INTRODUCTION
Gene mutation and methylation play essential roles in tumorigenesis and metastasis [1–3]. Tumor DNA contains specific somatic alterations that are crucial for the diagnosis and treatment of cancer. Due to the spatial and temporal intra-tumor heterogeneity, multi-sampling is needed to adequately characterize the somatic alterations [4–6]. Tissue biopsy, primarily the “gold standard” for diagnosis, however, is limited by the restricted access to sample and the challenges to recapitulate the tumor clonal diversity [7, 8]. Plasma circulating tumor DNA (ctDNA) is DNA fragments that contain the tumor-specific somatic alterations in the blood. Recent studies have demonstrated the application of plasma ctDNA detection in the tumor diagnosis and monitoring [9–11]. It is of non-invasive and can be collected repeatedly with minimal discomfort to the patient. It also reflects the total tumor burden and genetic heterogeneity.
Non-blood circulating tumor DNA are tumor DNA fragments presents in other body fluids, such as urine, saliva, sputum, stool, pleural fluid, and cerebrospinal fluid(CSF). Recent studies have demonstrated the presence of tumor DNA in these non-blood body fluids and their application to the diagnosis, screening, and monitoring of cancers [12] (Figure 1). The collection of these body fluids, such as urine, saliva, sputum, and stool, are relatively safe, non-invasive, economic and can be performed at home, without professional help [13, 14]. It also applies perfectly to patients with anemia, which is quite common in advanced stage cancer patients. Moreover, enrichment of tumor DNA of local cancers in non-blood body fluids may help to archive a higher sensitivity than in plasma. Non-blood body fluids are a viable alternative to blood samples as a source of DNA for tumor diagnosis and monitoring.
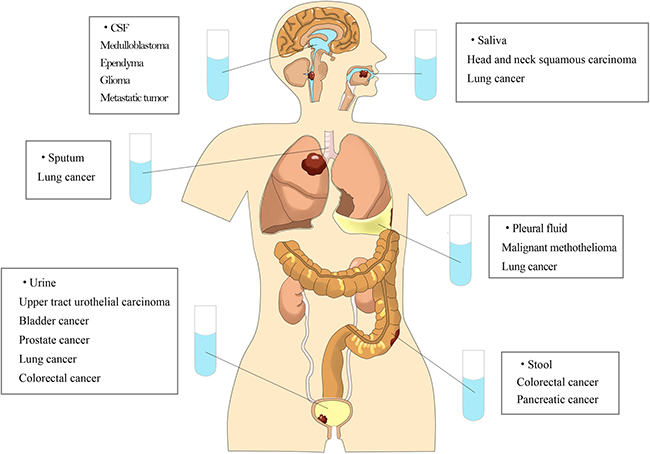
Figure 1: Tumor DNA can be detected in various kinds of non-blood body fluids.
Sources of non-blood tumor DNA
There are 2 types of non-blood tumor DNA: genomic DNA from local tumor cells that shed into the body fluid (cellular tumor DNA) and cell-free tumor DNA (cfDNA) from plasma cell-free DNA or from neighboring tumor cells due to necrosis or apoptosis (Figure 2).
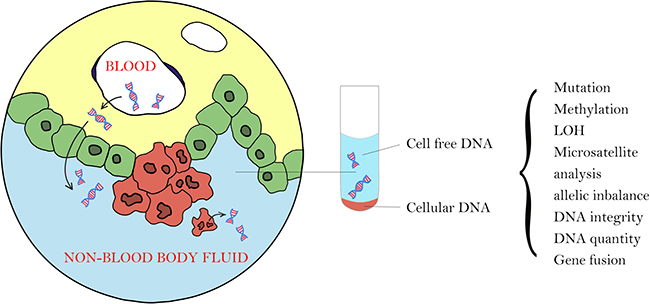
Figure 2: There are 2 types of tumor DNA in non-blood body fluid: cellular tumor DNA from local tumor cells that shed into the body fluid and cell-free tumor DNA from plasma cell-free DNA or from local tumor cells due to necrosis or apoptosis.
Urinary cfDNA mainly originates from plasma cfDNA that pass the kidney barrier [15]. Circulating tumor cfDNA is highly fragmented and primarily present in the blood as part of super molecule complexes, such as nucleosomes. During circulation, cfDNA was filtrated from the blood into the primary urine through the kidney barrier, which has been proved to be permeable to DNA molecules [16]. However, it is not known of the mechanism of DNA translocation from the bloodstream into the urine. Limited by the basal membrane and slit membranes between podocytes pedicles, only complexes smaller than 6.4 nm in diameter and with a molecular weight no greater than 70 kDa can pass through the kidney barrier and enter the nephron. It corresponds to DNA of about 100 bp in size, which is smaller than mononucleosome [15]. Moreover, the negatively charged cfDNA might face an additional barrier because of the negative charge of the glomerular basement membrane. It might be due to the non-globular shape or by the deformability of the DNA complexes. Another explanation is that cfDNA may be covered by liposomes, which make their penetration through the kidney barrier theoretically possible [17, 18]. Besides, renal permeability might increase for some physical and pathological conditions, such as pregnancy, cancer, and inflammation.
Ying et al. reported two size categories of urinary DNA: low molecular weight (MW) urine DNA and high MW urine DNA. Low MW class of urine DNA is between 150 to 250 bp and derived from the circulation, while the high MW urine DNA is greater than 1 kb and mostly from the cells shed into the urinary tract [13]. Similar findings were also reported in CSF. Size distribution of CSF cfDNA peaks at 160 and 340bp indicates an apoptotic source. CSF cellular DNA is of larger size and may origin from blood cells and tumor cells in CSF [19]. Saliva DNA can also be locally generated by cell necrosis, apoptosis, and exfoliation, or by active transport, passive diffusion or ultrafiltration from the plasma cfDNA [20]. The possible origins of pleural DNA also include ultrafiltration from the plasma (cfDNA) and local dying or apoptotic cells (cellular DNA) [21].
Non-blood tumor DNA and cancer
Urinary tumor DNA
Urinary cellular DNA origins from urological cancer cells shed into urine [22, 23]. In 1991, Sidransky et al. first identified p53 gene mutations in the urine sediment of bladder cancer patients [24]. In 2001, Carsten et al. demonstrated the presence of GSTP1 promoter hypermethylation in the urine sediments of prostate cancer patients [25]. Since then, gene mutations such as p53 [26], TERT [27, 28], FGFR3 [29, 30] mutations and gene hypermethylation changes such as GSTP1 [31], RARβ [31, 32] were reported in the urine of urothelial carcinomas patients (Table 1). In a 2-year follow-up of patients with superficial bladder transitional cell carcinoma after transurethral resection, Camille and colleagues assessed the diagnostic and prognostic performance of urinary cellular FGFR3 mutation analysis. Urinary cellular FGFR3 mutation has a sensitivity of 73% (95% CI, 0.58–0.89) and a specificity of 87% (95% CI, 0.82–0.93) in the diagnosis of cancer recurrence after transurethral resection, and a sensitivity of 70% (95% CI, 0.54–0.86) and specificity of 87% (95% CI, 0.76–0.98) in the prediction of recurrence within 2 years after surgery [38].
Table 1: Urinary tumor DNA detection in cancer
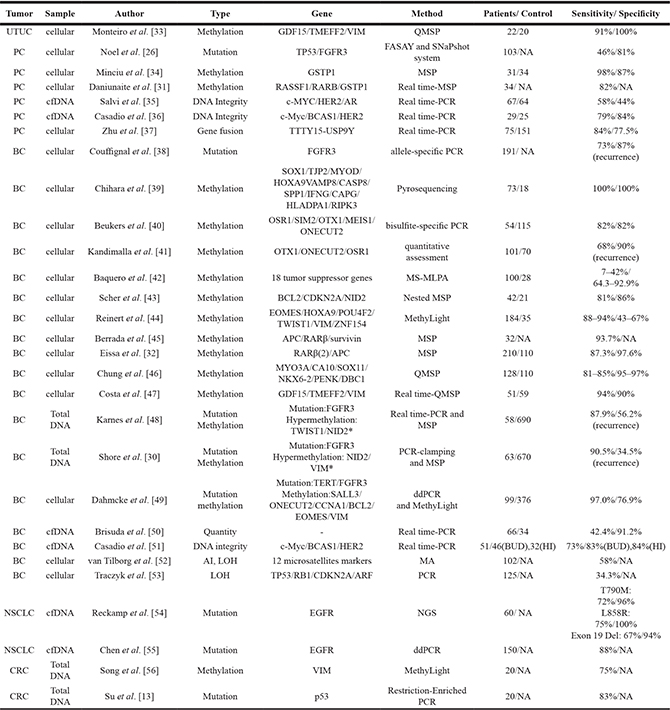
BC = bladder cancer; BUD = benign urogenital diseases; CRC = colorectal cancer; HI = healthy individuals; NSCLC = non-small cell lung cancer; PC = prostate cancer; UTUC = upper tract urothelial carcinoma; ddPCR = droplet digital polymerase chain reaction; MSP = methylation-specific PCR; NGS = next generation sequencing; QMSP = quantitative MSP; FASAY = functional analysis of separated allele in yeast; MS-MLPA = methylation-specific multiplex ligation-dependent probe amplification; MA = microsatellite analysis; AI = allelic imbalance; LOH = loss-of-heterozygosity; NA = not available.
*Panel includs a protein biomarker matrix metalloproteinase(MMP)-2.
Urinary cfDNA can be detected in non-urothelial carcinoma patients. The p53 mutation was found by Selena et al. in the urine of hepatocellular carcinoma patients and most of the p53 mutation was detected in the low MW urine DNA fraction [57]. Chen et al. detected EGFR mutations in urinary cfDNA in non-small cell lung cancer (NSCLC) patients with a concordance of 88% to their primary tumors [55].
Su et al. demonstrated that DNA methylation in urine is better in predicting recurrence than cytology and cystoscopy in bladder cancer patients after transurethral resection. By using a three-marker panel (SOX1, IRAK3, and L1-MET), they could predict tumor recurrence in 80% of patients, which is superior to cytology (35%) and cystoscopy (15%) [58]. Reckamp et al. detected the EGFR activating mutations and the T790M resistance mutation in urine and plasma of NSCLC patients by short footprint mutation enrichment next generation sequencing assays. They found that with a recommended specimen volume (90–100 mL), the sensitivity of urine and plasma are comparable [54]. Chen et al. conducted a serial monitoring trial to detect urinary cfDNA of EFGR mutation in NSCLC patients receiving EGFR-TKIs. They found a concordance in the quantity of urinary cfDNA and plasma cfDNA at baseline. During the treatment, a concordance in the decline of the quantity of cfDNA was also observed. Nevertheless, a more significant decrease in urinary cfDNA than plasma cfDNA was found during the early phase of monitoring as a result of treatment, which indicated that urinary cfDNA might potentially be of higher sensitivity [55].
Saliva tumor DNA
Saliva provides good-quality genomic DNA, which is comparable to blood as a template for genotyping [59, 60]. Salivary DNA has been used in germline mutations detection for various cancer screening, such as breast cancer [61] and braintumor [62].
In 2000, Liao et al. detected p53 gene mutation in the saliva of oral squamous cell carcinomas patients [63]. In 2001, El-Naggar et al. performed microsatellite analysis at chromosomal regions frequently altered in head and neck squamous cell carcinoma (HNSCC) on matched saliva and tumor samples. Their results showed a statistically significant correlation in loss of heterozygosity (LOH) between saliva and tumor with some sets of markers [64]. Salivary DNA promoter hypermethylation analysis has also been found to be an efficient tool for diagnosis of HNSCC [65–67] (Table 2). In 2014, Wei et al. developed a novel core technology, called electric field-induced release and measurement (EFIRM) to detect EGFR mutations directly in body fluids with a multiplexable electrochemical sensor. They demonstrated that EFIRM could detect EGFR mutations in the saliva of NSCLC patients, with an area under the curve (AUC) of 0.94 in the detection of exon 19 deletion and an AUC of 0.90 in the detection of L858R mutation [75]. Using EFIRM, Pu et al. detected the EFGR exon 19 deletion and p.L858 mutations in saliva and plasma samples of 17 lung adenocarcinoma patients. They found a perfect concordance between saliva and tumor samples, with an AUC of 1.0 [14]. It is therefore suggested that there is a link between the peripheral circulatory system and the salivary glands that translocate cfDNA from the bloodstream into saliva [14].
Table 2: Saliva tumor DNA detection in HNSCC
Author |
Type |
Gene |
Method |
Patients/Controls |
Sensitivity/Specificity |
|---|---|---|---|---|---|
Sun et al. [68] |
Methylation |
TIMP3 |
QMSP |
197/NA |
NA/NA |
Gaykalova et al. [69] |
Methylation |
ZNF14/ZNF160/ZND420 |
QMSP |
59/ NA |
57.6%/100% |
Ovchinnikov et al. [70] |
Methylation |
MED15/PCQAP |
MSP |
46/49(5′-CpGs) |
70%/63% (5′-CpGs) |
Rettori et al. [71] |
Methylation |
CCNA1/DAPK/DCC/MGMT/TIMP3 |
QMSP |
146/60 |
55%/76% |
Demokan et al. [72] |
Methylation |
KIF1A/EDNRB |
QMSP |
71/61 |
77.4%/93.1% |
Righini et al. [67] |
Methylation |
TIMP3/ECAD/p16INK4a/MGMT/DAPK/RASSF1 |
QMSP |
60/ NA |
78.3%/ NA |
Ovchinnikov et al. [73] |
Methylation |
RASSF1A/DAPK1/ p16INK4a |
MSP |
143/31 |
80%/87% |
Schussel et al. [74] |
Methylation |
EDNRB/DCC |
QMSP |
48/113 |
46%/72% |
MSP = Methylation-Specific PCR; QMSP = Quantitative MSP; NA = not available.
Sputum tumor DNA
Sputum contains cells from the lungs and lower respiratory tract and provides sufficient tumor DNA for detection. Numerous studies have shown that sputum tumor DNA could be a promising tool for early detection of lung cancer (Table 3). In 1994, Mao et al. detected the K-ras mutation and p53 mutation in sputum samples of lung cancer patients. Using a PCR-based assay, they detected mutant DNA in the sputum of 8 of 10 patients with oncogene mutations in their primary tumor prior to clinical diagnosis [88]. DNA hypermethylation in sputum also helps the diagnosis of lung cancer [89]. Wang et al. carried out a meta-analysis to comprehensively review the evidence for using sputum aberrant methylation DNA to detect NSCLC. They found that the combined sensitivity was 62% (95% CI: 0.59–0.65), and specificity was 73% (95% CI: 0.70–0.75) [90]. Miglio et al. demonstrated that MGMT promoter methylation was present in small cell lung cancer and cytological samples were perfectly adequate for methylation analysis [91]. Flow cytometric DNA analysis of sputum cells also showed good sensitivity in the diagnosis of lung cancer. Compared with cytologic morphology of sputum cells, sensitivity of sputum DNA heteroploidy analysis was significantly higher (82.8% vs 27.6%, P < 0.005) [92].
Table 3: Sputum tumor DNA detection in lung cancer
Author |
Type |
Gene |
Method |
Patients/Control |
Sensitivity/Specificity |
|---|---|---|---|---|---|
Konno et al. [76] |
Methylation |
p16INK4a/APC/RARβ |
MSP |
78/95 |
78%/79% |
Wang et al. [77] |
Methylation |
p16INK4a |
PCR |
34/21 |
32%/100% |
Belinsky et al. [78] |
Methylation |
p16INK4a /DAPK/H-cadherin/PAX5α/PAX5β/MGMT/RASSF1A |
MSP |
53/118 |
85%/35% |
Olaussen et al. [79] |
Methylation |
HOX/p16INK4a/MAGE/MAGE |
MSP |
22/56 |
96%/79% |
Cirincione et al. [80] |
Methylation |
RARβ2/p16INK4a/RASSF1A |
MSP |
18/112 |
50%/38% |
Georgiou et al. [81] |
Methylation |
p16INK4a |
MSP |
80/40 |
69%/76% |
Shivapurkar et al. [82] |
Methylation |
3-OST-2/RASSF1A/ p16INK4a /APC |
Quantitative Real time-PCR |
13/23 |
62%/100% |
van der Drift et al. [83] |
Methylation |
RASSF1A |
RT-globin PCR |
28/68 |
46%/99% |
Hwang et al. [84] |
Methylation |
HOXA9 |
MSP |
76/109 |
71%/55% |
Zhang et al. [85] |
Methylation |
p16INK4a |
MSP |
44/20 |
61% |
Destro et al. [86] |
Mutation methylation |
Mutation: K-ras |
PCR-RFLP and MSP |
50/100 |
75%/96% |
Wang et al. [87] |
Methylation, LOH, MSI |
methylation: p16INK4a/RARβ |
MSP LOH analyses |
79/22 |
81%/72% |
MSP = Methylation-Specific PCR; PCR-RFLP = PCR Restriction Fragment Length Polymorphism; MSI = microsatellite instability; LOH = loss of heterozygosity.
DNA analysis from sputum was consistent with that from plasma in patients with lung cancer. By using a fluorescent PCR-based approach, Castagnaro et al. were able to assess the consistency of DNA microsatellite analysis of induced sputum. They demonstrated a significant trend in the percentage of the genetic alterations, found both in induced sputum and in blood samples, from healthy subjects to heavy smokers and lung cancer patients [93]. LOH and microsatellite instability (MSI) in at least one locus was observed in 55% of patients, in 18% of smokers, and in 4.5% of healthy subjects. These results showed that sputum DNA provided data that were consistent with those from plasma [93].
Stool tumor DNA
Stool is the fecal discharge from the bowels. It is biologically rational to use stool as a non-invasive sample for colorectal cancer (CRC). Stool DNA detection has a marked improvement of the sensitivity when compared to fecal blood tests. Compared to colonoscopy, which is currently the dominant screening test, stool DNA detection is patient-friendly and free from unpleasant cathartic bowel preparations and diet or medication restrictions. Furthermore, colonoscopy is operator dependent and has been shown to be not as effective detecting proximal lesions [94]. Stool DNA testing detects proximal and distal colorectal neoplasms equally well.
According to a meta-analysis involving 7524 patients, the pool sensitivities of stool DNA testing for CRC were 48% for single-gene and 77.8% for multiple-gene assays, and the specificities were 97% and 92.7%, respectively [95]. In a cross-sectional study including 9989 participants, Thomas et al. evaluated the effectiveness of multitarget stool DNA (mt-sDNA) test in colorectal-cancer screening. The mt-sDNA test included quantitative molecular assays for K-ras mutations, aberrant NDRG4, and BMP3 methylation, and β-actin, plus a hemoglobin immunoassay. The sensitivity for detecting colorectal cancer was significantly higher with mt-sDNA test than immunochemical test (FIT) (92.3% vs. 73.8%, p = 0.002) [96].
Besides CRC, stool tumor DNA can also be detected in other digestive system neoplasms. In 1994, Caldas demonstrated the presence of K-ras mutation in the stool of pancreatic cancer patients [97]. In a more recent work, Kisiel et al found a sensitivity of 67% and a specificity of 90% in detecting pancreatic cancer with a combination of stool mutated K-ras and methylated BMP3 detection [98].
Plasma based DNA tests, especially marking the aberrant methylation of SEPT9 gene, have been evaluated as a potential screening tool for CRC and advanced adenomas [99]. Ahlquist et al. conducted a case-control study to compare the sensitivities of multimarker stool DNA test and plasma methylated SEPT9 test in identifying patients with large adenomas or CRC. Their results demonstrated that mt-sDNA test had a significantly greater level of sensitivity than the plasma methylated SEPT9 test for detection of both CRC and large adenomas (87% vs. 60% CRC sensitivity and 82% vs. 14% adenoma sensitivity) [100]. The high sensitivity of stool DNA test may be related to disproportionately copious exfoliation of cancer cells, remarkably large functional surface area of neoplasms, enhanced survival of shed dysplastic cells, and relative stability and informativeness of tumor-associated DNA changes [100].
Pleural tumor DNA
Pleural effusions arise from a variety of systemic, inflammatory, infectious and malignant conditions. Malignant pleural effusion (MPE) is a devastating complication caused by a series of cancers, including lung cancer and mesotheliomas. Positive cytologic or tissue confirmation of malignant cells is necessary to establish a diagnosis. However, the sensitivity of pleural fluid cytological analysis is relatively low. Molecular biology techniques, such as analyses of DNA mutation and methylation status, have provided novel diagnostic tools for MPEs.
In 2006, Kimura et al. assessed the pleural effusion of 43 known NSCLC patients and found mutated EGFR in 11 of 43 cases [101]. Using peptide nucleic acid (PNA)-mediated real-time PCR clamping, Yeo et al. detected the EGFR mutation in pleural effusion of NSCLC patients with a sensitivity of 89% and a specificity of 100% [102]. Benlloch et al. examined the promoter methylation status of 4 genes (DAPK, RASSF1A, RARβ, p16INK4a) in patients with pleural effusion. Abnormal DNA methylation was detected in 58.5% of malignant pleural effusions, while in 0% of patients with benign pleural effusions [103]. Fujii et al. detected hypermethylated RASSF1A, p16INK4a, RARβ in both malignant pleural mesothelioma (MPM) and lung cancer. They found that the methylation ratios for the three genes were significantly higher in lung cancer than in MPM, which suggested that pleural fluid DNA could be a possible marker for differentiating MPM from lung cancer [104].
Using high resolution melting (HRM) analysis, Lin et al. assessed the pleural cfDNA and pleural cellular DNA of 13 known NSCLC cancer patients with EGFR mutation in matched biopsy tumor tissues, and found mutated EGFR in 12 and 9 of 13 cases, respectively [105]. Similar results were reported by Liu et al. using amplification refractory mutation system. Higher sensitivity of pleural cfDNA might be due to the tumor cells damaged under high speed of centrifugation, and DNA fragments were released from the nucleus, making up the dominant components of the supernatant [106].
CSF tumor DNA
Circulating tumor DNA has been detected in a variety of cancers. However, it is rarely found in patients with isolated brain tumors, presumably owing to the blood-brain barrier [107]. CSF is a clear, colorless body fluid that bathes the brain and spinal cord. It circulates nutrients and chemicals filtered from the blood and removes waste products from the brain. Examining the fluid can be useful in diagnosing many diseases of the nervous system, including brain tumors.
CSF tumor DNA provides a minimally invasive method to assess the genomic alterations of the tumor and monitor the therapy effect that helps both diagnosis and treatment. Using next generation sequencing approach, Pentsova et al. sequenced 341 cancer-associated genes in CSF of 53 patients with suspected or known CNS cancers. They detected high-confidence somatic alterations in 63% (20 of 32) of patients with CNS metastases of solid tumors, 50% (6 of 12) of patients with primary brain tumors, and 0% (0 of 9) of patients without CNS involvement by cancer [108]. Wang et al. found that all medulloblastomas, ependymomas, and high-grade gliomas that abutted a CSF space or cortical surface were detectable (100% of 21 cases; 95% CI = 88–100%), whereas no CSF tumor DNA was detected in patients whose tumors were not directly adjacent to a CSF reservoir [109].
In De Mattos-Arruda et al.’s seminal work, CSF ctDNA was identified in brain primary and metastatic tumors and represented private mutations from CNS lesions. Furthermore, sensitivity of ctDNA for somatic mutations of the CNS was higher than plasma DNA in patients with a CNS-restricted disease (58% vs. 0%, p = 0.0006). While in patients with the abundant visceral disease, the sensitivity of CNS DNA and plasma DNA was comparable (60.5% vs. 55.5%). The investigators further monitored the change of mutant allelic frequency (MAF) of CSF DNA and plasma DNA in a serial study. MAFs of CSF DNA decreased with surgical resection and/or responses to systemic therapy and increased with tumor progression [7]. Similar findings were also presented by Pan et al. The median concentration of cfDNA in CSF is lower than that in plasma (2.1 ng/mL vs. 7.7 ng/mL). However, the ability to detect mutations in CSF is stronger than in plasma in brain tumor patients with low systemic metastatic burden [19]. EGFR mutation in CSF was also detected in a case with suspected leptomeningeal metastasis from EGFR mutant lung adenocarcinoma, which indicates the characterization of brain tumor genomic aberrations through CSF DNA analysis is possible. Very few cells are present in CSF under routine conditions (0–5 cells/L). The scarcity of cells in CSF may reduce the background noise from normal DNA when detecting mutations [19].
CONCLUSIONS
Non-blood circulating tumor DNA has an enormous potential for large-scale screening of local neoplasms because of its non-invasive nature, close proximity to the tumors, easiness and it is an economically viable option. It permits longitudinal assessments and allows sequential monitoring of response and progression [110]. The direct contact of cancerous cells and body fluid may facilitate the detection of tumor DNA, while vascular invasion likely happens at a later stage in tumorigenesis, which may explain the low sensitivity of plasma-based tests [111]. Furthermore, normal DNA always dilutes the ctDNA, which may be aggravated by inflammation and injury when very high amounts of normal DNA are released into the circulation [112]. Altogether, our review indicate that non-blood circulating tumor DNA presents an option where the disease can be tracked in a simple and less-invasive manner, allowing for serial sampling informing of the tumor heterogeneity and response to treatment.
ACKNOWLEDGMENTS AND FUNDING
The authors greatly appreciate Dr. Shiyu Chen, for her kindly help with the figures of the manuscript.
CONFLICTS OF INTEREST
There are not any financial/commercial conflicts of interests involving this study.
REFERENCES
1. Chen C, Peng H, Huang X, Zhao M, Li Z, Yin N, Wang X, Yu F, Yin B, Yuan Y, Lu Q. Genome-wide profiling of DNA methylation and gene expression in esophageal squamous cell carcinoma. Oncotarget. 2016; 7:4507–4521. https://doi.org/10.18632/oncotarget.6607.
2. Liu ZH, Liu JJ, Li SS, Yang XM. Association of RUNX3 Methylation with Clinical Outcome and Cell Migration/Invasion in Laryngeal Squamous Cell Carcinoma. Cancer Invest. 2016; 34:105–113.
3. Zhang S, Feng XL, Shi L, Gong CJ, He ZJ, Wu HJ, Ling TY. Genome-wide analysis of DNA methylation in tongue squamous cell carcinoma. Oncol Rep. 2013; 29:1819–1826.
4. Gerlinger M, Rowan AJ, Horswell S, Larkin J, Endesfelder D, Gronroos E, Martinez P, Matthews N, Stewart A, Tarpey P, Varela I, Phillimore B, Begum S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012; 366:883–892.
5. Sottoriva A, Spiteri I, Piccirillo SG, Touloumis A, Collins VP, Marioni JC, Curtis C, Watts C, Tavare S. Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc Natl Acad Sci USA. 2013; 110:4009–4014.
6. He C, Zhao X, Jiang H, Zhong Z, Xu R. Demethylation of miR-10b plays a suppressive role in ccRCC cells. Int J Clin Exp Pathol. 2015; 8:10595–10604.
7. De Mattos-Arruda L, Mayor R, Ng CK, Weigelt B, Martinez-Ricarte F, Torrejon D, Oliveira M, Arias A, Raventos C, Tang J, Guerini-Rocco E, Martinez-Saez E, Lois S, et al. Cerebrospinal fluid-derived circulating tumour DNA better represents the genomic alterations of brain tumours than plasma. Nat Commun. 2015; 6:8839.
8. Diao CY, Guo HB, Ouyang YR, Zhang HC, Liu LH, Bu J, Wang ZH, Xiao T. Screening for metastatic osteosarcoma biomarkers with a DNA microarray. Asian Pac J Cancer Prev. 2014; 15:1817–1822.
9. Murtaza M, Dawson SJ, Tsui DW, Gale D, Forshew T, Piskorz AM, Parkinson C, Chin SF, Kingsbury Z, Wong AS, Marass F, Humphray S, Hadfield J, et al. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature. 2013; 497:108–112.
10. Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, Dunning MJ, Gale D, Forshew T, Mahler-Araujo B, Rajan S, Humphray S, Becq J, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013; 368:1199–1209.
11. De Mattos-Arruda L, Weigelt B, Cortes J, Won HH, Ng CK, Nuciforo P, Bidard FC, Aura C, Saura C, Peg V, Piscuoglio S, Oliveira M, Smolders Y, et al. Capturing intra-tumor genetic heterogeneity by de novo mutation profiling of circulating cell-free tumor DNA: a proof-of-principle. Ann Oncol. 2014; 25:1729–1735.
12. Patel KM, Tsui DW. The translational potential of circulating tumour DNA in oncology. Clin Biochem. 2015; 48:957–961.
13. Su YH, Wang M, Brenner DE, Ng A, Melkonyan H, Umansky S, Syngal S, Block TM. Human urine contains small, 150 to 250 nucleotide-sized, soluble DNA derived from the circulation and may be useful in the detection of colorectal cancer. J Mol Diagn. 2004; 6:101–107.
14. Pu D, Liang H, Wei F, Akin D, Feng Z, Yan Q, Li Y, Zhen Y, Xu L, Dong G, Wan H, Dong J, Qiu X, et al. Evaluation of a novel saliva-based epidermal growth factor receptor mutation detection for lung cancer: A pilot study. Thorac Cancer. 2016; 7:428–436.
15. Bryzgunova OE, Laktionov PP. Extracellular Nucleic Acids in Urine: Sources, Structure, Diagnostic Potential. Acta Naturae. 2015; 7:48–54.
16. Botezatu I, Serdyuk O, Potapova G, Shelepov V, Alechina R, Molyaka Y, Ananev V, Bazin I, Garin A, Narimanov M, Knysh V, Melkonyan H, Umansky S, et al. Genetic analysis of DNA excreted in urine: a new approach for detecting specific genomic DNA sequences from cells dying in an organism. Clin Chem. 2000; 46:1078–1084.
17. Lichtenstein AV, Melkonyan HS, Tomei LD, Umansky SR. Circulating nucleic acids and apoptosis. Ann N Y Acad Sci. 2001; 945:239–249.
18. Foglieni C, Bragonzi A, Cortese M, Cantu L, Boletta A, Chiossone I, Soria MR, Monaco L. Glomerular filtration is required for transfection of proximal tubular cells in the rat kidney following injection of DNA complexes into the renal artery. Gene Ther. 2000; 7:279–285.
19. Pan W, Gu W, Nagpal S, Gephart MH, Quake SR. Brain tumor mutations detected in cerebral spinal fluid. Clin Chem. 2015; 61:514–522.
20. Kaufman E, Lamster IB. The diagnostic applications of saliva—a review. Crit Rev Oral Biol Med. 2002; 13:197–212.
21. Chan MH, Chow KM, Chan AT, Leung CB, Chan LY, Chow KC, Lam CW, Lo YM. Quantitative analysis of pleural fluid cell-free DNA as a tool for the classification of pleural effusions. Clin Chem. 2003; 49:740–745.
22. Utting M, Werner W, Dahse R, Schubert J, Junker K. Microsatellite analysis of free tumor DNA in urine, serum, and plasma of patients: a minimally invasive method for the detection of bladder cancer. Clin Cancer Res. 2002; 8:35–40.
23. Eisenberger CF, Schoenberg M, Enger C, Hortopan S, Shah S, Chow NH, Marshall FF, Sidransky D. Diagnosis of renal cancer by molecular urinalysis. J Natl Cancer Inst. 1999; 91:2028–2032.
24. Sidransky D, Von Eschenbach A, Tsai YC, Jones P, Summerhayes I, Marshall F, Paul M, Green P, Hamilton SR, Frost P, Vogelstein B. Identification of p53 gene mutations in bladder cancers and urine samples. Science. 1991; 252:706–709.
25. Goessl C, Muller M, Heicappell R, Krause H, Miller K. DNA-based detection of prostate cancer in blood, urine, and ejaculates. Ann N Y Acad Sci. 2001; 945:51–58.
26. Noel N, Couteau J, Maillet G, Gobet F, D'Aloisio F, Minier C, Pfister C. TP53 and FGFR3 Gene Mutation Assessment in Urine: Pilot Study for Bladder Cancer Diagnosis. Anticancer Res. 2015; 35:4915–4921.
27. Kinde I, Munari E, Faraj SF, Hruban RH, Schoenberg M, Bivalacqua T, Allaf M, Springer S, Wang Y, Diaz LA Jr, Kinzler KW, Vogelstein B, Papadopoulos N, et al. TERT promoter mutations occur early in urothelial neoplasia and are biomarkers of early disease and disease recurrence in urine. Cancer Res. 2013; 73:7162–7167.
28. Wang K, Liu T, Liu L, Liu J, Liu C, Wang C, Ge N, Ren H, Yan K, Hu S, Bjorkholm M, Fan Y, Xu D. TERT promoter mutations in renal cell carcinomas and upper tract urothelial carcinomas. Oncotarget. 2014; 5:1829–1836. https://doi.org/10.18632/oncotarget.1829.
29. Lamy A, Gobet F, Laurent M, Blanchard F, Varin C, Moulin C, Andreou A, Frebourg T, Pfister C. Molecular profiling of bladder tumors based on the detection of FGFR3 and TP53 mutations. J Urol. 2006; 176:2686–2689.
30. Shore ND, Fernandez CA, Shuber AP. Noninvasive multianalyte diagnostic assay for monitoring bladder cancer recurrence. Res Rep Urol. 2012; 4:49–56.
31. Daniunaite K, Berezniakovas A, Jankevicius F, Laurinavicius A, Lazutka JR, Jarmalaite S. Frequent methylation of RASSF1 and RARB in urine sediments from patients with early stage prostate cancer. Medicina (Kaunas). 2011; 47:147–153.
32. Eissa S, Swellam M, El-Khouly IM, Kassim SK, Shehata H, Mansour A, Esmat M, Nossier AI, Hamdy MA, Awad NM, El-Ahmady O. Aberrant methylation of RARbeta2 and APC genes in voided urine as molecular markers for early detection of bilharzial and nonbilharzial bladder cancer. Cancer Epidemiol Biomarkers Prev. 2011; 20:1657–1664.
33. Monteiro-Reis S, Leca L, Almeida M, Antunes L, Monteiro P, Dias PC, Morais A, Oliveira J, Henrique R, Jeronimo C. Accurate detection of upper tract urothelial carcinoma in tissue and urine by means of quantitative GDF15, TMEFF2 and VIM promoter methylation. Eur J Cancer. 2014; 50:226–233.
34. Minciu R, Dumache R, Gheorghe P, Daminescu L, Rogobete AF, Ionescu D. Molecular Diagnostic of Prostate Cancer From Body Fluids Using Methylation-Specific PCR (MS-PCR) Method. Clin Lab. 2016; 62:1183–1186.
35. Salvi S, Gurioli G, Martignano F, Foca F, Gunelli R, Cicchetti G, De Giorgi U, Zoli W, Calistri D, Casadio V. Urine Cell-Free DNA Integrity Analysis for Early Detection of Prostate Cancer Patients. Dis Markers. 2015; 2015:574120.
36. Casadio V, Calistri D, Salvi S, Gunelli R, Carretta E, Amadori D, Silvestrini R, Zoli W. Urine cell-free DNA integrity as a marker for early prostate cancer diagnosis: a pilot study. Biomed Res Int. 2013; 2013:270457.
37. Zhu Y, Ren S, Jing T, Cai X, Liu Y, Wang F, Zhang W, Shi X, Chen R, Shen J, Lu J, Xu C, Wang H, et al. Clinical utility of a novel urine-based gene fusion TTTY15-USP9Y in predicting prostate biopsy outcome. Urol Oncol. 2015; 33:384.e9–320.
38. Couffignal C, Desgrandchamps F, Mongiat-Artus P, Ravery V, Ouzaid I, Roupret M, Phe V, Ciofu C, Tubach F, Mentre F, Cussenot O, Grandchamp B. The Diagnostic and Prognostic Performance of Urinary FGFR3 Mutation Analysis in Bladder Cancer Surveillance: A Prospective Multicenter Study. Urology. 2015; 86:1185–1190.
39. Chihara Y, Kanai Y, Fujimoto H, Sugano K, Kawashima K, Liang G, Jones PA, Fujimoto K, Kuniyasu H, Hirao Y. Diagnostic markers of urothelial cancer based on DNA methylation analysis. BMC Cancer. 2013; 13:275.
40. Beukers W, Kandimalla R, van Houwelingen D, Kovacic H, Chin JF, Lingsma HF, Dyrskjot L, Zwarthoff EC. The use of molecular analyses in voided urine for the assessment of patients with hematuria. PLoS One. 2013; 8:e77657.
41. Kandimalla R, Masius R, Beukers W, Bangma CH, Orntoft TF, Dyrskjot L, van Leeuwen N, Lingsma H, van Tilborg AA, Zwarthoff EC. A 3-plex methylation assay combined with the FGFR3 mutation assay sensitively detects recurrent bladder cancer in voided urine. Clin Cancer Res. 2013; 19:4760–4769.
42. Garcia-Baquero R, Puerta P, Beltran M, Alvarez M, Sacristan R, Alvarez-Ossorio JL, Sanchez-Carbayo M. Methylation of a novel panel of tumor suppressor genes in urine moves forward noninvasive diagnosis and prognosis of bladder cancer: a 2-center prospective study. J Urol. 2013; 190:723–730.
43. Scher MB, Elbaum MB, Mogilevkin Y, Hilbert DW, Mydlo JH, Sidi AA, Adelson ME, Mordechai E, Trama JP. Detecting DNA methylation of the BCL2, CDKN2A and NID2 genes in urine using a nested methylation specific polymerase chain reaction assay to predict bladder cancer. J Urol. 2012; 188:2101–2107.
44. Reinert T, Borre M, Christiansen A, Hermann GG, Orntoft TF, Dyrskjot L. Diagnosis of bladder cancer recurrence based on urinary levels of EOMES, HOXA9, POU4F2, TWIST1, VIM, and ZNF154 hypermethylation. PLoS One. 2012; 7:e46297.
45. Berrada N, Amzazi S, Ameziane El Hassani R, Benbacer L, El Mzibri M, Khyatti M, Chafiki J, Abbar M, Al Bouzidi A, Ameur A, Attaleb M. Epigenetic alterations of adenomatous polyposis coli (APC), retinoic acid receptor beta (RARbeta) and survivin genes in tumor tissues and voided urine of bladder cancer patients. Cell Mol Biol (Noisy-le-grand). 2012; 58:OL1744–1751.
46. Chung W, Bondaruk J, Jelinek J, Lotan Y, Liang S, Czerniak B, Issa JP. Detection of bladder cancer using novel DNA methylation biomarkers in urine sediments. Cancer Epidemiol Biomarkers Prev. 2011; 20:1483–1491.
47. Costa VL, Henrique R, Danielsen SA, Duarte-Pereira S, Eknaes M, Skotheim RI, Rodrigues A, Magalhaes JS, Oliveira J, Lothe RA, Teixeira MR, Jeronimo C, Lind GE. Three epigenetic biomarkers, GDF15, TMEFF2, and VIM, accurately predict bladder cancer from DNA-based analyses of urine samples. Clin Cancer Res. 2010; 16:5842–5851.
48. Karnes RJ, Fernandez CA, Shuber AP. A noninvasive multianalyte urine-based diagnostic assay for urothelial cancer of the bladder in the evaluation of hematuria. Mayo Clin Proc. 2012; 87:835–842.
49. Dahmcke CM, Steven KE, Larsen LK, Poulsen AL, Abdul-Al A, Dahl C, Guldberg P. A Prospective Blinded Evaluation of Urine-DNA Testing for Detection of Urothelial Bladder Carcinoma in Patients with Gross Hematuria. Eur Urol. 2016.
50. Brisuda A, Pazourkova E, Soukup V, Horinek A, Hrbacek J, Capoun O, Svobodova I, Pospisilova S, Korabecna M, Mares J, Hanus T, Babjuk M. Urinary Cell-Free DNA Quantification as Non-Invasive Biomarker in Patients with Bladder Cancer. Urol Int. 2016; 96:25–31.
51. Casadio V, Calistri D, Tebaldi M, Bravaccini S, Gunelli R, Martorana G, Bertaccini A, Serra L, Scarpi E, Amadori D, Silvestrini R, Zoli W. Urine cell-free DNA integrity as a marker for early bladder cancer diagnosis: preliminary data. Urol Oncol. 2013; 31:1744–1750.
52. van Tilborg AA, Kompier LC, Lurkin I, Poort R, El Bouazzaoui S, van der Keur K, Zuiverloon T, Dyrskjot L, Orntoft TF, Roobol MJ, Zwarthoff EC. Selection of microsatellite markers for bladder cancer diagnosis without the need for corresponding blood. PLoS One. 2012; 7:e43345.
53. Traczyk M, Borkowska E, Jedrzejczyk A, Pietrusinski M, Rozniecki M, Marks P, Kaluzewski B. Detection of loss of heterozygosity in patients with urinary bladder carcinoma: neoplastic tissue vs. urine sediment cells. Cent European J Urol. 2011; 64:163–167.
54. Reckamp KL, Melnikova VO, Karlovich C, Sequist LV, Camidge DR, Wakelee H, Perol M, Oxnard GR, Kosco K, Croucher P, Samuelsz E, Vibat CR, Guerrero S, et al. A Highly Sensitive and Quantitative Test Platform for Detection of NSCLC EGFR Mutations in Urine and Plasma. J Thorac Oncol. 2016; 11:1690–1700.
55. Chen S, Zhao J, Cui L, Liu Y. Urinary circulating DNA detection for dynamic tracking of EGFR mutations for NSCLC patients treated with EGFR-TKIs. Clin Transl Oncol. 2016; 19:332–340.
56. Song BP, Jain S, Lin SY, Chen Q, Block TM, Song W, Brenner DE, Su YH. Detection of hypermethylated vimentin in urine of patients with colorectal cancer. J Mol Diagn. 2012; 14:112–119.
57. Lin SY, Dhillon V, Jain S, Chang TT, Hu CT, Lin YJ, Chen SH, Chang KC, Song W, Yu L, Block TM, Su YH. A locked nucleic acid clamp-mediated PCR assay for detection of a p53 codon 249 hotspot mutation in urine. J Mol Diagn. 2011; 13:474–484.
58. Su SF, de Castro Abreu AL, Chihara Y, Tsai Y, Andreu-Vieyra C, Daneshmand S, Skinner EC, Jones PA, Siegmund KD, Liang G. A panel of three markers hyper- and hypomethylated in urine sediments accurately predicts bladder cancer recurrence. Clin Cancer Res. 2014; 20:1978–1989.
59. Bahlo M, Stankovich J, Danoy P, Hickey PF, Taylor BV, Browning SR, Australian and New Zealand Multiple Sclerosis Genetics Consortium (ANZgene), Brown MA, Rubio JP. Saliva-derived DNA performs well in large-scale, high-density single-nucleotide polymorphism microarray studies. Cancer Epidemiol Biomarkers Prev. 2010; 19:794–798.
60. Hu Y, Ehli EA, Nelson K, Bohlen K, Lynch C, Huizenga P, Kittlelsrud J, Soundy TJ, Davies GE. Genotyping performance between saliva and blood-derived genomic DNAs on the DMET array: a comparison. PLoS One. 2012; 7:e33968.
61. Dean M, Boland J, Yeager M, Im KM, Garland L, Rodriguez-Herrera M, Perez M, Mitchell J, Roberson D, Jones K, Lee HJ, Eggebeen R, Sawitzke J, et al. Addressing health disparities in Hispanic breast cancer: accurate and inexpensive sequencing of BRCA1 and BRCA2. Gigascience. 2015; 4:50.
62. Adel Fahmideh M, Lavebratt C, Schuz J, Roosli M, Tynes T, Grotzer MA, Johansen C, Kuehni CE, Lannering B, Prochazka M, Schmidt LS, Feychting M. CCDC26, CDKN2BAS, RTEL1 and TERT Polymorphisms in pediatric brain tumor susceptibility. Carcinogenesis. 2015; 36:876–882.
63. Liao PH, Chang YC, Huang MF, Tai KW, Chou MY. Mutation of p53 gene codon 63 in saliva as a molecular marker for oral squamous cell carcinomas. Oral Oncol. 2000; 36:272–276.
64. El-Naggar AK, Mao L, Staerkel G, Coombes MM, Tucker SL, Luna MA, Clayman GL, Lippman S, Goepfert H. Genetic heterogeneity in saliva from patients with oral squamous carcinomas: implications in molecular diagnosis and screening. J Mol Diagn. 2001; 3:164–170.
65. Viet CT, Schmidt BL. Methylation array analysis of preoperative and postoperative saliva DNA in oral cancer patients. Cancer Epidemiol Biomarkers Prev. 2008; 17:3603–3611.
66. Carvalho AL, Jeronimo C, Kim MM, Henrique R, Zhang Z, Hoque MO, Chang S, Brait M, Nayak CS, Jiang WW, Claybourne Q, Tokumaru Y, Lee J, et al. Evaluation of promoter hypermethylation detection in body fluids as a screening/diagnosis tool for head and neck squamous cell carcinoma. Clin Cancer Res. 2008; 14:97–107.
67. Righini CA, de Fraipont F, Timsit JF, Faure C, Brambilla E, Reyt E, Favrot MC. Tumor-specific methylation in saliva: a promising biomarker for early detection of head and neck cancer recurrence. Clin Cancer Res. 2007; 13:1179–1185.
68. Sun W, Zaboli D, Wang H, Liu Y, Arnaoutakis D, Khan T, Khan Z, Koch WM, Califano JA. Detection of TIMP3 promoter hypermethylation in salivary rinse as an independent predictor of local recurrence-free survival in head and neck cancer. Clin Cancer Res. 2012; 18:1082–1091.
69. Gaykalova DA, Vatapalli R, Wei Y, Tsai HL, Wang H, Zhang C, Hennessey PT, Guo T, Tan M, Li R, Ahn J, Khan Z, Westra WH, et al. Outlier Analysis Defines Zinc Finger Gene Family DNA Methylation in Tumors and Saliva of Head and Neck Cancer Patients. PLoS One. 2015; 10:e0142148.
70. Ovchinnikov DA, Wan Y, Coman WB, Pandit P, Cooper-White JJ, Herman JG, Punyadeera C. DNA Methylation at the Novel CpG Sites in the Promoter of MED15/PCQAP Gene as a Biomarker for Head and Neck Cancers. Biomark Insights. 2014; 9:53–60.
71. Rettori MM, de Carvalho AC, Bomfim Longo AL, de Oliveira CZ, Kowalski LP, Carvalho AL, Vettore AL. Prognostic significance of TIMP3 hypermethylation in post-treatment salivary rinse from head and neck squamous cell carcinoma patients. Carcinogenesis. 2013; 34:20–27.
72. Demokan S, Chang X, Chuang A, Mydlarz WK, Kaur J, Huang P, Khan Z, Khan T, Ostrow KL, Brait M, Hoque MO, Liegeois NJ, Sidransky D, et al. KIF1A and EDNRB are differentially methylated in primary HNSCC and salivary rinses. Int J Cancer. 2010; 127:2351–2359.
73. Ovchinnikov DA, Cooper MA, Pandit P, Coman WB, Cooper-White JJ, Keith P, Wolvetang EJ, Slowey PD, Punyadeera C. Tumor-suppressor Gene Promoter Hypermethylation in Saliva of Head and Neck Cancer Patients. Transl Oncol. 2012; 5:321–326.
74. Schussel J, Zhou XC, Zhang Z, Pattani K, Bermudez F, Jean-Charles G, McCaffrey T, Padhya T, Phelan J, Spivakovsky S, Brait M, Li R, Bowne HY, et al. EDNRB and DCC salivary rinse hypermethylation has a similar performance as expert clinical examination in discrimination of oral cancer/dysplasia versus benign lesions. Clin Cancer Res. 2013; 19:3268–3275.
75. Wei F, Lin CC, Joon A, Feng Z, Troche G, Lira ME, Chia D, Mao M, Ho CL, Su WC, Wong DT. Noninvasive saliva-based EGFR gene mutation detection in patients with lung cancer. Am J Respir Crit Care Med. 2014; 190:1117–1126.
76. Konno S, Morishita Y, Fukasawa M, Shu Y, Wang D, Tanaka R, Minami Y, Iijima T, Noguchi M. Anthracotic index and DNA methylation status of sputum contents can be used for identifying the population at risk of lung carcinoma. Cancer. 2004; 102:348–354.
77. Wang X, Cao A, Peng M, Hu C, Liu D, Gu T, Liu H. The value of chest CT scan and tumor markers detection in sputum for early diagnosis of peripheral lung cancer. Zhongguo Fei Ai Za Zhi. 2004; 7:58–63.
78. Belinsky SA, Klinge DM, Dekker JD, Smith MW, Bocklage TJ, Gilliland FD, Crowell RE, Karp DD, Stidley CA, Picchi MA. Gene promoter methylation in plasma and sputum increases with lung cancer risk. Clin Cancer Res. 2005; 11:6505–6511.
79. Olaussen KA, Soria JC, Park YW, Kim HJ, Kim SH, Ro JY, Andre F, Jang SJ. Assessing abnormal gene promoter methylation in paraffin-embedded sputum from patients with NSCLC. Eur J Cancer. 2005; 41:2112–2119.
80. Cirincione R, Lintas C, Conte D, Mariani L, Roz L, Vignola AM, Pastorino U, Sozzi G. Methylation profile in tumor and sputum samples of lung cancer patients detected by spiral computed tomography: a nested case-control study. Int J Cancer. 2006; 118:1248–1253.
81. Georgiou E, Valeri R, Tzimagiorgis G, Anzel J, Krikelis D, Tsilikas C, Sarikos G, Destouni C, Dimitriadou A, Kouidou S. Aberrant p16 promoter methylation among Greek lung cancer patients and smokers: correlation with smoking. Eur J Cancer Prev. 2007; 16:396–402.
82. Shivapurkar N, Stastny V, Suzuki M, Wistuba II, Li L, Zheng Y, Feng Z, Hol B, Prinsen C, Thunnissen FB, Gazdar AF. Application of a methylation gene panel by quantitative PCR for lung cancers. Cancer Lett. 2007; 247:56–71.
83. van der Drift MA, Prinsen CF, Hol BE, Bolijn AS, Jeunink MA, Dekhuijzen PN, Thunnissen FB. Can free DNA be detected in sputum of lung cancer patients? Lung Cancer. 2008; 61:385–390.
84. Hwang SH, Kim KU, Kim JE, Kim HH, Lee MK, Lee CH, Lee SY, Oh T, An S. Detection of HOXA9 gene methylation in tumor tissues and induced sputum samples from primary lung cancer patients. Clin Chem Lab Med. 2011; 49:699–704.
85. Zhang W, Sun Y, Lu G. [The diagnostic value of determination of p16 methylation of sputum exfoliated cells for peripheral lung cancer]. [Article in Chinese]. Zhongguo Fei Ai Za Zhi. 2004; 7:46–49.
86. Destro A, Bianchi P, Alloisio M, Laghi L, Di Gioia S, Malesci A, Cariboni U, Gribaudi G, Bulfamante G, Marchetti A, Bosari S, Infante M, Ravasi G, et al. K-ras and p16(INK4A)alterations in sputum of NSCLC patients and in heavy asymptomatic chronic smokers. Lung Cancer. 2004; 44:23–32.
87. Wang YC, Hsu HS, Chen TP, Chen JT. Molecular diagnostic markers for lung cancer in sputum and plasma. Ann N Y Acad Sci. 2006; 1075:179–184.
88. Mao L, Hruban RH, Boyle JO, Tockman M, Sidransky D. Detection of oncogene mutations in sputum precedes diagnosis of lung cancer. Cancer Res. 1994; 54:1634–1637.
89. Hubers AJ, Heideman DA, Burgers SA, Herder GJ, Sterk PJ, Rhodius RJ, Smit HJ, Krouwels F, Welling A, Witte BI, Duin S, Koning R, Comans EF, et al. DNA hypermethylation analysis in sputum for the diagnosis of lung cancer: training validation set approach. Br J Cancer. 2015; 112:1105–1113.
90. Wang X, Ling L, Su H, Cheng J, Jin L. Aberrant methylation of genes in sputum samples as diagnostic biomarkers for non-small cell lung cancer: a meta-analysis. Asian Pac J Cancer Prev. 2014; 15:4467–4474.
91. Miglio U, Mezzapelle R, Paganotti A, Veggiani C, Mercalli F, Mancuso G, Gaudino E, Rena O, Buosi R, Boldorini R. Frequency of O(6)-methylguanine-DNA methyltransferase promoter methylation in cytological samples from small cell lung cancer. Diagn Cytopathol. 2015; 43:947–952.
92. Cao L, Zhai Z, Li J, Xu M, Xia H, Xu X, Ma D, Mei X, Cui W. [The diagnostic value of flow cytometric DNA analysis of sputum in lung cancer]. [Article in Chinese]. Zhongguo Fei Ai Za Zhi. 2004; 7:202–205.
93. Castagnaro A, Marangio E, Verduri A, Chetta A, D'Ippolito R, Del Donno M, Olivieri D, Di Cola G. Microsatellite analysis of induced sputum DNA in patients with lung cancer in heavy smokers and in healthy subjects. Exp Lung Res. 2007; 33:289–301.
94. Brenner H, Hoffmeister M, Arndt V, Stegmaier C, Altenhofen L, Haug U. Protection from right- and left-sided colorectal neoplasms after colonoscopy: population-based study. J Natl Cancer Inst. 2010; 102:89–95.
95. Zhai RL, Xu F, Zhang P, Zhang WL, Wang H, Wang JL, Cai KL, Long YP, Lu XM, Tao KX, Wang GB. The Diagnostic Performance of Stool DNA Testing for Colorectal Cancer: A Systematic Review and Meta-Analysis. Medicine (Baltimore). 2016; 95:e2129.
96. Imperiale TF, Ransohoff DF, Itzkowitz SH, Levin TR, Lavin P, Lidgard GP, Ahlquist DA, Berger BM. Multitarget stool DNA testing for colorectal-cancer screening. N Engl J Med. 2014; 370:1287–1297.
97. Caldas C, Hahn SA, Hruban RH, Redston MS, Yeo CJ, Kern SE. Detection of K-ras mutations in the stool of patients with pancreatic adenocarcinoma and pancreatic ductal hyperplasia. Cancer Res. 1994; 54:3568–3573.
98. Kisiel JB, Yab TC, Taylor WR, Chari ST, Petersen GM, Mahoney DW, Ahlquist DA. Stool DNA testing for the detection of pancreatic cancer: assessment of methylation marker candidates. Cancer. 2012; 118:2623–2631.
99. Church TR, Wandell M, Lofton-Day C, Mongin SJ, Burger M, Payne SR, Castanos-Velez E, Blumenstein BA, Rosch T, Osborn N, Snover D, Day RW, Ransohoff DF, et al. Prospective evaluation of methylated SEPT9 in plasma for detection of asymptomatic colorectal cancer. Gut. 2014; 63:317–325.
100. Berger BM, Ahlquist DA. Stool DNA screening for colorectal neoplasia: biological and technical basis for high detection rates. Pathology. 2012; 44:80–88.
101. Kimura H, Fujiwara Y, Sone T, Kunitoh H, Tamura T, Kasahara K, Nishio K. EGFR mutation status in tumour-derived DNA from pleural effusion fluid is a practical basis for predicting the response to gefitinib. Br J Cancer. 2006; 95:1390–1395.
102. Yeo CD, Kim JW, Kim KH, Ha JH, Rhee CK, Kim SJ, Kim YK, Park CK, Lee SH, Park MS, Yim HW. Detection and comparison of EGFR mutations in matched tumor tissues, cell blocks, pleural effusions, and sera from patients with NSCLC with malignant pleural effusion, by PNA clamping and direct sequencing. Lung Cancer. 2013; 81:207–212.
103. Benlloch S, Galbis-Caravajal JM, Martin C, Sanchez-Paya J, Rodriguez-Paniagua JM, Romero S, Mafe JJ, Massuti B. Potential diagnostic value of methylation profile in pleural fluid and serum from cancer patients with pleural effusion. Cancer. 2006; 107:1859–1865.
104. Fujii M, Fujimoto N, Hiraki A, Gemba K, Aoe K, Umemura S, Katayama H, Takigawa N, Kiura K, Tanimoto M, Kishimoto T. Aberrant DNA methylation profile in pleural fluid for differential diagnosis of malignant pleural mesothelioma. Cancer Sci. 2012; 103:510–514.
105. Lin J, Gu Y, Du R, Deng M, Lu Y, Ding Y. Detection of EGFR mutation in supernatant, cell pellets of pleural effusion and tumor tissues from non-small cell lung cancer patients by high resolution melting analysis and sequencing. Int J Clin Exp Pathol. 2014; 7:8813–8822.
106. Liu X, Lu Y, Zhu G, Lei Y, Zheng L, Qin H, Tang C, Ellison G, McCormack R, Ji Q. The diagnostic accuracy of pleural effusion and plasma samples versus tumour tissue for detection of EGFR mutation in patients with advanced non-small cell lung cancer: comparison of methodologies. J Clin Pathol. 2013; 66:1065–1069.
107. Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, Bartlett BR, Wang H, Luber B, Alani RM, Antonarakis ES, Azad NS, Bardelli A, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014; 6:224ra224.
108. Pentsova EI, Shah RH, Tang J, Boire A, You D, Briggs S, Omuro A, Lin X, Fleisher M, Grommes C, Panageas KS, Meng F, Selcuklu SD, et al. Evaluating Cancer of the Central Nervous System Through Next-Generation Sequencing of Cerebrospinal Fluid. J Clin Oncol. 2016; 34:2404–2415.
109. Wang Y, Springer S, Zhang M, McMahon KW, Kinde I, Dobbyn L, Ptak J, Brem H, Chaichana K, Gallia GL, Gokaslan ZL, Groves ML, Jallo GI, et al. Detection of tumor-derived DNA in cerebrospinal fluid of patients with primary tumors of the brain and spinal cord. Proc Natl Acad Sci U S A. 2015; 112:9704–9709.
110. Hegemann M, Stenzl A, Bedke J, Chi KN, Black PC, Todenhofer T. Liquid biopsy: ready to guide therapy in advanced prostate cancer? BJU Int. 2016.
111. Dickinson BT, Kisiel J, Ahlquist DA, Grady WM. Molecular markers for colorectal cancer screening. Gut. 2015; 64:1485–1494.
112. Heitzer E, Auer M, Ulz P, Geigl JB, Speicher MR. Circulating tumor cells and DNA as liquid biopsies. Genome Med. 2013; 5:73.

