INTRODUCTION
Medulloblastoma (MB) is the most common type of malignant solid tumors in the pediatric brain, which accounts for 20% of all pediatric tumors. It occurs in the external granular layer (EGL) of the cerebellum and may undergo metastasis (reviewed in [1]). Based on recent transcriptional profiling, MB can be classified into four subgroups: WNT subgroup, Sonic Hedgehog (SHH) subgroup, Group 3, and Group 4 [2]. The WNT subgroup accounts for ~10% of all MB and usually has good prognosis; this subgroup involves mutations in genes including TP53, CTNNB1, AXIN1, SMARCA4, and CREBBP. The SHH subgroup accounts for up to 30% of MB and contrarily shows poor prognosis. Loss of function mutations are found in genes PTC1/2 and SUFU whereas gain of function mutations are found in SMO and GLI1/2; furthermore, Notch and PI3K signaling (IGFRII) are activated in the SHH subgroup. Group 3 accounts for ~25% of all MB and carries no TP53 mutations while showing poor prognosis, immature histopathology, and highly metastatic property. Group 4 accounts for ~35% of all MB; similar to Group 3, Group 4 carries no TP53 mutations while mutations are often identified in genes including OTX2, N-MYC, FST, and CDK6.
The first MB model was SHH receptor Ptc1 heterozygote (+/−) mice which developed MB with low penetrance (< 7.4%) [3]. A conditional knockout mouse model for Ctnnb1, a critical component of Wnt signaling, was generated to reveal Zic1+ precursors in the dorsal brainstem as cells of origin of the WNT subtype MB [4]. In addition, a concomitant knockout mouse model for Tp53 and Wnt signaling resulted in MB that recapitulated the human MB subtype [4]. Recently, a mouse model for the most aggressive subgroup of human MB could be developed by enforced expression of Myc in Tp53-deficient cerebellar neuronal progenitors [5].
Retinoblastoma protein (Rb) exhibits tumor suppressive activity by binding and repressing a cell cycle activator, E2F (reviewed in [6]). Mutations of the Rb gene are rarely found in samples of human MB patients [7], and Rb knockout mice are predisposed primarily to the development of pituitary cancer rather than MB [8, 9]. However, simultaneous loss of function of Rb and Tp53, specifically in the EGL, had primarily shown typical MB development, strongly arguing that these genes and/or the signaling pathways regulated by these genes play critical roles in MB development [7]. The CDKN2A locus produces alternatively-spliced genes including p14ARF and p16Ink4a which up-regulate TP53 function by binding MDM2 or inhibit cyclin-dependent kinase 4 and 6 (CDK4 and CDK6) through the attenuation of Rb phosphorylation, respectively. The loss of function deletions of p14ARF and p16Ink4a have been well characterized in diverse types of tumors (reviewed in [10, 11]). The gene p15Ink4b, on the other hand, though it is encoded by a related locus CDKN2B and displays high similarity to p16Ink4a at the amino sequence level (~85%), remains unclear in various tumorigenic processes.
During the last decade or so, several zebrafish cancer models were successfully generated, which recapitulated human cancers such as leukemia, neuroblastoma, and melanoma [12]. In an unprecedented way, the identification of novel cancer signaling pathways and visualization of pathological processes has been made possible by advances in optical clarity of zebrafish for high resolution imaging, chemical screening using whole animals, and genetic manipulations to generate mutants using genome editing tools such as Transcription activator-like effector nucleases (TALEN) and CRISPR/Cas9, as well as transgenic animals using Tol2 system and I-SceI meganuclease [13–15]. Despite these advancements, a zebrafish model for the most frequently occurring pediatric tumor, MB, is currently unavailable. Very recently, zebrafish was employed in somatic inactivation of rb1 using TALEN to evaluate gene candidacy as a tumor suppressor [16]; however, the tumor types generated were not identified in detail, thus evaluating the interaction of two or more candidate tumor suppressor genes using somatic inactivation has not been reported yet, thus far.
In this study, we developed zebrafish cancer models by TALEN-mediated somatic inactivation of tumor suppressor genes with high efficiency. While cdkn2a/b gene inactivation accelerated development of Malignant Peripheral Nerve Sheath Tumors (MPNSTs) by TALEN injection in tp53 e7/e7 mutation background, TALEN-mediated somatic inactivation of rb1, which is a potential downstream target of CDKN2A/B, induced distinct brain tumors in the tp53 e7/e7 mutation background. Using RNA sequencing analysis together with histopathology and immunohistochemistry, we have demonstrated that brain tumors induced by rb1 somatic inactivation have a molecular feature of MB-like primitive neuroectodermal tumors (PNETs).
RESULTS
Somatic inactivation of cdkn2a/b gene by the injection of TALEN mRNA leads to MPNSTs in F0 founder tp53 e7/e7 mutant zebrafish
Synteny analysis demonstrated that zebrafish INK4a/ARF/INK4b locus was partially disrupted despite conservation of the adjacent gene block order (Supplementary Figure 1). Hence, zebrafish CDKN2A/B is the only encoded protein comparable to the three unique p19ARF, p16INK4b, and p15INK4b proteins which are expressed from INK4a/ARF/INK4b locus in humans. To investigate the role of cdkn2a/b in tumorigenesis of zebrafish, we performed genetic inactivation by TALEN-mediated genome editing. Two different TALENs targeting the first exon of zebrafish cdkn2a/b gene (designated as cdkn2a/b-TALENs1 and cdkn2a/b-TALENs2, respectively) were constructed and followed by microinjection of in vitro synthesized TALEN mRNAs into one-cell stage zebrafish embryos (Figure 1A). After 3–4 months, founder zebrafish were successfully obtained which transmitted germline mutations to F1 progeny. With the injection of cdkn2a/b-TALENs1 mRNA, eleven independent F1 heterozygous zebrafish were recovered in which seven different alleles (6 frameshifts and 1 in-frame) were mutated in the cdkn2a/b gene of zebrafish. After injection of cdkn2a/b-TALENs2 mRNA, sixteen independent F1 heterozygous zebrafish were recovered in which nine different alleles (4 frameshifts and 5 in-frames) were mutated in cdkn2a/b (Figure 1B). Homozygous cdkn2a/b mutant embryos that were generated by incrossing F1 heterozygous zebrafish (4bp deletion allele induced by cdkn2a/b-TALENs2) could not express cdkn2a/b transcript, which was quantified by real time RT-PCR with 5 days old embryos (Supplementary Figure 2).
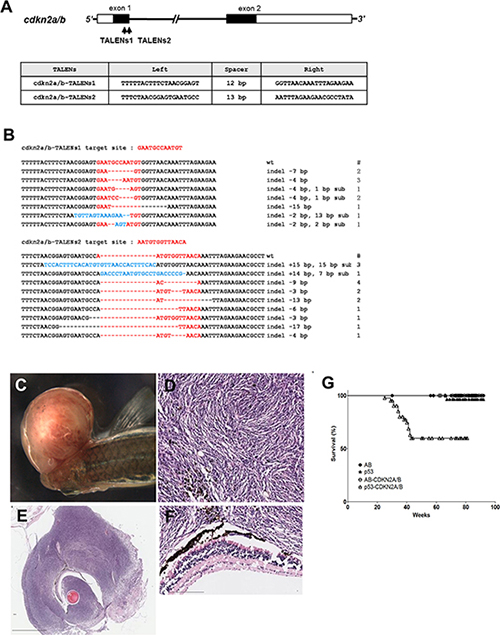
Figure 1: Somatic inactivation of cdkn2a/b gene by the injection of TALENs mRNA leads to MPNSTs in F0 founder zebrafish. (A) The targeted regions of two different TALENs (cdkn2a/b TALENs-1 and cdkn2a/b TALENs-2) are located in exon 1 of cdkn2a/b gene. The binding sequences of TALENs for cdkn2a/b gene are also shown. (B) By the injection of cdkn2a/b TALENs-1 mRNA, 7 different germline mutated alleles of cdkn2a/b gene were recovered at F1 generation. And 9 different alleles were also recovered at F1 generation after injection of cdkn2a/b TALENs-2 mRNA. (C) The large mass of tumor was observed in adjacent to eyes of the cdkn2a/b TALENs mRNA injected tp53 mutant zebrafish at 5.5 month post fertilization. (D–F) Histological image of cross sections of tumor tissue was visualized with Hematoxylin-Eosin staining. (D) High magnification image of tumor tissue shows the herringbone pattern that is a typical feature of malignant peripheral nerve sheath tumor. (F) There is no evident abnormality in the neuroepithelial layer of retina adjacent to tumor tissue. (G) Kaplan-Meier survival representation of wild type (AB), tp53 mutant, and cdkn2a/b TALENs mRNA injected AB or tp53 mutant zebrafish. The cdkn2a/b TALENs mRNA injected tp53 mutant zebrafish had only started to exhibit death with tumor bearing as early as about 5 months. Other adult zebrafish such as tp53 mutant and cdkn2a/b TALENs mRNA injected AB zebrafish did not show early death with tumor bearing. All data was acquired from two independent experiments. Scale bars: 2 mm (E) and 100 μm (D and F).
During analysis of cdkn2a/b mutants, we encountered an unexpected observation that adult tp53 mutant zebrafish via one-cell stage injection of cdkn2a/b-TALEN mRNA could develop tumor mass, mainly occurring adjacent to the eyes as early as 6 to 7 months of age (Figure 1C, Table 1). Previous studies have reported that malignant peripheral nerve sheath tumors (MPNSTs) spontaneously developed in tp53e7/e7 mutant zebrafish at 28% of tumor incidence rate by around 14 months of age [17, 18]. Similarly, our results showed that tp53 mutant zebrafish could develop MPNSTs at an average of 66 weeks post fertilization (wpf) with a tumor incidence frequency of about 15 % (Table 1). Under the same conditions, injection of cdkn2a/b-TALEN mRNA led to an increase in tumor incidence frequency up to about 39% and accelerated tumor onset as early as 35 to 37 wpf in tp53e7/e7 mutant zebrafish (Table 1). Kaplan–Meier survival curves also represented that cdkn2a/b-TALEN mRNA injection lead tp53e7/e7 mutant zebrafish to early death with tumor bearing compared to non injected mutant zebrafih, remarkably (Figure 1G, P < 0.0001).
Table 1: The frequency and occurrence regions of tumors induced by somatic gene inactivation with cdkn2a/b or rb1 TALENs in adult zebrafish
TALENs and genetic background |
Average age of appearance of tumors (weeks) |
Tumor frequency (%) |
Tumor occurrence regions and types |
|||
|---|---|---|---|---|---|---|
Ocular MPNSTs |
Abdominal MPNSTs |
MB-like PNETs |
Mixtures of MPNSTs & MB-like PNETs |
|||
WT a |
- |
0/39 (0) |
- |
- |
- |
- |
tp53 mutant b |
66 |
5/34 (14.7) |
1 |
1 |
- |
- |
WT-cdkn2a/b TALENs1 |
- |
0/26 (0) |
- |
- |
- |
- |
tp53 mutant -cdkn2a/b TALENs1c |
35 |
9/23 (39.1) |
6 |
3 |
- |
- |
WT-cdkn2a/b TALENs2 |
- |
0/29 (0) |
- |
- |
- |
- |
tp53 mutant-cdkn2a/b TALENs2d |
37 |
7/19 (36.8) |
4 |
3 |
- |
- |
WT-rb1 TALENs1 |
32 |
7/30 (23.3) |
- |
- |
7 |
- |
tp53 mutant-rb1 TALENs1e |
29 |
19/33 (57.5) |
1 |
1 |
15*1 |
2 |
tp53 mutant-cdkn2a/b TALENs2 &-rb1 TALENs1f |
31 |
20/42 (47.6) |
4 |
1 |
8 |
7*2 |
Tumor frequency was measured at from 20 until 90 weeks post fertilization (wpf) except for cdkn2a/b TALENs2 and rb1 TALENs1 double injection into tp53 mutant, which was measured for 60 wpff.a wild type zebrafish is Oregon AB genetic background.b Homozygous tp53e7/e7, tp53 M214K missense mutation in the p53 DNA binding domain preventing activation of p53.c, d, e Adult homozygous tp53e7/e7 zebrafish injected with about 200 pg of TALENs mRNAs for cdkn2a/b TALENs1, cdkn2a/b TALENs2, and rb1 TALENs1, respectively. Data were obtained by two independent injection experiments.*1 Actually, 8 of 17 adult zebrafish with tumors in brain region were confirmed with histopathological methods.*2 4 of 7 brain tumor-bearing zebrafish had ocular MPNSTs, and 3 of 7 brain tumor-bearing zebrafish had abdominal MPNSTs. Abbreviations: MPNST (Malignant peripheral nerve sheath tumors), MB (Medulloblstoma), and PNET (Primitive neuroectodermal tumors).
Histopathological and immunohistochemical analysis of MPNSTs induced by somatic inactivation of cdkn2a/b gene in tp53e7/e7 mutant zebrafish
Histopathological analysis by H & E staining of tumor tissues from cdkn2a/b-TALEN mRNA injected tp53e7/e7 mutant zebrafish showed that spindle-shaped hyperchromatic nuclei were densely arranged in wave-like manner (Figure 1D, 1E). There was no evident abnormality in the neuroepithelial layer of retina adjacent to the tumor tissue (Figure 1F); also herringbone pattern of tumor tissue could be observed in Figure 1E as a typical feature characteristic of MPNSTs. We further analyzed tumor tissues from cdkn2a/b-TALEN mRNA injected tp53e7/e7 mutant zebrafish with immunostaining methods.
Immunostaining with anti-phospho-Histone3(H3) antibodies demonstrated that tumor tissue of cdkn2a/b TALEN mRNA injected tp53e7/e7 mutant zebrafish had highly mitotic cells (Figure 2A, 2A’). As phospho-H3 positive mitotic cells were more evident in the outer regions, it appeared that tumor tissues had outgrown radially from the tumor. To discern neural and/or glial cell origin, tumor tissues were stained with anti-HuC/D and Zrf-1 antibodies. Despite clear staining in normal retinal layers near the tumoral tissue (inset box in Figure 2B’ and Figure 2C’), anti-HuC/D positive neuronal signatures and Zrf-1 positive glial characteristics were scarcely detected in the dense cellular region of tissue (Figure 2B, 2B’, 2C, and 2C’). Despite limited diagnostic utility and variable staining (of about 50–90% of tumors) [19], S100 has traditionally been regarded as the best marker for human MPNST. Immunostaining with epithelioid MPNST marker, S100β, showed strong expression in spindle-shaped tumor cells (Figure 2D, 2D’) while the mesenchymal marker, Vimentin, also showed extensive staining in tumor tissues of cdkn2a/b TALEN mRNA injected tp53e7/e7 mutant zebrafish (Figure 2E, 2E’). These data shows that TALEN-mediated somatic gene inactivation would be a useful strategy for tumor induction in zebrafish; targeted somatic inactivation of cdkn2a/b genes with TALENs efficiently induced early onset of MPNSTs with higher rate of incidence in tp53e7/e7 mutant zebrafish.
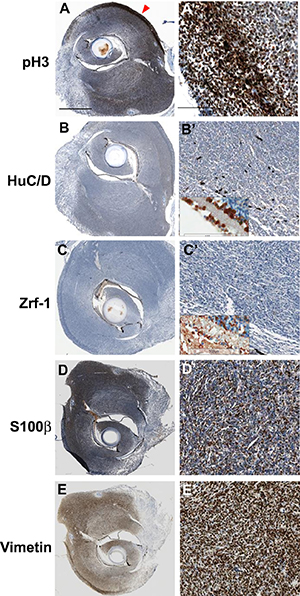
Figure 2: Immunohistochemical analysis of tumors from cdkn2a/b TALENs mRNA injected tp53 mutant zebrafish. (A) (A’) Immunostaining with anti- phospho-Histone 3 antibidy. The tumor tissue from cdkn2a/b TALENs mRNA injected tp53 mutant zebrafish is highly mitotic. The phospho-Histone 3 positive mitotic cells were more evident in the outer region of tumor tissue (red arrowhead in A). (B) (B’) Neuronal signature that was visualized by immunostaining with anti-HuC/D antibody was scarcely detected in densely cellular region of tumor tissue. (C) (C’) Immunostaining with zrf-1 which detect zebrafish GFAP protein. The tumor tissue induced with cdkn2a/b TALENs mRNA also did not show any glial characteristics. Normal staining of anti-HuC/D and zrf-1 antibody could be observed in retinal layers near to tumor tissues (Inset box in B’ and C’). (D) (D’) Epithelioid MPNST marker, S100β, is strongly expressed in tumor tissue. (E) (E’) Mesenchymal marker, Vimentin, is also positively stained in tumor tissue. Scale bars: 1 mm (A) and 100 μm (A’).
Somatic inactivation of rb1 by TALEN mRNA injection induced tumors in head regions of F0 founder zebrafish
As was introduced above, zebrafish cdkn2a/b gene is a unique orthologue for the human INK4a/ARF/INK4b locus. According to reports, zebrafish CDKN2A/B may regulate CDK4/6-mediated phosphorylation of retinoblastoma (Rb) family proteins given that p15INK4b and p16INK4a maintain Rb family proteins in a hypophosphorylated state to promote E2F binding and G1 cell cycle arrest [11]. Thus, it seemed worthwhile to examine how rb1 function in tumorigenesis compared to that of cdkn2a/b in zebrafish.
To examine the role of rb1 in tumorigenesis of zebrafish, TALENs were designed to target the first exon of zebrafish rb1 (designated as rb1-TALENs1). Somatic gene inactivation was performed using microinjection of rb1-TALENs1 mRNA into one-cell stage zebrafish embryos (Figure 3A). After injection, 13 independent germline-transmitted heterozygous zebrafish lines with frameshift mutations could be obtained in the F1 generation. Among them, 4 different deletion and 1 insertion mutated alleles of rb1 were successfully recovered (Figure 3B). Homozygous rb1 mutant larvae were generated by incrossing heterozygous F1 zebrafish with same frameshift mutated alleles. Because the expression of rb1 gene was scarcely detected compared to wild type embryos at 5 dpf, homozygous rb1 mutant had no functional Rb1 protein (Supplementary Figure 2). Homozygous mutants exhibited flattened swimming bladder, and enlarged liver at 7 day post fertilization (dpf) and did not survive until 10 dpf (Supplementary Figure 3).
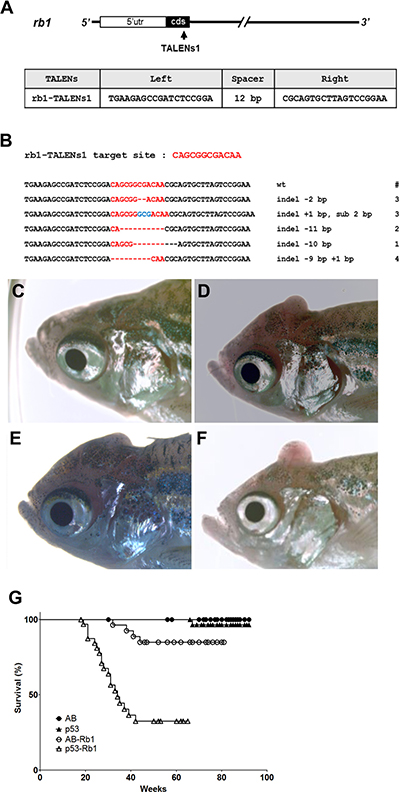

Figure 3: Somatic inactivation of rb1 by the injection of rb1-TALENs mRNA leads to medulloblastoma like PNETs in F0 founder zebrafish. (A) The targeted region of rb1-TALENs is located in 1st exon of zebrafish rb1 locus. The binding sequences of TALENs for rb1 are indicated. (B) After the injection of rb1-TALENs mRNA, 4 different deletion and 1 insertion germline mutated alleles of rb1 were successfully recovered at F1 generation. Hence, 13 independent lines which have frameshift mutation in rb1 could be obtained. (C–F) Morphological appearance of tumor bearing in head region of 5 month old zebrafish was presented. Protruding tumor mass were observed in rb1-TALENs mRNA injected zebrafish. The same aged wild type (C) and mutant (D–F) zebrafish are shown for comparison. (G) Kaplan-Meier survival representation of wild type, tp53 mutant, and rb1-TALENs injected wild type or tp53 mutant zebrafish. At about 6 month post fertilization, zebrafish which were injected with rb1-TALENs mRNA at one cell stage had started to die with tumors in the head region in wild type adult zebrafish. The tumor incidence which was induced by the injection of rb1-TALENs mRNA was more evident in tp53 mutant zebrafish. Adult zebrafish that showed abnormal swimming behavior started to die with tumor in head regions as early as about 20 weeks after injection of rb1-TALENs mRNA into one cell stage embryos.
Similar to cdkn2a/b inactivation of F0 tp53e7/e7 mutants, TALEN-mediated somatic inactivation of rb1 led to early onset of brain tumors in adult zebrafish. At around 4 to 5 months post fertilization, adult zebrafish injected at one-cell stage often displayed abnormal behaviors such as whirling or screwed swimming (data not shown). Additionally, tumor masses were visible starting from the posterior head region and protruding across the skull (Figure 3D–3F). The average age of tumor incidence was 7.5 months after birth, and 7 of 30 (23.3%) wild type zebrafish injected with rb1-TALEN mRNA at one-cell stage died bearing tumors in head regions (Table 1). Tumor induction was significantly accelerated by rb1-TALENs in the tp53e7/e7 mutant background. 19 of 33 (57.5 %) of rb1-TALEN mRNA injected tp53e7/e7 mutant zebrafish displayed tumors mainly in head regions while the average age of tumor incidence advanced about 1 month sooner than wild type background (Table 1). The Kaplan–Meier survival plot also showed decreased survival rate of tp53e7/e7 mutants compared to wild type zebrafish injected with rb1 TALEN mRNA at one cell stage (Figure 3G, P < 0.0001).
Targeted inactivation of rb1 resulted in the early onset of brain tumors in zebrafish with significantly increased frequency of tumor incidence. These data also demonstrated the effectiveness of tumor induction with rb1-TALEN mediated somatic gene inactivation.
Zebrafish tumors induced by somatic inactivation of rb1 displayed characteristics of medulloblastoma-like PNETs
H & E staining was performed to analyze tumor tissues from rb1-TALEN mRNA injected tp53e7/e7 mutant zebrafish. As reported previously [20] and shown in Figure 4A, the majority of H & E staining positive nuclei were located in the cerebellum of zebrafish brain. Normal cerebellar granule cells possess a unique feature characterized by small rounded nuclei. The stains showed nuclei gathered densely in the cerebellum granular layer (Figure 4B). Tumors from rb1-TALENs mRNA injected tp53e7/e7 mutant zebrafish mainly arose in the cerebellum and encompassed a majority of the medulla, hypothalamus, and brainstem; resulting in abnormal brain structures (Figure 4C, 4E, 4G). Tumor cells visualized with H & E staining exhibited essentially similar appearance among the different samples. Tumor tissues were composed of highly dense tumor cells with hyperchromatic and wedged nuclei while definite Homer-Wright like structures were also detected in most dissected sections (white arrows in Figure 4D, 4F, 4H). In some sections, we observed that partial tumor cells could spread and infiltrate into adjacent normal tissue (black arrowheads in Figure 4F).
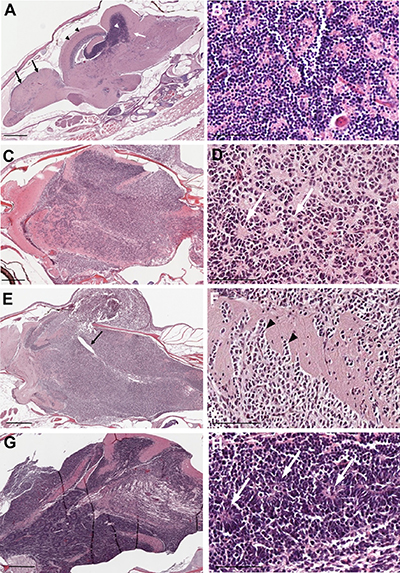
Figure 4: Histopathology of tumors from rb1-TALENs injected tp53 mutant zebrafish. (A) Sagittal section image of H & E staining of wild type zebrafish at 5 month post fertilization. Black arrows and arrow heads indicate forebrain and optic tectum, respectively. Cerebellum and cerebellar granule cells are indicated by white arrows and arrow heads, respectively. (B) High magnitude image of A. Normal cerebellar granule cells have a unique feature that is characterized with small rounded nuclei. (C–H) Sagittal section images of H & E staining of tumors from rb1-TALENs injected tp53 mutant zebrafish. (C) Tumors were mainly arising in cerebellum, medulla, and brainstem. (D) High magnitude image of C. Highly cellular tumor cells with wedged nuclei were observed. Rosette like structures was marked by white arrows. (E) Tumors were mainly arising in cerebellum encompassed a majority of medulla, hypothalamus, and brainstem. The forth ventricle which were surrounded with tumor cells were marked by black arrows. (F) High magnitude image of E. Infiltration of tumor cells was indicated by black arrowheads in dorsal brainstem. (H) High magnitude image of G. Homer-Wright rosettes which are seen in PNETs or medulloblastomas were observed distinctly (white arrows). Anterior is left of all images. Scale bars: 500 μm (A, C, E and G), and 50 μm (B, D, F and H).
Two independent brain tumor tissues used as the serial sections in Figure 4C and 4E were further analyzed via immunohistochemistry for tumor lineage. About 50% of the tumor cells of rb1-TALENs injected tp53e7/e7 mutant zebrafish displayed highly mitotic features (Figure 5B). We dismissed the possibility of glioma given that Zrf-1 positive glial characteristics were scarcely detected in the densely cellular region of rb1-TALEN induced brain tumor tissue (Figure 5D). Rather, the postmitotic neuronal marker, HuC/D antigen, was seen strongly expressed in this region of tumoral tissue (Figure 5F); and SOX2, a neural progenitor marker, was also detected. Taken together these data indicate that tumors from rb1-TALENs injected tp53e7/e7 mutant zebrafish may be medulloblastoma-like PNETs based on their incidence location, histopathological features, and neuronal/neural characteristics from immunohistochemistry.
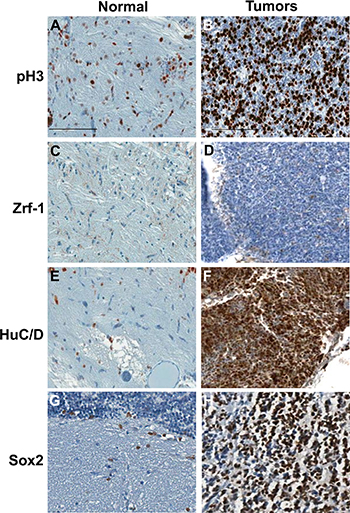
Figure 5: Immunohistochemical analysis of tumors from rb1-TALENs injected tp53 mutant zebrafish. Immunostaining was performed with lineage specific antibody with normal brain tissues (A, C, E, and G) and tumors from rb1-TALENs injected tp53 mutant zebrafish (B, D, F, and H) using anti phospho-Histone 3 (A and B), anti Zrf-1 (C and D), anti HuC/D (E and F) and anti Sox2 (G and H) antibodies. Highly mitotic tumor cells could be detected in tumor tissues from rb1-TALENs mRNA injected tp53 mutant zebrafish (B). Glial marker, Zrf-1 was scarcely stained in densely cellular region of tumor tissue (D). However, HuC/D antigen which is a postmitotic neuronal marker was strongly expressed in densely cellular region of tumor tissue (F). Neural progenitor marker, Sox2 expression was also detected in same cellular region of tumor tissue (H). Scale bars: 50 μm.
cdkn2a/b or rb1 were highly mutated in tumors induced from cdkn2a/b- or rb1 TALEN-mRNA injection
To evaluate mutation frequency of cdkn2a/b or rb1 in tumor tissues derived by TALEN injection, PCR products harboring each TALEN-targeted locus of genomic DNA were cloned and sequenced. The mutation frequency of cdkn2a/b or rb1 from the tissues was higher than that of heterozygous mutants in all tested tumors. Mutation frequency in tumor tissues from rb1 TALEN-injected F0 tp53e7/e7 zebrafish varied from 66.6% (32/48) to 83.3% (40/48), while the frequency in cdkn2a/b TALEN-injected F0 tp53e7/e7 zebrafish varied from 81.3% (39/48) to 91.6% (44/48) (Figure 6A).
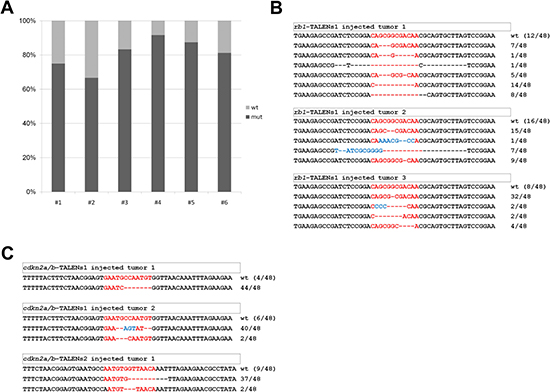
Figure 6: Mutational frequency and allele variation of rb1 or cdkn2a/b genes in tumor tissues derived by TALENs injected F0 adult zebrafish. (A) Mutational frequency of rb1 or cdkn2a/b genes was evaluated from sequencing analysis of cloned amplicon from genomic DNA of independent tumor tissues induced by rb1 or cdkn2a/b TALENs injected F0 tp53 mutant zebrafish. Mutational frequency of tumor tissues from rb1 TALENs injected F0 tp53 mutant zebrafish were varied from 66.6% (32/48) to 83.3% (40/48), and that of tumor tissues from cdkn2a/b TALENs injected F0 tp53 mutant zebrafish were varied from 81.3% (39/48) to 91.6% (44/48). (B) Mutation alleles of rb1’s exon 1 in tumor tissues from rb1 TALENs injected F0 tp53 mutant zebrafish. (C) Mutation alleles of cdkn2a/b’s exon 1 in tumor tissues from cdkn2a/b-TALENs1 or 2 injected F0 tp53 mutant zebrafish.
Mutation forms of rb1 in tumor tissues of rb1 TALEN-injected F0 tp53e7/e7 mutant zebrafish were analyzed. Total 16 mutations were identified from three independent tumor-bearing adult zebrafish. In two of the three tumors (rb1-TALEN-T2, rb1-TALEN-T3), all alleles showed frame-shift mutations; for the other tumor (rb1-TALEN-T1), 25% (9/36) of mutant alleles were in-frame mutations (Figure 6B). In the case of cdkn2a/b, most mutations in tumor tissues of cdkn2a/b TALEN-injected F0 tp53e7/e7 zebrafish were also frame-shifts, while some were in-frame mutations (Figure 6C).
Furthermore, cdkn2a/b and rb1 transcripts were also rarely expressed in tumors induced by cdkn2a/b and rb1 somatic inactivation by TALENs injection, respectively (Supplementary Figure 2).
These data show that TALEN-mediated tumors were induced by loss-of-function of targeted-genes and that these loss-of-function mutations were resulted from the highly efficient bi-allelic inactivation of target genes.
Molecular signatures of zebrafish tumors induced by rb1 somatic inactivation displayed a similar characteristic of human MB and/or PNETs
QuantSeq 3′ mRNA sequencing was performed in order to characterize the molecular features of tumor tissues induced by rb1-TALEN injection. For QuantSeq 3′ mRNA sequencing, normalization (i.e. RPKM or TPM) for sequencing depth and gene length was not required since only one fragment per transcript is generated and also the number of reads mapped to a gene is directly linked to its expression [21]. To identify differentially expressed genes, we selected genes for which fold change was higher than 1.50 and lower than 0.67. We finally obtained 1679 up- and 1183 down-regulated transcripts for the tumor tissues (Supplementary Table 1).
Gene Ontology Analysis was performed in order to understand the signatures of differentially expressed genes (DEGs). The analysis showed that cell cycle, DNA replication, cell division, and translation were significantly up-regulated while ATP metabolism and vesicle/protein transport were down-regulated in the tumor tissues. DNA binding in the nucleus, RNA binding, structural constituent of ribosome of molecular function and cellular component of gene ontology were also up-regulated in tumor tissues (Figure 7A–7C, Supplementary Table 2). Tumors induced by rb1 somatic inactivation displayed medulloblastoma like PNET characteristics in histopathology and immunohistochemistry analysis (Figure 4, Figure 5). Thus, we further analyzed pathway signatures of DEGs and focused especially on neurogenesis, WNT, SHH, and NOTCH pathways which are used as subgroup classifications of human medulloblastoma [2]. A variety of genes involved in cell cycle progression were highly expressed in the tumors (Figure 7D). Interestingly, expression levels of differentiated neuronal genes (i.e. elav3, isl1, etc.) and undifferentiated neuronal precursor genes (i.e. notch1a, sox2, nkx2.2a, etc.) were coincidentally up-regulated as well (Figure 7D). Although GESA for WNT, SHH, and NOTCH pathways scarcely showed statistical significance (data not shown), we could observe the up-regulation of some WNT component genes (such as tcf7l1a and tcf7l1b), SHH target genes (such as klf2c, invs, and gli2a), and NOTCH receptor or ligand genes (such as notch1a, dll4, dla, and dlb) and target genes (such as nrarpb) (Figure 7D). Thus, these data demonstrated that molecular signatures of tumors induced by rb1 somatic inactivation were characterized by high activation of cell cycle progression and protein synthesis and the up-regulation of neuronal genes including WNT, SHH, and NOTCH pathway components or target genes.
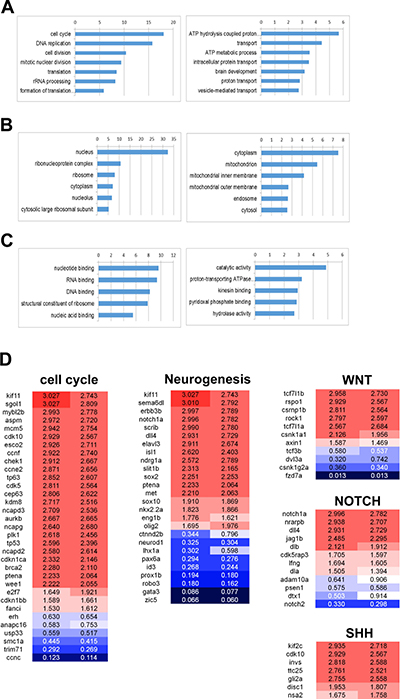
Figure 7: Gene ontology analysis of the differentially expressed genes in tumors induced by rb1 somatic inactivation. Gene Ontology Analysis was performed with the differentially expressed genes (DEG), after QuantSeq 3′ mRNA sequencing of tumors induced by rb1 somatic inactivation. (A–C) -log10 (P value) was indicated at the top. (A) The up or down-regulated in biological processes. The cell cycle, DNA replication, cell division, and translation process were significantly up-regulated however ATP metabolism and vesicle/protein transport were down-regulated. (B) The up or down-regulated in cellular component. The nucleus and ribonucleoprotein complex were up-regulated. (C) The up or down-regulated in molecular function. (D) The featured DEGs in the neurogenesis, WNT, SHH, and NOTCH pathway. Red and blue represent up- and down-regulation of genes with fold change.
Moreover, public microarray data of human medulloblastomas and PNETs were used to compare gene expression profiles of tumors induced by rb1 somatic inactivation (Table 2). As presented in Figure 8A, there were 57 up-regulated genes common to the tumors induced by rb1 somatic inactivation (zebrafish tumors) and human medulloblastomas and PNETs most of which were involved in cell cycle progression and mitosis (such as cdk1, aurka, plk1, etc., Supplementary Table 3). Of the up-regulated genes common to zebrafish tumors and human medulloblastomas, most of them are involved in cell cycle progression (ccnb1, cdk2, mycl, wee1, etc.) and/or DNA replication, initiation, and elongation (mcm2, mcm4, mcm7, etc., Figure 8A, Supplementary Table 3) and primarily annotated to medulloblastoma associated genes (DisGeNET, http://www.disgenet.org/web/DisGeNET/menu/home) [22]. Interestingly, there were more simultaneously up-regulated genes (157 genes) in zebrafish tumors and human medulloblastomas than there were (71 genes) in zebrafish tumors and human PNETs (Figure 8A, 8B). On the other hand, simultaneously down-regulated genes (40 genes) in zebrafish tumors and human medulloblastomas were fewer than those (81 genes) in zebrafish tumors and human PNETs (Figure 8A, 8C). These findings demonstrated that tumors induced by rb1-TALEN mediated somatic inactivation have both medulloblastoma and PNET molecular characteristics albeit a greater number of medulloblastoma associated up-regulated genes than PNET up-regulated genes.
Table 2: A list of the collected affymetrix human genome U133 Plus 2.0 array samples using the comparative analysis
GSE |
Raw data |
Normal sample |
Tumor sample |
PNET |
Medulloblastoma |
|---|---|---|---|---|---|
GSE19404 |
24 |
1 |
21 |
13 |
8 |
GSE50161 |
130 |
3 |
22 |
- |
22 |
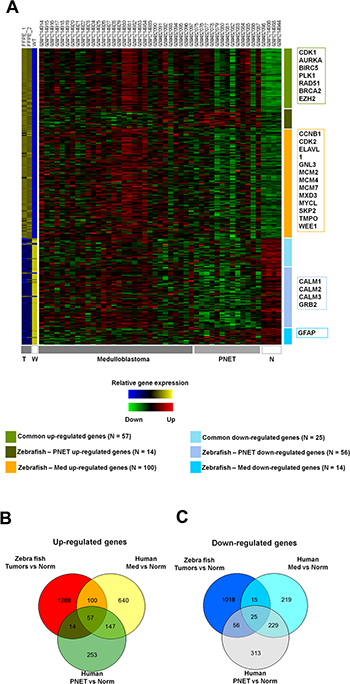
Figure 8: Comparative analysis between gene expression profiles of zebrafish tumors induced by rb1 somatic inactivation and public micro array data of human PNETs and medulloblastomas. (A) Heatmap of gene expression profiles. (T = Tumors, W = Wild type, and N = Normal tissue) The individual samples are indicated at the top of each column and the group of samples is indicated in the bottom. The representative genes are indicated in boxes at the left. (B, C). Venn diagrams showing differently expressed genes. (B) up-regulated and (C) down-regulated genes. (FDR < 0.05) (Norm; Normal sample, Med; medulloblastoma, PNET; primitive neuroectodermal tumor).
DISCUSSION
Somatic gene inactivation of tumor suppressor genes is a general feature of most human tumors [23–26]. In animal tumor modeling, however, most general methods for tumor suppressor gene inactivation involved germline-targeted knockout strategies [27, 28]. In the course of developing tumor suppressor gene knockout progeny, we observed that somatic gene inactivation of cdkn2a/b and/or rb1 efficiently induced tumors in the F0 founder generation. Our observations are consistent with previous reports that brain tumor modeling could be successfully performed with TALENs or CRISPR/Cas9 mediated somatic gene inactivation for tumor suppressor genes in mouse and zebrafish [16, 29]. The extremely high mutation frequency of cdkn2a/b or rb1 in tumor tissues derived by TALENs injection also supports the possibility of tumor induction by specific inactivation of cdkn2a/b or rb1 (Figure 6). Taken together, our findings showed that somatic inactivation of tumor suppressor genes by genome editing tools such as TALENs or CRISPR/Cas9 engineered nucleases could be a useful method for the development of animal tumor models.
Gene rb1 has important roles in regulating G1 to S phase entry by repressing the activity of E2Fs transcription factors and CDKs. Inactivation of rb1 is thought to promote either induction or progression events in a variety of cancers such as retinoblastoma, prostate cancer, small cell lung cancer, etc. [30–34]. In this study, brain tumors induced by somatic inactivation of rb1 showed incidence mainly in the cerebellum. Histopathology data exhibited distinct Homer-Wright like features [35] while immunohistochemistry elicited positive staining for neuronal/neural but not glial cell. For these reasons, we assumed that brain tumors induced by somatic inactivation of rb1 are medulloblastoma like PNETs, which molecular comparative analysis of RNA-seq results confirmed (Figure 8). Although rb1 mutation has not been widely detected in traditional human medulloblastomas, some reports from clinical case studies and animal tumor models have suggested that rb1 pathway participates in medulloblastoma development. In rare cases, pediatric patients with hereditary retinoblastoma harboring rb1 mutation are at an increased risk for pineoblastoma, supratentorial PNET, and medulloblastoma [36, 37]. Conditional knockout of rb1 in cerebellar external granular cells of tp53 knockouts induced development of medulloblastoma [7]. These reports are consistent with our findings that somatic inactivation of rb1 induced medulloblastoma like PNETs and that frequency of tumor induction was noticeably enhanced in tp53e7/e7 mutant zebrafish (Figure 3). A recent report showed that in glial cells of transgenic mice, overexpression of E2F1, a transcription factor downstream of rb1 pathway, also resulted in the induction of medulloblastomas and other PNETs [38]. Interestingly, conditional knockouts of upstream or downstream regulators of tp53e7/e7 - ATM, p19ARF (a mouse homolog of human p14ARF) or p21- were not able to elicit MB in Rb knockout background, indicating a specific role of Tp53 (such as inducing genomic instability, rather than cell cycle regulation or DNA damage response) in MB/PNET pathogenesis [39].
In mammals, p15Ink4b encoded by the cdkn2a locus neighboring cdkn2b locus, was presumed to be generated by gene duplication and believed to play a redundant role for p16Ink4a. In zebrafish, there seems to be only one cdkn2 gene, collectively called cdkn2a/b. Although mammalian p15Ink4b has been implicated primarily in leukemia and lymphoma [11], its involvement in MPNSTs has been suggested due to its expression and genetic status [40]. Consistent with this notion, vascular smooth muscle cells, the potential tumor cell origin for MPNST, underwent apoptosis in p15Ink4b knockout mice in a tp53-dependent manner [41]. Therefore, the acceleration of MPNSTs upon somatic inactivation of cdkn2a/b in tp53e7/e7 mutant background may possibly be due to a reduction in apoptosis of affected tumor cells of origin. The exact tumor types induced by cdkn2a/b mutations would be determined according to binding partner and/or biological context in normal and cancer cells. It was notable that cdkn2a/b and rb1 double somatic inactivation did not influence the overall tumor incidence rate, and merely lead to form the mixture of MPNSTs & MB-like PNETs in an each single tumor developing zebrafish (Table 1). These data mean that somatic inactivation of cdkn2a/b or rb1 did not show a synergic effect on tumor incidence rate or tumor types each other, and could be postulated that cdkn2a/b and rb1 have a distinct role in tumorigenesis.
The molecular features of zebrafish tumors induced by rb1 somatic inactivation were characterized by an up-regulation of genes for cell cycle progression, DNA replication, and translation, and a down-regulation of genes for ATP metabolism and vesicle/protein transport (Figure 7). These molecular characteristics are well-conserved features of a variety of tumors including medulloblastoma and PNETs. Consistent with the data, the signatures of tumor DEGs induced by rb1 somatic inactivation were closely associated with those of human medulloblastomas and PNETs (Figure 8). Hence, we could conclude that tumors induced by rb1 somatic inactivation are medulloblastoma-like PNETs. According to several reports [4, 42–45], medulloblastoma is classified into four molecularly distinct subgroups –WNT, SHH, Group C (MYC activated), and Group D. Zebrafish tumors induced by rb1 somatic inactivation displayed an apparent up-regulation of genes in neural development and NOTCH pathway. Some genes of the WNT and SHH pathways were specifically up-regulated in zebrafish tumors but not to statistical significance. Therefore, we inferred that zebrafish tumors induced by rb1 somatic inactivation are a primitive type of medulloblastoma-like PNET accompanied by indistinguishable subtypes.
In summary, we have presented the efficient development of a zebrafish tumor model by TALEN-mediated somatic inactivation of tumor suppressor genes. The development of MPNSTs was accelerated by cdkn2a/b gene inactivation via TALEN mRNA injection into fertilized eggs of tp53e7/e7 mutant background. Also, TALEN-mediated rb1 somatic inactivation led to the development of brain tumors in the zebrafish model. Using RNA sequencing analysis in addition to histopathological and immunohistochemistry, we showed that brain tumors induced by rb1 somatic inactivation have molecular features of medulloblastoma-like PNETs.
MATERIALS AND METHODS
Wild type and mutant zebrafish
Wild type zebrafish (Danio rerio) with Oregon AB genetic background were used. The tp53e7/e7 (tp53M214K) mutant was previously described [46]. Zebrafish were essentially maintained as described in [47], and all experimental procedures using zebrafish was approved by the Institutional Animal Care and Use Committee at National Cancer Center, Republic of Korea (permit number: NCC-15-116C).
TALEN construction, mRNA injection, and mutational analysis
TALEN vectors targeting the first exon of cdkn2a/b gene (Figure 1A) and rb1 (Figure 3A) were designed and constructed by ToolGen (http://toolgen.com/). TALEN vectors were linearized by PvuII, extracted with phenol/chloroform, and followed by ethanol precipitation. Subsequently, mRNAs encoding TALENs were synthesized using the mMESSAGE MACHINE T7 Transcription kit (Ambion) and purified as shown above; and about 200 pg of mRNAs were injected into one-cell stage zebrafish embryos.
Site-specific TALEN efficiency was examined by target-specific PCR with isolation of genomic DNA, and followed by T7 endonuclease I assay. In brief, the genomic DNA was extracted in DNA isolation buffer (10 mM Tris, 10 mM EDTA, 150 mM NaCl, 0.5% SDS and 0.1 mg/ml Proteinase K) for 2 h at 60°C, and PCRs were performed using the following primer pairs: cdkn1a/b-TALENs1&2 forward primer, 5′-CAGCGTTGAACTGATTGTTTTCG-3′; cdkn1a/b- TALENs1&2 reverse primer, 5′-TCCCATATAGTCAA ACAGGTGTG-3′; rb1-TALENs1 forward primer, 5′-CTG AGAGTGAACGCGCTCTTCT-3′; rb1-TALENs1 reverse primer, 5′-CGTCAGGTCGCTCTCTTCCTTCC-3′. Consequently, PCR products were purified by PCR/GEL purification mini kit (FAVORGEN) and analyzed by T7 endonuclease I assay or sequencing.
Histopathology and immunohistochemistry
For observation of histological features of normal and tumor tissues, adult zebrafish were anesthetized with 0.016% of tricane methanesulfonate (Sigma), fixed in 4% paraformaldehyde in PBS for 16 h at 4°C, and then decalcification was followed as described previously [48]. After sectioning (4 to 6 mm) and pretreatment of paraffin embedded tissue, tissue sections were stained with hematoxylin (DaKo) and eosin (Merck). Laser capture microdissection with LMD6 (Leica) was used for normal or tumor tissue preparation to examine the mutation allele of genomic DNA or analyze the differential expression level of RNAs with RNA-seq.
For immunostaining, antigen retrieval process and endogenous peroxidase blocking was performed with citrate buffer (pH 6.0) and 3% hydrogen peroxide, respectively. Tissue sections were incubated in 5% normal serum in PBSBD (1% BSA and 1%DMSO in PBS) for 1 h at room temperature and were reacted with primary antibody overnight at 4°C in a humidified chamber. Anti-phospho-Histone H3 (1/100 dilution, mouse monoclonal, Sigma), anti-GFAP (also known as Zrf-1, 1/2000, mouse monoclonal, Zebrafish International Resource Center), anti- HuC/HuD (1/2000, 16A11, mouse monoclonal, Sigma), anti-S100β (1/100, rabbit monoclonal, Abcam), anti-Vimentin (1/500, rabbit monoclonal, Abcam), and anti-Sox2(1/1000, rabbit monoclonal, Abcam) antibodies were used. VECTASTAIN Elite ABC HRP Kit (Vector Laboratories) and 3, 3′-diaminobenzidine (Vector Laboratories) as a chromogen were used for visualization of signals.
RNA isolation and library preparation and RNA sequencing
In order to characterize the molecular features of tumor tissues induced by rb1-TALENs injection, the tumor tissues were collected with laser captured microscope from paraffin embedded tissue section. Total RNA was isolated using Trizol reagent (Invitrogen). RNA quality was assessed by Agilent 2100 bioanalyzer using the RNA 6000 Nano Chip (Agilent Technologies), and RNA quantification was performed using ND-2000 Spectrophotometer (Thermo). Construction of the library was performed using SENSE 3′ mRNA-Seq Library Prep Kit (Lexogen) according to manufacturer instructions. In brief, reverse transcription reaction were performed with 500 ng total RNA and oligo-dT primer containing an Illumina-compatible sequence at its 5′ end. And then, synthesis of the second strand was initiated by random primer containing an Illumina-compatible linker sequence at its 5′ end, after degradation of the RNA template. The double-stranded library was purified by using magnetic beads to remove all reaction components, and amplified to add the complete adapter sequences required for cluster generation. High-throughput sequencing was performed as single-end 75 sequencing using NextSeq 500 (Illumina).
Data analysis
About 9 to 10 million sequence tags (11.6x106 for normal brain tissue, 9.8 and 8.9 × 106 for two independent tumor tissues) were obtained. SENSE 3′ mRNA-Seq reads were aligned using Bowtie2 version 2.1.0 [49]. Bowtie2 indices were either generated from genome assembly sequence or the representative transcript sequences for aligning to the genome and transcriptome. After mapping to the zebrafish reference sequence database for quantitative analysis of transcripts expression, 8329 genes for normal brain tissue, 5403 and 5688 genes for two independent tumor tissues were annotated. Differentially expressed genes were determined based on counts from unique and multiple alignments using EdgeR within R version 3.2.2 (R development Core Team, 2011) in BIOCONDUCTOR version 3.0 [50]. The RT (Read Count) data were processed based on Quantile normalization method using the Genowiz ™ version 4.0.5.6 (Ocimum Biosolutions). Cytoscape (version 2.7, http://www.cytoscape.org/), an open source bioinformatics platform developed by the Institute of Systems Biology, was used to construct network diagrams and to illustrate clustering of the genes in our dataset within specific pathways. Gene classification was based on searches done by DAVID (http://david.abcc.ncifcrf.gov/) and Medline databases (http://www.ncbi.nlm.nih.gov/).
Microarray data was collected for medulloblastoma, primitive neuroectodermal tumor and normal samples from Gene Expression Omnibus [51]. The gene expression study sample dataset included 30 medulloblastomas, 13 primitive neuroectodermal tumors, four normal brain samples. Scanned microarray data were background corrected and normalized using the Robust Multi-Array Average algorithm resulting in log2 gene expression values. Differently expressed genes were selected using false discovery rate < 0.05 between tumor and normal samples. Human gene symbols were converted to zebrafish gene symbols using The Zebrafish Model Organism Database [52]. Python (version 2.7.6) and the Pandas python library (version 0.15.2) were used for most data analysis. R (version 3.1.0) and MeV (version 4.9.0) [53] were used for image production. BIOCONDUCTOR version 3.0 [54] package was used for normalization of microarray data.
Authors’ contributions
The accession number for the RNA-seq data reported in this paper is GEO: GSE92260. JS, JHC, MHP, and YKB performed most of the experiments, analyzed the data. JSL, CHK, and YKB designed the study and manuscript writing. HNK, JHK, SYK, DWH, SSK, and JEL helped in data analysis, interpretation, and discussion.
ACKNOWLEDGMENTS
This work was supported in part by National Cancer Center, Korea (NCC-1310860 and NCC-1610390) and National Research Foundation of Korea (2014R1A2A1A11053562), National Research Council of Science & Technology (NST) grant by the Korea government (MSIP) (No. CRC-15-04-KIST), and KRIBB Research Initiative Program.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Gopalakrishnan V, Tao RH, Dobson T, Brugmann W, Khatua S. Medulloblastoma development: tumor biology informs treatment decisions. CNS Oncol. 2015; 4:79–89.
2. Northcott PA, Jones DT, Kool M, Robinson GW, Gilbertson RJ, Cho YJ, Pomeroy SL, Korshunov A, Lichter P, Taylor MD, Pfister SM. Medulloblastomics: the end of the beginning. Nat Rev Cancer. 2012; 12:818–834.
3. Goodrich LV, Milenkovic L, Higgins KM, Scott MP. Altered neural cell fates and medulloblastoma in mouse patched mutants. Science. 1997; 277:1109–1113.
4. Gibson P, Tong Y, Robinson G, Thompson MC, Currle DS, Eden C, Kranenburg TA, Hogg T, Poppleton H, Martin J, Finkelstein D, Pounds S, Weiss A, et al. Subtypes of medulloblastoma have distinct developmental origins. Nature. 2010; 468:1095–1099.
5. Kawauchi D, Robinson G, Uziel T, Gibson P, Rehg J, Gao C, Finkelstein D, Qu C, Pounds S, Ellison DW, Gilbertson RJ, Roussel MF. A mouse model of the most aggressive subgroup of human medulloblastoma. Cancer Cell. 2012; 21:168–180.
6. Henley SA, Dick FA. The retinoblastoma family of proteins and their regulatory functions in the mammalian cell division cycle. Cell Div. 2012; 7:10.
7. Marino S, Vooijs M, van Der Gulden H, Jonkers J, Berns A. Induction of medulloblastomas in p53-null mutant mice by somatic inactivation of Rb in the external granular layer cells of the cerebellum. Genes Dev. 2000; 14:994–1004.
8. Maandag EC, van der Valk M, Vlaar M, Feltkamp C, O’Brien J, van Roon M, van der Lugt N, Berns A, te Riele H. Developmental rescue of an embryonic-lethal mutation in the retinoblastoma gene in chimeric mice. EMBO J. 1994; 13:4260–4268.
9. Williams BO, Remington L, Albert DM, Mukai S, Bronson RT, Jacks T. Cooperative tumorigenic effects of germline mutations in Rb and p53. Nat Genet. 1994; 7:480–484.
10. Kim WY, Sharpless NE. The regulation of INK4/ARF in cancer and aging. Cell. 2006; 127:265–275.
11. Sharpless NE. INK4a/ARF: a multifunctional tumor suppressor locus. Mutat Res. 2005; 576:22–38.
12. White R, Rose K, Zon L. Zebrafish cancer: the state of the art and the path forward. Nat Rev Cancer. 2013; 13:624–636.
13. Zu Y, Tong X, Wang Z, Liu D, Pan R, Li Z, Hu Y, Luo Z, Huang P, Wu Q, Zhu Z, Zhang B, Lin S. TALEN-mediated precise genome modification by homologous recombination in zebrafish. Nat Methods. 2013; 10:329–331.
14. Hwang WY, Fu Y, Reyon D, Maeder ML, Tsai SQ, Sander JD, Peterson RT, Yeh JR, Joung JK. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol. 2013; 31:227–229.
15. Soroldoni D, Hogan BM, Oates AC. Simple and efficient transgenesis with meganuclease constructs in zebrafish. Methods Mol Biol. 2009; 546:117–130.
16. Solin SL, Shive HR, Woolard KD, Essner JJ, McGrail M. Rapid tumor induction in zebrafish by TALEN-mediated somatic inactivation of the retinoblastoma1 tumor suppressor rb1. Sci Rep. 2015; 5:13745.
17. Storer NY, Zon LI. Zebrafish models of p53 functions. Cold Spring Harb Perspect Biol. 2010; 2:a001123.
18. Zhang G, Hoersch S, Amsterdam A, Whittaker CA, Lees JA, Hopkins N. Highly aneuploid zebrafish malignant peripheral nerve sheath tumors have genetic alterations similar to human cancers. Proc Natl Acad Sci USA. 2010; 107:16940–16945.
19. Stasik CJ, Tawfik O. Malignant peripheral nerve sheath tumor with rhabdomyosarcomatous differentiation (malignant triton tumor). Arch Pathol Lab Med. 2006; 130:1878–1881.
20. Bae YK, Kani S, Shimizu T, Tanabe K, Nojima H, Kimura Y, Higashijima S, Hibi M. Anatomy of zebrafish cerebellum and screen for mutations affecting its development. Dev Biol. 2009; 330:406–426.
21. Moll P, Ante M, Seitz A, Reda T. QuantSeq 3′mRNA sequencing for RNA quantification. Nature Methods. 2014; 11.
22. Pinero J, Queralt-Rosinach N, Bravo A, Deu-Pons J, Bauer-Mehren A, Baron M, Sanz F, Furlong LI. DisGeNET: a discovery platform for the dynamical exploration of human diseases and their genes. Database (Oxford). 2015; 2015:bav028.
23. Campbell PJ, Stephens PJ, Pleasance ED, O’Meara S, Li H, Santarius T, Stebbings LA, Leroy C, Edkins S, Hardy C, Teague JW, Menzies A, Goodhead I, et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat Genet. 2008; 40:722–729.
24. Corradini P, Inghirami G, Astolfi M, Ladetto M, Voena C, Ballerini P, Gu W, Nilsson K, Knowles DM, Boccadoro M. Inactivation of tumor suppressor genes, p53 and Rb1, in plasma cell dyscrasias. Leukemia. 1994; 8:758–767.
25. Lindblad-Toh K, Tanenbaum DM, Daly MJ, Winchester E, Lui WO, Villapakkam A, Stanton SE, Larsson C, Hudson TJ, Johnson BE, Lander ES, Meyerson M. Loss-of-heterozygosity analysis of small-cell lung carcinomas using single-nucleotide polymorphism arrays. Nat Biotechnol. 2000; 18:1001–1005.
26. Peifer M, Fernandez-Cuesta L, Sos ML, George J, Seidel D, Kasper LH, Plenker D, Leenders F, Sun R, Zander T, Menon R, Koker M, Dahmen I, et al. Integrative genome analyses identify key somatic driver mutations of small-cell lung cancer. Nat Genet. 2012; 44:1104–1110.
27. Resor L, Bowen TJ, Wynshaw-Boris A. Unraveling human cancer in the mouse: recent refinements to modeling and analysis. Hum Mol Genet. 2001; 10:669–675.
28. Van Dyke T, Jacks T. Cancer modeling in the modern era: progress and challenges. Cell. 2002; 108:135–144.
29. Zuckermann M, Hovestadt V, Knobbe-Thomsen CB, Zapatka M, Northcott PA, Schramm K, Belic J, Jones DT, Tschida B, Moriarity B, Largaespada D, Roussel MF, Korshunov A, et al. Somatic CRISPR/Cas9-mediated tumour suppressor disruption enables versatile brain tumour modelling. Nat Commun. 2015; 6:7391.
30. Dimaras H, Khetan V, Halliday W, Orlic M, Prigoda NL, Piovesan B, Marrano P, Corson TW, Eagle RC Jr, Squire JA, Gallie BL. Loss of RB1 induces non-proliferative retinoma: increasing genomic instability correlates with progression to retinoblastoma. Hum Mol Genet. 2008; 17:1363–1372.
31. Jiang Z, Deng T, Jones R, Li H, Herschkowitz JI, Liu JC, Weigman VJ, Tsao MS, Lane TF, Perou CM, Zacksenhaus E. Rb deletion in mouse mammary progenitors induces luminal-B or basal-like/EMT tumor subtypes depending on p53 status. J Clin Invest. 2010; 120:3296–3309.
32. Meuwissen R, Linn SC, Linnoila RI, Zevenhoven J, Mooi WJ, Berns A. Induction of small cell lung cancer by somatic inactivation of both Trp53 and Rb1 in a conditional mouse model. Cancer Cell. 2003; 4:181–189.
33. Sharma A, Yeow WS, Ertel A, Coleman I, Clegg N, Thangavel C, Morrissey C, Zhang X, Comstock CE, Witkiewicz AK, Gomella L, Knudsen ES, Nelson PS, et al. The retinoblastoma tumor suppressor controls androgen signaling and human prostate cancer progression. J Clin Invest. 2010; 120:4478–4492.
34. Sutherland KD, Proost N, Brouns I, Adriaensen D, Song JY, Berns A. Cell of origin of small cell lung cancer: inactivation of Trp53 and Rb1 in distinct cell types of adult mouse lung. Cancer Cell. 2011; 19:754–764.
35. Katsetos CD, Liu HM, Zacks SI. Immunohistochemical and ultrastructural observations on Homer Wright (neuroblastic) rosettes and the “pale islands” of human cerebellar medulloblastomas. Hum Pathol. 1988; 19:1219–1227.
36. Elias WJ, Lopes MB, Golden WL, Jane JA Sr, Gonzalez-Fernandez F. Trilateral retinoblastoma variant indicative of the relevance of the retinoblastoma tumor-suppressor pathway to medulloblastomas in humans. J Neurosurg. 2001; 95:871–878.
37. Marcus DM, Brooks SE, Leff G, McCormick R, Thompson T, Anfinson S, Lasudry J, Albert DM. Trilateral retinoblastoma: insights into histogenesis and management. Surv Ophthalmol. 1998; 43:59–70.
38. Olson MV, Johnson DG, Jiang H, Xu J, Alonso MM, Aldape KD, Fuller GN, Bekele BN, Yung WK, Gomez-Manzano C, Fueyo J. Transgenic E2F1 expression in the mouse brain induces a human-like bimodal pattern of tumors. Cancer Res. 2007; 67:4005–4009.
39. Shakhova O, Leung C, van Montfort E, Berns A, Marino S. Lack of Rb and p53 delays cerebellar development and predisposes to large cell anaplastic medulloblastoma through amplification of N-Myc and Ptch2. Cancer Res. 2006; 66:5190–5200.
40. Endo M, Kobayashi C, Setsu N, Takahashi Y, Kohashi K, Yamamoto H, Tamiya S, Matsuda S, Iwamoto Y, Tsuneyoshi M, Oda, Y. Prognostic significance of p14ARF, p15INK4b, and p16INK4a inactivation in malignant peripheral nerve sheath tumors. Clin Cancer Res. 2011; 17:3771–3782.
41. Leeper NJ, Raiesdana A, Kojima Y, Kundu RK, Cheng H, Maegdefessel L, Toh R, Ahn GO, Ali ZA, Anderson DR, Miller CL, Roberts SC, Spin JM, et al. Loss of CDKN2B promotes p53-dependent smooth muscle cell apoptosis and aneurysm formation. Arterioscler Thromb Vasc Biol. 2013; 33:e1–e10.
42. Gendoo DM, Smirnov P, Lupien M, Haibe-Kains B. Personalized diagnosis of medulloblastoma subtypes across patients and model systems. Genomics. 2015; 106:96–106.
43. Northcott PA, Korshunov A, Witt H, Hielscher T, Eberhart CG, Mack S, Bouffet E, Clifford SC, Hawkins CE, French P, Rutka JT, Pfister S, Taylor MD. Medulloblastoma comprises four distinct molecular variants. J Clin Oncol. 2011; 29:1408–1414.
44. Parsons DW, Li M, Zhang X, Jones S, Leary RJ, Lin JC, Boca SM, Carter H, Samayoa J, Bettegowda C, Gallia GL, Jallo GI, Binder ZA, et al. The genetic landscape of the childhood cancer medulloblastoma. Science. 2011; 331:435–439.
45. Taylor MD, Northcott PA, Korshunov A, Remke M, Cho YJ, Clifford SC, Eberhart CG, Parsons DW, Rutkowski S, Gajjar A, Ellison DW, Lichter P, Gilbertson RJ, et al. Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol. 2012; 123:465–472.
46. Berghmans S, Murphey RD, Wienholds E, Neuberg D, Kutok JL, Fletcher CD, Morris JP, Liu TX, Schulte-Merker S, Kanki JP, Plasterk R, Zon LI, Look AT. tp53 mutant zebrafish develop malignant peripheral nerve sheath tumors. Proc Natl Acad Sci USA. 2005; 102:407–412.
47. Westerfield M. The zebrafish book. A guide for the laboratory use of zebrafish (Danio rerio). 4th ed., Univ. of Oregon Press, Eugene. 2000.
48. Moore JL, Aros M, Steudel KG, Cheng KC. Fixation and decalcification of adult zebrafish for histological, immunocytochemical, and genotypic analysis. Biotechniques. 2002; 32:296–298.
49. Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012; 9:357–359.
50. Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004; 5:R80.
51. Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Muertter RN, et al. NCBI GEO: archive for high-throughput functional genomic data. Nucleic Acids Res. 2009; 37:D885–890.
52. Howe DG, Bradford YM, Conlin T, Eagle AE, Fashena D, Frazer K, Knight J, Mani P, Martin R, Moxon SA, Paddock H, Pich C, Ramachandran S, et al. ZFIN, the Zebrafish Model Organism Database: increased support for mutants and transgenics. Nucleic Acids Res. 2013; 41:D854–860.
53. Howe EA, Sinha R, Schlauch D, Quackenbush J. RNA-Seq analysis in MeV. Bioinformatics. 2011; 27:3209–3210.
54. Gautier L, Cope L, Bolstad BM, Irizarry RA. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics. 2004; 20:307–315.

