INTRODUCTION
Coronary heart disease (CHD) is a major cause of death and disability worldwide [1]. In China, the CHD death rate increased from 141 per 10,000,000 person-years in 2003 to 229 per 10,000,000 person-years in 2010 [2, 3]. It has been well established that genetic and environmental factors, such as CHD family history, hypertension, diabetes mellitus, dyslipidemia, poor diet, advanced age and smoking habit, were associated with an increased risk of CHD [4].
CHD is associated with inflammation and considered as complex chronic diseases relating to multiple genes. And among these inflammatory cytokines, IL-6 is a circulating cytokine known as a regulation for inflammatory reaction, accelerates bone resorption [5] and plays a key role in atherogenesis and thrombosis [6, 7]. The IL-6 gene is mapped to chromosome 7 at p21 [8]. Vitro or vivo studies have identified single nucleotide polymorphisms (SNPs) in the promoter regions of the IL-6 gene as principle mediators for the circulation of mRNA and IL-6 levels and the status of inflammation [9, 10]. Several studies have reported the association between the IL-6 gene SNPs and CHD susceptibility in different populations [11–13]. However, these studies could not conclude a certain and definitive conclusion. Several meta-analyses [14, 15] were also carried out attempting to compromise the existing inconsistencies with elusive results. The SNPs in vitamin D receptor (VDR) were found to be a potential risk factor of CAD, which might be associated with a low level of vitamin D in CAD patients, but the exact mechanism underlying the influence of VDR polymorphisms on the pathogenesis of CHD was still unknown [16–18]. CHD risk was influenced by not only genetic factors, but also some environment factors and gene- environment interactions, and in all modifiable risk factors, tobacco smoking was an important environmental risk factor for CHD. But to date, the relationship of IL- 6 and VDR gene with CHD risk was not well known, and less study focused on the associations of gene- smoking interaction with CHD susceptibility. So the aim of this study was to investigate the impact of several SNPs within IL-6 and VDR gene, and additional gene-smoking interaction on CHD risk, based on a Chinese population.
RESULTS
A total of 1722 participants (880 males, 842 females) were selected, including 860 CHD patients and 862 control participants. The mean age of all participants was 63.8 ± 14.8 years. Table 1 shows the general characteristics in cases and controls. The mean of age and distributions of males and alcohol drinking were not significantly different between cases and controls. The means of TC, TG, LDL-C, BMI and WC, and the rate of smokers were higher in cases than controls. The mean of HDL-C was lower in cases than controls.
Table 1: General characteristics for all study participants in CHD cases and controls
Variables |
CHD cases (N=860) |
Controls (N=862) |
P-values |
|---|---|---|---|
Age (years) |
63.5±13.6 |
64.2±14.1 |
0.295 |
Males N (%) |
442 (51.4) |
438 (50.8) |
0.809 |
Drinking N (%) |
221(25.7) |
202(23.4) |
0.275 |
Smoke N (%) |
272 (31.6) |
192 (22.3) |
<0.001 |
WC (cm) |
86.5±15.2 |
84.1±15.8 |
<0.001 |
BMI (kg/m2) |
24.7±9.2 |
23.4±9.6 |
<0.001 |
TG (mmol/L) |
1.5±0.7 |
1.3± 0.8 |
<0.001 |
TC (mmol/L) |
4.8±1.2 |
4.2±1.3 |
<0.001 |
HDL-C (mmol/L) |
1.16±0.4 |
1.31±0.5 |
<0.001 |
LDL-C (mmol/L) |
2.51 ±0.72 |
2.71±0.76 |
<0.001 |
T2DM, N (%) |
145 (16.9) |
- |
- |
Hypertension, N (%) |
338 (39.3) |
- |
- |
WC, waist circumference; BMI, body mass index; N, number.
No significant difference in genotype frequencies from the Hardy–Weinberg equilibrium test was noted for any tested SNPs in the controls. The frequencies for the C allele of rs1800796 within IL-6 and T allele of rs2228570 within VDR gene are significantly higher in CHD cases than controls (30.8% vs20.3%, 29.0% vs19.5%). Logistic regression analysis showed that CHD risk was significantly higher in carriers with the C allele of rs1800796 within IL-6 gene than those with GG genotype (GC+ CC versus GG), adjusted OR (95%CI) =1.62 (1.19-2.23), and higher in carriers with T allele of rs2228570 within VDR gene than those with CC genotype (CT+ TT versus CC), adjusted OR (95%CI) = 1.68 (1.26-2.17). However, we did not find any direct association of the others SNPs in IL- 6 and VDR gene with CHD risk after covariates adjustment. (Table 2).
Table 2: Genotype and allele frequencies of 6 SNPs between case and control group
SNP |
Genotypes and alleles |
Frequencies N (%) |
OR (95%CI)* |
P- values |
P- values for HWE test in controls |
|
|---|---|---|---|---|---|---|
|
|
Control (n=862) |
Case (n=860) |
|
|
|
IL- 6 gene |
||||||
rs1800795-174G/C |
||||||
|
Co-dominant |
|
|
|
|
|
|
GG |
503 (58.4) |
450 (52.3) |
1.00 (ref) |
|
0.994 |
|
GC |
311 (36.1) |
338 (39.3) |
1.26 (0.82-1.85) |
0.287 |
|
|
CC |
48 (5.6) |
72 (8.4) |
1.50 (0.77-2.31) |
0.516 |
|
|
Dominant |
|
|
|
|
|
|
GG |
503 (58.4) |
450 (52.3) |
1.00 (ref) |
|
|
|
GC+CC |
359 (41.6) |
410 (47.7) |
1.34 (0.80-1.94) |
0.489 |
|
|
Allele, C (%) |
407 (23.6) |
482 (28.0) |
|
|
|
rs1800796-572G>C |
||||||
|
Co-dominant |
|
|
|
|
|
|
GG |
555 (64.4) |
427 (49.6) |
1.00 (ref) |
|
0.118 |
|
GC |
264 (30.6) |
337 (39.2) |
1.38 (1.10-1.77) |
0.0002 |
|
|
CC |
43 (5.0) |
96 (11.2) |
2.18 (1.49-3.07) |
<0.001 |
|
|
Dominant |
|
|
|
|
|
|
GG |
555 (64.4) |
427 (49.6) |
1.00 (ref) |
|
|
|
GC+CC |
307 (35.6) |
433 (50.4) |
1.62 (1.19-2.23) |
<0.001 |
|
|
Allele, C (%) |
350 (20.3) |
529 (30.8) |
|
|
|
rs1800797-597G/A |
||||||
|
Co-dominant |
|
|
|
|
|
|
GG |
494 (57.3) |
441 (51.3) |
1.00 (ref) |
|
0.128 |
|
GA |
306 (35.5) |
331 (38.5) |
1.21 (0.80-1.78) |
0.308 |
|
|
AA |
62 (7.2) |
88 (10.2) |
1.45 (0.83-2.10) |
0.672 |
|
|
Dominant |
|
|
|
|
|
|
GG |
494 (57.3) |
441 (51.3) |
1.00 (ref) |
|
|
|
GA+AA |
368 (42.7) |
419 (48.7) |
1.25 (0.80-1.87) |
0.356 |
|
|
Allele, A (%) |
430 (24.9) |
507 (29.5) |
|
|
|
VDR gene |
||||||
ApaI (rs7975232) |
||||||
|
Co-dominant |
|
|
|
|
|
|
GG |
499 (57.9) |
445 (51.7) |
1.00 (ref) |
|
0.104 |
|
GT |
302 (35.0) |
328 (38.1) |
1.16 (0.78-1.63) |
0.401 |
|
|
TT |
61 (7.1) |
87 (10.1) |
1.56 (0.71-2.44) |
0.532 |
|
|
Dominant |
|
|
|
|
|
|
GG |
499 (57.9) |
445 (51.7) |
1.00 (ref) |
|
|
|
GT+TT |
363 (42.1) |
415 (48.3) |
1.25 (0.76-1.89) |
0.482 |
|
|
Allele, T (%) |
424 (24.6) |
502 (29.2) |
|
|
|
FokI (rs2228570) |
||||||
|
Co-dominant |
|
|
|
|
|
|
CC |
564 (65.4) |
442 (51.4) |
1.00 (ref) |
|
0.254 |
|
CT |
260 (30.2) |
338 (39.3) |
1.47 (1.22-1.85) |
<0.001 |
|
|
TT |
38 (4.4) |
80 (9.3) |
2.12 (1.43-2.88) |
<0.001 |
|
|
Dominant |
|
|
|
|
|
|
CC |
564 (65.4) |
442 (51.4) |
1.00 (ref) |
|
|
|
CT+TT |
298 (34.6) |
418 (48.6) |
1.68 (1.26-2.17) |
<0.001 |
|
|
Allele, T (%) |
336 (19.5) |
498 (29.0) |
|
|
|
BsmI (rs1544410) |
||||||
|
Additive |
|
|
|
|
0.115 |
|
AA |
548 (63.6) |
484 (56.3) |
1.00 (ref) |
|
|
|
AG |
269 (31.2) |
301 (35.0) |
1.30 (0.90-1.79) |
0.213 |
|
|
GG |
45 (5.2) |
75 (8.7) |
1.50 (0.81-2.31) |
0.426 |
|
|
Dominant |
|
|
|
|
|
|
AA |
548 (63.6) |
484 (56.3) |
1.00 (ref) |
|
|
|
AG+GG |
314 (36.4) |
376 (43.7) |
1.36 (0.88-1.85) |
0.384 |
|
|
Allele, G (%) |
359 (20.8) |
451 (26.2) |
|
|
|
*Adjusted for gender, age, smoking and alcohol status, BMI and WC.
We also investigate the impact of the interaction among 6 SNPs within IL- 6 and VDR gene and gene- smoking interaction on CHD risk by using GMDR model. Table 3 summarized the results obtained from GMDR analysis. We found a significant two-locus model (p=0.0010) involving rs1800796 and smoking, indicating a potential interaction between rs1800796 and smoking on CHD risk. Overall, the cross-validation consistency of this two- locus model was 10/ 10, and the testing accuracy was 60.11%. We also conducted stratified analysis for the significant models in the GMDR analysis by using logistic regression. We found that current smokers with rs1800796- GC+CC genotype have the highest CHD risk, compared to never- smokers with rs1800796- GG within IL- 6 genotype, OR (95%CI) = 2.57 (1.74 -3.46), after covariates adjustment (Table 4).
Table 3: GMDR analysis on the best gene–gene and gene- smoking interaction models
Locus no. |
Best combination |
Cross-validation consistency |
Testing accuracy |
P-values |
|---|---|---|---|---|
Gene-gene interactions* |
||||
2 |
rs1800796 rs2228570 |
8/10 |
0.5399 |
0.0547 |
3 |
rs1800796 rs2228570 rs1800797 |
7/10 |
0.5399 |
0.1719 |
4 |
rs1800796 rs2228570 rs1800797 rs1544410 |
6/10 |
0.5399 |
0.3770 |
5 |
rs1800796 rs2228570 rs1800797 rs1544410 rs7975232 |
6/10 |
0.4958 |
0.4258 |
6 |
rs1800796 rs2228570 rs1800797 rs1544410 rs7975232 rs1800795 |
5/10 |
0.4958 |
0.6230 |
Gene- smoking interactions ** |
||||
2 |
rs1800796 Smoking |
10/10 |
0.6011 |
0.0010 |
3 |
rs1800796 rs2228570 Smoking |
8/10 |
0.5399 |
0.0547 |
4 |
rs1800796 rs2228570 rs1800797 Smoking |
7/10 |
0.5399 |
0.1719 |
5 |
rs1800796 rs2228570 rs1800797 rs1544410 Smoking |
7/10 |
0.4958 |
0.3770 |
6 |
rs1800796 rs2228570 rs1800797 rs1544410 rs7975232 Smoking |
6/10 |
0.4958 |
0.4258 |
7 |
rs1800796 rs2228570 rs1800797 rs1544410 rs7975232 rs1800795 Smoking |
4/10 |
0.4958 |
0.6230 |
*Adjusted for gender, age, smoking, drinking and BMI for gene- gene interaction analysis.
**Adjusted for gender, age, drinking and BMI for gene- smoking interaction analysis.
Table 4: Stratified analysis for gene- smoking interaction by using logistic regression
rs1800796 |
Smoking |
OR (95% CI)* |
P-values |
|---|---|---|---|
GG |
Never |
1.00 |
- |
GC+CC |
Never |
1.33(1.02 -1.76) |
0.032 |
GG |
Current |
1.46 (1.15-1.85) |
0.001 |
GC+CC |
Current |
2.57 (1.74 -3.46) |
<0.001 |
*Adjusted for gender, age, drinking and BMI.
DISCUSSION
In this study, we found that both the C allele of rs1800796 within IL-6 and T allele of rs2228570 within VDR gene were significantly associated with increased CHD risk. Several similar studies have been performed on association between IL-6 SNP and CHD risk; however these studies concluded inconsistent results. Gigante [19] indicate that IL6R haplotypes regulate the circulating levels of inflammatory biomarkers. But lack of association with the risk of CHD may be explained by the combined effect of SNPs with opposite effect on the CHD risk. A meta- analysis conducted by Song et al [20] suggested that the IL-6 -572G>C polymorphism may be linked with risk of CHD in a protective model. Another Chinese study [21] indicated that the -572G/C polymorphism in the IL-6 gene promoter region was involved in the pathogenesis and progression of CHD in the Han Chinese. But some other studies concluded different results with these studies. Tong et al [22] indicated that IL-6 SNPs were associated with increased CAD risk in the Chinese population, and may be useful predictive markers for CAD susceptibility. A meta- analysis [23] indicated that IL-6 gene −174 G/C polymorphism is associated with increased CHD risk among Asians. Another Chinese case- control study [24] also suggested that IL-6-572C/G polymorphism was associated with susceptibility to CHD after adjustment for other risk factors such as smoking, hypertension, hyperlipidema, obesity and alcohol drinking, they concluded that IL-6-572C/G polymorphism might be a potential risk factor for CHD in Chinese population. The potential mechanism for the association between IL-6-572C/G and CHD was not well known, but studies have indicated that inflammation plays an important role in the progression of atherosclerosis [25]. IL-6 functions as a multifunctional cytokine, playing a critical role in the inflammatory response in relation to the etiology of CHD [26].
The gene encoding the VDR is located at chromosome 12q and has 4 common polymorphisms. BsmI (rs1544410), ApaI (rs7975232) and FokI (rs2228570) were three common SNPs in VDR gene polymorphism. More and more recent attention has been focused on the possible role of VDR gene polymorphisms in the development of a range of diseases, including diabetes [27], as well as CAD [28–30]. Some previous studies also concluded inconsistent results on association between VDR gene and CHD risk. Pan et al [18] indicated that there was no significant difference in the genotype distribution or the allele frequencies of VDR FokI and BsmI between case and control groups in a Chinese population. Another study [31] also indicated that low vitamin D level was associated with an enhanced risk for incident CAD, but VDR genotypes did not show any association with either vitamin D levels or CAD. However the opposed results indicated that the haplotype comprising the minor allele of BsmI, major allele of ApaI and minor allele of TaqI of VDR (AAC) was associated with an increased risk of CAD in type 2 diabetes patients [30]. A meta- analysis [29] suggested that the Fok1 polymorphism may play a protective role in CAD, and the possible protective role in Apa1 CA genotype in CAD patients with T2DM still needed more studies to support. The Taq1 polymorphism is found to be associated with a significant increase in CAD risk based on our analysis. Moreover, increased risk in Apa1 variation in CAD patients without T2DM and Bsm1 variation in Caucasian group is also detected. But another meta- study [16] concluded that VDR FokI polymorphisms appear to be associated with CHD, but Apa1 and Taq1 was not associated with CHD risk. The SNPs in VDR were found to be a potential risk factor of CAD, which might be associated with a low level of vitamin D in CAD patients, but the exact mechanism underlying the influence of VDR polymorphisms on the pathogenesis of CAD is still unknown. Several frequent polymorphisms in the VDR gene have been reported to be associated with a variety of physiological and pathological phenotypes, including intrauterine and early postnatal growth, body weight and body height, as well as insulin secretion, insulin sensitivity, glucose tolerance and susceptibility to both type 1 and type 2 diabetes [32–34].
CHD susceptibility was influenced by both genetic and environmental factors, and previously several environmental factors associated with CHD have been reported, and in these risk factors, cigarette smoking has been suggested to play a crucial role in increasing the risk of CHD risk. In current study, the rate of smoking was higher in CHD cases than controls, so in this study we not only investigated gene- gene interaction on CHD, but also gene- environment interaction between SNPs and smoking. We found a significant interaction between rs1800796 and smoking on CHD risk, current smokers with rs1800796- GC or CC genotype have the highest CHD risk, compared to never- smokers with rs1800796- GG within IL-6 genotype.
There several limitations in our study. Firstly, more SNPs within IL-6 and VDR gene should been included in the analysis, not only for just 6 SNPs. Secondly, some others environmental risk factors should be investigated in the gene- environment interaction analysis. Thirdly, haplotype analysis should be performed in the future studies. Lastly, both the CHD and control groups were recruited from one hospital, and the controls were selected using convenient sampling. These selected controls could not represent the general population, because these people received physical examination routinely pay more attention to their health.
In conclusion, we found that the C allele of rs1800796 within IL-6 and T allele of rs2228570 within VDR gene, interaction between rs1800796 and smoking were all associated with increased CHD risk.
MATERIALS AND METHODS
Subjects
All study participants were recruited between March 2011 and July 2015 from the 2nd Affiliated Hospital of Wenzhou Medical University. The CHD patients were diagnosed by at least two experienced cardiologists and were confirmed by coronary angiography (>50 % diameter stenosis in at least one of the major coronary arteries) according to the World Health Organization criteria for the confirmation of CHD. The controls were randomly selected from those, who received physical examination in our hospital and 1:1 matched to cases on the basis of age (±3 years) and sex, all selected controls received electrocardiographic testing and coronary angiography, and those with abnormal electrocardiographic and coronary angiography profile were excluded. Participants with diabetes, hypertension, and others CHD related risks were excluded from the controls (Figure 1). Both the CHD cases and controls were unrelated Han Chinese population. Questionnaire investigation was conducted for all participants, and data on demographic information, clinical and biochemical index for all participants were obtained. Body weight, height and waist circumference (WC) were measured. Current cigarette smokers were those who self-reported smoking cigarettes at least once a day for 1 year or more, and still smoked at the time of the interview, and individuals with no history of cigarette smoking were considered as never smokers. Alcohol consumption was expressed as the sum of milliliters of alcohol per week from wine, beer, and spirits. Blood samples were collected from each participant in the morning after at least 8 hours of fasting. Informed consents were obtained from all participants.
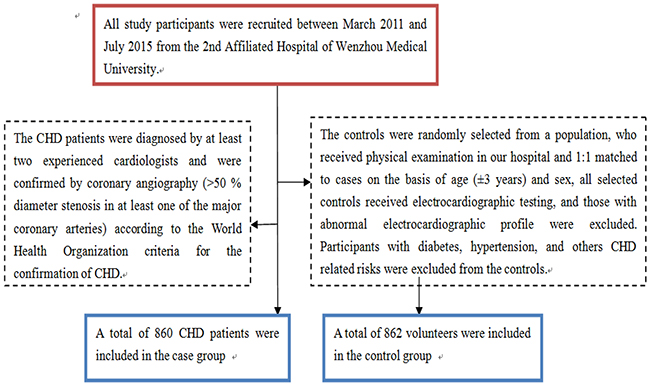
Figure 1: A flowchart on study population selection and exclusion.
Genomic DNA extraction and genotyping
The SNPs were selected based on the NCBI database (http://www.ncbi.nlm.nih.gov/projects/SNP), those SNPs with a minor allele frequency (MAF) of at least 5%, which have been reported in previous studies, but were not well studied, were selected for genotyping, including: rs1800796, rs1800797 and rs1800795 within IL-6 gene; rs2228570, rs1544410 and rs7975232 within VDR gene. Genomic DNA from participants was extracted from EDTA-treated whole blood, using the DNA Blood Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Genotyping for all 6 SNPs was performed by polymerase chain reaction (PCR) restriction fragment length polymorphism (PCR-RELP) method. The specific primers for the 6 SNPs listed in Table 5.
Table 5: Description and probe sequence for 6 SNPs used for PCR analysis
SNP ID |
Chromosome |
Functional consequence |
Major/ minor alleles |
Restriction enzyme |
Primer sequences |
IL- 6 gene |
|||||
rs1800795-174G /C |
7:22727026 |
Intron variant, upstream variant 2KB |
G/ C |
NlaIII |
F: 5’- tgacttcagctttactctttgt-3’ |
rs1800796-572G>C |
7:22726627 |
Intron variant, nc transcript variant, upstream variant 2KB |
G / C |
BsrBI |
F: 5’- gagacgccttgaagtaactg-3’ |
rs1800797-597G/A |
7:22726602 |
Intron variant, nc transcript variant, upstream variant 2KB |
G/ A |
FokI |
F: 5’- ctcctctaagtgggctgaag-3’ |
VDR gene |
|||||
FokI (rs2228570) |
12:47879112 |
Missense |
C/ T |
FokI |
F: GCACTGACTCTGGCTCTGAC |
BsmI (rs1544410) |
12:47846052 |
Intron variant, upstream variant 2KB |
A/ G |
FspI |
F: GGAGACACAGATAAGGAAATAC |
ApaI (rs7975232) |
12:47845054 |
Intron variant, upstream variant 2KB |
G/ T |
ApaI |
F: TGGGCACGGGGATAGAGAAG |
Genotyping for rs1800796, rs1800797 and rs1800795 within IL-6 gene: a 25 μl reaction mixture was amplified by PCR, containing 100 ng genomic DNA, 1.5 mM MgCl2, 0.2 mM dNTPs, 1 mM each primer, reaction buffer, and 0.625 U Taq polymerase. PCR conditions were as follows: initial denaturation at 94o C for 3 min, followed by 35 cycles of 94°C for 30 s, 56° C for 30 s (52° C for 30 s for rs1800795), 72° C for 20 s, and a final extension of 72° C for 5 min. Genotyping for rs2228570, rs1544410 and rs7975232 within VDR gene: The reaction volume was 25μl, containing Taq DNA polymerase, dNTPs mix and reaction buffer (Shanghai, China). A total of 0.5 μM of each primer, 2μl of template cDNA, 9μl 2.5* Mastermix, and 8.5 μl distilled water, (ddH2O) were used in all reactions (in triplicate). PCR conditions were as follows: denaturation 95 ° C, 15 s; annealing 60° C, 60 s; elongation 72° C, 45 s) consisted of 40 cycles to ensure a linear range of the PCR reaction.
Statistical analysis
All statistical analyses were performed using the SPSS 22.0 software package (SPSS Inc, Chicago) for Windows 7. Categorical variables were presented as absolute values and percentages, and continuous variables were expressed as mean ± standard deviation (SD). Student’s t test was used to compare continuous variables, while Chi-square test was used to compare categorical variables between cases and controls. Hardy-Weinberg equilibrium (HWE) examination was used by SNPstats (http://bioinfo.iconcologia.net/SNPstats). Generalized multifactor dimensionality reduction (GMDR) model was used to analyze the gene- gene interaction, some parameters including cross-validation consistency, the testing balanced accuracy and the sign test were calculated, a sign test or a permutation test (providing empirical p-values) for prediction accuracy can be used to measure the significance of an identified model. Logistic regression was performed to investigate association between 6 SNPs within IL-6 and VDR gene and CHD, and additional stratified analysis for gene- smoking interaction on CHD risk. Two sided test with P < 0.05 was considered statistically significant.
ACKNOWLEDGMENTS
We thank the investigators and staffs from the 2nd Affiliated Hospital of Wenzhou Medical University, and all the partners and staffs who help us in the process of this study. The authors disclosed receipt of the following financial supports for the research and authorship of this paper: Research Project of Wenzhou Municipal Science and Technology Bureau, No. Y20160029.
CONFLICTS OF INTEREST
There is no conflicts of interest.
FUNDING
Research Project of Wenzhou Municipal Science and Technology Bureau (Y20160029).
REFERENCES
1. Lopez AD, Mathers CD, Ezzati M, Jamison DT, Murray CJ. Global and regional burden of disease and risk factors, 2001: systematic analysis of population health data. Lancet. 2006; 367:1747–1757.
2. Daka B, Olausson J, Larsson CA, Hellgren MI, Råstam L, Jansson PA, Lindblad U. Circulating concentrations of endothelin-1 predict coronary heart disease in women but not in men: a longitudinal observational study in the Vara-Skovde Cohort. BMC Cardiovasc Disord. 2015; 15:146.
3. Hu Z, Yuan X, Rao K, Zheng Z, Hu S. National trend in congenital heart disease mortality in China during 2003 to 2010: a population-based study. J Thorac Cardiovasc Surg. 2014; 148:596–602.e1.
4. Lusk CM, Dyson G, Clark AG, Ballantyne CM, Frikke-Schmidt R, Tybjærg-Hansen A, Boerwinkle E, Sing CF. Validated context-dependent associations of coronary heart disease risk with genotype variation in the chromosome 9p21 region: the Atherosclerosis Risk in Communities study. Hum Genet. 2014; 133:1105–1116.
5. Ishimi Y, Miyaura C, Jin CH, Akatsu T, Abe E, Nakamura Y, Yamaguchi A, Yoshiki S, Matsuda T, Hirano T. IL-6 is produced by osteoblasts and induces bone resorption. J Immunol. 1990; 145:3297–3303.
6. Ikeda U, Ito T, Shimada K. Interleukin-6 and acute coronary syndrome. Clin Cardiol. 2001; 24:1574–1583.
7. Yudkin JS, Kumari M, Humphries SE, Mohamed-Ali V. Inflammation, obesity, stress and coronary heart disease: is interleukin-6 the link? Atherosclerosis. 2000; 148:209–214.
8. Ray A, LaForge KS, Sehgal PB. On the mechanism for efficient repression of the interleukin-6 promoter by glucocorticoids: enhancer, TATA box, and RNA start site (Inrmotif) occlusion. Mol Cell Biol. 1990; 10:5736–5746.
9. Yeh KY, Li YY, Hsieh LL, Chen JR, Tang RP. The -174 G/C polymorphism in interleukin-6 (IL-6) promoter region is associated with serum IL-6 and carcinoembryonic antigen levels in patients with colorectal cancers in Taiwan. J Clin Immunol. 2010; 30:53–59.
10. Pereira DS, Garcia DM, Narciso FM, Santos ML, Dias JM, Queiroz BZ, Souza ER, Nóbrega OT, Pereira LS. Effects of 174 G/C polymorphism in the promoter region of the interleukin-6 gene on plasma IL-6 levels and muscle strength in elderly women. Braz J Med Biol Res. 2011; 44:123–129.
11. Basso F, Lowe GD, Rumley A, McMahon AD, Humphries SE. Interleukin-6 -174G[C polymorphism and risk of coronary heart disease in West of Scotland coronary prevention study (WOSCOPS). Arterioscler Thromb Vasc Biol. 2002; 22:599–604.
12. Humphries SE, Luong LA, Ogg MS, Hawe E, Miller GJ. The interleukin-6 -174 G/C promoter polymorphism is associated with risk of coronary heart disease and systolic blood pressure in healthy men. Eur Heart J. 2001; 22:2243–2252.
13. Wei YS, Lan Y, Liu YG, Tang RG, Lan JS. Relationship between interleukin-6 gene polymorphism and coronary heart disease and its effect on plasma lipid levels. Zhongguo Wei Zhong Bing Ji Jiu Yi Xue. 2006; 18:233–236.
14. Yin YW, Hu AM, Sun QQ, Liu HL, Wang Q, Zeng YH, Xu RJ, Hou ZZ, Zhang ZD, Zhang SJ, Zhou MQ. Association between interleukin-6 gene −174 G/C polymorphism and the risk of coronary heart disease: a meta-analysis of 20 studies including 9619 cases and 10,919 controls. Gene. 2012; 503:25–30.
15. Zheng GH, Chen HY, Xiong SQ. Polymorphisms of 2174G.C and 2572G.C in the interleukin 6 (IL-6) gene and coronary heart disease risk: a meta-analysis of 27 research studies. PLoS One. 2012; 7:e34839.
16. He L, Wang M. Association of vitamin d receptor-a gene polymorphisms with coronary heart disease in Han Chinese. Int J Clin Exp Med. 2015; 8:6224–6229.
17. Abu el Maaty MA, Hassanein SI, Sleem HM, Gad MZ. Vitamin D receptor gene polymorphisms (TaqI and ApaI) in relation to 25-hydroxyvitamin D levels and coronary artery disease incidence. J Recept Signal Transduct Res. 2015; 35:391–395.
18. Pan XM, Li DR, Yang L, Wang EY, Chen TY, Liu YJ, Liu M, Liao ZG. No association between vitamin D receptor polymorphisms and coronary artery disease in a Chinese population. DNA Cell Biol. 2009; 28:521–525.
19. Gigante B, Strawbridge RJ, Velasquez IM, Golabkesh Z, Silveira A, Goel A, Baldassarre D, Veglia F, Tremoli E, Clarke R, Watkins H, Hamsten A, Humphries SE, de Faire U. Analysis of the role of interleukin 6 receptor haplotypes in the regulation of circulating levels of inflammatory biomarkers and risk of coronary heart disease. PLoS One. 2015; 10:e0119980.
20. Song C, Liu B, Yang D, Diao H, Zhao L, Lu Y, Yu Y, Guo Z, Zhang J, Liu J, Zaho Z, Zhang X. Association between interleukin-6 gene 2572G>C polymorphism and coronary heart disease. Cell Biochem Biophys. 2015; 71:359–365.
21. Jia X, Tian Y, Wang Y, Deng X, Dong Z, Scafa N, Zhang X. Association between the interleukin-6 gene -572G/C and -597G/A polymorphisms and coronary heart disease in the Han Chinese. Med Sci Monit. 2010; 16:CR103–108.
22. Tong Z, Li Q, Zhang J, Wei Y, Miao G, Yang X. Association between interleukin 6 and interleukin 16 gene polymorphisms and coronary heart disease risk in a Chinese population. J Int Med Res. 2013; 41:1049–1056.
23. Yin YW, Hu AM, Sun QQ, Liu HL, Wang Q, Zeng YH, Xu RJ, Hou ZZ, Zhang ZD, Zhang SJ, Zhou MQ. Association between interleukin-6 gene −174 G/C polymorphism and the risk of coronary heart disease: a meta-analysis of 20 studies including 9619 cases and 10,919 controls. Gene. 2012; 503:25–30.
24. Fan WH, Liu DL, Xiao LM, Xie CJ, Sun SY, Zhang JC. Coronary heart disease and chronic periodontitis: is polymorphism of interleukin-6 gene the common risk factor in a Chinese population? Oral Dis. 2011; 17:270–276.
25. Ross R. Atherosclerosis–an inflammatory disease. N Engl J Med. 1999; 340:115–126.
26. Plutzky J. Inflammatory pathways in atherosclerosis and acute coronary syndromes. Am J Cardiol. 2001; 88:10K–15K.
27. Rivera-Leon EA, Palmeros-Sanchez B, Llamas-Covarrubias IM, Fernandez S, Armendariz- Borunda J, Gonzalez-Hita M, Bastidas-Ramirez BE, Zepeda-Moreno A, Sanchez-Enriquez S. Vitamin-D receptor gene polymorphisms (TaqI and ApaI) and circulating osteocalcin in type 2 diabetic patients and healthy subjects. Endokrynol Pol. 2015; 66:329–333.
28. Abu El Maaty MA, Hassanein SI, Gad MZ. Genetic variation in vitamin D receptor gene (Fok1:rs2228570) is associated with risk of coronary artery disease. Biomarkers. 2016; 21:68–72.
29. Lu S, Guo S, Hu F, Guo Y, Yan L, Ma W, Wang Y, Wei Y, Zhang Z, Wang Z. The associations between the polymorphisms of vitamin D receptor and coronary artery disease: a systematic review and meta-analysis. Medicine (Baltimore). 2016; 95:e3467.
30. Ferrarezi DA, Bellili-Muñoz N, Dubois-Laforgue D, Cheurfa N, Lamri A, Reis AF, Le Feuvre C, Roussel R, Fumeron F, Timsit J, Marre M, Velho G. Allelic variations of the vitamin D receptor (VDR) gene are associated with increased risk of coronary artery disease in type 2 diabetics: the DIABHYCAR prospective study. Diabetes Metab. 2013; 39:263–270.
31. Shanker J, Maitra A, Arvind P, Nair J, Dash D, Manchiganti R, Rao VS, Radhika KN, Hebbagodi S, Kakkar VV. Role of vitamin D levels and vitamin D receptor polymorphisms in relation to coronary artery disease: the Indian atherosclerosis research study. Coron Artery Dis. 2011; 22:324–332.
32. Ferrarezi DA, Bellili-Munoz N, Nicolau C, Cheurfa N, Guazzelli IC, Frazzatto E, Velho G, Villares SM. Allelic variations in the vitamin D receptor gene, insulin secretion and parents’ heights are independently associated with height in obese children and adolescents. Metabolism. 2012; 61:1413–1421.
33. Jain R, von Hurst PR, Stonehouse W, Love DR, Higgins CM, Coad J. Association of vitamin D receptor gene polymorphisms with insulin resistance and response to vitamin D. Metabolism. 2012; 61:293–301.
34. Reis AF, Hauache OM, Velho G. Vitamin D endocrine system and the genetic susceptibility to diabetes, obesity and vascular disease. A review of evidence. Diabetes Metab. 2005; 31:318–325.

