expression level of miR-18b in hepatocellular carcinoma is associated with the grade of malignancy and prognosis. BMC Cancer. 2013;13:99.
13. Zhang Y, Guo X, Li Z, Li B, Li Z, Li R, Guo Q, Xiong L, Yu L, Zhao J, Lin N. A systematic investigation based on microRNA-mediated gene regulatory network reveals that dysregulation of microRNA-19a/Cyclin D1 axis confers an oncogenic potential and a worse prognosis in human hepatocellular carcinoma. RNA Biol. 2015; 12:643–57.
14. Hung CL, Yen CS, Tsai HW, Su YC, Yen CJ. Upregulation of MicroRNA-19b predicts good prognosis in patients with hepatocellular carcinoma presenting with vascular invasion or multifocal disease. BMC Cancer. 2015; 15:665.
15. Fan MQ, Huang CB, Gu Y, Xiao Y, Sheng JX, Zhong L. Decrease expression of microRNA-20a promotes cancer cell proliferation and predicts poor survival of hepatocellular carcinoma. J Exp Clin Cancer Res. 2013; 32:21.
16. Ma CL, Qiao S, Wang XF, Sun RJ, Zhang X, Li YC, Liu JG. High expression of miR-20a predicated poor prognosis of hepatocellular carcinoma. Int J Clin Exp Med. 2016; 9:3699–3704.
17. Xue TM, Tao LD, Zhang M, Zhang J, Liu X, Chen GF, Zhu YJ, Zhang PJ. Clinicopathological Significance of MicroRNA-20b Expression in Hepatocellular Carcinoma and Regulation of HIF-1 alpha and VEGF Effect on Cell Biological Behaviour. Dis Markers. 2015; 2015:325176.
18. von Felden J, Heim D, Schulze K, Krech T, Ewald F, Nashan B, Lohse AW, Wege H. High expression of micro RNA-135A in hepatocellular carcinoma is associated with recurrence within 12 months after resection. BMC Cancer. 2017;17:60.
19. Tomimaru Y, Eguchi H, Nagano H, Wada H, Tomokuni A, Kobayashi S, Marubashi S, Takeda Y, Tanemura M, Umeshita K, Doki Y, Mori M. MicroRNA-21 induces resistance to the anti-tumour effect of interferon-α/5-fluorouracil in hepatocellular carcinoma cells. Br J Cancer. 2010; 103:1617–1626.
20. Karakatsanis A, Papaconstantinou I, Gazouli M, Lyberopoulou A, Polymeneas G, Voros D. Expression of microRNAs, miR-21, miR-31, miR-122, miR-145, miR-146a, miR-200c, miR-221, miR-222, and miR-223 in patients with hepatocellular carcinoma or intrahepatic cholangiocarcinoma and its prognostic significance. Mol Carcinog. 2013; 52:297–303.
21. He X, Li J, Guo W, Liu W, Yu J, Song W, Dong L, Wang F, Yu S, Zheng Y, Chen S, Kong Y, Liu C. Targeting the microRNA-21/AP1 axis by 5-fluorouracil and pirarubicin in human hepatocellular carcinoma. Oncotarget. 2015; 6:2302–2314. https://doi.org/10.18632/oncotarget.2955.
22. Wang WY, Zhang HF, Wang L, Ma YP, Gao F, Zhang SJ, Wang LC. miR-21 expression predicts prognosis in hepatocellular carcinoma. Clin Res Hepatol Gastroenterol. 2014; 38:715–719.
23. Huang CS, Yu W, Cui H, Wang YJ, Zhang L, Han F, Huang T. Increased expression of miR-21 predicts poor prognosis in patients with hepatocellular carcinoma. Int J Clin Exp Pathol. 2015; 8:7234–7138.
24. Shi KQ, Lin Z, Chen XJ, Song M, Wang YQ, Cai YJ, Yang NB, Zheng MH, Dong JZ, Zhang L, Chen YP. Hepatocellular carcinoma associated microRNA expression signature: integrated bioinformatics analysis, experimental validation and clinical significance. Oncotarget. 2015; 6:25093–25108. https://doi.org/10.18632/oncotarget.4437.
25. Hu L, Ye H, Huang G, Luo F, Liu Y, Liu Y, Yang X, Shen J, Liu Q, Zhang J. Long noncoding RNA GAS5 suppresses the migration and invasion of hepatocellular carcinoma cells via miR-21. Tumour Biol. 2016; 37:2691–2702.
26. Zhang J, Yang Y, Yang T, Liu Y, Li A, Fu S, Wu M, Pan Z, Zhou W. microRNA-22, downregulated in hepatocellular carcinoma and correlated with prognosis, suppresses cell proliferation and tumourigenicity. Br J Cancer. 2010; 103:1215–1220.
27. Zhou L, He J, Zhang Y. MicroRNA-22 expression in hepatocellular carcinoma and its correlation with ezrin protein. J Int Med Res. 2013; 41:1009–1016.
28. Bao L, Zhao J, Dai X, Wang Y, Ma R, Su Y, Cui H, Niu J, Bai S, Xiao Z, Yuan H, Yang Z, Li C, et al. Correlation between miR-23a and onset of hepatocellular carcinoma. Clin Res Hepatol Gastroenterol. 2014; 38:318–330.
29. Grossi I, Arici B, Portolani N, De Petro G, Salvi A. Clinical and biological significance of miR-23b and miR-193a in human hepatocellular carcinoma. Oncotarget. 2017; 8:6955–6969. https://doi.org/10.18632/oncotarget.14332.
30. Liu YX, Long XD, Xi ZF, Ma Y, Huang XY, Yao JG, Wang C, Xing TY, Xia Q. MicroRNA-24 modulates aflatoxin B1-related hepatocellular carcinoma prognosis and tumorigenesis. Biomed Res Int. 2014;2014:482926.
31. Su ZX, Zhao J, Rong ZH, Geng WM, Wu YG, Qin CK. Upregulation of microRNA-25 associates with prognosis in hepatocellular carcinoma. Diagn Pathol. 2014; 9:47.
32. Liu B, Yang J, Wang G, Zhang Q, Li J. Over-expression of microRNA-25 promotes cell proliferation and induces cell apoptosis in patients with hepatocellular carcinoma. Int J Clin Exp Pathol. 2016; 9:3396–3402.
33. Yang X, Liang L, Zhang XF, Jia HL, Qin Y, Zhu XC, Gao XM, Qiao P, Zheng Y, Sheng YY, Wei JW, Zhou HJ, Ren N, et al. MicroRNA-26a suppresses tumor growth and metastasis of human hepatocellular carcinoma by targeting interleukin-6-Stat3 pathway. Hepatology. 2013; 58: 158–170.
34. Wang Y, Sun B, Zhao X, Zhao N, Sun R, Zhu D, Zhang Y, Li Y, Gu Q, Dong X, Wang M, An J. Twist1-related miR-26b-5p suppresses epithelial-mesenchymal transition, migration and invasion by targeting SMAD1 in hepatocellular carcinoma. Oncotarget. 2016; 7:24383–24401. https://doi.org/10.18632/oncotarget.8328.
35. Sun XF, Sun JP, Hou HT, Li K, Liu X, Ge QX. MicroRNA-27b exerts an oncogenic function by targeting Fbxw7 in human hepatocellular carcinoma. Tumour Biol. 2016; 37:15325–15332.
36. Zhou SL, Hu ZQ, Zhou ZJ, Dai Z, Wang Z, Cao Y, Fan J, Huang XW, Zhou J. miR-28-5p-IL-34-macrophage feedback loop modulates hepatocellular carcinoma metastasis. Hepatology. 2016; 63:1560–1575.
37. Zhu HT, Dong QZ, Sheng YY, Wei JW, Wang G, Zhou HJ, Ren N, Jia HL, Ye QH, Qin LX. MicroRNA-29a-5p is a novel predictor for early recurrence of hepatitis B virus-related hepatocellular carcinoma after surgical resection. PLoS One. 2012; 7:e52393.
38. Parpart S, Roessler S, Dong F, Rao V, Takai A, Ji J, Qin LX, Ye QH, Jia HL, Tang ZY, Wang XW. Modulation of miR-29 expression by α-fetoprotein is linked to the hepatocellular carcinoma epigenome. Hepatology. 2014; 60:872–883.
39. Xiong Y, Fang JH, Yun JP, Yang J, Zhang Y, Jia WH, Zhuang SM. Effects of microRNA-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology. 2010; 51:836–845.
40. Bae HJ, Noh JH, Kim JK, Eun JW, Jung KH, Kim MG, Chang YG, Shen Q, Kim SJ, Park WS, Lee JY, Nam SW. MicroRNA-29c functions as a tumor suppressor by direct targeting oncogenic SIRT1 in hepatocellular carcinoma. Oncogene. 2014; 33:2557–2567.
41. Dong CW, Wang YX, Du FT, Ding W, Hu SY. Low miR-29c expression is a prognostic marker in hepatocellular carcinoma. Genet Mol Res. 2016; 15.
42. Huang WT, Chen ZX, He RQ, Wu YZ, Yin SY, Liang XN, Chen G, Yang H, Peng ZG, Yang LH. Clinicopathological role of miR-30a-5p in hepatocellular carcinoma tissues and prediction of its function with bioinformatics analysis. Onco Targets Ther. 2016; 9: 5061–5071.
43. Liu Z, Tu K, Liu Q. Effects of microRNA-30a on migration, invasion and prognosis of hepatocellular carcinoma. FEBS Lett. 2014; 588:3089–3097.
44. Qin X, Chen J, Wu L, Liu Z. MiR-30b-5p acts as a tumor suppressor, repressing cell proliferation and cell cycle in human hepatocellular carcinoma. Biomed Pharmacother. 2017; 89:742–750.
45. Kim HS, Lee KS, Bae HJ, Eun JW, Shen Q, Park SJ, Shin WC, Yang HD, Park M, Park WS, Kang YK, Nam SW. MicroRNA-31 functions as a tumor suppressor by regulating cell cycle and epithelial-mesenchymal transition regulatory proteins in liver cancer. Oncotarget. 2015; 6:8089–8102. https://doi.org/10.18632/oncotarget.3512.
46. Han SY, Han HB, Tian XY, Sun H, Xue D, Zhao C, Jiang ST, He XR, Zheng WX, Wang J, Pang LN, Li XH, Li PP. MicroRNA-33a-3p suppresses cell migration and invasion by directly targeting PBX3 in human hepatocellular carcinoma. Oncotarget. 2016; 7:42461–42473. https://doi.org/10.18632/oncotarget.9886.
47. Li XY, Wen JY, Jia CC, Wang TT, Li X, Dong M, Lin QU, Chen ZH, Ma XK, Wei LI, Lin ZX, Ruan DY, Chen J, et al. MicroRNA-34a-5p enhances sensitivity to chemotherapy by targeting AXL in hepatocellular carcinoma MHCC-97L cells. Oncol Lett. 2015; 10:2691–2698.
48. Yang F, Li QJ, Gong ZB, Zhou L, You N, Wang S, Li XL, Li JJ, An JZ, Wang DS, He Y, Dou KF. MicroRNA-34a targets Bcl-2 and sensitizes human hepatocellular carcinoma cells to sorafenib treatment. Technol Cancer Res Treat. 2014; 13:77–86.
49. Cui X, Wu Y, Wang Z, Liu X, Wang S, Qin C. MicroRNA-34a expression is predictive of recurrence after radiofrequency ablation in early hepatocellular carcinoma. Tumour Biol. 2015; 36:3887–3893.
50. Xu X, Chen W, Miao R, Zhou Y, Wang Z, Zhang L, Wan Y, Dong Y, Qu K, Liu C. miR-34a induces cellular senescence via modulation of telomerase activity in human hepatocellular carcinoma by targeting FoxM1/c-Myc pathway. Oncotarget. 2015; 6:3988–4004. https://doi.org/10.18632/oncotarget.2905.
51. Zhang X, Zhu Q, Li J, Huang L, Li J, Yan J, Yan Y. MicroRNA-34b/c regulated by p53 is associated with unfavorable prognosis in patients with early hepatocellular carcinoma. Int J Clin Exp Pathol. 2016; 9:6117–6126.
52. Xiao CZ, Wei W, Guo ZX, Zhang MY, Zhang YF, Wang JH, Shi M, Wang HY, Guo RP. MicroRNA-34c-3p promotes cell proliferation and invasion in hepatocellular carcinoma by regulation of NCKAP1 expression. J Cancer Res Clin Oncol. 2017; 143:263–273.
53. Yang W, Dou C, Wang Y, Jia Y, Li C, Zheng X, Tu K. MicroRNA-92a contributes to tumor growth of human hepatocellular carcinoma by targeting FBXW7. Oncol Rep. 2015; 34:2576–2584.
54. Ohta K, Hoshino H, Wang J, Ono S, Iida Y, Hata K, Huang SK, Colquhoun S, Hoon DS. MicroRNA-93 activates c-Met/PI3K/Akt pathway activity in hepatocellular carcinoma by directly inhibiting PTEN and CDKN1A. Oncotarget. 2015; 6:3211–3224. https://doi.org/10.18632/oncotarget.3085.
55. Zhou W, Zou B, Liu L, Cui K, Gao J, Yuan S, Cong N. MicroRNA-98 acts as a tumor suppressor in hepatocellular carcinoma via targeting SALL4. Oncotarget. 2016; 7:74059–74073. https://doi.org/10.18632/oncotarget.12190.
56. Li D, Liu X, Lin L, Hou J, Li N, Wang C, Wang P, Zhang Q, Zhang P, Zhou W, Wang Z, Ding G, Zhuang SM, et al. MicroRNA-99a inhibits hepatocellular carcinoma growth and correlates with prognosis of patients with hepatocellular carcinoma. J Biol Chem. 2011; 286:36677–36685.
57. Zhang J, Jin H, Liu H, Lv S, Wang B, Wang R, Liu H, Ding M, Yang Y, Li L, Zhang J, Fu S, Xie D, et al. MiRNA-99a directly regulates AGO2 through translational repression in hepatocellular carcinoma. Oncogenesis. 2014; 3:e97.
58. Yang J, Liu X, Yuan X, Wang Z. miR-99b promotes metastasis of hepatocellular carcinoma through inhibition of claudin 11 expression and may serve as a prognostic marker. Oncol Rep. 2015; 34:1415–1423.
59. Chen P, Zhao X, Ma L. Downregulation of microRNA-100 correlates with tumor progression and poor prognosis in hepatocellular carcinoma. Mol Cell Biochem. 2013; 383:49–58.
60. Zhou HC, Fang JH, Luo X, Zhang L, Yang J, Zhang C, Zhuang SM. Downregulation of microRNA-100 enhances the ICMT-Rac1 signaling and promotes metastasis of hepatocellular carcinoma cells. Oncotarget. 2014; 5: 12177–12188. https://doi.org/10.18632/oncotarget.2601.
61. Zhang Y, Guo X, Xiong L, Kong X, Xu Y, Liu C, Zou L, Li Z, Zhao J, Lin N. MicroRNA-101 suppresses SOX9-dependent tumorigenicity and promotes favorable prognosis of human hepatocellular carcinoma. FEBS Lett. 2012; 586:4362–4370.
62. Lv X, Li J, Yang B. Clinical effects of miR-101 on prognosis of hepatocellular carcinoma and carcinogenic mechanism of anti-miR-101. Oncol Rep. 2016; 36:2184–2192.
63. Ma YS, Wu TM, Lv ZW, Lu GX, Cong XL, Xie RT, Yang HQ, Chang ZY, Sun R, Chai L, Cai MX, Zhong XJ, Zhu J, et al. High expression of miR-105-1 positively correlates with clinical prognosis of hepatocellular carcinoma by targeting oncogene NCOA1.Oncotarget. 2017; 8: 11896–11905. https://doi.org/10.18632/oncotarget.14435.
64. Li BK, Huang PZ, Qiu JL, Liao YD, Hong J, Yuan YF. Upregulation of microRNA-106b is associated with poor prognosis in hepatocellular carcinoma. Diagn Pathol. 2014; 9:226.
65. Yen CS, Su ZR, Lee YP, Liu IT, Yen CJ. miR-106b promotes cancer progression in hepatitis B virus-associated hepatocellular carcinoma. World J Gastroenterol. 2016; 22:5183–5192.
66. Wu Q, Liu HO, Liu YD, Liu WS, Pan D, Zhang WJ, Yang L, Fu Q, Xu JJ, Gu JX. Decreased expression of hepatocyte nuclear factor 4α (Hnf4α)/microRNA-122 (miR-122) axis in hepatitis B virus-associated hepatocellular carcinoma enhances potential oncogenic GALNT10 activity. J Biol Chem. 2015; 290:1170–1185.
67. Xu Q, Zhang M, Tu J, Pang L, Cai W, Liu X. MicroRNA-122 affects cell aggressiveness and apoptosis by targeting PKM2 in human hepatocellular carcinoma. Oncol Rep. 2015; 34:2054–2064.
68. Xu L, Dai W, Li J, He L, Wang F, Xia Y, Chen K, Li S, Liu T, Lu J, Zhou Y, Wang Y, Guo C. Methylation-regulated miR-124-1 suppresses tumorigenesis in hepatocellular carcinoma by targeting CASC3. Oncotarget. 2016; 7:26027–26041. https://doi.org/10.18632/oncotarget.8266.
69. Zheng F, Liao YJ, Cai MY, Liu YH, Liu TH, Chen SP, Bian XW, Guan XY, Lin MC, Zeng YX, Kung HF, Xie D. The putative tumour suppressor microRNA-124 modulates hepatocellular carcinoma cell aggressiveness by repressing ROCK2 and EZH2. Gut. 2012; 61:278–289.
70. Bi Q, Tang S, Xia L, Du R, Fan R, Gao L, Jin J, Liang S, Chen Z, Xu G, Nie Y, Wu K, Liu J, et al. Ectopic expression of MiR-125a inhibits the proliferation and metastasis of hepatocellular carcinoma by targeting MMP11 and VEGF. PLoS One. 2012;7:e40169.
71. Li W, Xie L, He X, Li J, Tu K, Wei L, Wu J, Guo Y, Ma X, Zhang P, Pan Z, Hu X, Zhao Y, et al. Diagnostic and prognostic implications of microRNAs in human hepatocellular carcinoma. Int J Cancer. 2008; 123:1616–1622.
72. Tsang FH, Au V, Lu WJ, Shek FH, Liu AM, Luk JM, Fan ST, Poon RT, Lee NP. Prognostic marker microRNA-125b inhibits tumorigenic properties of hepatocellular carcinoma cells via suppressing tumorigenic molecule eIF5A2. Dig Dis Sci. 2014; 59:2477–2487.
73. Chen H, Miao R, Fan J, Han Z, Wu J, Qiu G, Tang H, Peng Z. Decreased expression of miR-126 correlates with metastatic recurrence of hepatocellular carcinoma. Clin Exp Metastasis. 2013; 30:651–658.
74. Huang CY, Huang XP, Zhu JY, Chen ZG, Li XJ, Zhang XH, Huang S, He JB, Lian F, Zhao YN, Wu GB. miR-128-3p suppresses hepatocellular carcinoma proliferation by regulating PIK3R1 and is correlated with the prognosis of HCC patients. Oncol Rep. 2015; 33:2889–2898.
75. Ma N, Chen F, Shen SL, Chen W, Chen LZ, Su Q, Zhang LJ, Bi J, Zeng WT, Li W, Huang XH, Wang Q. MicroRNA-129-5p inhibits hepatocellular carcinoma cell metastasis and invasion via targeting ETS1. Biochem Biophys Res Commun. 2015; 461:618–623.
76. Liu Z, Dou C, Yao B, Xu M, Ding L, Wang Y, Jia Y, Li Q, Zhang H, Tu K, Song T, Liu Q. Methylation-mediated repression of microRNA-129-2 suppresses cell aggressiveness by inhibiting high mobility group box 1 in human hepatocellular carcinoma. Oncotarget. 2016; 7:36909–36923. https://doi.org/10.18632/oncotarget.9377.
77. Li B, Huang P, Qiu J, Liao Y, Hong J, Yuan Y. MicroRNA-130a is down-regulated in hepatocellular carcinoma and associates with poor prognosis. Med Oncol. 2014; 31:230.
78. Wang WY, Zhang HF, Wang L, Ma YP, Gao F, Zhang SJ, Wang LC. High expression of microRNA-130b correlates with poor prognosis of patients with hepatocellular carcinoma. Diagn Pathol. 2014; 9:160.
79. Liu S, Guo W, Shi J, Li N, Yu X, Xue J, Fu X, Chu K, Lu C, Zhao J, Xie D, Wu M, Cheng S, et al. MicroRNA-135a contributes to the development of portal vein tumor thrombus by promoting metastasis in hepatocellular carcinoma. J Hepatol. 2012; 56:389–396.
80. Liu LL, Lu SX, Li M, Li LZ, Fu J, Hu W, Yang YZ, Luo RZ, Zhang CZ, Yun JP. FoxD3-regulated microRNA-137 suppresses tumour growth and metastasis in human hepatocellular carcinoma by targeting AKT2. Oncotarget. 2014; 5:5113–5124. https://doi.org/10.18632/oncotarget.2089.
81. Sakabe T, Azumi J, Umekita Y, Toriguchi K, Hatano E, Hirooka Y, Shiota G. Prognostic relevance of miR-137 in patients with hepatocellular carcinoma. Liver Int. 2017; 37:271–279.
82. Wang Z, Ding Q, Li Y, Liu Q, Wu W, Wu L, Yu H. Reanalysis of microRNA expression profiles identifies novel biomarkers for hepatocellular carcinoma prognosis. Tumour Biol. 2016; 37:14779–14787.
83. Wong CC, Wong CM, Tung EK, Au SL, Lee JM, Poon RT, Man K, Ng IO. The microRNA miR-139 suppresses metastasis and progression of hepatocellular carcinoma by down-regulating Rho-Kinase 2. Gastroenterology. 2011; 140:322–331.
84. Yang H, Fang F, Chang R, Yang L. MicroRNA-140-5p suppresses tumor growth and metastasis by targeting transforming growth factor β receptor 1 and fibroblast growth factor 9 in hepatocellular carcinoma. Hepatology. 2013; 58:205–217.
85. Law PT, Ching AK, Chan AW, Wong QW, Wong CK, To KF, Wong N. MiR-145 modulates multiple components of the insulin-like growth factor pathway in hepatocellular carcinoma. Carcinogenesis. 2012; 33:1134–1141.
86. Zhang Z, Zhang Y, Sun XX, Ma X, Chen ZN. microRNA-146a inhibits cancer metastasis by downregulating VEGF through dual pathways in hepatocellular carcinoma. Mol Cancer. 2015; 14:5.
87. Heo MJ, Kim YM, Koo JH, Yang YM, An J, Lee SK, Lee SJ, Kim KM, Park JW, Kim SG. microRNA-148a dysregulation discriminates poor prognosis of hepatocellular carcinoma in association with USP4 overexpression. Oncotarget. 2014; 5:2792–2806. https://doi.org/10.18632/oncotarget.1920.
88. Li L, Liu Y, Guo Y, Liu B, Zhao Y, Li P, Song F, Zheng H, Yu J, Song T, Niu R, Li Q, Wang XW, et al. Regulatory MiR-148a-ACVR1/BMP circuit defines a cancer stem cell-like aggressive subtype of hepatocellular carcinoma. Hepatology. 2015; 61:574–584.
89. Pan L, Huang S, He R, Rong M, Dang Y, Chen G. Decreased expression and clinical significance of miR-148a in hepatocellular carcinoma tissues. Eur J Med Res. 2014;19:68.
90. Zhang Z, Zheng W, Hai J. MicroRNA-148b expression is decreased in hepatocellular carcinoma and associated with prognosis. Med Oncol. 2014;31:984.
91. Zhang JG, Shi Y, Hong DF, Song M, Huang D, Wang CY, Zhao G. MiR-148b suppresses cell proliferation and invasion in hepatocellular carcinoma by targeting WNT1/β-catenin pathway. Sci Rep. 2015;5:8087.
92. Zhang Y, Guo X, Xiong L, Yu L, Li Z, Guo Q, Li Z, Li B, Lin N. Comprehensive analysis of microRNA-regulated protein interaction network reveals the tumor suppressive role of microRNA-149. Mol Cancer. 2014;13:253.
93. Luo G, Chao YL, Tang B, Li BS, Xiao YF, Xie R, Wang SM, Wu YY, Dong H, Liu XD, Yang SM. miR-149 represses metastasis of hepatocellular carcinoma by targeting actin-regulatory proteins PPM1F. Oncotarget. 2015; 6: 37808–37823. https://doi.org/10.18632/oncotarget.5676.
94. Lin L, Zhang YD, Chen ZY, Chen Y, Ren CP. The clinicopathological significance of miR-149 and PARP-2 in hepatocellular carcinoma and their roles in chemo/radiotherapy. Tumour Biol. 2016; 37:12339–12346.
95. Sun W, Zhang Z, Wang J, Shang R, Zhou L, Wang X, Duan J, Ruan B, Gao Y, Dai B, Qu S, Liu W, Ding R, et al. MicroRNA-150 suppresses cell proliferation and metastasis in hepatocellular carcinoma by inhibiting the GAB1-ERK axis. Oncotarget. 2016; 7:11595–11608. https://doi.org/10.18632/oncotarget.7292.
96. Tan YL, Shan S, Bai ZG, Sun LY, Zhu ZJ, Wei L, Qu W, Zeng ZG, Liu Y, Ma XM, Yang Y, Wang TT, Zou WL, et al. Expression of miR-151 in hepatocellular carcinoma tissues and its clinicopathological significance. Int J Clin Exp Pathol. 2017; 10:829–834.
97. Dang YW, Zeng J, He RQ, Rong MH, Luo DZ, Chen G. Effects of miR-152 on cell growth inhibition, motility suppression and apoptosis induction in hepatocellular carcinoma cells. Asian Pac J Cancer Prev. 2014; 15: 4969–4976.
98. Tang B, Lei B, Qi G, Liang X, Tang F, Yuan S, Wang Z, Yu S, He S. MicroRNA-155-3p promotes hepatocellular carcinoma formation by suppressing FBXW7 expression. J Exp Clin Cancer Res. 2016; 35:93.
99. Han ZB, Chen HY, Fan JW, Wu JY, Tang HM, Peng ZH. Up-regulation of microRNA-155 promotes cancer cell invasion and predicts poor survival of hepatocellular carcinoma following liver transplantation. J Cancer Res Clin Oncol. 2012; 138:153–161.
100. Zhang L, Wang W, Li X, He S, Yao J, Wang X, Zhang D, Sun X. MicroRNA-155 promotes tumor growth of human hepatocellular carcinoma by targeting ARID2. Int J Oncol. 2016; 48:2425–2434.
101. Wang J, Li J, Shen J, Wang C, Yang L, Zhang X. MicroRNA-182 downregulates metastasis suppressor 1 and contributes to metastasis of hepatocellular carcinoma. BMC Cancer. 2012; 12:227.
102. Liang Z, Gao Y, Shi W, Zhai D, Li S, Jing L, Guo H, Liu T, Wang Y, Du Z. Expression and significance of microRNA-183 in hepatocellular carcinoma. ScientificWorldJournal. 2013; 2013:381874.
103. Zhi Q, Zhu J, Guo X, He S, Xue X, Zhou J, Hu B, Li H, Chen S, Zhao H, Kuang Y. Metastasis-related miR-185 is a potential prognostic biomarker for hepatocellular carcinoma in early stage. Biomed Pharmacother. 2013; 67:393–398.
104. Dou C, Liu Z, Xu M, Jia Y, Wang Y, Li Q, Yang W, Zheng X, Tu K, Liu Q. miR-187-3p inhibits the metastasis and epithelial-mesenchymal transition of hepatocellular carcinoma by targeting S100A4. Cancer Lett. 2016; 381:380–390.
105. Fang F, Chang RM, Yu L, Lei X, Xiao S, Yang H, Yang LY. MicroRNA-188-5p suppresses tumor cell proliferation and metastasis by directly targeting FGF5 in hepatocellular carcinoma. J Hepatol. 2015; 63:874–885.
106. He Y, Cui Y, Wang W, Gu J, Guo S, Ma K, Luo X. Hypomethylation of the hsa-miR-191 locus causes high expression of hsa-miR-191 and promotes the epithelial-to-mesenchymal transition in hepatocellular carcinoma. Neoplasia. 2011; 13:841–853.
107. Ge Y, Yan X, Jin Y, Yang X, Yu X, Zhou L, Han S, Yuan Q, Yang M. MiRNA-192 [corrected] and miRNA-204 Directly Suppress lncRNA HOTTIP and Interrupt GLS1-Mediated Glutaminolysis in Hepatocellular Carcinoma. PLoS Genet. 2015; 11:e1005726.
108. Lian J, Jing Y, Dong Q, Huan L, Chen D, Bao C, Wang Q, Zhao F, Li J, Yao M, Qin L, Liang L, He X. miR-192, a prognostic indicator, targets the SLC39A6/SNAIL pathway to reduce tumor metastasis in human hepatocellular carcinoma. Oncotarget. 2016; 7:2672–2683. https://doi.org/10.18632/oncotarget.6603.
109. Zhao Y, Li F, Zhang X, Liu A, Qi J, Cui H, Zhao P. MicroRNA-194 acts as a prognostic marker and inhibits proliferation in hepatocellular carcinoma by targeting MAP4K4. Int J Clin Exp Pathol. 2015; 8:12446–12454.
110. Wang R, Zhao N, Li S, Fang JH, Chen MX, Yang J, Jia WH, Yuan Y, Zhuang SM. MicroRNA-195 suppresses angiogenesis and metastasis of hepatocellular carcinoma by inhibiting the expression of VEGF, VAV2, and CDC42. Hepatology. 2013; 58:642–653.
111. Wang M, Zhang J, Tong L, Ma X, Qiu X. MiR-195 is a key negative regulator of hepatocellular carcinoma metastasis by targeting FGF2 and VEGFA. Int J Clin Exp Pathol. 2015; 8:14110–14120.
112. Wang H, Su X, Yang M, Chen T, Hou J, Li N, Cao X. Reciprocal control of miR-197 and IL-6/STAT3 pathway reveals miR-197 as potential therapeutic target for hepatocellular carcinoma. Oncoimmunology. 2015; 4:e1031440.
113. Li B, He L, Zuo D, He W, Wang Y, Zhang Y, Liu W, Yuan Y. Mutual Regulation of MiR-199a-5p and HIF-1α Modulates the Warburg Effect in Hepatocellular Carcinoma. J Cancer. 2017; 8:940–949.
114. Wang P, Chen S, Fang H, Wu X, Chen D, Peng L, Gao Z, Xie C. miR-214/199a/199a* cluster levels predict poor survival in hepatocellular carcinoma through interference with cell-cycle regulators. Oncotarget. 2016; 7:929–945. https://doi.org/10.18632/oncotarget.6137.
115. Wang C, Song B, Song W, Liu J, Sun A, Wu D, Yu H, Lian J, Chen L, Han J. Underexpressed microRNA-199b-5p targets hypoxia-inducible factor-1α in hepatocellular carcinoma and predicts prognosis of hepatocellular carcinoma patients. J Gastroenterol Hepatol. 2011; 26:1630–1637.
116. Xiao F, Zhang W, Zhou L, Xie H, Xing C, Ding S, Chen K, Zheng S. microRNA-200a is an independent prognostic factor of hepatocellular carcinoma and induces cell cycle arrest by targeting CDK6. Oncol Rep. 2013; 30:2203–2210.
117. Feng J, Wang J, Chen M, Chen G, Wu Z, Ying L, Zhuo Q, Zhang J, Wang W. miR-200a suppresses cell growth and migration by targeting MACC1 and predicts prognosis in hepatocellular carcinoma. Oncol Rep. 2015; 33:713–720.
118. Yang X, Wang J, Qu S, Zhang H, Ruan B, Gao Y, Ma B, Wang X, Wu N, Li X, Dou K, Li H. MicroRNA-200a suppresses metastatic potential of side population cells in human hepatocellular carcinoma by decreasing ZEB2. Oncotarget. 2015; 6:7918–7929. https://doi.org/10.18632/oncotarget.3486.
119. Chen HY, Han ZB, Fan JW, Xia J, Wu JY, Qiu GQ, Tang HM, Peng ZH. miR-203 expression predicts outcome after liver transplantation for hepatocellular carcinoma in cirrhotic liver. Med Oncol. 2012; 29:1859–1865.
120. Liu Y, Ren F, Rong M, Luo Y, Dang Y, Chen G. Association between underexpression of microrna-203 and clinicopathological significance in hepatocellular carcinoma tissues. Cancer Cell Int. 2015; 15:62.
121. Wan D, Shen S, Fu S, Preston B, Brandon C, He S, Shen C, Wu J, Wang S, Xie W, Chen B, Liya A, Guo Y, et al. miR-203 suppresses the proliferation and metastasis of hepatocellular carcinoma by targeting oncogene ADAM9 and oncogenic long non-coding RNA HULC. Anticancer Agents Med Chem. 2016; 16:414–423.
122. Zhao J, Xu G, Qian YW, Li YW. Down-regulation of miR-205 promotes stemness of hepatocellular carcinoma cells by targeting PLCβ1 and increasing CD24 expression. Neoplasma. 2015; 62:567–573.
123. Kai AK, Chan LK, Lo RC, Lee JM, Wong CC, Wong JC, Ng IO. Down-regulation of TIMP2 by HIF-1α/miR-210/HIF-3α regulatory feedback circuit enhances cancer metastasis in hepatocellular carcinoma. Hepatology. 2016; 64:473–487.
124. Yang Y, Zhang J, Xia T, Li G, Tian T, Wang M, Wang R, Zhao L, Yang Y, Lan K, Zhou W. MicroRNA-210 promotes cancer angiogenesis by targeting fibroblast growth factor receptor-like 1 in hepatocellular carcinoma. Oncol Rep. 2016; 36:2553–2562.
125. Deng B, Qu L, Li J, Fang J, Yang S, Cao Z, Mei Z, Sun X. MiRNA-211 suppresses cell proliferation, migration and invasion by targeting SPARC in human hepatocellular carcinoma. Sci Rep. 2016; 6:26679.
126. Dou C, Wang Y, Li C, Liu Z, Jia Y, Li Q, Yang W, Yao Y, Liu Q, Tu K. MicroRNA-212 suppresses tumor growth of human hepatocellular carcinoma by targeting FOXA1. Oncotarget. 2015; 6:13216–13228. https://doi.org/10.18632/oncotarget.3916.
127. Tu H, Wei G, Cai Q, Chen X, Sun Z, Cheng C, Zhang L, Feng Y, Zhou H, Zhou B, Zeng T. MicroRNA-212 inhibits hepatocellular carcinoma cell proliferation and induces apoptosis by targeting FOXA1. Onco Targets Ther. 2015; 8:2227–2235.
128. Wang J, Li J, Wang X, Zheng C, Ma W. Downregulation of microRNA-214 and overexpression of FGFR-1 contribute to hepatocellular carcinoma metastasis. Biochem Biophys Res Commun. 2013; 439:47–53.
129. Liu FY, Zhou SJ, Deng YL, Zhang ZY, Zhang EL, Wu ZB, Huang ZY, Chen XP. MiR-216b is involved in pathogenesis and progression of hepatocellular carcinoma through HBx-miR-216b-IGF2BP2 signaling pathway. Cell Death Dis. 2015; 6:e1670.
130. Tu K, Li C, Zheng X, Yang W, Yao Y, Liu Q. Prognostic significance of miR-218 in human hepatocellular carcinoma and its role in cell growth. Oncol Rep. 2014; 32:1571–1577.
131. Huang N, Lin J, Ruan J, Su N, Qing R, Liu F, He B, Lv C, Zheng D, Luo R. MiR-219-5p inhibits hepatocellular carcinoma cell proliferation by targeting glypican-3. FEBS Lett. 2012; 586:884–891.
132. Gramantieri L, Fornari F, Ferracin M, Veronese A, Sabbioni S, Calin GA, Grazi GL, Croce CM, Bolondi L, Negrini M. MicroRNA-221 targets Bmf in hepatocellular carcinoma and correlates with tumor multifocality. Clin Cancer Res. 2009; 15:5073–5081.
133. Yoon SO, Chun SM, Han EH, Choi J, Jang SJ, Koh SA, Hwang S, Yu E. Deregulated expression of microRNA-221 with the potential for prognostic biomarkers in surgically resected hepatocellular carcinoma. Hum Pathol. 2011; 42:1391–1400.
134. Rong M, Chen G, Dang Y. Increased MiR-221 expression in hepatocellular carcinoma tissues and its role in enhancing cell growth and inhibiting apoptosis in vitro. BMC Cancer. 2013; 13:21.
135. Chen F, Li XF, Fu DS, Huang JG, Yang SE. Clinical potential of miRNA-221 as a novel prognostic biomarker for hepatocellular carcinoma. Cancer Biomark. 2017; 18:209–214.
136. Wong QW, Ching AK, Chan AW, Choy KW, To KF, Lai PB, Wong N. MiR-222 overexpression confers cell migratory advantages in hepatocellular carcinoma through enhancing AKT signaling. Clin Cancer Res. 2010; 16:867–875.
137. Gyöngyösi B, Végh É, Járay B, Székely E, Fassan M, Bodoky G, Schaff Z, Kiss A. Pretreatment MicroRNA Level and Outcome in Sorafenib-treated Hepatocellular Carcinoma. J Histochem Cytochem. 2014; 62:547–555.
138. Wang L, Bo X, Zheng Q, Xiao X, Wu L, Li B. miR-296 inhibits proliferation and induces apoptosis by targeting FGFR1 in human hepatocellular carcinoma. FEBS Lett. 2016; 590:4252–4262.
139. Chen YL, Xu QP, Guo F, Guan WH. MicroRNA-302d downregulates TGFBR2 expression and promotes hepatocellular carcinoma growth and invasion. Exp Ther Med. 2017; 13:681–687.
140. Li H, Huang W, Luo R. The microRNA-325 inhibits hepatocellular carcinoma progression by targeting high mobility group box 1. Diagn Pathol. 2015; 10:117.
141. Lu M, Kong X, Wang H, Huang G, Ye C, He Z. A novel microRNAs expression signature for hepatocellular carcinoma diagnosis and prognosis. Oncotarget. 2017; 8:8775–8784. https://doi.org/10.18632/oncotarget.14452.
142. Zhou J, Li W, Guo J, Li G, Chen F, Zhou J. Downregulation of miR-329 promotes cell invasion by regulating BRD4 and predicts poor prognosis in hepatocellular carcinoma. Tumour Biol. 2016; 37:3561–3569.
143. Hu X, Feng Y, Sun L, Qu L, Sun C. Roles of microRNA-330 and Its Target Gene ING4 in the Development of Aggressive Phenotype in Hepatocellular Carcinoma Cells. Dig Dis Sci. 2017; 62:715–722.
144. Chang RM, Yang H, Fang F, Xu JF, Yang LY. MicroRNA-331-3p promotes proliferation and metastasis of hepatocellular carcinoma by targeting PH domain and leucine-rich repeat protein phosphatase. Hepatology. 2014; 60:1251–1263.
145. Nie H, Li J, Yang XM, Cao QZ, Feng MX, Xue F, Wei L, Qin W, Gu J, Xia Q, Zhang ZG. Mineralocorticoid receptor suppresses cancer progression and the Warburg effect by modulating the miR-338-3p-PKLR axis in hepatocellular carcinoma. Hepatology. 2015; 62:1145–1159.
146. Wang YL, Chen CM, Wang XM, Wang L. Effects of miR-339-5p on invasion and prognosis of hepatocellular carcinoma. Clin Res Hepatol Gastroenterol. 2016; 40:51–56.
147. Chen Z, Huang Z, Ye Q, Ming Y, Zhang S, Zhao Y, Liu L, Wang Q, Cheng K. Prognostic significance and anti-proliferation effect of microRNA-365 in hepatocellular carcinoma. Int J Clin Exp Pathol. 2015; 8:1705–1711.
148. Xu WP, Yi M, Li QQ, Zhou WP, Cong WM, Yang Y, Ning BF, Yin C, Huang ZW, Wang J, Qian H, Jiang CF, Chen YX, et al. Perturbation of MicroRNA-370/Lin-28 homolog A/nuclear factor kappa B regulatory circuit contributes to the development of hepatocellular carcinoma. Hepatology. 2013; 58:1977–1991.
149. Gu H, Guo X, Zou L, Zhu H, Zhang J. Upregulation of microRNA-372 associates with tumor progression and prognosis in hepatocellular carcinoma. Mol Cell Biochem. 2013; 375:23–30.
150. Wu G, Wang Y, Lu X, He H, Liu H, Meng X, Xia S, Zheng K, Liu B. Low mir-372 expression correlates with poor prognosis and tumor metastasis in hepatocellular carcinoma. BMC Cancer. 2015;15:182.
151. Zhou N, Wu J, Wang X, Sun Z, Han Q, Zhao L. Low-level expression of microRNA-375 predicts poor prognosis in hepatocellular carcinoma. Tumour Biol. 2016; 37: 2145–2152.
152. Chen L, Guan H, Gu C, Cao Y, Shao J, Wang F. miR-383 inhibits hepatocellular carcinoma cell proliferation via targeting APRIL. Tumour Biol. 2016; 37:2497–2507.
153. Yang H, Zheng W, Shuai X, Chang RM, Yu L, Fang F, Yang LY. MicroRNA-424 inhibits Akt3/E2F3 axis and tumor growth in hepatocellular carcinoma. Oncotarget. 2015; 6:27736–27750. https://doi.org/10.18632/oncotarget.4811.
154. Vaira V, Roncalli M, Carnaghi C, Faversani A, Maggioni M, Augello C, Rimassa L, Pressiani T, Spagnuolo G, Di Tommaso L, Fagiuoli S, Rota Caremoli E, Barberis M, et al. MicroRNA-425-3p predicts response to sorafenib therapy in patients with hepatocellular carcinoma. Liver Int. 2015; 35:1077–1086.
155. Huang XY, Yao JG, Huang HD, Wang C, Ma Y, Xia Q, Long XD. MicroRNA-429 Modulates Hepatocellular Carcinoma Prognosis and Tumorigenesis. Gastroenterol Res Pract. 2013; 2013:804128.
156. Huang JY, Zhang K, Chen DQ, Chen J, Feng B, Song H, Chen Y, Zhu Z, Lu L, De W, Wang R, Chen LB. MicroRNA-451: epithelial-mesenchymal transition inhibitor and prognostic biomarker of hepatocelluar carcinoma. Oncotarget. 2015; 6:18613–18630. https://doi.org/10.18632/oncotarget.4317.
157. Zheng Z, Liu J, Yang Z, Wu L, Xie H, Jiang C, Lin B, Chen T, Xing C, Liu Z, Song P, Yin S, Zheng S, et al. MicroRNA-452 promotes stem-like cells of hepatocellular carcinoma by inhibiting Sox7 involving Wnt/β-catenin signaling pathway. Oncotarget. 2016; 7:28000–28012. https://doi.org/10.18632/oncotarget.8584.
158. Zhou L, Qu YM, Zhao XM, Yue ZD. Involvement of miR-454 overexpression in the poor prognosis of hepatocellular carcinoma. Eur Rev Med Pharmacol Sci. 2016; 20:825–829.
159. Qin L, Zhang Y, Lin J, Shentu Y, Xie X. MicroRNA-455 regulates migration and invasion of human hepatocellular carcinoma by targeting Runx2. Oncol Rep. 2016; 36: 3325–3332.
160. Huang XP, Hou J, Shen XY, Huang CY, Zhang XH, Xie YA, Luo XL. MicroRNA-486-5p, which is downregulated in hepatocellular carcinoma, suppresses tumor growth by targeting PIK3R1. FEBS J. 2015; 282:579–594.
161. Lin Y, Liu J, Huang Y, Liu D, Zhang G, Kan H. microRNA-489 Plays an Anti-Metastatic Role in Human Hepatocellular Carcinoma by Targeting Matrix Metalloproteinase-7. Transl Oncol. 2017; 10:211–220.
162. Chuang KH, Whitney-Miller CL, Chu CY, Zhou Z, Dokus MK, Schmit S, Barry CT. MicroRNA-494 is a master epigenetic regulator of multiple invasion-suppressor microRNAs by targeting ten eleven translocation 1 in invasive human hepatocellular carcinoma tumors. Hepatology. 2015; 62:466–480.
163. Zhang L, Yu Z, Xian Y, Lin X. microRNA-497 inhibits cell proliferation and induces apoptosis by targeting YAP1 in human hepatocellular carcinoma. FEBS Open Bio. 2016; 6:155–164.
164. Xiao F, Zhang W, Chen L, Chen F, Xie H, Xing C, Yu X, Ding S, Chen K, Guo H, Cheng J, Zheng S, Zhou L. MicroRNA-503 inhibits the G1/S transition by downregulating cyclin D3 and E2F3 in hepatocellular carcinoma. J Transl Med. 2013;11:195.
165. Shao J, Cao J, Liu Y, Mei H, Zhang Y, Xu W. MicroRNA-519a promotes proliferation and inhibits apoptosis of hepatocellular carcinoma cells by targeting FOXF2. FEBS Open Bio. 2015; 5:893–899.
166. Tu K, Liu Z, Yao B, Han S, Yang W. MicroRNA-519a promotes tumor growth by targeting PTEN/PI3K/AKT signaling in hepatocellular carcinoma. Int J Oncol. 2016; 48:965–974.
167. Kan H, Guo W, Huang Y, Liu D. MicroRNA-520g induces epithelial-mesenchymal transition and promotes metastasis of hepatocellular carcinoma by targeting SMAD7. FEBS Lett. 2015; 589:102–109.
168. Shi YH, Qi BB, Liu XB, Ding HM. Upregulation of miR-522 is associated with poor outcome of hepatocellular carcinoma. Eur Rev Med Pharmacol Sci. 2016; 20: 3194–3198.
169. Li JJ, Yang X, Lin XM, He RQ, Lin XG, Luo YH, Wang HL, Luo FF, Wen X, Dang YW, Chen G. Downregulation of microRNA-542-5p and its relationship with EGFR and clinicopathological features in hepatocellular carcinoma. Int J Clin Exp Pathol. 2016; 9:920–929.
170. Liu Z, Dou C, Yao B, Xu M, Ding L, Wang Y, Jia Y, Li Q, Zhang H, Tu K, Song T, Liu Q. Ftx non coding RNA-derived miR-545 promotes cell proliferation by targeting RIG-I in hepatocellular carcinoma. Oncotarget. 2016; 7:25350–25365. https://doi.org/10.18632/oncotarget.8129.
171. Zhang X, Jiang P, Shuai L, Chen K, Li Z, Zhang Y, Jiang Y, Li X. miR-589-5p inhibits MAP3K8 and suppresses CD90+ cancer stem cells in hepatocellular carcinoma. J Exp Clin Cancer Res. 2016;35:176.
172. Jia YY, Zhao JY, Li BL, Gao K, Song Y, Liu MY, Yang XJ, Xue Y, Wen AD, Shi L. miR-592/WSB1/HIF-1α axis inhibits glycolytic metabolism to decrease hepatocellular carcinoma growth.Oncotarget. 2016; 7:35257–35269. https://doi.org/10.18632/oncotarget.9135.
173. Wang K, Liang Q, Wei L, Zhang W, Zhu P. MicroRNA-608 acts as a prognostic marker and inhibits the cell proliferation in hepatocellular carcinoma by macrophage migration inhibitory factor. Tumour Biol. 2016; 37:3823–3830.
174. Zeng XC, Liu FQ, Yan R, Yi HM, Zhang T, Wang GY, Li Y, Jiang N. Downregulation of miR-610 promotes proliferation and tumorigenicity and activates Wnt/β-catenin signaling in human hepatocellular carcinoma. Mol Cancer. 2014;13:261.
175. Song WH, Feng XJ, Gong SJ, Chen JM, Wang SM, Xing DJ, Zhu MH, Zhang SH, Xu AM. microRNA-622 acts as a tumor suppressor in hepatocellular carcinoma. Cancer Biol Ther. 2015; 16:1754–1763.
176. Zhou X, Zhang CZ, Lu SX, Chen GG, Li LZ, Liu LL, Yi C, Fu J, Hu W, Wen JM, Yun JP. miR-625 suppresses tumour migration and invasion by targeting IGF2BP1 in hepatocellular carcinoma. Oncogene. 2015; 34:965–977.
177. Chen WX, Zhang ZG, Ding ZY, Liang HF, Song J, Tan XL, Wu JJ, Li GZ, Zeng Z, Zhang BX, Chen XP. MicroRNA-630 suppresses tumor metastasis through the TGF-β-miR-630-Slug signaling pathway and correlates inversely with poor prognosis in hepatocellular carcinoma. Oncotarget. 2016; 7:22674–22686. https://doi.org/10.18632/oncotarget.8047.
178. Zhang CZ, Cao Y, Fu J, Yun JP, Zhang MF. miR-634 exhibits anti-tumor activities toward hepatocellular carcinoma via Rab1A and DHX33. Mol Oncol. 2016; 10:1532–1541.
179. Cheng J, Chen Y, Zhao P, Liu X, Dong J, Li J, Huang C, Wu R, Lv Y. Downregulation of miRNA-638 promotes angiogenesis and growth of hepatocellular carcinoma by targeting VEGF. Oncotarget. 2016; 7:30702–30711. https://doi.org/10.18632/oncotarget.8930.
180. Zhang Y, Zhang D, Jiang J, Dong L. Loss of miR-638 promotes invasion and epithelial-mesenchymal transition by targeting SOX2 in hepatocellular carcinoma. Oncol Rep. 2017; 37:323–332.
181. Tan YL, Bai ZG, Zou WL, Ma XM, Wang TT, Guo W, Liu J, Li JS, Jie-Yin, Zang YJ, Zhang ZT. miR-744 is a potential prognostic marker in patients with hepatocellular carcinoma. Clin Res Hepatol Gastroenterol. 2015; 39: 359–365.
182. Zhang Z, Yin J, Yang J, Shen W, Zhang C, Mou W, Luo J, Yan H, Sun P, Luo Y, Tian Y, Xiang R. miR-885-5p suppresses hepatocellular carcinoma metastasis and inhibits Wnt/β-catenin signaling pathway. Oncotarget. 2016; 7:75038–75051. https://doi.org/10.18632/oncotarget.12602.
183. Jia B, Tan L, Jin Z, Jiao Y, Fu Y, Liu Y. MiR-892a Promotes Hepatocellular Carcinoma Cells Proliferation and Invasion Through Targeting CD226. J Cell Biochem. 2017; 118:1489–1496.
184. Yuan B, Liang Y, Wang D, Luo F. MiR-940 inhibits hepatocellular carcinoma growth and correlates with prognosis of hepatocellular carcinoma patients. Cancer Sci. 2015; 106:819–824.
185. Ding D, Zhang Y, Yang R, Wang X, Ji G, Huo L, Shao Z, Li X. miR-940 Suppresses Tumor Cell Invasion and Migration via Regulation of CXCR2 in Hepatocellular Carcinoma. Biomed Res Int. 2016;2016:7618342.
186. He Z, Xu H, Meng Y, Kuang Y. miR-944 acts as a prognostic marker and promotes the tumor progression in endometrial cancer. Biomed Pharmacother. 2017; 88:902–910.
187. Tan G, Wu L, Tan J, Zhang B, Tai WC, Xiong S, Chen W, Yang J, Li H. MiR-1180 promotes apoptotic resistance to human hepatocellular carcinoma via activation of NF-κB signaling pathway. Sci Rep. 2016;6:22328.
188. Sun Z, Meng C, Wang S, Zhou N, Guan M, Bai C, Lu S, Han Q, Zhao RC. MicroRNA-1246 enhances migration and invasion through CADM1 in hepatocellular carcinoma. BMC Cancer. 2014;14:616.
189. Lu YL, Yao JG, Huang XY, Wang C, Wu XM, Xia Q, Long XD. Prognostic significance of miR-1268a expression and its beneficial effects for post-operative adjuvant transarterial chemoembolization in hepatocellular carcinoma. Sci Rep. 2016;6:36104.
190. Gan TQ, Tang RX, He RQ, Dang YW, Xie Y, Chen G. Upregulated MiR-1269 in hepatocellular carcinoma and its clinical significance. Int J Clin Exp Med. 2015; 8:714–721.
191. Law PT, Qin H, Ching AK, Lai KP, Co NN, He M, Lung RW, Chan AW, Chan TF, Wong N. Deep sequencing of small RNA transcriptome reveals novel non-coding RNAs in hepatocellular carcinoma. J Hepatol. 2013; 58:1165–1173.
192. Jiang J, Zhang Y, Guo Y, Yu C, Chen M, Li Z, Tian S, Sun C. MicroRNA-3127 promotes cell proliferation and tumorigenicity in hepatocellular carcinoma by disrupting of PI3K/AKT negative regulation. Oncotarget. 2015; 6: 6359–6372. https://doi.org/10.18632/oncotarget.3438.
193. Tang D, Sun B, Yu H, Yang Z, Zhu L. Tumor-suppressing effect of miR-4458 on human hepatocellular carcinoma. Cell Physiol Biochem. 2015; 35:1797–1807.
194. Bo W, Hu Y, Feng X, Zhang H, Tian L, Liu A. The tumor suppressor role of miR-4782-3p in hepatocellular carcinoma. Oncol Rep. 2016; 35:2107–2112.
195. Köberle V, Kronenberger B, Pleli T, Trojan J, Imelmann E, Peveling-Oberhag J, Welker MW, Elhendawy M, Zeuzem S, Piiper A, Waidmann O. Serum microRNA-1 and microRNA-122 are prognostic markers in patients with hepatocellular carcinoma. Eur J Cancer. 2013; 49: 3442–3449.
196. Yoon EL, Yeon JE, Ko E, Lee HJ, Je JH, Yoo YJ, Kang SH, Suh SJ, Kim JH, Seo YS, Yim HJ, Byun KS. An Explorative Analysis for the Role of Serum miR-10b-3p Levels in Predicting Response to Sorafenib in Patients with Advanced Hepatocellular Carcinoma. J Korean Med Sci. 2017; 32:212–220.
197. Tomimaru Y, Eguchi H, Nagano H, Wada H, Kobayashi S, Marubashi S, Tanemura M, Tomokuni A, Takemasa I, Umeshita K, Kanto T, Doki Y, Mori M. Circulating microRNA-21 as a novel biomarker for hepatocellular carcinoma. J Hepatol. 2012; 56:167–175.
198. Liu M, Liu J, Wang L, Wu H, Zhou C, Zhu H, Xu N, Xie Y. Association of serum microRNA expression in hepatocellular carcinomas treated with transarterial chemoembolization and patient survival. PLoS One. 2014;9:e109347.
199. Wang X, Zhang J, Zhou L, Lu P, Zheng ZG, Sun W, Wang JL, Yang XS, Li XL, Xia N, Zhang N, Dou KF. Significance of serum microRNA-21 in diagnosis of HCC: clinical analyses of patients and an HCC rat model. Int J Clin Exp Pathol. 2015; 8:1466–1478.
200. Cho HJ, Kim SS, Nam JS, Kim JK, Lee JH, Kim B, Wang HJ, Kim BW, Lee JD, Kang DY, Kim JH, Jae YM, Hwang JC, et al. Low levels of circulating microRNA-26a/29a as poor prognostic markers in patients with hepatocellular carcinoma who underwent curative treatment. Clin Res Hepatol Gastroenterol. 2017; 41:181–189.
201. Meng FL, Wang W, Jia WD. Diagnostic and prognostic significance of serum miR-24-3p in HBV-related hepatocellular carcinoma. Med Oncol. 2014; 31:177.
202. Zhu HT, Hasan AM, Liu RB, Zhang ZC, Zhang X, Wang J, Wang HY, Wang F, Shao JY. Serum microRNA profiles as prognostic biomarkers for HBV-positive hepatocellular carcinoma. Oncotarget. 2016; 7:45637–45648. https://doi.org/10.18632/oncotarget.10082.
203. Chen Y, Dong X, Yu D, Wang X. Serum miR-96 is a promising biomarker for hepatocellular carcinoma in patients with chronic hepatitis B virus infection. Int J Clin Exp Med. 2015; 8:18462–18468.
204. Zheng F, Liao YJ, Cai MY, Liu TH, Chen SP, Wu PH, Wu L, Bian XW, Guan XY, Zeng YX, Yuan YF, Kung HF, Xie D. Systemic delivery of microRNA-101 potently inhibits hepatocellular carcinoma in vivo by repressing multiple targets. PLoS Genet. 2015;11:e1004873.
205. Cho HJ, Kim JK, Nam JS, Wang HJ, Lee JH, Kim BW, Kim SS, Noh CK, Shin SJ, Lee KM, Cho SW, Cheong JY. High circulating microRNA-122 expression is a poor prognostic marker in patients with hepatitis B virus-related. Clin Biochem. 2015;48:1073–1078.
206. Xu Y, Bu X, Dai C, Shang C. High serum microRNA-122 level is independently associated with higher overall survival rate in hepatocellular carcinoma patients. Tumour Biol. 2015; 36:4773–4776.
207. Kim SS, Nam JS, Cho HJ, Won JH, Kim JW, Ji JH, Yang MJ, Park JH, Noh CK, Shin SJ, Lee KM, Cho SW, Cheong JY. Plasma micoRNA-122 as a predictive marker for treatment response following transarterial chemoembolization in patients with hepatocellular carcinoma. J Gastroenterol Hepatol. 2017; 32:199–207.
208. Ng KT, Lo CM, Wong N, Li CX, Qi X, Liu XB, Geng W, Yeung OW, Ma YY, Chan SC, Man K. Early-phase circulating miRNAs predict tumor recurrence and survival of hepatocellular carcinoma patients after liver transplantation. Oncotarget. 2016; 7:19824–19839. https://doi.org/10.18632/oncotarget.7627.
209. Wang L, Liu M, Zhu H, Rong W, Wu F, An S, Liu F, Feng L, Wu J, Xu N. Identification of recurrence-related serum microRNAs in hepatocellular carcinoma following hepatectomy. Cancer Biol Ther. 2015; 16:1445–1452.
210. Zhuang L, Xu L, Wang P, Meng Z. Serum miR-128-2 serves as a prognostic marker for patients with hepatocellular carcinoma. PLoS One. 2015; 10:e0117274.
211. Wang F, Ying H, He B, Pan Y, Sun H, Wang S. Circulating miR-148/152 family as potential biomarkers in hepatocellular carcinoma. Tumour Biol. 2016; 37:4945–4953.
212. Yu F, Lu Z, Chen B, Dong P, Zheng J. microRNA-150: a promising novel biomarker for hepatitis B virus-related hepatocellular carcinoma. Diagn Pathol. 2015; 10:129.
213. Nishida N, Arizumi T, Hagiwara S, Ida H, Sakurai T, Kudo M. MicroRNAs for the Prediction of Early Response to Sorafenib Treatment in human Hepatocellular Carcinoma. Liver Cancer. 2017; 6:113–125.
214. Chen L, Chu F, Cao Y, Shao J, Wang F. Serum miR-182 and miR-331-3p as diagnostic and prognostic markers in patients with hepatocellular carcinoma. Tumour Biol. 2015; 36:7439–7447.
215. Zhan M, Li Y, Hu B, He X, Huang J, Zhao Y, Fu S, Lu L. Serum microRNA-210 as a predictive biomarker for treatment response and prognosis in patients with hepatocellular carcinoma undergoing transarterial chemoembolization. J Vasc Interv Radiol. 2014; 25:1279–1287.e1.
216. Yang L, Xu Q, Xie H, Gu G, Jiang J. Expression of serum miR-218 in hepatocellular carcinoma and its prognostic significance. Clin Transl Oncol. 2016; 18:841–847.
217. Li J, Wang Y, Yu W, Chen J, Luo J. Expression of serum miR-221 in human hepatocellular carcinoma and its prognostic significance. Biochem Biophys Res Commun. 2011; 406:70–73.
218. Zhuang LP, Meng ZQ. Serum miR-224 reflects stage of hepatocellular carcinoma and predicts survival. Biomed Res Int. 2015; 2015:731781.
219. Cui L, Hu Y, Bai B, Zhang S. Serum miR-335 Level is Associated with the Treatment Response to Trans-Arterial Chemoembolization and Prognosis in Patients with Hepatocellular Carcinoma. Cell Physiol Biochem. 2015; 37:276–283.
220. Yao H, Liu X, Chen S, Xia W, Chen X. Decreased expression of serum miR-424 correlates with poor prognosis of patients with hepatocellular carcinoma. Int J Clin Exp Pathol. 2015; 8:14830–14835.
221. Hu T, Li J, Zhang C, Lv X, Li S, He S, Yan H, Tan Y, Lei M, Wen M, Zuo J. The potential value of microRNA-4463 in the prognosis evaluation in hepatocellular. Genes & Diseases. 2017.
222. Bruix J, Gores GJ, Mazzaferro V. Hepatocellular carcinoma: clinical frontiers and perspectives. Gut. 2014; 63:844–855.
223. Forner A, Llovet JM, Bruix J. Hepatocellular carcinoma. Lancet. 2012; 379:1245–1255.
224. Dhanasekaran R, Limaye A, Cabrera R. Hepatocellular carcinoma: current trends in worldwide epidemiology, risk factors, diagnosis, and therapeutics. Hepat Med. 2012; 4:19–37.
225. Mazzola A, Costantino A, Petta S, Bartolotta TV, Raineri M, Sacco R, Brancatelli G, Cammà C, Cabibbo G. Recurrence of hepatocellular carcinoma after liver transplantation: an update. Future Oncol. 2015; 11:2923–2936.
226. Palmer DH, Johnson PJ. Evaluating the role of treatment-related toxicities in the challenges facing targeted therapies for advanced hepatocellular carcinoma. Cancer Metastasis Rev. 2015; 34:497–509.
227. Yates LA, Norbury CJ, Gilbert RJ. The long and short of microRNA. Cell. 2013; 153:516–519.
228. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Q, Li X, Wang W, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancerand other diseases. Cell Res. 2008; 18:997–1006.
229. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O'Briant KC, Allen A, Lin DW, Urban N, Drescher CW, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008; 105:10513–10518.
230. Mao B, Wang G. MicroRNAs involved with hepatocellular carcinoma (Review). Oncol Rep. 2015; 34:2811–2820.
231. Giordano S, Columbano A. MicroRNAs: new tools for diagnosis, prognosis, and therapy in hepatocellular carcinoma? Hepatology. 2013; 57:840–847.
232. Wong CM, Wong CC, Lee JM, Fan DN, Au SL, Ng IO. Sequential alterations of microRNA expression in hepatocellular carcinoma development and venous metastasis. Hepatology. 2012; 55:1453–1461.
233. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials. 2007; 8:16.

