INTRODUCTION
The identification of activated forms of the epidermal growth factor receptor (EGFR) as oncogenic drivers in non–small cell lung cancer (NSCLC) has rendered this receptor a key target in precision medicine [1]. Activation of wild-type (WT) EGFR in response to ligand binding is mediated by receptor homodimerization and autophosphorylation. In contrast, activating mutations of EGFR present in some NSCLC tumors result in receptor dimerization in the absence of ligand and in constitutive activation of the receptor tyrosine kinase. Compared with standard platinum-based chemotherapy doublets, EGFR tyrosine kinase inhibitors (TKIs) such as gefitinib, erlotinib, and afatinib show more pronounced antitumor effects in NSCLC patients who harbor such activating EGFR mutations, which include in-frame deletions in exon 19 and an L858R point mutation in exon 21 [2–5]. Although the response rate of such patients to EGFR-TKIs is ~80%, it follows that these drugs have no effect in ~20% of mutation-positive patients. The response rate was only 8.9% in NSCLC patients WT for EGFR who received erlotinib after one or two prior chemotherapy regimens [6]. EGFR-TKIs also prolong survival in a subset of patients with colon, pancreatic, or head and neck cancer WT for EGFR [7–9]. Evidence thus suggests that the efficacy of such treatment varies among individuals regardless of EGFR mutation status and may also reflect EGFR activation not attributable to mutation.
Polymerase chain reaction (PCR)–based assays are usually adopted for detection of EGFR mutations [10]. However, a method for detection of EGFR activation that is not based on mutation identification has not been established in the clinical setting. We have now applied a proximity ligation assay (PLA) to visualize and quantitate EGFR homodimerization. We also examined the relation of EGFR dimerization determined by PLA analysis to EGFR autophosphorylation.
RESULTS
Detection of EGFR homodimers by PLA analysis
We first attempted to detect EGFR homodimers in seven NSCLC cell lines (Supplementary Table 1) positive or negative for activating EGFR mutations with the use of PLA probes derived from a monoclonal antibody to EGFR. PLA signals were detected in EGFR mutation–positive PC9 cells in a manner dependent on the addition of both PLUS and MINUS probes (Figure 1A), with annealing of the probes and consequent generation of the fluorescence signal indicating the presence of EGFR homodimers. PLA analysis also detected EGFR homodimers in EGFR mutation–positive HCC827 cells and to a much lesser extent in EGFR-WT A549 cells (Figure 1B), suggesting that the level of homodimerization is related to EGFR activation.
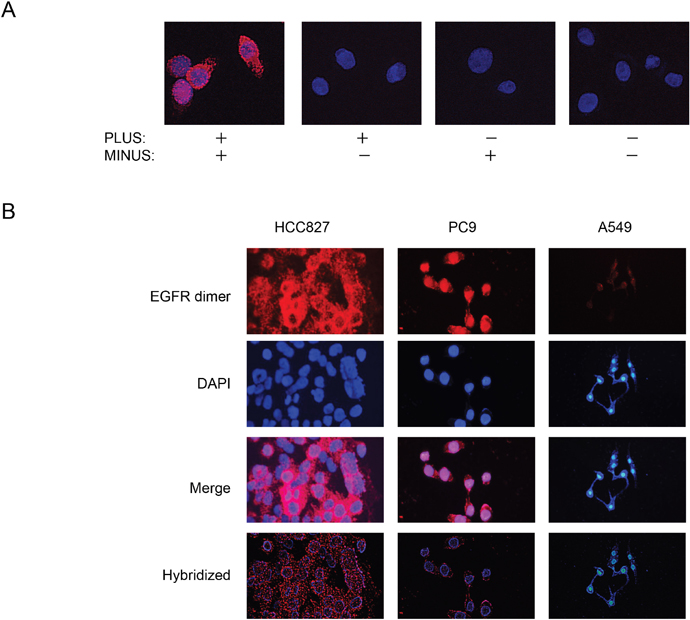
Figure 1: Detection of EGFR homodimers in NSCLC cell lines by PLA analysis. (A) PLA analysis of PC9 cells performed in the absence or presence of PLUS and MINUS probes as indicated. Red signals corresponding to EGFR homodimers were detected only in the presence of both probes. Nuclei were stained blue with 4', 6-diamidino-2-phenylindole (DAPI). (B) PLA analysis of EGFR mutation–positive (PC9, HCC827) or –negative (A549) NSCLC cell lines.
Relation between EGFR homodimerization and autophosphorylation
We next examined the effect of EGF on EGFR homodimerization in NSCLC cell lines. Whereas immunohistochemistry revealed no substantial effect of EGF on the pattern of EGFR expression in HCC827 or A549 cells (Figure 2A), PLA analysis showed that EGF induced EGFR homodimerization in A549 and H2228 cells (both of which are WT for EGFR) but not in EGFR mutation–positive HCC827 cells (Figure 2B). Furthermore, immunoblot and PLA analyses revealed phosphorylation (Figure 2C) and homodimerization (Figure 2D) of exogenous EGFR in transfected Ba/F3 cells. We then examined the relation between EGFR homodimerization (Figure 2E) and autophosphorylation (Figure 2F) in NSCLC cell lines positive (HCC827, PC9, 11_18) or negative (A549, H2228, H157, SBC5) for EGFR mutations. PLA analysis revealed that EGF induced a marked increase in the extent of EGFR homodimerization in all cell lines with the exception of HCC827 and H157. Similarly, immunoblot analysis showed that EGF markedly increased the extent of EGFR phosphorylation in most cell lines, although HCC827 showed a high basal level of such phosphorylation. These data thus suggested that the extent of EGFR homodimerization as determined by PLA analysis is related to the extent of EGFR phosphorylation in NSCLC cell lines.
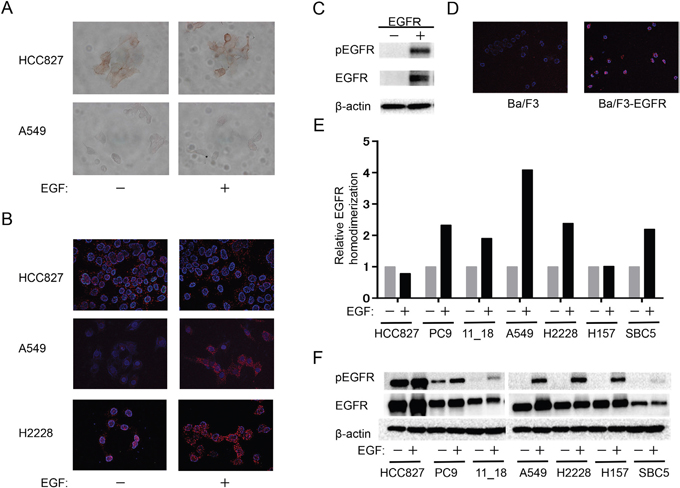
Figure 2: Relation between EGFR homodimerization and phosphorylation in NSCLC cell lines. (A, B) The indicated cell lines were deprived of serum overnight, exposed (or not) to EGF (100 ng/ml) for 10 min, and then subjected either to immunohistochemistry with antibodies to EGFR (A) or to PLA analysis of EGFR homodimers (B). (C, D) Parental Ba/F3 cells or Ba/F3 cells transfected with an EGFR expression plasmid (Ba/F3-EGFR) and then cultured for 12 h were subjected either to immunoblot analysis of phosphorylated (p) or total forms of EGFR as well as of β-actin (loading control) (C) or to PLA analysis of EGFR homodimerization (D). (E, F) The indicated cell lines were deprived of serum and stimulated with EGF as in A. They were then subjected both to quantitative PLA analysis of EGFR homodimers (E) and to immunoblot analysis of EGFR phosphorylation (F). Data in E are expressed relative to the corresponding value for nonstimulated cells and are means from a representative experiment.
PLA analysis of NSCLC tissue
Finally, we applied the PLA method to NSCLC tissue obtained by transbronchial lung biopsy from 15 patients harboring EGFR mutations and 14 patients WT for EGFR. Consistent with the cell line data, the extent of EGFR homodimerization was significantly higher in tumors harboring EGFR mutations than in those WT for EGFR (Figure 3). These results thus indicated that the detection of EGFR homodimers by PLA analysis is also feasible for tissue samples from NSCLC patients.
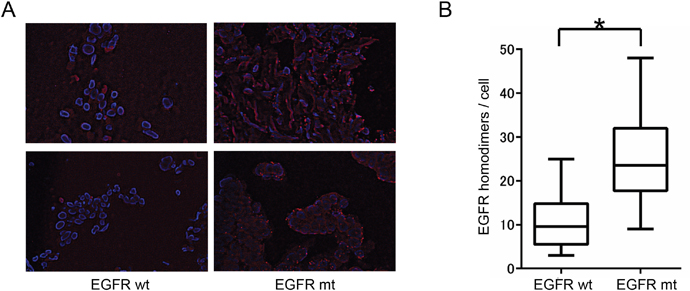
Figure 3: Relation between EGFR homodimerization and EGFR mutation in NSCLC tissue specimens. (A) PLA analysis of EGFR homodimers in tumor tissue from two patients positive for EGFR mutations (EGFR mt) and two patients WT for EGFR (EGFR wt). (B) Box-and-whisker plots for the extent of EGFR homodimerization determined as in A for 15 patients with and 14 without EGFR mutations. *P < 0.05 (Mann-Whitney U test).
DISCUSSION
We have here demonstrated the detection of EGFR homodimers in NSCLC cells with a PLA.
PLA signals for EGFR homodimers tended to be higher in NSCLC cells harboring EGFR mutations than in those WT for EGFR. We further found that EGF increased EGFR homodimerization in association with induction of EGFR autophosphorylation in NSCLC cells WT for EGFR. Recent studies also found that PLA signals corresponding to EGFR-related interactions such as those between EGFR and GRB2 or between nonphosphorylated and phosphorylated forms of EGFR were higher in NSCLC cell lines positive for EGFR mutations than in those WT for EGFR [11–12]. EGFR belongs to the HER family, the members of which include EGFR (HER1), HER2, HER3, and HER4 and form heterodimers with each other as well as homodimers [13]. We also detected the formation of EGFR-HER2 heterodimers in PC9 cells by PLA analysis (Supplementary Figure 1A). Furthermore, we detected PLA signals corresponding to the EML4-ALK fusion protein in NSCLC cells harboring the EML4-ALK fusion oncogene (Supplementary Figure 1B), suggesting that this method will be applicable to the detection of other such fusion oncoproteins.
The PLA method also detected EGFR homodimerization in fixed tumor tissue from NSCLC patients. This analysis was performed with small NSCLC tissue specimens obtained by transbronchial lung biopsy, suggesting that the method should be applicable to the detection of EGFR homodimers in tissue microarrays. Given that EGFR-TKI therapy confers a survival benefit in some patients with colon, pancreatic, or head and neck tumors that are WT for EGFR, the amount of activated EGFR may be an important marker for the response to TKI therapy in such tumors. Whereas there is currently no commercially available antibody that is suitable for detection of EGFR activation by immunohistochemistry, our results now suggest that detection of EGFR homodimers in tissue samples by PLA offers an alternative approach to prediction of the response to EGFR-TKIs.
In conclusion, we have detected EGFR homodimers by PLA analysis in a quantitative manner in both NSCLC cell lines and tissue specimens obtained by transbronchial lung biopsy. The PLA signal for EGFR homodimerization appeared to be related to the extent of EGFR phosphorylation. PLA analysis may thus provide a new tool for visualization and quantitation of EGFR dimerization and activation.
MATERIALS AND METHODS
Cell culture and reagents
Cell lines were obtained and maintained as previously described [14], with the exception that SBC5 cells were provided by Japanese Collection of Research Bioresources. For PLA, cells were grown to 60% confluence on 12-mm-diameter cover glasses (Matsunami) placed in a 24-well plate (Greiner bio-one). EGF (Sigma-Aldrich) was dissolved in acetic acid (Wako) and stored at 4°C for up to 1 month.
Patient specimens
Tumor specimens were obtained from 29 patients with lung adenocarcinoma (15 with an activating EGFR mutation, 14 WT for EGFR) who had undergone transbronchial lung biopsy between January 2007 and January 2016 at Kyushu University Hospital. EGFR mutations were identified by the peptide nucleic acid–locked nucleic acid PCR clamp method [10]. Specimens were fixed in 10% neutral buffered formalin, embedded in paraffin, sectioned, and treated as previously described [14]. The present study conforms to the tenets of the Declaration of Helsinki and was approved by the ethics review board of Kyushu University.
In situ PLA and microscopy analysis
Cells were fixed for 20 min with 4% paraformaldehyde in phosphate-buffered saline (PBS) and permeabilized for 15 min with 0.2% Triton X-100 in PBS. An in situ PLA for detection of EGFR homodimers was performed with Duolink II PLA probes, Probemaker, and detection reagents (Olink Bioscience). Rabbit monoclonal antibodies to EGFR (Abcam) were conjugated to PLUS or MINUS PLA oligonucleotide arms with the use of Probemaker. Cells grown on cover glasses or tumor sections were incubated overnight at 4°C with the antibody-oligonucleotide complexes (PLA probes). Annealing of the PLUS and MINUS PLA probes occurs when two EGFR monomers are in close proximity, and repeat sequences in the annealed oligonucleotide complexes are amplified and then recognized by a fluorescently labeled oligonucleotide probe. PLA signals were detected with a Keyence BZ-8100 fluorescence microscope and were quantified with BZ Analyzer software (Keyence); 20 cells for each cell line and all cells in each image for tumor sections were examined for determination of the number of PLA signals per cell.
Immunohistochemical staining
Cells were incubated overnight at 4°C with rabbit polyclonal antibodies to EGFR (Cell Signaling Technology), and immune complexes were detected with secondary antibodies, the streptavidin-biotin-peroxidase system, and 3, 3'-diaminobenzidine (Nichirei).
Immunoblot analysis
Immunoblot analysis was performed as previously described [14]. Rabbit polyclonal antibodies to human Tyr1068-phosphorylated EGFR, to EGFR, and to β-actin were obtained from Cell Signaling Technology.
Plasmid transfection
A Cell Line Nucleofector Kit V was obtained from Lonza. Ba/F3 cells (2 × 106) were suspended in 100 μl of Nucleofector solution containing 2 μg of an expression vector for human EGFR [15] and were then subjected to electroporation according to program X-001 with a Nucleofector II Device (Amaxa Biosystems).
Author contributions
KO performed the in vitro analyses and wrote the initial draft of the manuscript. TH designed the study. All of the authors contributed to data collection and interpretation and to critical review of the manuscript. The final version of the manuscript was approved by all authors.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
This work was supported in part by the Fukuoka Foundation for Sound Health Cancer Research Fund.
REFERENCES
1. Oxnard GR, Binder A, Janne PA. New targetable oncogenes in non-small-cell lung cancer. J Clin Oncol. 2013; 31:1097-1104.
2. Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H, Gemma A, Harada M, Yoshizawa H, Kinoshita I, Fujita Y, Okinaga S, Hirano H, et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med. 2010; 362:2380-2388.
3. Takano T, Ohe Y, Sakamoto H, Tsuta K, Matsuno Y, Tateishi U, Yamamoto S, Nokihara H, Yamamoto N, Sekine I, Kunitoh H, Shibata T, Sakiyama T, et al. Epidermal growth factor receptor gene mutations and increased copy numbers predict gefitinib sensitivity in patients with recurrent non-small-cell lung cancer. J Clin Oncol. 2005; 23:6829-6837.
4. Zhou C, Wu YL, Chen G, Feng J, Liu XQ, Wang C, Zhang S, Wang J, Zhou S, Ren S, Lu S, Zhang L, Hu C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011; 12:735-742.
5. Yang JC, Wu YL, Schuler M, Sebastian M, Popat S, Yamamoto N, Zhou C, Hu CP, O'Byrne K, Feng J, Lu S, Huang Y, Geater SL, et al. Afatinib versus cisplatin-based chemotherapy for EGFR mutation-positive lung adenocarcinoma (LUX-Lung 3 and LUX-Lung 6): analysis of overall survival data from two randomised, phase 3 trials. Lancet Oncol. 2015; 16:141-151.
6. Shepherd FA, Rodrigues Pereira J, Ciuleanu T, Tan EH, Hirsh V, Thongprasert S, Campos D, Maoleekoonpiroj S, Smylie M, Martins R, van Kooten M, Dediu M, Findlay B, et al. Erlotinib in previously treated non-small-cell lung cancer. N Engl J Med. 2005; 353:123-132.
7. Jonker DJ, O'Callaghan CJ, Karapetis CS, Zalcberg JR, Tu D, Au HJ, Berry SR, Krahn M, Price T, Simes RJ, Tebbutt NC, van Hazel G, Wierzbicki R, et al. Cetuximab for the treatment of colorectal cancer. N Engl J Med. 2007; 357:2040-2048.
8. Moore MJ, Goldstein D, Hamm J, Figer A, Hecht JR, Gallinger S, Au HJ, Murawa P, Walde D, Wolff RA, Campos D, Lim R, Ding K, et al. Erlotinib plus gemcitabine compared with gemcitabine alone in patients with advanced pancreatic cancer: a phase III trial of the National Cancer Institute of Canada Clinical Trials Group. J Clin Oncol. 2007; 25:1960-1966.
9. Vermorken JB, Mesia R, Rivera F, Remenar E, Kawecki A, Rottey S, Erfan J, Zabolotnyy D, Kienzer HR, Cupissol D, Peyrade F, Benasso M, Vynnychenko I, et al. Platinum-based chemotherapy plus cetuximab in head and neck cancer. N Engl J Med. 2008; 359:1116-1127.
10. Nagai Y, Miyazawa H, Huqun, Tanaka T, Udagawa K, Kato M, Fukuyama S, Yokote A, Kobayashi K, Kanazawa M, Hagiwara K. Genetic heterogeneity of the epidermal growth factor receptor in non-small cell lung cancer cell lines revealed by a rapid and sensitive detection system, the peptide nucleic acid-locked nucleic acid PCR clamp. Cancer Res. 2005; 65:7276-7282.
11. Smith MA, Hall R, Fisher K, Haake SM, Khalil F, Schabath MB, Vuaroqueaux V, Fiebig HH, Altiok S, Chen YA, Haura EB. Annotation of human cancers with EGFR signaling-associated protein complexes using proximity ligation assays. Sci Signal. 2015; 8:ra4.
12. Chen TC, Liu YW, Huang YH, Yeh YC, Chou TY, Wu YC, Wu CC, Chen YR, Cheng HC, Lu PJ, Lai JM, Huang CY. Protein phosphorylation profiling using an in situ proximity ligation assay: phosphorylation of AURKA-elicited EGFR-Thr654 and EGFR-Ser1046 in lung cancer cells. PLoS One. 2013; 8:e55657.
13. Graus-Porta D, Beerli RR, Daly JM, Hynes NE. ErbB-2, the preferred heterodimerization partner of all ErbB receptors, is a mediator of lateral signaling. EMBO J. 1997; 16:1647-1655.
14. Ota K, Azuma K, Kawahara A, Hattori S, Iwama E, Tanizaki J, Harada T, Matsumoto K, Takayama K, Takamori S, Kage M, Hoshino T, Nakanishi Y, Okamoto I. Induction of PD-L1 expression by the EML4-ALK oncoprotein and downstream signaling pathways in non-small cell lung cancer. Clin Cancer Res. 2015; 21:4014-4021.
15. Furuyama K, Harada T, Iwama E, Shiraishi Y, Okamura K, Ijichi K, Fujii A, Ota K, Wang S, Li H, Takayama K, Giaccone G, Nakanishi Y. Sensitivity and kinase activity of epidermal growth factor receptor (EGFR) exon 19 and others to EGFR-tyrosine kinase inhibitors. Cancer Sci. 2013; 104:584-589.

