INTRODUCTION
Male gametes are produced during spermatogenesis occurring in seminiferous tubules inside the testis. Spermatogenesis is a complex process through which spermatogonial stem cells (SSCs) differentiate into spermatozoa. This process involves three phases: mitotic phase (spermatogonial proliferation and differentiation), meiotic phase (chromosomal reduction in spermatocytes via meiosis), and spermatid differentiation into spermatozoa [1]. The successful differentiation of primordial germ cells into mature spermatozoa is mainly controlled by hormones released by the hypothalamic–pituitary–testis axis and growth factors [2]. In addition to the hypothalamic–pituitary–testis axis, the pineal gland is essential for male reproduction [3, 4].
Melatonin is the main neurohormone secreted by the pineal gland, and this hormone is involved in diverse physiological functions, such as male reproduction [5]. In male reproduction, melatonin plays regulatory roles in the testicular development of seasonal breeding animals and affects the release of gonadotropins, which regulate spermatogenesis and testosterone synthesis [6]. Melatonin as a potent free radical scavenger influences sperm quality [7, 8] and performs testicular regulatory roles possibly mediated by receptors in testicular tissues of different species [9, 10]. This hormone can also alter the expression profile of miRNAs in mouse testis, as indicated by indirect evidence [11]. The regulatory roles of melatonin in testicular cells have been further investigated in detail. Sertoli cells play an essential role in spermatogenesis regulation by providing physical support for germ cells and thus form the blood–testis barrier and secrete protein products [12]. Sertoli cells are sensitive to hormonal actions, and the proliferation and metabolism of these cells are regulated by hormones [13]. For example, bovine Sertoli cells contain melatonin receptors 1 and 2, and the expression of these receptors and spermatogenesis-regulated genes is influenced by melatonin in a time- and dose-dependent manner [14]. The metabolites of Sertoli cells are also vital for germ cell survival, and the metabolism of Sertoli cells may be mediated by hormones, such as estrogen and androgen [13]. In an in vitro model, the glycolytic profile of Sertoli cells can be altered by melatonin [15]. Thus, spermatogenesis may be affected by melatonin through the regulation of Sertoli cell functions. For instance, spermatogenesis successfully occurs through the renewal and differentiation of spermatogonial stem cells [16]. To confirm the role of melatonin in spermatogenesis, researchers cultured dissociated cells and seminiferous tubular fragments from sheep testis for 30 days and found that melatonin significantly increases testosterone concentrations and expression of luteinizing hormone receptor (LHR) and steroidogenic acute regulatory protein (StAR) [17]. Melatonin also significantly promotes the differentiation of cultured SSCs into haploid germ cells; therefore, melatonin is implicated in spermatogenesis [17]. Our previous study provided direct evidence showing that melatonin promotes the proliferation of GC-1 spg cells by downregulating miR-16 expression [11].
Melatonin elicits beneficial effects on male reproduction. For instance, this hormone protects the testis from oxidative damage and promotes germ cell growth. However, relevant molecular mechanisms remain unknown. Various molecular mechanisms involved in spermatogenesis have been identified through different methods [18, 19], and some of these mechanisms are regulated by different signaling pathways, including phosphatidylinositol 3-kinase (PI3K), extracellular signal–regulated kinases 1 and 2 (ERK1/2), and transforming growth factor-beta (TGF-beta) pathways [20–22].
Although studies have revealed that exposure to melatonin alters the microRNA expression in the testis, studies have yet to determine whether melatonin can regulate the changes in transcriptome in germ cells. Melatonin-regulated cellular functions, including growth and apoptosis [23, 24], involve ERK1/2 signaling pathway. Nevertheless, the interaction of melatonin and ERK1/2 signaling pathway in germ cells has yet to be examined in detail. To obtain further insights into melatonin-promoted proliferation of GC-1 spg cells, we investigated melatonin-regulated genes through transcriptome sequencing and observed the interaction of melatonin and ERK1/2 signaling pathway.
RESULTS
The effect of melatonin on the proliferation of GC-1 spg cells was determined with CCK8 assays after these cells were treated with different melatonin concentrations (1–0.01 μM) for 24 hrs. Consistent with previous findings [11], our results confirmed that melatonin induces a statistically beneficial effect on the proliferation of GC-1 spg cells (Figure 1). Melatonin of 10nM was used for the follow-up experiments.
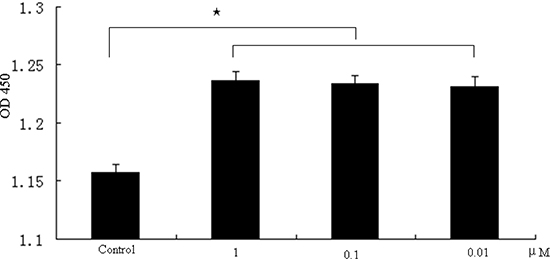
Figure 1: Promotion of cell proliferation by melatonin in GC-1 spg cells. GC-1 spg cells were treated with different concentrations of melatonin (1–0.01 μM). Data represent the mean ± SD of the results in three independent experiments. ⋆ indicates significant differences (p < 0.05).
In the current experiment, transcriptome sequencing arrays were used to determine the mRNA expression profile of GC-1 spg cells induced by melatonin, which could account for the significant proliferation effect. After 24 hrs stimulation on GC-1 spg cell, different patterns of mRNA expression were observed (Figure 2). RNA sequencing alignments revealed 18,526 genes. Of these genes, 11 were upregulated and 17 were downregulated. These 28 genes were differentially expressed between melatonin-stimulated and control groups (Supplementary Table 1).
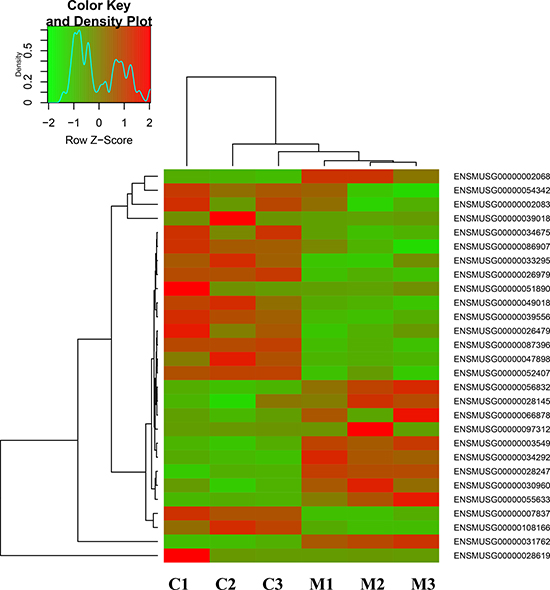
Figure 2: The distinct expression patterns of transcriptome in melatonin-regulated GC-1 spg cells. Hierarchical clustering of transcriptome that showed the different gene expression profiles in GC-1 spg cells after melatonin treatment. Expression levels are defined by the color key on the top-left side of the figure. C indicates control, M indicates melatonin treatment.
The differentially expressed genes were classified into 3 GO categories and 34 terms through gene ontology analysis (Figure 3). In the biological process category, the differentially expressed genes were classified into 16 terms, including cellular process, biological regulation, growth, reproduction, and et al. In the cellular component categories, the differentially expressed genes were classified into 13 terms, including cell part, organelle part, membrane, macromolecular complex, and et al. In the molecular function category, the differentially expressed genes were classified into 5 terms, including binding, transporter activity, molecular transducer activity, molecular function regulator, catalytic activity, and et al.
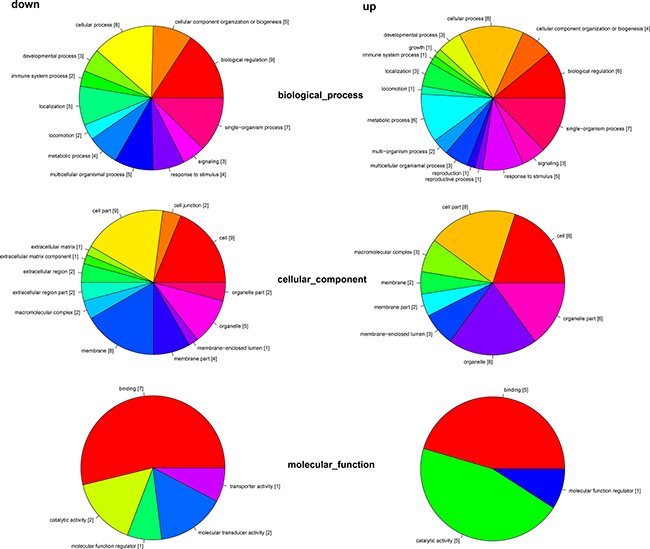
Figure 3: Distribution of transcriptome by gene ontology (GO). All sequences with Blast2GO matches were assigned to three GO categories (biological process, cellular component, and molecular function) and classified into 34 functional terms. Data are shown for both the downregulated (left) and upregulated (right) transcriptome regulated by melatonin.
The differentially expressed genes were compared with the KEGG database using BLASTX, and the corresponding pathways were established. Among the differentially expressed genes assigned to the KEGG pathway 1, 2, 8, 7, 8 and 15 were assigned to metabolism, genetic information, environmental information processing, cellular process, organismal systems, and human diseases, respectively (Figure 4). These annotations provided valuable information to investigate specific processes, functions, and pathways influencing the role of melatonin in male reproduction.
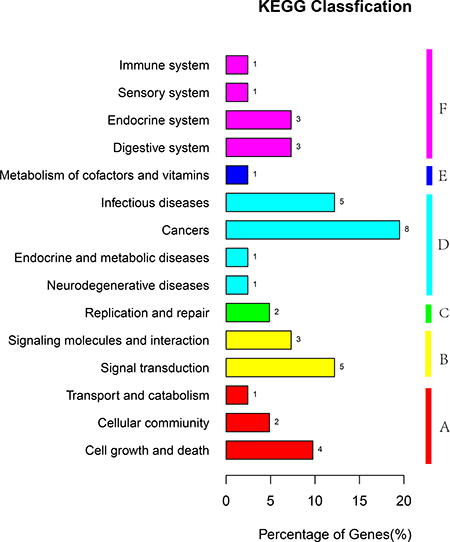
Figure 4: KEGG classification of unigenes. The unigenes were summarized into six main categories (a: cellular processes, b: environmental information processing, c: genetic information processing, d: human diseases, e: metabolism, and f: organismal systems,). The x-axis represents the unigenes’ respective categories, whereas the y-axis represents the percentage of unigenes.
The differentially expressed mRNAs were subjected real-time quantitative PCR to validate the mRNA sequencing data. Five genes, namely, two downregulated and three upregulated genes, associated with DNA replication and repair and cell cycle were quantified. Real-time quantitative PCR data were consistent with mRNA sequencing results (Figure 5). The Mt2 expression was significantly increased after the GC-1 spg cells were treated with melatonin.
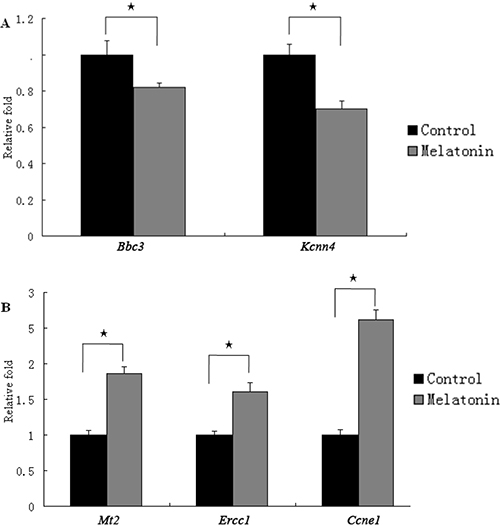
Figure 5: The expression of genes regulated by melatonin in GC-1 spg cells. Five differentially expressed genes identified by transcriptome sequencing were determined using qRT-PCR. The mRNA level of Bbc3 and Kcnn4 was downregulated by melatonin (A). The mRNA level of Mt2, Ercc1, and Ccne1 was upregulated by melatonin (B). ⋆ indicates significant differences (p < 0.05).
The GC-1 spg cells were transfected with Mt2 expression plasmid (pCMV6-Mt2) to investigate whether Mt2 is an essential mediator of the effect of melatonin on the proliferation of GC-1 spg cells. The results showed that the expression of Mt2 was remarkably increased after transfection with pCMV6-Mt2 (Figure 6A, 6B). The CCK8 assay demonstrated that cell proliferation of the Mt2-transfected cells grew more rapidly than the control cells did (Figure 6C). Real-time PCR revealed that the expression of genes associated with cell proliferation was also increased (Figure 6D). Flow cytometric analyses showed that the percentage of GC-1 spg cells in the S phase was higher than that of the control cells because of the overexpression of Mt2 (Figure 7A). Double-staining with Annexin V and propidium iodide showed that Mt2 decreased the percentage of apoptotic cells relative to that of the control groups (Figure 7B).
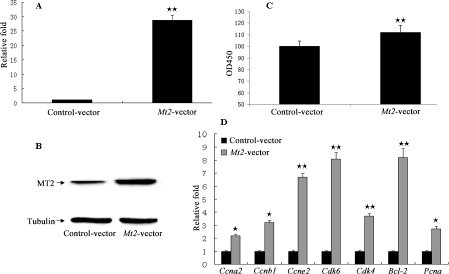
Figure 6: The cell proliferation assay after Mt2 overexpression in GC-1 spg cells. Overexpression plasmid of Mt2 was constructed using pCMV6 plasmid. Plasmid was transfected into GC-1 spg cells with Lipofectamine 2000. Both mRNA (A) and protein (B) of Mt2 were increased in cells with Mt2 overexpression. Furthermore, cell proliferation assay showed that Mt2 overexpression promoted cell proliferation and the expression of genes associated with cell proliferation (C, D). ⋆ indicates significant differences (p < 0.05). ⋆⋆ indicates extremely significant differences (p < 0.01).
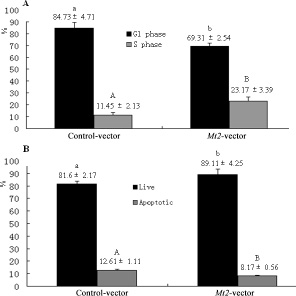
Figure 7: Cell cycle and apoptosis after Mt2 overexpression in GC-1 spg cells. The percentage of cells in the G1 phase was decreased, and the percentage of cells in the S phase was increased with Mt2 overexpression (A). Meanwhile, the percentage of apoptotic cells was decreased when GC-1 spg cells were transfected with Mt2 plasmid (B). Different letters indicate significant differences (p < 0.05). In apoptosis assay, the percentage of apoptotic cells included both early and late apoptosis.
A complementation experiment was then performed. GC-1 spg cells were transfected with Mt2-specific siRNA or a nonspecific siRNA. The transfection significantly decreased the Mt2 expression in GC-1 spg cells transfected with Mt2 siRNA than in the cells in the control siRNA group (Figure 8A, 8B). CCK8 assay demonstrated that the proliferation of GC-1spg cells was significantly inhibited after Mt2 siRNA transfection was performed (Figure 8C). The Mt2 knockdown in GC-1 spg cells significantly downregulated the expression of genes associated with cell proliferation (Figure 8D). Furthermore, Mt2 knockdown in GC-1 spg cells decreased the percentage of cells in the S phase (Figure 9A) and increased the percentage of apoptotic cells relative to those in the control group (Figure 9B).
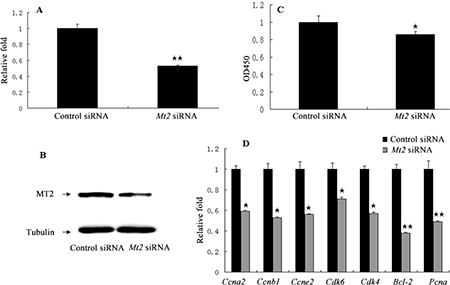
Figure 8: The cell proliferation after Mt2 knockdown in GC-1 spg cells. GC-1 spg cells were transfected with Mt2 siRNA. Both mRNA (A) and protein (B) of Mt2 were decreased in cells with Mt2 siRNA. Cell proliferation assay showed that Mt2 knockdown inhibited cell proliferation (C). The expression of genes involved in cell proliferation was decreased with Mt2 siRNA (D). ⋆ indicates significant differences (p < 0.05). ⋆⋆ indicates extremely significant differences (p < 0.01).
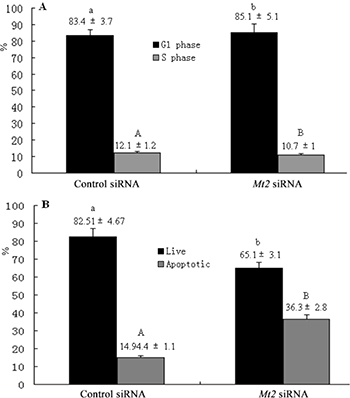
Figure 9: The cell cycle and apoptosis after Mt2 knockdown in GC-1 spg cells. The percentage of cells in the G1 phase was increased, and the percentage of cells in the S phase was decreased with Mt2 siRNA (A). Meanwhile, the percentage of apoptotic cells was increased after Mt2 knockdown in cells (B). Different letters indicate significant differences (p < 0.05).
Mitogen-activated protein kinases (MAPKs) are implicated in cell growth. Thus, we investigated whether melatonin treatment affects MAPK signaling in melatonin-induced proliferation of GC-1 spg cells. Results showed that cell treatment with melatonin at different time points (2, 6, and 12 h) activated the phosphorylation of ERK1/2 (Figure 10A). We then investigated whether MAPK signaling is involved in melatonin-induced proliferation of GC-1 spg cells. The cells were treated with ERK1/2 inhibitor (PD098059) and subjected to melatonin treatment. Results revealed that ERK1/2 inhibitor significantly inhibited the melatonin-induced proliferation in GC-1 spg cells (Figure 10B), downregulated the expression of Mt2 (Figure 10C), decreased the percentage of cells in the S phase (Figure 11A), and increased the number of apoptotic cells (Figure 11B). Thus, melatonin mediated the upregulation of Mt2 and subsequently increased the proliferation of GC-1spg cells by activating the MAPK signaling cascade.
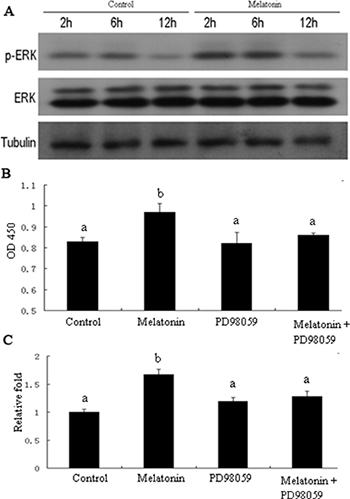
Figure 10: ERK1/2 signaling pathway involved in melatonin-regulated Mt2 expression and cell proliferation in GC-1 spg cell. GC-1 spg cells were treated with melatonin of 10nM for 2 hrs, 6 hrs, and 12 hrs. The level of ERK1/2 phosphorylation was increased after melatonin treatment and detected by Western blot (A). Cells were treated ERK1/2 signaling inhibitor (PD98059, S1805, Beyotime, 30μM) for 2 hrs, then followed by melatonin treatment for 24 hrs. PD98059 inhibited the melatonin-induced proliferation of GC-1 spg cells (B). The qRT-PCR showed that Mt2 expression induced by melatonin was also decreased after PD98059 treatment (C). Different letters indicate significant differences (p < 0.05).
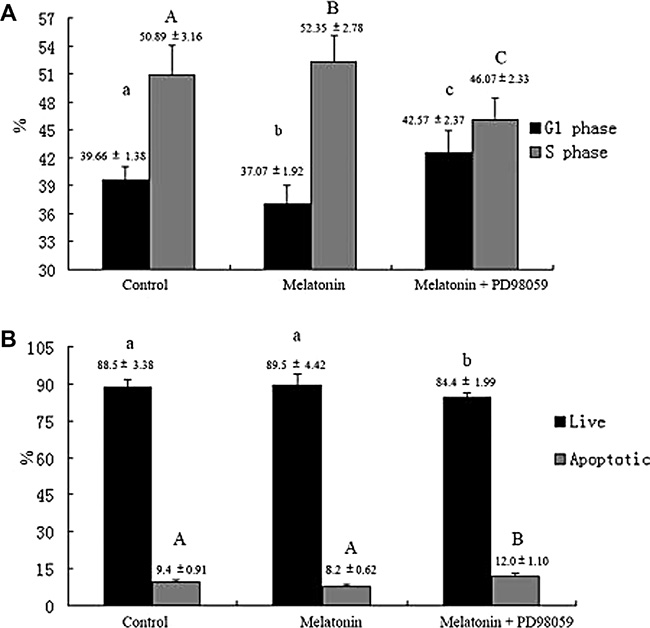
Figure 11: ERK1/2 signaling pathway involved in melatonin-regulated cell cycle and apoptosis in GC-1 spg cells. PD98059 decreased the percentage of cells in the S phase and increased the percentage of cells in the G1 phase (A). Furthermore, PD98059 increased the percentage of apoptotic cells (B). Different letters indicate significant differences (p < 0.05).
DISCUSSION
Germ cells in spermatogenesis contain the same genome, but only some genes are used in each stage of spermatogenesis. Some of these genes are expressed when necessary or stimulated by other factors [25]. Spermatogenesis can be elucidated by determining how genes are controlled and expressed at the exact time and what genes function in specific stages. Different methods, such as RT-PCR, Northern blot, and in situ hybridization, have been employed to detect genes [26–28], but these methods can focus on some genes at a time. As such, advanced approaches have been utilized to detect gene expression profiles and analyze gene expression with high throughput [29, 30]. For instance, cDNA microarray technology has been applied to investigate gene expression profiles in mouse spermatogenic cells in different stages of spermatogenesis [18]. Numerous genes have been detected. 181 genes in primitive type A spermatogonia, 256 genes in type B spermatogonia, 221 genes in preleptotene spermatocytes, 160 genes in pachytene spermatocytes, 141 genes in round spermatids, and 126 genes in elongating spermatids; these findings provide additional information for spermatogenesis research [18]. RNA sequencing technology has also revealed that gene expression patterns have significantly differed between mouse type B spermatogonia and primary spermatocytes; therefore, the basic molecular mechanism of early spermatogenesis is strongly related to the dynamics of cell junction [19].
In the present study, GC-1 spg cells were used as a model to investigate the role of melatonin in spermatogenesis. The physiological concentration of melatonin significantly promoted the proliferation of GC-1 spg cells. Melatonin can also enhance the differentiation of spermatogonial stem cells into haploid germ cells in sheep testis in vitro by activating melatonin membrane receptors and spermatogenesis-related gene expression [17]. Accumulated studies revealed that melatonin processes a beneficial function on human sperm motility [31], ram and pig sperm quality [32, 33]. Melatonin also improves the quality of thawed bovine serum by decreasing lipid peroxidation and enhancing the activity of antioxidant enzymes [34]. Intraperitoneal injection with 50 mg/kg melatonin significantly decreased levels of oxidative stress enzymes and lipid peroxidation in a rat model of testicular torsion induced by ischemia-reperfusion (I/R) injury [35]. Anticancer drugs can cause testicular toxicity, melatonin administration helped protect against drug-induced testicular toxicity [36, 37]. The protective roles of melatonin in testicular damage suggest that it may have clinical applications as a free radical scavenger and indirect antioxidant in the treatment of testicular damage.
Our previous study showed that melatonin can influence the testicular functions by regulating the differential expression of microRNAs in mouse testis [11]. Furthermore, melatonin can promote the proliferation of GC-1 spg cells by downregulating the expression of miR-16 and thus indicated that melatonin may regulate spermatogenesis [11]. To further explore the mechanism of melatonin in regulating spermatogenesis, we performed transcriptome sequencing and provided an in-depth analysis of the transcriptome of GC-1 spg cells exposed to melatonin. GO and KEGG analyses showed that differentially expressed genes, including Ccne1, Ercc1, and Mt2, were implicated in cell proliferation. Ccne1 is a positive regulator of G1/S phase transition and important factor in cell cycle [38]. The knockdown of Ccne1 inhibits cell growth [39]. Ccne1 is overexpressed in most tumor cells [40]. The antitumor effects of melatonin are associated with the downregulated Ccne1 expression [41]. Ercc1 is essential for the repair of nucleotide excision and DNA double-strand breaks [42]. The disruption of mouse Ercc1 causes cell cycle abnormalities in the liver and kidney before weaning [43]. Interestingly, Ercc1 is expressed in the testis at high levels, and the knockout of Ercc1 in mice increases the level of DNA strand breaks; oxidative DNA damage is also observed in Ercc1-deficient testis, and increased apoptosis is noted in male germ cells [44]. Melatonin elicits beneficial effects on the testis by preventing oxidative stress-evoked DNA damage in human spermatozoa and alleviating Cd-induced cellular stress and germ cell apoptosis, which may be associated with the melatonin-regulated expression of Ercc1 [45]. As another important upregulated gene, Mt2 is a member of metal-binding proteins, such as metallothioneins (Mts) [46]. MTs are divided into four subgroups that show tissue-specific expression and various functions. Mt genes are also actively expressed in male germ cells of mice [47] and rats [48]. The Mt1 and Mt2 expression levels in the testes from sexually mature adults are 10-fold higher than those in the livers of control adults [49]. The increased expression of Mt1 and Mt2 induced by melatonin in the testis mainly sequesters nonessential toxic heavy metals [50, 51]. However, studies have yet to provide evidence supporting high Mt2 levels in GC-1 spg cells associated with melatonin-promoted cell proliferation. The overexpression of Mt2 promoted the proliferation of GC-1 spg cells, whereas the knockdown of Mt2 significantly suppressed the proliferation of GC-1 spg cells and induced their apoptosis. These results indicated that Mt2 might be involved in melatonin-induced cell proliferation. The elevated level of Mts has been found in rapidly proliferating cells and various tumors [52–54]. In the analysis of the cycle of human colonic cancer cells, the maximum concentration of Mt is in successive late G phases and G1/S transition [53]. However, silencing Mt2 in breast cancer cells inhibits the progression the cell cycle from G1 phase to S phase [55]. Therefore, Mt2 could modulate cell proliferation by regulating cell cycle progression.
ERK1/2 signaling pathway mediates various cellular behaviors, including cell growth and apoptosis, in response to extracellular stimulation [56, 57]. The ERK1/2 pathway may also control the actions of melatonin [58, 59]. To identify whether ERK1/2 pathway is involved in the melatonin-promoted proliferation of GC-1 spg cells, we examined the ERK1/2 signaling pathway through Western blot and found that melatonin activated the phosphorylation of ERK1/2 in GC-1 spg cells. However, the suppression of ERK1/2 pathway significantly inhibited the proliferation of GC-1 spg cells and downregulated the expression of Mt2 induced by melatonin.
In summary, our study revealed the expression profile of genes in GC-1 spg cells stimulated by melatonin. Melatonin could induce the differential expression of genes in GC-1 spg cells. The melatonin-promoted proliferation of GC-1 spg cells might be associated with the upregulation of genes involved in cell proliferation. The functional characterization of genes demonstrated that Mt2 was regulated by melatonin and mediated melatonin-induced cell proliferation. The melatonin-induced expression of Mt2 was mediated by the ERK1/2 pathway during GC-1 spg cell proliferation. These findings revealed that melatonin might be essential for spermatogenesis. However, the effects of melatonin on germ cell differentiation have yet to be observed because of the absence of a melatonin null model. Hence, further studies using genetically inactivated mice may help clarify their effects on this process.
MATERIALS AND METHODS
Cell culture
The mouse-derived spermatogonial cell line, GC-1 spg cells, (ATCC, VA, USA) were cultured under controlled conditions (37°C, 5% CO2) and grown in Roswell Park Memorial Institute (RPMI-1640, Hyclone, MA, USA ) medium supplemented with 10% FBS (Gibco, CA, USA) and antibiotics (penicillin 100 IU/mL and streptomycin 100 μg/mL, Hyclone, MA, USA ). At 80% confluence, the cells were subcultured with 0.05% trypsin (Gibco, CA, USA) for further experiments.
Cell proliferation assay
The proliferation of GC-1 spg cells were evaluated by cell counting kit 8 (CCK8) assay (Beyotime Biotech, Haimen, China). In brief, these cells were seeded in 96-well plates at a density of 1 × 104 cells per well with the complete growth medium. 12 hrs after seeding, cells were serum starved for 12 hrs then different melatonin concentrations in complete growth medium were added for another 24 hrs treatment. At the end of treatment time, 10 μL of the CCK8 solution was added to each well and incubated at 37°C for 2 hrs. The number of cells was identified in triplicates by determining the absorbance in a 96-well microplate reader (ELx800, BioTek, Inc., IL, USA) at a wavelength of 450 nm (OD450).
High-resolution mRNA sequencing and data processing
GC-1 spg cells were seeded in a 6-well plate at a density of 1 × 106. Cells were serum free starved for 12 hrs when they reached 50% confluence. Then 10 nM melatonin in complete growth medium was replaced for another 24 hrs treatment. Total RNA was isolated by using TRIzol reagent according to the manufacturer’s instructions (Invitrogen, CA, USA). To establish a paired-end RNA-seq library for transcriptome analysis, we isolated 10 μg of total poly(A) mRNA by using Sera-Mag Magnetic Oligo(dT) Beads (Sigma, MO, USA), broke the isolated mRNA into short fragments (Multiskan Spectrum; Thermo Scientific Multiskan), and added adapters. An appropriate size of the fragments was selected and amplified through PCR. Fragmentation buffer was added to divide the mRNA into short fragments. These short fragments were used as templates and a random hexamer primer was utilized to synthesize the first-strand cDNA. The second-strand cDNA was also synthesized with 20 μL of the second-strand buffer (Invitrogen, CA, USA), 10 mM of dNTP Mix, 5 U/μL of RNaseH, and 10 U/μL of DNA polymerase I. The short fragments were purified with a Qiaquick PCR purification kit (Qiagen, Dusseldorf, Germany) and resolved with EB buffer for the end reparation and poly(A) addition. The short fragments were then connected with sequencing adapters. After agarose gel electrophoresis was conducted, suitable fragments as templates were selected for PCR amplification. The paired-end (PE) RNA-seq libraries were prepared and sequenced on an Illumina HiSeq™ 2500 platform according to the manufacturer’s protocols. Data analysis and base calling were performed with Illumina instrument software. High-throughput sequencing was carried out with Beijing ori-gene science and technology corp., ltd (Beijing, China). Differential expression (DE) results were based on the fold changes of the expression levels (p < 0.05) obtained by using Cufflinks (version 2.0.2). The DE genes were clustered through COG analysis by performing Fisher’s and chi-square tests.
The unigenes were used for BLASTX and annotation against protein databases, including nonredundant (nr), SwissPort, COG, KEGG, and GO, with a cutoff E-value of 0.00001. After the GO annotation for each unigene was obtained, GO functional classification for all unigenes was conducted and the distribution of gene functions was elucidated in WEGO. The KEGG pathway annotation was performed using blastall software instead of the KEGG database.
Real-time quantitative PCR
GC-1 spg cells were seeded in a 12-well plate at a density of 5 × 105. Cells were serum free starved for 12 hrs when they reached 50% confluence. Then 10 nM melatonin in complete growth medium was replaced for another 24 hrs treatment. Total RNA was extracted using a Total Cell RNA isolation kit (TIANGEN, Beijing, China). First-strand cDNA was synthesized using a PrimeScript first-strand cDNA synthesis kit (TaKaRa Biotechnology, Dalian, China) according to the manufacturer’s protocol. Real-time quantitative PCR was performed in triplicate in 96-well plates in an ABI Prism 7500 sequence detection system (Applied Biosystems, CA, USA). Reverse transcription was performed at 25°C for 10 min, 42°C for 20 min, and 85°C for 5 min. PCR was performed at 95°C for 5 min, followed by 40 cycles of 95°C for 15 s and 59°C for 30 s. Relative expression was normalized to Hprt RNA via ΔΔCt method. The sequences of the primers used for qRT-PCR are shown in Table 1.
Table 1: List of primers used for qRT-PCR
Primer name |
Sequence 5′-3′ |
|---|---|
Bbc3-F |
CCCAGAAATGGAGCCCAACT |
Bbc3-R |
CAAGGCTGGCAGTCCAGTAT |
Kcnn4-F |
CAACAAGGCGGAGAAACACG |
Kcnn4-R |
GTCCTTCCTTCGAGTGTGCT |
Mt2-F |
TCCATCACGCTCCTAGAACTC |
Mt2-R |
CAGGATCCATCGGAGGCACA |
Ercc1-F |
TAGCATCATCGTGAGCCCG |
Ercc1-R |
GCCCAGCACATAATCGGGAA |
Ccne1-F |
AGGCGAGGATGAGAGCAGTT |
Ccne1-R |
GGTGCAACTTTGGAGGGTAGA |
Hprt-F |
TCCTCCTCAGACCGCTTTT |
Hprt-R |
CCTGGTTCATCATCGCTAATC |
Ccna2-F |
CCTACCTCAAAGCGCCACAA |
Ccna2-R |
CTCCGAGCGACTCCTGTTTC |
Ccnb1-F |
TTGTGTGCCCAAGAAGATGCT |
Ccnb1-R |
GTACATCTCCTCATATTTGCTTGCA |
Ccne2-F |
CTGCTGCCGCCTTATGTCAT |
Ccne2-R |
TACACACTGGTGACAGCTGC |
Cdk6-F |
TCTCACAGAGTAGTGCATCGT |
Cdk6-R |
CGAGGTAAGGGCCATCTGAAAA |
Cdk4-F |
TTGTGCAGGTAGGAGTGCTG |
Cdk4-R |
TGCCAGAGATGGAGGAGTCT |
Bcl-2-F |
TGGAGAGCGTCAACAGGGAGA |
Bcl-2-R |
GCCAGGAGAAATCAAACAGAGGT |
Pcna-F |
TAAAGAAGAGGAGGCGGTAA |
Pcna-R |
TAAGTGTCCCATGTCAGCAA |
Western blot analysis
Protein lysates were prepared with lysis buffer supplemented with 1 mM PMSF (Beyotime Biotech, Haimen, China). Normalized proteins (20 μg) from each sample were electrophoresed with SDS-PAGE gels. Western blot analysis was performed with primary antibodies specific for MT2 (BS7352, Bioworld, Nanjing, China), ERK1/2 (BS1112, Bioworld, Nanjing, China), p-ERK1/2 (BS4621, Bioworld, Nanjing, China), or tubulin (AF0001, Beyotime Biotech, Haimen, China). After the reaction with HRP-conjugated goat anti-rabbit secondary antibody (A0208, Beyotime Biotech, Haimen, China) occurred, the membranes were detected with BeyoECL Plus kit (Beyotime Biotech, Haimen, China).
Cell cycle analysis
Flow cytometric analyses were performed to define the cell cycle distribution. In brief, GC-1 spg cells were seeded in a 12-well plate at a density of 5 × 105. Cells were serum free starved for 12 hrs when they reached 50% confluence. Then 10nM melatonin in complete growth medium was replaced for another 24 hrs treatment. Cells were harvested with trypsin and fixed with 70% ethanol at 4°C for 12 hrs. After ethanol was removed through centrifugation, the cells were washed with PBS and stained with 50 μg/mL propidium iodide and 50 μg/mL RNase A (Beyotime Biotech, Haimen, China) at 37°C for 30 min. Cell cycle distribution was analyzed by using a Cytomics FC500 flow cytometer (Beckman Coulter, Inc., CA, USA).
Apoptosis assay
Apoptosis was assessed by using an Annexin V-FITC apoptosis detection kit (Beyotime Biotech, Haimen, China). GC-1 spg cells were seeded in a 12-well plate at a density of 5 × 105. When cells occupied 50% confluence, 12 hrs serum free starvation followed by 24 hrs melatonin of 10nM with complete growth medium treatment was performed. Cells were collected by centrifuge, resuspended in 100 μL of buffer with 5 μL of Annexin V and 1 μL of propidium iodide, and incubated for 15 min at 25°C in the dark. Buffer (400 μL) was added to obtain the final volume of 500 μL. The cells were immediately analyzed with a Cytomics FC500 flow cytometer (Beckman Coulter Inc., CA, USA).
Plasmids, siRNAs, and transfection
pCMV6-empty plasmid vector (OriGene Technologies, MD, USA) and pCMV6 carrying the full-length mouse Mt2 cDNA (gene ID: 17750) were prepared by using a ClonExpress II one-step cloning kit (Vzayme, Nanjing, China). The sequences of siRNAs (RiboBio, Guangzhou, China) for Mt2 were used in the knockdown experiments. For the transient transfection of plasmids (200ng for 96-well plate and 1500ng for 12-well plate) or siRNAs (5pmol for 96-well plate and 40 pmol for 12-well plate), GC-1 spg cells were cultured in 96- or 12-well plates with complete growth medium until 70% confluence was reached. The cells were washed once with PBS and then transfected in a serum-free RPMI-1640 medium by using Lipofectamine 2000 (Invitrogen, CA, USA) according to the manufacturer’s instructions. After 12 hrs transfection, medium was changed to complete growth medium for another 24 hrs incubation.
Statistical analysis
Experiments were performed at least thrice. Differences between groups were compared through one-way ANOVA, followed by Dunnett’s multiple range or chi-square test for multiple comparisons in SPSS. Results were expressed as mean ± standard deviation. A p-value of < 0.05 was considered statistically significant.
Abbreviations
Mt2, metallothionein-2; ERK1/2, extracellular-signal-regulated kinase 1/2; KEGG, Kyoto Encyclopedia of Genes and Genomes; GO, Gene ontology; CCK8, cell counting kit 8; siRNA, small interfering RNA; MAPK, mitogen-activated protein kinase; Ccne1, cyclin E1; Ercc1, ERCC excision repair 1.
Author contributions
Chunjin Li and Xu Zhou conceived and designed the experiments, and wrote manuscript. Xiaoling Zhu and Shuxiong Chen performed the experiments, data mining, functional predictions, and analyzed the data. Lu Chen, Yun Zhao, Yanwen Jiang, Shan Gao, Fengge Wang, Zhuo Liu, Rong Fan, Liting Sun performed experimental work and contributed critical feedback on manuscript. All the authors have reviewed the manuscript.
ACKNOWLEDGMENTS
This work was supported by the National Natural Science Foundation of China [31672417, 31301224, 31301969] and National key research and development program of China [2016YFD0500503].
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
REFERENCES
1. Chianese R, Cobellis G, Chioccarelli T, Ciaramella V, Migliaccio M, Fasano S, Pierantoni R, Meccariello R. Kisspeptins, Estrogens and Male Fertility. Curr Med Chem. 2016; 23:4070–91.
2. Corradi PF, Corradi RB, Greene LW. Physiology of the Hypothalamic Pituitary Gonadal Axis in the Male. Urol Clin North Am. 2016; 43:151–62. https://doi.org/10.1016/j.ucl.2016.01.001.
3. Rudolph LM, Bentley GE, Calandra RS, Paredes AH, Tesone M, Wu TJ, Micevych PE. Peripheral and Central Mechanisms Involved in the Hormonal Control of Male and Female Reproduction. J Neuroendocrinol. 2016; 28. https://doi.org/10.1111/jne.12405.
4. Li C, Zhou X. Melatonin and male reproduction. Clin Chim Acta. 2015; 446:175–80. https://doi.org/10.1016/j.cca.2015.04.029.
5. Yelamanchi SD, Kumar M, Madugundu AK, Gopalakrishnan L, Dey G, Chavan S, Sathe G, Mathur PP, Gowda H, Mahadevan A, Shankar SK, Prasad TS. Characterization of human pineal gland proteome. Mol Biosyst. 2016; 12:3622–32. https://doi.org/10.1039/c6mb00507a.
6. Diaz Lopez B, Diaz Rodriguez E, Urquijo C, Fernandez Alvarez C. Melatonin influences on the neuroendocrine-reproductive axis. Ann N Y Acad Sci. 2005; 1057:337–64. https://doi.org/10.1196/annals.1356.026.
7. Succu S, Berlinguer F, Pasciu V, Satta V, Leoni GG, Naitana S. Melatonin protects ram spermatozoa from cryopreservation injuries in a dose-dependent manner. J Pineal Res. 2011; 50:310–8. https://doi.org/10.1111/j.1600-079X.2010.00843.x.
8. du Plessis SS, Hagenaar K, Lampiao F. The in vitro effects of melatonin on human sperm function and its scavenging activities on NO and ROS. Andrologia. 2010; 42:112–6.
9. Pandi-Perumal SR, Trakht I, Srinivasan V, Spence DW, Maestroni GJ, Zisapel N, Cardinali DP. Physiological effects of melatonin: role of melatonin receptors and signal transduction pathways. Prog Neurobiol. 2008; 85:335–53. https://doi.org/10.1016/j.pneurobio.2008.04.001.
10. Cebrian-Perez JA, Casao A, Gonzalez-Arto M, dos Santos Hamilton TR, Perez-Pe R, Muino-Blanco T. Melatonin in sperm biology: breaking paradigms. Reprod Domest Anim. 2014; 49 Suppl 4:11–21. https://doi.org/10.1111/rda.12378.
11. Li C, Chen S, Li H, Chen L, Zhao Y, Jiang Y, Liu Z, Liu Y, Gao S, Wang F, Yu J, Wang H, Rao J, et al. MicroRNA-16 Modulates Melatonin-Induced Cell Growth in the Mouse-Derived Spermatogonia Cell Line GC-1 spg Cells by Targeting Ccnd1. Biol Reprod. 2016; 95:57. https://doi.org/10.1095/biolreprod.115.138313.
12. De Gendt K, Swinnen JV, Saunders PTK, Schoonjans L, Dewerchin M, Devos A, Tan K, Atanassova N, Claessens F, Lecureuil C, Heyns W, Carmeliet P, Guillou F, et al. A Sertoli cell-selective knockout of the androgen receptor causes spermatogenic arrest in meiosis. Proc Natl Acad Sci U S A. 2004; 101:1327–32. https://doi.org/10.1073/pnas.0308114100.
13. Alves MG, Rato L, Carvalho RA, Moreira PI, Socorro S, Oliveira PF. Hormonal control of Sertoli cell metabolism regulates spermatogenesis. Cell Mol Life Sci. 2013; 70:777–93. https://doi.org/10.1007/s00018-012-1079-1.
14. Yang WC, Tang KQ, Fu CZ, Riaz H, Zhang Q, Zan LS. Melatonin regulates the development and function of bovine Sertoli cells via its receptors MT1 and MT2. Anim Reprod Sci. 2014; 147:10–6. https://doi.org/10.1016/j.anireprosci.2014.03.017.
15. Rocha CS, Martins AD, Rato L, Silva BM, Oliveira PF, Alves MG. Melatonin alters the glycolytic profile of Sertoli cells: implications for male fertility. Mol Hum Reprod. 2014; 20:1067–76. https://doi.org/10.1093/molehr/gau080.
16. Komeya M, Ogawa T. Spermatogonial stem cells: Progress and prospects. Asian J Androl. 2015; 17:771–5. https://doi.org/10.4103/1008-682X.154995.
17. Deng SL, Chen SR, Wang ZP, Zhang Y, Tang JX, Li J, Wang XX, Cheng JM, Jin C, Li XY, Zhang BL, Yu K, Lian ZX, et al. Melatonin promotes development of haploid germ cells from early developing spermatogenic cells of Suffolk sheep under in vitro condition. J Pineal Res. 2016; 60:435–47. https://doi.org/10.1111/jpi.12327.
18. Yu Z, Guo R, Ge Y, Ma J, Guan J, Li S, Sun X, Xue S, Han D. Gene expression profiles in different stages of mouse spermatogenic cells during spermatogenesis. Biol Reprod. 2003; 69:37–47. https://doi.org/10.1095/biolreprod.102.012609.
19. Zhang XJ, Hao LL, Meng LJ, Liu ML, Zhao LN, Hu F, Ding CB, Wang Y, He BL, Pan YX, Fang W, Chen J, Hu SN, et al. Digital Gene Expression Tag Profiling Analysis of the Gene Expression Patterns Regulating the Early Stage of Mouse Spermatogenesis. Plos One. 2013; 8. https://doi.org/ARTN e5868010.1371/journal.pone.0058680.
20. Kim ST, Omurtag K, Moley KH. Decreased Spermatogenesis, Fertility, and Altered Slc2A Expression in Akt1-/- and Akt2-/- Testes and Sperm. Reprod Sci. 2012; 19:31–42. https://doi.org/10.1177/1933719111424449.
21. Almog T, Naor Z. Mitogen activated protein kinases (MAPKs) as regulators of spermatogenesis and spermatozoa functions. Mol Cell Endocrinol. 2008; 282:39–44. https://doi.org/10.1016/j.mce.2007.11.011.
22. Young JC, Wakitani S, Loveland KL. TGF-beta superfamily signaling in testis formation and early male germline development. Semin Cell Dev Biol. 2015; 45:94–103. https://doi.org/10.1016/j.semcdb.2015.10.029.
23. Shi D, Xiao X, Wang J, Liu L, Chen W, Fu L, Xie F, Huang W, Deng W. Melatonin suppresses proinflammatory mediators in lipopolysaccharide-stimulated CRL1999 cells via targeting MAPK, NF-kappaB, c/EBPbeta, and p300 signaling. J Pineal Res. 2012; 53:154–65. https://doi.org/10.1111/j.1600-079X.2012.00982.x.
24. Jin H, Wang Y, Zhou L, Liu L, Zhang P, Deng W, Yuan Y. Melatonin attenuates hypoxic pulmonary hypertension by inhibiting the inflammation and the proliferation of pulmonary arterial smooth muscle cells. J Pineal Res. 2014; 57:442–50. https://doi.org/10.1111/jpi.12184.
25. Shima JE, McLean DJ, McCarrey JR, Griswold MD. The murine testicular transcriptome: characterizing gene expression in the testis during the progression of spermatogenesis. Biol Reprod. 2004; 71:319–30. https://doi.org/10.1095/biolreprod.103.026880.
26. Singer-Sam J, Robinson MO, Bellve AR, Simon MI, Riggs AD. Measurement by quantitative PCR of changes in HPRT, PGK-1, PGK-2, APRT, MTase, and Zfy gene transcripts during mouse spermatogenesis. Nucleic Acids Res. 1990; 18:1255–9.
27. Marchand O, Govoroun M, D’Cotta H, McMeel O, Lareyre JJ, Bernot A, Laudet V, Guiguen Y. DMRT1 expression during gonadal differentiation and spermatogenesis in the rainbow trout, Oncorhynchus mykiss. Biochim Biophys Acta. 2000; 1493:180–7.
28. Niederberger C, Agulnik AI, Cho Y, Lamb D, Bishop CE. In situ hybridization shows that Dazla expression in mouse testis is restricted to premeiotic stages IV-VI of spermatogenesis. Mamm Genome. 1997; 8:277–8.
29. Shiraki T, Kondo S, Katayama S, Waki K, Kasukawa T, Kawaji H, Kodzius R, Watahiki A, Nakamura M, Arakawa T, Fukuda S, Sasaki D, Podhajska A, et al. Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage. Proc Natl Acad Sci U S A. 2003; 100:15776–81. https://doi.org/10.1073/pnas.2136655100.
30. Ak H, Zeybek B, Atay S, Askar N, Akdemir A, Aydin HH. Microarray gene expression analysis of uterosacral ligaments in uterine prolapse. Clin Biochem. 2016; 49:1238–42. https://doi.org/10.1016/j.clinbiochem.2016.08.004.
31. Ortiz A, Espino J, Bejarano I, Lozano GM, Monllor F, Garcia JF, Pariente JA, Rodriguez AB. High endogenous melatonin concentrations enhance sperm quality and short-term in vitro exposure to melatonin improves aspects of sperm motility. J Pineal Res. 2011; 50:132–9. https://doi.org/10.1111/j.1600-079X.2010.00822.x.
32. Casao A, Vega S, Palacin I, Perez-Pe R, Lavina A, Quintin FJ, Sevilla E, Abecia JA, Cebrian-Perez JA, Forcada F, Muino-Blanco T. Effects of melatonin implants during non-breeding season on sperm motility and reproductive parameters in Rasa Aragonesa rams. Reprod Domest Anim. 2010; 45:425–32. https://doi.org/10.1111/j.1439-0531.2008.01215.x.
33. Jang HY, Kim YH, Kim BW, Park IC, Cheong HT, Kim JT, Park CK, Kong HS, Lee HK, Yang BK. Ameliorative effects of melatonin against hydrogen peroxide-induced oxidative stress on boar sperm characteristics and subsequent in vitro embryo development. Reprod Domest Anim. 2010; 45:943–50. https://doi.org/10.1111/j.1439-0531.2009.01466.x.
34. Ashrafi I, Kohram H, Ardabili FF. Antioxidative effects of melatonin on kinetics, microscopic and oxidative parameters of cryopreserved bull spermatozoa. Anim Reprod Sci. 2013; 139:25–30. https://doi.org/10.1016/j.anireprosci.2013.03.016.
35. Parlaktas BS, Atilgan D, Ozyurt H, Gencten Y, Akbas A, Erdemir F, Uluocak N. The biochemical effects of ischemia-reperfusion injury in the ipsilateral and contralateral testes of rats and the protective role of melatonin. Asian J Androl. 2014; 16:314–8. https://doi.org/10.4103/1008-682X.122202.
36. Lee KM, Lee IC, Kim SH, Moon C, Park SH, Shin DH, Park SC, Kim HC, Kim JC. Melatonin attenuates doxorubicin-induced testicular toxicity in rats. Andrologia. 2012; 44:796–803. https://doi.org/10.1111/j.1439-0272.2011.01269.x.
37. Chabra A, Shokrzadeh M, Naghshvar F, Salehi F, Ahmadi A. Melatonin ameliorates oxidative stress and reproductive toxicity induced by cyclophosphamide in male mice. Hum Exp Toxicol. 2014; 33:185–95. https://doi.org/10.1177/0960327113489052.
38. Wang F, Fu X-D, Zhou Y, Zhang Y. Down-regulation of the cyclin E1 oncogene expression by microRNA-16-1 induces cell cycle arrest in human cancer cells. BMB Rep. 2009; 42:725–30.
39. Zhang X, Hu S, Wang L, Yan B, Zhao J, Yang A, Zhang R. MicroRNA-7 arrests cell cycle in G1 phase by directly targeting CCNE1 in human hepatocellular carcinoma cells. Biochem Biophys Res Commun. 2014; 443:1078–84. https://doi.org/10.1016/j.bbrc.2013.12.095.
40. Luo Q, Li X, Li J, Kong X, Zhang J, Chen L, Huang Y, Fang L. MiR-15a is underexpressed and inhibits the cell cycle by targeting CCNE1 in breast cancer. Int J Oncol. 2013; 43:1212–8.
41. Shen CJ, Chang CC, Chen YT, Lai CS, Hsu YC. Melatonin Suppresses the Growth of Ovarian Cancer Cell Lines (OVCAR-429 and PA-1) and Potentiates the Effect of G1 Arrest by Targeting CDKs. Int J Mol Sci. 2016; 17. https://doi.org/10.3390/ijms17020176.
42. Olaussen KA, Dunant A, Fouret P, Brambilla E, Andre F, Haddad V, Taranchon E, Filipits M, Pirker R, Popper HH, Stahel R, Sabatier L, Pignon JP, et al. DNA repair by ERCC1 in non-small-cell lung cancer and cisplatin-based adjuvant chemotherapy. N Engl J Med. 2006; 355:983–91.
43. Weeda G, Donker I, de Wit J, Morreau H, Janssens R, Vissers CJ, Nigg A, van Steeg H, Bootsma D, Hoeijmakers JH. Disruption of mouse ERCC1 results in a novel repair syndrome with growth failure, nuclear abnormalities and senescence. Curr Biol. 1997; 7:427–39.
44. Hsia K-T, Millar MR, King S, Selfridge J, Redhead NJ, Melton DW, Saunders PTK. DNA repair gene Ercc1 is essential for normal spermatogenesis and oogenesis and for functional integrity of germ cell DNA in the mouse. Development. 2003; 130:369–78.
45. Ji YL, Wang H, Meng C, Zhao XF, Zhang C, Zhang Y, Zhao M, Chen YH, Meng XH, Xu DX. Melatonin alleviates cadmium-induced cellular stress and germ cell apoptosis in testes. J Pineal Res. 2012; 52:71–9. https://doi.org/10.1111/j.1600-079X.2011.00921.x.
46. Palumaa P, Tammiste I, Kruusel K, Kangur L, Jornvall H, Sillard R. Metal binding of metallothionein-3 versus metallothionein-2: lower affinity and higher plasticity. Biochim Biophys Acta. 2005; 1747:205–11. https://doi.org/10.1016/j.bbapap.2004.11.007.
47. De SK, Enders GC, Andrews GK. High levels of metallothionein messenger RNAs in male germ cells of the adult mouse. Mol Endocrinol. 1991; 5:628–36. https://doi.org/10.1210/mend-5-5-628.
48. Suzuki JS, Kodama N, Molotkov A, Aoki E, Tohyama C. Isolation and identification of metallothionein isoforms (MT-1 and MT-2) in the rat testis. Biochem J. 1998; 334:695–701.
49. Salehi-Ashtiani K, Widrow RJ, Markert CL, Goldberg E. Testis-specific expression of a metallothionein I-driven transgene correlates with undermethylation of the locus in testicular DNA. Proc Natl Acad Sci USA. 1993; 90:8886–90.
50. Reiter RJ, Mayo JC, Tan D-X, Sainz RM, Alatorre-Jimenez M, Qin L. Melatonin as an antioxidant: under promises but over delivers. J Pineal Res. 2016; 61:253–78.
51. Alonso-Gonzalez C, Mediavilla D, Martinez-Campa C, Gonzalez A, Cos S, Sanchez-Barcelo EJ. Melatonin modulates the cadmium-induced expression of MT-2 and MT-1 metallothioneins in three lines of human tumor cells (MCF-7, MDA-MB-231 and HeLa). Toxicol Lett. 2008; 181:190–5. https://doi.org/10.1016/j.toxlet.2008.07.021.
52. Nagel WW, Vallee BL. Cell cycle regulation of metallothionein in human colonic cancer cells. Proc Natl Acad Sci U S A. 1995; 92:579–83.
53. Cherian MG, Apostolova MD. Nuclear localization of metallothionein during cell proliferation and differentiation. Cell Mol Biol (Noisy-le-grand). 2000; 46:347–56.
54. Yap X, Tan HY, Huang J, Lai Y, Yip GW, Tan PH, Bay BH. Over-expression of metallothionein predicts chemoresistance in breast cancer. J Pathol. 2009; 217:563–70. https://doi.org/10.1002/path.2489.
55. Lai Y, Lim D, Tan PH, Leung TKC, Yip GWC, Bay BH. Silencing the Metallothionein-2A gene induces entosis in adherent MCF-7 breast cancer cells. Anat Rec (Hoboken). 2010; 293:1685–91.
56. Bonni A, Brunet A, West AE, Datta SR, Takasu MA, Greenberg ME. Cell survival promoted by the Ras-MAPK signaling pathway by transcription-dependent and -independent mechanisms. Science. 1999; 286:1358–62.
57. Mebratu Y, Tesfaigzi Y. How ERK1/2 activation controls cell proliferation and cell death is subcellular localization the answer? Cell Cycle. 2009; 8:1168–75.
58. Chen X, Li X, Du Z, Shi W, Yao Y, Wang C, He K, Hao A. Melatonin promotes the acquisition of neural identity through extracellular-signal-regulated kinases 1/2 activation. J Pineal Res. 2014; 57:168–76.
59. Cui P, Yu M, Luo Z, Dai M, Han J, Xiu R, Yang Z. Intracellular signaling pathways involved in cell growth inhibition of human umbilical vein endothelial cells by melatonin. J Pineal Res. 2008; 44:107–14. https://doi.org/10.1111/j.1600-079X.2007.00496.x.

