INTRODUCTION
Lung cancer is a common malignant cancer worldwide, which makes up about 18.2% of all malignant tumor mortality in the world [1, 2]. It is estimated that 222,500 new cases will be diagnosed with lung cancer and caused about 155,870 people’s death according to the cancer statistics 2017 [3]. Non-small cell lung cancer (NSCLC), which consists predominantly of squamous cell carcinoma (SCC) and adenocarcinoma (AD), is the most common type of lung cancer [4]. Despite surgical resection, systemic chemotherapy and targeted drugs can treat lung cancer patients successfully, the prognosis of NSCLC still remains poor owing to the late diagnosis of patients [5, 6]. Therefore, it is of great importance to discover markers specific for lung cancer in order to provide better individualized treatment.
Silent mating-type information regulation 2 homologue 1 (sirtuin1; SIRT1) is the closest mammalian homologue of yeast Sir2 and is an NAD-dependent protein deacetylase, which regulates life span in accordance with nutritional provisions under conditions of stress [7]. It is reported that SIRT1 played important roles in many biological processes, including aging, DNA repair, apoptosis and cell senescence [8–10]. Nowadays, many studies demonstrated that aberrant expression of SIRT1 was found in various cancers, containing hepatocellular carcinoma [11, 12], gastric carcinoma [13], breast carcinoma [14] and colorectal carcinoma [15, 16], indicating that SIRT1plays important roles in the initial and progress of cancers.
To date, abundant researches have confirmed that SIRT1 was abnormally expressed in NSCLC. The overexpression of SIRT1 is significantly correlated with high pathological T-stage and lymph node metastasis in NSCLC [17]. SIRT1 increases vessel density through facilitating endothelial cell branching and proliferation and promotes lung tumor growth through down-regulation of DLL4/Notch signaling and deacetylation of Notch1 intracellular domain [18], the Inactivation of SIRT1 can exert its antitumor activities through the tumor suppressor p27 in NSCLC [19]. Moreover, SIRT1 could contribute to the development of lung cancer through TNF-α/β-catenin axis [20]. Recent studies have revealed that abnormally expressed SIRT1was associated with prognosis of NSCLC [17, 21–26]. Therefore, a hypothetical that there might be a significant correlation between the expression of SIRT1 and the clinical outcomes of NSCLC was speculated.
With the aim to evaluate the association between the SIRT1 expression and NSCLC clinical outcomes comprehensively, we carried out this meta-analysis of the published articles to explore the clinical value of SIRT1 in NSCLC.
RESULTS
Study selection and characteristics
Figure 1 showed the selection process of included studies. The electronic search identified 76 records from PubMed, EMBASE, and Web of Science and 24 articles were excluded due to duplicated publication. After we screened the titles and abstracts, 32 irrelevant articles were excluded. Subsequently, the 20 remaining full-text articles were assessed and 13 studies were further excluded according to the exclusion criteria. Finally, seven articles were included in the current meta-analysis [17, 21, 22, 24–27]. All the included studies used FFPE tissue to detect the expression of SIRT1. As shown in Table 1, the studies were published from 2013 to 2016, and included 979 patients with the highest number of 295 and the smallest number of 40. All the studies used immunolytic enzyme histochemistry technology to evaluate the expression level of SIRT1. Among the seven studies, three are from China [17, 25, 27], one from Japan [24], one from USA [21], one from Iran [22] and one from Spain [26].
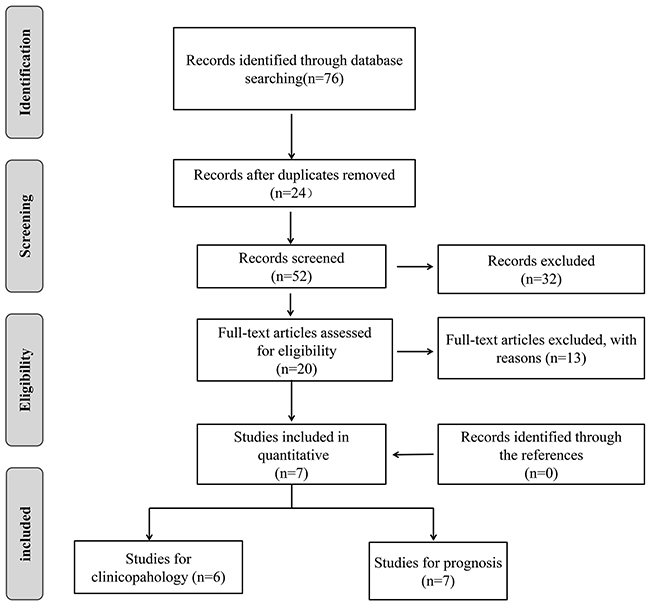
Figure 1: Flow diagram of study search and selection process.
Table 1: Characteristics of the included studies in this meta-analysis
Author |
Year |
Country |
Total number |
Histologic grade (W and M/Poorly) |
Method |
Type of survival data |
Follow up time (month) |
HR |
NOS score |
|---|---|---|---|---|---|---|---|---|---|
Noh |
2013 |
USA |
144 |
105/39 |
IHC |
OS |
NR |
Reported |
7 |
Gharabaghi |
2016 |
Iran |
40 |
22/18 |
IHC |
OS |
50 |
Reported |
7 |
Lin |
2016 |
Japan |
260 |
61/199 |
IHC |
OS |
125 |
Reported |
8 |
Li |
2015 |
China |
75 |
- |
IHC |
OS |
120 |
Reported |
8 |
Zhang |
2013 |
China |
295 |
242/53 |
IHC |
OS |
25 |
Reported |
7 |
Grbesa |
2015 |
Spain |
105 |
42/57 |
IHC |
OS |
144 |
Survival Curve |
8 |
Wang |
2016 |
China |
60 |
43/17 |
IHC |
OS |
100 |
Reported |
7 |
IHC: immunohistochemistry; HR: hazard ratio; W, well; M, moderate
Prognostic value of SIRT1 expession in NSCLC
There are 7 studies [17, 21, 22, 24–27] investigated the relationship betweenSIRT1 expression and overall survival (OS) with a total number of 979 NSCLC patients. We then analyzed the association between the expression of the SIRT1 and patient survival. The pooled HR for OS showed that SIRT1 was associated with poor prognosis in NSCLC patients (HR=1.99, 95% CI: 1.33-2.98, P=0.0009 random-effects) (Figure 2A). Subsequent analyses of subgroup based on Asian and non-Asian were performed. Result revealed that SIRT1 expression was significantly related to OS in Asian (HR= 2.41, 95% CI=1.50-3.89, P<0.01, random-effect), but not in non-Asian (HR=1.22, 95% CI=0.75-1.98, P=0.42, random-effect). (Figure 2B).
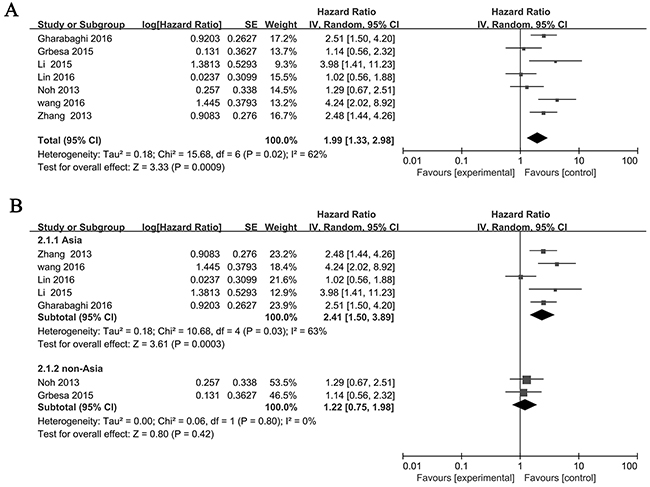
Figure 2: Forest plot of HRs for the association of SIRT1 expression in NSCLC with OS. (A) All NSCLC patients, (B) Asian and non-Asian NSCLC patients. The point estimate is bounded by a 95% confidence interval, and the perpendicular line represents no increased risk for the outcome. NSCLC: non-small cell lung cancer.
Association between SIRT1 and histological grade
Six studies investigated the correlation between the expression of SIRT1 and histological grade, so we explored whether an association between SIRT1 expression and histological grade was existed. The pooled OR from 383 poorly and 515 well and moderately differented is shown in Figure 3A (OR= 2.00, 95% CI= 1.05–3.78, P= 0.03, random-effects). Subgroup analyses were done based on on Asian and non-Asian. We found that SIRT1 expression was more frequent in patients that are poorly differented in Asian (OR= 2.51, 95% CI= 1.55–4.07, P<0.01) but not tend to be frequent in non-Asian (OR= 1.31, 95% CI= 0.28–6.20, P= 0.74) (Figure 3B).
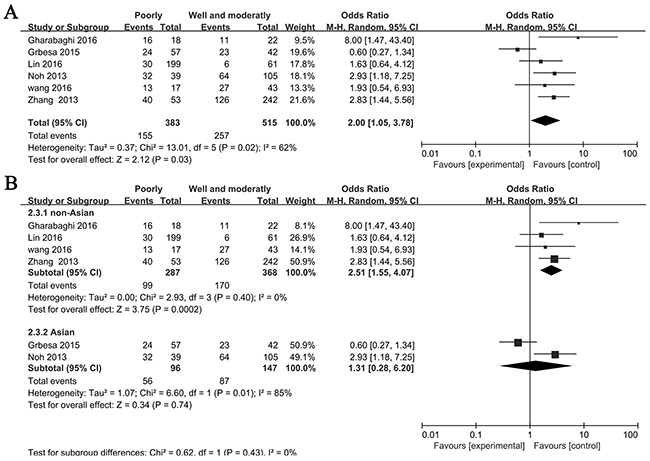
Figure 3: Forest plot for association between SIRT1 and histological grade. (A) All NSCLC patients, (B) Asian and non-Asian NSCLC patients.
Sensitivity analyses and publication bias
Stata11.0 software was used to perform sensitivity analysis to assess the stability of our results. The results indicated that individual study had little influence on our final results, which validated the stablity and crediblity of our results (Figure 4). Both Begg’s test and Egger’s test were used to evaluate the publication bias. The results indicated no publication bias existed in all subgroups since all the values of P > 0.05 (Figure 5).
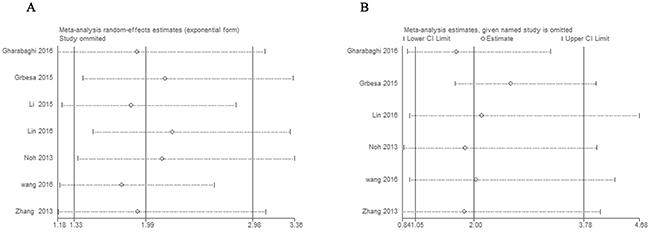
Figure 4: Sensitivity analyses of the studies. (A) Overall survival, (B) histological grade.
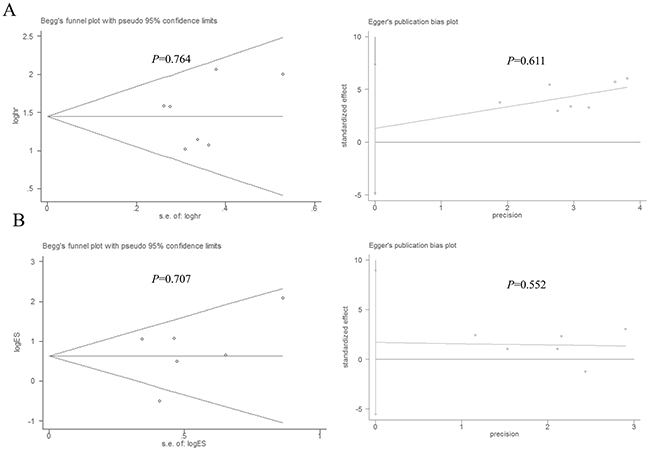
Figure 5: Begg’s and Egger’s test for publication bias. (A) Overall survival, (B) histological grade.
DISCUSSION
NSCLC is one of the leading causes of cancer-related death worldwide [28]. The overall prognosis of NSCLC is poor and it has a high risk of recurrence after surgical resection [29]. Thus it is urgently needed to identify new biomarkers for NSCLC. SIRT1, homologue of the yeast Sir2 protein, is the most researched sirtuin in the mammalian sirtuin family [30]. It has been commonly accepted that SIRT1 plays critical roles in many important biological processes such as cell proliferation, cell death, senescence and stress response [26]. Numrous studies reported the involvement of SIRT1 in NSCLC [31–33]. Xie et al. showed that overexpression of SIRT1 in transgenic mice facilitated endothelial cell branching, thus increasing vessel density and lung tumor growth [18]. Cha et al. [34] found that celecoxib and sulindac can inhibit TGF-β1-induced EMT and suppress lung cancer cell migration and invasion via downregulation of SIRT1. In addition, aberant expresion of SIRT1 was reported to be associated with clinical and progress of NSCLC patients.
The role of SIRT1 in NSCLC is debatable. A large number of studies have reported that SIRT1 was high expressed in NSCLC. For exmple, SIRT1 was reported to be a tumor promoter in lung adenocarcinoma [23] and can enhance β-catenin accumulation then to activate TNF-α transcription, leading to sustained inflammation and lung cancer development [20]. On the other hand, SIRT1 activation has been reported to hamper lung cancer metastasis [33] and Beane et al. [35] validated that the activity of SIRT1 was down-regulated in tumors from smokers. Owing to the controversial role of SIRT1 in NSCLC, it is of great value to perform this meta-analysis of the current literature to gain a better understanding between SIRT1 and NSCLC.
In the present meta-analysis, we found that the increased expressions of SIRT1 were associated with poor prognosis, which is in consistent with four of the included studies. However, the remaning three articles showed no corelation betwwen SIRT1 and OS of NSCLC. When we did subgroup analysis, we found that SIRT1 expression was related to OS in Asian rather than in non-Asian, indicating that the discrepency may be due to different origin of the samples. Then, we explored the relation between SIRT1 expression and histological grade. We found that SIRT1 tend to be highly expressed in poorly differented NSCLC patients, indicating a tumorgenesis role of SIRT1 in NSCLC.
It should be noted that there were limitations in our analysis. Firstly, the definition of SIRT1 positive expression varied between the selected studies, which may reduce the reliability of our study; secondly, since only English language papers were included, the data collection may be incomplete; thirdly, the HR value from 1 study was got from KaplaneMeier survival curve instead of the given data, which may generate inaccurate results; fourthly, some studies distinguished the expression location of SIRT1 but somes didn’t, which may reduce the credibility of the study; fifthly, the sample size maybe too small to fully confirm the association between SIRT1 and clinicopathological characteristics, which needs more studies. Despite the limitations mentioned above, our study is the first meta-analysis to explore the association between SIRT1 expression and clinical features of NSCLC. Importantly, we utilized a statistical approach to combine the results of multiple studies, achieving an objective result on the basis of strict inclusion and exclusion criteria.
In summary, our study was the first to evaluate the correlation between SIRT1 and clinical values of patients with NSCLC. We revealed that SIRT1 expression was related to histological grade. Besides, we found that SIRT1 can be used as potential prognostic markers for NSCLC. However, considering the limitations of individual study, in future, large-scale and and good quality studies were needed to confirm our results.
MATERIALS AND METHODS
Search strategy
PubMed, EMBASE, and Web of Science were systematically searched in February 2017 using the search terms: “SIRT1”, “silent mating type information regulation 2 homolog-1”, “sirtuin 1” and “NSCLC”, “non-small cell lung cancer“, “LAD”, “lung adenocarcinoma”, “LSCC”, “lung squamous cell cancer”. Additionally, references of retrieved relevant articles were screened to identify potentially eligible literature. We did not limit our search by country, race and date.
Inclusion and exclusion criteria
Inclusion criteria: According to the principle of PICO (participant, intervention, comparison and outcomes) [36], studies were selected for the following criteria:
• Participants: patients diagnosed with NSCLC.
• Intervention: SIRT1 positive or highly expressed.
• Comparison: SIRT1 negative or low expressed.
• Outcomes: shorten in overall survival time, histological grade reviewed as poorly.
The exclusion criteria included: (1) studies without usable data; (2) duplicate publications; (3) sample cases fewer than 30; (4) reviews, letters, single case reports; (5) animal studies.
Data extraction and quality assessment
Two investigators extracted the data from eligible studies independently and the following information was collected: name of the first author, year of publication, number of cases, histopathological grade of tumors (moderately and well vs poorly), method, information about OS (HR value and follow-up duration). In our study, we defined OS as duration of time from Starting from randomization to death caused by any reason. Data for the characteristics of the studies were summarized and organized into a table format.
Newcastle-Ottawa Scale score (NOS) was used to assess the quality of the studies. The scale allocates a maximum of nine points for the quality of selection, comparability, exposure, and outcomes for study participants. The total scores ranged from 0 to 9, and a study with a score of 7 or more represents a good quality [37].
Statistical analysis
All analyses were conducted using the STATA software version 11.0 (Stata Corporation, College Station, Texas, USA) or Review Manager Version 5.3. HR and 95% CIs were used to assess the association between SIRT1expresion and OS in NSCLC, and ORs and 95% CIs were used to evaluate the relation between SIRT1 expression and clinical features. When the I2≤50%, the fixed effect model was used, otherwise, a random effect model was applied [38]. HRs with their 95% CIs were got directly from data in articles or from Kaplan–Meier survival curves using Engauge Digitizer version 4.1 [39]. Sensitivity analysis was conducted by omitting the study sequentially. Begg’s and Egger’s test were used for assessing the publication bias. Two-sided P < 0.05 was considered statistically significant.
Author contributions
Yifei Chen, Tao Wang and Wei Wang contributed for study search, data extraction and drafting; Jiahao Hu, Ruiting Li and Shaojun He carried out the statistical analysis; Jiong Yang critically revised the manuscript. All authors confirmed the final edition.
ACKNOWLEDGMENTS
This study was supported by National Natural Science Foundation of China (NSFC 81670024, to Yang J). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest. The authors alone are responsible for the content and writing of the paper.
REFERENCES
1. Torre LA, Siegel RL, Jemal A. Lung cancer statistics. Adv Exp Med Biol. 2016; 893: 1-19. https://doi.org/10.1007/978-3-319-24223-1_1.
2. Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010; 127: 2893-2917. https://doi.org/10.1002/ijc.25516.
3. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017; 67: 7-30. https://doi.org/10.3322/caac.21387.
4. Niklinski J, Niklinska W, Chyczewski L, Becker HD, Pluygers E. Molecular genetic abnormalities in premalignant lung lesions: biological and clinical implications. Eur J Cancer Prev. 2001; 10: 213-226.
5. Deng J, Liang Y, Liu C, He S, Wang S. The up-regulation of long non-coding RNA AFAP1-AS1 is associated with the poor prognosis of NSCLC patients. Biomed Pharmacother. 2015; 75: 8-11. https://doi.org/10.1016/j.biopha.2015.07.003.
6. Youlden DR, Cramb SM, Baade PD. The International Epidemiology of Lung Cancer: geographical distribution and secular trends. J Thorac Oncol. 2008; 3: 819-831. https://doi.org/10.1097/JTO.0b013e31818020eb.
7. Imai S, Armstrong CM, Kaeberlein M, Guarente L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature. 2000; 403: 795-800. https://doi.org/10.1038/35001622.
8. Bordone L, Guarente L. Calorie restriction, SIRT1 and metabolism: understanding longevity. Nat Rev Mol Cell Biol. 2005; 6: 298-305. https://doi.org/10.1038/nrm1616.
9. O’Hagan HM, Wang W, Sen S, Destefano Shields C, Lee SS, Zhang YW, Clements EG, Cai Y, Van Neste L, Easwaran H, Casero RA, Sears CL, Baylin SB. Oxidative damage targets complexes containing DNA methyltransferases, SIRT1, and polycomb members to promoter CpG Islands. Cancer Cell. 2011; 20: 606-619. https://doi.org/10.1016/j.ccr.2011.09.012.
10. Liu T, Liu PY, Marshall GM. The critical role of the class III histone deacetylase SIRT1 in cancer. Cancer Res. 2009; 69: 1702-1705. https://doi.org/10.1158/0008-5472.can-08-3365.
11. Hao C, Zhu PX, Yang X, Han ZP, Jiang JH, Zong C, Zhang XG, Liu WT, Zhao QD, Fan TT, Zhang L, Wei LX. Overexpression of SIRT1 promotes metastasis through epithelial-mesenchymal transition in hepatocellular carcinoma. BMC Cancer. 2014; 14: 978. https://doi.org/10.1186/1471-2407-14-978.
12. Li Y, Xu S, Li J, Zheng L, Feng M, Wang X, Han K, Pi H, Li M, Huang X, You N, Tian Y, Zuo G, et al. SIRT1 facilitates hepatocellular carcinoma metastasis by promoting PGC-1alpha-mediated mitochondrial biogenesis. Oncotarget. 2016; 7: 29255-29274. https://doi.org/10.18632/oncotarget.8711.
13. Noguchi A, Kikuchi K, Zheng H, Takahashi H, Miyagi Y, Aoki I, Takano Y. SIRT1 expression is associated with a poor prognosis, whereas DBC1 is associated with favorable outcomes in gastric cancer. Cancer Med. 2014; 3: 1553-1561. https://doi.org/10.1002/cam4.310.
14. Wu M, Wei W, Xiao X, Guo J, Xie X, Li L, Kong Y, Lv N, Jia W, Zhang Y, Xie X. Expression of SIRT1 is associated with lymph node metastasis and poor prognosis in both operable triple-negative and non-triple-negative breast cancer. Med Oncol. 2012; 29: 3240-3249. https://doi.org/10.1007/s12032-012-0260-6.
15. Chen X, Sun K, Jiao S, Cai N, Zhao X, Zou H, Xie Y, Wang Z, Zhong M, Wei L. High levels of SIRT1 expression enhance tumorigenesis and associate with a poor prognosis of colorectal carcinoma patients. Sci Rep. 2014; 4: 7481. https://doi.org/10.1038/srep07481.
16. Jung W, Hong KD, Jung WY, Lee E, Shin BK, Kim HK, Kim A, Kim BH. SIRT1 expression is associated with good prognosis in colorectal cancer. Korean J Pathol. 2013; 47: 332-339. https://doi.org/10.4132/KoreanJPathol.2013.47.4.332.
17. Zhang T, Rong N, Chen J, Zou C, Jing H, Zhu X, Zhang W. SIRT1 expression is associated with the chemotherapy response and prognosis of patients with advanced NSCLC. PLoS One. 2013; 8: e79162. https://doi.org/10.1371/journal.pone.0079162.
18. Xie M, Liu M, He CS. SIRT1 regulates endothelial Notch signaling in lung cancer. PLoS One. 2012; 7: e45331. https://doi.org/10.1371/journal.pone.0045331.
19. Zhu L, Chiao CY, Enzer KG, Stankiewicz AJ, Faller DV, Dai Y. SIRT1 inactivation evokes antitumor activities in NSCLC through the tumor suppressor p27. Mol Cancer Res. 2015; 13: 41-49. https://doi.org/10.1158/1541-7786.mcr-14-0239.
20. Lu J, Zhang M, Huang Z, Sun S, Zhang Y, Zhang L, Peng L, Ma A, Ji P, Dai J, Cui T, Liu H, Gao J. SIRT1 in B[a]P-induced lung tumorigenesis. Oncotarget. 2015; 6: 27113-27129. doi: 10.18632/oncotarget.4729.
21. Noh SJ, Baek HA, Park HS, Jang KY, Moon WS, Kang MJ, Lee DG, Kim MH, Lee JH, Chung MJ. Expression of SIRT1 and cortactin is associated with progression of non-small cell lung cancer. Pathol Res Pract. 2013; 209: 365-370. https://doi.org/10.1016/j.prp.2013.03.011.
22. Gharabaghi MA. Diagnostic investigation of BIRC6 and SIRT1 protein expression level as potential prognostic biomarkers in patients with non-small cell lung cancer. Clin Respir J. 2016. https://doi.org/10.1111/crj.12572.
23. Chen X, Hokka D, Maniwa Y, Ohbayashi C, Itoh T, Hayashi Y. Sirt1 is a tumor promoter in lung adenocarcinoma. Oncol Lett. 2014; 8: 387-393. https://doi.org/10.3892/ol.2014.2057.
24. Lin SY, Peng F. Association of SIRT1 and HMGA1 expression in non-small cell lung cancer. Oncol Lett. 2016; 11: 782-788. https://doi.org/10.3892/ol.2015.3914.
25. Li C, Wang L, Zheng L, Zhan X, Xu B, Jiang J, Wu C. SIRT1 expression is associated with poor prognosis of lung adenocarcinoma. Onco Targets Ther. 2015; 8: 977-984. https://doi.org/10.2147/ott.s82378.
26. Grbesa I, Pajares MJ, Martinez-Terroba E, Agorreta J, Mikecin AM, Larrayoz M, Idoate MA, Gall-Troselj K, Pio R, Montuenga LM. Expression of sirtuin 1 and 2 is associated with poor prognosis in non-small cell lung cancer patients. PLoS One. 2015; 10: e0124670. https://doi.org/10.1371/journal.pone.0124670.
27. Wang J, Wang C. Prognostic and predictive role of sirtuin1 expression in lung adenocarcinoma. Clin Lab. 2016; 62: 1989-1994. https://doi.org/10.7754/Clin.Lab.2016.160317.
28. Lindskog C, Edlund K, Mattsson JS, Micke P. Immunohistochemistry-based prognostic biomarkers in NSCLC: novel findings on the road to clinical use? Expert Rev Mol Diagn. 2015; 15: 471-490. https://doi.org/10.1586/14737159.2015.1002772.
29. Rami-Porta R, Crowley JJ, Goldstraw P. The revised TNM staging system for lung cancer. Ann Thorac Cardiovasc Surg. 2009; 15: 4-9.
30. Houtkooper RH, Pirinen E, Auwerx J. Sirtuins as regulators of metabolism and healthspan. Nat Rev Mol Cell Biol. 2012; 13: 225-238. https://doi.org/10.1038/nrm3293.
31. Liang Z, Yang Y, Wang H, Yi W, Yan X, Yan J, Li Y, Feng Y, Yu S, Yang J, Jin Z, Duan W, Chen W. Inhibition of SIRT1 signaling sensitizes the antitumor activity of silybin against human lung adenocarcinoma cells in vitro and in vivo. Mol Cancer Ther. 2014; 13: 1860-1872. https://doi.org/10.1158/1535-7163.mct-13-0942.
32. Sun Y, Sun D, Li F, Tian L, Li C, Li L, Lin R, Wang S. Downregulation of Sirt1 by antisense oligonucleotides induces apoptosis and enhances radiation sensitization in A549 lung cancer cells. Lung Cancer. 2007; 58: 21-29. https://doi.org/10.1016/j.lungcan.2007.05.013.
33. Sun L, Li H, Chen J, Dehennaut V, Zhao Y, Yang Y, Iwasaki Y, Kahn-Perles B, Leprince D, Chen Q, Shen A, Xu Y. A SUMOylation-dependent pathway regulates SIRT1 transcription and lung cancer metastasis. J Natl Cancer Inst. 2013; 105: 887-898. https://doi.org/10.1093/jnci/djt118.
34. Cha BK, Kim YS, Hwang KE, Cho KH, Oh SH, Kim BR, Jun HY, Yoon KH, Jeong ET, Kim HR. Celecoxib and sulindac inhibit TGF-beta1-induced epithelial-mesenchymal transition and suppress lung cancer migration and invasion via downregulation of sirtuin 1. Oncotarget. 2016; 7: 57213-57227. https://doi.org/10.18632/oncotarget.11127.
35. Beane J, Cheng L, Soldi R, Zhang X, Liu G, Anderlind C, Lenburg ME, Spira A, Bild AH. SIRT1 pathway dysregulation in the smoke-exposed airway epithelium and lung tumor tissue. Cancer Res. 2012; 72: 5702-5711. https://doi.org/10.1158/0008-5472.can-12-1043.
36. Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA Statement. Open Med. 2009; 3: e123-130.
37. Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol. 2010; 25: 603-605. https://doi.org/10.1007/s10654-010-9491-z.
38. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7: 177-188.
39. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials. 2007; 8: 16. https://doi.org/10.1186/1745-6215-8-16.

