INTRODUCTION
Acute myeloid leukemia (AML), one of the most common types of hematopoietic malignancy, is the leading cause of leukemia death in USA [1]. According to the statistics, the annual incidence rate of AML was 4.14 per 100,000 in USA (2009–2013) [1], and 4.39 per 100,000 in UK (2004–2013) [2]. There was no detailed prevalence of AML in China, but it was reported that the annual incidence rate of leukemia was 5.00 per 100,000 in China (2013) [3], and the annual incidence rates of AML was 1.35 per 100,000 in Nanjing (2003–2007) [4]. AML is more common in adults with the median age being 67 years old at diagnosis, and about 4% patients are children and adolescents [5]. With the development of chemotherapy and hematopoietic stem cell transplantation, the clinical outcomes of AML patients have been improved greatly in the past few decades [6, 7]. However, relapsed and refractory leukemia remain as the major causes of mortality in AML patients [8, 9]. Several criteria have been proposed to classify leukemia subtypes and predict prognosis, based on morphological features or different biomarkers identified via serology analysis, immunophenotyping or genetic profiling [10, 11]. These methods and criteria are important for the clinical diagnosis and treatment of leukemia [12]; however, one biomarker is not sufficient enough to distinguish leukemia subtypes for personalized therapeutic strategies. Therefore, it is of great importance to explore more reliable prognostic biomarkers and use them as combinations for accurate diagnosis and medical decision-making for AML patients.
MicroRNAs (miRNAs) are a class of highly conserved small noncoding RNAs (range from 18 to 24 nucleotides) in a variety of organisms, which regulate target gene expressions via degradation or inhibition of the target mRNAs translation upon binding to 3′-untranslated regions of target mRNAs [13–16]. Recently, accumulating investigations have identified the distinct expression patterns and biological functions of miRNAs [17–19], as well as their potential roles as diagnostic and prognostic biomarkers in leukemia [20, 21]. The microRNA-181 (miR-181) family includes 4 members (miR-181a, miR-181b, miR-181c and miR-181d), which are highly evolutionarily conserved across almost all vertebrates [22, 23]. Human miR-181a and miR-181b genes are located on chromosomes 1 (miR-181a1 and miR-181b1) and 9 (miR-181a2 and miR-181b2), whereas miR-181c and miR-181d genes are clustered closely on chromosome 19 [24]. A single nucleotide polymorphism (SNP), rs78086449, is existed in human pre-miRNA regions of miR-181b2 [25]. Recently, it has been demonstrated that the expression of miR-181 is related to the prognosis of AML patients [24, 26]. Li et al. showed that increased expression of miR-181a or miR-181b was significantly associated with longer overall survival (OS) in cytogenetically abnormal AML (CA-AML) patients [27], which was consistent with Schwind’s findings in cytogenetically normal AML (CN-AML) patients [28]. In contrast, Zhi et al. and Xiang et al. reported that high miR-181b expression was associated with poorer prognosis, as revealed by higher OS or reduced complete remission (CR), in AML patients [29, 30]. In addition, Butrym et al. also found that AML patients with lower expression of miR-181a showed longer survival rates [31]. The contradictory results among these studies made it difficult to assess the prognostic effect of miR-181 in AML patients.
To overcome the discrepancy and low reproducibility of individual studies evaluating prognostic value of miR-181 in AML patients, a meta-analysis was performed in the present study. Eligible studies on the roles of miR-181 in AML patients were identified and retrieved from databases of PubMed, Embase and Cochrane Library. Moreover, hazard ratios (HRs) and their corresponding 95% confidence intervals (CIs) were extracted from individual study and pooled together in overall meta-analysis to elucidate the prognostic value of miR-181 in AML patients. Our results indicate that miR-181 can be used as a prognostic biomarker in AML patients based on the origin of patient population.
RESULTS
Summary of analyzed studies
A flowchart of detailed searching process was illustrated in Figure 1. Using the three-step literature searching strategy, a total of 183 articles were initially identified from PubMed, Embase, and Cochrane Library databases. Following removal of 40 duplicate records, another 97 records that are conference abstracts, meta-analysis, reviews or non-relevant studies were excluded based on manual screening of the abstracts and titles. Full-text of the remaining 46 articles were further evaluated, and 6 articles were selected for the present meta-analysis based on our inclusion and exclusion criteria described in Methods. The public repositories including GEO, EBI ArrayExpress, and TCGA were also searched, but no eligible datasets were founded.
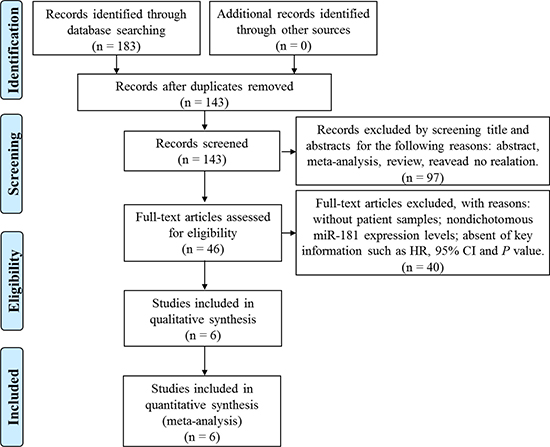
Figure 1: Flow diagram of the screening and selection process of studies.
The main characteristics and basic information of eligible studies were listed in Table 1. Due to the inability to acquire information concerning the race and ethnicity of the patients, these studies were analyzed according to the country of the patients. A total of 815 patients diagnosed as AML, CA-AML, CN-AML in China [29, 30, 32], Poland [31] and USA [27, 28] were enrolled in these studies. The clinical data on these patients were shown in Supplementary Table 1. Four studies used quantitative PCR (qPCR) to measure the expression levels of miR-181 (TaqMan: 2 and SYBR Green PCR: 2), and another 2 studies employed the microarray method. Li et al. analyzed all members of miR-181 family, and investigated two independent populations as a training set and a validation set [27]. Schwind et al. examined miR-181a expression in 187 younger (< 60 years) adults with CN-AML and 122 CN-AML patients with FLT3-internal tandem duplication (FLT3-ITD) and/or NPM1wt haplotypes (≥ 60 years) [28]. Zhi et al. estimated the OS of AML patients based on the expression levels of miR-181b-5p (high versus low expression group) in a training set, a validation set, and the combined set [29], and data of the combined set were extracted for analysis herein. Among these eligible studies, the prognostic role of miR-181 was evaluated in 13 datasets for OS, 3 datasets for CR, 2 datasets for disease-free survival (DFS), and 2 datasets for relapse-free survival (RFS).
Table 1: Main characteristics of eligible studies
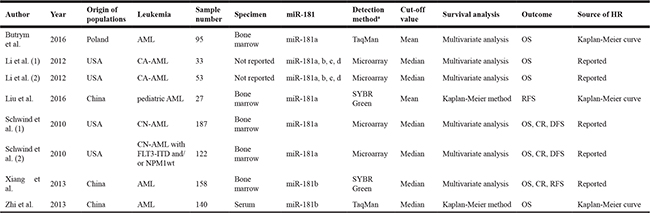
AML: acute myeloid leukemia; CA-AML: cytogenetically abnormal acute myeloid leukemia; CN-AML: cytogenetically normal acute myeloid leukemia; HR, hazard ratio; OS: overall survival; CR: complete remission; DFS: disease-free survival; RFS: relapse-free survival. aDetection method: both TaqMan and SYBR Green are quantitative real-time PCR methods.
Study quality was assessed by the Newcastle-Ottawa Quality Assessment Scale (NOS), which comprises the following three parameters of quality: the selection, the comparability and the outcome [33]. All of the selected 6 studies scored 1 or higher on each of the 3 parameters, and total scores ranged from 6 to 8 (Table 2). The study quality is considered as high if the total score is greater than 5 [34]. Hence, these studies are of a relatively high quality and are included in the final analysis.
Table 2: Quality assessment based on the newcastle–ottawa quality assessment scale
Author |
Year |
Selection |
Comparability |
Outcome |
Total score |
|---|---|---|---|---|---|
Butrym et al. |
2016 |
4 |
1 |
1 |
6 |
Li et al. |
2012 |
4 |
2 |
1 |
7 |
Liu et al. |
2016 |
4 |
1 |
1 |
6 |
Schwind et al. |
2010 |
4 |
2 |
2 |
8 |
Xiang et al. |
2013 |
4 |
1 |
1 |
6 |
Zhi et al. |
2013 |
4 |
1 |
1 |
6 |
Association of OS with miR-181 expression
The main results of this meta-analysis were displayed in Figure 2. Thirteen datasets including 788 patients were investigated to understand the relationship between miR-181 expression and OS in AML patients. As some HRs in studies were not directly reported and could not be obtained from the authors, Kaplan-Meier curves were used to extract data [35, 36]. For these datasets, the pooled HR and its 95% CI were calculated by a random-effects model (Figure 2A). Furthermore, seven subgroup analyses of OS were performed through classifying patients based on origin of population, specimen, member of miR-181 family, detection method, cut-off value, survival analysis and source of HR (Table 3). Subgroup analysis by origin of population showed that high miR-181 levels were significantly associated with a favorable OS in American patients (pooled HR: 0.74, 95% CI: 0.67–0.82, P < 0.01), but suggested a worse OS in Chinese patients (pooled HR: 1.78, 95% CI: 1.35–2.34, P < 0.01) (Table 3 and Figure 2B). There was no significant difference in relationship between expression levels of individual miR-181 family member and OS in the subgroup analysis in American patients (Figure 2C). Since there were only miR-181a expression in Polish patients and only miR-181b expression in Chinese patients, the subgroup analysis based on members of miR-181 family was not performed in Polish and Chinese patients.
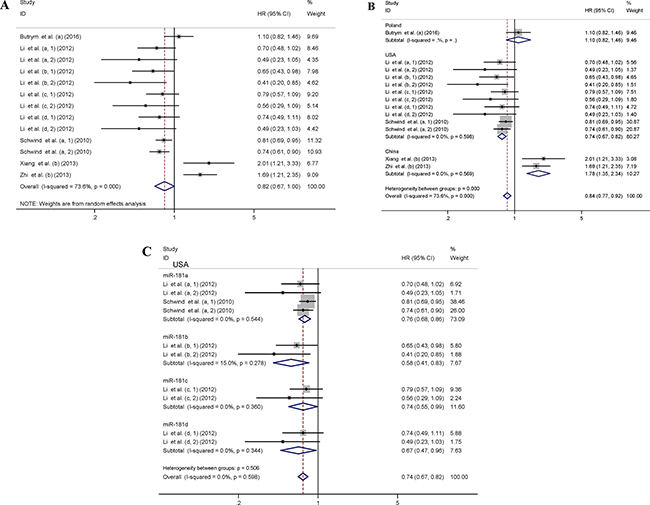
Figure 2: Forest plots of studies evaluating the pooled HR of elevated miR-181 levels for OS. Forest plot of the relationship between miR-181 expression and OS using a random-effects model (A), by subgroup analysis based on origin of population using a fixed-effect model (B), by subgroup analysis based on members of miR-181 family in American patients using a fixed-effect model (C). The lower-case letters a, b, c, d represent that miR-181a, miR-181b, miR-181c and miR-181d were analyzed separately in the studies.
Table 3: Subgroup analyses of the relationship between miR-181 expression and OS
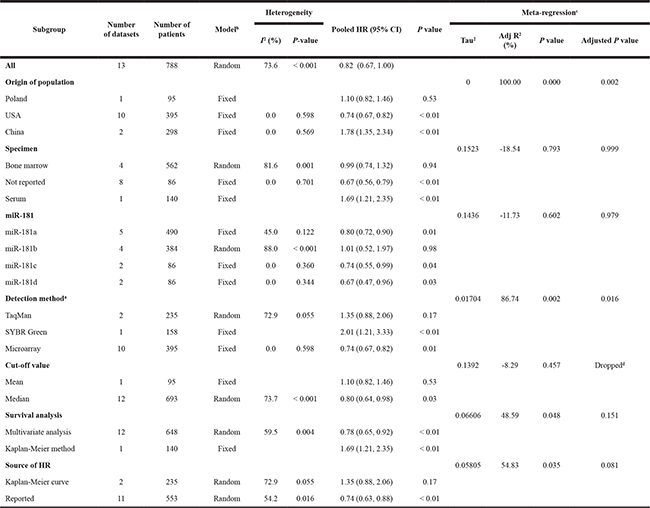
HR, hazard ratio; CI, confidence interval. aDetection method: both TaqMan and SYBR Green are quantitative real-time PCR methods. bModel: Fixed, fixed-effect model; Random, random-effects model. cMeta-regression was performed to explore possible explanations for heterogeneity with Monte Carlo permutation test (Permutations = 10,000). dSource of HR dropped due to collinearity in meta-regression with Monte Carlo permutation test.
Heterogeneity analysis
Obvious heterogeneity (I2 = 73.6%, P < 0.001) was discovered among the selected datasets evaluating the correlation between OS and miR-181 (Table 3 and Figure 2A). The higher significant heterogeneity was found in datasets with miR-181b (I2 = 88.0%, P < 0.001) and bone marrow specimen type (I2 = 81.6%, P = 0.001) (Table 3). However, the heterogeneity was significantly decreased in the subgroup analysis by origin of population. A meta-regression analysis was performed to investigate the sources of the heterogeneity in OS analysis group (Table 3), based on origin of population, specimen, member of miR-181 family, detection method, cut-off value, survival analysis and source of HR as covariates. All covariates were entered into the meta-regression model simultaneously, and the Monte Carlo permutation tests were performed for 10,000 times to acquire higher precision. We found that the origin of population (adjusted P = 0.002) and detection method (adjusted P = 0.016) together contributed to the heterogeneity to one degree or another (Table 3).
Association of CR, DFS, and RFS with miR-181 expression
Three datasets including 467 patients investigated the prognostic role of miR-181 for CR (Figure 3A). A random-effects model was utilized to calculate the pooled HR and its 95% CI, due to the high heterogeneity among these datasets (I2 = 88.1%, P < 0.001). The meta-regression was not used for heterogeneity analysis of the CR, due to smaller size of the qualified datasets (< 10). Two datasets including 250 American patients evaluated the relationship between miR-181 and DFS (Figure 3B), and two datasets including 185 Chinese patients studied the relationship between miR-181 and RFS (Figure 3C). A fixed-effect model was applied to assess the association between miR-181 and DFS/RFS, respectively, since no heterogeneity was found among the datasets of each group. As origin of population in AML patients contributed to the heterogeneity in datasets evaluating the correlation between OS and miR-181, the datasets evaluating the correlation between CR, DFS, RFS and miR-181 were also investigated by subgroup analysis based on origin of population. Altogether, our results showed that high miR-181 level predicted favorable responses in terms of CR (pooled HR: 1.77, 95% CI: 1.25–2.52), DFS (pooled HR: 0.78, 95% CI: 0.67–0.91) in American AML patients (P < 0.01), and inferior outcomes with CR (HR: 0.54, 95% CI: 0.34–0.86), RFS (pooled HR: 2.07, 95% CI: 1.41–3.04) in Chinese AML patients (P < 0.01) (Figure 3). Collectively, our results support the prognostic prediction of miR-181 in AML patients based on the origin of population.
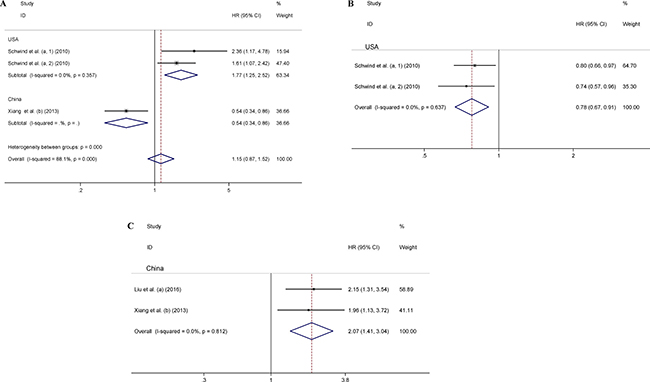
Figure 3: Forest plots of the relationship between miR-181 expression and CR (A), DFS (B), and RFS (C). The lower-case letters a, b represent that miR-181a and miR-181b were analyzed separately in the studies.
Sensitivity analysis
To investigate the effect of individual study on the overall meta-analysis estimate, sensitivity analysis was performed by calculating the pooled HRs with successive exclusion of one study. The result showed that pooled HRs did not change substantially with removal of any study (Figure 4), indicating a more reliable results of this meta-analysis.
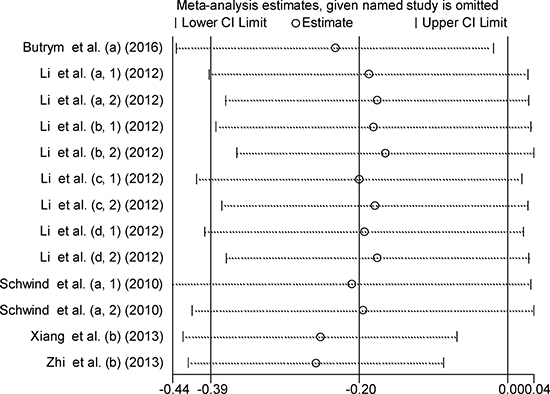
Figure 4: Sensitivity analysis of the relationship between miR-181 expression and OS. The lower-case letters a, b, c, d represent that miR-181a, miR-181b, miR-181c and miR-181d were analyzed separately in the studies.
Publication bias
Begg’s and Egger’s tests were used to assess the potential publication bias of the enrolled datasets that evaluated the correlation between OS and miR-181. The P values for Begg’s and Egger’s test were 0.127 and 0.680, respectively. Moreover, the funnel plot of the OS analysis based on miR-181 revealed no obvious asymmetry (Figure 5).
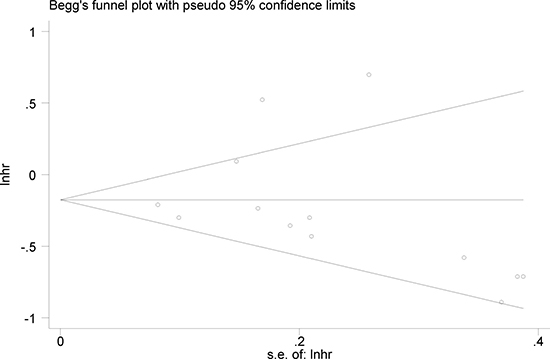
Figure 5: Begg’s funnel plot of miR-181 and OS for publication bias testing.
DISCUSSION
Even though accumulating evidence has indicated an association between miR-181 expression and hematological malignancies [37, 38], the prognostic role of miR-181 for survival in AML patients remains controversial. The inconsistent results among published studies could be due to the following factors: diverse origins of population, different platforms of miRNA profiling, and/or relatively small sample size in individual research. Lin et al. reported that miR-181a/b was significantly associated with OS in hematological malignancies (pooled HR: 0.72, 95% CI: 0.63–0.82) [39]. They analyzed hematological malignancies in 2 AML studies in 330 American patients and 2 chronic lymphoid leukemia (CLL) studies in 86 American and 150 Chinese patients. As the heterogeneity was found in AML and CLL, American and Chinese patients, the accuracy of their conclusion requires further confirmation. In our meta-analysis, six published articles were enrolled to evaluate the relationships between all members of miR-181 family and OS, CR, DFS, and RFS in AML patients. Overall, we found a positive correlation between miR-181 expression and survival in American AML patients (P < 0.01), with pooled HR of 0.74 (95% CI: 0.67–0.82) for OS, 1.77 (95% CI: 1.25–2.52) for CR, 0.78 (95% CI: 0.67–0.91) for DFS, and a negative correlation between miR-181 expression and survival in Chinese AML patients (P < 0.01), with pooled HR of 1.78 (95% CI: 1.35–2.34) for OS, 0.54 (95% CI: 0.34–0.86) for CR, 2.07 (95% CI: 1.41–3.04) for RFS. The result in Polish AML patients still requires further confirmation, as there is only one study evaluating the association between OS and miR-181 (HR: 1.10, 95% CI: 0.82–1.46). The incidence rate of adult is much higher than that of children and adolescent, indicating that different pathogenesis may exist between adult and pediatric AML patients. Since differences are existed between adult and pediatric AML patients, the prognostic values of miR-181 may be inconsistent. Liu et al. detected the expression of miR-181a in pediatric AML patients in China [32], and analyzed the correlation between the level of miR-181a and RFS (HR: 2.15, 95% CI: 1.31–3.54), which was similar with Xiang’s result (HR: 1.96, 95% CI: 1.13–3.72) in Chinese adult AML patients [30]. As there was only one dataset evaluating the prognostic value of miR-181 in pediatric AML patients, the results may require further validation with large datasets. Meanwhile, due to the limited samples size involved in CR, DFS, and RFS, our results may require further validation with large datasets. To our knowledge, this is the first meta-analysis that provides a systemic evaluation of the prognostic valueof miR-181 in AML patients.
There are several reported mechanisms supporting the protective role of miR-181 in AML patients. Nanbakhsh et al. indicated that overexpression of miR-181a in AML blasts resulted in the attenuation of their resistance to chemotherapy and NK-cell-mediated killing [40]. In addition, it was found that miR-181a could directly downregulate KRAS, NRAS and MAPK1, resulting in longer survival in a murine AML model treated with miR-181a mimics [41]. Lu et al. demonstrated that miR-181b increased the sensitivity of K562/A02 and HL-60/ADM cells to cytotoxic chemotherapeutic agents and promoted drug-induced apoptosis by inhibiting HMGB1 and MCL-1 expression [42]. Moreover, Bai et al. also proved that overexpression of miR-181a in HL-60/Ara-C cells could induce cytochrome C release and caspase 9/caspase 3 activation by directly targeting BCL-2 [43]. However, Bräuer-Hartmann et al. proved that overexpression of miR-181a/b inhibited granulocytic differentiation by targeting tumor suppressor gene RASSF1A and regulating the cell-cycle regulator cyclin D1 in NB4 cells [37]. Liu et al. found that overexpression of miR-181a increased the ratio of S-phase cells and significantly promoted cell proliferation by directly decreasing the tumor suppressor ataxia telangiectasia mutated (ATM) in NB4 and K562 cells [32]. Su et al. also demonstrated that miR-181 inhibited granulocytic and macrophage-like differentiation of HL-60 cells by directly inhibiting the expression of protein kinase C delta (PRKCD), CTD small phosphatase like (CTDSPL) and calcium/calmodulin dependent protein kinase kinase 1 (CAMKK1) [22]. These results not only highlight the importance of miR-181 as a prognostic biomarker in AML patients, but also indicate that miR-181 may serve as a potential therapeutic target for AML treatment.
Prognostic biomarkers are important for understanding the biological processes of diseases, designing accurate therapeutic strategies, and evaluating the prognosis for patients [44]. There are many prognostic biomarkers in leukemia, including leukocyte morphology and size test in blood, bone marrow or spleen [45], circulating cytokine levels [46], genetic variations [47], and epigenetics characteristics [48]. These distinct (epi)genetic and biological features are also associated with clinical responses to different therapeutic regiments. The clinical outcomes of AML patients are affected by white blood cell count [49], metabolic status [50], and genetic alterations (i.e. chromosomal rearrangements [51] and genetic mutations [52, 53]). For example, Ma et al. found that gene mutations of IDH1, IDH2 and high IDH1 expression were associated with a poor prognosis in CN-AML patients with shorter OS and event-free survival [54]. Chen et al. showed that a panel of 6 serum metabolite markers (including Lactate, 2-Oxoglutarate, Pyruvate) demonstrated prognostic value in CN-AML patients [50]. Recently, researchers have paid increasing attention to epigenetic biomarkers that contribute to the leukemogenesis [55], which include DNA methylation [56] and miRNA dysregulation [26]. The etiology of leukemia is complex, and combinational assessment of different biomarkers is expected to provide more reliable methods to assist medical decision-making and predict the clinical outcomes. In the present study, we confirmed the role of miR-181 in the prognosis of AML patients, but further studies are required to evaluate whether miR-181 influences the patients’ responses to therapeutic regimens in the clinic.
In conclusion, this meta-analysis indicates that higher miR-181 levels are positively associated with the prognostic outcome in American AML patients, and indicate a worse prognostic outcome in Chinese AML patients. Our findings contribute to the better understanding of epigenetic modifiers in tumorigenesis of AML, and pinpoint a novel biomarker and potentially a new therapeutic target in AML patients. We are aware of several limitations in this study. First, a high heterogeneity (I2 = 73.6%, P < 0.001) was found among the datasets evaluating the correlation between OS and miR-181. Sensitivity analysis showed no single study profoundly influenced the stability of the results. However, both subgroup analyses and meta-regression indicated the origin of patients contributed to the heterogeneity. Second, several HRs with 95% CI were extrapolated based on the data extracted from the Kaplan-Meier curves, which could slightly differ from the exact HRs. The samples in these studies were obtained from different tissues (bone marrow, plasma or unknown specimen), and the expression levels of miR-181 were detected by different methods (TaqMan PCR, SYBR Green PCR, or microarray), which might influence the accuracy of miR-181 expression. Moreover, the information of the race and ethnicity of AML patients was unable to be acquired from the enrolled studies, so further studies are required to elucidate the association between miR-181 and patient origin. Finally, the number of studies in sample types and subtype analyses was relatively small, and well-designed investigations with larger sample size are required for future validation of our findings. Furthermore, the role of miR-181 in the outcome of each French, American, and British (FAB) classification and/or cytogenetic subtype is of great interest for future studies.
MATERIALS AND METHODS
Search strategy
Published studies were systematically searched in PubMed, Embase and Cochrane Library databases up to May 15, 2017 without language restrictions by two independent researchers (Qiang Guo and Junwen Luan). The following keywords were applied simultaneously: (miR-181* OR miRNA-181* OR microRNA-181* OR miR181* OR miRNA181* OR microRNA181*) AND (“leukemia, myeloid, acute” OR “AML” OR “acute myeloid leukemia” OR “acute myeloid leukaemia”). Moreover, potentially related studies were also collected from the reference lists of the screened full-text articles above.
Inclusion and exclusion criteria
The eligible studies should follow these inclusive criteria: (1) the study subjects were patients with AML; (2) miR-181 expression levels were measured; (3) the association between a member of miR-181 family and clinical outcome, such as OS, CR, DFS, or RFS, was reported. Studies were excluded if they were: (1) abstracts, meta-analysis, or review articles; (2) without patient samples; (3) nondichotomous miR-181 expression levels; (4) lack of key information such as HR, 95% CI and P value. When duplicate publications were identified, only the newest or most informative single article was selected. We also contacted authors of some studies for additional information and data needed for the present meta-analysis. The entire searching was conducted independently by two authors (Qiang Guo and Junwen Luan), and supervised by a third author (Xia Li). Any disagreement was checked and resolved by discussion among the authors.
Quality assessment
A quality assessment was independently performed for all of the included studies by two authors (Xiaoxiao Zhu and Lin Zhao) using the NOS [33], and any disagreement was resolved by discussion and consensus. The NOS comprises the following three parameters of quality: the selection of the study groups (maximum score is 4), the comparability of the groups (maximum score is 2), and the outcome of interest (maximum score is 3). All of the three parameters were appraised according to the criteria of NOS, with compliance of one NOS standard = 1. The selection score indicates the degree of representativeness of the selected study, the comparability evaluates the comparability of selected studies based on its design or analysis, while the outcome assesses the comprehensiveness of the research contents. The total score is the sum of all three values, and represents the overall quality assessment of the selected paper. The lowest score is 0 and the highest is 9. Studies with a score greater than 5 were considered to be indicative of high-quality studies [34], and were enrolled in the final analysis in the present study.
Data extraction
The following information was extracted from all eligible studies: (1) publication details: first author’s last name and publication year; (2) characteristics of studied subject: ethnicity/origin, leukemia types, number of patients and specimen used for miR-181 detection; (3) miR-181 assessment methods and the cut-off definition; (4) survival analysis; and (5) HR of elevated miR-181 for OS (CR, DFS, or RFS), as well as their 95% CI and P value. If the HRs were not directly reported in the publication, the corresponding authors were contacted for additional data. If only Kaplan-Meier curves were available, Engauge Digitizer version 9.7 was used to extract data from the graphical survival plots, and HRs and 95% CIs were then calculated using the described method [35, 36]. In studies with both univariate analysis and multivariate analysis, HR obtained from multivariate analyses were preferably extracted as inclusion of other variables in the analysis tends to generate more accurate results. The data extraction was performed independently by two authors (Ni Li and Zhen Zhang), with consultation of a third author (Guosheng Jiang) for disagreements.
Statistical methods
The pooled HR with 95% CI was used to evaluate the correlation of miR-181 expression and the survival outcome of AML patients. The heterogeneity among all included studies was examined using Cochran’s Q test and Higgins I-square (I2) statistic. In case of no or moderate heterogeneity (P > 0.1 or I2 < 50%), the fixed-effect model (Mantel-Haenszel test) was applied; otherwise, the random-effects model (Der Simonian and Laird method) was used. Subgroup analysis and meta-regression were carried out to further explore possible explanations for heterogeneity [57]. In addition, Begg’s funnel plot and Egger’s bias were used to evaluate the potential publication bias [58, 59]. If a publication bias did exist, the Duval and Tweedie nonparametric Trim and Fill method was used to adjust the results [60]. Sensitivity analysis was performed by removing one study at each time to assess its influence on the pooled HR. All analyses were conducted by STATA package version 12.0 (Stata Corporation, College Station, Texas, USA). A P value < 0.05 was considered to be statistically significant.
Abbreviations
AML, acute myeloid leukemia; ATM, ataxia telangiectasia mutated; CA-AML, cytogenetically abnormal acute myeloid leukemia; CAMKK1, calcium/calmodulin dependent protein kinase kinase 1; CI, confidence interval; CLL, chronic lymphoid leukemia; CN-AML, cytogenetically normal acute myeloid leukemia; CR, complete remission; CTDSPL, CTD small phosphatase like; DFS, disease-free survival; FAB, French, American, and British; FLT3-ITD, FLT3-internal tandem duplication; HR, hazard ratio; miRNA, microRNA; miR-181, microRNA-181; NOS, Newcastle-Ottawa Quality Assessment Scale; OS, overall survival; PRKCD, protein kinase C delta; qPCR, quantitative PCR; RFS, relapse-free survival; SNP, single nucleotide polymorphism.
Authors’ contributions
Q.G., and X.L. designed this study; Q.G., J.W.L. and X.L. searched databases and collected full-text papers; N. L., Z.Z. and G.S.J. extracted the data; X.X.Z. and L.Z. performed the quality assessment; Q.G., J.W.L., Z.Z. and L.L.S. analyzed the data; R.W., Y.S., X.Q.Y. and N.D. contributed resources to facilitate the analyses; Q.G., and X.L. wrote the manuscript. All authors reviewed the manuscript.
ACKNOWLEDGMENTS
We greatly appreciated Dr. Yutao Diao in Shandong Academy Medical Sciences for data analysis.
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
FUNDING
This work was supported by the Natural Science Foundation of China (81373670, 81401314, 81573467), the Natural Science Foundation of Shandong Province (ZR2014YL028, ZR2015HM014), the Key Project of Science and Technology of Shandong Province (2015ZDJS04003), and the Project for Laureate of Taishan Scholar (No. ts201511075), the Primary Research & Development Plan of Shandong Province (2016GSF202016), the Project of Transformation in High-tech Achievements (2013ZHZX2A0405), the Project for Shandong Medical and Health Science and Technology Plan (2015WS0191), the Science and Technology Development Grant of the State Administration of TCM of Shandong Province (2013–2016, 2013–218), Family Planning Committee of Shandong Province ([2014]14), the National Science Foundation for Postdoctoral Scientist of China (2014M560566, 2015T80728) and the Innovation Project of Shandong Academy of Medical Sciences.
REFERENCES
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017; 67:7–30.
2. Roman E, Smith A, Appleton S, Crouch S, Kelly R, Kinsey S, Cargo C, Patmore R. Myeloid malignancies in the real-world: Occurrence, progression and survival in the UK’s population-based Haematological Malignancy Research Network 2004–15. Cancer Epidemiol. 2016; 42:186–98.
3. Chen W, Zheng R, Zhang S, Zeng H, Xia C, Zuo T, Yang Z, Zou X, He J. Cancer incidence and mortality in China, 2013. Cancer Letters. 2017; 401:63–71.
4. Chen BA, Huang ZH, Zhang XP, Ou-Yang J, Li JY, Zhai YP, Sun XM, Xu YL, Lu Q, Wang JM, Li D, Liao H, Shen ZX, et al. An epidemiological investigation of leukemia incidence between 2003 and 2007 in Nanjing, China. J Hematol Oncol. 2010; 3:21.
5. Miller KD, Siegel RL, Lin CC, Mariotto AB, Kramer JL, Rowland JH, Stein KD, Alteri R, Jemal A. Cancer treatment and survivorship statistics, 2016. CA Cancer J Clin. 2016; 66:271–89.
6. Bao H, Wu D. Current status of leukemia cytotherapy - exploitation with immune cells. Curr Stem Cell Res Ther. 2017; 12:188–96.
7. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016; 66:7–30.
8. Hackl H, Astanina K, Wieser R. Molecular and genetic alterations associated with therapy resistance and relapse of acute myeloid leukemia. J Hematol Oncol. 2017; 10:51.
9. Bose P, Vachhani P, Cortes JE. Treatment of relapsed/refractory acute myeloid leukemia. Curr Treat Options Oncol. 2017; 18:17.
10. Baccarani M, Deininger MW, Rosti G, Hochhaus A, Soverini S, Apperley JF, Cervantes F, Clark RE, Cortes JE, Guilhot F, Hjorth-Hansen H, Hughes TP, Kantarjian HM, et al. European LeukemiaNet recommendations for the management of chronic myeloid leukemia: 2013. Blood. 2013; 122:872–84.
11. Suzuki K, Okuno Y, Kawashima N, Muramatsu H, Okuno T, Wang X, Kataoka S, Sekiya Y, Hamada M, Murakami N, Kojima D, Narita K, Narita A, et al. MEF2D-BCL9 fusion gene is associated with high-risk acute B-cell precursor lymphoblastic leukemia in adolescents. J Clin Oncol. 2016; 34:3451–9.
12. Oldenhuis CN, Oosting SF, Gietema JA, de Vries EG. Prognostic versus predictive value of biomarkers in oncology. Eur J Cancer. 2008; 44:946–53.
13. Yates LA, Norbury CJ, Gilbert RJ. The long and short of microRNA. Cell. 2013; 153:516–9.
14. Mendell JT, Olson EN. MicroRNAs in stress signaling and human disease. Cell. 2012; 148:1172–87.
15. Di Leva G, Garofalo M, Croce CM. MicroRNAs in cancer. Annu Rev Pathol. 2014; 9:287–314.
16. Lu L, McCurdy S, Huang S, Zhu X, Peplowska K, Tiirikainen M, Boisvert WA, Garmire LX. Time Series miRNA-mRNA integrated analysis reveals critical miRNAs and targets in macrophage polarization. Sci Rep. 2016; 6:37446.
17. Balatti V, Pekarky Y, Croce CM. Role of microRNA in chronic lymphocytic leukemia onset and progression. J Hematol Oncol. 2015; 8:12.
18. Balatti V, Acunzo M, Pekarky Y, Croce CM. Novel mechanisms of regulation of miRNAs in CLL. Trends Cancer. 2016; 2:134–43.
19. Agatheeswaran S, Pattnayak NC, Chakraborty S. Identification and functional characterization of the miRNA-gene regulatory network in chronic myeloid leukemia lineage negative cells. Sci Rep. 2016; 6:32493.
20. Marcucci G, Radmacher MD, Maharry K, Mrozek K, Ruppert AS, Paschka P, Vukosavljevic T, Whitman SP, Baldus CD, Langer C, Liu CG, Carroll AJ, Powell BL, et al. MicroRNA expression in cytogenetically normal acute myeloid leukemia. N Engl J Med. 2008; 358:1919–28.
21. Wang Y, Li Z, He C, Wang D, Yuan X, Chen J, Jin J. MicroRNAs expression signatures are associated with lineage and survival in acute leukemias. Blood Cells Mol Dis. 2010; 44:191–7.
22. Su R, Lin HS, Zhang XH, Yin XL, Ning HM, Liu B, Zhai PF, Gong JN, Shen C, Song L, Chen J, Wang F, Zhao HL, et al. MiR-181 family: regulators of myeloid differentiation and acute myeloid leukemia as well as potential therapeutic targets. Oncogene. 2015; 34:3226–39.
23. Wang H, Tao T, Yan W, Feng Y, Wang Y, Cai J, You Y, Jiang T, Jiang C. Upregulation of miR-181s reverses mesenchymal transition by targeting KPNA4 in glioblastoma. Sci Rep. 2015; 5:13072.
24. Weng H, Lal K, Yang FF, Chen J. The pathological role and prognostic impact of miR-181 in acute myeloid leukemia. Cancer Genet. 2015; 208:225–9.
25. Gong J, Liu C, Liu W, Wu Y, Ma Z, Chen H, Guo AY. An update of miRNASNP database for better SNP selection by GWAS data, miRNA expression and online tools. Database (Oxford). 2015; 2015:bav029.
26. Liao Q, Wang B, Li X, Jiang G. miRNAs in acute myeloid leukemia. Oncotarget. 2017; 8:3666–82. https://doi.org/10.18632/oncotarget.12343.
27. Li Z, Huang H, Li Y, Jiang X, Chen P, Arnovitz S, Radmacher MD, Maharry K, Elkahloun A, Yang X, He C, He M, Zhang Z, et al. Up-regulation of a HOXA-PBX3 homeobox-gene signature following down-regulation of miR-181 is associated with adverse prognosis in patients with cytogenetically abnormal AML. Blood. 2012; 119:2314–24.
28. Schwind S, Maharry K, Radmacher MD, Mrozek K, Holland KB, Margeson D, Whitman SP, Hickey C, Becker H, Metzeler KH, Paschka P, Baldus CD, Liu S, et al. Prognostic significance of expression of a single microRNA, miR-181a, in cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. J Clin Oncol. 2010; 28:5257–64.
29. Zhi F, Cao X, Xie X, Wang B, Dong W, Gu W, Ling Y, Wang R, Yang Y, Liu Y. Identification of circulating microRNAs as potential biomarkers for detecting acute myeloid leukemia. PLoS One. 2013; 8:e56718.
30. Xiang L, Li M, Liu Y, Cen J, Chen Z, Zhen X, Xie X, Cao X, Gu W. The clinical characteristics and prognostic significance of MN1 gene and MN1-associated microRNA expression in adult patients with de novo acute myeloid leukemia. Ann Hematol. 2013; 92:1063–9.
31. Butrym A, Rybka J, Baczynska D, Poreba R, Mazur G, Kuliczkowski K. Expression of microRNA-181 determines response to treatment with azacitidine and predicts survival in elderly patients with acute myeloid leukaemia. Oncol Lett. 2016; 12:2296–300.
32. Liu X, Liao W, Peng H, Luo X, Luo Z, Jiang H, Xu L. miR-181a promotes G1/S transition and cell proliferation in pediatric acute myeloid leukemia by targeting ATM. J Cancer Res Clin Oncol. 2016; 142:77–87.
33. Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol. 2010; 25:603–5.
34. Zhang M, Dong Y, Liu H, Wang Y, Zhao S, Xuan Q, Wang Y, Zhang Q. The clinicopathological and prognostic significance of PD-L1 expression in gastric cancer: a meta-analysis of 10 studies with 1,901 patients. Sci Rep. 2016; 6:37933.
35. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials. 2007; 8:16.
36. Guyot P, Ades AE, Ouwens MJ, Welton NJ. Enhanced secondary analysis of survival data: reconstructing the data from published Kaplan-Meier survival curves. BMC Med Res Methodol. 2012; 12:9.
37. Brauer-Hartmann D, Hartmann JU, Wurm AA, Gerloff D, Katzerke C, Verga Falzacappa MV, Pelicci PG, Muller-Tidow C, Tenen DG, Niederwieser D, Behre G. PML/RARα-regulated miR-181a/b cluster targets the tumor suppressor RASSF1A in acute promyelocytic leukemia. Cancer Res. 2015; 75:3411–24.
38. Saleh LM, Wang W, Herman SE, Saba NS, Anastas V, Barber E, Corrigan-Cummins M, Farooqui M, Sun C, Sarasua SM, Zhao Z, Abousamra NK, Elbaz O, et al. Ibrutinib downregulates a subset of miRNA leading to upregulation of tumor suppressors and inhibition of cell proliferation in chronic lymphocytic leukemia. Leukemia. 2017; 31:340–9.
39. Lin S, Pan L, Guo S, Wu J, Jin L, Wang JC, Wang S. Prognostic role of microRNA-181a/b in hematological malignancies: a meta-analysis. PLoS One. 2013; 8:e59532.
40. Nanbakhsh A, Visentin G, Olive D, Janji B, Mussard E, Dessen P, Meurice G, Zhang Y, Louache F, Bourhis JH, Chouaib S. miR-181a modulates acute myeloid leukemia susceptibility to natural killer cells. Oncoimmunology. 2015; 4:e996475.
41. Huang X, Schwind S, Santhanam R, Eisfeld AK, Chiang CL, Lankenau M, Yu B, Hoellerbauer P, Jin Y, Tarighat SS, Khalife J, Walker A, Perrotti D, et al. Targeting the RAS/MAPK pathway with miR-181a in acute myeloid leukemia. Oncotarget. 2016; 7:59273–86. https://doi.org/10.18632/oncotarget.11150.
42. Lu F, Zhang J, Ji M, Li P, Du Y, Wang H, Zang S, Ma D, Sun X, Ji C. miR-181b increases drug sensitivity in acute myeloid leukemia via targeting HMGB1 and Mcl-1. Int J Oncol. 2014; 45:383–92.
43. Bai H, Cao Z, Deng C, Zhou L, Wang C. miR-181a sensitizes resistant leukaemia HL-60/Ara-C cells to Ara-C by inducing apoptosis. J Cancer Res Clin Oncol. 2012; 138:595–602.
44. Pfirrmann M, Lauseker M, Hoffmann VS, Hasford J. Prognostic scores for patients with chronic myeloid leukemia under particular consideration of competing causes of death. Ann Hematol. 2015; 94:S209–18.
45. Hasford J, Baccarani M, Hoffmann V, Guilhot J, Saussele S, Rosti G, Guilhot F, Porkka K, Ossenkoppele G, Lindoerfer D, Simonsson B, Pfirrmann M, Hehlmann R. Predicting complete cytogenetic response and subsequent progression-free survival in 2060 patients with CML on imatinib treatment: the EUTOS score. Blood. 2011; 118:686–92.
46. Nievergall E, Reynolds J, Kok CH, Watkins DB, Biondo M, Busfield SJ, Vairo G, Fuller K, Erber WN, Sadras T, Grose R, Yeung DT, Lopez AF, et al. TGF-α and IL-6 plasma levels selectively identify CML patients who fail to achieve an early molecular response or progress in the first year of therapy. Leukemia. 2016; 30:1263–72.
47. Patel JP, Gonen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J, Van Vlierberghe P, Dolgalev I, Thomas S, Aminova O, Huberman K, Cheng J, Viale A, et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med. 2012; 366:1079–89.
48. Qu X, Davison J, Du L, Storer B, Stirewalt DL, Heimfeld S, Estey E, Appelbaum FR, Fang M. Identification of differentially methylated markers among cytogenetic risk groups of acute myeloid leukemia. Epigenetics. 2015; 10:526–35.
49. Radhi M, Meshinchi S, Gamis A. Prognostic factors in pediatric acute myeloid leukemia. Curr Hematol Malig Rep. 2010; 5:200–6.
50. Chen WL, Wang JH, Zhao AH, Xu X, Wang YH, Chen TL, Li JM, Mi JQ, Zhu YM, Liu YF, Wang YY, Jin J, Huang H, et al. A distinct glucose metabolism signature of acute myeloid leukemia with prognostic value. Blood. 2014; 124:1645–54.
51. Grimwade D, Ivey A, Huntly BJ. Molecular landscape of acute myeloid leukemia in younger adults and its clinical relevance. Blood. 2016; 127:29–41.
52. Rampal R, Figueroa ME. Wilms tumor 1 mutations in the pathogenesis of acute myeloid leukemia. Haematologica. 2016; 101:672–9.
53. Medinger M, Lengerke C, Passweg J. Novel prognostic and therapeutic mutations in acute myeloid leukemia. Cancer Genomics Proteomics. 2016; 13:317–29.
54. Ma QL, Wang JH, Wang YG, Hu C, Mu QT, Yu MX, Wang L, Wang DM, Yang M, Yin XF, Chen FF, Lu SS, Chen J, et al. High IDH1 expression is associated with a poor prognosis in cytogenetically normal acute myeloid leukemia. Int J Cancer. 2015; 137:1058–65.
55. Chen J, Odenike O, Rowley JD. Leukaemogenesis: more than mutant genes. Nat Rev Cancer. 2010; 10:23–36.
56. Qu X, Othus M, Davison J, Wu Y, Yan L, Meshinchi S, Ostronoff F, Estey EH, Radich JP, Erba HP, Appelbaum FR, Fang M. Prognostic methylation markers for overall survival in cytogenetically normal patients with acute myeloid leukemia treated on SWOG trials. Cancer. 2017.
57. Thompson SG, Sharp SJ. Explaining heterogeneity in meta-analysis: a comparison of methods. Stat Med. 1999; 18:2693–708.
58. Begg CB, Mazumdar M. Operating characteristics of a rank correlation test for publication bias. Biometrics. 1994; 50:1088–101.
59. Egger M, Smith GD, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629–34.
60. Duval S, Tweedie R. Trim and fill: a simple funnel-plot-based method of testing and adjusting for publication bias in meta-analysis. Biometrics. 2000; 56:455–63.

