INTRODUCTION
Coronary artery disease (CAD) is the most common form of heart disease. As a major cause of morbidity, mortality, and disability worldwide, it imposes a tremendous social and economic burden on society [1]. CAD is characterized by atherosclerosis, a process of cumulative deposition of low-density lipoprotein (LDL) cholesterol in the arteries supplying blood to the heart that eventually leads to impaired or absent blood supply and myocardial infarction [2]. CAD is a complex, multifactorial disease. Among its risk factors are smoking, advanced age, diabetes mellitus, high blood pressure, high-fat diet, obesity, infectious agents, increased total and LDL cholesterol in plasma, increased plasma triglycerides, and decreased plasma high-density lipoprotein (HDL) cholesterol [3, 4]. In recent years, considerable effort has been devoted to identifying genes and inherited DNA sequence variants that contribute to CAD risk [5, 6]. In addition, gender differences in the incidence of CAD have caught the interest of many clinical researchers. In the current study, we focused on several CAD-related genes: CDKN2B-AS1, CYP1A1, PSRC1, APOC1, SH2B3, and SMARCA4, previously addressed through genome-wide association studies (GWAS) and large-scale replication studies [7–9].
The SH2B3 gene, located on chromosome 12 (12q24), is a member of the SH2B adaptor family. The SH2B3 protein has been associated with negative regulation of cytokine signaling. However, the specific functions of SH2B3 remain unclear. Variants in this region have been shown to be associated with various other traits including blood pressure [10, 11], blood lipids [12], platelet count [13], and type-1 diabetes mellitus [14]. Correlations between the SH2B3 gene and CAD susceptibility have been addressed by GWAS [15–17]. The SMARCA4 gene is located on chromosome 19 (19q13) and is a member of the SWI/SNF family of proteins. Recent reports suggested that mutations in this gene cause small cell carcinoma of the ovary of hypercalcemic type [18], lung cancer [19], and other diseases [20, 21]. Combining GWAS data and case-control studies, we reported the association of SMARCA4 with CAD risk [22–24]. To date, however, no studies investigated the possible correlations between SH2B3 and SMARCA4 genes and CAD susceptibility in the Chinese Han population.
To test the hypothesis that SH2B3 and SMARCA4 gene polymorphisms contribute differentially to coronary vascular pathology in men and women, we conducted a case-control study to examine the associations between single nucleotide polymorphisms (SNPs) in those genes and CAD risk in Chinese Han individuals.
RESULTS
The demographic characteristics of the study population, including gender and age, are summarized in Table 1. The study included 456 CAD cases (291 males and 165 females) and 685 controls (385 males and 300 females). The mean age of males and females in the case group was 59.56 ± 12.18 years and 64.01 ± 10.74 years, respectively. In the control group, the mean age of males and females was 47.55 ± 10.66 years and 49.93 ± 7.74 years, respectively. There was a significant difference between genders’ ages in the case group (p = 0.000), but not in the control group (p = 0.063).
Table 1: Characteristics of the subjects
Characteristics |
Cases (n = 456) |
p |
Controls (n = 685) |
p |
||
|---|---|---|---|---|---|---|
Male |
Female |
Male |
Female |
|||
Count |
291 |
165 |
385 |
300 |
||
Age (mean ± SD years) |
59.56±12.18 |
64.01±10.74 |
0.00* |
47.55±10.66 |
49.93±7.74 |
0.063 |
*p < 0.05 indicates statistical significance
Basic information of candidate SNPs, such as chromosomal position, gene, allele, HWE test results, as well as minor allele frequency (MAF) by gender are shown in Table 2. Rs11879293 and rs12232780 in SMARCA4 were associated with CAD risk (p = 0.036 and p = 0.001, respectively) in the male population only. In this group, rs2072382 was also significantly associated with CAD but was excluded due to significant deviation from HWE (p = 0.03). Meanwhile, an almost significant association was observed between SH2B3 and SMARCA4 genes and CAD risk in the women population.
Table 2: Sex-specific association of individual SNPs with CAD
SNP |
Chr |
Allele A#/B |
Gene |
Minor allele frequency, % |
|||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Men |
Women |
||||||||||||||||
HWE |
Cases |
Controls |
OR |
95% |
CI |
p |
HWE |
Case |
Control |
OR |
95% |
CI |
p |
||||
rs3742003 |
12 |
G/A |
SH2B3 |
0.75 |
0.12 |
0.09 |
1.36 |
0.95 |
1.95 |
0.090 |
1.00 |
0.09 |
0.09 |
0.97 |
0.61 |
1.55 |
0.903 |
rs12580300 |
12 |
A/G |
SH2B3 |
0.46 |
0.41 |
0.42 |
0.98 |
0.78 |
1.22 |
0.831 |
0.72 |
0.37 |
0.42 |
0.81 |
0.61 |
1.07 |
0.134 |
rs7309325 |
12 |
G/T |
SH2B3 |
0.50 |
0.11 |
0.08 |
1.38 |
0.96 |
1.98 |
0.082 |
0.15 |
0.09 |
0.09 |
0.98 |
0.61 |
1.59 |
0.946 |
rs2078863 |
12 |
T/C |
SH2B3 |
0.76 |
0.53 |
0.50 |
1.11 |
0.90 |
1.38 |
0.324 |
0.91 |
0.55 |
0.49 |
1.25 |
0.95 |
1.63 |
0.107 |
rs7296313 |
12 |
T/C |
SH2B3 |
0.34 |
0.11 |
0.09 |
1.29 |
0.90 |
1.85 |
0.172 |
0.49 |
0.09 |
0.09 |
0.99 |
0.62 |
1.58 |
0.969 |
rs11879293 |
19 |
A/G |
SMARCA4 |
0.70 |
0.22 |
0.27 |
0.76 |
0.59 |
0.98 |
0.036* |
0.17 |
0.27 |
0.26 |
1.08 |
0.79 |
1.46 |
0.634 |
rs12232780 |
19 |
A/G |
SMARCA4 |
0.78 |
0.18 |
0.25 |
0.64 |
0.49 |
0.84 |
0.001* |
0.14 |
0.23 |
0.19 |
1.23 |
0.88 |
1.70 |
0.220 |
rs2072382 |
19 |
T/C |
SMARCA4 |
0.03ψ |
0.34 |
0.28 |
1.30 |
1.03 |
1.65 |
0.026* |
0.19 |
0.31 |
0.28 |
1.15 |
0.86 |
1.55 |
0.343 |
rs1529729 |
19 |
C/T |
SMARCA4 |
0.66 |
0.22 |
0.22 |
0.98 |
0.76 |
1.28 |
0.904 |
0.14 |
0.22 |
0.23 |
0.93 |
0.67 |
1.28 |
0.638 |
rs1122608 |
19 |
T/G |
SMARCA4 |
0.41 |
0.08 |
0.10 |
0.74 |
0.51 |
1.08 |
0.120 |
1.00 |
0.09 |
0.08 |
1.22 |
0.76 |
1.96 |
0.411 |
SNP = single nucleotide polymorphism; CAD = coronary artery disease; OR = odds ratio;
95% CI = 95% confidence interval; Chr = chromosome; HWE = Hardy–Weinberg equilibrium
#Minor allele
p values were calculated from two-sided chi-square tests for either allele frequency
ψSite with HWE P < 0.05 is excluded in the men population
* p < 0.05 indicates statistical significance
The association between each SNP and CAD risk was further assessed using unconditional logistic regression analysis including five genetic models: codominant, dominant, recessive, overdominant, and additive (Table 3). The minor allele “A” of rs12580300 in SH2B3 was associated with CAD risk in the female population under the codominant (p = 0.043), dominant (AA + AG vs. GG: OR = 0.59, 95% CI: 0.35-0.97; p = 0.039), and additive (OR = 0.63, 95% CI: 0.44-0.91; p = 0.012) models. In this same population, the allele “T” in SH2B3 rs2078863 was associated with decreased odds of CAD risk in the codominant (p = 0.022), recessive (CC vs. TT + TC: OR = 0.44, 95% CI: 0.24-0.82; p = 0.007), and additive (OR = 0.64, 95% CI: 0.44-0.91; p = 0.012) models. Interestingly, in the SH2B3 gene, rs3742003 in the codominant and overdominant models, and rs7309325 in the overdominant model, were associated with increased CAD risk in the male population. Meanwhile, the SMARCA4 SNPs rs11879293 in the codominant, dominant, and additive models, and rs12232780 in the co-dominant, dominant, recessive, overdominant, and additive models, were associated with decreased CAD risk in this population
Table 3: Genotypes of SH2B3 and SMARCA4 polymorphisms and CAD risk associations (age-adjusted)
Men |
Women |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|
SNP |
Model |
Genotype |
Control |
Case |
OR (95% CI) |
p |
Control |
Case |
OR (95% CI) |
p |
rs3742003 |
Codominant |
A/A |
319 (83.1%) |
224 (77%) |
1 |
0.076 |
246 (82%) |
136 (82.4%) |
1 |
0.960 |
G/A |
63 (16.4%) |
67 (23%) |
1.57 (1.01-2.42) |
52 (17.3%) |
28 (17%) |
0.99 (0.53-1.85) |
||||
G/G |
2 (0.5%) |
0 (0%) |
0.00 (0.00-NA) |
2 (0.7%) |
1 (0.6%) |
0.59 (0.02-20.73) |
||||
Dominant |
A/A |
319 (83.1%) |
224 (77%) |
1 |
0.054 |
246 (82%) |
136 (82.4%) |
1 |
0.950 |
|
G/A-G/G |
65 (16.9%) |
67 (23%) |
1.53 (0.99-2.36) |
54 (18%) |
29 (17.6%) |
0.98 (0.53-1.81) |
||||
Recessive |
A/A-G/A |
382 (99.5%) |
291 (100%) |
1 |
0.300 |
298 (99.3%) |
164 (99.4%) |
1 |
0.770 |
|
G/G |
2 (0.5%) |
0 (0%) |
0.00 (0.00-NA) |
2 (0.7%) |
1 (0.6%) |
0.59 (0.02-20.72) |
||||
Overdominant |
A/A-G/G |
321 (83.6%) |
224 (77%) |
1 |
0.041* |
248 (82.7%) |
137 (83%) |
1 |
0.990 |
|
G/A |
63 (16.4%) |
67 (23%) |
1.57 (1.02-2.43) |
52 (17.3%) |
28 (17%) |
0.99 (0.53-1.85) |
||||
Log-additive |
--- |
--- |
--- |
1.47 (0.96-2.23) |
0.074 |
--- |
--- |
0.97 (0.53-1.75) |
0.910 |
|
rs12580300 |
Codominant |
G/G |
134 (35%) |
98 (33.7%) |
1 |
0.820 |
99 (33%) |
63 (38.2%) |
1 |
0.043* |
G/A |
179 (46.7%) |
147 (50.5%) |
1.08 (0.73-1.59) |
150 (50%) |
82 (49.7%) |
0.66 (0.39-1.13) |
||||
A/A |
70 (18.3%) |
46 (15.8%) |
0.93 (0.56-1.55) |
51 (17%) |
20 (12.1%) |
0.39 (0.18-0.85) |
||||
Dominant |
G/G |
134 (35%) |
98 (33.7%) |
1 |
0.840 |
99 (33%) |
63 (38.2%) |
1 |
0.039* |
|
G/A-A/A |
249 (65%) |
193 (66.3%) |
1.04 (0.72-1.49) |
201 (67%) |
102 (61.8%) |
0.59 (0.35-0.97) |
||||
Recessive |
G/G-G/A |
313 (81.7%) |
245 (84.2%) |
1 |
0.610 |
249 (83%) |
145 (87.9%) |
1 |
0.046 |
|
A/A |
70 (18.3%) |
46 (15.8%) |
0.89 (0.56-1.41) |
51 (17%) |
20 (12.1%) |
0.50 (0.24-1.00) |
||||
Overdominant |
G/G-A/A |
204 (53.3%) |
144 (49.5%) |
1 |
0.570 |
150 (50%) |
83 (50.3%) |
1 |
0.590 |
|
G/A |
179 (46.7%) |
147 (50.5%) |
1.11 (0.78-1.56) |
150 (50%) |
82 (49.7%) |
0.87 (0.54-1.42) |
||||
Log-additive |
--- |
--- |
--- |
0.98 (0.77-1.26) |
0.890 |
--- |
--- |
0.63 (0.44-0.91) |
0.012* |
|
rs7309325 |
Codominant |
T/T |
320 (83.5%) |
226 (77.7%) |
1 |
0.100 |
248 (82.7%) |
137 (83.5%) |
1 |
0.880 |
G/T |
62 (16.2%) |
65 (22.3%) |
1.55 (1.00-2.40) |
52 (17.3%) |
26 (15.8%) |
1.02 (0.54-1.92) |
||||
G/G |
1 (0.3%) |
0 (0%) |
0.00 (0.00-NA) |
0 (0%) |
1 (0.6%) |
NA (0.00-NA) |
||||
Dominant |
T/T |
320 (83.5%) |
226 (77.7%) |
1 |
0.055 |
248 (82.7%) |
137 (83.5%) |
1 |
0.930 |
|
G/T-G/G |
63 (16.4%) |
65 (22.3%) |
1.53 (0.99-2.37) |
52 (17.3%) |
27 (16.5%) |
1.03 (0.55-1.93) |
||||
Recessive |
T/T-G/T |
382 (99.7%) |
291 (100%) |
1 |
0.420 |
300 (100%) |
163 (99.4%) |
1 |
0.620 |
|
G/G |
1 (0.3%) |
0 (0%) |
0.00 (0.00-NA) |
0 (0%) |
1 (0.6%) |
NA (0.00-NA) |
||||
Overdominant |
T/T-G/G |
321 (83.8%) |
226 (77.7%) |
1 |
0.047* |
248 (82.7%) |
138 (84.2%) |
1 |
0.960 |
|
G/T |
62 (16.2%) |
65 (22.3%) |
1.56 (1.01-2.41) |
52 (17.3%) |
26 (15.8%) |
1.02 (0.54-1.91) |
||||
Log-additive |
--- |
--- |
--- |
1.49 (0.97-2.29) |
0.066 |
--- |
--- |
1.04 (0.56-1.94) |
0.900 |
|
T/T |
94 (24.4%) |
78 (26.8%) |
1 |
72 (24%) |
44 (26.7%) |
1 |
0.022* |
|||
Codominant |
C/T |
196 (50.9%) |
150 (51.5%) |
0.80 (0.53-1.21) |
0.480 |
152 (50.7%) |
93 (56.4%) |
0.82 (0.46-1.47) |
||
C/C |
95 (24.7%) |
63 (21.6%) |
0.76 (0.46-1.25) |
76 (25.3%) |
28 (17%) |
0.39 (0.19-0.81) |
||||
Dominant |
T/T |
94 (24.4%) |
78 (26.8%) |
1 |
0.240 |
72 (24%) |
44 (26.7%) |
1 |
0.140 |
|
C/T-C/C |
291 (75.6%) |
213 (73.2%) |
0.79 (0.53-1.17) |
228 (76%) |
121 (73.3%) |
0.66 (0.38-1.15) |
||||
Recessive |
T/T-C/T |
290 (75.3%) |
228 (78.3%) |
1 |
0.560 |
224 (74.7%) |
137 (83%) |
1 |
0.007* |
|
rs2078863 |
C/C |
95 (24.7%) |
63 (21.6%) |
0.89 (0.59-1.34) |
76 (25.3%) |
28 (17%) |
0.44 (0.24-0.82) |
|||
Overdominant |
T/T-C/C |
189 (49.1%) |
141 (48.5%) |
1 |
0.580 |
148 (49.3%) |
72 (43.6%) |
1 |
0.330 |
|
C/T |
196 (50.9%) |
150 (51.5%) |
0.91 (0.64-1.28) |
152 (50.7%) |
93 (56.4%) |
1.27 (0.78-2.06) |
||||
Log-additive |
--- |
--- |
--- |
0.87 (0.68-1.12) |
0.270 |
--- |
--- |
0.64 (0.44-0.91) |
0.012* |
|
rs11879293 |
G/G |
205 (53.5%) |
176 (60.5%) |
1 |
0.018* |
160 (53.3%) |
86 (52.1%) |
1 |
0.760 |
|
Codominant |
G/A |
148 (38.6%) |
101 (34.7%) |
0.68 (0.47-0.99) |
125 (41.7%) |
68 (41.2%) |
0.83 (0.50-1.37) |
|||
A/A |
30 (7.8%) |
14 (4.8%) |
0.41 (0.19-0.89) |
15 (5%) |
11 (6.7%) |
0.93 (0.35-2.46) |
||||
Dominant |
G/G |
205 (53.5%) |
176 (60.5%) |
1 |
0.012* |
160 (53.3%) |
86 (52.1%) |
1 |
0.490 |
|
G/A-A/A |
178 (46.5%) |
115 (39.5%) |
0.64 (0.45-0.91) |
140 (46.7%) |
79 (47.9%) |
0.84 (0.52-1.37) |
||||
Recessive |
G/G-G/A |
353 (92.2%) |
277 (95.2%) |
1 |
0.049* |
285 (95%) |
154 (93.3%) |
1 |
0.990 |
|
A/A |
30 (7.8%) |
14 (4.8%) |
0.48 (0.23-1.02) |
15 (5%) |
11 (6.7%) |
1.01 (0.39-2.60) |
||||
Overdominant |
G/G-A/A |
235 (61.4%) |
190 (65.3%) |
1 |
0.110 |
175 (58.3%) |
97 (58.8%) |
1 |
0.470 |
|
G/A |
148 (38.6%) |
101 (34.7%) |
0.75 (0.52-1.07) |
125 (41.7%) |
68 (41.2%) |
0.84 (0.51-1.37) |
||||
Log-additive |
--- |
--- |
--- |
0.66 (0.50-0.89) |
0.005* |
--- |
--- |
0.90 (0.61-1.32) |
0.580 |
|
rs12232780 |
Codominant |
G/G |
215 (56.1%) |
198 (68%) |
1 |
0.002* |
0.970 |
|||
G/A |
146 (38.1%) |
84 (28.9%) |
0.56 (0.39-0.82) |
191 (63.7%) |
94 (57%) |
1 |
||||
A/A |
22 (5.7%) |
9 (3.1%) |
0.33 (0.13-0.84) |
102 (34%) |
67 (40.6%) |
1.03 (0.62-1.71) |
||||
Dominant |
G/G |
215 (56.1%) |
198 (68%) |
1 |
0.001* |
7 (2.3%) |
4 (2.4%) |
0.86 (0.19-4.01) |
0.930 |
|
G/A-A/A |
168 (43.9%) |
93 (32%) |
0.53 (0.37-0.76) |
191 (63.7%) |
94 (57%) |
1 |
||||
Recessive |
G/G-G/A |
361 (94.3%) |
282 (96.9%) |
1 |
0.047* |
109 (36.3%) |
71 (43%) |
1.02 (0.62-1.68) |
0.840 |
|
A/A |
22 (5.7%) |
9 (3.1%) |
0.41 (0.16-1.03) |
293 (97.7%) |
161 (97.6%) |
1 |
||||
Overdominant |
G/G-A/A |
237 (61.9%) |
207 (71.1%) |
1 |
0.007* |
7 (2.3%) |
4 (2.4%) |
0.85 (0.19-3.92) |
0.880 |
|
G/A |
146 (38.1%) |
84 (28.9%) |
0.60 (0.42-0.88) |
198 (66%) |
98 (59.4%) |
1 |
||||
Log-additive |
--- |
--- |
--- |
0.57 (0.41-0.78) |
0.000* |
102 (34%) |
67 (40.6%) |
1.04 (0.63-1.72) |
||
Codominant |
C/C |
188 (49.5%) |
129 (44.3%) |
1 |
0.013* |
152 (50.7%) |
78 (47.3%) |
1 |
0.670 |
|
T/C |
171 (45%) |
128 (44%) |
1.05 (0.73-1.52) |
130 (43.3%) |
73 (44.2%) |
1.15 (0.69-1.90) |
||||
T/T |
21 (5.5%) |
34 (11.7%) |
2.69 (1.36-5.32) |
18 (6%) |
14 (8.5%) |
1.47 (0.59-3.66) |
||||
rs2072382 |
Dominant |
C/C |
188 (49.5%) |
129 (44.3%) |
1 |
0.270 |
152 (50.7%) |
78 (47.3%) |
1 |
0.470 |
T/C-T/T |
192 (50.5%) |
162 (55.7%) |
1.21 (0.86-1.72) |
148 (49.3%) |
87 (52.7%) |
1.20 (0.74-1.94) |
||||
Recessive |
C/C-T/C |
359 (94.5%) |
257 (88.3%) |
1 |
0.003 |
282 (94%) |
151 (91.5%) |
1 |
0.470 |
|
T/T |
21 (5.5%) |
34 (11.7%) |
2.62 (1.36-5.06) |
18 (6%) |
14 (8.5%) |
1.38 (0.57-3.32) |
||||
Overdominant |
C/C-T/T |
209 (55%) |
163 (56%) |
1 |
0.620 |
170 (56.7%) |
92 (55.8%) |
1 |
0.740 |
|
T/C |
171 (45%) |
128 (44%) |
0.91 (0.65-1.30) |
130 (43.3%) |
73 (44.2%) |
1.09 (0.67-1.77) |
||||
Log-additive |
--- |
--- |
--- |
1.35 (1.02-1.77) |
0.034 |
--- |
--- |
1.19 (0.81-1.73) |
0.380 |
|
OR = odds ratio; 95 % CI = 95 % confidence interval
p values were calculated by unconditional logistic regression adjusted for age
*p < 0.05 indicates statistical significance
We also observed that some haplotypes of the SH2B3 gene were associated with CAD risk. However, the specific haplotypes involved were different in the men and women. One block (rs7309325, rs2078863, rs7296313) showed to be associated with CAD risk only in women (Figure 1, Table 4). This association resulted from a modest but significant decrease in the frequency of the “TTC” haplotype in female patients compared with controls (OR = 0.59, 95% CI: 0.40-0.86; p = 0.007). Meanwhile, the haplotype “GTT” in another block (rs12580300, rs7309325, rs2078863) was associated with increased susceptibility to CAD (OR = 1.59, 95% CI: 1.01-2.50; p = 0.047) in the men population (Table 5, Figure 2). All of the above results were adjusted by age.
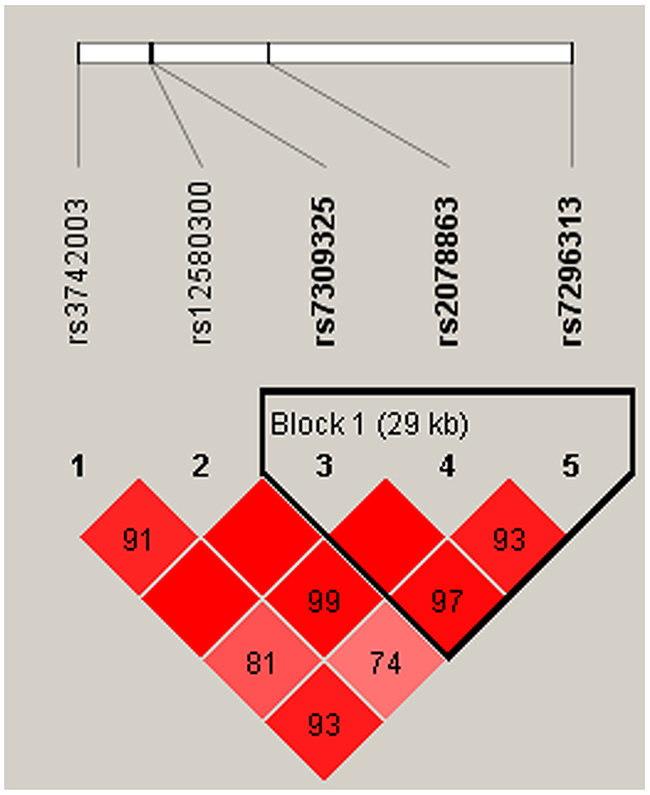
Figure 1: Haplotype block map for SNPs of the SH2B3 gene in the women population.
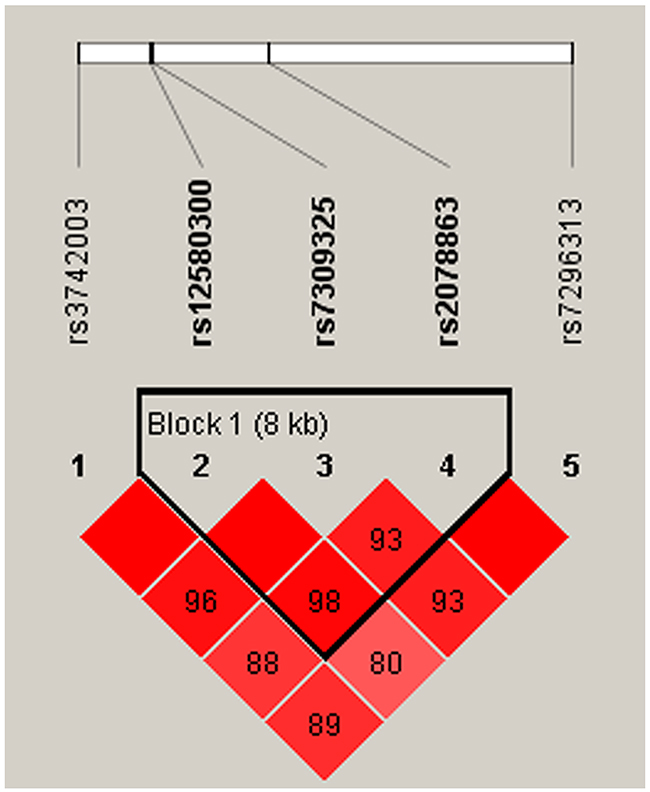
Figure 2: Haplotype block map for SNPs of the SH2B3 gene in the men population.
Table 4: SH2B3 haplotype frequencies and CAD risk association in the women group
Gene |
SNP |
Haplotype |
Frequency (%) |
OR 95%CI |
p |
|
|---|---|---|---|---|---|---|
Cases |
Controls |
|||||
SH2B3 |
rs7309325|rs2078863|rs7296313 |
TCC |
0.545 |
0.492 |
1 |
--- |
TTC |
0.361 |
0.451 |
0.59 (0.40 - 0.86) |
0.007* |
||
GTT |
0.082 |
0.085 |
0.79 (0.41 - 1.53) |
0.490 |
||
--- |
--- |
--- |
0.97 (0.18 - 5.15) |
0.970 |
||
OR = odds ratio; 95 % CI = 95 % confidence interval
p values were calculated using two-sided Chi-square test without adjustment for age
*p < 0.05 indicates statistical significance
Table 5: SH2B3 haplotype frequencies and CAD risk association in the men group
Gene |
SNP |
Haplotype |
Frequency (%) |
OR 95%CI |
p |
|
|---|---|---|---|---|---|---|
Cases |
Controls |
|||||
SH2B3 |
rs12580300|rs7309325|rs2078863 |
GTC |
0.471 |
0.495 |
1 |
--- |
ATT |
0.41 |
0.413 |
1.07 (0.82 - 1.39) |
0.61 |
||
GTT |
0.108 |
0.081 |
1.59 (1.01 - 2.50) |
0.047* |
||
--- |
--- |
--- |
0.82 (0.24 - 2.79) |
0.75 |
||
OR = odds ratio; 95 % CI = 95 % confidence interval
p value from were calculated from two-sided Chi-squared test without adjusted by age
*p < 0.05 indicates statistical significance
Next, SH2B3 and SMARCA4 gene polymorphisms were analyzed to establish their potential associations with study subjects’ lipid profiles, including triglyceride (TG), total cholesterol (TC), apolipoprotein A1/apolipoprotein B (APOA1/APOB), and HDL and LDL cholesterol (Table 6). Results showed that LDL cholesterol levels in rs3742003 G allele carriers were higher in cases versus controls in the male population (p = 0.049). In addition, LDL cholesterol levels were associated with rs7309325 (p = 0.048), rs7296313 (p = 0.040), and rs11879293 (p = 0.013) in the male population. In the female group, in contrast, we detected associations between rs3742003, rs7309325 and rs7296313, and APOA1/APOB levels. There were no significant differences between cases and controls in the other lipids profiled (Table 6).
Table 6: Lipid levels measured for SNP genotypes in the men and women populations
SNP |
TG (mmol/L) |
p |
TC (mmol/L) |
p |
HDL (mmol/L) |
p |
LDL (mmol/L) |
p |
APOA1/APOB |
p |
|---|---|---|---|---|---|---|---|---|---|---|
Men population (n = 291) |
||||||||||
rs3742003 |
||||||||||
A/A (n = 221) |
1.8687 ± 1.798 |
0.211 |
3.929 ± 1.14474 |
0.455 |
1.0817 ± 0.247 |
0.055 |
1.8005 ± 0.682 |
0.049* |
1.438 ± 0.744 |
0.726 |
G/A (n = 64) |
1.5745 ± 0.962 |
4.0567 ± 1.321 |
1.1503 ± 0.257 |
2.0514 ± 1.364 |
1.473 ± 0.609 |
|||||
rs12580300 |
||||||||||
G/G (n = 92) |
1.6597 ± 1.1596 |
0.587 |
3.99 ± 1.193 |
0.610 |
1.108 ± 0.264 |
0.768 |
2.0148 ± 1.163 |
0.107 |
1.487 ± 0.988 |
0.463 |
A/A (n = 41) |
1.8168 ± 1.3119 |
4.0898 ± 1.038 |
1.1102 ± 0.273 |
1.8382 ± 0.597 |
1.327 ± 0.480 |
|||||
G/A (n = 139) |
1.8881 ± 1.9825 |
3.8954 ± 1.2286 |
1.0868 ± 0.234 |
1.7623 ± 0.742 |
1.457 ± 0.542 |
|||||
rs7309325 |
||||||||||
T/T (n = 213) |
1.8592 ± 1.7928 |
0.271 |
3.9294 ± 1.1429 |
0.451 |
1.0824 ± 0.2458 |
0.062 |
1.8017 ± 0.6819 |
0.048* |
1.437 ± 0.7411 |
0.709 |
G/T (n = 62) |
1.5976 ± 0.9665 |
4.0595 ± 1.3322 |
1.1498 ± 0.2611 |
2.0555 ± 1.3812 |
1.476 ± 0.6161 |
|||||
rs7296313 |
||||||||||
C/C (n = 215) |
1.8807 ± 1.7968 |
0.125 |
3.9401 ± 1.1415 |
0.628 |
1.0864 ± 0.2548 |
0.158 |
1.8005 ± 0.6803 |
0.040* |
1.442 ± 0.7444 |
0.864 |
T/C (n = 60) |
1.5122 ± 0.8676 |
4.0247 ± 1.3458 |
1.138 ± 0.2319 |
2.0682 ± 1.400 |
1.46 ± 0.0772 |
|||||
rs11879293 |
||||||||||
G/G (n = 172) |
1.933 ± 1.9368 |
0.224 |
3.8893 ± 1.0763 |
0.140 |
1.0934 ± 0.2457 |
0.324 |
1.7395 ± 0.6508 |
0.013* |
1.485 ± 0.7881 |
0.511 |
A/A (n = 13) |
1.6231 ± 0.9815 |
3.6023 ± 1.2987 |
1.01 ± 0.1292 |
1.9146 ± 0.7138 |
1.369 ± 0.5040 |
|||||
G/A (n = 90) |
1.5721 ± 0.9515 |
4.1438 ± 1.3499 |
1.1184 ± 0.2708 |
2.079 ± 1.2209 |
1.383 ± 0.5787 |
|||||
rs2072382 |
||||||||||
C/C (n = 119) |
1.745 ± 1.1847 |
0.573 |
3.9791 ± 1.2366 |
0.081 |
1.0771 ± 0.2341 |
0.257 |
1.8449 ± 0.8019 |
0.081 |
1.457 ± 0.6025 |
0.021* |
T/C (n = 122) |
1.9077 ± 1.9746 |
4.0543 ± 1.1807 |
1.1254 ± 0.2679 |
1.9537 ± 1.0251 |
1.354 ± 0.5 |
|||||
T/T (n = 34) |
1.6082 ± 1.7521 |
3.5421 ± 0.9466 |
1.0697 ± 0.2378 |
1.5679 ± 0.5779 |
1.735 ± 1.3715 |
|||||
Women population (n = 165) |
||||||||||
rs3742003 |
||||||||||
A/A (n = 127) |
1.6872 ± 0.7916 |
0.069 |
4.3562 ± 1.0572 |
0.304 |
1.2002 ± 0.2547 |
0.550 |
2.0403 ± 0.6762 |
0.085 |
1.405 ± 0.5733 |
0.036* |
G/A (n = 27) |
2.3096 ± 2.5019 |
4.153 ± 1.0764 |
1.1448 ± 0.3397 |
1.8841 ± 0.8282 |
1.785 ± 1.1370 |
|||||
rs12580300 |
||||||||||
G/G (n = 62) |
1.9269 ± 1.7576 |
0.574 |
4.2015 ± 0.8815 |
0.464 |
1.1535 ± 0.2472 |
0.080 |
1.9276 ± 0.6519 |
0.368 |
1.434 ± 0.6677 |
0.285 |
A/A (n = 19) |
1.6637 ± 0.7636 |
4.3642 ± 1.2542 |
1.1268 ± 0.2492 |
2.1495 ± 0.7979 |
1.274 ± 0.5184 |
|||||
G/A (n = 73) |
1.72 ± 0.8241 |
4.4273 ± 1.1479 |
1.2365 ± 0.2505 |
2.0682 ± 0.7359 |
1.551 ± 0.7881 |
|||||
rs7309325 |
||||||||||
T/T (n = 127) |
1.6842 ± 0.7911 |
0.060 |
4.3565 ± 1.0572 |
0.305 |
1.2004 ± 0.2546 |
0.560 |
2.0382 ± 0.6769 |
0.094 |
1.408 ± 0.5727 |
0.036* |
G/T (n = 26) |
2.3358 ± 2.5476 |
4.1492 ± 1.0976 |
1.145 ± 0.2445 |
1.8919 ± 0.8436 |
1.796 ± 1.1581 |
|||||
rs7296313 |
||||||||||
C/C (n = 127) |
1.694 ± 0.7912 |
0.097 |
4.3457 ± 1.0468 |
0.373 |
1.1967 ± 0.2546 |
0.753 |
2.032 ± 0.6697 |
0.113 |
1.402 ± 0.511 |
0.029* |
T/C (n = 27) |
2.2774 ± 2.5105 |
4.2022 ± 1.1328 |
1.1611 ± 0.2437 |
1.9233 ± 0.8599 |
1.796 ± 0.218 |
|||||
rs11879293 |
||||||||||
G/G (n = 83) |
1.8205 ± 1.5942 |
0.929 |
4.1941 ± 0.9307 |
0.221 |
1.1711 ± 0.2454 |
0.550 |
1.9389 ± 0.6324 |
0.051 |
1.473 ± 0.6894 |
0.323 |
A/A (n = 11) |
1.6664 ± 0.8972 |
4.3755 ± 1.1683 |
1.2436 ± 0.3385 |
1.75 ± 0.6879 |
1.764 ± 1.1986 |
|||||
G/A (n = 61) |
1.7857 ± 0.7488 |
4.5048 ± 1.1942 |
1.2057 ± 0.2444 |
2.1839 ± 0.7899 |
1.411 ± 0.6327 |
|||||
rs2072382 |
||||||||||
C/C (n = 74) |
1.9289 ± 1.5378 |
0.358 |
4.3726 ± 0.9984 |
0.892 |
1.1814 ± 0.2459 |
0.303 |
1.9684 ± 0.6406 |
0.609 |
1.432 ± 0.6319 |
0.803 |
T/C (n = 70) |
1.7254 ± 1.0477 |
4.2901 ± 1.1109 |
1.2163 ± 0.2589 |
2.0529 ± 0.7795 |
1.496 ± 0.6449 |
|||||
T/T (n = 13) |
1.4443 ± 0.4935 |
4.2929 ± 1.2073 |
1.1057 ± 0.2394 |
2.1507 ± 0.7514 |
1.543 ± 1.2924 |
|||||
Triglyceride = TG; Total Cholesterol = TC; Apolipoprotein A1/ Apolipoprotein B = APOA1/APOB;
High-density lipoprotein = HDL; Low-density lipoprotein = LDL
p values were calculated using t test
*p < 0.05 indicates statistical significance
DISCUSSION
In the present case-control study we confirmed striking sex-related differences in the association of polymorphisms of the SH2B3 and SMARCA4 genes with CAD risk. Thus, certain SH2B3 SNPs, i,e, rs12580300 and rs2078863, were associated with CAD risk in women. In addition, we also underscored associations between SMARCA4 gene SNPs and CAD risk in men. These observations may explain differences in the propensity for early CAD development in men and women and may have potential sex-specific therapeutic implications.
The SH2B3 gene encodes SH2B adaptor protein 3, a member of the Src homology 2-B (SH2B) adaptor family. For a long time, the functions of the SH2B3 gene remained poorly understood, until it was shown to affect several traits linked to CAD, including regulation of hematopoiesis and cytokine signaling. A recent study demonstrated that a SH2B3 polymorphism was associated with both lower LDL cholesterol and HDL cholesterol concentration [25], whereas another study pointed the association of CAD risk loci in the SH2B3 gene with regulation of blood pressure [10].
Two SNPs in the SH2B3 gene showed an association with CAD risk in our study; one was associated with decreased risk in women and the other with increased risk. rs12580300, an intron-variant in the SH2B3 gene, was found to be associated with decreased CAD risk in the Chinese population for the first time. However, such association is not entirely clear. In contrast, rs2078863, also located in an intronic region of the SH2B3 gene, increased CAD risk and showed the strongest, so far unreported, association signal with CAD for SH2B3. Both loci were selected randomly. Therefore, further studies should be performed to verify the association.
SMARCA4 encodes a protein that belongs to the SWI/SNF protein family. It is the central catalytic component of the SWI/SNF complex, which involves multiple domains including an evolutionarily conserved catalytic ATPase domain, a conserved C-terminal bromodomain, an AT-hook motif, and the less characterized N-terminal region, all of which participate in the recognition of modified histone proteins, DNA binding, or recruitment of SWI/SNF [26, 27]. SMARCA4 is involved in various cellular processes including transcriptional regulation, cell cycle control, proliferation, DNA repair, and recombination [28]. Recently, the association of SMARCA4 with CAD risk has been highlighted by GWAS and other methods [22, 24, 29, 30]. Accordingly, our analysis shows that SMARCA4 rs11879293 and rs12232780 are associated with reduced risk of CAD in men, a result consistent with previous research by Martinelli and colleagues [23].
Our study is the first to demonstrate that CAD pathogenesis is influenced by sex differences in polymorphisms in SMARCA4 and SH2B3 genes. Although this study had sufficient statistical power, there were still some intrinsic limitations. First, we investigated only 5 genetic variants in each gene, and cannot exclude the possibility that other polymorphisms might also be associated with CAD risk. Also, we did not found an association between rs1122608 in SMARCA4 and CAD risk, as suggested by previous research [23]. Second, the association between SMARCA4 and SH2B3 polymorphisms and clinical information was not evaluated in this study. In addition, no significant associations were observed between the SNPs and the risk of CAD after Bonferroni correction. This may be due to our relatively small sample size, the SNP selection criteria (minor allele frequency > 5%), and inherent weaknesses of the Bonferroni correction itself (the interpretation of the results depends upon the number of comparisons performed). Multiple independent studies with large sample sizes are required to validate our findings.
MATERIALS AND METHODS
Subjects
The study included 456 Chinese CAD patients (291 men and 165 women) and 685 controls (385 men and 300 women) enrolled from September 2014 to October 2015 at the First hospital of Xi’an, China. The subjects presented no within-group relatedness. The Judkins approach [31] was used by highly skilled physicians to perform all coronary angiography procedures. A cardiologist diagnosed each patient according to applicable diagnostic criteria such as typical ischemic discomfort, electrocardiographic changes, increases in cardiac markers including creatine kinase-MB and troponin T, and coronary angiography outcomes (coronary diameter ≥ 2 mm; stenosis ≥ 50%). At least two experienced imaging specialists interpreted the coronary angiography findings, and the final CAD diagnosis was made based on the angiography report. Exclusion criteria included additional syndromes, multiple developmental abnormalities, or known chromosomal abnormalities. CAD patients were also excluded if their mothers had maternal diabetes mellitus, phenylketonuria, and teratogen or therapeutic drug exposure during pregnancy. In addition, 685 age-matched Chinese control subjects (ages 50–80 years) without signs or symptoms of CAD were recruited from the same hospital over the same time period. All control participants underwent a coronary angiogram confirming absence of coronary artery stenoses, and showed no clinical or electrocardiographic evidence of myocardial infarction or CAD. We recorded detailed information about the participants, including their personal medical history (hypertension, diabetes mellitus, etc.), familial medical history, reproductive history, menopausal status, and lifestyle habits (smoking, drinking, etc.). All subjects underwent a clinical examination at which a blood sample was drawn for routine blood analysis, biochemical tests, coagulation function, and genetic analyses. Informed consents were obtained from all participants. This study was approved by the ethical committee of Xi’an Jiaotong University, School of Medicine, Xi’an, China.
SNP selection and genotyping
This study selected two CAD-susceptibility genes, SH2B3 and SMARCA4, identified in previous research [25, 32]. SNPs within these genes were selected randomly from the HapMap database. Ten candidate SNPs (rs3742003, rs12580300, rs7309325, rs2078863, rs7296313, rs11879293, rs12232780, rs2072382, rs1529729, and rs1122608) in the SH2B3 and SMARCA4 genes with minor allele frequencies (MAF) >5 % in Asians were finally selected for genotyping.
We collected 5 ml venous blood samples in EDTA-containing tubes and stored them at -20˚C. DNA was obtained from whole blood leukocytes using the Gold Mag-Mini Whole Blood Genomic DNA Purification Kit (version 3.0; TaKaRa, Japan) [33] and its purity and concentration were determined spectrophotometrically by measuring the absorbance at 260 and 280 nm. Finally, genotypes derived from pure, integrated, and high-quality DNA samples were analyzed.
Genotyping was conducted using a Sequenom MassARRAY RS1000 (Sequenom, Inc.) following the manufacturer’s instructions. Briefly, it included a polymerase chain reaction (PCR) amplification assay, designing of primers and probes, purification with Shrimp Alkaline Phosphatase (SAP), addition of primers and extension of the basic group, followed by stimulating co-crystallization by combining sample analyte and chip substrates. The primers were designed using Sequenom MassARRAY Assay Design 3.0 software [34]. The overall success rate of all the genotyping assays was over 98%.
Statistical methods
All statistical analyses were performed using SPSS version 17.0 for Windows (SPSS, Chicago, IL). Differences in categorical and continuous variables between cases and controls were tested using the chi-square (χ2) test and the Student’s t-test, respectively. χ2 test for genotype distribution was conducted to evaluate deviation from Hardy–Weinberg equilibrium (HWE) for the ten SNPs. Genotype and allele distributions for SNPs between controls and CAD groups were also compared by χ2 test. All of the minor alleles were regarded as risk alleles for CAD susceptibility. Unconditional logistic regression, with adjustment for age, was used to estimate relative risk of CAD for each of the tested genotypes in the form of odds ratio (OR) and 95 % confidence interval (CI). All statistical tests were 2-sided and a p < 0.05 was considered as statistically significant. Linkage disequilibrium analysis and SNP haplotypes were done using the Haploview software package (version 4.2) and the SHEsis software platform (http://analysis.bio-x.cn/myAnalysis.php).
CONCLUSION
This is the first study examining shared genetic influences on CAD risk and revealing a strong sex-dependence for these associations. With basis on these findings, further studies on the contribution of germline genetic variants in the SH2B3 and SMARCA4 genes to CAD risk are warranted.
ACKNOWLEDGMENTS
This work was supported by the Science and Technology Research and Development Program of Shaanxi Province of China (No. 2014K11-03-01-02), the Science and Technology Planning Project of Xi’an (No. SF1416(4)), China Postdoctoral Science Foundation (No. 2016M592835) and Medical and Health Research Program of Yan’an (No. 2015KW-04). The authors are also grateful to the patients and control individuals for their participation in the study. We thank the clinicians and hospital staff who contributed to sampling and data collection for this study.
CONFLICTS OF INTEREST
The authors declare that there are no conflicts of interest.
REFERENCES
1. Zou JG, Ma YT, Xie X, Yang YN, Pan S, Adi D, Liu F, Chen BD. The association between CYP1A1 genetic polymorphisms and coronary artery disease in the Uygur and Han of China. Lipids in health and disease. 2014; 13:145.
2. Mackay J, Mensah GA, Mendis S, Greenlund K. The atlas of heart disease and stroke. Geneva: World Health Organization; 2004.
3. Kibos A, Guerchicoff A. Susceptibility genes for coronary heart disease and myocardial infarction. Acute cardiac care. 2011; 13:136-142.
4. Lopez AD, Mathers CD, Ezzati M, Jamison DT, Murray CJ. Global and regional burden of disease and risk factors, 2001: systematic analysis of population health data. The Lancet. 2006; 367:1747-1757.
5. Wang B, Fu ZY, Ma YT, Huang D, Liu F, Dong CL, Wang T, Meng YJ. Identification of a CYP19 Gene Single-Nucleotide Polymorphism Associated with a Reduced Risk of Coronary Heart Disease. Genetic testing and molecular biomarkers. 2016; 20:2-10.
6. Winkelmann BR, Hager J. Genetic variation in coronary heart disease and myocardial infarction: methodological overview and clinical evidence. Pharmacogenomics. 2000; 1:73-94.
7. Harismendy O, Notani D, Song X, Rahim NG, Tanasa B, Heintzman N, Ren B, Fu XD, Topol EJ, Rosenfeld MG. 9p21 DNA variants associated with coronary artery disease impair interferon-[ggr] signalling response. Nature. 2011; 470:264-268.
8. Willer CJ, Sanna S, Jackson AU, Scuteri A, Bonnycastle LL, Clarke R, Heath SC, Timpson NJ, Najjar SS, Stringham HM. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nature genetics. 2008; 40:161-169.
9. Deloukas P, Kanoni S, Willenborg C, Farrall M, Assimes TL, Thompson JR, Ingelsson E, Saleheen D, Erdmann J, Goldstein BA, Stirrups K, Konig IR, Cazier JB, et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat Genet. 2013; 45:25-33.
10. Ehret GB, Munroe PB, Rice KM, Bochud M, Johnson AD, Chasman DI, Smith AV, Tobin MD, Verwoert GC, Hwang SJ, Pihur V, Vollenweider P, O’Reilly PF, et al. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature. 2011; 478:103-109.
11. Wain LV, Verwoert GC, O’Reilly PF, Shi G, Johnson T, Johnson AD, Bochud M, Rice KM, Henneman P, Smith AV, Ehret GB, Amin N, Larson MG, et al. Genome-wide association study identifies six new loci influencing pulse pressure and mean arterial pressure. Nat Genet. 2011; 43:1005-1011.
12. Teslovich TM, Musunuru K, Smith AV, Edmondson AC, Stylianou IM, Koseki M, Pirruccello JP, Ripatti S, Chasman DI, Willer CJ, Johansen CT, Fouchier SW, Isaacs A, et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature. 2010; 466:707-713.
13. Qayyum R, Snively BM, Ziv E, Nalls MA, Liu Y, Tang W, Yanek LR, Lange L, Evans MK, Ganesh S, Austin MA, Lettre G, Becker DM, et al. A meta-analysis and genome-wide association study of platelet count and mean platelet volume in african americans. PLoS genetics. 2012; 8:e1002491.
14. Barrett JC, Clayton DG, Concannon P, Akolkar B, Cooper JD, Erlich HA, Julier C, Morahan G, Nerup J, Nierras C, Plagnol V, Pociot F, Schuilenburg H, et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet. 2009; 41:703-707.
15. van Setten J, Isgum I, Smolonska J, Ripke S, de Jong PA, Oudkerk M, de Koning H, Lammers JW, Zanen P, Groen HJ, Boezen HM, Postma DS, Wijmenga C, et al. Genome-wide association study of coronary and aortic calcification implicates risk loci for coronary artery disease and myocardial infarction. Atherosclerosis. 2013; 228:400-405.
16. Erbilgin A, Civelek M, Romanoski CE, Pan C, Hagopian R, Berliner JA, Lusis AJ. Identification of CAD candidate genes in GWAS loci and their expression in vascular cells. Journal of lipid research. 2013; 54:1894-1905.
17. Dichgans M, Malik R, König IR, Rosand J, Clarke R, Gretarsdottir S, Thorleifsson G, Mitchell BD, Assimes TL, Levi C. Shared genetic susceptibility to ischemic stroke and coronary artery disease A genome-wide analysis of common variants. Stroke. 2014; 45:24-36.
18. Conlon N, Silva A, Guerra E, Jelinic P, Schlappe BA, Olvera N, Mueller JJ, Tornos C, Jungbluth AA, Young RH, Oliva E, Levine D, Soslow RA. Loss of SMARCA4 Expression Is Both Sensitive and Specific for the Diagnosis of Small Cell Carcinoma of Ovary, Hypercalcemic Type. The American journal of surgical pathology. 2016; 40:395-403.
19. Medina PP, Romero OA, Kohno T, Montuenga LM, Pio R, Yokota J, Sanchez-Cespedes M. Frequent BRG1/SMARCA4–inactivating mutations in human lung cancer cell lines. Human mutation. 2008; 29:617-622.
20. Medina PP, Sanchez-Cespedes M. Involvement of the chromatin-remodeling factor BRG1/SMARCA4 in human cancer. Epigenetics. 2008; 3:64-68.
21. Kidder BL, Palmer S. Examination of transcriptional networks reveals an important role for TCFAP2C, SMARCA4, and EOMES in trophoblast stem cell maintenance. Genome research. 2010; 20:458-472.
22. Lin Y, Guo X, Zhao B, Liu J, Da M, Wen Y, Hu Y, Ni B, Zhang K, Yang S, Xu J, Dai J, Wang X, et al. Association analysis identifies new risk loci for congenital heart disease in Chinese populations. Nature communications. 2015; 6:8082.
23. Martinelli N, Girelli D, Lunghi B, Pinotti M, Marchetti G, Malerba G, Pignatti PF, Corrocher R, Olivieri O, Bernardi F. Polymorphisms at LDLR locus may be associated with coronary artery disease through modulation of coagulation factor VIII activity and independently from lipid profile. Blood. 2010; 116:5688-5697.
24. Lee SE, Kim HS. Unraveling new therapeutic targets of coronary artery disease by genetic approaches. Circulation journal. 2015; 79:8-14.
25. Deloukas P, Kanoni S, Willenborg C, Farrall M, Assimes TL, Thompson JR, Ingelsson E, Saleheen D, Erdmann J, Goldstein BA. Large-scale association analysis identifies new risk loci for coronary artery disease. Nature genetics. 2013; 45:25-33.
26. Shen W, Xu C, Huang W, Zhang J, Carlson JE, Tu X, Wu J, Shi Y. Solution structure of human Brg1 bromodomain and its specific binding to acetylated histone tails. Biochemistry. 2007; 46:2100-2110.
27. Singh M, D’Silva L, Holak TA. DNA-binding properties of the recombinant high-mobility-group-like AT-hook-containing region from human BRG1 protein. Biological chemistry. 2006; 387:1469-1478.
28. Trotter KW, Archer TK. The BRG1 transcriptional coregulator. Nuclear receptor signaling. 2008; 6:e004.
29. Brænne I, Civelek M, Vilne B, Di Narzo A, Johnson AD, Zhao Y, Reiz B, Codoni V, Webb TR, Asl HF. Prediction of causal candidate genes in coronary artery disease loci. Arteriosclerosis, thrombosis, and vascular biology. 2015; 35:2207-2217.
30. Jamaldini SH, Babanejad M, Mozaffari R, Nikzat N, Jalalvand K, Badiei A, Sanati H, Shakerian F, Afshari M, Kahrizi K. Association of polymorphisms at LDLR locus with coronary artery disease independently from lipid profile. Acta Medica Iranica. 2014; 52:352-359.
31. Franchi E, Marino P, Biondi-Zoccai GG, De Luca G, Vassanelli C, Agostoni P. Transradial versus transfemoral approach for percutaneous coronary procedures. Current cardiology reports. 2009; 11:391-397.
32. Kathiresan S, Voight BF, Purcell S, Musunuru K, Ardissino D, Mannucci PM, Anand S, Engert JC, Samani NJ, Schunkert H. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nature genetics. 2009; 41:334-341.
33. Carracedo A, ed. Forensic DNA typing protocols. Totowa (NJ): Humana Press; 2005.
34. Gabriel S, Ziaugra L, Tabbaa D. SNP genotyping using the Sequenom MassARRAY iPLEX platform. Current protocols in human genetics. 2009; Chapter 2:Unit 2.12.

