INTRODUCTION
The majority of humans have experienced mild to moderate pruritus. Mosquito biting evoked-acute itching is one of the most common types of moderate pruritus. Some patients with serious skin diseases such as atopic dermatitis, advanced liver diseases or renal diseases, will experience moderate to severe itching. Obstinate pruritus is a pathological condition that affects skin sensory processing [1–4]. The mechanisms responsible for intractable pruritus are poorly understood, but refractory itching seems to be enhanced by a state of spinal hypersensitivity. It was reported that the spinothalamic tract implicated the spinal cord as a critical site for scratch-evoked suppression of itch [5–8]. Study of Sun et al has indicated that the gastrin-releasing peptide receptor (GRPR) in the dorsal horn of the spinal cord represent the first molecule that is dedicated to mediating the itch sensation [9]. It is well-known that GRPR-positive neurons in the spinal cord constitute a long-sought labeled line for itch not pain sensation [10]. In our previous research, the results of proton nuclear magnetic resonance (1H-NMR) studies of the spinal cord extracts revealed that the metabolites showed very different patterns for pruritogen and algogenic substance [11]. Given the high prevalence of itch in some patients with skin diseases [12–14], it is pivotal to know whether the genes are dysregulated in the spinal cord in different itch models.
Previous studies from pain data indicate that identification of whole genome messenger RNA (mRNA) expression profiles provide insight into the widespread factors that contribute to the development of more effective treatments for intractable pain [15–17]. Analysis using expression microarrays has revealed that many genes are regulated following noxious stimuli such as pruritus. In the present study, whole genome mRNA expression profiles in spinal cords of mice with compound 48/80 evoked-itching were analyzed to identify as unique target genes. Further, these candidate genes were examined in different itch models so as to provide central therapeutic targets for antipruritic drug development.
RESULTS
mRNA expression profiling revealed itch-related genes in lower cervical spinal cord from compound 48/80-evoked pruritus mice
To identify altered genes that might contribute to spinal hypersensitivity of acute itch, we conducted itch mRNA profiling experiment on mouse acute itching models by compound 48/80. The mRNA expressions of lower cervical spinal cord (C5-C8) of the animals were examined using Affymetrix Mouse Genome 430 2.0 microarrays that include 10,000 probe sets. The gene expression profiles in the 48/80 group were compared with the corresponding data of control group. We identified that a total of 15 probe sets were differentially expressed between control group and 48/80 group by the microarray data analysis, in which 9 probe sets were up-regulated and 6 probe sets were down-regulated (Figure 2). The maximal and minimal fold change was 5.16 and 2.01, respectively.
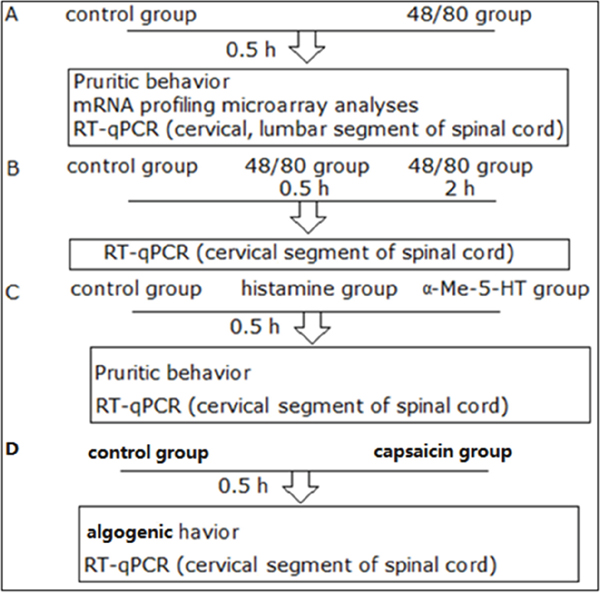
Figure 1: An illustration of the experimental design.
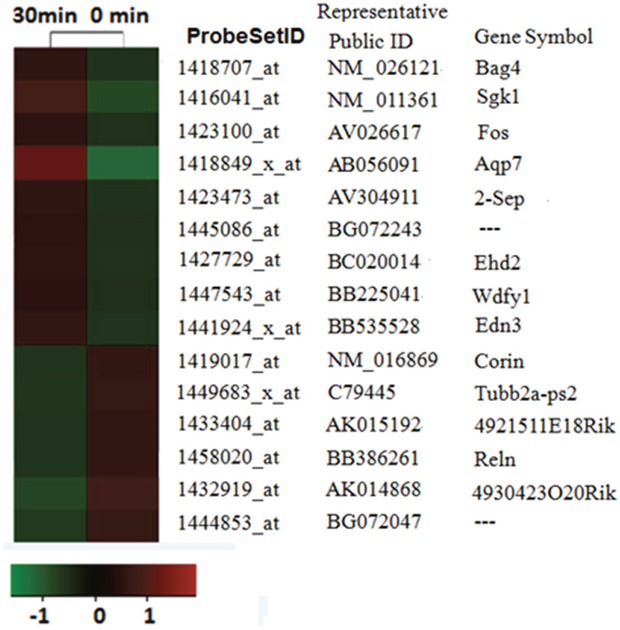
Figure 2: mRNA profiling analysis reveals itch-related altered genes. The probe sets whose expressions were altered in compound 48/80 evoked-acute itching were identified by microarray analysis. The expression levels of each probe set (each line represents a single probe set) were displayed as a log2 ratio of their expression values divided by their expression value in sham mice.
Real-time quantitative PCR (qPCR) validation of mRNA expression in compound 48/80-evoked pruritus mice
To validate the reliability of the microarray results, we analyzed these differentially expressed (DE) mRNAs, including 8 up-regulated mRNA and 4 down-regulated mRNA, by qPCR. The spinal cord (C5-C8) tissues were collected from control group (naive mice) and itch group (compound 48/80 treated mice). Six up-regulated mRNA, including Sgk1, Bag4, Fos, Ehd2, Edn3 and Wdfy were significantly increased, and three down-regulated mRNA, including Corin, 4921511E18Rik and 4930423020Rik were significantly decreased. qPCR results of three mRNA, including Aqp7, 2-sep and Reln were not consistent with data from microarray.
The expression of 9 mRNA in the upper cervical spinal cord (C1-C4) from compound 48/80-evoked pruritus mice
To examine that if there is the differential expression of 9 mRNA between C1-C4 and C5-C8 in spinal cord, we validated the expression of 9 mRNA in the upper cervical spinal cord (C1-C4) from compound 48/80-evoked pruritus mice. Our results showed that the expression of mRNA Ehd2 (d), 4921511E18Rik (h) and 4930423O20Rik (i) was significantly down-regulated in 48/80 group (Figure 3), and the expression of 9 mRNA Sgk1 (a), Bag4 (b), Fos (c), Edn3 (e), Wdfy1 (f) and Corin (g) had no statistically different between control group and 48/80 group (Figure 3).
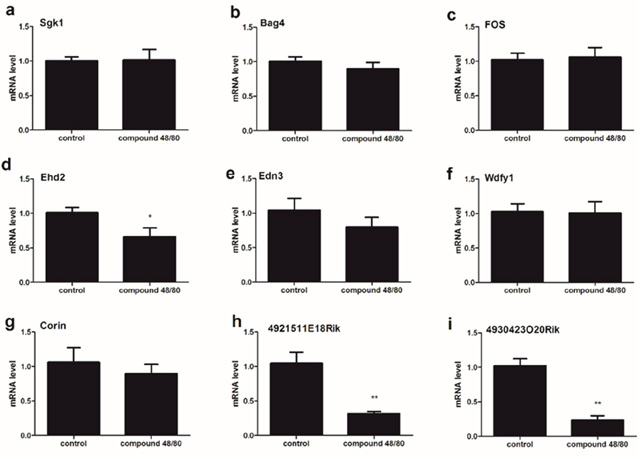
Figure 3: The expression of 9 mRNA in the spinal cord (C1-C4) from compound 48/80-evoked pruritus mice. The expression of mRNA Ehd2 (d), 4921511E18Rik (h) and 4930423O20Rik (i) was significantly down-regulated. The expression of 9 mRNA Sgk1 (a), Bag4 (b), Fos (c), Edn3 (e), Wdfy1 (f) and Corin (g) had no statistically different between control group and itch group. Mann-Whitney test.
The expression of 9 mRNA in the lumbar enlargement of spinal cord from compound 48/80-evoked pruritus mice
To examine if there is the differential expression of 9 mRNA between C5-C8 and the lumbar enlargement of spinal cord, we validated the expression of 9 mRNA in the lumbar enlargement of spinal cord (L3-L5) from compound 48/80-evoked pruritus mice. Our results showed that the expression of 9 mRNA had no statistically different between control group and itch group (Figure 4).
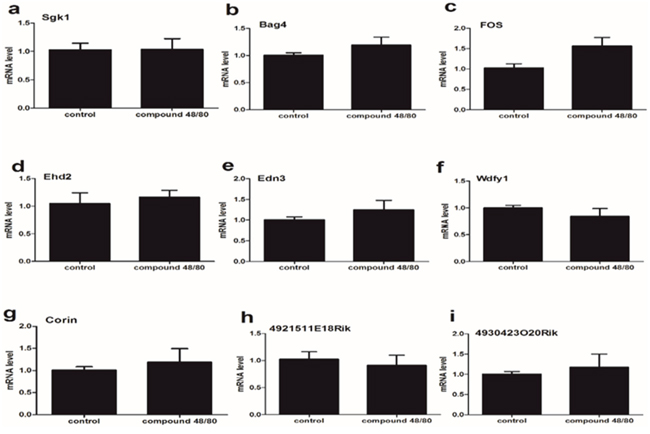
Figure 4: The expression of 9 mRNA in the spinal cord (lumbar enlargement) from compound 48/80-evoked pruritus mice. (a-i) The expression of 9 mRNA had no statistically different between control group and itch group. Mann-Whitney test.
The expression of 9 mRNAs in the spinal cord (C5-C8) at different time points (0.5h/2h) after compound 48/80 injection
Obviously, gene expressions after compound 48/80 injection are varied in different time points. We collected spinal tissue sample from C5-C8 0.5h vs 2h after compound 48/80 injection forRT-qPCR validation. Our results indicated that the expressions of mRNA Sgk1 (a), Bag4 (b), Fos (c), Ehd2 (d), Edn3 (e), Wdfy1 (f) and Reln (l) were significantly up-regulated underlying both 0.5h group and 2h group (Figure 5). The expression of mRNA Corin (g), 4921511E18Rik (h) and 4930423O20Rik (i) was significantly down-regulated underlying both 0.5h group and 2h group. One-way ANOVA (Dunnelt: Compare all columns vs. naive column) (Figure 5).
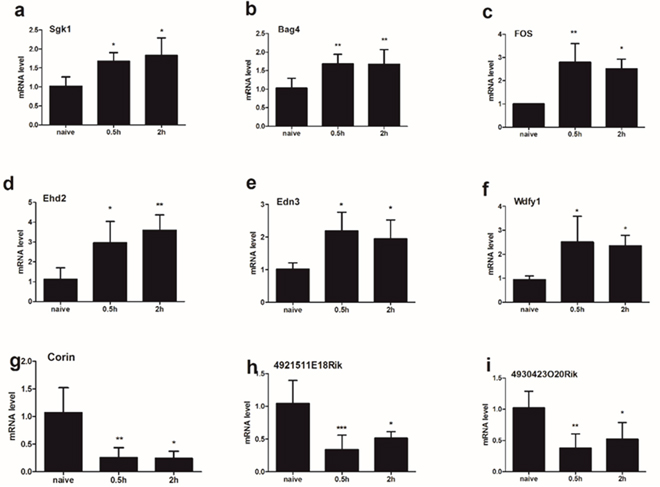
Figure 5: The expression of 9 mRNA were analyzed in the spinal cord (C5-C8) at different time points (0.5h/2h) after compound 48/80 injection. The expressions of mRNA Sgk1 (a), Bag4 (b), Fos (c), Ehd2 (d), Edn3 (e), and Wdfy1 (f) were significantly up-regulated underlying both 0.5h group and 2h group. The expression of mRNA Corin (g), 4921511E18Rik (h) and 4930423O20Rik (i) was significantly down-regulated underlying both 0.5h group and 2h group. One-way ANOVA (Dunnelt: Compare all columns vs. naive column). *P < 0.05, **P < 0.01, ***P < 0.001.
RT-qPCR validation of mRNA expression in the spinal cord (C5-C8) from different itch models
We further analyzed the expression of the 9 mRNAs which were in line with the array results in other two acute itch model including histamine (HA)-treated and α-Me-5-HT-treated mice (Figure 6). The expression of mRNA Fos and Ehd2 (Figure 6c, 6d) were significantly increased in HA group and α-Me-5-HT group. The expression of mRNA Bag4, Corin, 4921511E18Rik and 4930423O20Rik (Figure 6b, 6g, 6h, 6i) were significantly decreased in HA group and α-Me-5-HT group. The expression of mRNA Sgk1, Edn3 and Wdfy1 (Figure 6a, 6e, 6f) had not statistically different between control group and itch group.
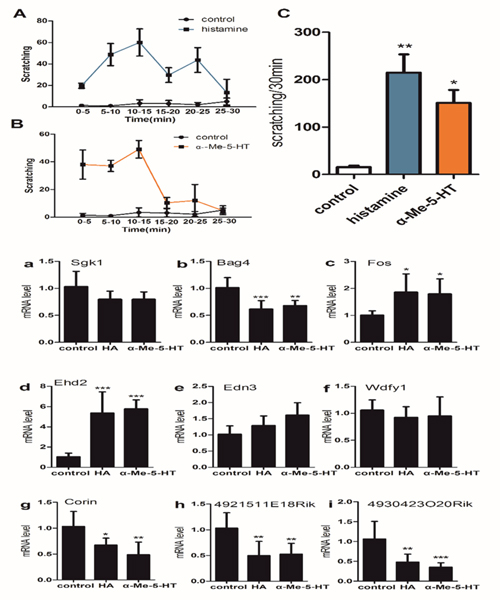
Figure 6: (upper) Pruritogen-evoked scratching behavior in histamine-treated (800μg/50μl), α-Me-5-HT-treated (35μg/50 μl) and saline-treated (control) mice (n = 6/group). (C) Graph by recording the total numbers of scratching bout at the 0-30min observation period by an intradermal microinjection of saline (15.67±5.13), histamine (215.00±66.57) and a-Me-5-HT (151.00±47.00). (A, B) Bar graph plots mean number of scratch bouts recorded at 5 min intervals. One-way ANOVA (Dunnelt: Compare all columns vs. control column). *P < 0.05, **P < 0.01.
(lower) The expressions of 9 mRNA were analyzed in histamine (HA)-treaded and α-Me-5-HT-treaded mice. The expression of mRNA Fos and Ehd2 (c, d) were both significantly increased in HA group and α-Me-5-HT group. The expression of mRNA Bag4 (b), Corin (g), 4921511E18Rik (h) and 4930423O20Rik (i) were both significantly decreased in HA group and α-Me-5-HT group. The expression of mRNA Sgk1 (a), Edn3 (e) and Wdfy1 (f) had no statistically different between control group and itch group. One-way ANOVA (Dunnelt). *P < 0.05, **P < 0.01, ***P < 0.001.
The expression of 9 mRNA in the spinal cord (C5-C8) from capsaicin-treated mice
The time course of the wipes behaviors was compared between control group and capsaicin group. The total numbers of wipes -like behaviors during the 30-min test period was calculated in Figure 7. The difference between the capsaicin group and the control group was significant. RT-qPCR results that the expressions of mRNA Ehd2 (d), Edn3 (e) and Corin (g) were significantly up-regulated in capsaicin group whereas the expression of mRNA 4921511E18Rik (h) was significantly down-regulated, and the expression of mRNA Sgk1 (a), Bag4 (b), Fos (c), Wdfy1 (f) and 4930423O20Rik (i) hadn’t statistically different between control group and capsaicin group (Figure 7).
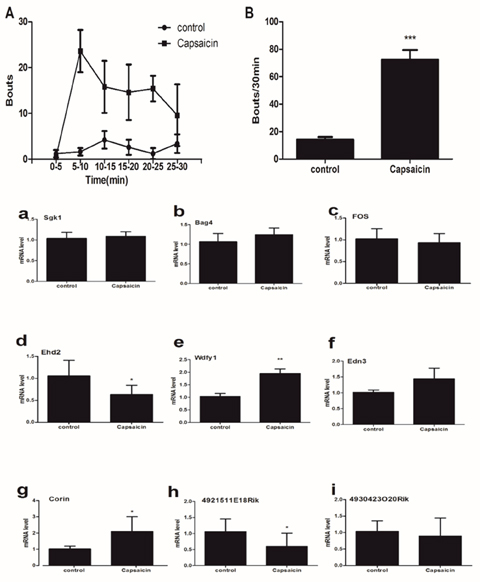
Figure 7: (upper) Bouts directed toward the site of injection of capsaicin (40μg/50 μl) into the nape of the neck. (A) The time course shows bouts at 5-min intervals. (B) Mean total number of bouts during the 30 min period after injection of saline (15.67±5.13) and capsaicin (72.40±7.096). Unpaired t test. ***P < 0.001.
(lower) The expression of 9 mRNA in the spinal cord (C5-C8) from capsaicin-treated mice. The expressions of mRNA Ehd2 (d), Edn3 (e) and Corin (g) were significantly up-regulated in capsaicin group whereas the expression of mRNA 4921511E18Rik (h) was significantly down-regulated. The expression of mRNA Sgk1 (a), Bag4 (b), Fos (c), Wdfy1 (f) and 4930423O20Rik (i) had no statistically different between control group and capsaicin group. Mann-Whitney test. *P < 0.05, **P < 0.01.
DISCUSSION
Here, we identified that compound 48/80, histamine and α-Me-5-HT produced profound scratching behaviors, and these results were consistent with previous reports describing marked effects in mice responding to intradermal injection of a variety of pruritogens [18–26].
It is well-known that regulations of gene expression are varied in time course [27–30], and some gene could be regulated rapidly such as Fos, whereas some might be slow. 0.5h vs 2h after compound 48/80 injection, C5-C8 spinal tissue samples were collected forRT-qPCR validation. Our results indicated that there was no difference between 0.5h group and 2h group, and the expressions of 6 mRNA (Sgk1, Bag4, Fos, Ehd2, Edn3, Wdfy1 and Reln) were significantly up-regulated underlying 0.5h group and 2h group, whereas the expression of 3 mRNA (Corin, 4921511E18Rik and 4930423O20Rik) was significantly down-regulated. Further, some reports previously described that the change of scratching bout took place in the 0-0.5h observation period after pruritogen injection [6, 31]. Thus, in the present acute itching study, the spinal cord tissue samples of animals were detected 0.5 hour after the injection.
In the present report, we address if the intradermal injection of compound 48/80 induce a nonspecific response, since the blood absorption of compound 48/80 from the injection site may lead to the wide changes in gene expression in the whole spinal cord. Therefore, RT-qPCR validation between C1-C4 and lumbar spinal cord tissue was used to exclude the nonspecific response as a control. Our results showed that the expression of 9 mRNA (gk1, Bag4, Fos, Ehd2, Edn3, Wdfy, Corin, 4921511E18Rik and 4930423020Rik) in the spinal cord (lumbar enlargement) was no significant difference between control group and compound 48/80 group, suggesting that compound 48/80 injection don’t induce a nonspecific response by its blood absorption. Furthermore, the expression of 3 mRNA [Ehd2 (d), 4921511E18Rik (h) and 4930423O20Rik (i)] in the spinal cord (C1-C4) was significantly down-regulated, whereas the expression of 6 mRNA [Sgk1 (a), Bag4 (b), Fos (c), Edn3 (e), Wdfy1 (f) and Corin (g)] had no statistically different between control group and compound 48/80 group, suggesting that C1-C4 spinal cord segment may not be primarily involved in neuronal regulation of the injection site.
The spinal cord is a key hub for pruriceptive sensation and its signaling to the central nervous system (CNS) [32]. Pruritogen-triggered hyperexcitability in spinal cord is considered a major source of obstinate itching [33, 34]. The cellular hyperexcitability, in turn, is thought to be implicated in transcriptional switching in special cell body [16]. It is believed that analysis of microarray data of spinal cord may determine every mRNA transcript in a particular sample [35–39]. We selected 12 genes for RT-qPCR validation, and found that 75 % of the expression changes that were initially identified by the microarray data analysis were confirmed by RT-qPCR. These data revealed that the results from these microarray results had a high discovery rate. On comparing three itch models, we found that the various pruritogens yielded consistent results, and in three itch models, Fos and Ehd2 were up-regulated whereas Corin, 4921511E18Rik and 4930423020Rik were down-regulated. Furthermore, Corin and 4930423020Rik were down-regulated in itch model group compared to capsaicin group. A further study is needed in fact to address special function of these mRNA during acute itching.
In conclusion, the present results reveal that the application of microarray technique, coupled with RT-qPCR validation, further explain the mechanism behind itching evoked by pruritic compounds. Our data provides a powerful a useful baseline to begin functional experimentation involved in acute itching. It can thus contribute to our understanding of pharmacological methods for prevention or treatment of pruritus.
MATERIALS AND METHODS
Animal care
Male C57BL/6J mice (8-10 weeks old) were provided by the Experimental Animal Research Center of Hubei Province (license number: 42000600018764). The animals were maintained in a climate controlled room on a 12-hlight/dark cycle (light on at 07:00 h). Mice were housed (5/cage), but they were individually caged during each experiment. Food and water were available ad libitum. Experimental protocols were approved by the Institutional Animal Care and Use Committee of Tongji Hospital (No.TJ-A20150803). All testing and surgeries were performed in accordance with the policies and recommendations of the National Guides for the Care and Use of Laboratory Animals.
Experimental design
Experiment A Mice were randomly assigned to two groups: (1) control group (saline 100μl, n = 18); (2) compound 48/80 group (50 μg/50μl, n = 18). After injection, pruritic behavior was immediately measured every 5 min. 0.5h after injection, cervical segment of spinal cord (C5-C8) tissue were prepared for mRNA profiling microarray analyses, and cervical (C1-C4, C5-C8) and lumbar segment of spinal cord for Real-Time quantitative PCR (RT-qPCR).
Experiment B Mice were divided into saline group (control group, n=12) and compound 48/80 group (48/80 group, n=12). 0.5h and 2h after injection, C5-C8 tissue was prepared for RT-qPCR.
Experiment C Mice were randomly assigned to three groups: (1) control group (saline 100μl, n = 6); (2) histamine group (800μg/50μl, n = 6); (3) α-Me-5-HT group (35 μg/50μl, n = 6). 0.5h after injection, C5-C8 tissue was prepared for RT-qPCR.
Experiment D Mice were randomly assigned to two groups: (1) control group (saline 100μl, n = 6); (2) capsaicin group (40μg/50μl, n = 6). Algogenic behavior was immediately measured every 5 min after injection. 0.5h after injection, C5-C8 tissue was prepared for RT-qPCR.
The protocols of each experiment were presented in the parts of figure legends. All procedures and behavioral assay were conducted in an isolated quiet room to reduce variance. An illustration of the experimental design above is displayed in Figure 1.
Pruritic/wipes behavior
To induce acute itching on the skin, we followed a previously reported procedure [6, 31, 40–42]. Animals were received an intradermal microinjection of 100 μl saline, compound 48/80 (50μg/50μl), histamine (800μg/50μl), α-Me-5-HT group (35μg/50μl) or capsaicin (40μg/50μl) in the nape of the neck (via a 0.3 ml insulin syringe), respectively. Compound 48/80, histamine, α-Me-5-HT and capsaicin were purchased from Sigma-Aldrich (St. Louis, MO, USA). All drugs were dissolved in saline. All pruritic behaviors were evaluated as previously described with minor modifications [6, 43, 44]. The tests of algogenic behavior by capsaicin were performed similar with previous researches [11, 43].
Tissue preparation
After pruritic behavior assay, the animals were immediately decapitated by cervical dislocation. Following decapitation, cervical (C1-C4, C5-C8) or lumbar segments in spinal cord were dissected using a dissection microscope and taken for subsequent analysis. The tissue was flash frozen in liquid nitrogen. Total RNA was isolated using Trizol® reagent (Invitrogen, Carlsbad CA). RNA samples were performed by Ambion mirVana miRNA Isolation Kit for purity and concentration.
mRNA profiling microarray analyses
High quality samples containing 200 ng of total RNA were used on microarray chips according to the manufacturer’s instructions. RNA was collected from six animals for each experimental condition. Gene profiling of C5-C8 spinal cord from control group and 48/80 group were carried out using standard Affymetrix protocols and hybridized to Affymetrix Mouse Genome 430 2.0 Array as described previously [45]. All subsequent analyses were conducted using SAM3.0 software (Stanford University, Stanford CA; http://www-stat.stanford.edu/~tibs/SAM).
Real-time quantitative PCR
2 μg of total RNA was extracted from mouse spinal cords (C1-C4, C5-C8 or lumbar segment) using TRIzol reagent (Invitrogen, USA) according to the manufacturer’s protocol and quantified using a spectrophotometer (BioPhotometer, Eppendorf, Hamburg, Germany)[27, 46–49]. The threshold cycle (CT) was used to estimate the amount of target mRNA. The comparative CT method with the formula for relative fold-change = 2-ΔΔCTwas used to quantify the amplified transcripts. The specific forward (F) and reverse (R) primer sequences (Table 1) were designed [1]. Experiments were evaluated in triplicate.
Table 1: Primer sequences for RT-qPCR
Gene |
Forward (5′ to 3′) |
Reverse (5′ to 3′) |
|---|---|---|
Sgk1 |
TGCCACCCTGGATCTATAACTG |
GGCCTCAAAGTCTGACTCCC |
Bag4 |
CCAAACACCTACCGTTCACCT |
TGTGGTCGTCCAGTCCCTC |
Aqp7 |
AAGGGCTTTCGTGCATCAGTA |
ACTCCTATCCAGAAAACCGTCAA |
Sept2 |
GGGTGGTGACAGTGACAGCG |
CCTTCCTTCCACAGGGCTAA |
Edn3 |
GCTTGCGTTGTACTTGTATGGG |
GGTGGGCTTTATCTGTCCTTGA |
Wdfy1 |
ATCAAGACCTATCCAGCCCACC |
AAGCCCACGACGAGAAGAAG |
Corin |
AAAAGCGACCGAGATAAGAGTG |
AAGCGCAGCAAGTTAGCAGT |
Tubb2a-ps2 |
TGCATTTTGATGCCTTAGAAGT |
CGCATGGTGCCTGGTTAG |
Reln |
CTTTGATGGCTTGCTGGTGA |
GGTTGGTTGTAGGCAGGTGA |
Fos |
GCCCCTTCTCAACGACCC |
CATCCCCAAGGAATTGCTGT |
Ehd2 |
CGCAAGCTCAACGACCTAGT |
TGAAGTCATGTGCCATCAACAG |
4930423O20Rik |
CTCAGCACCGACTCTTACACG |
TGCTCTTGCTTCTTGCTCCTA |
4921511E18 |
CGTCCCTGACCCCTACTCC |
AAACTAGCAAGTGGCCCGTTA |
β-actin |
CACGATGGAGGGGCCGGACTCATC |
TAAAGACCTCTATGCCAACACAGT |
Statistics and data analysis
Behavioral tests were performed by observers blinded to the treatments of the animals used. All quantification data are presented as mean ± SEM, and error bars represent SEM. Statistical comparisons were performed with Mann-Whitney test. The statistical analyses were done using t test, and P < 0.05 was considered statistically significant.
ACKNOWLEDGMENTS
This work was supported by grants from National Natural Science Foundation of China (81670240 and 81271766), and National Natural Science Foundation of Hubei Province (2016CFB625 and 2016CFB324).
CONFLICTS OF INTEREST
The authors have no conflicts of interest related to this paper.
REFERENCES
1. He ZG, Liu BW, Li ZX, Liu C, Xiang HB. Altered expression profiling of spinal genes modulated by compound 48/80 in a mouse itch model. J Anesth Perioper Med. 2017; 4. https://doi.org/10.24015/JAPM.2017.0020.
2. Zhao ZQ, Huo FQ, Jeffry J, Hampton L, Demehri S, Kim S, Liu XY, Barry DM, Wan L, Liu ZC, Li H, Turkoz A, Ma K, et al. Chronic itch development in sensory neurons requires BRAF signaling pathways. J Clin Invest. 2013; 123:4769-4780.
3. Nattkemper LA, Zhao ZQ, Nichols AJ, Papoiu AD, Shively CA, Chen ZF, Yosipovitch G. Overexpression of the gastrin-releasing peptide in cutaneous nerve fibers and its receptor in the spinal cord in primates with chronic itch. J Invest Dermatol. 2013; 133:2489-2492.
4. Liu T, Xu ZZ, Park CK, Berta T, Ji RR. Toll-like receptor 7 mediates pruritus. Nat Neurosci. 2010; 13:1460-1462.
5. Davidson S, Zhang X, Khasabov SG, Simone DA, Giesler GJ Jr. Relief of itch by scratching: state-dependent inhibition of primate spinothalamic tract neurons. Nat Neurosci. 2009; 12:544-546.
6. Akiyama T, Merrill AW, Zanotto K, Carstens MI, Carstens E. Scratching behavior and Fos expression in superficial dorsal horn elicited by protease-activated receptor agonists and other itch mediators in mice. J Pharmacol Exp Ther. 2009; 329:945-951.
7. Davidson S, Zhang X, Yoon CH, Khasabov SG, Simone DA, Giesler GJ Jr. The itch-producing agents histamine and cowhage activate separate populations of primate spinothalamic tract neurons. J Neurosci. 2007; 27:10007-10014.
8. Paus R, Schmelz M, Biro T, Steinhoff M. Frontiers in pruritus research: scratching the brain for more effective itch therapy. J Clin Invest. 2006; 116:1174-1186.
9. Sun YG, Chen ZF. A gastrin-releasing peptide receptor mediates the itch sensation in the spinal cord. Nature. 2007; 448:700-703.
10. Sun YG, Zhao ZQ, Meng XL, Yin J, Liu XY, Chen ZF. Cellular basis of itch sensation. Science. 2009; 325:1531-1534.
11. Liu T, He Z, Tian X, Kamal GM, Li Z, Liu Z, Liu H, Xu F, Wang J, Xiang H. Specific patterns of spinal metabolites underlying alpha-Me-5-HT-evoked pruritus compared with histamine and capsaicin assessed by proton nuclear magnetic resonance spectroscopy. Biochim Biophys Acta. 2017; 1863:1222-1230.
12. Akiyama T, Carstens MI, Carstens E. Enhanced scratching evoked by PAR-2 agonist and 5-HT but not histamine in a mouse model of chronic dry skin itch. Pain. 2010; 151:378-383.
13. Akiyama T, Iodi Carstens M, Carstens E. Transmitters and pathways mediating inhibition of spinal itch-signaling neurons by scratching and other counterstimuli. PLoS One. 2011; 6:e22665.
14. Valtcheva MV, Samineni VK, Golden JP, Gereau RW, Davidson S. Enhanced nonpeptidergic intraepidermal fiber density and an expanded subset of chloroquine-responsive trigeminal neurons in a mouse model of dry skin itch. J Pain. 2015; 16:346-356.
15. Maratou K, Wallace VC, Hasnie FS, Okuse K, Hosseini R, Jina N, Blackbeard J, Pheby T, Orengo C, Dickenson AH, McMahon SB, Rice AS. Comparison of dorsal root ganglion gene expression in rat models of traumatic and HIV-associated neuropathic pain. Eur J Pain. 2009; 13:387-398.
16. Persson AK, Gebauer M, Jordan S, Metz-Weidmann C, Schulte AM, Schneider HC, Ding-Pfennigdorff D, Thun J, Xu XJ, Wiesenfeld-Hallin Z, Darvasi A, Fried K, Devor M. Correlational analysis for identifying genes whose regulation contributes to chronic neuropathic pain. Mol Pain. 2009; 5:7.
17. Dominguez CA, Carlstrom KE, Zhang XM, Al Nimer F, Lindblom RP, Ortlieb Guerreiro-Cacais A, Piehl F. Variability in C-type lectin receptors regulates neuropathic pain-like behavior after peripheral nerve injury. Mol Pain. 2014; 10:78.
18. Xiang CX, Zhang KH, Johnson RL, Jacquin MF, Chen ZF. The transcription factor, Lmx1b, promotes a neuronal glutamate phenotype and suppresses a GABA one in the embryonic trigeminal brainstem complex. Somatosens Mot Res. 2012; 29:1-12.
19. Sikand P, Dong X, LaMotte RH. BAM8-22 peptide produces itch and nociceptive sensations in humans independent of histamine release. J Neurosci. 2011; 31:7563-7567.
20. Patel KN, Liu Q, Meeker S, Undem BJ, Dong X. Pirt, a TRPV1 modulator, is required for histamine-dependent and -independent itch. PLoS One. 2011; 6:e20559.
21. Patel KN, Dong X. Itch: cells, molecules, circuits. ACS Chem Neurosci. 2011; 2:17-25.
22. Liu XY, Liu ZC, Sun YG, Ross M, Kim S, Tsai FF, Li QF, Jeffry J, Kim JY, Loh HH, Chen ZF. Unidirectional cross-activation of GRPR by MOR1D uncouples itch and analgesia induced by opioids. Cell. 2011; 147:447-458.
23. Liu Q, Weng HJ, Patel KN, Tang Z, Bai H, Steinhoff M, Dong X. The distinct roles of two GPCRs, MrgprC11 and PAR2, in itch and hyperalgesia. Sci Signal. 2011; 4:ra45.
24. Xiang C, Zhang KH, Yin J, Arends JJ, Erzurumlu RS, Jacquin MF, Chen ZF. The transcription factor, Lmx1b, is necessary for the development of the principal trigeminal nucleus-based lemniscal pathway. Mol Cell Neurosci. 2010; 44:394-403.
25. Reddy VB, Shimada SG, Sikand P, Lamotte RH, Lerner EA. Cathepsin S elicits itch and signals via protease-activated receptors. J Invest Dermatol. 2010; 130:1468-1470.
26. Patel KN, Dong X. An itch to be scratched. Neuron. 2010; 68:334-339.
27. Ke C, Gao F, Tian X, Li C, Shi D, He W, Tian Y. Slit2/Robo1 mediation of synaptic plasticity contributes to bone cancer pain. Mol Neurobiol. 2017; 54:295-307.
28. Harrison BJ, Venkat G, Hutson T, Rau KK, Bunge MB, Mendell LM, Gage FH, Johnson RD, Hill C, Rouchka EC, Moon L, Petruska JC. Transcriptional changes in sensory ganglia associated with primary afferent axon collateral sprouting in spared dermatome model. Genom Data. 2015; 6:249-252.
29. Knowlton WM, Palkar R, Lippoldt EK, McCoy DD, Baluch F, Chen J, McKemy DD. A sensory-labeled line for cold: TRPM8-expressing sensory neurons define the cellular basis for cold, cold pain, and cooling-mediated analgesia. J Neurosci. 2013; 33:2837-2848.
30. Yang W, Rudick CN, Hoxha E, Allsop SA, Dimitrakoff JD, Klumpp DJ. Ca(2+)/calmodulin-dependent protein kinase II is associated with pelvic pain of neurogenic cystitis. Am J Physiol Renal Physiol. 2012; 303:F350-356.
31. Akiyama T, Merrill AW, Carstens MI, Carstens E. Activation of superficial dorsal horn neurons in the mouse by a PAR-2 agonist and 5-HT: potential role in itch. J Neurosci. 2009; 29:6691-6699.
32. Weber HC. Gastrointestinal peptides and itch sensation. Curr Opin Endocrinol Diabetes Obes. 2015; 22:29-33.
33. Akiyama T, Tominaga M, Takamori K, Carstens MI, Carstens E. Roles of glutamate, substance P, and gastrin-releasing peptide as spinal neurotransmitters of histaminergic and nonhistaminergic itch. Pain. 2014; 155:80-92.
34. Sukhtankar DD, Ko MC. Physiological function of gastrin-releasing peptide and neuromedin B receptors in regulating itch scratching behavior in the spinal cord of mice. PLoS One. 2013; 8:e67422.
35. LaCroix-Fralish ML, Austin JS, Zheng FY, Levitin DJ, Mogil JS. Patterns of pain: meta-analysis of microarray studies of pain. Pain. 2011; 152:1888-1898.
36. Zhao B, Yu Q, Li H, Guo X, He X. Characterization of microRNA expression profiles in patients with intervertebral disc degeneration. Int J Mol Med. 2014; 33:43-50.
37. Raju HB, Englander Z, Capobianco E, Tsinoremas NF, Lerch JK. Identification of potential therapeutic targets in a model of neuropathic pain. Front Genet. 2014; 5:131.
38. Liu TT, Liu BW, He ZG, Feng L, Liu SG, Xiang HB. Delineation of the central melanocortin circuitry controlling the kidneys by a virally mediated transsynaptic tracing study in transgenic mouse model. Oncotarget. 2016; 7:69256-69266. https://doi.org/10.18632/oncotarget.11956.
39. Sakurai E, Kurihara T, Kouchi K, Saegusa H, Zong S, Tanabe T. Upregulation of casein kinase 1epsilon in dorsal root ganglia and spinal cord after mouse spinal nerve injury contributes to neuropathic pain. Mol Pain. 2009; 5:74.
40. He ZG, Zhang DY, Liu SG, Feng L, Feng MH, Xiang HB. Neural circuits of pain and itch processing involved in anterior cingulate cortex. Int J Clin Exp Med. 2016; 9:22976-22984.
41. Liu C, Liu TT, He ZG, Shu B, Xiang HB. Inhibition of itch-related responses by selectively ablated serotonergic signals at the rostral ventromedial medulla in mice. Int J Clin Exp Pathol. 2014; 7:8917-8921.
42. Yu YQ, Barry DM, Hao Y, Liu XT, Chen ZF. Molecular and neural basis of contagious itch behavior in mice. Science. 2017; 355:1072-1076.
43. Shimada SG, LaMotte RH. Behavioral differentiation between itch and pain in mouse. Pain. 2008; 139:681-687.
44. Liu Q, Tang Z, Surdenikova L, Kim S, Patel KN, Kim A, Ru F, Guan Y, Weng HJ, Geng Y, Undem BJ, Kollarik M, Chen ZF, et al. Sensory neuron-specific GPCR Mrgprs are itch receptors mediating chloroquine-induced pruritus. Cell. 2009; 139:1353-1365.
45. Bottomly D, Walter NA, Hunter JE, Darakjian P, Kawane S, Buck KJ, Searles RP, Mooney M, McWeeney SK, Hitzemann R. Evaluating gene expression in C57BL/6J and DBA/2J mouse striatum using RNA-Seq and microarrays. PLoS One. 2011; 6:e17820.
46. Guan XH, Fu QC, Shi D, Bu HL, Song ZP, Xiong BR, Shu B, Xiang HB, Xu B, Manyande A, Cao F, Tian YK. Activation of spinal chemokine receptor CXCR3 mediates bone cancer pain through an Akt-ERK crosstalk pathway in rats. Exp Neurol. 2015; 263:39-49.
47. Bu H, Shu B, Gao F, Liu C, Guan X, Ke C, Cao F, Hinton AO Jr, Xiang H, Yang H, Tian X, Tian Y. Spinal IFN-gamma-induced protein-10 (CXCL10) mediates metastatic breast cancer-induced bone pain by activation of microglia in rat models. Breast Cancer Res Treat. 2014; 143:255-263.
48. He Z, Fan X, Liu TG, Xiang H. Practical management for fast atrial fibrillation during video-assisted thoracoscopic thymic surgery in patients with coronary heart disease: a case report. Int J Clin Exp Med. 2016; 9:3735-3740.
49. Fu Q, Shi D, Zhou Y, Zheng H, Xiang H, Tian X, Gao F, Manyande A, Cao F, Tian Y, Ye D. MHC-I promotes apoptosis of GABAergic interneurons in the spinal dorsal horn and contributes to cancer induced bone pain. Exp Neurol. 2016; 286:12-20.

