INTRODUCTION
Non-small cell lung cancer (NSCLC), a leading neoplastic disease [1] with the highest incidence rate and cancer-related mortality worldwide, has a 5-year survival rate under 10% due to high metastasis rate and poor prognosis. Resection has been the main therapy by far [2].
Targeted therapy for oncogene has been an emerging therapeutic method in recent years, whereas patient survival rate has not been improved due to late detection and therapy. Thus it was essential to find cancer marker guidance of early discovery, diagnosis and treatment, which led to biomarker becoming one of the most valuable area in cancer research. NSCLC is hardly characterized by one marker for heterogeneous disorder with a variety of biomarker [3], making it hard to find effective NSCLC marker.
With rapid development of high-throughput biotechnology, biological data have been obtained from protein complex and gene expression profile, which helped to clarifying gene function and resource acquisition. Some scholars observed differentially expressed genes with microarray between Lung adenocarcinoma and adjacent normal tissue and established protein-protein interaction (PPI) network with gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) for enhancement of lung adenocarcinoma knowledge [4]. PPI search tool of network construction and interaction gene STRING database provided mutual information for experiment and prediction. To get gene-related information, studying topological features of disease gene product in PPI network has become a new way of gene predicting.
Closely related core genes could be found by means of gene interaction network starting with known NSCLC gene. PARP1 had a high connectivity, namely as a basic core, and directly attracted with typical oncogene EGFR and ALK; mutating EGFR in NSCLC constitutively activated encoding protein and caused the occurrence of abnormal function and constant signal transduction out of control, which was in correction with NSCLC development [5]; as a representative molecularly targeted agent, ALK inhibitor could activate recombination through mutation, gene amplification and chromosome abnormality to increase carcinogenic driver expression [6].
Furthermore, TCGA indicated that PARP1 was highly expressed in NSCLC, suggesting that PARP1 had a great influence on NSCLC. Coded protein in eukaryotic cells of PARP1 catalyzes poly-ADP-ribosylation and is involved in the process of DNA single-strand damage repair. The main achievement of PARP1 in cancer is stimulation for proliferation of colorectal cancerous cells, whereas there is no research on PARP1 functional mechanism in NSCLC.
The study firstly surveyed PARP1’s effect on NSCLC cells and discussed its functionary mechanism, which found that PARP1 was expressed strongly in metastatic NSCLC; and high PARP1 expression was relevant to prognosis and facilitated NSCLC cells migration and invasion, which provided not only corresponding foundation of experiment medicine for later targeted treatment studies but theory support of PARP1 expression within NSCLC and possible mechanism research.
RESULTS
The known genes of non-small cell lung cancer
OMIM (Online Mendelian Inheritance in Man) is a comprehensive database involved in human gene and genetic disease [7]. There were 3358 annotation of genes related to mutation phenotype of Mendelian inherited disease by March, 16th, 2015. Seven genes were identified from the search with the key word “non-small cell lung cancer”.
The database COSMIC (Catalogue of Somatic Mutations in Cancer) [8] embodied cancer-related gene extracted in high-throughput experimental data which was from the reported papers and cancer genome plan of Sanger laboratory. 19 genes were identified from the search with the key word “non-small cell lung cancer” in this database.
GAD (The Genetic Association Database) [9] is a comprehensive database collecting complex disease of human. The reported literature and pathogenic gene of complex disease was annotated in GWAS experimental data. 12 genes were identified from the search with the key word “non-small cell lung cancer” in this database.
The database DisGeNET [10] annotate pathogenic gene by integrating common database and gene-disease relationship in reported literature. Currently 381,056 gene-disease relationships included in this database (including 16,666 genes and 13,172 kinds of diseases). There are 39 genes were identified from the search with the key word “non-small cell lung cancer” in this database. In summary, 62 genes were identified through searching database in total (no repetition). The specific gene name and database source were included in the Supplementary Table 1.
Though many databases were developed to collect pathogenic gene, there were some differences in the data with disease phenotype covered and various orientation of genotype due to the data types of different database. Meanwhile, the collection of information from above database might be incomplete considering the lag characteristic in the maintaining process of database.
Thus, there was a need for pathogenic gene through artificial screening to further analyze in terms of specific disease phenotype. Detection of the relationship between gene and non-small cell lung cancer conducted in Pubmed, gene name and non-small cell lung cancer as the searched key words, that is (“gene symbol” or “gene”) and “non-small cell lung cancer”. The literature would be recorded if the two key words appeared both in title and abstract of the same literature, as the evidence of detection of the association between gene and non-small cell lung cancer. Finally, 81 pathogenic genes were gained from the literature, specific gene name included in the Supplementary Table 1.
Shortest path analysis of genes
A non-directional network chart with the PPI data gained from STRING was established, and the connection of their shortest path through Dijkstra's algorithm was calculated (Figure 1). Then the Betweenness value of internal node in these paths was counted. 385 genes whose Betweenness value over 0 were gained, and these genes were listed out in the Supplementary Table 2. Permutation test was employed to check and calculate their permutation FDR in terms of further selection of these genes, similarly the results showed in the Supplementary Table 2.
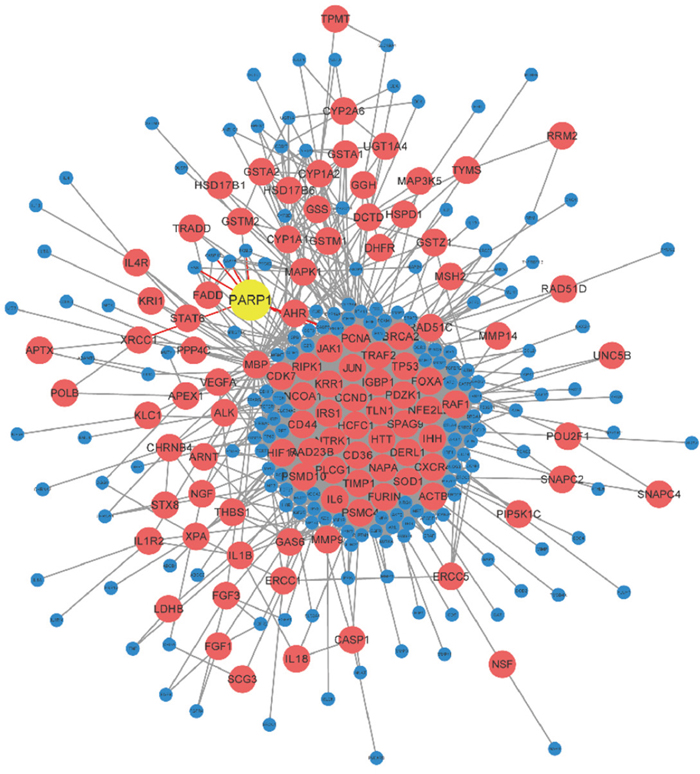
Figure 1: Network figure of NSCLC genes. The PPI data of NSCLC was gained from STRING, and the connection of their shortest path was calculated through Dijkstra's algorithm. It indicated that PARP1 involved in the occurrence and development of NSCLC.
Among them, permutation FDR of 103 genes were less than 0.05. There was a strong correlation between these genes and NSCLC, which could be used in the subsequent analysis. A gene PARP1 with high betweenness value except for the famous oncogene with high connected degree was found, which indicated it played at a relatively core role in NSCLC gene network, yet it was unknown of the role of this gene in lung cancer.
There was direct gene connections between PARP1 and the famous NSCLC genes, EGFR and ALK. Thus it was possible that PARP1 involved in the occurrence and development of NSCLC. Therefore, functional experiment was further adopted to detect the effect of PARP1 on NSCLC.
The high expression of PARP1 in non-small cell lung cancer and metastatic non-small cell lung cancer
The expression of PARP1 in lung cancer with the expression data of TCGA was detected firstly. The data of squamous cell carcinoma and adenocarcinoma in TCGA indicated significant higher expression of PARP1 in tumor tissues (Figure 2).
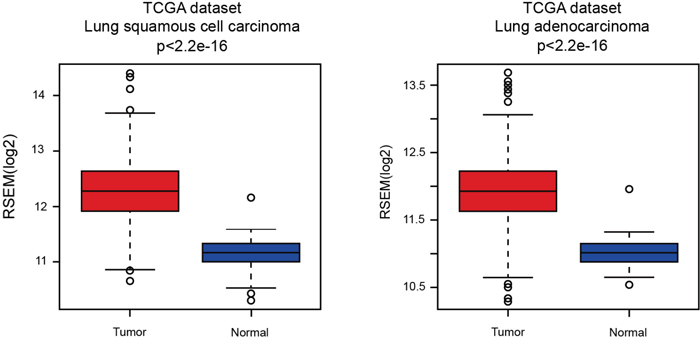
Figure 2: Expression of PARP1 in lung cancer. The expression of PARP1 of squamous cell carcinoma (left) and adenocarcinoma (right) was gained from TCGA. The data of squamous cell carcinoma and adenocarcinoma in TCGA indicated significant higher expression of PARP1 in tumor tissues.
Next the detection was conducted in the 75 patients with NSCLC disease. Among the 75 patients (40 patients of non-metastatic NSCLC and 35 patients of metastatic NSCLC) no differences was shown in their age, sex and smoking status.
The PARP1 expression in serum samples was done by q-PCR, which indicated that the expression of PARP1 in metastatic NSCLC was significantly higher than that in non-metastatic NSCLC in blood samples (Figure 3).
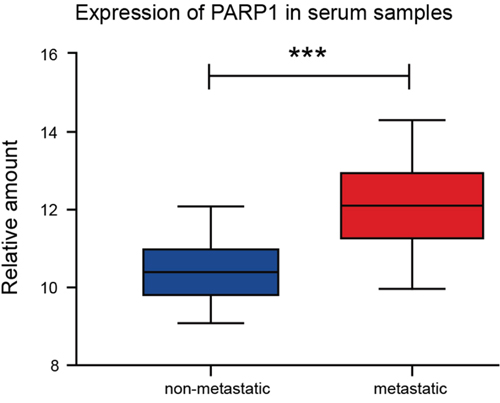
Figure 3: Higher expression of PARP1 in the serum samples ofmetastatic-NSCLC patients. Q-PCR was used to detect the PARP1 expression of serum samples of 75 NSCLC patients. The results were analyzed using t-test. *p < 0.05; **p < 0.01; ***p < 0.001. Red, metastatic group; blue, non-metastatic group.
The high expression of PARP1 in metastatic non-small cell lung cancer
We explored the expression of PARP1 in blood samples and tissue samples of NSCLC which showed a high consistency in its expression of blood and tissue samples (Figure 4A). It was indicated that the expression of PARP1 in blood could instruct the expression in tissue.
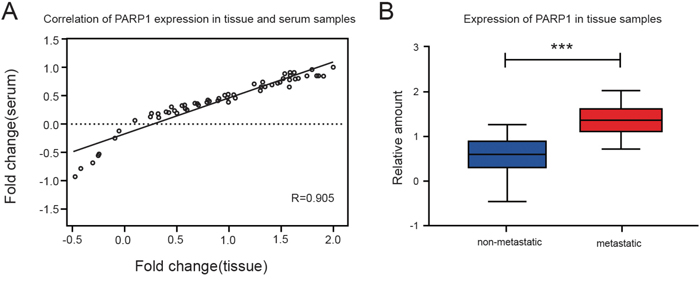
Figure 4: The expression of PARP1 in the tissue samples of NSCLC patients. Q-PCR was used to detect the PARP1 expression of tissue samples of 75 NSCLC patients. (A) The PARP1 expression of Pearson correlation scatter diagram in the matching tissue of 75 NSCLC patients and serum samples. A high consistency was found in its expression of blood and tissue samples. (B) The expression of PARP1 in metastatic NSCLC was higher than that in the non-metastatic NSCLC tissue. The results were analyzed using t-test. *p < 0.05; **p < 0.01; ***p < 0.001. Red, metastatic group; blue, non-metastatic group.
The results indicated that the expression of PARP1 in metastatic NSCLC tissues was higher (columnar section) than that in metastatic NSCLC tissue and non-metastatic NSCLC tissue (Figure 4B).
Efficiency of PARP1 silence and overexpression in NSCLC cell line
A549, H1975 and PC9 were chosen to represent the typical NSCLC cell line in this study. Function analysis was conducted in vitro cell line to study the effects of PARP1 on NSCLC invasion by adopting overexpression and inhibition of PARP1 expression. The overexpression and silence cell line of PARP1 gene with NSCLC cell line A549, H1975 and PC9 was established, then the PARP1 expression was detected in the 3 cell lines.
The q-PCR results showed that the expression of PARP1 in silence cell line decreased 70-80% compared with that of the control group, and the expression of PARP1 in overexpression cell line increased (Figure 5).
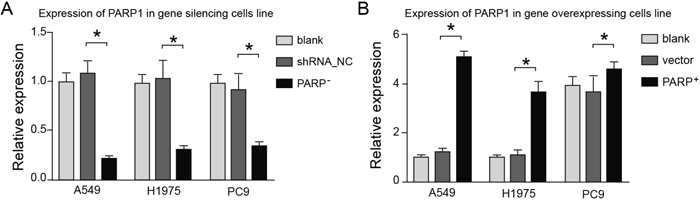
Figure 5: The effects of overexpressing or silencing plasmids on the PARP1 expression. The expression of PARP1 in silence cell line decreased 70-80% compared with that of the control group (left), whereas the expression of PARP1 in overexpression cell line increased significantly (right).
Promoting effect of PARP1 on proliferation, migration and invasion of NSCLC cell
Higher expression of PARP1 was showed in NSCLC patients and metastatic NSCLC patients in the above study, which indicated PARP1 involved in the metastatic process of NSCLC. Thus, PARP1 was necessary for the cell proliferation, migration and invasion during the metastatic process.
Thus, firstly the effects of PARP1 on the cell proliferation of A549, H1975 and PC9 were tested through MTT assays. The result indicated that PARP1 gene silence could significantly inhibit the cell growth of A549, H1975 and PC9, while PARP1 overexpression promoted the growth of A549 cell significantly (Figure 6). Secondly, cell scratch test would be conducted to identify whether PARP1 could promote the metastatic capacity of NSCLC. The result showed that PARP1 silence inhibited cell migration significantly shown in cell scratch test, while PARP1 overexpression promoted migration of NSCLC cell significantly (Figure 7). Finally, trans-well assays (Figure 8) was adopted to confirm these results, with the same conclusion drawn.
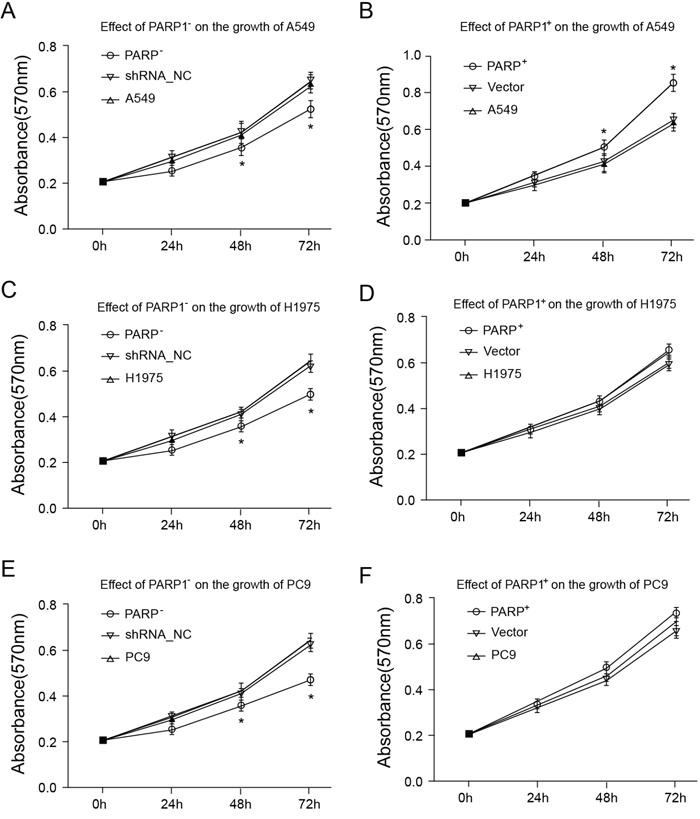
Figure 6: Cell proliferation was assessed with an MTT assay. 2 × 104/ml cell suspension fluid was inoculated in 96-well plates, and cultured for 96 h, respectively. Following that, the plates were added with MTT solution, and incubated for 4 h, then added with DMSO to fully resolve MTT pyrolysis products. Optical density (OD) was measured by immunoassay analyzer at the wavelength of 570 nm. (A, B) Effect of PARP1 on the growth of A549; (C, D) effect of PARP1 on the growth of H1975; (E, F) effect of PARP1 on the growth of PC9.
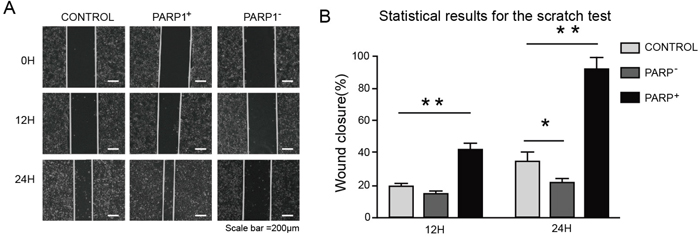
Figure 7: Promoting effect of PARP1 on migration of NSCLC cell. For the scratch test, cells were plated at 2×105 cells/well in a 6-well plate and grown overnight under standard conditions. After 12 h, the confluent monolayer was scratched manually with a plastic 200 μl pipette tip and after washing with PBS. Then the culture was continued in 2% serum at 37 °C for 24 h. The wound area were imaged using an inverted microscope at 100× magnification. The distance was measured using ImageJ2x. (A) Images of wounds of A549 cells transfected with the indicated plasmids taken 0, 12 and 24 h afer the wounds were inflicted (× 100). (B) Overexpressing PARP1 promoted the wound-healing process in A549 cells, whereas PARP1 silencing did not contribute to wound healing. Wound-healing results for A549 cells transfected with the indicated plasmids (analyzed by t-tests). *p < 0.05; **p < 0.01;***p < 0.001.
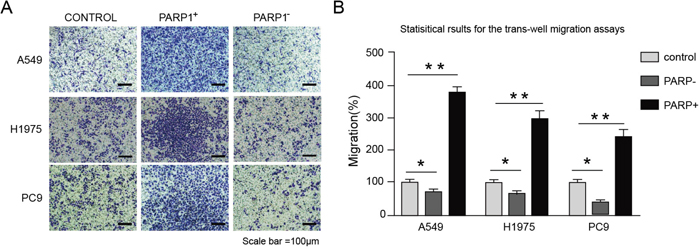
Figure 8: Overexpressing PARP1 promotes the metastasis of NSCLC cells lines, whereas PARP1 silencing inhibited the processes. 1 × 105 cells were plated onto 24-well cell chambers (pore size: 8 μm). The cells were plated in medium without serum, and medium with 20% serum was used as a chemoattractant. The cells were incubated at 37 °C for 16 h. Cells in 10 random fields of view at 100× magnification were counted. Images were acquired using an inverted phase-contrast microscope at 100× magnification. (A) Images from the Trans-well migration assays of A549, H1975 and PC9 cells transfected with indicated plasmids (× 100). (B) Statistical results for the trans-well migration assays of A549, H1975 and PC9 cells transfected with indicated plasmids (analyzed by t-tests); %migration = [mean number of cells invading the membrane/mean number of cells migrating through the control insert membrane] × 100. *p < 0.05; **p < 0.01; ***p < 0.001.
PARP1 as prognostic gene
The survival relationship between PARP1 and patients of NSCLC was analyzed with the data in GEO database. Kaplan-meier survival curve was made using the online survival analysis tool, PROGgene. It was found that the expression of PARP1 was related to the prognosis or recurrence of patients through Kaplan-meier survival curve: The overall survival (OS) in PARP1 high expression group was significantly lower than that in PARP1 low expression group (Figure 9).
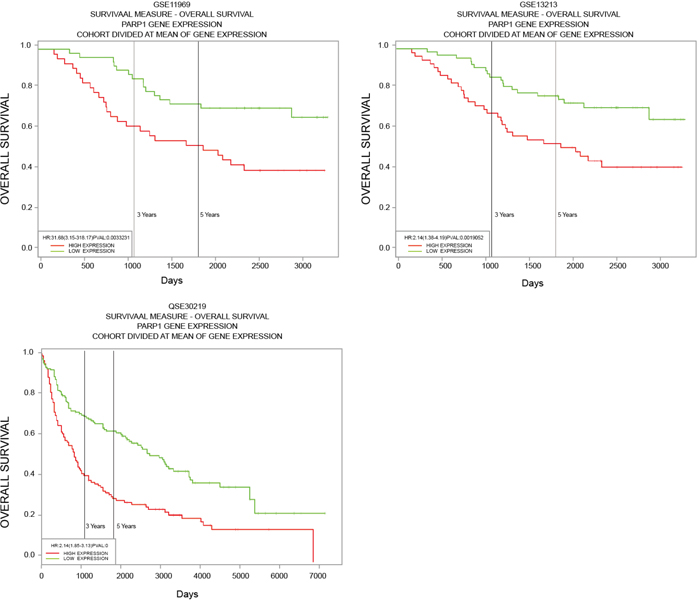
Figure 9: Survival relationship between PARP1 and patients of NSCLC. Data from GSE11969, 13213 and 30219 was gained from GEO database, then relationship between expression of PARP1 and the prognosis or recurrence of patients was tested through Kaplan-Meier survival curve using the online survival analysis tool, PROGgene.
DISCUSSION
Complex diseases were generally affected by the interaction of multiple genes [11]. Generally, the common influence of interaction in interacting genes could not only decrease the complex degree of biological network, but also benefit to explore meaningful biological information for the researchers, which further provide scientific basis for therapy and study of disease. Previous studies indicated that many interacting genes had similar or same effects, or involved in the same biochemical pathways. Therefore, the known key gene in an established interaction network of genes would be found easily, and the functions of interacting genes can be preliminary predicted and annotated based on this gene. In this study, NSCLC-related gene was searched from database OMIM, COSMIC, GAD and DisGeNET, then a network chart was established with the adoption of PPI data from STRING, followed by the shortest path analysis and Permutation test. There is a total of 385 genes whose Betweenness value was greater than 0 gained through Dijkstra's algorithm. Permutation test showed 103 genes whose Permutation FDR was less than 0.05. There was a strong correlation between these genes and NSCLC, which could be used in the subsequent analysis.
Almost all the NSCLC patients would recurred in different stages, and finally suffered from disease metastasis and death [12]. Even though there was a new development of immunotherapy and targeted therapy of tumors, once the systematic metastasis of cancer cell occurred, the five-year survival rate would decrease at lessby 10% [13]. For that it was still a challenge to expand the survival rate of NSCLC patients by inhibiting the cell migration of cancer cells. Therefore, it was of significance for the study of molecular markers and potential therapeutic small molecules in clinical stage. PARP1 played an important role in DNA damage repair, especially in base excision repair (BER) pathway [14]. It was further verified that PARP1 played a key role in DNA repair [15–17], which made PARP1 a valuable therapeutic target in the research of disease drug. Recently, it was found that PARP1 presented high expression in a series of cancers, which provided a high-value target for the study of tumor detection, stage and biological characteristics [18–23]. The overexpression of PARP1 results from much DNA damage in cancer cells with unstable genes, instead of the induction of the activated specific carcinogenic pathway [24]. Byers, et al. demonstrated that PARP1 could be a new target molecule for NSCLC by combined transcriptome and proteome analysis [25]. Meanwhile, a new study claimed that PARP1 could promote the metastasis and recurrence of lung adenocarcinoma towards brain and bones by regulating several steps in the metastasis process, which was not related to DNA repair. There were several ways of PARP1 enhancing the metastasis of lung adenocarcinoma towards brain: promoting the invasion of lung adenocarcinoma cells, anoikis resistance, exosmosis, and self-updating, as well as regulating cerebral microenvironment [26].
In the current study, the expression of PARP1 in the blood samples showed a high consistency with that in the tissue samples, which indicated that the expression of PARP1 in blood could instruct the expression in tissue. Because of that, the related information about the development of tumor could be obtained through the detection of PARP1 expression in blood at the early stage of NSCLC tumor. The results of MTT assays indicated that lockdown of PARP1 gene significantly inhibited the cell growth of A549, H1975 and PC9, while PARP1 overexpression promoted the growth of A431 cell significantly. It indicated that PARP1 gene promoted the proliferation of NSCLC, whose role was consistent in others malignant tumor [27, 28]. There were many extensive researches on the gene which influence NSCLC cell migration, mainly including MALAT-1 [29], MTA1 [30], BRAF [31], EGFR and so on. However, the role of PARP1 in NSCLC migration remains still unknown. Our experiments revealed that: expression of PARP1 was upregulated in NSCLC and showed higher expression in metastatic NSCLC; PARP1 silence inhibited cell migration significantly, while PARP1 overexpression promoted NSCLC cell migration significantly [26]. It has demonstrated that knockout or inhibition of PARP1 could decrease the migration of lung adenocarcinoma significantly, which indicated PARP1 was involved in the migration of NSCLC. In addition, there was a direct association between PARP1 and the key cancer gene of NSCLC: EGFR, ALK in the interaction network of genes. Previous studies have suggested that EGFR takes part in DNA repair through PI3K signaling pathway [32–33]. In the cells with ALK fusion, it has illustrated that dimers can be activated abnormally by aberrant receptors, which can activate the downstream signaling pathway PI3K-AKT and drive aberrant proliferation of tumor cells [34–35]. Based on these findings, it can be see that ALK and EGFR after requiring functional mutation could drive the growth of tumor cells via PI3K-AKT pathway. So we hypothesized that the effects of PARP1 on NSCLC tumor migration may be associated with the PI3K-AKT pathway. However, the mechanism needs further exploration.
It is of directive significance to understand the effect of prognostic factors on the prediction, diagnosis, and treatment of diseases. According to the study, PARP1 may be an independent prognostic factor of multiple malignant tumors. And PARP1 Val762Ala polymorphism may be an independent prognostic factor for cervical cancer, which might play a role in the prediction of clinical results [36]. Collectively, the expression of PARP1 might be considered as an independent prognostic factor for the patients of breast cancer [37]. While in the limited stage of SCLC, the expression of PARP1 was related to the progression free survival of tumor cells [38]. The COX regression analysis conducted by KJ Xie, et al. [39] indicated that PARP1 was an independent adverse prognostic factor of NSCLC patients. According to Kaplan-meier survival curve, the higher expression of PARP1, the lower survival rate of patients, which preliminary suggested that PARP1 might be an independent adverse prognostic factor of NSCLC. Consistently with the above result we found that PARP1 presented high expression in NSCLC, and the survival rate of patients was decreased accompanied by increased PARP1 expression. In summary, based on gene interaction network, the shortest paths method was adopted in NSCLC gene network to find a number of candidate oncogenes of NSCLC. We conducted the functional verification on PARP1, which provided convenience for the following discussion and study of the mechanisms related to the occurrence and development of NSCLC. We found the role of PARP1 in the migration and prognosis of NSCLC, laying foundations for the following clinical experiments, and providing theoretical basis for revealing the effects of PARP1 on NSCLC.
Based on the known NSCLC genes, this study found novel NSCLC gene PARP1 by using gene interaction network. and verified the promoting effect of PARP1 gene on the migration of non-small cell lung cancer.
MATERIALS AND METHODS
Network construction and shortest path algorithm
Data collection: taking “non-small cell lung cancer” as key word to retrieve and gain NSCLC-related gene from database of OMIM (Online Mendelian Inheritance in Man) (Rashbass, 1995), COSMIC (Catalogue of Somatic Mutations in Cancer) (Forbes et al., 2011) and GAD (The Genetic Association Database) (Becker, Barnes, Bright, & Wang, 2004).
Then the data acquired from STRING (version 9.0) (http://string.embl.de/) was used to build up initial weighted PPI network diagram in which all connected lines weight depended separately on confidence levels of interdependency between two genes.
We could make sure the shortest path of each pair of correlatively known gene of lung cancer and genes on the path by Dijkstra's algorithm: calculating optimal path of one node to all the other nodes, moreover, arraying these genes by Betweenness value that stood for internal nodes of certain gene contained all optimal paths found.
Permutation test
No matter what kind of gene was included in our gene network and how the real node or gene was distributed, it was always easy to find the genes with high Betweenness value for some Betweenness value might sway by network basically structural models, yet these universal genes was not our wanted genes found from special genetic network. So we adopted permutation test to further screen NSCLC genes with optimal path in specific net.
From gene network, we randomly selected simulated genes involved with a number of NSCLC-associated genes and then calculated the shortest path within genes (a total of 500 simulations); after we counted once, the Betweenness value in stimulation was bigger than that of actual one. After 500 randomized trials, each gene would get a frequency that was defined as permutation FDR of shortest-path gene of which small value (FDR < 0.05) meant a significantly NSCLC-involved gene of the shortest path.
Patients and clinical samples
There were serum and tumor biopsy samples of 75 patients with NSCLC, including 40 metastatic and 35 non-metastatic, staging by pathological diagnosis and International Association for the Study of Lung Cancer (IASLC) TNM Classification.
5 ml peripheral blood was drawn and put in vascular separator after admission. The serum was extracted within 2h and kept at -80°C.
Inclusive criteria: the experiment have met with medical ethics of Baoji Central Hospital approval; all patients without treatment before a month have sighed informed consent and got ECOG score ranging from 0-1 during hospitalization.
Total RNA extraction
Total RNA from 8 cell lines, blood samples, 75 cancer tissues of NSCLC and the adjacent normal tissues were extracted in TRIZOL RNA extraction reagent (Invitrogen) by manual. The experiment was performed as follows: the tissues were added with 500 ul Trizol and 200 ul chloroform in order, after vibrating and standing, then were placed into centrifugal machine for 10 min, mixing with isopropanol and ethanol to discard supernatant by centrifugation; following that, the tissues were added with 20-50 ul DEPC-ddH2O to dissolving dissolve RNA using 20-50ul DEPC-ddH2O and preserved saving with absolute ethanol in refrigerator at -70°C.
Reverse transcription PCR and QPCR
SuperScript III First-Strand Synthesis System kit (Invitrogen) was used to synthesize cDNA, with the detailed processes shown in brochure. The fluorescent primers were designed using primer premier 5.0 according to the sequence of genes. Real-time PCR measurements were performed on CFX96 (Bio-Rad). Each PCR reaction contained SYBR® Premix Ex Taq (2x) 12.5 μl, 10 μmol/L upstream and downstream primers each 1 ul, cDNA template 2 μl, dH2O 10μl. The temperature process was 95°C for 30 sec followed by 40 cycles of amplification (95°C for 5 sec, 60°C for 30 sec, and 72°C for 30 sec). 2-ΔΔt was adopted for date processing.
NSCLC cell lines and culture of cells
NSCLC cell lines A549, H1975 and PC9, purchased from The Shanghai Institutes for Biological Sciences of the Chinese Academy of Sciences, were taken out and inoculated to RPMI-1640 culture solution complementary with 10% fetal bovine serum, and cultured at under 37°C with a humidity of 5%CO2 saturation, with liquid exchanging and passaging every 3-4 days.
Construction of PARP1 gene silencing and overexpression of stable transgenic lines
Completed PARP1 cDNA would be acquired with primers (Table 1), and then we cloned completed PARP1 cDNA into pEGFP-N2vector to construct gene overexpression plasmid. As for the PARP1 gene silencing, shRNA and negative-control shRNA (shRNA-NC) of PRAP1 were designed according the sequence of PARP1 (NM_007415) from GenBank (Table1). After annealing, shRNA of PRAP1 were ligated into pGPU6/GFP/Neo vector to build gene silencing recombinant plasmid. And then recombinant vectors were transfected into cell lines by liposome Lipofectamine 3000. Meanwhile, the blank vector transfection group was taken as the control group. After transfection for 2 days, there were part of green fluorescence cells which were converted to use 400 μg/ml G418 DMEM medium, screened for a month to attain gene silencing and overexpression stable transgenic lines, culture and cryopreserved in DMEM + 10% FBS + P.S. medium.
Table 1: Primers used for the detection and vector construction of PARP1
Usage |
Forward |
Reverse |
|---|---|---|
qRT-PCR |
GCAGAGTATGCCAAGTCCAACAG |
ATCCACCTCATCGCCTTTTC |
cDNA |
GAAGATCTTCGCCATG GCGGAGTCTTCGGATAAGCT |
CCGCTCGAGCGGCCA CAGGGAGGTCTTAAAAT |
shRNA |
CACCGGACCAAGTGTATGGTCAAGATTCA AGAGATCTTGACCA TACACTTGGTCCTTTTTTG |
GATCCAAAAAAGGACCAA GTGTATGGTCAAGATCTCTTGA ATCTTGACCATACACTTGGTCC |
shNC |
CACCGTTCTCCGAACGTGT CACGTCAAGAGATTACGTGA CACGTTCGGAGAATTTTTTG |
GATCCAAAAAATTCTCCGAACGTGTCACGT AATCTCTTGACGTGACACGTTCGGAGAAC |
Cell proliferation detection with MTT method
2 × 104/ml cell suspension fluid was inoculated in 96-well plates, with each well having 3 parallel wells and about 100 μl fluid, and cultured in an incubator with 5% CO2 at 37°C to 24, 48, 72, 96 h, respectively. Following that, the plates were taken out, added with 50 μl MTT solution each well that was configured into 1 g/L with normal saline, and incubated at 37 °C for 4 h to discard the supernatant fluid, then added with dimethylformamide (DMSO) (100 μl per well), shaken for 10 min to fully resolve MTT pyrolysis products. Optical density (OD) was measured by immunoassay analyzer at the wavelength of 570 nm, drawing cell growth curve.
The scratch test and trans-well experiment for cell migration detection
For the scratch test, cells were plated at 2×105 cells per well in a 6-well plate and grown overnight under standard conditions. After 12 h, the confluent monolayer was scratched manually with a plastic 200 μl pipette tip and after washing with PBS. Then the culture was continued in 2% serum at 37°C for 24 h. Cells that had migrated into the wound area were imaged using an inverted microscope at 100× magnification (Olympus, CKX41). The distance was measured using ImageJ2x.
Trans-well experiment was carried out manually. In brief, 1 × 105 cells were plated onto 24-well cell chambers with a non-coated membrane (pore size: 8 μm, BD Biosciences). The cells were plated in medium without serum, and medium with 20% serum was used as a chemoattractant in the lower chamber. The cells were incubated at 37 °C for 16 h, and cells that did not migrate through the pores were removed with a cotton swab. Cells on the lower surface of the membrane were fixed with 100% methanol for 5 min and stained with 0.1% crystal violet for 2 min (Sigma). Cells in 10 random fields of view at 100× magnification were counted and expressed as the average number of cells per field of view. Images were acquired using an inverted phase-contrast microscope at 100× magnification (Olympus, CKX41).
Statistical analysis
All statistical analyses were performed using SPSS 21.0 statistical software. Measurement data were presented by mean ± standard deviation (AV ± SD). Unpaired Student's t-test for parametric data or Mann-Whitney rank sum test for nonparametric data were applied to analyze mean value between two groups. Chi-square tests were applied to determine count data expressed in n (%). In all cases, p < 0.05 was considered statistically significant.
Abbreviations
PARP1: poly (ADP-ribose) polymerase 1; NSCLC: non-small cell lung cancer; GO: gene ontology; KEGG: Kyoto encyclopedia of genes and genomes; OMIM: Online Mendelian Inheritance in Man; COSMIC: Catalogue of Somatic Mutations in Cancer; GAD: The Genetic Association Database; IASLC: International Association for the Study of Lung Cancer.
Author contributions
Kai Chen and Yajie Li participated in the conception and design of the study. JL, Hui Xu and Chunfeng Zhang performed the analysis and interpretation of data. MYL, Zhiqiang Li and Wei Wang contributed to drafting the article. Baofeng Wang and Kai Chen revised it critically for important intellectual content. Yajie Li is the GUARANTOR for the article who accepts full responsibility for the work and the conduct of the study, had access to the data, and oversaw the decision to publish.
ACKNOWLEDGMENTS
We would like to give our sincere appreciation to the reviewers for their helpful comments on this article.
CONFLICTS INTEREST
The authors have declared that no competing interests exist.
FUNDING
None.
REFERENCES
1. Reck M, Heigener DF, Mok T, Soria JC, Rabe KF. Management of non-small-cell lung cancer: recent developments. Lancet. 2013; 382:709-719.
2. Coleman MH, Bueno R. Role of adjuvant chemotherapy in NSCLC (stages I to III). Surg Oncol Clin N Am. 2011; 20:757-767.
3. Socinski MA, Pennell NA. Best practices in treatment selection for patients with advanced NSCLC. Cancer Control. 2016; 23:2-14.
4. Xu H, Ma J, Wu J, Chen L, Sun F, Qu C, Zheng D, Xu S. Gene expression profiling analysis of lung adenocarcinoma. Braz J Med Biol Res. 2016; 49.
5. Kuykendall A, Chiappori A. Advanced EGFR mutation-positive non-small-cell lung cancer: case report, literature review, and treatment recommendations. Cancer Control. 2014; 21:67-73.
6. Shaw AT, Kim DW, Mehra R, Tan DS, Felip E, Chow LQ, Camidge DR, Vansteenkiste J, Sharma S, De Pas T, Riely GJ, Solomon BJ, Wolf J, et al. Ceritinib in ALK-rearranged non-small-cell lung cancer. N Engl J Med. 2014; 370:1189-1197.
7. Hamosh A, Scott AF, Amberger J, Bocchini C, Valle D, McKusick VA. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 2002; 30:52-55.
8. Forbes SA, Bindal N, Bamford S, Cole C, Kok CY, Beare D, Jia M, Shepherd R, Leung K, Menzies A, Teague JW, Campbell PJ, Stratton MR, Futreal PA. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 2011; 39:D945-950.
9. Becker KG, Barnes KC, Bright TJ, Wang SA. The genetic association database. Nat Genet. 2004; 36:431-432.
10. Bauer-Mehren A, Rautschka M, Sanz F, Furlong LI. DisGeNET: a cytoscape plugin to visualize, integrate, search and analyze gene-disease networks. Bioinformatics. 2010; 26:2924-2926.
11. Mackay TF. Quantitative trait loci in Drosophila. Nat Rev Genet. 2001; 2:11-20.
12. Herbst RS, Prager D, Hermann R, Fehrenbacher L, Johnson BE, Sandler A, Kris MG, Tran HT, Klein P, Li X, Ramies D, Johnson DH, Miller VA. TRIBUTE: a phase III trial of erlotinib hydrochloride (OSI-774) combined with carboplatin and paclitaxel chemotherapy in advanced non-small-cell lung cancer. J Clin Oncol. 2005; 23:5892-5899.
13. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015; 65:5-29.
14. De Vos M, Schreiber V, Dantzer F. The diverse roles and clinical relevance of PARPs in DNA damage repair: current state of the art. Biochem Pharmacol. 2012; 84:137-146.
15. Bouchard VJ, Rouleau M, Poirier GG. PARP-1, a determinant of cell survival in response to DNA damage. Exp Hematol. 2003; 31:446-454.
16. Rouleau M, Patel A, Hendzel MJ, Kaufmann SH, Poirier GG. PARP inhibition: PARP1 and beyond. Nat Rev Cancer. 2010; 10:293-301.
17. Ellisen LW. PARP inhibitors in cancer therapy: promise, progress, and puzzles. Cancer Cell. 2011; 19:165-167.
18. Bieche I, de Murcia G, Lidereau R. Poly (ADP-ribose) polymerase gene expression status and genomic instability in human breast cancer. Clin Cancer Res. 1996; 2:1163-1167.
19. Chow JP, Man WY, Mao M, Chen H, Cheung F, Nicholls J, Tsao SW, Li Lung M, Poon RY. PARP1 is overexpressed in nasopharyngeal carcinoma and its inhibition enhances radiotherapy. Mol Cancer Ther. 2013; 12:2517-2528.
20. Dziaman T, Ludwiczak H, Ciesla JM, Banaszkiewicz Z, Winczura A, Chmielarczyk M, Wisniewska E, Marszalek A, Tudek B, Olinski R. PARP-1 expression is increased in colon adenoma and carcinoma and correlates with OGG1. PLoS One. 2014; 9:e115558.
21. Galia A, Calogero AE, Condorelli R, Fraggetta F, La Corte A, Ridolfo F, Bosco P, Castiglione R, Salemi M. PARP-1 protein expression in glioblastoma multiforme. Eur J Histochem. 2012; 56:e9.
22. Salemi M, Galia A, Fraggetta F, La Corte C, Pepe P, La Vignera S, Improta G, Bosco P, Calogero AE. Poly (ADP-ribose) polymerase 1 protein expression in normal and neoplastic prostatic tissue. Eur J Histochem. 2013; 57:e13.
23. Green AR, Caracappa D, Benhasouna AA, Alshareeda A, Nolan CC, Macmillan RD, Madhusudan S, Ellis IO, Rakha EA. Biological and clinical significance of PARP1 protein expression in breast cancer. Breast Cancer Res Treat. 2015; 149:353-362.
24. Ossovskaya V, Koo IC, Kaldjian EP, Alvares C, Sherman BM. Upregulation of poly (ADP-Ribose) polymerase-1 (PARP1) in triple-negative breast cancer and other primary human tumor types. Genes Cancer. 2010; 1:812-821.
25. Byers LA, Wang J, Nilsson MB, Fujimoto J, Saintigny P, Yordy J, Giri U, Peyton M, Fan YH, Diao L, Masrorpour F, Shen L, Liu W, et al. Proteomic profiling identifies dysregulated pathways in small cell lung cancer, novel therapeutic targets including PARP1. Cancer Discov. 2012; 2:798-811.
26. Choi EB, Yang AY, Kim SC, Lee J, Choi JK, Choi C, Kim MY. PARP1 enhances lung adenocarcinoma metastasis by novel mechanisms independent of DNA repair. Oncogene. 2016; 35:4569-4579.
27. Xu KW, Qin CJ, Chen ZH, Song XM. AB38. Expression of PARP1 in human colorectal cancer and its relationship with tumor cell proliferation and apoptosis of SW620 cells. Transl Gastrointest Cancer. 2015.
28. Wu WQ, Kong ZZ, Zhu HL, Duan XL, Li SJ. Effects and underlying mechanism of siRNA targeting PARP1 on the proliferation of PC3 cells. Chin J Urol. 2013:130-134.
29. Gutschner T, Hammerle M, Eissmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, Zornig M, MacLeod AR, Spector DL, Diederichs S. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013; 73:1180-1189.
30. Li Y, Chao Y, Fang Y, Wang J, Wang M, Zhang H, Ying M, Zhu X, Wang H. MTA1 promotes the invasion and migration of non-small cell lung cancer cells by downregulating miR-125b. J Exp Clin Cancer Res. 2013; 32:33.
31. Sun M, Liu XH, Wang KM, Nie FQ, Kong R, Yang JS, Xia R, Xu TP, Jin FY, Liu ZJ, Chen JF, Zhang EB, De W, Wang ZX. Downregulation of BRAF activated non-coding RNA is associated with poor prognosis for non-small cell lung cancer and promotes metastasis by affecting epithelial-mesenchymal transition. Mol Cancer. 2014; 13:68.
32. Golding SE, Morgan RN, Adams BR, Hawkins AJ, Povirk LF, Valerie K. Pro-survival AKT and ERK signaling from EGFR and mutant EGFRvIII enhances DNA double-strand break repair in human glioma cells. Cancer Biol Ther. 2009; 8:730-738.
33. Fischer OM, Hart S, Gschwind A, Ullrich A. EGFR signal transactivation in cancer cells. Biochem Soc Trans. 2003; 31:1203-1208.
34. Yang L, Li G, Zhao L, Pan F, Qiang J, Han S. Blocking the PI3K pathway enhances the efficacy of ALK-targeted therapy in EML4-ALK-positive nonsmall-cell lung cancer. Tumour Biol. 2014; 35:9759-9767.
35. Seo M, Kim JH, Suk K. Role of the p55-gamma subunit of PI3K in ALK-induced cell migration: RNAi-based selection of cell migration regulators. Cell Adh Migr. 2017; 11:205-210.
36. Nogueira A, Assis J, Faustino I, Pereira D, Catarino R, Medeiros R. Base excision repair pathway: PARP1 genotypes as modulators of therapy response in cervical cancer patients. Biomarkers. 2016:1-7.
37. Mazzotta A, Partipilo G, De Summa S, Giotta F, Simone G, Mangia A. Nuclear PARP1 expression and its prognostic significance in breast cancer patients. Tumour Biol. 2016; 37:6143-6153.
38. Kim HC, Song JS, Lee JC, Lee DH, Kim SW, Lee JS, Kim WS, Rho JK, Kim SY, Choi CM. Clinical significance of NQO1 polymorphism and expression of p53, SOD2, PARP1 in limited-stage small cell lung cancer. Int J Clin Exp Pathol. 2014; 7:6743-6751.
39. Xie KJ, He HE, Sun AJ, Liu XB, Sun LP, Dong XJ. Expression of ERCC1, MSH2 and PARP1 in non-small cell lung cancer, prognostic value in patients treated with platinum-based chemotherapy. Asian Pac J Cancer Prev. 2014; 15:2591-2596.

