INTRODUCTION
Apoptosis or programmed cell death is an important process in maintaining proper development and tissue homeostasis in multicellular organisms [1]. The apoptotic program is triggered by signaling events that result in activation of cysteine proteases called caspases. Eukaryotic cells constitutively express all components of apoptotic machinery, and caspase activation must be regulated by a very precise and sensitive mechanism. Proteolytic cascades mediated via activator dimerization cleave initiator caspases to activate effector caspases, which consequentially cleave multiple protein substrates, ultimately advancing the cell to destruction [2, 3]. In vertebrates, the core programmed cell death pathway is mediated via cytochrome-c release from the mitochondria which activates adaptor protein Apaf-1 to form a heptameric apoptosome complex [4, 5]. The Apaf-1 apoptosome complex facilitates autocatalytic cleavage of the caspase-9 zymogen and, as such, exhibits two orders of magnitude higher catalytic activity than free caspase-9 [6–8]. However, in Drosophila, the analogous pathway appears to have no dependence on free cytochrome-c [9–11]. Instead, the Drosophila Apaf-1 homolog, death-associated Apaf1-related killer (Dark) is permanently expressed and can form an octameric apoptosome complex which then activates the key initiator caspase death regulator Nedd2-like caspase (Dronc) [12–14]. Unlike caspase-9, the activated Dronc caspase domain is dissociated from the apoptosome but exhibits robust protease activity toward effector caspases [15–17]. Initiation of apoptosis in Drosophila is solely dependent on caspase inhibitors known as inhibitor of apoptosis proteins (IAPs) [18, 19]. A major Drosophila IAP, DIAP1, is an E3 ligase whose activity mediates the anti-apoptotic function via ubiquitination of caspases [16, 20, 21]. This protein is constantly expressed in Drosophila cells and eliminates active caspases, either inducing their degradation [22] or acting via nondegradative mechanisms [23–26]. Apoptotic stimuli block DIAP1’s activity mainly via transcriptional activation of a family of IAP antagonists, including Reaper (Rpr), Hid, and Grim (RHG proteins or IAP antagonists) [27–29], which bind DIAP1’s BIR domains, preventing their interaction with caspases [30, 31]. Contrary to DIAP1, much less is known about how other central proapoptotic protein, Dark, and the related apoptosome complex are regulated in Drosophila [12, 32]. It is hypothesized that Dark can also be regulated by ubiquitination, leading to its subsequent inactivation or degradation [15]. The process of protein elimination is often antagonized by specific deubiquitinases (DUBs) which remove the ubiquitin chains from target proteins, thereby stabilizing their activity.
Based on domain structure prediction, there are about 100 highly conserved human proteins which possess deubiquitinating activity, and they can be classified into six subfamilies [33, 34]. They are structurally diverse isopeptidases that specifically cleave ubiquitin conjugates at the ubiquitin carboxyl end. DUBs mediate the maintenance of the free ubiquitin pool in cells by processing ubiquitin precursors and recycling ubiquitin from proteins committed to proteasomal elimination [33]. An important function of many DUBs is to modulate protein stability by removing ubiquitin from target proteins. DUBs are involved in the regulation of several key apoptotic regulators, including IAPs and Bcl-2 family proteins [35–37]. Recently, it has been shown that deubiquitinase DUBAI stabilizes DIAP1, serving as an additional regulator inhibiting Drosophila apoptosis [38]. Deubiquitinase DUBA has also been identified as an enzyme associated with regulation of Dronc function [39]. However, the functions of the majority of predicted DUBs remain unknown. Herein, identification of a novel deubiquitinase, DUSP31/CG30421, that regulates proapoptotic components of the Drosophila cell death pathway is described.
RESULTS AND DISCUSSION
Genetic screening identified CG30421/DUSP31 as a Dronc phenotype suppressor
To assess whether any DUB enzymes are involved in regulation of proapoptotic components, such as Dark and Dronc, available double-stranded RNA (dsRNA) lines were screened for their ability to suppress the Dronc gain-of-function phenotype in fly eyes (GMR-Gal4 driver). Unlike Dark, overexpression of wild type Dronc protein in photoreceptor cells caused a mild apoptotic phenotype featured by an extensive number of depigmented photoreceptor cells (Figure 1b) [32, 40, 41]. Under this condition, Dark tends to be activated by an excess of Dronc, probably via a positive feedback loop triggered by effector caspases [42]. In a control experiment, inactivation of Dark in GMR-Gal4/UAS-Dronc, UAS-DarkdsRNA/+ genetic background leads to complete suppression of the Dronc phenotype (Figure 1e). Removal of the corresponding Dark or Dronc activity by some unknown factors would be manifested by a reduced number of depigmented eye facets. The modified genetic screen of the available DUBs was designed to identify UAS-dsRNA lines able to suppress the Dronc phenotype (screen cross: GMR-Gal4/UAS-Dronc x UAS-DUB’sdsRNA/+). The results of this screen showed the CG30421 dsRNA line suppresses the Dronc phenotype (Figure 1c versus 1b). In addition, the heterozygous KG04149 allele of this gene could also substantially suppress the Dronc phenotype (Figure 1d). Analysis of the KG04149 allele confirmed that the P-element insertion in the 5’-untranslated region of CG30421 leads to complete lack of its long isoform expression and significantly reduces that of the short transcripts (Figure 2C and discussed below). None of the other screened DUB dsRNA (16 genes) showed suppression of the Dronc phenotype but instead, showed often enhancement, featured by depigmentation and roughness of the eyes (example CG14619, Figure 1h, and Supplementary Table 1). This data suggests that the protein encoded by CG30421 is able to positively affect some proapoptotic proteins.
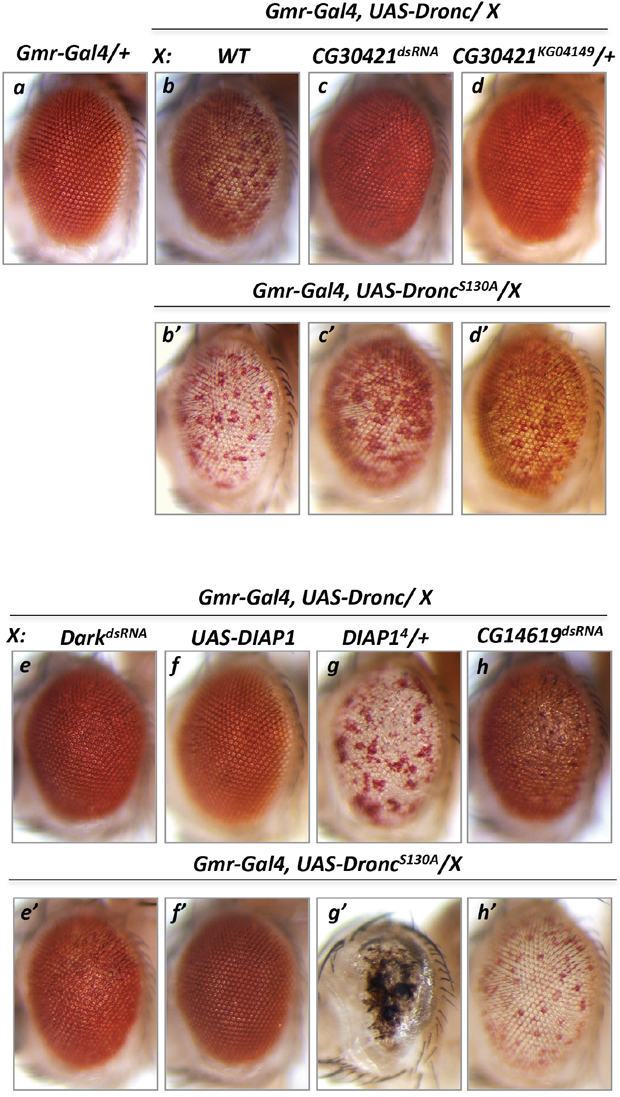
Figure 1: Modifier genetic screen identified deubiquitinase CG30421 loss-of-function alleles as suppressors of Dronc gain-of-function phenotype. Overexpression of Dronc induced apoptosis seen as depigmentation of a large number of Drosophila eye facets (b, Gmr-Gal4/+, UAS-Dronc/+) compared to Gmr-Gal4/+ control eyes (a). dsRNA-mediated inactivation of DUSP31 (c, Gmr-Gal4/UAS-DUSP31dsRNA, UAS-Dronc/+) or the heterozygous DUSP31 allele (d, Gmr-Gal4/DUSP31KG04149, UAS-Dronc/+), but not dsRNA-mediated inactivation of CG14619 (h, Gmr-Gal4/UAS-CG14619dsRNA, UAS-Dronc/+), suppressed the apoptotic Dronc phenotype in eye photoreceptor cells (b, Gmr-Gal4/+, UAS-Dronc/+). Phospho-mutant Dronc S130A induced stronger cell death seen as increased eye depigmentation (b’, Gmr-Gal4/+, UAS-Dronc S130A/+). Similar to wild type Dronc, inactivation of DUSP31 (c’, Gmr-Gal4/ UAS-DUSP31dsRNA, UAS-Dronc S130A /+) and the heterozygous allele of DUSP31 (d’, Gmr-Gal4/DUSP31KG04149, UAS-Dronc S130A /+), but not inactivation of CG14619 (h’, Gmr-Gal4/UAS-CG14619dsRNA, UAS-Dronc S130A /+), substantially suppressed the phenotype induced by the activated DroncS130A mutant. In the positive control, dsRNA-mediated inactivation of Dark (e, Gmr-Gal4/UAS-DarkdsRNA, UAS-Dronc/+ and e’, Gmr-Gal4/UAS-DarkdsRNA, UAS-Dronc S130A /+) or overexpression of DIAP1 (f, Gmr-Gal4/UAS-DIAP1, UAS-Dronc/+ and f’, Gmr-Gal4/UAS-DIAP1, UAS-Dronc S130A /+) suppressed Dronc and Dronc S130A overexpression phenotypes, while the heterozygous DIAP14 allele enhanced Dronc (g, Gmr-Gal4/+, UAS-Dronc/DIAP14) and DroncS130A (g’, Gmr-Gal4/+, UAS-Dronc S130A /DIAP14) phenotypes.
Phosphorylation of Dronc at S130 has been shown to mediate suppression of Dronc activation upon metabolic stimulation by glucose-6-phosphate dehydrogenase via increased levels of NADPH [43]. S130 phosphorylation does not affect Dronc’s catalytic activity but inhibits its interaction with Dark [43]. DroncS130A mutants possess constant and enhanced protease activity (Figure 1b’) [43] that can be substantially suppressed by inactivation of Dark (Figure 1e’) and completely suppressed by ectopic DIAP1 (Figure 1f’) compared with the control genetic background (Figure 1b’). In addition, whether the CG30421 alleles could also suppress the DroncS130A phenotype was also assessed. Importantly, RNA interference-mediated inactivation and the heterozygous CG30421 allele also substantially suppressed the DroncS130A phenotype (Figure 1c’ and 1d’ versus 1b’). Thus, these data additionally suggest that DUBs encoded by CG30421 are able to positively regulate some proapoptotic proteins. CG30421 is hereafter referred to as the Drosophila ubiquitin specific peptidase 31 (DUSP31) gene, as it has the closest homology to mammalian USP31 (Figure 2B).
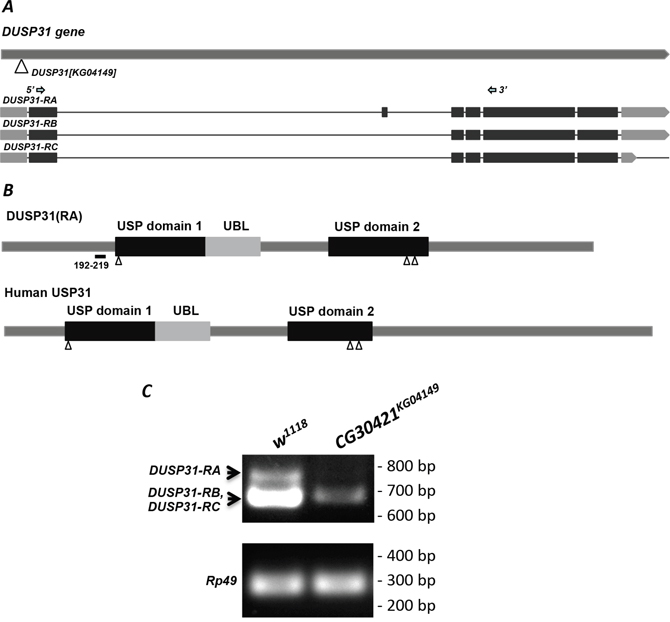
Figure 2: DUSP31KG04149 is a hypomorph P-element allele. (A) Scheme of the DUSP31 gene. The triangle indicates the location of P-element insertion into the DUSP31KG04149 allele. Three types of mRNA encoding DUSP31-RA, -RB, and -RC are shown (darker boxes are protein coding exons). Arrows indicate locations of forward and reverse primers used for RT-PCR. (B) Representation of DUSP31 and human USP31 protein domain structure. These DUBs have two highly conserved USP-catalytic and one ubiquitin-like (UBL) domains. Triangles indicate predicted active sites. The amino acid sequence from 192-219 denotes polypeptide differences between DUSP31-RA and DUSP31-RB/-RC isoforms. (C) Abnormal size and significantly reduced mRNA levels of DUSP31 revealed by semiquantitative RT-PCR of CG30421KG04149 homozygotes compared with wild type tissues (w1118). Arrows indicate amplified fragments of DUSP31-RA and DUSP31-RB/-RC isoforms. Rp49 was used as a loading control.
Also, inactivation of DUSP31 or Dark was consistently unable to completely suppress the DroncS130A phenotype (Figure 1c’ and 1e’), suggesting that some portion of the Dronc protein pool was not dependent on Dark’s ability to induce effector caspase activation. Furthermore, the same level of heterozygous DIAP1 (DIAP14/+) potentiated the strongest phospho-mutant DroncS130A activity (almost complete loss of eye) compared with mild enhancement of the wild type Dronc phenotype by this DIAP14 allele (Figure 1g’ versus 1g). Thus, DUSP31 is also required for a positive control of metabolically regulated Dark/Dronc activities.
The level of DUSP31 transcripts is dramatically reduced in DUSP31KG04149 transposon allele homozygotes, as evidenced by RT-PCR analysis (Figure 2C). Accordingly, three predicted transcripts (RA, and almost identical RB and RC) encoding two different protein isoforms of DUSP31 could be amplified (Figure 2A and 2C). The long transcript, DUSP31-RA, is not detected at all, while levels of shorter DUSP31-RB or -RC transcripts are more than 20-times lower in homozygous DUSP31KG04149 flies (Figure 2C), indicating this allele is a strong hypomorph. These homozygous flies are fertile and viable, while ubiquitous inactivation of DUSP31 with dsRNA (tub-Gal4) is lethal, suggesting that there is likely maternal contribution of this gene. Expectedly, expression of DUSP31dsRNA with Gmr-Gal4 does not affect eye development.
DUSP31 deficiency suppresses Grim-induced apoptosis
Next it was assessed whether DUSP31 deficiency was sufficient to alter apoptosis induced by RHG proteins or IAP antagonists acting upstream of Dark and DIAP1. Overexpressing one copy of Grim (Gmr-Gal4/+, Gmr-Grim/+ genetic background) caused depigmentation, mild roughness, and slight reduction in the size of fly eyes (Figure 3Ab versus 3Aa) [29, 40]. As expected, DUSP31 dsRNA strongly suppressed Grim-induced apoptosis in the eye (Figure 3Ac). In controls, inactivation of Dark strongly suppressed the Grim-induced phenotype (Figure 3Ad). On the other hand, hetero- and homozygous DUSP31KG04149 alleles were found to mildly but substantially suppress the strong apoptotic phenotype induced by overexpression of two copies of Gmr-Grim constructs (Gmr-Gal4/+, Gmr-Grim/Gmr-Grim genetic background) (Figure 3Bb and 3Bc) compared with the wild type background (Figure 3Ba). This observation is consistent with the notion that DUSP31 function is required to positively regulate apoptotic signals induced by one of RHG proteins, possibly involving stabilization of some proapoptotic factor. Interestingly, dsRNA and heterozygous DUSP31KG04149 alleles of DUSP31 do not affect the Reaper-induced eye phenotype (Gmr-Gal4/+, UAS-Rpr/+) (Figure 4Ac versus 4Ab). In addition, similar to the strong Gmr-Grim phenotype, DUSP31dsRNA and heterozygous DUSP31KG04149 alleles could only mildly suppress the apoptotic phenotype induced by Gmr-Gal4/UAS-DIAP1dsRNA, as seen by a slight increase in eye size (Supplementary Figure 1). This data suggests that DUSP31 plays a role as a fine-tune regulator of proapoptotic factors because strong signals induced by Rpr, doubled amounts of Grim, or DIAP1 activity removal (Supplementary Figure 1) caused robust activation of Dark and Dronc. This is also consistent with observations that inactivation of DIAP1 caused caspase-independent cell death, and DIAP1 may be involved in ubiquitination of Dark itself [15]. Dark is constitutively active and triggers activation of Dronc that is not bound by DIAP1. It has also been shown that excessive amounts of Dronc and Dark are degraded through a feedback inhibition mechanism mediated by the catalytic activity of Dronc, as well as by the C-terminal of Dark and DIAP1 [32]. Although the mechanism of Dark degradation is not clear, it likely involves ubiquitination. It has also been shown that overexpression of DUSP31 in S2 cells does not affect the levels of DIAP1 expression nor its stabilization [38]. Thus this suggests that DUSP31 acts independently of DIAP1, likely via association with Grim-induced signals, and regulates downstream Dark/Dronc proapoptotic transduction sites. Altogether, these results suggest DUSP31 functions as a regulator of the apoptosome complex downstream of RHG proteins and DIAP1.
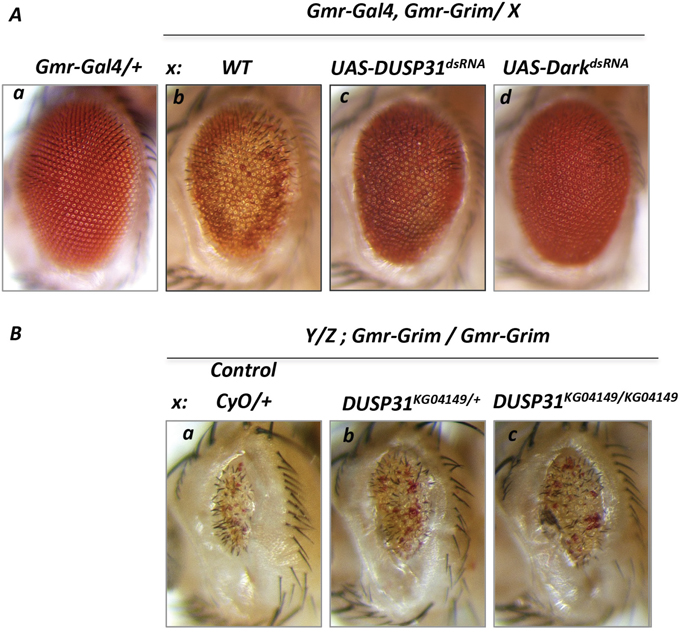
Figure 3: Deficiency in DUSP31 function suppressed apoptosis induced by overexpression of Grim protein. Image of wild type Drosophila eye (Aa, Gmr-Gal4/+). Overexpression of Grim (Ab, Gmr-Gal4/+, UAS-Grim/+) produced a cell death phenotype evidenced by depigmentation and slightly reduced eye size. dsRNA-mediated inactivation of DUSP31 (Ac, Gmr-Gal4/UAS-DUSP31dsRNA, Gmr-Grim/+) substantially suppressed depigmentation of eye photoreceptor cells caused by Grim-induced apoptosis. Inactivation of Dark almost completely suppressed the Grim-induced phenotype (Ad, Gmr-Gal4/+, Gmr-Grim/UAS-DarkdsRNA). (B) Deficiency in DUSP31 only mildly suppressed apoptosis caused by high levels of apoptotic induction. Overexpression of two copies of Grim (Ba, Gmr-Grim/Gmr-Grim) produced a strong cell death phenotype characterized by depigmentation, roughness, and a significant reduction in eye size. Heterozygous (Bb, DUSP31KG04149/CyO, Gmr-Grim/Gmr-Grim) and homozygous (Bc, DUSP31KG04149/DUSP31KG04149, Gmr-Grim/Gmr-Grim) DUSP31 alleles slightly suppressed the Grim-induced apoptotic phenotype compared with control (Ba, CyO/+, Gmr-Grim/Gmr-Grim).
DUSP31 is involved in positive regulation of Dark
The above experiments suggest that DUSP31 may be involved in regulation of Dark function. As seen by residual and mild eye roughness, inactivation of Dark by dsRNA did not completely suppress Rpr-induced cell death (Figure 4Ad and 4Ad’) compared with wild type and Rpr-mutant eyes (Figure 4Aa), suggesting residual Dark activity in Gmr-Gal4, UAS-Rpr, UAS-DarkdsRNA mutants. If DUSP31 protects Dark from degradation, then inactivating DUSP31 should further suppress Dark-mediated cell death. In agreement with this prediction, inactivation of DUSP31 on this genetic background completely suppressed cell death in the eye (Gmr-Gal4, UAS-Rpr, UAS- DarkdsRNA, UAS-DUSP31dsRNA) (Figure 4Ae, 4Ae’ versus 4Ad, 4Ad’). As a control, Ubpy DUB dsRNA did not suppress apoptosis in Gmr-Gal4, UAS-Rpr, UAS- DarkdsRNA, UAS-UpbydsRNA genetic background (Figure 4Bc versus 4Bb). This suggests that DUSP31 preferably acts as a positive regulator of Dark. To substantiate this idea further, a similar experiment was performed in which Dronc (instead of Dark) was inactivated in the Gmr-Gal4/UAS-Rpr background (Gmr-Gal4, UAS-Rpr, UAS- DroncdsRNA, UAS-DUSP31dsRNA). Similarly, reduction of Dronc levels led to strong but not complete suppression of Rpr-induced apoptosis characterizing by roughness in the eye (Figure 4Af versus 4Aa). In fact, if this residual activity of Dronc can be regulated by DUSP31, then it is possible that complete removal by inactive DUSP31 would completely suppress apoptosis, as shown for Dark. However, in contrast to the former, inactivation of DUSP31 on Gmr-Gal4, UAS-Rpr, UAS- DroncdsRNA, UAS-DUSP31dsRNA background did not suppress the residual apoptosis (Figure 4Ag, 4Ag’ versus 4Af, 4Af’). Thus, this genetic data strongly suggest that DUSP31 acts as a modifier regulating Dark activity.
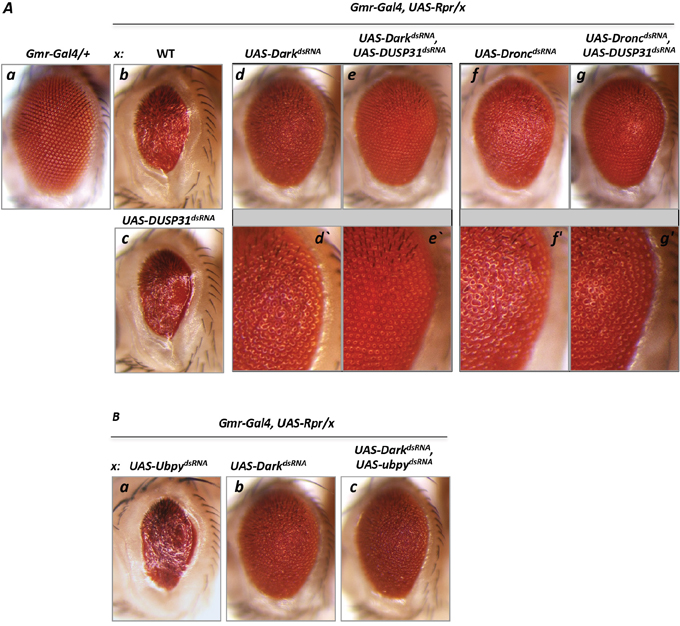
Figure 4: Inactivation of DUSP31 strongly suppressed the apoptotic phenotype caused by overexpression of Rpr and Dark dsRNA in Drosophila eyes. (A) Overexpression of Rpr (Ab, Gmr-Gal4/+, UAS-Rpr/+) induced a strong apoptosis phenotype manifested by glassines and a significantly reduced eye size compared with control (Aa, Gmr-Gal4/+). Inactivation of DUSP31 did not suppress the apoptotic phenotype caused by overexpression of Rpr (Ac, Gmr-Gal4/UAS-DUSP31dsRNA, UAS-Rpr/+). dsRNA-mediated inactivation of Dark significantly (but not completely) suppressed the Rpr-induced phenotype (Ad, Gmr-Gal4/UAS-DarkdsRNA, UAS-Rpr/+) evidenced by glassines and a rough eye phenotype (Ad’). Additional inactivation of DUSP31 in the Gmr-Gal4/UAS-DUSP31dsRNA, UAS-Rpr/UAS-DarkdsRNA background led to complete suppression of the apoptotic phenotype (Ae and Ae’). Conversely, inactivation of DUSP31 was not sufficient to suppress the phenotype caused by overexpression of Rpr and Dronc dsRNA (Ag and Ag’, Gmr-Gal4/UAS-DUSP31dsRNA, UAS-Rpr/UAS-DroncdsRNA) compared with Dronc inactivation alone (Af and Af’, Gmr-Gal4/+, UAS-Rpr/UAS-DroncdsRNA). (B) Control with an additional DUB, Ubpy. Inactivation of Ubpy did not affect the Rpr-induced phenotype (Ba, Gmr-Gal4/UAS-UbpydsRNA, UAS-Rpr/+ versus Ab). Ubpy inactivation did not suppress the phenotype in fly eyes caused by overexpression of both Rpr and dsRNA of Dark (Bc, Gmr-Gal4/UAS-UbpydsRNA, UAS-Rpr/UAS-DarkdsRNA) compared with the control (Bb, Gmr-Gal4/+, UAS-Rpr/UAS-DarkdsRNA).
On the other hand, to exclude the possibility that DUSP31 stabilizes downstream effector Death caspase-1 (Dcp-1), the presence of DUSP31 deficiency upon Dcp-1 overexpression in the eye was analyzed. Overexpression of Dcp1 with Gmr-Gal4 caused a robust apoptotic phenotype characterized by vast depigmentation in the eye (Figure 5Ab) [44–46] similar to the phenotype induced by overexpression of mutant DroncS130A. Surprisingly, inactivation of Dark also suppressed the Dcp-1 overexpression phenotype (Figure 5Ac versus 5Ab). Accordingly, inactivation of Dark was not predicted to affect the phenotype caused by overexpression of the downstream Dcp-1 enzyme. However, such was not the case; this can be explained by the dependency of Dcp-1 activation upon the activity of Dark and the apoptosome, which are probably activated via a positive feedback loop upon Dcp-1 overexpression. This notion is consistent with a previous report showing that an effector caspase can activate the Grim protein located upstream in the pathway [42]. Importantly, both DUSP31 alleles phenocopied the Dark phenotype in Gmr-Gal4/+, UAS-Dcp1/+ genetic background, thereby suppressing the Dcp-1 overexpression phenotype (Figure 5Ad and 5Ae). In controls, dsRNA-mediated inactivation of another DUB, CG7288, did not suppress the Dcp-1 overexpression phenotype (Figure 5Af). Thus, DUSP31 likely positively regulates Dark activity in this particular context. Taken together, these genetic data additionally suggest that DUSP31 acts as a modifier, activating Dark and/or its positive regulators.
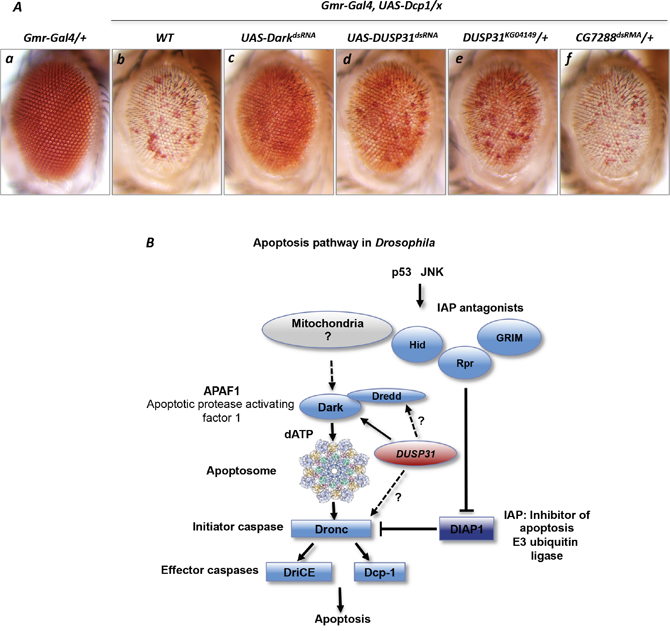
Figure 5: Inactivation of DUSP31 suppressed the apoptotic phenotype caused by Dcp-1 overexpression in Drosophila eyes. Overexpression of Dcp1 induced apoptosis in fly eyes manifested as depigmentation of a vast majority of eye facets (Ab, Gmr-Gal4/+, UAS-Dcp1/+) compared with Gmr-Gal4/+ control eyes (Aa). dsRNA-mediated inactivation of Dark (Ac, Gmr-Gal4/UAS-DarkdsRNA, UAS-Dcp1/+), DUSP31 (Ad, Gmr-Gal4/UAS-DUSP31dsRNA, UAS-Dcp1/+), and the heterozygous DUSP31 allele (Ae, Gmr-Gal4/UAS-DUSP31KG04149, UAS-Dcp1/+), but not dsRNA-mediated inactivation of CG7288 (Af, Gmr-Gal4/UAS-CG14619dsRNA, UAS-Dronc/+), suppressed the apoptotic Dcp-1 phenotype in eye photoreceptor cells (Ab, Gmr-Gal4/+, UAS-Dcp1/+). (B) Schematic representation of the Drosophila apoptosis pathway. The proapoptotic function of DUSP31 is associated with positive regulation of the Dark apoptosome protein or probably with apical caspases Dronc and Dredd (indicated by arrows).
To date, the exact mechanism of Dark protein activation and apoptosome complex assembly in Drosophila programmed cell death pathways remains poorly understood. In contrast to mammalian Apaf1, Dark associates with cytochrome-c; however, this interaction does not induce apoptosome formation [9]. On the other hand, dATP is important in the formation of this complex in both invertebrate and vertebrate systems [14]. It has also been shown that apical caspase Dredd interacts with Dark, mediating its activation [13]. It has not yet been shown whether Dark can be regulated by any post-translational modifications, including ubiquitination with subsequent proteasomal degradation. Further biochemical studies are required to answer this question. Dredd can be regulated by ubiquitination [47], and it is hypothesized that DUSP31 antagonizes this process, acting as proapoptotic factor. In the Dark apoptosome complex, the caspase recruitment domains interact with nucleotide-binding domains, leading to conformational changes within the complex to facilitate Dronc recruitment and activation [48–50]. Dronc is constantly ubiquitinated via DIAP1 action. Therefore, the possibility of DUSP31 playing a role during this step and activating the caspase cannot be excluded. Current genetic data described above indicate that DUSP31 could be involved in the regulation of all of these proapoptotic proteins (summarized in Figure 5B).
Drosophila DUSP31 encodes ubiquitin C-terminal hydrolase of family 2, called peptidase C19. These DUBs are large proteins ranging in size from 100 to 200 kDa and have two USP catalytic domains, Cys- and His-boxes, and one ubiquitin-like domain [51]. The three predicted and isolated isoforms of DUSP31 have minor differences in a short sequence lying near the Cys-box catalytic center of the protein’s N-terminus (Figure 2B); thus, these isoforms might have functional differences. However, only initial descriptions of the human homologs of DUSP31, USP31 and USP43, are available [33, 52, 53]. For example, it was reported that human USP31 ubiquitinase activity is involved in the regulation of nuclear factor-kappa B activation via components of the tumor necrosis factor signaling pathway [54]. It has also been reported that DUBs can be regulated by allosteric regulation within large multimolecular complexes by phosphorylation, ubiquitination, sumoylation, proteolytic cleavage, and direct stress stimuli, such as reactive oxygen species [33, 55–57]. Recently, several Drosophila DUBs were shown to be involved in regulation of apoptosis by stabilizing both anti- and proapoptotic cell death machinery proteins [38, 39, 58, 59]. DUBAI deubiquitinase positively controls the stability of central anti-apoptotic E3 ligase DIAP1. Another Drosophila DUB which is homologous to human USP36, called scny/et, functions as a direct or indirect positive regulator of DIAP1 [38, 58]. The DUBA enzyme (also known as OTUD5), belonging to the OTU class of DUBs, possesses Dronc-controlling activity, whose catalytic activity is positively regulated by phosphorylation [39]. The present genetic study identified a proapoptotic function for the deubiquitinase DUSP31, which is involved in the positive regulation of Dark/Dronc apoptosome components. This is only an initial step in characterization of this DUB’s function. Further biochemical analyses will decipher the molecular mechanisms of DUSP31 activity and regulation in Drosophila programmed cell death, as well as define a possible function for mammalian USP31 in vertebrate apoptosis.
MATERIALS AND METHODS
Drosophila stocks and genetics
w1118 flies were used as wild-type controls. Flies were maintained on standard Drosophila cornmeal/sucrose/yeast medium at 20, 25, or 29 °C as necessary. Details about the Drosophila stocks used are available in Supplementary Tables 1 and 2. The combined and screening fly stocks were made with the two-balancer stock (Stock # 3703, Drosophila Bloomington Stock Center). Genotypes of progeny from each cross are described in each figure legend. The modifier genetic screen was developed based on the UAS/Gal4 expression system [60] using the late developmental driver GMR-Gal4 in the fly eye at 25°C or other specified temperatures. Micrographs of the eyes of age-matched female progeny are shown.
Semiquantitative RT–PCR
DUSP31 transcripts were quantified using a one-step RT–PCR reaction kit (Invitrogen). For each reaction, the DUSP31 forward (5’-GCACCTGAG-CTCAACGGTTAA-3’) and reverse (5’-CTGTTGTTGCACCGGATGTGA-3’) primers were used in conjunction with 80 ng of total RNA. Rp49 was amplified with primers described in [61]. RNA was isolated from fly heads with TRIzolR (Invitrogen) and a DNA-free RNA kit (ZYMO Research). The PCR reaction was carried out for 30 cycles, with each cycle consisting of denaturation (94 °C for 30 s), annealing (55 °C for 30 s), and extension (72 °C for 1min).
ACKNOWLEDGMENTS
I thank Chihsheng Yang for critical comments during the study and manuscript preparation. I am grateful to Ook Joon Yoo for UAS-Dcp-1 flies, and Hyung Don Ryoo and Pascal Meier for the UAS-DIAP1dsRNA fly strain. I thank Sally Kornbluth, and Eric Spana for research support, laboratory resources and discussions. I thank Alexey N. Tomilin for laboratory resources and support. This work was supported by grants from NIH (R01 GM080333) and the Russian Science Foundation (14-15-00068), and the Federal Agency of Scientific Organizations (FASO Russia) for basic research.
CONFLICTS OF INTEREST
The author declares no conflicts of interest.
REFERENCES
1. Fuchs Y, Steller H. Programmed cell death in animal development and disease. Cell. 2011; 147:742-758.
2. Kornbluth S, White K. Apoptosis in Drosophila: neither fish nor fowl (nor man, nor worm). J Cell Sci. 2005; 118:1779-1787.
3. Kuida K, Haydar TF, Kuan CY, Gu Y, Taya C, Karasuyama H, Su MS, Rakic P, Flavell RA. Reduced apoptosis and cytochrome c-mediated caspase activation in mice lacking caspase 9. Cell. 1998; 94:325-337.
4. Riedl SJ, Li W, Chao Y, Schwarzenbacher R, Shi Y. Structure of the apoptotic protease-activating factor 1 bound to ADP. Nature. 2005; 434:926-933.
5. Zou H, Li Y, Liu X, Wang X. An APAF-1. cytochrome c multimeric complex is a functional apoptosome that activates procaspase-9. J Biol Chem. 1999; 274:11549-11556.
6. Cecconi F, Alvarez-Bolado G, Meyer BI, Roth KA, Gruss P. Apaf1 (CED-4 homolog) regulates programmed cell death in mammalian development. Cell. 1998; 94:727-737.
7. Li P, Nijhawan D, Budihardjo I, Srinivasula SM, Ahmad M, Alnemri ES, Wang X. Cytochrome c and dATP-dependent formation of Apaf-1/caspase-9 complex initiates an apoptotic protease cascade. Cell. 1997; 91:479-489.
8. Srinivasula SM, Ahmad M, Fernandes-Alnemri T, Alnemri ES. Autoactivation of procaspase-9 by Apaf-1-mediated oligomerization. Mol Cell. 1998; 1:949-957.
9. Abdelwahid E, Yokokura T, Krieser RJ, Balasundaram S, Fowle WH, White K. Mitochondrial disruption in Drosophila apoptosis. Dev Cell. 2007; 12:793-806.
10. Dorstyn L, Kumar S. A biochemical analysis of the activation of the Drosophila caspase DRONC. Cell Death Differ. 2008; 15:461-470.
11. Dorstyn L, Read S, Cakouros D, Huh JR, Hay BA, Kumar S. The role of cytochrome c in caspase activation in Drosophila melanogaster cells. J Cell Biol. 2002; 156:1089-1098.
12. Pang Y, Bai XC, Yan C, Hao Q, Chen Z, Wang JW, Scheres SH, Shi Y. Structure of the apoptosome: mechanistic insights into activation of an initiator caspase from Drosophila. Genes Dev. 2015; 29:277-287.
13. Rodriguez A, Oliver H, Zou H, Chen P, Wang X, Abrams JM. Dark is a Drosophila homologue of Apaf-1/CED-4 and functions in an evolutionarily conserved death pathway. Nat Cell Biol. 1999; 1:272-279.
14. Cheng TC, Akey IV, Yuan S, Yu Z, Ludtke SJ, Akey CW. A near-atomic structure of the Dark apoptosome provides insight into assembly and activation. Structure. 2017; 25:40-52.
15. Igaki T, Yamamoto-Goto Y, Tokushige N, Kanda H, Miura M. Down-regulation of DIAP1 triggers a novel Drosophila cell death pathway mediated by Dark and DRONC. J Biol Chem. 2002; 277:23103-23106.
16. Muro I, Hay BA, Clem RJ. The Drosophila DIAP1 protein is required to prevent accumulation of a continuously generated, processed form of the apical caspase DRONC. J Biol Chem. 2002; 277:49644-49650.
17. Rodriguez A, Chen P, Oliver H, Abrams JM. Unrestrained caspase-dependent cell death caused by loss of Diap1 function requires the Drosophila Apaf-1 homolog, Dark. EMBO J. 2002; 21:2189-2197.
18. Orme M, Meier P. Inhibitor of apoptosis proteins in Drosophila: gatekeepers of death. Apoptosis. 2009; 14:950-960.
19. Hay BA, Wassarman DA, Rubin GM. Drosophila homologs of baculovirus inhibitor of apoptosis proteins function to block cell death. Cell. 1995; 83:1253-1262.
20. Wang SL, Hawkins CJ, Yoo SJ, Muller HA, Hay BA. The Drosophila caspase inhibitor DIAP1 is essential for cell survival and is negatively regulated by HID. Cell. 1999; 98:453-463.
21. Yokokura T, Dresnek D, Huseinovic N, Lisi S, Abdelwahid E, Bangs P, White K. Dissection of DIAP1 functional domains via a mutant replacement strategy. J Biol Chem. 2004; 279:52603-52612.
22. Broemer M, Meier P. Ubiquitin-mediated regulation of apoptosis. Trends Cell Biol. 2009; 19:130-140.
23. Herman-Bachinsky Y, Ryoo HD, Ciechanover A, Gonen H. Regulation of the Drosophila ubiquitin ligase DIAP1 is mediated via several distinct ubiquitin system pathways. Cell Death Differ. 2007; 14:861-871.
24. Wilson R, Goyal L, Ditzel M, Zachariou A, Baker DA, Agapite J, Steller H, Meier P. The DIAP1 RING finger mediates ubiquitination of Dronc and is indispensable for regulating apoptosis. Nat Cell Biol. 2002; 4:445-450.
25. Kamber Kaya HE, Ditzel M, Meier P, Bergmann A. An inhibitory mono-ubiquitylation of the Drosophila initiator caspase Dronc functions in both apoptotic and non-apoptotic pathways. PLoS Genet. 2017; 13:1-24.
26. Lee TV, Fan Y, Wang S, Srivastava M, Broemer M, Meier P, Bergmann A. Drosophila IAP1-mediated ubiquitylation controls activation of the initiator caspase DRONC independent of protein degradation. PLoS Genet. 2011; 7:1-13.
27. White K, Tahaoglu E, Steller H. Cell killing by the Drosophila gene reaper. Science. 1996; 271:805-807.
28. Grether ME, Abrams JM, Agapite J, White K, Steller H. The head involution defective gene of Drosophila melanogaster functions in programmed cell death. Genes Dev. 1995; 9:1694-1708.
29. Chen P, Nordstrom W, Gish B, Abrams JM. grim, a novel cell death gene in Drosophila. Genes Dev. 1996; 10:1773-1782.
30. Goyal L, McCall K, Agapite J, Hartwieg E, Steller H. Induction of apoptosis by Drosophila reaper, hid and grim through inhibition of IAP function. EMBO J. 2000; 19:589-597.
31. Zachariou A, Tenev T, Goyal L, Agapite J, Steller H, Meier P. IAP-antagonists exhibit non-redundant modes of action through differential DIAP1 binding. EMBO J. 2003; 22:6642-6652.
32. Shapiro PJ, Hsu HH, Jung H, Robbins ES, Ryoo HD. Regulation of the Drosophila apoptosome through feedback inhibition. Nat Cell Biol. 2008; 10:1440-1446.
33. Komander D, Clague MJ, Urbe S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 2009; 10:550-563.
34. Fraile JM, Quesada V, Rodriguez D, Freije JM, Lopez-Otin C. Deubiquitinases in cancer: new functions and therapeutic options. Oncogene. 2012; 31:2373-2388.
35. Vucic D, Dixit VM, Wertz IE. Ubiquitylation in apoptosis: a post-translational modification at the edge of life and death. Nat Rev Mol Cell Biol. 2011; 12:439-452.
36. Lee EW, Seong D, Seo J, Jeong M, Lee HK, Song J. USP11-dependent selective cIAP2 deubiquitylation and stabilization determine sensitivity to Smac mimetics. Cell Death Differ. 2015; 22:1463-1476.
37. Weber A, Heinlein M, Dengjel J, Alber C, Singh PK, Hacker G. The deubiquitinase Usp27x stabilizes the BH3-only protein Bim and enhances apoptosis. EMBO Rep. 2016; 17:724-738.
38. Yang CS, Sinenko SA, Thomenius MJ, Robeson AC, Freel CD, Horn SR, Kornbluth S. The deubiquitinating enzyme DUBAI stabilizes DIAP1 to suppress Drosophila apoptosis. Cell Death Differ. 2014; 21:604-611.
39. Koerver L, Melzer J, Roca EA, Teichert D, Glatter T, Arama E, Broemer M. The de-ubiquitylating enzyme DUBA is essential for spermatogenesis in Drosophila. Cell Death Differ. 2016; 23:2019-2030.
40. Hawkins CJ, Yoo SJ, Peterson EP, Wang SL, Vernooy SY, Hay BA. The Drosophila caspase DRONC cleaves following glutamate or aspartate and is regulated by DIAP1, HID, and GRIM. J Biol Chem. 2000; 275:27084-27093.
41. Quinn LM, Dorstyn L, Mills K, Colussi PA, Chen P, Coombe M, Abrams J, Kumar S, Richardson H. An essential role for the caspase dronc in developmentally programmed cell death in Drosophila. J Biol Chem. 2000; 275:40416-40424.
42. Yeh TC, Bratton SB. Caspase-dependent regulation of the ubiquitin-proteasome system through direct substrate targeting. Proc Natl Acad Sci U S A. 2013; 110:14284-14289.
43. Yang CS, Thomenius MJ, Gan EC, Tang W, Freel CD, Merritt TJ, Nutt LK, Kornbluth S. Metabolic regulation of Drosophila apoptosis through inhibitory phosphorylation of Dronc. EMBO J. 2010; 29:3196-3207.
44. Xu D, Wang Y, Willecke R, Chen Z, Ding T, Bergmann A. The effector caspases drICE and dcp-1 have partially overlapping functions in the apoptotic pathway in Drosophila. Cell Death Differ. 2006; 13:1697-1706.
45. Song Z, McCall K, Steller H. DCP-1, a Drosophila cell death protease essential for development. Science. 1997; 275:536-540.
46. Kim YI, Ryu T, Lee J, Heo YS, Ahnn J, Lee SJ, Yoo O. A genetic screen for modifiers of Drosophila caspase Dcp-1 reveals caspase involvement in autophagy and novel caspase-related genes. BMC cell biology. 2010; 11:1-16.
47. Meinander A, Runchel C, Tenev T, Chen L, Kim CH, Ribeiro PS, Broemer M, Leulier F, Zvelebil M, Silverman N, Meier P. Ubiquitylation of the initiator caspase DREDD is required for innate immune signalling. The EMBO journal. 2012; 31:2770-2783.
48. Yu X, Acehan D, Menetret JF, Booth CR, Ludtke SJ, Riedl SJ, Shi Y, Wang X, Akey CW. A structure of the human apoptosome at 12.8 A resolution provides insights into this cell death platform. Structure. 2005; 13:1725-1735.
49. Wu CC, Bratton SB. DARK apoptosome secrets come to light. Structure. 2011; 19:4-6.
50. Yuan S, Yu X, Topf M, Dorstyn L, Kumar S, Ludtke SJ, Akey CW. Structure of the Drosophila apoptosome at 6.9 a resolution. Structure. 2011; 19:128-140.
51. Tsou WL, Sheedlo MJ, Morrow ME, Blount JR, McGregor KM, Das C, Todi SV. Systematic analysis of the physiological importance of deubiquitinating enzymes. PLoS One. 2012; 7:1-11.
52. Lockhart PJ, Hulihan M, Lincoln S, Hussey J, Skipper L, Bisceglio G, Wilkes K, Farrer MJ. Identification of the human ubiquitin specific protease 31 (USP31) gene: structure, sequence and expression analysis. DNA sequence: the journal of DNA Seq. 2004; 15:9-14.
53. Quesada V, Diaz-Perales A, Gutierrez-Fernandez A, Garabaya C, Cal S, Lopez-Otin C. Cloning and enzymatic analysis of 22 novel human ubiquitin-specific proteases. Biochem Biophys Res Commun. 2004; 314:54-62.
54. Tzimas C, Michailidou G, Arsenakis M, Kieff E, Mosialos G, Hatzivassiliou EG. Human ubiquitin specific protease 31 is a deubiquitinating enzyme implicated in activation of nuclear factor-kappaB. Cell Signal. 2006; 18:83-92.
55. Asaoka T, Ikeda F. New insights into the role of ubiquitin networks in the regulation of antiapoptosis pathways. Int Rev Cell Mol Biol. 2015; 318:121-158.
56. Cao MN, Zhou YB, Gao AH, Cao JY, Gao LX, Sheng L, Xu L, Su MB, Cao XC, Han MM, Wang MK, Li J. Curcusone D, a novel ubiquitin-proteasome pathway inhibitor via ROS-induced DUB inhibition, is synergistic with bortezomib against multiple myeloma cell growth. Biochim Biophys Acta. 2014; 1840:2004-2013.
57. Huang OW, Cochran AG. Regulation of deubiquitinase proteolytic activity. Curr Opin Struct Biol. 2013; 23:806-811.
58. Ribaya JP, Ranmuthu M, Copeland J, Boyarskiy S, Blair AP, Hay B, Laski FA. The deubiquitinase emperor's thumb is a regulator of apoptosis in Drosophila. Dev Biol. 2009; 329:25-35.
59. Kovacs L, Nagy O, Pal M, Udvardy A, Popescu O, Deak P. Role of the deubiquitylating enzyme DmUsp5 in coupling ubiquitin equilibrium to development and apoptosis in Drosophila melanogaster. PLoS One. 2015; 10:1-19.
60. Brand AH, Perrimon N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 1993; 118:401-415.
61. Sinenko SA, Mathey-Prevot B. Increased expression of Drosophila tetraspanin, Tsp68C, suppresses the abnormal proliferation of ytr-deficient and Ras/Raf-activated hemocytes. Oncogene. 2004; 23:9120-9128.

