INTRODUCTION
As an activator of mitogen-activated protein kinase (MAPK)/ERK kinase (MEK) pathway and an effector of Ras, Raf is involved in many fundamental cellular processes such as cell proliferation, differentiation, cell death and survival, metabolism and motility [1-3]. The mammalian Raf kinase family includes A-Raf, B-Raf, and Raf1 (C-Raf). These proteins all have auto-inhibitory, regulatory, and catalytic domains, and contain multiple phosphorylation sites [4]. Among the three members, Raf1 has drawn the most attention since it was identified 30 years ago [5]. Raf1 has been widely reported as a key effector of the small G protein Ras, and after activation, Raf can phosphorylate MEK, which then activates ERK. ERK then phosphorylates an impressive roster of membrane, cytosolic, and nuclear targets to regulate numerous cell functions [6].
However, unlike B-Raf, Raf1 is not essential for ERK activation. During tumorigenesis, Raf1 interacts with different proteins to allow cross-talk between signaling pathways [7]. Protein phosphatase 2A (PP2A)/PP1 and members of the 14-3-3 family can interact with Raf1 to control its enzymatic activity [8, 9]. Heat shock protein 90 (HSP90) and its co-chaperone CDC37 have also been identified as Raf-associated proteins that are crucial for the maturation and activation of Raf1 [10]. Further research has identified more proteins that interact with and regulate the activity of Raf1. These proteins include p21-activated kinase (PAK3), serine/threonine kinase 3 (STK3), and protein kinase C (PKC) [11, 12]. However, the proteins interacting with Raf1 have not been fully elucidated.
To explore the biological functions of Raf1, we aimed to create an overview of the Raf1 interactome. We enriched Raf1-interacting proteins under stable, MAPK-inactive conditions using co-immunoprecipitation. Then, we identified these interacting proteins using liquid chromatography–mass spectrometry (LC-MS). We uncovered 198 Raf1-interacting proteins and confirmed 12 of these interactions by western blotting. Gene ontology (GO) and pathway enrichment analysis indicated that these Raf1-protein interactions regulated six signaling pathways, and were involved in the ten known hallmarks of cancer. Our findings have deepened our understanding of Raf1 function, and offer potential oncological targets for studying specific Raf1 functions in the future.
RESULTS
Raf1 co-immunoprecipitated with 198 proteins
To identify Raf1 interacting proteins, we induced VSV-Raf1 expression with doxycycline in Flp-in To-REx HEK293 cells containing VSV-Raf1 (Figure 1A). After doxycycline induction, cell lysates were precipitated with anti-VSV-glycoprotein-agarose beads. Exogenous VSV-Raf1 was effectively pulled down and very little VSV-Raf1 remained in the flow through (FT) line (Figure 1B). Immunoprecipitated proteins were separated by SDS-PAGE then observed by silver staining. As shown in Figure 1C, most of the Raf1 specific interacting proteins are concentrated in the ranges of 25~35 KDa and 70~130 KDa. Immunoprecipitated proteins were analyzed by LC−MS. In three independent experiments, 441, 297, and 478 Raf1 interacting proteins were identified. Comparison of these three data sets (Figure 1D) revealed 198 common putative Raf1 interacting proteins. This number was comparable with previously published work [13, 14]. The full list and details of these 198 proteins is presented in Supplementary Table 1.
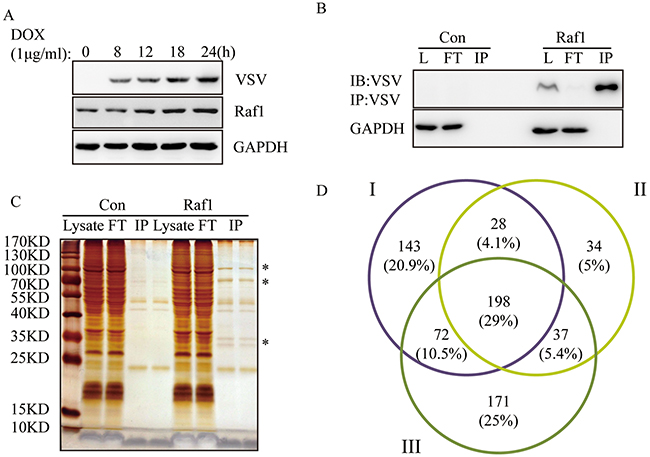
Figure 1: Identification of Raf1 interacting proteins in three independent experiments. (A) Expression of VSV-tagged-Raf1 in Flp-In To-REx HEK293 cells. Cells were induced with 1μg/ml of doxycycline for 8, 12, 18, and 24 hours. VSV-RAF-1 and Raf1 (exogenous and endogenous) were detected using anti-VSV and anti-Raf1 antibodies, respectively. (B) VSV-Raf1 was immunoprecipitated with mouse anti-VSV-G antibody covalently bound to agarose beads and detected using an VSV antibody (L: lysate, FT: flow through). Flp-In To-REx cells not expressing VSV-Raf1 were used as negative controls (Con). (C) Samples from (B) were resolved by SDS-PAGE and observed by silver staining. Flp-In To-REx cells not expressing VSV-Raf1 were used as negative controls (Con). Specific Raf1 interacting proteins are indicated by a star*, two bands of IP were the some samples with two loading repeats (10μl/5μl). (D) Venn diagram corresponding to proteins identified in three independent Raf1 immunoprecipitations. The overlapping 198 proteins correspond to proteins common to the three data sets (http://bioinfogp.cnb.csic.es/tools/venny/).
182 out of these 198 proteins are novel Raf1-interacting proteins
Based on the Raf1 mapping information obtained from the STRING database, we exported 69 experimentally validated interacting proteins that had a combined score of >0.9 (Supplementary Table 2). These interacting proteins included H-RAS, N-RAS, K-RAS, AKT, and PAK1/2/3/4. We found that 16 of our 198 identified interaction partners had been reported previously. These were YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, HSP90AA1, HSP90AB1, CDC37, KRAS, NRAS, HRAS, PPP2R1A, and BRAF (gene names) (Figure 2).
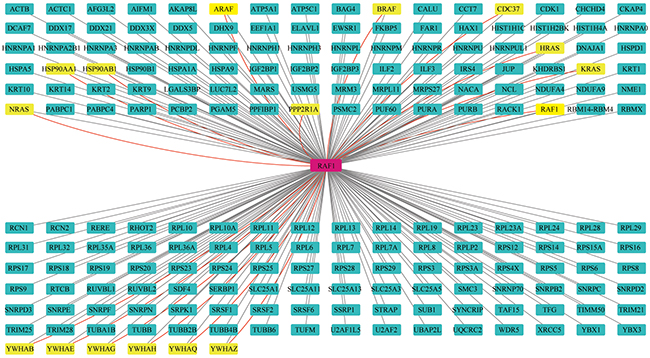
Figure 2: Interaction network of proteins from the Raf1 interactome. Novel interaction partners are identified by black lines and green nodes. Previously identified interaction partners are shown as red lines and yellow nodes.
Western blotting confirmed the reliability of Raf1-interacting proteins
To confirm the reliability of these 198 proteins, we selected 20 interacting proteins of different functions and examined the interaction by western blotting. We were able to confirm an interaction with Raf1 for 17 out of 20 (~85%) selected Raf1 interacting proteins. We could not detect an interaction of AKAP8L, NDUFA4 or HAX1 with Raf1 by western blotting (Figure 3).
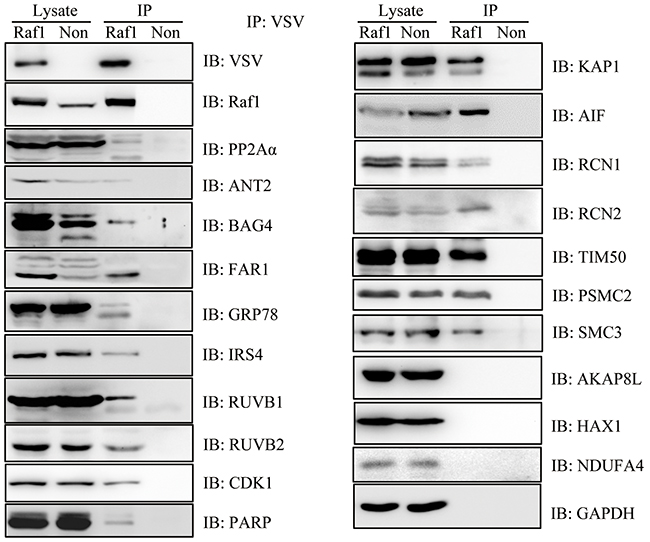
Figure 3: Western blotting confirmed the interaction of 12 proteins with Raf1. Twenty-four hours after doxycycline induction, VSV-Raf1 protein was immunoprecipitated from Flp-In To-Rex HEK293 cells expressing VSV-Raf1 (Raf1) with anti-VSV-G agarose beads. VSV-Raf1 associated proteins were detected by western blotting using the indicated antibodies. Flp-In To-Rex HEK293 cells containing the pcDNA5 empty vector (Non) were used as controls. GAPDH was immunoprecipitated as a negative binding control (LC-MS data showed that GAPDH did not interact with Raf1) and as a reference of equal loading.
Novel Raf1 interacting proteins were involved in many biological processes and distributed in different cellular components
DAVID software was utilized to identify GO categories and KEGG pathways in the 198 Raf1-interacting proteins. GO analysis showed that certain MFs were enriched for Raf1 interacting proteins, including pyrophosphatase activity, enzyme binding, protein kinase binding, GTPase activity, and heat shock protein binding. In addition, Raf1 interacting proteins were enriched in BPs, including positive regulation of metabolism, programmed cell death, cellular stress responses, chromosome organization, and cell-cell adhesion. In addition, GO CC analysis demonstrated significant enrichment for cell junction, mitochondrion, methyltransferase complex, nucleolus, and extracellular matrix proteins (Supplementary Tables 3-5). In Ingenuity Diseases and Bio Functions analysis, Raf1 interacting proteins were enriched to 15 bio functions which were all covered by GO analysis results. However, inductions and descriptions are slightly different between these two analyses (Supplementary Table 7).
Raf1 interacting proteins contribute to all hallmarks of cancer
Ten hallmarks of cancer have been described by Hanahan and Weinberg (outlined in Table 1) [15, 16]. These describe the acquired biological capabilities of cancer cells. We investigated the contribution of Raf1 interacting proteins to the hallmarks and states of cancer based on GO enrichment (BPs and MFs). Table 1 shows specific GO terms that apply to cancer hallmarks, including regeneration (GO: 0031099) and the ERBB2 signaling pathway (GO:0038128), which correlate with “evading growth suppressors”. We confirmed that Raf1 interacting proteins are involved in all ten hallmarks of cancer.
Table 1: Classification of Raf1 interacting proteins based on the hallmarks of cancer
Hallmark |
Term |
GO term and function |
|---|---|---|
|
Sustaining proliferative signaling |
GO: 0030518~intracellular steroid hormone receptor signaling pathway |
|
Evading growth suppression |
GO: 0031099~regeneration |
|
Resisting cell death |
GO: 0012501~programmed cell death |
|
Enabling replicative immortality |
GO: 1903047~mitotic cell cycle process |
|
Inducing angiogenesis |
GO: 0031100~organ regeneration |
|
Activating invasion and metastasis |
GO: 0098609~cell-cell adhesion |
|
Genome instability and mutation |
GO: 0006281~DNA repair |
|
Avoiding immune destruction |
GO: 0002253~activation of immune response |
|
Tumor-promoting inflammation |
GO: 0071353~cellular response to interleukin-4 |
|
Deregulating cellular energetics |
GO: 0044248~cellular catabolic process |
Raf1 interacting proteins were enriched in 6 signal pathways and 15 diseases
Supplementary Table 6 shows the pathways in which Raf1 interacting proteins were most significantly enriched based on KEGG pathway analysis. The results were divided into three categories: biological functions, cellular signaling pathways, and diseases. Interacting proteins were most enriched in the following biological functions: cell cycle, oocyte meiosis, gap junctions, long-term potentiation, long-term depression, progesterone-mediated oocyte maturation, and pathogenic Escherichia coli infection. Raf1 interacting proteins were significantly enriched in the ErbB, FoxO, PI3K-Akt, Hippo, insulin, and estrogen signaling pathways. Regarding human diseases, Raf1-interacting proteins were mainly enriched in different cancers, including endometrial cancer, renal cell carcinoma, glioma, prostate cancer, thyroid cancer, melanoma, bladder cancer, leukemia, and non-small cell lung cancer. Viral infections were also enriched, including hepatitis C/B, Epstein-Barr virus, and viral carcinogenesis.
According to Ingenuity Canonical Pathways analysis and Ingenuity Diseases analysis, Raf1 interacting proteins were enriched to 157 signal pathways and 18 diseases due to its broad classification criteria (Supplementary Tables 7, 9).
DISCUSSION
The goal of the present work was to decode Raf1 functions and get a global review of its biological role in cell processes, especially in carcinogenesis. We identified 198 Raf1 interacting proteins using in vitro VSV-tagging (Figure 2) and 14 of these had been reported previously reported to interact with Raf1. Among these 14 proteins, six belong to the 14-3-3 family (gene names: YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, and YWHAZ), which is well known to regulate Raf1 activity by phosphorylation and which is a highly conserved protein family that regulates many cellular processes such as proliferation, differentiation, apoptosis, and the cell cycle [17]. Three of the 14 previously identified binding proteins belong to the molecular chaperones (gene name: HSP90AA1, HSP90AB1, and CDC37), which are necessary for stabilizing the tertiary structure of Raf1 [18]. As the signal generator of Raf1, K/N/HRAS proteins were found to interact with Raf1 at high abundance in the interaction databases. The protein phosphatase PP2Aα, which is responsible for Raf1 inactivation, was also identified as an important Raf1-interacting protein [19]. The activation of Raf1 relies on Ras-dependent dimerization, including both homo- and hetero-dimerization [20]. Thus, we have also detected all three Raf variants among the Raf1-interacting proteins. Actually, two addition Raf1-interacting proteins recorded in Ingenuity Knowledge Base are A-Raf and Raf1 itself (Supplementary Table 8). These data suggest we may find new Raf1 functions from newly identified 182 proteins.
We verified our LC-MS data by confirming the interaction of selected binding partners with Raf1 by western blotting. To avoid selection bias, we chose proteins with different cellular locations, including the plasma membrane (IRS4, BAG4, CDK1 and PSMC2), mitochondria (AIF and ANT2), endoplasmic reticulum (GRP78 and RCN1/2), nucleus (KAP1, RUVB1/2, SMC3, PARP and TIM50), and peroxisomes (FAR1). In addition, selected proteins had independent roles in apoptosis, cell metabolism, cell proliferation, cell adhesion, DNA repair, protein scaffolding, and embryonic development [21-29]. Furthermore, we had access to good quality antibodies to detect these proteins. We confirmed an interaction with Raf1 for all selected proteins except AKAP8L, NDUFA4 and HAX1. This may be explained by the low sensitivity of the western blotting assay.
GO enrichment analysis revealed that many Raf1 interacting partners are involved in programmed cell death, cellular stress responses, and cell-cell adhesion. These findings may help to elucidate how Raf1 regulates these processes. We also found that Raf1 interacts with FAR1 and IRS4, suggesting that Raf1 may participate in lipid metabolism by protein interacting [28]. KEGG pathway analysis (Supplementary Table 6) indicated that Raf1 is involved in 10 biological functions, six cell signaling pathways, and 14 human diseases. Some of these findings were not reported previously and may provide novel insights into Raf1 functions.
To gain further insight into the role of Raf1 in cancer, we investigated whether the identified Raf1 interacting proteins are involved in the ten hallmarks of cancer (described in Table 1). Our findings showed that Raf1 sustains cell proliferation through interactions with proteins involved in hormone responses, growth signal transduction, and evasion of growth suppressors. Furthermore, Raf1 promotes resistance to apoptosis and autophagy in cancer cells, and enhances replicative immortality by regulating telomere maintenance. Raf1 has also been implicated in organ regeneration, therefore may induce angiogenesis, which is essential for tumor growth. The invasion and metastasis of cancer cells is controlled by changes in cell-cell adhesion. Therefore, the regulation of cell-cell adhesion by Raf1-interacting proteins may explain how Raf1 promotes tumor malignancy. Genome instability and gene mutations are hallmarks of cancer. Our findings indicated that Raf1 can promote DNA repair and chromosome organization through interactions with novel binding partners. We also observed that Raf1 interacting partners are involved in immune response activation and the response to interleukins. These interactions may allow host cells to avoid immune destruction and respond to tumor-promoting inflammation. Furthermore, Raf1 interactions control cellular catabolism and ATP metabolism to maintain the strong proliferative ability of cancer cells. Taken together, these findings suggest that Raf1 plays a more widespread role in carcinogenesis than was previously reported.
KEGG pathway analysis showed that novel Raf1-interacting proteins are also involved in cell signaling pathways and the development of various diseases, including endometrial cancer, which has not been reported previously. We found that the Raf1-interacting proteins AIFM1, DDX17, DDX5, DNAJA1, HNRNPU, HSPD1, KRAS, PARP1, RBM14, RPL32, and YWHAH (gene name) may contribute to the response to steroid hormones, especially estrogen. This suggests a mechanism for Raf1 in endometrial cancer.
In conclusion, we have identified 198 Raf1 interacting proteins and verified 17 of these by western blotting. The identified Raf1 interacting proteins were involved in all hallmarks of cancer, highlighting the importance of Raf1 in carcinogenesis. Much remains to be learned about Raf1 functions and the present study has identified putative candidates for future investigations. Furthermore, our findings provide a useful overview of Raf1 function from a systematic biology perspective.
MATERIALS AND METHODS
Materials
Doxycycline, monoclonal mouse anti-VSV-G antibody, and anti-VSV-G agarose beads were purchased from Sigma-Aldrich (Gillingham, United Kingdom). Lipofectamine 2000 transfection reagent was purchased from Invitrogen (Paisley, United Kingdom). Protease inhibitor cocktail tablets were purchased from Millipore (Massachusetts, United States). Anti-BAG4, anti-FAR1, anti-PP2Aα, anti-TIM50, anti-AKAP8L, anti-PSMC2, anti-SMC3 and anti-GAPDH antibodies were purchased from Santa Cruz (Heidelberg, Germany). Anti-Raf1, anti-AIF, anti-HAX1, and anti-GRP78 antibodies were purchased from BD Biosciences (Oxford, United Kingdom). The anti-ANT2 antibody was purchased from Cell Signaling Technology (Hitchin, Hertfordshire, United Kingdom). Anti-IRS4, anti-KAP1, anti-RCN1, anti-NDUFA4 and anti-RCN2 antibodies were purchased from Abcam (Cambridge, United Kingdom). Anti-VSV antibody was from sigma Sigma-Aldrich (Gillingham, United Kingdom). Anti-RUVB1 and anti- RUVB 2 antibodies were from Proteintech (Illinois, United States).
Cell culture
Flp-In To-Rex HEK293 cells were maintained in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% (vol/vol) fetal bovine serum (FBS), 100 μg/mL zeocin, and 15 μg/mL blasticidin. Stable Flp-In To-Rex HEK293 cells inducibly expressing VSV-Raf1 were maintained in DMEM supplemented with 10% (vol/vol) FBS, 50 μg/mL hygromycin, and 15 μg/mL blasticidin.
Generation of stable Flp-In TO-REx HEK293 cells inducibly expressing VSV-Raf1
VSV-Raf1 was amplified by PCR and inserted between BamHI and XhoI sites of pcDNA5/FRT/TO. The primers were as follows: Forward (VSV-G sequence and BamHI site underlined) CG GGATCC GCC ACC ATG TAC ACC GAT ATA GAG ATG AAC CGC CTT GGA AAG GAG CACAT ACA GGG AGC TTG GAA; reverse (XhoI site underlined) CCG CTCGAG CTA GAA GAC AGG CAG CCT CGG. To generate stable Flp-In To-Rex HEK293 cells that inducibly express VSV-Raf1, cells were transfected with a mixture of VSV-Raf1 cDNA in pcDNA5/FRT/TO vector and the pOG44 vector (1:9) using Lipofectamine 2000 transfection reagent, according to the manufacturer’s instructions. Resistant clones were selected by replacing zeocin with 200 μg/mL hygromycin B. VSV-Raf1 expression was induced by treating cells with 1 μg/mL doxycycline for 24 h and confirmed by western blotting using an anti-VSV antibody.
Identification of VSV-Raf1-interacting proteins by mass spectrometry
VSV-Raf1 was induced by 1 μg of doxycycline/ml in Flp-in To-REx HEK293 cells. After 24 hours of induction, cells were harvested and resuspended in immunoprecipitation buffer (150 mM NaCl, 0.01 mM NaPO4, 2 mM EDTA, 0.5% Triton X-100, 5% glycerol, and protease inhibitor cocktail tablets). Doxycycline induction was performed in Flp-In To-REx 293 cells transfected with a pcDNA5 empty vector as a negative control. The cell pellets were lysed and centrifuged for 15 min at 20,000 g at 4°C, and the supernatant was transferred to a fresh tube. Equal amounts of protein were incubated with anti-VSV-G agarose beads (Sigma) at 4°C for 3 h on a rotating wheel. After incubation, samples were washed four times with immunoprecipitation buffer. Proteins were eluted from the beads by vortexing and boiling for 7 min in SDS buffer. Raf1 interacting proteins were detected and analyzed as previously described. Briefly, protein samples were digested with trypsin overnight at 37°C. Then, the peptide mixture was desalted and concentrated using a Peptide Microtrap (MW0.5–50 kDa, 0.5mm × 2 mm, Michrom Bioresources, CA, USA). The prepared sample (10 μL) was loaded onto a hybrid linear ion trap (LTQ) Orbitrap mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA) equipped with ADVANCE Spray Source (Michrom Bioresources, CA) and reversed-phase capillary column (0.1 mm × 150 mm, packed with 5 μm 100Å Magic C18 resin, Michrom Bioresources, CA, USA) with an auto-sampler (HTS-PAL, CTC Analytics, Zwingen, Switzerland) at a flow rate of 1 μL/min for 15 min. Other parameters and conditions were as previously described [30].
Western blot analysis
We performed SDS-PAGE and immunoblotting as previously described [31]. For each sample, 30 μg of protein was separated by SDS-PAGE on 10–15% gels and transferred onto polyvinylidene difluoride membranes. Membranes were blocked in skimmed milk overnight at 4˚C before incubating with primary antibodies followed by peroxidase-conjugated anti-mouse or anti-rabbit IgGs. The epitopes were detected using an enhanced chemiluminescence western blot detection kit.
GO enrichment and KEGG pathway analysis
GO enrichment and KEGG pathway analysis were performed using the DAVID (https://david.ncifcrf.gov/) online tool. A P value of <0.05 was considered statistically significant. GO analysis revealed a significant enrichment in biological processes (BP), molecular function (MF), and cell component (CC) terms. KEGG (http://www.genome.jp/kegg/) systematically analyzed gene functions and linked genomic information with higher-order functional information. Additional functional annotations of the identified proteins were performed using the Ingenuity pathway analysis software.
Integration of the protein–protein interaction (PPI) network
The Search Tool for the Retrieval of Interacting Genes (STRING) database is an online tool designed for evaluating protein–protein interaction (PPI) information. STRING (version 10.0) covers 9,643,763 proteins from 2031 organisms. To evaluate the interactive relationships of Raf1-interacting proteins, we mapped Raf1 to STRING. We only experimentally validated interactions with a combined score of >0.9, which were selected as significant. Then, PPI networks were constructed using Cytoscape software (version 3.4.0).
Abbreviations
Raf, RAF proto-oncogene serine/threonine-protein kinase; MAPK, mitogen-activated protein kinase; MEK, mitogen-activated protein kinase kinase; ERK, extracellular signal regulated kinase; PP2A, protein phosphatase 2A; HSP90, heat shock protein 90; CDC37, cell division cycle 37; PAK, p21-activated kinase; STK3, serine/threonine kinase 3; PKC, protein kinase C; LC-MS, liquid chromatograph mass spectrometer; AKT, serine/threonine kinase 1; AIF, apoptosis inducing factor; FAR1, fatty acyl-CoA reductase1; KAP1, transcription intermediary factor 1-beta; RCN1/2, reticulocalbin-1/2; TIM50, mitochondrial import inner membrane translocase subunit TIM50; ANT2, ADP/ATP translocase 2; BAG4, BAG family molecular chaperone regulator 4; GRP78, 78 kDa glucose-regulated protein; IRS4, insulin receptor substrate 4; RUVB1, RuvB-like 1; RUVB2, RuvB-like 2; AKAP8L, A-kinase anchor protein 8-like; HAX1, HCLS1-associated protein X-1; CDK1, cyclin-dependent kinase 1; PARP, poly (ADP-ribose) polymerase 1; PSMC2, 26S protease regulatory subunit 7; SMC3, structural maintenance of chromosomes protein 3; NDUFA4, cytochrome c oxidase subunit NDUFA4.
Author contributions
YH and TX initiated and wrote most of the paper; XG, BH and XMZ contributed the revision of the paper; SA, XYZ, YY, YL and QH took part in writing and language editing of the paper. All authors approved the final vision of the manuscript.
ACKNOWLEDGMENTS
This work was funded by NSFC funding (U1302225, 81460417, 81460253, 81473342, 81560455), High-End Talent Grant of Yunnan Province, China (2012HA008) and Scientific Research Grant of Education Department of Yunnan Province, China (2015Y046). The work is partial conducted in University Based Provincial Key Laboratory of Screening and Utilization of Targeted Drugs.
CONFLICTS OF INTEREST
No conflicts of interest was declared.
REFERENCES
1. Kern F, Niault T, Baccarini M. Ras and Raf pathways in epidermis development and carcinogenesis. Br J Cancer. 2011; 104:229-234.
2. Odabaei G, Chatterjee D, Jazirehi AR, Goodglick L, Yeung K, Bonavida B. Raf-1 kinase inhibitor protein: structure, function, regulation of cell signaling, and pivotal role in apoptosis. Adv Cancer Res. 2004; 91:169-200.
3. Maurer G, Tarkowski B, Baccarini M. Raf kinases in cancer-roles and therapeutic opportunities. Oncogene. 2011; 30:3477-3488.
4. Lavoie H, Therrien M. Regulation of RAF protein kinases in ERK signalling. Nat Rev Mol Cell Biol. 2015; 16:281-298.
5. Kozak C, Gunnell MA, Rapp UR. A new oncogene, c-raf, is located on mouse chromosome 6. J Virol. 1984; 49:297-299.
6. Kyriakis JM, App H, Zhang XF, Banerjee P, Brautigan DL, Rapp UR, Avruch J. Raf-1 activates MAP kinase-kinase. Nature. 1992; 358:417-421.
7. Desideri E, Cavallo AL, Baccarini M. Alike but different: RAF paralogs and their signaling outputs. Cell. 2015; 161:967-970.
8. Rommel C, Radziwill G, Lovric J, Noeldeke J, Heinicke T, Jones D, Aitken A, Moelling K. Activated Ras displaces 14-3-3 protein from the amino terminus of c-Raf-1. Oncogene. 1996; 12:609-619.
9. Jaumot M, Hancock JF. Protein phosphatases 1 and 2A promote Raf-1 activation by regulating 14-3-3 interactions. Oncogene. 2001; 20:3949-3958.
10. Grammatikakis N, Lin JH, Grammatikakis A, Tsichlis PN, Cochran BH. p50(cdc37) acting in concert with Hsp90 is required for Raf-1 function. Mol Cell Biol. 1999; 19:1661-1672.
11. Zang M, Hayne C, Luo Z. Interaction between active Pak1 and Raf-1 is necessary for phosphorylation and activation of Raf-1. J Biol Chem. 2002; 277:4395-4405.
12. Romano D, Matallanas D, Weitsman G, Preisinger C, Ng T, Kolch W. Proapoptotic kinase MST2 coordinates signaling crosstalk between RASSF1A, Raf-1, and Akt. Cancer Res. 2010; 70:1195-1203.
13. Liu R, Liu T, Wei W, Guo K, Yang N, Tian S, Zhu J, Liu Y, Zhou W, Yang H. Novel interacting proteins identified by tandem affinity purification coupled to nano LC-MS/MS interact with ribosomal S6 protein kinase 4 (RSK4) and its variant protein (RSK4m). Int J Biol Macromol. 2017; 96:421-428.
14. Liu B, Zheng Y, Wang TD, Xu HZ, Xia L, Zhang J, Wu YL, Chen GQ, Wang LS. Proteomic identification of common SCF ubiquitin ligase FBXO6-interacting glycoproteins in three kinds of cells. J Proteome Res. 2012; 11:1773-1781.
15. Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000; 100:57-70.
16. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011; 144:646-674.
17. Fischer A, Baljuls A, Reinders J, Nekhoroshkova E, Sibilski C, Metz R, Albert S, Rajalingam K, Hekman M, Rapp UR. Regulation of RAF activity by 14-3-3 proteins: RAF kinases associate functionally with both homo- and heterodimeric forms of 14-3-3 proteins. J Biol Chem. 2009; 284:3183-3194.
18. Du J, Zeng J, Ou X, Ren X, Cai S. Methylglyoxal downregulates Raf-1 protein through a ubiquitination-mediated mechanism. Int J Biochem Cell Biol. 2006; 38:1084-1091.
19. Raabe T, Rapp UR. Ras signaling: PP2A puts Ksr and Raf in the right place. Curr Biol. 2003; 13:R635-R637.
20. Lavoie H, Li JJ, Thevakumaran N, Therrien M, Sicheri F. Dimerization-induced allostery in protein kinase regulation. Trends Biochem Sci. 2014; 39:475-486.
21. Ikink GJ, Boer M, Bakker ER, Hilkens J. IRS4 induces mammary tumorigenesis and confers resistance to HER2-targeted therapy through constitutive PI3K/AKT-pathway hyperactivation. Nat Commun. 2016; 7:13567.
22. Gao SP, Sun HF, Jiang HL, Li LD, Hu X, Xu XE, Jin W. Loss of TIM50 suppresses proliferation and induces apoptosis in breast cancer. Tumour Biol. 2016; 37:1279-1287.
23. Du Y, Huang Y, Gao Y, Song B, Mao J, Chen L, Bai L, Tang J. Annexin A7 modulates BAG4 and BAG4-binding proteins in mitochondrial apoptosis. Biomed Pharmacother. 2015; 74:30-34.
24. Kramann N, Hasenfuss G, Seidler T. B-RAF and its novel negative regulator reticulocalbin 1 (RCN1) modulates cardiomyocyte hypertrophy. Cardiovasc Res. 2014; 102:88-96.
25. Cook KL, Clarke PA, Clarke R. Targeting GRP78 and antiestrogen resistance in breast cancer. Future Med Chem. 2013; 5:1047-1057.
26. Iyengar S, Farnham PJ. KAP1 protein: an enigmatic master regulator of the genome. J Biol Chem. 2011; 286:26267-26276.
27. Izumi N, Yamashita A, Iwamatsu A, Kurata R, Nakamura H, Saari B, Hirano H, Anderson P, Ohno S. AAA+ proteins RUVBL1 and RUVBL2 coordinate PIKK activity and function in nonsense-mediated mRNA decay. Sci Signal. 2010; 3:ra27.
28. Honsho M, Asaoku S, Fujiki Y. Posttranslational regulation of fatty acyl-CoA reductase 1, Far1, controls ether glycerophospholipid synthesis. J Biol Chem. 2010; 285:8537-8542.
29. Joza N, Pospisilik JA, Hangen E, Hanada T, Modjtahedi N, Penninger JM, Kroemer G. AIF: not just an apoptosis-inducing factor. Ann N Y Acad Sci. 2009; 1171:2-11.
30. Zhang X, Wang T, Zhang H, Han B, Wang L, Kang J. Profiling of drug binding proteins by monolithic affinity chromatography in combination with liquid chromatography-tandem mass spectrometry. J Chromatogr A. 2014; 1359:84-90.
31. Huang Y, Yang X, Xu T, Kong Q, Zhang Y, Shen Y, Wei Y, Wang G, Chang KJ. Overcoming resistance to TRAIL-induced apoptosis in solid tumor cells by simultaneously targeting death receptors, c-FLIP and IAPs. Int J Oncol. 2016; 49:153-163.











