INTRODUCTION
Tyrosine kinase inhibitors (TKIs) targeting the epidermal growth factor receptor (EGFR) have substantially improved the quality of life and survival of advanced non-small-cell lung cancer (NSCLC) patients [1–5]. However, in daily clinical practice, many questions concerning the management of patients undergoing TKI therapies remain unanswered. For example, conventional monitoring of TKI-treated patients relies on morphologic tumor changes, which are generally assessed by computed tomography (CT) scans and response evaluation criteria in solid tumors (RECIST). However, several studies have indicated that RECIST assessment may have some limitations when measuring tumor response to TKIs [6–8]. Oncologists usually continue to treat NSCLC patients harbouring sensitizing EGFR mutation with TKIs for extended periods after RECIST progression [9–11] although this may be continuing ineffective treatments. Indeed, it has been reported that the p.T790M resistance mutation can be effectively detected in plasma of advanced lung cancer patients several months prior to clinical progression [12, 13]. In the same way, dose readjustment due to toxicity is performed empirically and whether drug efficacy is compromised by dose reduction is difficult to measure in daily clinical practice.
This study assessed dynamic changes in EGFR mutation quantification using digital PCR (dPCR) methodology in longitudinally collected plasma samples from NSCLC patients with activating EGFR mutations. We evaluated the clinical information that EGFR mutation levels in plasma samples can provide in the context of daily clinical practice, not only in terms of tumor response, but also with respect to pharmacokinetics and drug interactions and individualization of treatment.
RESULTS
Study cohort
This study reports daily clinical practice data obtained from 18 NSCLC patients. Routine follow-up examinations were performed by a medical oncologist every 3 weeks for the first 3 months, and every 12 weeks thereafter or as required according to the oncologist’s criteria. The pathological characteristics of the study population are summarized in Table 1.
Table 1: Clinico-pathological characteristics of the study population
Median age at diagnosis |
59 years (40–73) |
N |
% |
|---|---|---|---|
Sex |
Women |
8 |
44% |
Men |
10 |
55% |
|
Smoking status |
Former/quit |
10 |
55% |
Never |
8 |
44% |
|
Histology |
Adenocarcinoma |
18 |
100% |
UICC Stage |
III |
1 |
5% |
IV |
17 |
94% |
|
EGFR mutation |
exon 19 deletion |
12 |
67% |
exon 20 insertion |
1 |
5% |
|
L858R |
5 |
28% |
|
Number of treatment lines monitored |
1 line |
3 |
17% |
throughout the study |
2 lines |
10 |
55% |
3 lines |
5 |
28% |
An average of 10 cfDNA samples were analysed per patient. cfDNA from all blood samples was analysed for the amount of sensitizing EGFR mutation. The p.T790M mutation was concomitantly analysed in all cfDNA samples. ctDNA fluctuations measured as mutated copies/ml or as the ratio mutant allele fraction showed similar patterns. According to our data, in positive case samples the ratio of mutant DNA molecules vs total DNA molecules ranged from 0.10% to 18.5% and the number of mutated copies in positive samples ranged from 144 to 328570 copies/ml.
EGFR mutation levels to monitor non-standard therapies
Overall, changes in EGFR mutation levels correlated with treatment responses observed in radiology assessments. Our results consistently showed that the detection of the p.T790M resistant mutation in blood, as well as an increment in the quantification of the original sensitizing EGFR mutation in serial plasma samples correlated with the assessment of progressive disease (PD) whereas the opposite occurred in responding patients (Supplementary Figure 1). Importantly, a substantial decrease of p.T790M and the sensitizing mutation levels were notice in a patient (03LSM) treated with cetuximab plus afatinib, who maintained in stable disease (according to radiological assessments) 493 days after the initiation of treatment, suggesting that EGFR mutation levels in plasma might be useful to monitor non-standardized treatments (Supplementary Figure 1). Of note, a total of 19 samples were obtained and analysed from patient 03LSM, which implied obtaining an average of one sample every 26 days during the 493 days of follow up of this patient. This close disease monitoring would be unthinkable if performing exclusively imaging tests.
EGFR mutation levels tracking as a tool to monitor dose reduction and drug interactions
Dose adjustments were required in 5 patients (04BAA, 07AB, 18MACC, 13CGG and 39YPG). In four of them (04BAA, 07AB, 07MACC and 13CGG), increased EGFR mutation levels were detected after TKI dose readjustment (Figures 1 and 2), and correlated with a subsequent diagnosis of PD. In contrast, no sensitizing mutation increase was observed in the remaining patient (39YPG) after TKI dose reduction and the patient remained in partial response (PR) (Figure 1). In addition, in three cases (04BAA, 07AB and 07MACC) a drop in EGFR mutation levels were observed following the reintroduction of TKI at full dose.
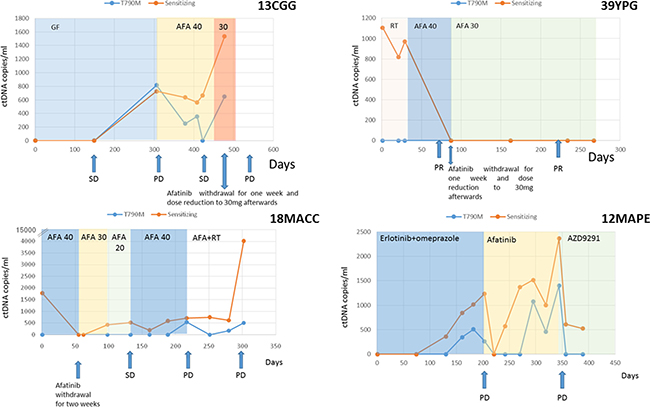
Figure 1: ctDNA levels (copies/ml) of patients 13CGG, 29YPG, 18MACC and 12MAPE. Therapies are denoted by different colored shading. Disease status as ascertained on a CT-scan at different time points is marked with arrows. In cases 13CGG and 18MACC an increase in ctDNA is observed after TKI dose reduction correlating with subsequent tumor progression. Increasing levels on ctDNA in case 12MAPE suggest a possible interaction between erlotinib and omeprazole. (Abbreviations: SD = stable disease, PR = partial response, PD = progressive disease, GF = gefitinib, AFA = afatinib, PMT = Pemetrexed, E = erlotinib, RT = radiotherapy).
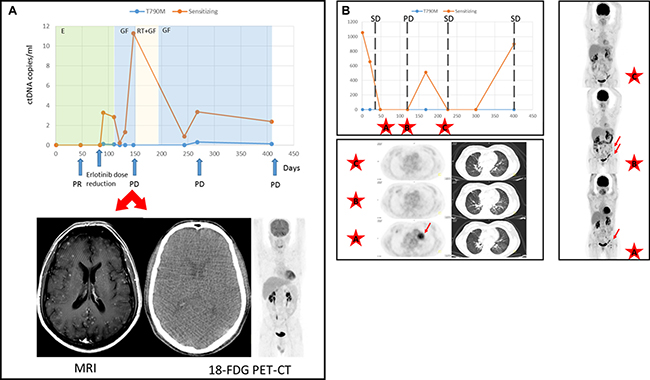
Figure 2: Discrepancies in the assessment of tumor response between ctDNA and TAC. (A) ctDNA levels (copies/ml) of patient 07AB. Therapies are denoted by different colored shading. Disease status as ascertained on a CT-scan at different time points is marked with arrows. As shown, the 18-FDG PET-CT scan at day 150 failed to detect brain metastases that were however diagnosed by MRI. (Abbreviations: PR = partial response, PD = progressive disease, GF = gefitinib, RT = radiotherapy). (B) Mutation tracking profile of case 11MCMS and treatment outcome assessed according to RECIST criteria v.1.1. SD = stable disease, PD = progressive disease. 18-FDG PET-CT scans at time points are marked with a discontinuous line. Images from data points A, B and C are presented. Lung tumor and adenopathies are denoted by arrows.
Finally, one case (12MAPE) corresponded to a patient receiving erlotinib who was concomitantly taking omeprazole without medical indication, and increasing levels of EGFR mutation were detected during co-administration of both drugs. As shown in Figure 1, a reduction to undetectable levels in EGFR mutation levels were demonstrated immediately after omeprazole withdrawal although these levels increased again soon after.
EGFR mutation levels for monitoring oligometastatic disease treatment
During the study, eight patients (02RGB, 07AB, 17VVM, 05JBP, 18MACC, 19MLCG, 30MMP and 32LICV) received loco-regional radiotherapy (RT). As shown in Supplementary Figure 1, changes in EGFR mutation levels were observed during RT, which correlated with responses ascertained by the radiologist. Importantly, EGFR mutation levels in plasma dropped after RT in all patients (Table 2) except for one case (18MACC) who progressed after 64 days. The short time to progression in the latter case suggests a systemic rather than oligometastatic disease. This could potentially be suspected from the increasing EGFR mutation levels in plasma after RT. Note that in patient 07AB, RT was given with palliative intent, due to the patient´s diagnosis of leptomeningeal carcinomatosis.
Table 2: ctDNA fluctuations in patients undergoing radiotherapy (RT) and time to progression after RT (days)
Patient |
Progression after RT |
Time to progression (days) |
ctDNA fluctuation |
|---|---|---|---|
02RGB |
no |
- |
↓ |
07AB |
yes |
98 |
↓ |
17VVM |
no |
- |
↓ |
05JBP |
yes |
223 |
↓ |
18MACC |
yes |
64 |
↑ |
19MLCG |
no |
- |
↓ |
30MMP |
no |
- |
↓ |
32LICV |
no |
↓ |
EGFR mutation levels in patients with doubtful diagnoses
Discrepancies with the imaging studies helped to resolve a doubtful diagnosis in patient 07AB, in which an 18FDG-PET/CT scan failed to detect tumor progression in the brain after second-line treatment with full-dose gefitinib. However, a head MRI confirmed multiple brain metastases which correlated with increased plasma levels of EGFR mutation levels within the same time frame (Figure 2A). Similarly, in patient 11MCMS, 18FDG-PET/CT assessments showed a decrease in size and metabolic activity of the pulmonary nodule that coincided with EGFR mutation quantification. Nevertheless, axillary, inguinal and retroperitoneal lymphadenopathies increased in size and number, and progressive disease was diagnosed after radiological imaging (Figure 2B). Remarkably, pathological evaluation of the inguinal adenopathy revealed an absence of malignancy. Notably, this patient was diagnosed with Sjogren’s syndrome and essential mixed cryoglobulinemia.
Body fluids to improve diagnostic yield of ctdna
In four cases, paired samples were collected from body fluid and plasma. Specifically the cytological evaluation of the pleural and pericardial effusions reveal the presence of lung adenocarcinoma cells in all three cases. Similarly, according to pathologist report the presence of lung adenocarcinoma metastasis in cerebrospinal fluid was confirmed. As shown in Table 3, EGFR mutation levels was always higher in the body fluids compared to plasma. In one case (24MAG) (Supplementary Figure 1) p.T790M mutation was not detected in re-biopsy FFPE or in the corresponding plasma sample. However, p.T790M mutation was detected in the ctDNA isolated from malignant pleural effusion at progression and sensitizing levels dramatically went down after treatment initiation with osimertinib.
Table 3: ctDNA mutant allele fraction (%) and ctDNA copies/ml in paired samples from plasma and body fluids
Patient |
cthNA Source |
p.T790M mutant allele fraction (%) |
p.T790M copies/ml |
Sensitizing mutant allele fraction (%) |
Sensitizing copies/ml |
|---|---|---|---|---|---|
16JLPG |
Plasma |
ND |
ND |
0.345 |
1804.5 |
CSF |
ND |
ND |
6.431 |
785.25 |
|
27FJAR |
Plasma |
0.144 |
436.812 |
0.524 |
676.062 |
PE |
10.253 |
334355.500 |
20.74 |
510110 |
|
24 MAG |
Plasma |
ND |
ND |
0.427 |
409.625 |
PE |
3.18 |
62225.17 |
9.66 |
23080.38 |
|
35MJSS |
Plasma |
0.11 |
155.56 |
0.80 |
1259.87 |
PCE |
9.66 |
3360850.33 |
26.88 |
9607335.10 |
Abbreviations: ND = not detected, CSF = cerebrospinal fluid, PE = pleural effusion, PEC pericardial fluid).
DISCUSSION
The majority of previous liquid biopsy studies in patients with EGFR mutations have primarily reported the feasibility of detecting resistance and sensitizing EGFR mutations and how much earlier tumor progression can be predicted using liquid biopsy than CT-scans [12–16]. However, the usefulness of EGFR mutation detection in plasma may not be limited to the early detection of progression, as it could also assist clinicians in daily clinical practice in terms of dose readjustments and drug interactions, among other aspects. For example, where there are no differences in the clinical effectiveness of different TKIs, in most cases the choice relies either on the oncologist’s experience and personal decision, or the expected toxicity. In this respect, we know that although the frequency of all-cause severe adverse effects is somewhat similar among TKIs, they may have differing toxicity profiles [17, 18]. Until now, whether dose reductions after adverse events could influence patient survival was unknown, since these events are relatively rare (less than 6%). From our data, it can be speculated that EGFR mutation level quantification can, at least partially, assist clinicians to lower treatment dose minimizing undesired outcomes. As presented, increasing EGFR mutation levels anticipated disease progression after TKI dose reduction whereas EGFR mutation levels dropped when patient maintained response despite dose reduction. Similarly, we report the clinical utility of EGFR mutation levels in plasma to detect drug-drug interactions that we know might occur even with some common, commercially available, soft drinks [19, 20]. As shown in Figure 1, increasing plasma levels of EGFR mutation were found in a patient (12MAPE) who was concomitantly taking erlotinib and omeprazole, suggesting that omeprazole may reduce effectivity of erlotinib upon co-administration.
We similarly consider that EGFR mutation levels in plasma could help to better redefine the use of determined treatment combinations, as occurred with patient 03LSM, who was treated with afatinib plus cetuximab after progression to afatinib. The rationale of this combination is based on solid preclinical data indicating that concomitant administration of the two drugs can eventually overcome p.T790M-mediated resistance [21]. However, in a phase Ib trial, dual inhibition of EGFR with afatinib and cetuximab was modestly active among patients with acquired resistance with an objective response rate of 29% and median progression free survival of 4.7 months (95% confidence interval, 4.3–6.4) [22]. Consequently, this combination has not been extended to routine clinical practice. Patient 03LSM exemplifies how tumor monitoring through liquid biopsy can open up new perspectives for the use of new therapies, anti-EGFR re-challenge, or even N-of-1 trials [23]. On one hand, liquid biopsy reduces the time for treatment outcome monitoring (from 4 weeks when based in imaging tests results to 5 days when based in liquid biopsy tests results, in our institution). On the other, changes in EGFR mutation levels in plasma can be observed within days (data not shown). This enables a closer real time follow up, getting ahead of the radiological response, and therefore improving decision-making regarding treatment selection, modification or continuation.
Finally, our results indicate that cancer associated body fluids are certainly a suitable source for biomarker testing that can extend EGFR mutation detection to bio-fluids other than blood (Table 3). As proved with patient 24MAG, who presented a p.T790M negative biopsy both in the solid tumor and blood, but was positive in the malignant pleural effusion. Malignant pleural effusions are frequently observed in advanced lung adenocarcinomas [24] and thoracentesis may be necessary for diagnosis and treatment. However, pleural fluid is usually discarded after the removal of cell component for cytological examination despite being helpful for EGFR mutation detection.
In conclusion, sensitizing and resistance mutation quantification in plasma throughout the course of a NSCLC cancer allows us to resolve doubts that frequently arise in daily clinical practice and reduce time for decision, which constitutes a major step towards personalized medicine.
MATERIALS AND METHODS
Study population and data management
A total of 18 cases from NSCLC patients were included in the study. Written informed consent was obtained from every patient. Eligible patients were both male and female patients with a pathologically confirmed diagnosis of stage IIIB-IV NSCLC tumor with an EGFR mutation in primary tumor tissue. A complete staging workup was performed prior to recruitment into the study. Blood samples were collected as follows: at diagnosis, when patients returned for re-evaluation and when appointed by a medical oncologist due to uncertainties about a patient’s clinical status, radiological assessments, toxicity events or TKI dose reduction. Demographic characteristics, clinico-pathological features, tumor mutation status, vital status, disease status, drug dose adjustments or discontinuation of medication were collected.
The study was approved by the Hospital Puerta de Hierro Ethics Committee (internal code PI/144-14) and was conducted in accordance with the precepts of the Code of Ethics of The World Medical Association (Declaration of Helsinki).
Laboratory procedures
For EGFR mutation levels quantification, peripheral whole blood was collected from each subject in a 5 ml EDTA tube containing a gel barrier (PPT™, BECTON DICKINSON) to separate the plasma from blood cells after centrifugation. All samples were processed as previously described [14, 25]. cfDNA was extracted using a starting volume of 1 ml of plasma with a Maxwell® RSC instrument (Promega), using the Maxwell® RSC ccfDNA Plasma Kit, as specified by the manufacturer and was eluted in 50 μl of the supplied buffer. In addition, biological fluids with malignant abnormal cytology, including malignant pleural (N = 2) and pericardial (N = 1) effusions and cerebrospinal fluid (N = 1) were also analysed for EGFR mutation levels. Biological fluids were centrifuged and the supernatant was used for cfDNA isolation using the same protocol. Germline DNA was obtained from blood leukocytes with a MagNA Pure LC total nucleic acid extraction kit in a MagNA Pure LC instrument (Roche Diagnostics, Penzberg, Germany). cfDNA samples were then analysed by dPCR using Rare Mutation Assays for p.T790M (AHRSROS), p.L858R (AHRSRSV), p.G719A (AHABH29), p.G719C (AH0JEWC), p.G719S (AHZAGP4), p.H773_V774insH (AH5I7PA), p.D770_N771insG (AH7031Q), p.L747_T751 > P (AHFA92K), p.L747_A750 > P (AHS1PY0), p.E746_T751 > A (AHHS6E0), p.E746_A750delELREA (AHLJ0XO), p.L747_T751delLREAT (AHCTDP3) and p.L747_S752delLREATS (AHGJ78R ) on a QuantStudio®3D Digital PCR System (Applied Biosystems, South San Francisco, CA), as previously described [14]. A wt control DNA was included in every run.
Tumor response evaluation
Computed tomography (CT) measurements and magnetic resonance imaging (MRI) were obtained as clinically indicated. The clinical response was evaluated according to RECIST criteria v1.1 combined with a blinded medical judgment of benefits from the treatment. Additionally, whole body 18F-fluoro-2-deoxy-D-glucose-positron emission tomography (18FDG-PET)- CT scans were performed as clinically indicated using a Siemens Biograph 6 True Point PET-CT (Siemens). A 350-450MBq 18F-FDG dose was administered 55-65 min prior to image acquisition. Reconstruction was performed using an iterative method and attenuation/scatter correction.
ACKNOWLEDGMENTS
The authors wish to thank Martin Hadley-Adams for assisting with the English language and preparation of the manuscript.
CONFLICTS OF INTEREST
The authors have declare no conflicts of interest.
FUNDING
This study was supported by Carlos III Institute of Health, Spanish Ministry of Science and Innovation, and European Regional Development Fund (grant number: PI16/01818 and PIE14/00064), A Romero is supported by Joan Rodés fellowship (grant number: JR14/00017) and CP pre-doctoral studies are supported by Jose Luís Castaño Foundation.
REFERENCES
1. Kris MG, Natale RB, Herbst RS, Lynch TJ Jr, Prager D, Belani CP, Schiller JH, Kelly K, Spiridonidis H, Sandler A, Albain KS, Cella D, Wolf MK, et al. Efficacy of gefitinib, an inhibitor of the epidermal growth factor receptor tyrosine kinase, in symptomatic patients with non-small-cell lung cancer. JAMA. 2003; 290:2149–58.
2. Zhou C, Wu YL, Chen G, Feng J, Liu XQ, Wang C, Zhang S, Wang J, Zhou S, Ren S, Lu S, Zhang L, Hu C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011; 12:735–42.
3. Taron M, Ichinose Y, Rosell R, Mok T, Massuti B, Zamora L, Mate JL, Manegold C, Ono M, Queralt C, Jahan T, Sanchez JJ, Sanchez-Ronco M, et al. Activating mutations in the tyrosine kinase domain of the epidermal growth factor receptor are associated with improved survival in gefitinib-treated chemorefractory lung adenocarcinomas. Clin Cancer Res. 2005; 11:5878–85.
4. Pao W, Miller V, Zakowski M, Doherty J, Politi K, Sarkaria I, Singh B, Heelan R, Rusch V, Fulton L, Mardis E, Kupfer D, Wilson R, et al. EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. ProcNatlAcadSci USA. 2004; 101:13306–13311.
5. Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small cell lung cancer to gefitinib. N Engl J Med. 2004; 350:2129–2139.
6. Shim SS, Lee KS, Kim BT, Chung MJ, Lee EJ, Han J, Choi JY, Kwon OJ, Shim YM, Kim S. Non-small cell lung cancer: prospective comparison of integrated FDG PET/CT and CT alone for preoperative staging. Radiology. 2005; 236:1011–9.
7. Wolchok JD, Hoos A, O'Day S, Weber JS, Hamid O, Lebbé C, Maio M, Binder M, Bohnsack O, Nichol G, Humphrey R, Hodi FS. Guidelines for the evaluation of immune therapy activity in solid tumors: immune-related response criteria. Clin Cancer Res. 2009; 15:7412–7420.
8. Nishino M, Jagannathan JP, Krajewski KM, O'Regan K, Hatabu H, Shapiro G, Ramaiya NH. Personalized Tumor Response Assessment in the Era of Molecular Medicine: Cancer-specific and Therapy-specific Response Criteria to Complement Pitfalls of RECIST. AJR Am J Roentgenol. 2012; 198:737–745.
9. Riely GJ, Kris MG, Zhao B, Akhurst T, Milton DT, Moore E, Tyson L, Pao W, Rizvi NA, Schwartz LH, Miller VA. Prospective assessment of discontinuation and reinitiation of erlotinib or gefitinib in patients with acquired resistance to erlotinib or gefitinib followed by the addition of everolimus. Clin Cancer Res. 2007; 13:5150–5155.
10. Jackman D, Pao W, Riely GJ, Engelman JA, Kris MG, Jänne PA, Lynch T, Johnson BE, Miller VA. Clinical definition of acquired resistance to epidermal growth factor receptor tyrosine kinase inhibitors in non-small-cell lung cancer. J Clin Oncol. 2010; 28:357–360
11. Mok TS. Living with imperfection. J Clin Oncol. 2010; 28: 191–19.
12. Sorensen BS, Wu L, Wei W, Tsai J, Weber B, Nexo E, Meldgaard P. Monitoring of epidermal growth factor receptor tyrosine kinase inhibitor-sensitizing and resistance mutations in the plasma DNA of patients with advanced non-small cell lung cancer during treatment with erlotinib. Cancer. 2014; 120: 3896–901.
13. Zheng D, Ye X, Zhang MZ, Sun Y, Wang JY, Ni J, Zhang HP, Zhang L, Luo J, Zhang J, Tang L, Su B, Chen G, et al. Plasma EGFR T790M ctDNA status is associated with clinical outcome in advanced NSCLC patients with acquired EGFR-TKI resistance. Sci Rep. 2016; 6:20913.
14. García-Sáenz JA, Ayllon P, Laig M, Acosta-Eyzaguirre D, García-Esquinas M, Montes M, Sanz J, Barquin M, Moreno F, Garcia-Barberan V, Díaz-Rubio E, Caldés T, Romero A. Tumor burden monitoring using cell-free tumor DNA could be limited by tumor heterogeneity in advanced breast cancer and should be evaluated together with radiographic imaging. BMC Cancer. 2017; 17:210.
15. Sacher AG, Paweletz C, Dahlberg SE, Alden RS, O'Connell A, Feeney N, Mach SL, Jänne PA, Oxnard GR. Prospective of rapid plasma genotyping for the detection of EGFR and KRAS mutations in advanced lung cancer. JAMA Oncol. 2016; 2:1014–22.
16. Yanagita M, Redig AJ, Paweletz CP, Dahlberg SE, O'Connell A, Feeney N, Taibi M, Boucher D, Oxnard GR, Johnson BE, Costa DB, Jackman DM, Jänne PA. A prospective evaluation of circulating tumor cells and cell-free DNA in EGFR mutant non-small cell lung cancer patients treated with erlotinib on a phase II trial. Clin Cancer Res. 2016; 22:6010–6020.
17. Park K, Tan EH, O'Byrne K, Zhang L, Boyer M, Mok T, Hirsh V, Yang JC, Lee KH, Lu S, Shi Y, Kim SW, Laskin J, et al. Afatinib versus gefitinib as first-line treatment of patients with EGFR mutation-positive non-small-cell lung cancer (LUX-Lung 7): a phase 2B, open-label, randomised controlled trial. Lancet Oncol. 2016; 17:577–89.
18. Hirsh V, Blais N, Burkes R, Verma S, Croitoru K. Management of diarrhea induced by epidermal growth factor receptor tyrosine kinase inhibitors. Curr Oncol. 2014; 21:329–36.
19. van Leeuwen RW, Peric R, Hussaarts KG, Kienhuis E, IJzerman NS, de Bruijn P, van der Leest C, Codrington H, Kloover JS, van der Holt B, Aerts JG, van Gelder T, Mathijssen RH. Influence of the acidic beverage cola on the absorption of erlotinib in patients with non-small-cell lung cancer. J Clin Oncol. 2016; 34:1309–1314.
20. Duong S, Leung M. Should the concomitant use of erlotinib and acid-reducing agents be avoided? The drug interaction between erlotinib and acid-reducing agents. J Oncol Pharm Pract. 2011; 17:448–52.
21. Regales L, Gong Y, Shen R, de Stanchina E, Vivanco I, Goel A, Koutcher JA, Spassova M, Ouerfelli O, Mellinghoff IK, Zakowski MF, Politi KA, Pao W. Dual targeting of EGFR can overcome a major drug resistance mutation in mouse models of EGFR mutant lung cancer. J Clin Invest. 2009; 119:3000–10.
22. Janjigian YY, Smit EF, Groen HJ, Horn L, Gettinger S, Camidge DR, Riely GJ, Wang B, Fu Y, Chand VK, Miller VA, Pao W. inhibition of EGFR with afatinib and cetuximab in kinase inhibitor-resistant EFGR-mutant lung cancer with and without T790M mutations. Cancer Discov 2014; 4:1036–45.
23. Schork NJ. Personalized medicine: time for one-person trials. Nature. 2015; 520:609–611.
24. Heffner JE, Klein JS. Recent advances in the diagnosis and management of malignant pleural effusions. Mayo Clin Proc. 2008; 83:235–50.
25. Pérez-Barrios C, Nieto-Alcolado I, Torrente M, Jiménez-Sánchez C, Calvo V, Gutierrez-Sanz L, Palka M, Donoso-Navarro E, Provencio M, Romero A. Comparison of methods for circulating cell-free DNA isolation using blood from cancer patients: impact on biomarker testing. Transl Lung Cancer Res. 2016; 5:665–672.

