INTRODUCTION
With the completion of the Human Genome Project (HGP) and beginning of the post-genomic era, non-coding RNA (ncRNA) had aroused great interest and attention in various areas of research. After the Nobel Prize was awarded for the discovery of small interfering RNA (siRNA) in 2006 [1], ncRNAs, including long non-coding RNAs (LncRNAs) gradually gained a dominant position in biomedical research. Thousands of LncRNAs have been discovered in the human genome over the past three years, implying that LncRNAs may play an irreplaceable role in cellular functioning. Although LncRNAs do not encode functional proteins, they are involved in many physiological processes, playing essential roles in maintaining cell proliferation and differentiation [2, 3]. Increasing evidence supports the hypothesis that LncRNAs participate in physiological and pathological processes by modulating gene expression at the epigenetic, transcriptional, and posttranscriptional levels [4, 5]. Moreover, abnormal expression of LncRNAs has been associated with a variety of human diseases, including psoriasis [6], coronary artery disease [7, 8], diabetes mellitus (DM) [9, 10], tumors [11–13], etc. However, their comprehensive bio-functions and molecular mechanisms of LncRNAs in human diseases still remain elusive. Therefore, more efforts should be made to explore the LncRNA world.
DM is a class of metabolic disorders characterized by hyperglycaemia resulting from the relatively reduced insulin secretion and occurrence of insulin resistance, which can be either due to inherited or environmental factors [14, 15]. Chronic hyperglycaemia of DM is associated with long-term damage, dysfunction and failure of multiple organ systems, especially the kidneys, eyes, blood vessels, heart, and nerves [16, 17]. DM has rapidly become a global health problem affecting more than 400 million people worldwide [18], in fact it has become the third most prevalent non-infective disease (NCD) exceeded only by cardiovascular diseases and cancer [19]. The global prevalence of DM is expected to increase from 4% in 1995 to 5.4% by the year 2025 [20], and that number of patients will reach 642 million by 2040 [21]. Currently, the countries with the largest number of diabetic patients are India, China, and United States [22]. Inspiringly, the morbidity and mortality caused by DM can be reduced by regular screening, early detection, and appropriate treatment of chronic complications [23, 24]. Therefore, there is a great urgency to prevent and treat diabetes and its associated complications. On account of the widespread use of new technologies, several LncRNAs have been proved as novel regulatory players in the molecular biology of DM, which may provide new strategies for the prevention, early diagnosis and treatment of DM. In this review, we summarize the origin and overview function of LncRNAs; highlighted the roles of LncRNAs in DM; and outlined the molecular mechanisms of LncRNAs in DM. Further, the application of LncRNAs as biomarkers in the prevention, early diagnosis, and treatment of DM has also been discussed.
Definition and situation of LncRNAs
The Human Genome Project revealed that protein-coding genes represent less than 2% of the total genome sequence [25], the remaining greater portion of DNA sequences do not code for proteins, which are regarded as “junk DNAs” that have accumulated because of the process of evolution [26, 27]. In addition, most of the “junk DNAs” are intron DNAs in animals, and are termed as non-coding DNAs (ncDNAs) [28, 29]. Generally, based on the length of a transcript, ncRNAs are divided into two categories: small ncRNAs (fewer than 200 nucleotides) and long ncRNAs (more than 200 nucleotides) [30, 31]. With the progress of the whole-genome resequencing efforts, the latest GENCODE release (version 25) (http://www.gencodegenes.org) indicated that 75–90% of the human genome was transcribed to generate a series of LncRNAs [32, 33]. Based on previous studies, the comprehensive definition of LncRNA is as follows: LncRNAs are a subset of RNAs first found in a eukaryotic cell and have a transcribing length of 200–100000 nt; they lack a complete functional open reading frame (ORF), rarely encode a functional short peptide, and are located in either the nucleus or cytoplasm [34, 35]. LncRNAs usually function as primary transcribed or as a spliced RNA [36]. Further, most of them are similar to mRNAs with respect to structural features like poly(A) tails, 5′-caps, and promoter structure. They are transcribed by RNA polymerase II and often alternatively spliced to be finally polyadenylation [37, 38].
Interestingly, the LncRNA sequences are conserved, and they exhibit strong tissue- and cell-specific expression patterns in humans [39, 40]. LncRNAs have recently gained widespread attention over microRNAs (miRNAs) and siRNAs. With the development of RNA-seq using next-generation sequencing, a growing number of LncRNAs have been discovered and defined. The total number of functional LncRNAs thus far identified has reached 184 for human disease (http://lncrnadb.org/) [41]; nevertheless, this is only the tip of the iceberg when compared to the total number of LncRNAs predicted by bio-informatics software analysis. By elucidating the relationship between LncRNAs and disease, we can further understand the complicated and multilayer regulatory system in human, and provide novel therapeutic strategies using LncRNAs as molecular markers and potential drug targets.
Classification and functions of LncRNAs
With advancement the sequencing technologies, especially RNA-seq technology of next-generation sequencing, an increasing number of LncRNAs have been found [42, 43]. LncRNAs have displayed characteristics that can be described as ‘three more’, namely more types, more patterns and more quantity. Although many LncRNAs have been discovered, still their origin is not completely understood. Possible origin includes gene mutations, tandem duplication events, chromatin rearrangement, retrotransposition, and insertion of a transposable element [44]. LncRNAs can be classified into five categories according to their genomic proximity with neighbouring transcripts: (1) sense strand synthesis, which are located on the sense strand of annotated transcription units; (2) antisense strand synthesis, which are located on the antisense strand of annotated transcription units; (3) intronic synthesis, which are from introns of annotated genes; (4) bidirectional synthesis, which arise from both sense and antisense directions of transcription start areas; and (5) intergenic synthesis (also known as lincRNAs), whose LncRNA transcripts are from introns of annotated genes [45, 46].
Accumulating evidence supports that LncRNAs are an important class of regulatory molecules in the human genome. The long nucleotide chain of LncRNAs can either form a complex spatial structure and interact with protein factors, or provide a large segment for the concurrent binding of many molecules that collectively participate in epigenetic regulation, X-chromosome silencing, genomic imprinting, nuclear and cytoplasmic trafficking, transcriptional activation and interference, mRNA splicing and degradation, etc. [36, 46] Although there is little knowledge about LncRNAs at present, they have been shown to have crucial roles in regulating various biological processes, such as cell proliferation, differentiation, apoptosis, senescence and death, etc. [47, 48]. Recently, it has been accepted that the molecular functions of LncRNAs at the epigenetic, transcriptional and post-transcriptional can be subdivided as follows (Figure 1): (1) recruiting and interacting with proteins: For example, LncRNA RMRP acts as a scaffold to interact with DEAD-box helicase 5 (DDX5) and Retinoid-related orphan receptor gamma (RORγ) to control the transcriptional program in Th17 cells [49]; (2) acting as a co-regulator or a co-repressor: For example, An antisense LncRNA was found to localize at the 5’ long-term repeats (LTR) of HIV promoter, and suppress transcription of viral genes by recruiting enhancer of zeste homolog-2 (EZH2), histone deacetylase 1 (HDAC1), and DNA-methyltransferase 3 alpha (Dnmt3a) to form a transcriptional repressor complex [50]; (3) acting as a decoy: For example, p50-associated COX-2 extragenic RNA (PACER) acts as a decoy molecule to promote the transcription of cytochrome c oxidase subunit II (COX2) in the NF-κB signalling pathway [51]; (4) acting as host genes for miRNA: For example, LncRNA H19 serves as the reservoir of miR-675-3p and miR-675-5p, which are induced during skeletal muscle differentiation [52]; (5) interacting with miRNA: For example, Long intergenic non-protein coding RNA, muscle differentiation 1 (Linc-MD1) acts as a competing endogenous lncRNA (ceRNA) to sponge miR-133 and thus regulates muscle cell differentiation in skeletal muscle [53].
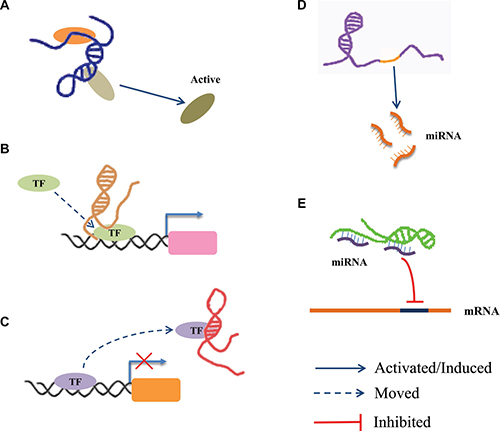
Figure 1: The regulatory mechanism of LncRNAs. (A) LncRNA acts as a scaffold via recruiting and interacting with proteins and regulate the activity of proteins. (B) LncRNA acts as a guide to promote the gene expression via recruiting transcription factors (TF) to the region of the gene promoter. (C) LncRNA acts as a decoy via interacting with TF to inhibit transcriptional regulation. (D) LncRNA acts as host genes to promote the production of miRNA. (E) LncRNA serves as ceRNA to interact with miRNAs.
Diabetes related-LncRNA
With the rapid increase in population, aging, urbanization and the prevalence of obesity and physical inactivity, DM has become a global health problem in recent years. The development and progression of DM is a complex process that involves multiple factors and steps. Too much food and too little physical exercises are just two standard factors. The development and progression of DM is closely related to environmental factors, inherited factors, microbial infections and a decreased immune system response caused by other diseases. At present, the treatment of DM consists of five main aspects: diabetes education, self-monitoring of blood glucose, diet therapy, exercise and drug treatment. Only these five aspects of the treatment go hand in hand and can effectively control the blood glucose, thereby delaying the progress of DM. Based on pathogenesis, DM is divided into four types: type 1 diabetes mellitus (T1DM), type 2 diabetes mellitus (T2DM), gestational diabetes mellitus, and other specific types of diabetes mellitus. Amongst these, T2DM is the most common type throughout the world, exceeding 90% of all cases of diabetes [54]. T2DM is mainly influenced by lifestyle factors and genetic components [55]. The development of DM is accompanied by a series of complications, such as diabetic nephropathy (DN), diabetic retinopathy (DR), diabetic cardiomyopathy (DCM) and diabetic neuropathic pain (DNP).
LncRNAs, as multifunctional molecules, have recently gained widespread attention. Tremendous amount of data indicate that the DM susceptibility loci is associated with abnormal expression of LncRNAs [56–73], and part of these LncRNAs play a crucial role in the progression of DM and its associated complications [73–77], involving in β islet cell disorders, insulin resistance and epigenetic regulation, etc. However, our understanding about LncRNAs and their role in DM is still in its infancy. Through Literature search was performed in PubMed, Embase, and Web of Science with the following search terms: “Diabetes Mellitus” or “Diabetes” or “Diabetic” AND “Long Noncoding RNA” or “ LncRNA” or “ Long Non-Coding RNA” or “Long Non Protein Coding RNA” or “Long Untranslated RNA” or “Long Intergenic Non Protein Coding RNA” or “LincRNAs”. After excluding meta-analysis, letters, comments, case reports, reviews and duplicate publications, a total of 132 papers were found on the relations of LncRNAs with DM by the end of March 28, 2017. The number of relevant publications showed an explosive increase since 2012, indicating increasing focus of researchers on the relationship between DM and LncRNAs. By integrating the published literature and analysing the LncRNA Disease database (http://www.cuilab.cn/lncrnadisease) and LncRNAWiki database (http://lncrna.big.ac.cn/index.php/Main_Page) we found that till date, a total of twenty-eight LncRNAs have been associated with DM. Amongst these nineteen have been detected in human DM. An overview of the human lncRNAs that are associated with DM is shown in Table 1.
Table 1: LncRNAs are associated with human diabetes mellitus
LncRNAs Name |
RNA Description |
Aliases |
Genomic Location |
Diabetes Type |
Dysfunction Type |
Ref. |
|---|---|---|---|---|---|---|
HYMAI |
Hydatidiform mole associated and imprinted |
NCRNA00020 |
6q24.2 |
Transient neonatal diabetes |
Expression |
[67] |
CDKN2B-AS1 |
Cyclin dependent kinase inhibitor 2B (CDKN2B) antisense RNA 1 |
ANRIL,DKN2BAS,CDKN2B-ASC, T12,p15AS NCRNA00089PCA |
9p21.3 |
Diabetes |
Mutation |
[68] |
Expression |
[69] |
|||||
Type 2 diabetes |
Locus/Mutation |
[70] |
||||
Mutation |
||||||
IGF2-AS |
Insulin like growth factor 2 antisense RNA |
IGF2-AS1, IGF2AS, PEG8 |
11p15.5 |
Type 1 diabetes |
Mutation |
[74] |
LINC00271 |
Long intergenic non-protein coding RNA 271 |
C6orf217, NCRNA00271 |
6q23.3 |
Type 2 diabetes |
Mutation |
[75] |
MALAT1 |
Metastasis associated lung adenocarcinoma transcript 1 |
HCN, LINC00047, NCRNA00047NEA,T2 PRO2853 |
11q13.1 |
Diabetes |
Regulation |
[85] |
MEG3 |
Maternally expressed 3 |
FP504, GTL2, LINC00023, NCRNA00023, PRO0518,PRO2160, |
14q32.2 |
Type 1 diabetes |
Locus |
[76] |
PDZRN3-AS1 |
PDZ domain containing ring finger 3(PDZRN3) antisense RNA 1 |
N/A |
3p13 |
Type 2 diabetes |
Mutation |
[77] |
PTV1 |
Plasmacytoma variant translocation 1 |
LINC00079, |
8q24.21 |
Type 1 diabetes |
Mutation |
[78] |
Type 2 diabetes |
Mutation |
|||||
RNCR3 |
Retinal non-coding RNA3 |
LINC00599 |
8p23.1 |
Diabetes |
Expression |
[86] |
H19 |
H19, imprinted maternally expressed transcript |
ASM, ASM1, BWS, D11S813E, LINC00008, NCRNA00008,WT2 |
11p15.5 |
Type 2 diabetes |
Locus |
[80] |
Regulation |
[81] |
|||||
PLUT |
PDX1 associated lncRNA,upregulator of transcription |
PLUTO; HI-LNC71; PDX1-AS1 |
13q12.2 |
Type 2 diabetes |
Regulation |
[87] |
MIAT |
Myocardial infarction associated transcript |
C22orf35, RNCR2, GOMAFU, LINC00066, NCRNA00066, lncRNA-MIAT |
22q12.1 |
Diabetes |
Regulation |
[88] |
Type 2 diabetes |
Expression |
[80] |
||||
LINC01611 |
Long intergenic non-protein coding RNA 1611 |
RP1-90L14.1, TCONS_l2_00025430 |
6q14.3 |
Diabetes |
Locus |
[82] |
GAS5 |
Growth arrest specific 5 |
NCRNA00030, SNHG2 |
1q25.1 |
Type 2 diabetes |
Expression |
[83] |
LINC01370 |
long intergenic non-protein coding RNA 1370 |
HILNC25, HI-LNC25 |
20q12 |
Type 2 diabetes |
Regulation |
[84] |
LINC00673 |
Long intergenic non-protein coding RNA 673 |
HI-LNC75 |
17q24.3 |
Locus/ Expression |
||
LINC01512 |
Long intergenic non-protein coding RNA 1512 |
LOC100132354, TCONS_00011120, HI-LNC77 |
6p21.1 |
Locus |
||
LINC01574 |
Long intergenic non-protein coding RNA 1574 |
HI-LNC12, TCONS_00009551 |
5q35.2 |
Locus |
||
TUNAR |
TCL1 upstream neural differentiation-associated RNA |
TUNA, HI-LNC78 |
14q32.2 |
Locus |
NOTE: Mutation—Genetic variation in lncRNA genes region on chromosome causes disease and influences susceptibility; Expression—the dysfunction expression of LncRNAs were associated with diabetes; Locus—A genetic susceptibility locus; Regulation—Overexpression or depletion of a LncRNA will regulate other molecules expression to lead to the progression of diabetes. Diabetes—its associated complications, involving in diabetic retinopathy (DR), diabetic nephropathy(DN), diabetic cardiomyopathy (DCM) and diabetic neuropathic pain (DNP).
Dysregulation and roles of LncRNAs in DM
LncRNAs and pancreatic β-cells
Pancreatic islets consist of a class of hormone-releasing cells, and about 70% of pancreatic cells are β-cells. Pancreatic β-cells play a central role in maintaining blood glucose homeostasis through insulin secretion [55, 78]. In turn, insulin controls the carbohydrate and lipid metabolism in the human body. Therefore, reduced secretion of insulin, due to β-cell dysfunction and/or loss, leads to different forms of DM [79]. Recent studies demonstrated that the expression profile of LncRNAs changes spatiotemporally during the maturation process of pancreatic β-cells [73, 76]. Abnormal expression and dysregulation of these LncRNAs, accompanied with the dysfunction of pancreatic β-cells influences cell survival, proliferation, differentiation, or function (especially insulin secretion) and apoptosis. For example, using a whole-genome transcriptome map, Morán et al reported that a series of islet lncRNAs (e.g., HI-LNC25, HI-LNC75, HI-LNC12, and HI-LNC78) were dynamically regulated. They showed that these LncRNAs were not activated in human embryonic pancreatic progenitor cells and were activated in mature pancreatic cells, thus suggesting that these LncRNAs are an integral component of the β cell differentiation and maturation program [73]. Another report demonstrated that a β cell-specific LncRNA, HI-LNC25 downregulates GLIS family zinc finger 3 (GLIS3) mRNA that is an islet transcription factor [80] via a gene regulatory function to influence pancreatic β-cell programing. Moreover, Zou et al. [81] demonstrated that the expression of Sox2 and other stem cell markers (such as Oct4 and Nanog) was downregulated in human amniotic epithelial stem cells (HuAECs) after overexpression of a LincRNA-regulator of reprogramming (LincRNA-ROR) specific siRNA. Meanwhile, miR-145 can play a role in the expression of LncRNA-ROR and Sox2, which is deduced from the results of a luciferase reporter assay. Additionally, LincRNA-ROR effectively maintains Sox2 gene expression through competitive binding with miR-145 to maintain the β islet-like cell differentiation efficiency. During proliferation of pancreatic β-cells, Mutskov et al. [82] showed that the expression of insulin like growth factor 2 antisense RNA (IGF2-AS) was upregulated after β cells were stimulated with high concentration of glucose, thus potentially regulating β cell proliferation. Inflammatory microenvironment is an important factor that results in β cell apoptosis [83]. Motterle et al. [84] showed that the upregulation of mouse islet LncRNA-1 promoted nuclear factor-kappa B (NF-κB) expression that in turn induces the sensitive β cells to undergo apoptosis in pre-diabetic nonobese diabetic (NOD) mice. Taken together, the dysregulation of miRNAs may lead to diabetes mellitus by regulating the biology and functions of pancreatic β-cells.
LncRNA and insulin resistance
Insulin resistance is characterised by impaired cellular response to insulin, and the inability of physiological levels of insulin to achieve glucose homeostasis. This pathological state of reduced insulin sensitivity or reactivity in the target tissues or cells is a hallmark of T2DM [18]. The process of insulin resistance is associated with defects in insulin signalling downstream of insulin receptor (INSR) [85], insulin Receptor Substrate 1/2 (IRS-1/2) [86], Phosphoinositide 3-Kinase (PI3K)/AKT Serine/Threonine Kinase (AKT) [87] and Glucose Transporter 4 (GLUT4) [88]. Recently, a growing body of evidence indicates that LncRNAs may act as a link between insulin signalling and insulin resistance. They act as key regulators of gene expression and play significant role in regulating the function of insulin-target tissues, especially liver [89, 90]. Zhu et al. [90] demonstrated that LncRNA maternally expressed gene 3 (MEG3) was upregulated in livers of high-fat diet fed and ob/ob mice. Consequently, they enhanced hepatic insulin resistance via increased expression of forkhead box O1 (FoxO1), a critical regulator of hepatic glucose and lipid metabolism via its ability to regulate the expression of G6pc and Pepck in gluconeogenesis [91]. Moreover, Yan and colleagues [89] reported that the expression of metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) is increased in livers of ob/ob mice, and promoted hepatic insulin resistance by increasing the stability nuclear sterol regulatory element binding transcription factor 1c (SREBP-1c). Therefore, inhibition of the hepatic insulin resistance-associated LncRNAs may be a potential strategy against hyperglycaemia associated with T2DM.
LncRNAs and diabetic epigenetic modification
Epigenetics is related to heritable changes in gene expression caused by mechanisms that do not affect the DNA sequence itself. Epigenetic alterations have been repeatedly proposed as a likely molecular mechanism linking birth weight and later determinant of health in childhood or even later in life [92]. Epigenetic modifications can regulate DM-related genes by activating or inhibiting their transcription, thereby affecting glucose homeostasis, β-cell function, insulin secretion and vasculopathy, etc. Moreover, LncRNAs can be localized in the nucleus and participate in the assembly of long noncoding RNA-dependent nuclear bodies by forming chromatin remodelling complexes [93, 94]. An emerging concept suggests that not only LncRNAs regulate gene expression at the epigenetic level, but also their expression could regulated by epigenetic modifications [95]. Epigenetic modifications occur mainly by two most widely studied mechanisms, namely DNA methylation and histone modifications, which are known to be crucial for proper control of gene expression [96, 97]. DNA methylation occurs mainly in sequences enriched in CpG dinucleotides, or the so-called CpG islands that are mainly located in the proximal region of promoters and are underrepresented in the rest of the genome. Methylation of CpG islands is generally associated with gene silencing whereas hypomethylation is linked to hyperactivation of gene expression [79, 96]. A study [98] showed that the abnormal expression of hydatidiform mole associated and imprinted (HYMAI), a special LncRNA that was expressed in paternal alleles [99], is closely associated with transient neonatal diabetes mellitus (TNDM), and such TNDM co-occurs with defective DNA methylation. The DNA methylation at the insulin like growth factor 2 (IGF2)/H19 gene locus, that may be induced by intrauterine hyperglycaemia [100], and is a cornerstone that links birth weight and foetal metabolic programming of late onset obesity [101]. In addition, abnormal DNA methylation of LncRNA MEG3 gene was associated with T1DM [65] and T2DM [102]. However, LncRNAs are also known to regulate DNA methylation. For instance, Dhawan et al. [103] demonstrated that MALAT1 can regulate the level of methyl-CpG binding protein 2 (MeCP2) [104], that plays a role in regulating the methylation of aristaless related homeobox (Arx), to maintain the differentiated status of β cells, and hence, the dynamic equilibrium of glucose. Histone modifications refer to post-transcriptional changes at the amino-terminal of histone subunits. Some histone modifications, such as acetylation, are labile and associated with gene activation, while others, such as methylation, are stable and lead to gene inactivation [79, 96]. For example, Long et al. [105] showed that the LncRNA taurine-upregulated 1 (Tug1) expression is decreased in diabetic podocytes. However, since Tug1 regulates the transcription coactivator PPARγ coactivator 1α (PGC-1α) expression by epigenetically enhancing PGC-1α promoter activity, it influences chronic kidney disease (CKD) development. Meanwhile, Zhuo et al. [106] demonstrated that H19 overexpression induced by administration of lentivirus pcDNA-H19 can inhibit the expression of autophagy-related genes (e.g. LC3-II, ATG7b and BECN1) in diabetic rats. Furthermore, H19 could directly bind to EZH2 in an RNA-binding protein immunoprecipitation (RIP) assay, and the complex of H19/EZH2 could inhibit autophagy by epigenetically silencing the DIRAS family GTPase 3 (DIRAS3) in cardiomyocytes, as seen in a chromatin immunoprecipitation (ChIP) assay. However, the mechanism of and relationship between LncRNAs and diabetic epigenetic modifications is still at an exploratory stage, and the detailed mechanisms still need further research.
LncRNAs and diabetic complications
lncRNA and diabetic nephropathy (DN)
Diabetic nephropathy (DN) is the leading cause of end-stage renal disease (ESRD) in DM, with an incidence of 20%–40%, and is thus a major cause of death and disability in DM [107, 108]. DN is characterized by progressive renal interstitial fibrosis [15] that results in a series of pathological changes, including excessive accumulation of extracellular matrix (ECM), mesangial expansion, thickening of glomerular and tubular basement membranes and increased production of mesangial matrix [109, 110]. Recent studies reported that LncRNAs play a crucial role in the development of DN, and regulate ECM accumulation. Importantly, the gene locus of plasmacytoma variant translocation 1 (PVT1) was reported to be associated with ESRD, which in turn is attributed to T1DM [111] and T2DM [69]. Alvarez et al. [67] reported that PVT1 expression was significantly upregulated in response to glucose treatment in human mesangial cells. The upregulated PVT1 promotes the level of the major ECM proteins, like fibronectin 1 (FN1) and collagen type IV alpha 1 chain (COL4A1), and two key regulators of ECM proteins, namely, transforming growth factor beta 1 (TGFβ1) and plasminogen activator inhibitor-1 (PAI-1). Moreover, they also found that PVT1 acts as a host gene to drive the production of miR-1207-5p which can directly target glucose-6-phosphate dehydrogenase (G6PD), prostate transmembrane protein, androgen induced 1 (PMEPA1), 3-phosphoinositide dependent protein kinase 1 (PDPK1) and SMAD family member 7 (SMAD7), and increase the expression of TGF-β1, PAI-1, and FN1 to regulate the pathogenesis of DN [112]. Furthermore, using miRNA microarray analysis and a luciferase reporter assay, Duan et al. [113] demonstrated that miRNA-377 was remarkably up-regulated in db/db DN mice and LncRNA Tug1 was a direct target of miR-377, respectively. LncRNA Tug1 also acts as an endogenous sponge of miR-377 and downregulates miR-377 expression levels as measured in quantitative real time PCR (qRT-PCR); this relieves the inhibition of its target gene PPARγ, alleviating PAI-1 and TGF-β1 accumulation in mesangial cells (MC). In another report, LincRNA-Gm4419 was shown to participate in NF-κB/NLRP3 inflammasome-mediated inflammation in db/db DN mice [114]. In addition, Zhou et al. [115] demonstrated that LncRNA Myocardial infarction associated transcript (MIAT)- Nrf2 axis may serve as an important signalling pathway for high glucose induced renal tubular epithelial injury, and also in the pathological process of acute kidney injury (AKI).
LncRNAs and diabetic retinopathy (DR)
Diabetic retinopathy (DR) is one of the most common complications of DM and a leading cause of catastrophic vision loss in developed nations [116]. DR-induced deterioration of vision is usually accompanied by inflammation, retinal ischemia, neovascularization, vascular hyperpermeability, and vascular cell dysfunction [117, 118]. Therefore, structural or functional abnormality of retinal microvasculature is an important characteristic of DR [119, 120]. Based on the proliferative status of retinal neovasculature, DR can be divided into two categories: non-proliferative diabetic retinopathy (NPDR) and proliferative diabetic retinopathy (PDR) [121, 122]. Through DR has obvious clinical and pathological characteristics, identifying biomarkers to predict DR or to determine therapeutic response is important because of the complex pathogenesis and ambiguous risk factors [123, 124]. Recently, accumulating studies reported that LncRNAs play an important role in the development of DR, especially in retinal microvascular dysfunction. Importantly, MALAT1 was the earliest reported aberrantly expressed LncRNA in DR [125], and first identified to be crucial for the angiogenic response of endothelial cells as well as for vascularization in vivo [120, 126]. Soon afterwards, a series of LncRNAs have been reported to be associated with DR [127], such as retinal non-coding RNA3 (RNCR3) [128, 129], MIAT [77, 130], MEG3 [131], etc. Interestingly, the regulatory mechanism involving LncRNA-miRNA-mRNA manner. For example, Shan et al. [128, 129] revealed that LncRNA RNCR3 is significantly up-regulated in diabetic mice or in RF/6A cells, which were exposed to high glucose. This was measured by qRT-PCR. RNCR3 can act as ceRNA with miR-185-5p to regulate its target gene kruppel-like factor 2 (KLF2) as observed by using a luciferase reporter assay and the gene knockdown technique to silence the expression of RNCR3. The RNCR3/miR-185-5p/KLF2 complex was also involved in atherosclerosis and DM-induced retinal microvascular abnormalities. Moreover, LncRNA MIAT has been related closely with DM-induced retinal microvascular dysfunction in vivo and in vitro, via its interaction with miR-150-5p [77] and miR-29b [130]. Furthermore, LncRNAs are able to take part in regulating retinal ganglion cell (RGC) injury, which is another important pathological feature of DR [132]. Li et al. [133] demonstrated that LncRNA SOX2 overlapping transcript (LncRNA-Sox2OT) knockdown can protect RGCs against high glucose-induced injury and play neuroprotective role in DM-related retinal neurodegeneration in vivo. Meanwhile, Liu et al. [75] revealed that RNCR3 knockdown alleviates DM-induced retinal neurodegeneration by reducing the expression of glial reactivity-related genes including glial fibrillary acidic protein (GFAP) and vimentin. These studies suggest LncRNAs have a strong potential to act as therapeutic targets for treating DR.
LncRNAs and diabetic cardiomyopathy (DCM)
Diabetic cardiomyopathy (DCM) is one of the prominent cardiovascular complications of DM, and carries a substantial risk for the subsequent development of heart failure and increased mortality [134]. DCM is defined as myocardial dysfunction occurring in patients with diabetes in the absence of coronary artery disease, hypertension, or valvular heart disease [135, 136]. The pathogenesis of DCM may involve inflammation, oxidative stress, mitochondrial dysfunction, impaired calcium handling, renin-angiotensin system activation, cardiomyocyte apoptosis [137]. Recently, increasing evidence demonstrated that LncRNAs play a crucial role in regulating cardiac muscle [138], such as growth arrest-specific 5 (GAS5) [139], cardiac mesoderm enhancer-associated noncoding RNA (CARMEN) [140], urothelial carcinoma-associated 1 (UCA1) [141],etc. However, currently only two lncRNAs have been associated with DCM, namely, MALAT1 [142, 143], and H19 [106, 144]. MALAT1 expression is upregulated in patients with myocardial infarction (MI) [58] and in cardiac tissue of diabetic rats [142, 143]. Two studies by Zhang and colleagues [142, 143] revealed that MALAT1 knockdown can reduce diabetes-induced myocardial inflammation as well as cardiomyocyte apoptosis, and consequently improve left ventricular function in diabetic rats. Compared with MALAT1, the regulatory mechanism of H19 in DCM was shown to be more complex. The H19 LncRNA encodes a 2.6 kb capped, spliced and polyadenylated noncoding RNA that is predominantly cytoplasmic, with a minor fraction being localised in the nucleus [145]. However, H19, that is downregulated by acute hyperinsulinemia [70], can exert its biological function in the nucleus by binding and recruiting the histone methyltransferase EZH2 at the promoter of DIRAS3. EZH2 recruitment in turn leads to the epigenetic silencing of DIRAS3, and therefore inhibiting DIRAS3-induced autophagy, and protecting cardiomyocytes exposed to high glucose [106]. In an alternative mechanism, cytoplasmic H19 can act as miRNA sponge to sequester miR-106a as well as the miR-let7 family members [146, 147]. Moreover, H19 can serve as a precursor of miR-675 that will in turn, post-transcriptionally regulate a number of target genes involved in cell proliferation and differentiation [52, 148]. Zhang et al. [144] demonstrated that miR-675 expression was decreased after transfection with H19 siRNA in cardiomyocytes. As shown by a luciferase reporter assay, miR-675 can directly target the voltage-dependent anion channel 1 (VDAC1), thereby forming the H19/miR-675/VDAC1 axis to inhibit apoptosis induced by high glucose. This finding may provide a novel therapeutic strategy for the treatment of DCM.
LncRNAs and diabetic neuropathic pain (DNP)
Diabetic neuropathic pain (DNP) is one of the most common chronic complications of T2DM, fifty percent of diabetic patients suffer from DNP [149]. Typical symptoms of DNP include pathological nerve pain, including spontaneous pain, hyperalgesia (increased pain perception to noxious stimuli), and allodynia (pain to normally innocuous stimuli) [149, 150]. Patients with DNP are predominantly characterized by sensory symptoms like “glove-and-stocking” distribution and have burning/ lancing (stabbing) sensations, tingling (“pins and needles” or paraesthesia), or shooting (electric shock) [149, 150]. DNP has become a substantial problem in the field of intractable pain therapy [151], which is frequently accompanied by diminished quality of life in diabetic patients [152]. Recently, a few studies show that LncRNAs play a crucial role in progress of DNP. LncRNA-NONRATT021972, a blood glucose regulator [153], is a typical representative associated with DNP. On one hand, NONRATT021972 was showed to be related with diabetic autonomic neuropathy (DAN) [154] by Xu and colleagues [155]. Their study reported that NONRATT021972 induces the expression of tumour necrosis factor-α (TNF-α) and serine phosphorylation of insulin receptor substrate 1 (IRS1) in superior cervical ganglia (SCG) to mediate heart rate variability (HRV) in diabetic rats. Moreover, NONRATT021972 is associated with the abnormalities of function in the dorsal root ganglia (DRG) [149] that is a clinical characteristic of DNP. Liu et al. [156] reported that NONRATT021972 siRNA treatment can decrease the expression levels of purinergic receptor P2X 7 (P2X7) and inflammatory factors TNF-α to inhibit the excitability of DRG neurons, reducing mechanical and thermal hyperalgesia in T2DM rats. Moreover, Peng et al. [157] demonstrated that NONRATT021972 is a potential induction factor to promote the development of DNP via the P2X3 receptor expression in rat DRG. In another study, LncRNA uc.48+ was also reported to mediate the P2X3 receptor expression, thereby promote the excitatory transmission in DRG [158]. Overall, these studies provide novel insights for the treatment of DNP.
LncRNAs and diabetic other complications associated with DM
LncRNAs also display a role other complications associated with DM. For instance, in the aspects of inflammatory complications of DM, Reddy et al. [159] reported that lncRNA-E330013P06 was upregulated in macrophages from db/db mice and T2DM patients, and induced the expression of several genes to heighten the inflammatory response. In addition, Puthanveetil et al. [160] demonstrated that glucose stimulates human umbilical vein endothelial cells (HUVECs) and can increase the expression of MALAT1 and serum amyloid A 3 (SAA3) as measured in a qRT-PCR. The expression of MALAT1, SAA3 and inflammatory mediators IL-6 and TNF-α were down-regulated after transfection of the MALAT1-specific siRNA, which may eventually promote diabetes-induced micro and macrovascular complications. Further, antisense noncoding RNA for the INK4 locus (ANRIL) was shown to upregulate vascular endothelial growth factor (VEGF) expression and promote angiogenesis by activating NF-κB inflammation signalling pathway in DM combined cerebral infarction (CI) rats [161]. In the aspects of diabetes-associated cognitive impairment, Li et al. [162] demonstrated that PVT1-mediated autophagy may protect hippocampal neurons from impairment of synaptic plasticity and apoptosis, and then ameliorate cognitive impairment in streptozotocin (STZ)-induced diabetic mice with the cognitive impairment induced by 3-methyladenine (3-MA). In the aspects of diabetes-associated abnormal immunity, LncRNAs show a potential regulatory function in T1DM. T1DM is an organ-specific autoimmune disease characterized by chronic and progressive apoptotic destruction of pancreatic β cells. Research in recent years indicates that LncRNAs modulate the apoptosis of sensitive β cell and they may emerge as novel biomarkers in diagnosis and as targets for development of new therapies for T1DM [84, 163].
Potential clinical applications of LncRNAs in DM
Through the tremendous progress in the therapy of DM, the morbidity and mortality caused by DM is still high in worldwide. Inspirationally, numerous studies showed that the therapeutic efficacy for DM patients could be improved via the early detection of DM, followed by timely intervention with particular attention to blood pressure control (thus limiting proteinuria), glycaemic control and cardiovascular risk control, etc. [23, 24]. Therefore, there is a great urgency to prevent and treat diabetes by searching for novel molecular markers and drug targets. With the application of next-generation sequencing in RNA-seq technology [164], changes in the expression of LncRNAs have been found to be prevalent in the progress of DM and its associated complications. Because of their unique structure, LncRNAs are more stable in a disease, tissue, cell, or development phase specific manner [45]. Further, LncRNAs are easy to extract and detect with higher specificity compared with proteins. Detecting LncRNAs by RT-PCR and in-situ hybridization is also more specific and sensitive than detecting proteins by an antigen-antibody reaction [165]. There is also a greater possibility to screen for sensitive and specific molecular biomarkers from thousands of LncRNAs with regard to the relatively small number of miRNAs.
Recent substantial evidence indicates that the expression of many LncRNAs varies in normal/diabetic animal models [125, 154, 166] and healthy/diabetic patients’ clinical files [69, 72]. Therefore, LncRNAs have served as novel potential biomarkers for diagnosis and prognosis of DM. For example, GAS5 has been proposed as a prognostic biomarker [72], receiver operating characteristic (ROC) curve analysis revealed that GAS5 expression was a suitable candidate to distinguish between diabetic and non-diabetic samples (sensitivity 85.1%, specificity 67.3%). The area under curve (AUC) of ROC were 0.81 [OR (Odds Ratio) = 11.79, 95% confidence interval (CI): 3.97, 37.26), p < 0.001], respectively (Table 2). In conclusion, the use of LncRNAs outweighs the use of current DM biomarkers in many fields, and they could be promising molecular biomarkers and new targets for future drugs. LncRNAs, novel molecules that recently come to our attention, are no longer considered anomalies or noise. On the contrary, they have become a popular research subject, after siRNAs and miRNAs, owing to their important functions [150]. We believed that further investigation into the characteristic and function of LncRNAs will be of great value in preventing and treating diabetes.
Table 2: Application index of LncRNA in human DM
LncRNAs |
Diabetes Type |
Sample numbers (non-diabetic/diabetic) |
AUC |
Sensitivity |
Specificity |
OR (95%CI*) |
Ref. |
|---|---|---|---|---|---|---|---|
GAS5 |
Type 2 diabetes |
49/47 |
0.81 |
85.1% |
67.3% |
11.79 (3.97, 37.26) |
[83] |
CONCLUSIONS AND PROSPECTIVE
To summarize, increasing studies showed that LncRNAs have close connection with the progression of DM and its associated complications, such as pancreatic β-cells disorder, insulin resistance and epigenetic regulation, etc., by forming a complex regulatory network (Figure 2). LncRNAs, as novel regulatory biomarkers, participate in multiple steps of DM by modulating the expression of related-genes at the epigenetic, transcriptional and posttranscriptional levels. According to the role and features of DM-related LncRNAs, we may silence or active the LncRNAs in DM patients via an exogenous means (e.g., gene knock in, RNA interference and gene supplement) to achieve the prevention and treatment of DM. Although many important biological functions of lncRNAs have been discovered during the past three years, an overwhelming majority of LncRNAs have not been well-characterized, and there is still a long to go before we can identify, characterize, and elucidate the actual functions of LncRNAs in the pathogenesis of DM at the molecular level and use them in clinical practice and for therapeutic intervention. Currently, the research of LncRNAs faces two challenges. On one hand, there is need to sequentially validate the LncRNAs derived from the RNA sequencing in human tissue or cells to demonstrate whether they have function. In addition, there is a need to explore the functional LncRNAs to establish whether they are associated with specifically one or more diseases and by what molecular mechanisms. Although our current knowledge on LncRNAs is only the tip of an iceberg, novel methods and technology will eventually lift the mysterious veil covering LncRNAs, thereby providing a novel strategy for the prevention, early diagnosis and treatment of DM.
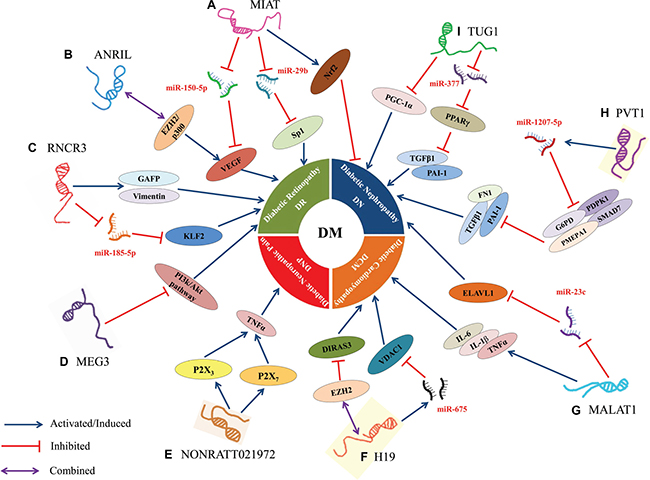
Figure 2: Dysregulation and functional roles of LncRNAs in DM. (A) MIAT regulated microvascular dysfunction via a ceRNA regulatory network MIAT/miR-150-5p/VEGF in diabetic retinas and endothelial cells; MIAT promoted cell apoptosis by absorbing miR-29b and subsequently relieving its repressive effect on the targeting genes Sp1 in high glucose stimulated rat retinal Müller cells (rMC-1); MIAT suppressed high glucose-induced tubular damage by promoting the expression of Nrf2 in human renal tubular epithelial cell line (HK-2). (B) ANRIL regulated VEGF expression prompted the development of DN by directly binding the p300 and EZH2 complex. (C) RNCR3 promoted DM-induced retinal neurodegeneration because of retinal cell apoptosis and Müller glial cell proliferation via the increased expression of glial reactivity-related genes including GFAP and vimentin; RNCR3 regulated endothelial cell function through RNCR3/miR-185-5p/KLF2 regulatory network to promote DM-induced retinal microvascular abnormalities. (D) MEG3 inhibited retinal endothelial cell proliferation, migration, and tube formation by suppressing the activation of PI3k/Akt signalling in the retinas of STZ-induced diabetic mice. (E) NONRATT021972 could interact with P2X3 or P2X7 to induce the release of inflammatory factors (TNF-α), thereby resulting in the excitability of DRG neurons to increase the risk of DNP. (F) H19 inhibited autophagy in cardiomyocytes exposed to high glucose by recruiting EZH2 and subsequently suppressing DIRAS3 expression; H19, a host gene of miR-675, also suppressed high glucose-induced cardiomyocyte apoptosis by reducing the expression of miR-675 target gene VDAC1. (G) MALAT1 attenuated the diabetes-induced myocardial inflammation in myocardial tissue of diabetic rats by reducing the levels of inflammatory markers such as TNF-α, IL-1β and IL-6; MALAT1 facilitated renal tubular epithelial pyroptosis by modulated miR-23c targeting of ELAVL1 in STZ-induced diabetic rats. (H) PVT1, a host gene of miR-1207-5p, suppressed the expression of miR-1207-5p targeting genes (G6PD, PMEPA1, PDPK1, and SMAD7), thereby increasing the expression of ECM-related genes FN1, PAI-1 and TGF-β1 in mesangial cells. (I) TUG1 regulated the metabolism of DN by epigenetic targeting of expression of PGC-1α in podocytes; TUG1 suppressed ECM accumulation via TUG1/miR-377/PPARγ signalling, and was also involved in reciprocally suppressing miR-377 and increasing the expression of targeted gene PPARγ to decrease the ECM accumulation in mesangial cell. In this diagram, the symbol “ ”represents activated or induced; the symbol “
”represents activated or induced; the symbol “ ”represents inhibited; and the symbol “
”represents inhibited; and the symbol “ ” represents combined or sponged.
” represents combined or sponged.
Highlights
LncRNAs have emerged as an essential regulator in almost every aspect of biology, Misexpression of LncRNAs alters the progression of diabetes mellitus(DM) and its associated complications, involving pancreatic β-cell disorder, insulin resistance, epigenetic regulation, etc. LncRNAs serve as a promising target for DM prevention, early diagnosis, and therapy.
Abbreviations
LncRNAs , long non-coding RNAs; ORF, open reading frame; DM, diabetes mellitus; HGP, human Genome Project; siRNA, small interfering RNA ; NCD, non-infective diseases; RMRP, RNA component of mitochondrial RNA processing endoribonuclease ; DDX5, DEAD-box helicase 5; RORγ, Retinoid-related orphan receptor gamma; EZH2, enhancer of zeste homolog-2; HDAC1, histone deacetylase 1; Dnmt3a, DNA-methyltransferase 3 alpha; PACER, p50-associated COX-2 extragenic RNA; COX2, cytochrome c oxidase subunit II; Linc-MD1, Long intergenic non-protein coding RNA, muscle differentiation 1; ceRNA, competing endogenous lncRNA; T1DM, type 1 diabetes mellitus; T2DM, type 2 diabetes mellitus; DN, diabetic nephropathy; DR, diabetic retinopathy; DCM, diabetic cardiomyopathy; DNP, diabetic neuropathic pain; GLIS3, GLIS family zinc finger 3; LincRNA-ROR, LincRNA-regulator of reprogramming; IGF2-AS, insulin like growth factor 2 antisense RNA; NOD, nonobese diabetic; NF-κB, nuclear factor-kappa B; INSR, insulin receptor; IRS-1/2, insulin receptor substrate 1/2; PI3K, phosphoinositide 3-Kinase; AKT, AKT Serine/Threonine kinase; GLUT4, glucose transporter 4; MEG3, maternally expressed gene 3; FoxO1, forkhead box O1; MALAT1, metastasis-associated lung adenocarcinoma transcript 1; SREBP-1c, sterol regulatory element binding transcription factor 1c; HYMAI, hydatidiform mole associated and imprinted; TNDM, transient neonatal diabetes mellitus; IGF2, insulin like growth factor 2 ; MeCP2, methyl-CpG binding protein 2; Arx, aristaless related homeobox; Tug1, taurine-upregulated 1; PGC-1α, PPARγ coactivator 1α; CKD, chronic kidney disease; DIRAS3, DIRAS family GTPase 3; ESRD, end-stage renal disease; ECM, extracellular matrix; PVT1, plasmacytoma variant translocation 1; FN1, fibronectin 1; COL4A1, collagen type IV alpha 1 chain; TGFB1, transforming growth factor beta 1 ; PAI-1, plasminogen activator inhibitor-1; G6PD, glucose-6-phosphate dehydrogenase; PMEPA1, prostate transmembrane protein, androgen induced 1; PDPK1, 3-phosphoinositide dependent protein kinase 1; SMAD7, SMAD family member 7; MIAT, myocardial infarction–associated transcript; AKI, acute kidney injury; NPDR, non-proliferative diabetic retinopathy; PDR, proliferative diabetic retinopathy; RNCR3, retinal non-coding RNA3; KLF2, kruppel like factor 2 ; RGC, retinal ganglion cell; LncRNA-Sox2OT, LncRNA SOX2 overlapping transcript; GFAP, glial fibrillary acidic protein; GAS5, growth arrest-specific 5; CARMEN, cardiac mesoderm enhancer-associated noncoding RNA; UCA1, urothelial carcinoma-associated 1; MI, myocardial infarction; VDAC1, voltage dependent anion channel 1; DAN, diabetic autonomic neuropathy; TNF-α, tumor necrosis factor α; IRS, insulin receptor substrate; SCG, superior cervical ganglia; HRV, heart rate variability; DRG, dorsal root ganglia; P2X7, purinergic receptor P2X 7; SAA3, serum amyloid A 3; HUVECs, human umbilical vein endothelial cells; ANRIL, antisense noncoding RNA in the INK4 locus; VEGF, vascular endothelial growth factor; 3-MA, 3-methyladenine; STZ, streptozotocin; ROC, receiver operating characteristic; AUC, area under curve; OR, Odds Ratio; CI, confidence interval.
ACKNOWLEDGMENTS
We thank the Research Center of Guilin Medical University, and the help of laboratory workers. This work was supported by the National Natural Science Foundation of China (grant no. 81260134, 81560148)
CONFLICTS OF INTEREST
None.
REFERENCES
1. Paulson H, Gonzalez-Alegre P. RNAi gets its prize. Lancet Neurol. 2006; 5:997–999.
2. Hirano T, Yoshikawa R, Harada H, Harada Y, Ishida A, Yamazaki T. Long noncoding RNA, CCDC26, controls myeloid leukemia cell growth through regulation of KIT expression. Mol Cancer. 2015; 14:90.
3. Yin Y, Yan P, Lu J, Song G, Zhu Y, Li Z, Zhao Y, Shen B, Huang X, Zhu H, Orkin SH, Shen X. Opposing Roles for the lncRNA Haunt and Its Genomic Locus in Regulating HOXA Gene Activation during Embryonic Stem Cell Differentiation. Cell Stem Cell. 2015; 16:504–516.
4. Gong Z, Zhang S, Zhang W, Huang H, Li Q, Deng H, Ma J, Zhou M, Xiang J, Wu M, Li X, Xiong W, Li X, et al. Long non-coding RNAs in cancer. Sci China Life Sci. 2012; 55:1120–1124.
5. Peng L, Yuan X, Jiang B, Tang Z, Li GC. LncRNAs: key players and novel insights into cervical cancer. Tumour Biol. 2016; 37:2779–2788.
6. Széll M, Danis J, Bata-Csörgő Z, Kemény L. PRINS, a primate-specific long non-coding RNA, plays a role in the keratinocyte stress response and psoriasis pathogenesis. Pflugers Arch. 2016; 468:935–943.
7. Yang Y, Cai Y, Wu G, Chen X, Liu Y, Wang X, Yu J, Li C, Chen X, Jose PA, Zhou L, Zeng C. Plasma long non-coding RNA, CoroMarker, a novel biomarker for diagnosis of coronary artery disease. Clin Sci (Lond). 2015; 129:675–685.
8. Wu G, Cai J, Han Y, Chen J, Huang ZP, Chen C, Cai Y, Huang H, Yang Y, Liu Y, Xu Z, He D, Zhang X, et al. LincRNA-p21 regulates neointima formation, vascular smooth muscle cell proliferation, apoptosis, and atherosclerosis by enhancing p53 activity. Circulation. 2014; 130:1452–1465.
9. Wang M, Wang S, Yao D, Yan Q, Lu W. A novel long non-coding RNA CYP4B1-PS1-001 regulates proliferation and fibrosis in diabetic nephropathy. Mol Cell Endocrinol. 2016; 426:136–145.
10. Kaur S, Mirza AH, Brorsson CA, Fløyel T, Størling J, Mortensen HB, Pociot F, Hvidoere International Study Group. The genetic and regulatory architecture of ERBB3-type 1 diabetes susceptibility locus. Mol Cell Endocrinol. 2016; 419:83–91.
11. Jin Y, Cui Z, Li X, Jin X, Peng J. Upregulation of long non-coding RNA PlncRNA-1 promotes proliferation and induces epithelial-mesenchymal transition in prostate cancer. Oncotarget. 2017; 8:26090–99. https://doi.org/10.18632/oncotarget.15318.
12. He X, Zheng Y, Zhang Y, Gan Y, Zhou Y, Liang H, Wu D, Ge W, Deng J, Xu X. Long non-coding RNA AK058003, as a precursor of miR-15a, interacts with HuR to inhibit the expression of γ-synuclein in hepatocellular carcinoma cells. Oncotarget. 2017; 8:9451–65. https://doi.org/10.18632/oncotarget.14276.
13. Wang Q, Yang L, Hu X, Jiang Y, Hu Y, Liu Z, Liu J, Wen T, Ma Y, An G, Feng G. Upregulated NNT-AS1, a long noncoding RNA, contributes to proliferation and migration of colorectal cancer cells in vitro and in vivo. Oncotarget. 2017; 8:3441–3453. https://doi.org/10.18632/oncotarget.13840.
14. Alegre-Díaz J, Herrington W, López-Cervantes M, Gnatiuc L, Ramirez R, Hill M, Baigent C, McCarthy MI, Lewington S, Collins R, Whitlock G, Tapia-Conyer R, Peto R, et al. Diabetes and Cause-Specific Mortality in Mexico City. N Engl J Med. 2016; 375:1961–1971.
15. Valk EJ, Bruijn JA, Bajema IM. Diabetic nephropathy in humans: pathologic diversity. Curr Opin Nephrol Hypertens. 2011; 20:285–289.
16. Daneshgari F, Moore C. Diabetic uropathy. Semin Nephrol. 2006; 26:182–185.
17. Mitrović-Perisić N, Antić S. [Risk factors for coronary heart disease and actual diagnostic criteria for diabetes mellitus]. [Article in Serbian]. Vojnosanit Pregl. 2009; 66:973–78.
18. Feng J, Xing W, Xie L. Regulatory Roles of MicroRNAs in Diabetes. Int J Mol Sci. 2016; 17:17.
19. Wang L, Kong L, Wu F, Bai Y, Burton R. Preventing chronic diseases in China. Lancet. 2005; 366:1821–1824.
20. King H, Aubert RE, Herman WH. Global burden of diabetes, 1995–2025: prevalence, numerical estimates, and projections. Diabetes Care. 1998; 21:1414–1431.
21. Cefalu WT, Buse JB, Tuomilehto J, Fleming GA, Ferrannini E, Gerstein HC, Bennett PH, Ramachandran A, Raz I, Rosenstock J, Kahn SE. Update and Next Steps for Real-World Translation of Interventions for Type 2 Diabetes Prevention: Reflections From a Diabetes Care Editors’ Expert Forum. Diabetes Care. 2016; 39:1186–1201.
22. Duan JG, Chen XY, Wang L, Lau A, Wong A, Thomas GN, Tomlinson B, Liu R, Chan JC, Leung TW, Mok V, Wong KS. Sex differences in epidemiology and risk factors of acute coronary syndrome in Chinese patients with type 2 diabetes: a long-term prospective cohort study. PLoS One. 2015; 10:e122031.
23. Paul P, Chakraborty A, Sarkar D, Langthasa M, Rahman M, Bari M, Singha RS, Malakar AK, Chakraborty S. Interplay between miRNAs and human diseases. J Cell Physiol. 2017; 9999:1–12.
24. Eriksson JG, Laine MK. Insulin therapy in the elderly with type 2 diabetes. Minerva Endocrinol. 2015; 40:283–295.
25. Lander ES. Initial impact of the sequencing of the human genome. Nature. 2011; 470:187–197.
26. Wright MW, Bruford EA. Naming ‘junk’: human non-protein coding RNA (ncRNA) gene nomenclature. Hum Genomics. 2011; 5:90–98.
27. Biscotti MA, Olmo E, Heslop-Harrison JS. Repetitive DNA in eukaryotic genomes. Chromosome Res. 2015; 23:415–420.
28. Nowak R. Mining treasures from ‘junk DNA’. Science. 1994; 263:608–610.
29. Wong GK, Passey DA, Huang Y, Yang Z, Yu J. Is “junk” DNA mostly intron DNA? Genome Res. 2000; 10:1672–78.
30. Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011; 10:38.
31. Brosnan CA, Voinnet O. The long and the short of noncoding RNAs. Curr Opin Cell Biol. 2009; 21:416–425.
32. Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012; 22:1775–1789.
33. Harrow J, Frankish A, Gonzalez JM, Tapanari E, Diekhans M, Kokocinski F, Aken BL, Barrell D, Zadissa A, Searle S, Barnes I, Bignell A, Boychenko V, et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 2012; 22:1760–1774.
34. Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012; 482:339–346.
35. Bazzini AA, Johnstone TG, Christiano R, Mackowiak SD, Obermayer B, Fleming ES, Vejnar CE, Lee MT, Rajewsky N, Walther TC, Giraldez AJ. Identification of small ORFs in vertebrates using ribosome footprinting and evolutionary conservation. EMBO J. 2014; 33:981–993.
36. Beaulieu YB, Kleinman CL, Landry-Voyer AM, Majewski J, Bachand F. Polyadenylation- dependent control of long noncoding RNA expression by the poly(A)-binding protein nuclear 1. PLoS Genet. 2012; 8:e1003078.
37. Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, et al. The transcriptional landscape of the mammalian genome. Science. 2005; 309:1559–1563.
38. Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermüller J, Hofacker IL, Bell I, Cheung E, Drenkow J, et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007; 316:1484–1488.
39. Sun J, Lin Y, Wu J. Long non-coding RNA expression profiling of mouse testis during postnatal development. PLoS One. 2013; 8:e75750.
40. Ríos-Barrera LD, Gutiérrez-Pérez I, Domínguez M, Riesgo-Escovar JR. acal is a long non-coding RNA in JNK signaling in epithelial shape changes during drosophila dorsal closure. PLoS Genet. 2015; 11:e1004927.
41. Quek XC, Thomson DW, Maag JL, Bartonicek N, Signal B, Clark MB, Gloss BS, Dinger ME. lncRNAdb v2.0: expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res. 2015; 43:D168–D173.
42. Ilott NE, Ponting CP. Predicting long non-coding RNAs using RNA sequencing. Methods. 2013; 63:50–59.
43. Guenzl PM, Barlow DP. Macro lncRNAs: a new layer of cis-regulatory information in the mammalian genome. RNA Biol. 2012; 9:731–741.
44. Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009; 136:629–641.
45. Gutschner T, Diederichs S. The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol. 2012; 9:703–719.
46. He Y, Meng XM, Huang C, Wu BM, Zhang L, Lv XW, Li J. Long noncoding RNAs: Novel insights into hepatocelluar carcinoma. Cancer Lett. 2014; 344:20–27.
47. Di Gesualdo F, Capaccioli S, Lulli M. A pathophysiological view of the long non-coding RNA world. Oncotarget. 2014; 5:10976–96. https://doi.org/10.18632/oncotarget.2770.
48. Orom UA, Derrien T, Beringer M, Gumireddy K, Gardini A, Bussotti G, Lai F, Zytnicki M, Notredame C, Huang Q, Guigo R, Shiekhattar R. Long noncoding RNAs with enhancer-like function in human cells. Cell. 2010; 143:46–58.
49. Huang W, Thomas B, Flynn RA, Gavzy SJ, Wu L, Kim SV, Hall JA, Miraldi ER, Ng CP, Rigo F, Meadows S, Montoya NR, Herrera NG, et al. DDX5 and its associated lncRNA Rmrp modulate TH17 cell effector functions. Nature. 2015; 528:517–522.
50. Saayman S, Ackley A, Turner AM, Famiglietti M, Bosque A, Clemson M, Planelles V, Morris KV. An HIV-Encoded Antisense Long Noncoding RNA Epigenetically Regulates Viral Transcription. Mol Ther. 2014; 22:1164–1175.
51. Krawczyk M, Emerson BM. p50-associated COX-2 extragenic RNA (PACER) activates COX-2 gene expression by occluding repressive NF-κB complexes. eLife. 2014; 3:e01776.
52. Dey BK, Pfeifer K, Dutta A. The H19 long noncoding RNA gives rise to microRNAs miR-675-3p and miR-675-5p to promote skeletal muscle differentiation and regeneration. Genes Dev. 2014; 28:491–501.
53. Cesana M, Cacchiarelli D, Legnini I, Santini T, Sthandier O, Chinappi M, Tramontano A, Bozzoni I. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell. 2011; 147:358–369.
54. Ma RC, Chan JC. Type 2 diabetes in East Asians: similarities and differences with populations in Europe and the United States. Ann N Y Acad Sci. 2013; 1281:64–91.
55. Ashcroft FM, Rorsman P. Diabetes mellitus and the beta cell: the last ten years. Cell. 2012; 148:1160–1171.
56. Mackay DJ, Coupe AM, Shield JP, Storr JN, Temple IK, Robinson DO. Relaxation of imprinted expression of ZAC and HYMAI in a patient with transient neonatal diabetes mellitus. Hum Genet. 2002; 110:139–144.
57. Cunnington MS, Santibanez Koref M, Mayosi BM, Burn J, Keavney B. Chromosome 9p21 SNPs Associated with Multiple Disease Phenotypes Correlate with ANRIL Expression. PLoS Genet. 2010; 6:e1000899.
58. Vausort M, Wagner DR, Devaux Y. Long noncoding RNAs in patients with acute myocardial infarction. Circ Res. 2014; 115:668–677.
59. Pasmant E, Sabbagh A, Vidaud M, Bièche I. ANRIL, a long, noncoding RNA, is an unexpected major hotspot in GWAS. FASEB J. 2011; 25:444–448.
60. Zeggini E, Weedon MN, Lindgren CM, Frayling TM, Elliott KS, Lango H, Timpson NJ, Perry JR, Rayner NW, Freathy RM, Barrett JC, Shields B, Morris AP, et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science. 2007; 316:1336–1341.
61. Scott LJ, Mohlke KL, Bonnycastle LL, Willer CJ, Li Y, Duren WL, Erdos MR, Stringham HM, Chines PS, Jackson AU, Prokunina-Olsson L, Ding CJ, Swift AJ, et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science. 2007; 316:1341–1345.
62. Broadbent HM, Peden JF, Lorkowski S, Goel A, Ongen H, Green F, Clarke R, Collins R, Franzosi MG, Tognoni G, Seedorf U, Rust S, Eriksson P, et al, and PROCARDIS consortium. Susceptibility to coronary artery disease and diabetes is encoded by distinct, tightly linked SNPs in the ANRIL locus on chromosome 9p. Hum Mol Genet. 2008; 17:806–814.
63. Todd JA, Walker NM, Cooper JD, Smyth DJ, Downes K, Plagnol V, Bailey R, Nejentsev S, Field SF, Payne F, Lowe CE, Szeszko JS, Hafler JP, et al., and Genetics of Type 1 Diabetes in Finland, and Wellcome Trust Case Control Consortium. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat Genet. 2007; 39:857–864.
64. Salonen JT, Uimari P, Aalto JM, Pirskanen M, Kaikkonen J, Todorova B, Hyppönen J, Korhonen VP, Asikainen J, Devine C, Tuomainen TP, Luedemann J, Nauck M, et al. Type 2 diabetes whole-genome association study in four populations: the DiaGen consortium. Am J Hum Genet. 2007; 81:338–345.
65. Wallace C, Smyth DJ, Maisuria-Armer M, Walker NM, Todd JA, Clayton DG. The imprinted DLK1-MEG3 gene region on chromosome 14q32.2 alters susceptibility to type 1 diabetes. Nat Genet. 2010; 42:68–71.
66. Murea M, Lu L, Ma L, Hicks PJ, Divers J, McDonough CW, Langefeld CD, Bowden DW, Freedman BI. Genome-wide association scan for survival on dialysis in African-Americans with type 2 diabetes. Am J Nephrol. 2011; 33:502–509.
67. Alvarez ML, Distefano JK. Functional characterization of the plasmacytoma variant translocation 1 gene (PVT1) in diabetic nephropathy. PLoS One. 2011; 6:e18671.
68. Hanson RL, Craig DW, Millis MP, Yeatts KA, Kobes S, Pearson JV, Lee AM, Knowler WC, Nelson RG, Wolford JK. Identification of PVT1 as a candidate gene for end-stage renal disease in type 2 diabetes using a pooling-based genome-wide single nucleotide polymorphism association study. Diabetes. 2007; 56:975–983.
69. de Gonzalo-Calvo D, Kenneweg F, Bang C, Toro R, van der Meer RW, Rijzewijk LJ, Smit JW, Lamb HJ, Llorente-Cortes V, Thum T. Circulating long-non coding RNAs as biomarkers of left ventricular diastolic function and remodelling in patients with well-controlled type 2 diabetes. Sci Rep. 2016; 6:37354.
70. Gao Y, Wu F, Zhou J, Yan L, Jurczak MJ, Lee HY, Yang L, Mueller M, Zhou XB, Dandolo L, Szendroedi J, Roden M, Flannery C, et al. The H19/let-7 double-negative feedback loop contributes to glucose metabolism in muscle cells. Nucleic Acids Res. 2014; 42:13799–13811.
71. Awata T, Yamashita H, Kurihara S, Morita-Ohkubo T, Miyashita Y, Katayama S, Mori K, Yoneya S, Kohda M, Okazaki Y, Maruyama T, Shimada A, Yasuda K, et al. A genome-wide association study for diabetic retinopathy in a Japanese population: potential association with a long intergenic non-coding RNA. PLoS One. 2014; 9:e111715.
72. Carter G, Miladinovic B, Patel AA, Deland L, Mastorides S, Patel NA. Circulating long noncoding RNA GAS5 levels are correlated to prevalence of type 2 diabetes mellitus. BBA Clin. 2015; 4:102–107.
73. Jang WI, Kim MS, Kang SH, Jo AJ, Kim YJ, Tchoe HJ, Park CM, Kim HJ, Choi JA, Choi HJ, Paik EK, Seo YS, Yoo HJ, et al. Association between metformin use and mortality in patients with type 2 diabetes mellitus and localized resectable pancreatic cancer: a nationwide population-based study in korea. Oncotarget. 2017; 8:9587–9596. https://doi.org/10.18632/oncotarget.14525.
74. Li X, Zeng L, Cao C, Lu C, Lian W, Han J, Zhang X, Zhang J, Tang T, Li M. Long noncoding RNA MALAT1 regulates renal tubular epithelial pyroptosis by modulated miR-23c targeting of ELAVL1 in diabetic nephropathy. Exp Cell Res. 2017; 350:327–335.
75. Liu C, Li CP, Wang JJ, Shan K, Liu X, Yan B. RNCR3 knockdown inhibits diabetes mellitus-induced retinal reactive gliosis. Biochem Biophys Res Commun. 2016; 479:198–203.
76. Akerman I, Tu Z, Beucher A, Rolando DM, Sauty-Colace C, Benazra M, Nakic N, Yang J, Wang H, Pasquali L, Moran I, Garcia-Hurtado J, Castro N, et al. Human Pancreatic beta Cell lncRNAs Control Cell-Specific Regulatory Networks. Cell Metab. 2017; 25:400–411.
77. Yan B, Yao J, Liu JY, Li XM, Wang XQ, Li YJ, Tao ZF, Song YC, Chen Q, Jiang Q. lncRNA-MIAT regulates microvascular dysfunction by functioning as a competing endogenous RNA. Circ Res. 2015; 116:1143–1156.
78. Horak M, Novak J, Bienertova-Vasku J. Muscle-specific microRNAs in skeletal muscle development. Dev Biol. 2016; 410:1–13.
79. Guay C, Jacovetti C, Nesca V, Motterle A, Tugay K, Regazzi R. Emerging roles of non-coding RNAs in pancreatic beta-cell function and dysfunction. Diabetes Obes Metab. 2012; 14:12–21.
80. Senée V, Chelala C, Duchatelet S, Feng D, Blanc H, Cossec JC, Charon C, Nicolino M, Boileau P, Cavener DR, Bougnères P, Taha D, Julier C. Mutations in GLIS3 are responsible for a rare syndrome with neonatal diabetes mellitus and congenital hypothyroidism. Nat Genet. 2006; 38:682–687.
81. Zou G, Liu T, Guo L, Huang Y, Feng Y, Huang Q, Duan T. miR-145 modulates lncRNA-ROR and Sox2 expression to maintain human amniotic epithelial stem cell pluripotency and beta islet-like cell differentiation efficiency. Gene. 2016; 591:48–57.
82. Mutskov V, Felsenfeld G. The human insulin gene is part of a large open chromatin domain specific for human islets. Proc Natl Acad Sci USA. 2009; 106:17419–17424.
83. Eizirik DL, Colli ML, Ortis F. The role of inflammation in insulitis and beta-cell loss in type 1 diabetes. Nat Rev Endocrinol. 2009; 5:219–226.
84. Motterle A, Gattesco S, Caille D, Meda P, Regazzi R. Involvement of long non-coding RNAs in beta cell failure at the onset of type 1 diabetes in NOD mice. Diabetologia. 2015; 58:1827–1835.
85. Stumvoll M, Goldstein BJ, van Haeften TW. Type 2 diabetes: principles of pathogenesis and therapy. Lancet. 2005; 365:1333–1346.
86. Fernandez-Twinn DS, Alfaradhi MZ, Martin-Gronert MS, Duque-Guimaraes DE, Piekarz A, Ferland-McCollough D, Bushell M, Ozanne SE. Downregulation of IRS-1 in adipose tissue of offspring of obese mice is programmed cell-autonomously through post-transcriptional mechanisms. Mol Metab. 2014; 3:325–333.
87. Xu G, Ji C, Song G, Zhao C, Shi C, Song L, Chen L, Yang L, Huang F, Pang L, Zhang N, Zhao Y, Guo X. MiR-26b modulates insulin sensitivity in adipocytes by interrupting the PTEN/PI3K/AKT pathway. Int J Obes. 2015; 39:1523–1530.
88. Chen YH, Heneidi S, Lee JM, Layman LC, Stepp DW, Gamboa GM, Chen BS, Chazenbalk G, Azziz R. miRNA-93 inhibits GLUT4 and is overexpressed in adipose tissue of polycystic ovary syndrome patients and women with insulin resistance. Diabetes. 2013; 62:2278–2286.
89. Yan C, Chen J, Chen N. Long noncoding RNA MALAT1 promotes hepatic steatosis and insulin resistance by increasing nuclear SREBP-1c protein stability. Sci Rep. 2016; 6:22640.
90. Zhu X, Wu YB, Zhou J, Kang DM. Upregulation of lncRNA MEG3 promotes hepatic insulin resistance via increasing FoxO1 expression. Biochem Biophys Res Commun. 2016; 469:319–325.
91. Haeusler RA, Kaestner KH, Accili D. FoxOs function synergistically to promote glucose production. J Biol Chem. 2010; 285:35245–35248.
92. Smith AC, Choufani S, Ferreira JC, Weksberg R. Growth regulation, imprinted genes, and chromosome 11p15.5. Pediatr Res. 2007; 61:43R–47R.
93. Kawaguchi T, Hirose T. Chromatin remodeling complexes in the assembly of long noncoding RNA-dependent nuclear bodies. Nucleus. 2015; 6:462–67.
94. Ip JY, Nakagawa S. Long non-coding RNAs in nuclear bodies. Dev Growth Differ. 2012; 54:44–54.
95. Wu SC, Kallin EM, Zhang Y. Role of H3K27 methylation in the regulation of lncRNA expression. Cell Res. 2010; 20:1109–1116.
96. Chuang JC, Jones PA. Epigenetics and microRNAs. Pediatr Res. 2007; 61:24R–29R.
97. Villeneuve LM, Natarajan R. The role of epigenetics in the pathology of diabetic complications. Am J Physiol Renal Physiol. 2010; 299:F14–F25.
98. Gardner RJ, Mackay DJ, Mungall AJ, Polychronakos C, Siebert R, Shield JP, Temple IK, Robinson DO. An imprinted locus associated with transient neonatal diabetes mellitus. Hum Mol Genet. 2000; 9:589–596.
99. Iglesias-Platas I, Martin-Trujillo A, Cirillo D, Court F, Guillaumet-Adkins A, Camprubi C, Bourc’his D, Hata K, Feil R, Tartaglia G, Arnaud P, Monk D. Characterization of novel paternal ncRNAs at the Plagl1 locus, including Hymai, predicted to interact with regulators of active chromatin. PLoS One. 2012; 7:e38907.
100. Su R, Wang C, Feng H, Lin L, Liu X, Wei Y, Yang H. Alteration in Expression and Methylation of IGF2/H19 in Placenta and Umbilical Cord Blood Are Associated with Macrosomia Exposed to Intrauterine Hyperglycemia. PLoS One. 2016; 11:e148399.
101. St-Pierre J, Hivert MF, Perron P, Poirier P, Guay SP, Brisson D, Bouchard L. IGF2 DNA methylation is a modulator of newborn’s fetal growth and development. Epigenetics. 2012; 7:1125–1132.
102. Briggs TA, Lokulo-Sodipe K, Chandler KE, Mackay DJ, Temple IK. Temple syndrome as a result of isolated hypomethylation of the 14q32 imprinted DLK1/MEG3 region. Am J Med Genet A. 2016; 170A:170–175.
103. Dhawan S, Georgia S, Tschen SI, Fan G, Bhushan A. Pancreatic beta cell identity is maintained by DNA methylation-mediated repression of Arx. Dev Cell. 2011; 20:419–429.
104. Maxwell SS, Pelka GJ, Tam PP, El-Osta A. Chromatin context and ncRNA highlight targets of MeCP2 in brain. RNA Biol. 2013; 10:1741–1757.
105. Long J, Badal SS, Ye Z, Wang Y, Ayanga BA, Galvan DL, Green NH, Chang BH, Overbeek PA, Danesh FR. Long noncoding RNA Tug1 regulates mitochondrial bioenergetics in diabetic nephropathy. J Clin Invest. 2016; 126:4205–4218.
106. Zhuo C, Jiang R, Lin X, Shao M. LncRNA H19 inhibits autophagy by epigenetically silencing of DIRAS3 in diabetic cardiomyopathy. Oncotarget. 2017; 8:1429–1437. https://doi.org/10.18632/oncotarget.13637.
107. Schernthaner G, Mogensen CE, Schernthaner GH. The effects of GLP-1 analogues, DPP-4 inhibitors and SGLT2 inhibitors on the renal system. Diab Vasc Dis Res. 2014; 11:306–323.
108. Zhang MH, Feng L, Zhu MM, Gu JF, Jiang J, Cheng XD, Ding SM, Wu C, Jia XB. The anti-inflammation effect of Moutan Cortex on advanced glycation end products-induced rat mesangial cells dysfunction and High-glucose-fat diet and streptozotocin-induced diabetic nephropathy rats. J Ethnopharmacol. 2014; 151:591–600.
109. Kanwar YS, Wada J, Sun L, Xie P, Wallner EI, Chen S, Chugh S, Danesh FR. Diabetic nephropathy: mechanisms of renal disease progression. Exp Biol Med (Maywood). 2008; 233:4–11.
110. Abe H. [Recent progress in understanding the molecular pathogenesis of diabetic nephropathy]. [Article in Japanese]. Rinsho Byori. 2011; 59:179–86.
111. Millis MP, Bowen D, Kingsley C, Watanabe RM, Wolford JK. Variants in the plasmacytoma variant translocation gene (PVT1) are associated with end-stage renal disease attributed to type 1 diabetes. Diabetes. 2007; 56:3027–3032.
112. Alvarez ML, Khosroheidari M, Eddy E, Kiefer J. Role of microRNA 1207-5P and its host gene, the long non-coding RNA Pvt1, as mediators of extracellular matrix accumulation in the kidney: implications for diabetic nephropathy. PLoS One. 2013; 8:e77468.
113. Duan LJ, Ding M, Hou LJ, Cui YT, Li CJ, Yu DM. Long noncoding RNA TUG1 alleviates extracellular matrix accumulation via mediating microRNA-377 targeting of PPARgamma in diabetic nephropathy. Biochem Biophys Res Commun. 2017; 484:598–604.
114. Yi H, Peng R, Zhang LY, Sun Y, Peng HM, Liu HD, Yu LJ, Li AL, Zhang YJ, Jiang WH, Zhang Z. LincRNA-Gm4419 knockdown ameliorates NF-kappaB/NLRP3 inflammasome-mediated inflammation in diabetic nephropathy. Cell Death Dis. 2017; 8:e2583.
115. Zhou L, Xu DY, Sha WG, Shen L, Lu GY, Yin X. Long non-coding MIAT mediates high glucose-induced renal tubular epithelial injury. Biochem Biophys Res Commun. 2015; 468:726–732.
116. Yau JW, Rogers SL, Kawasaki R, Lamoureux EL, Kowalski JW, Bek T, Chen SJ, Dekker JM, Fletcher A, Grauslund J, Haffner S, Hamman RF, Ikram MK, et al. Global prevalence and major risk factors of diabetic retinopathy. Diabetes Care. 2012; 35:556–564.
117. Campochiaro PA. Ocular neovascularization. J Mol Med (Berl). 2013; 91:311–321.
118. Brownlee M. Biochemistry and molecular cell biology of diabetic complications. Nature. 2001; 414:813–820.
119. Schalkwijk CG, Stehouwer CD. Vascular complications in diabetes mellitus: the role of endothelial dysfunction. Clin Sci (Lond). 2005; 109:143–159.
120. Michalik KM, You X, Manavski Y, Doddaballapur A, Zörnig M, Braun T, John D, Ponomareva Y, Chen W, Uchida S, Boon RA, Dimmeler S. Long noncoding RNA MALAT1 regulates endothelial cell function and vessel growth. Circ Res. 2014; 114:1389–1397.
121. Curtis TM, Gardiner TA, Stitt AW. Microvascular lesions of diabetic retinopathy: clues towards understanding pathogenesis? Eye (Lond). 2009; 23:1496–1508.
122. Wu L, Fernandez-Loaiza P, Sauma J, Hernandez-Bogantes E, Masis M. Classification of diabetic retinopathy and diabetic macular edema. World J Diabetes. 2013; 4:290–294.
123. Ting DS, Tan KA, Phua V, Tan GS, Wong CW, Wong TY. Biomarkers of Diabetic Retinopathy. Curr Diab Rep. 2016; 16:125.
124. Dhoot DS, Avery RL. Vascular Endothelial Growth Factor Inhibitors for Diabetic Retinopathy. Curr Diab Rep. 2016; 16:122.
125. Yan B, Tao ZF, Li XM, Zhang H, Yao J, Jiang Q. Aberrant expression of long noncoding RNAs in early diabetic retinopathy. Invest Ophthalmol Vis Sci. 2014; 55:941–951.
126. Liu JY, Yao J, Li XM, Song YC, Wang XQ, Li YJ, Yan B, Jiang Q. Pathogenic role of lncRNA-MALAT1 in endothelial cell dysfunction in diabetes mellitus. Cell Death Dis. 2014; 5:e1506.
127. Jae N, Dimmeler S. Long noncoding RNAs in diabetic retinopathy. Circ Res. 2015; 116:1104–1106.
128. Shan K, Li CP, Liu C, Liu X, Yan B. RNCR3: A regulator of diabetes mellitus-related retinal microvascular dysfunction. Biochem Biophys Res Commun. 2017; 482:777–783.
129. Shan K, Jiang Q, Wang XQ, Wang YN, Yang H, Yao MD, Liu C, Li XM, Yao J, Liu B, Zhang YY, Yong J, Yan B. Role of long non-coding RNA-RNCR3 in atherosclerosis-related vascular dysfunction. Cell Death Dis. 2016; 7:e2248.
130. Zhang J, Chen M, Chen J, Lin S, Cai D, Chen C, Chen Z. Long noncoding RNA MIAT acts as a biomarker in diabetic retinopathy by absorbing of miR-29b and regulating cell apoptosis. Biosci Rep. 2017; 37.
131. Qiu GZ, Tian W, Fu HT, Li CP, Liu B. Long noncoding RNA-MEG3 is involved in diabetes mellitus-related microvascular dysfunction. Biochem Biophys Res Commun. 2016; 471:135–141.
132. Kern TS, Barber AJ. Retinal ganglion cells in diabetes. J Physiol. 2008; 586:4401–4408.
133. Li CP, Wang SH, Wang WQ, Song SG, Liu XM. Long Noncoding RNA-Sox2OT Knockdown Alleviates Diabetes Mellitus-Induced Retinal Ganglion Cell (RGC) injury. Cell Mol Neurobiol. 2017; 37:361–369.
134. Bahtiyar G, Gutterman D, Lebovitz H. Heart Failure: a Major Cardiovascular Complication of Diabetes Mellitus. Curr Diab Rep. 2016; 16:116.
135. Aneja A, Tang WH, Bansilal S, Garcia MJ, Farkouh ME. Diabetic cardiomyopathy: insights into pathogenesis, diagnostic challenges, and therapeutic options. Am J Med. 2008; 121:748–757.
136. Falcao-Pires I, Leite-Moreira AF. Diabetic cardiomyopathy: understanding the molecular and cellular basis to progress in diagnosis and treatment. Heart Fail Rev. 2012; 17:325–344.
137. Bugger H, Abel ED. Molecular mechanisms of diabetic cardiomyopathy. Diabetologia. 2014; 57:660–671.
138. Pang L, Hu J, Zhang G, Li X, Zhang X, Yu F, Lan Y, Xu J, Pang B, Han D, Xiao Y, Li X. Dysregulated long intergenic non-coding RNA modules contribute to heart failure. Oncotarget. 2016; 7:59676–59690. https://doi.org/10.18632/oncotarget.10834.
139. Wang YN, Shan K, Yao MD, Yao J, Wang JJ, Li X, Liu B, Zhang YY, Ji Y, Jiang Q, Yan B. Long Noncoding RNA-GAS5: A Novel Regulator of Hypertension-Induced Vascular Remodeling. Hypertension. 2016; 68:736–748.
140. Ounzain S, Micheletti R, Arnan C, Plaisance I, Cecchi D, Schroen B, Reverter F, Alexanian M, Gonzales C, Ng SY, Bussotti G, Pezzuto I, Notredame C, et al. CARMEN, a human super enhancer-associated long noncoding RNA controlling cardiac specification, differentiation and homeostasis. J Mol Cell Cardiol. 2015; 89:98–112.
141. Liu Y, Zhou D, Li G, Ming X, Tu Y, Tian J, Lu H, Yu B. Long non coding RNA-UCA1 contributes to cardiomyocyte apoptosis by suppression of p27 expression. Cell Physiol Biochem. 2015; 35:1986–1998.
142. Zhang M, Gu H, Chen J, Zhou X. Involvement of long noncoding RNA MALAT1 in the pathogenesis of diabetic cardiomyopathy. Int J Cardiol. 2016; 202:753–755.
143. Zhang M, Gu H, Xu W, Zhou X. Down-regulation of lncRNA MALAT1 reduces cardiomyocyte apoptosis and improves left ventricular function in diabetic rats. Int J Cardiol. 2016; 203:214–216.
144. Li X, Wang H, Yao B, Xu W, Chen J, Zhou X. lncRNA H19/miR-675 axis regulates cardiomyocyte apoptosis by targeting VDAC1 in diabetic cardiomyopathy. Sci Rep. 2016; 6:36340.
145. Gabory A, Jammes H, Dandolo L. The H19 locus: role of an imprinted non-coding RNA in growth and development. BioEssays. 2010; 32:473–480.
146. Imig J, Brunschweiger A, Brümmer A, Guennewig B, Mittal N, Kishore S, Tsikrika P, Gerber AP, Zavolan M, Hall J. miR-CLIP capture of a miRNA targetome uncovers a lincRNA H19-miR-106a interaction. Nat Chem Biol. 2015; 11:107–114.
147. Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L, Lu L, Liu C, Yi JS, Zhang H, Min W, Bennett AM, Gregory RI, et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell. 2013; 52:101–112.
148. Keniry A, Oxley D, Monnier P, Kyba M, Dandolo L, Smits G, Reik W. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat Cell Biol. 2012; 14:659–665.
149. Callaghan BC, Cheng HT, Stables CL, Smith AL, Feldman EL. Diabetic neuropathy: clinical manifestations and current treatments. Lancet Neurol. 2012; 11:521–534.
150. Davies M, Brophy S, Williams R, Taylor A. The prevalence, severity, and impact of painful diabetic peripheral neuropathy in type 2 diabetes. Diabetes Care. 2006; 29:1518–1522.
151. Tesfaye S, Selvarajah D. Advances in the epidemiology, pathogenesis and management of diabetic peripheral neuropathy. Diabetes Metab Res Rev. 2012; 28:8–14.
152. Morales-Vidal S, Morgan C, McCoyd M, Hornik A. Diabetic peripheral neuropathy and the management of diabetic peripheral neuropathic pain. Postgrad Med. 2012; 124:145–153.
153. Song M, Zou L, Peng L, Liu S, Wu B, Yi Z, Gao Y, Zhang C, Xu H, Xu Y, Tang M, Wang S, Xue Y, et al. LncRNA NONRATT021972 siRNA normalized the dysfunction of hepatic glucokinase through AKT signaling in T2DM rats. Endocr Res. 2017; 42:180–90.
154. Li G, Sheng X, Xu Y, Jiang H, Zheng C, Guo J, Sun S, Yi Z, Qin S, Liu S, Gao Y, Zhang C, Xu H, et al. Co-expression changes of lncRNAs and mRNAs in the cervical sympathetic ganglia in diabetic cardiac autonomic neuropathic rats. J Neurosci Res. 2017; 95:1690–99.
155. Xu H, Liu C, Rao S, He L, Zhang T, Sun S, Wu B, Zou L, Wang S, Xue Y, Jia T, Zhao S, Li G, et al. LncRNA NONRATT021972 siRNA rescued decreased heart rate variability in diabetic rats in superior cervical ganglia. Auton Neurosci. 2016; 201:1–7.
156. Liu S, Zou L, Xie J, Xie W, Wen S, Xie Q, Gao Y, Li G, Zhang C, Xu C, Xu H, Wu B, Lv Q, et al. LncRNA NONRATT021972 siRNA regulates neuropathic pain behaviors in type 2 diabetic rats through the P2X7 receptor in dorsal root ganglia. Mol Brain. 2016; 9:44.
157. Peng H, Zou L, Xie J, Wu H, Wu B, Zhu G, Lv Q, Zhang X, Liu S, Li G, Xu H, Gao Y, Xu C, et al. lncRNA NONRATT021972 siRNA Decreases Diabetic Neuropathic Pain Mediated by the P2X3 Receptor in Dorsal Root Ganglia. Mol Neurobiol. 2017; 54:511–523.
158. Wu B, Zhang C, Zou L, Ma Y, Huang K, Lv Q, Zhang X, Wang S, Xue Y, Yi Z, Jia T, Zhao S, Liu S, et al. LncRNA uc.48+ siRNA improved diabetic sympathetic neuropathy in type 2 diabetic rats mediated by P2X7 receptor in SCG. Auton Neurosci. 2016; 197:14–18.
159. Reddy MA, Chen Z, Park JT, Wang M, Lanting L, Zhang Q, Bhatt K, Leung A, Wu X, Putta S, Sætrom P, Devaraj S, Natarajan R. Regulation of inflammatory phenotype in macrophages by a diabetes-induced long noncoding RNA. Diabetes. 2014; 63:4249–4261.
160. Puthanveetil P, Chen S, Feng B, Gautam A, Chakrabarti S. Long non-coding RNA MALAT1 regulates hyperglycaemia induced inflammatory process in the endothelial cells. J Cell Mol Med. 2015; 19:1418–1425.
161. Zhang B, Wang D, Ji TF, Shi L, Yu JL. Overexpression of lncRNA ANRIL up-regulates VEGF expression and promotes angiogenesis of diabetes mellitus combined with cerebral infarction by activating NF-kappaB signaling pathway in a rat model. Oncotarget. 2017; 8:17347–59. https://doi.org/10.18632/oncotarget.
162. Li Z, Hao S, Yin H, Gao J, Yang Z. Autophagy ameliorates cognitive impairment through activation of PVT1 and apoptosis in diabetes mice. Behav Brain Res. 2016; 305:265–277.
163. Sun C, Xue L, Zhu Z, Zhang F, Yang R, Yuan X, Jia Z, Liu Q. Insights from lncRNAs Profiling of MIN6 Beta Cells Undergoing Inflammation. Mediators Inflamm. 2016; 2016:9275106.
164. Lee JH, Gao C, Peng G, Greer C, Ren S, Wang Y, Xiao X. Analysis of transcriptome complexity through RNA sequencing in normal and failing murine hearts. Circ Res. 2011; 109:1332–1341.
165. Wang Y, Mo Y, Gong Z, Yang X, Yang M, Zhang S, Xiong F, Xiang B, Zhou M, Liao Q, Zhang W, Li X, Li X, et al. Circular RNAs in human cancer. Mol Cancer. 2017; 16:25.
166. Chen S, Dong C, Qian X, Huang S, Feng Y, Ye X, Miao H, You Q, Lu Y, Ding D. Microarray analysis of long noncoding RNA expression patterns in diabetic nephropathy. J Diabetes Complications. 2017; 31:569–576.

