Introduction
Bladder cancer is the most common malignant tumor of urinary system with an increasing incidence [1]. Papillary and non-papillary carcinomas are the two different types of bladder cancer with unique, yet overlapping clinical and pathological features [2]. The urothelial carcinomas of the bladder are generally called bladder cancer or bladder transitional cell carcinoma and are most common. Nearly 80% of urothelial carcinomas are non-invasive urothelial papillomas with high recurrence rates after resection without infiltrating the bladder wall or distant metastasis. The rest 20% are muscle-invasive bladder cancers that are highly invasive with distant metastasis [3]. Although patients with superficial papillary lesions usually experience multiple recurrences, only 10%-30% of them develop into high-grade invasive tumors. On the contrary, most patients with high-grade invasive bladder cancers do not show superficial papillary lesions. The 5-year survival rate of muscle-invasive bladder cancer patients is low and has shown no improvement inspite of great advances in prognosis and surgical methods. The bladder cancer stem cells (BCSCs) are probably responsible for the high recurrence rates of bladder cancer, tumor heterogeneity and other complex biological behaviors.
The cancer stem cells (CSCs) are defined as a subset of cells with the ability to self-renew and differentiate into hierarchical tumor cells, thereby contributing to tumor heterogeneity [4]. Cancer stem cells (CSCs) were first identified in hematopoietic malignancies as a subgroup of cells that demonstrated stemness and differentiated into heterogenous populations of tumor cells [5, 6]. CSCs have also been identified in solid cancers like colorectal carcinomas [7]and ovarian [8] and liver cancers [9]. They also demonstrate self-renewal and differentiation properties [10] and are responsible for tumor heterogeneity and high recurrence rates of many cancers [4].
Bladder cancer is highly recurrent, metastatic and heterogenous, thereby resulting in poor prognosis [11]. It is postulated that bladder cancer stem cells are partly responsible for the clinical characteristics and complex biological behavior of bladder cancer [12]. Therefore, understanding the role of CSCs in bladder cancer is critical to gain insights into the mechanisms responsible for high recurrence and metastasis. Additionally, it would help identify novel therapeutic avenues and facilitate better prognosis. In this review, we focus on the new frontiers and progress in the study of bladder cancer stem cells.
Origin of bladder cancer stem cells
Bladder cancer types
The 2 types of bladder cancers, namely non-muscle and muscular bladder cancers originate from two different pathways [13]. Studies with transgenic mice revealed that normal stem cells with HRAS or FGFR3 mutations can transform into BCSCs that develop into superficial non-muscle invasive bladder cancer, whereas stem cells with p53/Rb/PTEN gene mutations transform into BCSCs that initiate muscular invasive bladder cancer [14]. Further, transgenic mice with continuous activation of HRAS gene developed non-invasive bladder cancer, whereas transgenic mice with UPII promoter-SV40T gene in combination with persistent inactivation of p53 and pRb gene developed invasive bladder cancer [15]. These studies postulated that BCSCs probably originated from transformed transgenic stem cells.
The origin of bladder cancer stem cells
The topic about the origin of CSCs is still controversial [16, 17]. It is widely assumed that CSCs may arise from normal stem cells having suffered gene mutations [18] and the generation of CSCs from normal stem cells is complex [17]. By single cell sequencing, Yang et al showed that bladder cancer stem cells (BCSCs) originated from bladder cancer stem cells (BCSCs) or bladder cancer non-stem cells (BCNSCs) with clonal homogeneity among BCSCs and BESCs or BCSCs and BCNSCs [19]. Apart from stem cells, CSCs can also originate from progenitor cells or differentiated cells that undergo de-differentiation or tumor cells that gain stem cell properties or fusion cells [16].
Several common markers of BCSCs including CD44+, EMA-, 67LR+, BCMab1+ [20, 21]are located in the basal cell layer of bladder tumor leading to more debates regarding the source of BCSCs. Theoretically, if all the markers are from a specific cell type in bladder cancer, it is assumed that BCSCs may have originated from mutated normal stem cells. On the other hand, if the markers are expressed on different normal cell types, then the BCSCs may be derived from progenitors or differentiated cells that acquired de-differentiation properties due to mutations, thereby leading to different subgroups of BCSCs (Figure 1).
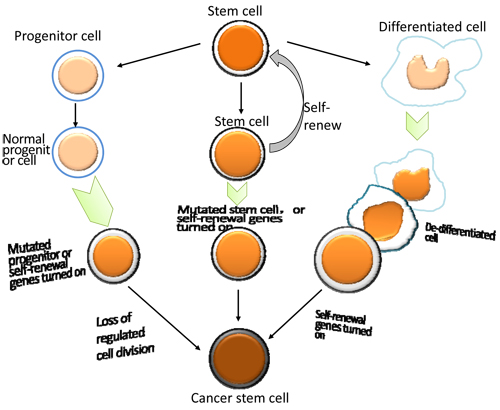
Figure 1: The origin of bladder cancer stem cells.
Normal urothelial stem cells
Colon cancer stem cells generally originate from intestinal epithelial stem cells expressing Lgr5 [22]. BrdU pulse-chase labeling assays suggested that the urothelial stem cells were located in the basal cell layer [23]. This was further confirmed by mitochondria DNA mutation experiments [24]. Nitrosamine induced bladder cancer model confirmed that invasive bladder cancer originated from stem cells from basal cell layer [25]. These studies suggested that BCSCs originated from urothelial stem cells in the basal cell layer.
Urothelial stem cell
Urothelial, bone marrow and adipose stem cells are all capable of repairing bladder damage [26]. Therefore, these stem cells are possible sources of BCSCs. The gram-negative bacterium, Helicobacter pylori is a carcinogen that recruits bone marrow derived cells (BMDCs) in gastric cancer [27]. However, in chemical induced bladder cancer, BMDCs are associated with inflammation in response to tumor and not related to tumorigenesis [28].
Basal cells
BCSCs were found to be CD44+CK5+CK20- that were characteristic basal cell markers [5]. Yang et al. showed that CD44+ cells were in the basal cell layer of normal urothelium and urothelial carcinoma [20]. Shin et al demonstrated that muscular invasive bladder cancer were derived specifically from Sonic hedgehog (SHh) expressing basal cells [25].
Intermediate stratum cells
The cells within the intermediate layer express different levels of CD44, which has been identified also as the BCSC marker [20]. Lineage tracing experiments in a mouse tumor model demonstrated that papillary cancer cells mainly originated from the intermediate layer [20]. Further, Brant et al thought that mutations of the fibroblast growth receptor FGFR3 in intermediate layer cells might help intermediate stratum bladder cells transform into malignant low-grade papillary carcinoma and urinary epithelial hyperplasia [12]. These experiments showed that non-muscle invasive bladder stem cells may originate from the intermediate layer cells.
Umbrella cells
The muscular invasive bladder cancer of intracavity type showed aberrant expression of transcription factors PPARG, ESR1, and FGFR3 [22]. Also, they expressed umbrella cell markers such as keratin 20 [29]. This suggested that BCSCs that develop into muscle invasive bladder cancer are derived from umbrella cells.
Bladder cancer cells
Cancer stemness is affected by genotype heterogeneity, epigenetic alterations and tumor microenvironment [30]. The interaction of tumor cells with tumor associated fibroblasts, macrophages, perivascular stroma and endothelial cells is critical for their survival in hypoxic and low nutritional conditions. The CSC-like phenotype of bladder cancer is observed during late stages of tumor development suggesting that the early bladder cancer cells may transform into CSCs through mechanisms such as epithelial mesenchymal transformation (EMT), dedifferentiation, and hypoxia [31].
Identification of bladder cancer stem cells
BCSC surface markers
Bladder cancer stem cells were identified for the first time in 2009 via the markers used to isolate normal stem cells [32]. So cell surface markers are traditionally used to isolate BCSCs. Since the biological behavior and phenotypes of tumor cell lines may have changed due to long-term in vitro culturing, primary or early passage tumor cell lines are ideal for isolating and identifying of BCSCs.
Chan et al. demonstrated that 40% of more than 300 bladder transitional cell carcinoma patient samples contained CD44+ cells. They further showed CD44 expressing subpopulation of cells in serial xenograft experiments with fresh patient samples and the tumorigenicity of CD44+ bladder cancer cells was found to be 10-200 times higher than CD44- bladder cancer cells when transplanted in immunodeficient mice [5]. The CD44+ cells maintained heterogeneity of the primary tumor after sequential transplantation, thereby complying with the functional standard for stem cells [5]. In vitro clonal sphere formation studies have been widely used to evaluate the stemness of tumor cells. In one such experiment, the CD44 splice isomer (CD44v6) was used to separate the CD44v6+ epithelial membrane antigen negative (EMA-) stem cell subtype from bladder tumors [33].
Further, the 67LR+CEACAM- BCSCs were identified using two surface markers, namely, the 67kDa basal layer laminin receptor (67LR) and carcinoembryonic antigen related cell adhesion factor 6 (CEACAM6); 67LR is expressed in the junction of tumor stroma and found in 80% of high-grade invasive bladder cancer; CEACAM6 is a non-specific poor reaction antigen [33]. In both experiments, the stem cell subgroups with special tumor characteristics were isolated using cell surface markers that were normally expressed in the urinary epithelial basal cells.
ALDH1A1 is another widely used bladder cancer stem cells marker. By in vitro sphere formation assays and in vivo xenograft experiments, it was shown that ALDH1A1+ cells had better colony formation and tumorigenicity characteristics than ALDH1A1- cells [34]. Moreover, the colony formation and tumorigenicity properties of BCSCs were significantly reduced by shRNA knockdown of the ALDH1A1 gene. Also, the ALDH1A1+ cells were subtypes of CD44+ cells and may represent more primitive BCSCs [34].
Since the CSC markers are complex, BCSCs have also been isolated using other markers. For example, Li et al. combined 2 monoclonal antibodies of human bladder cancer against BCMab1 and CD44 to isolate the BCMab1+CD44+ subgroup with strong proliferative and self-renewal properties that were associated with stem cells [35].
Side population cell
The biological features of stem cells can also be used to isolate CSCs. Stem cells highly express the ATP-binding cassette transporter in order to pump out drugs. This characteristic has been utilized to sort CSCs based on their ability to efficiently efflux the DNA fluorescent dye Hoechst 33342 and isolate the side population cells by FACS. Compared to other cells, the side population cells demonstrate colony formation, self-renewal and multi-directional differentiation characteristics that are typical of CSCs [36].
Molecular mechanisms of bladder cancer stem cells
Both normal stem cells and CSCs are characterized by their ability to self-renew. Signaling pathways related to the stemness maintenance and plasticity regulation of normal stem cells such as Hedgehog, Notch and Wnt are also involved in the stemness maintenance of CSCs including BCSCs. In addition, the tyrosine kinase receptor signaling pathway, interleukin-6 and tyrosine kinase 1 signaling pathways also play an important role in regulating the stemness of solid CSCs.
Hedgehog signaling pathway
Hedgehog signaling pathway (Figure 2) plays an important role in regulating embryogenesis and includes three homologous hedgehog genes namely Sonic Hedgehog (SHh), Indian Hedgehog (IHh) and Desert Hedgehog (DHh). Among these, SHh is highly expressed [37]. The hedgehog ligand binds to the transmembrane protein receptor Patched 1 (PTCH) and activates both the GLI transcription factor and the transmembrane protein receptor, Patched 1, which are inhibited by Smoothened (Smo) [38]. Aberrant activation of this pathway is central to development of a variety of tumors and their chemotherapy resistance [39]. Hence, inhibition of any of these factors may inhibit CSCs and therefore is a potential therapeutic target.
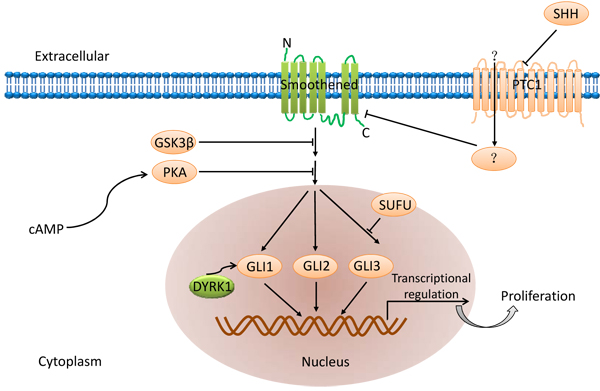
Figure 2: Hedgehog signaling pathway.
Notch signaling pathway
Notch signaling pathway (Figure 3) plays an important role in intercellular communication and cell fate determination during embryogenesis and in adult cells [40]. The pathway includes four receptors (Notch1~Notch4) and five ligands that regulate expression of multiple target genes [41]. When the Notch ligand binds to its receptor, the extracellular and transmembrane domains of the receptor are degraded by metalloproteinases and the secretory enzyme, respectively, thereby releasing the notch intracellular domain [42]. The soluble intracellular domain is imported to the nucleus and activates its target genes. The Notch signaling pathway is hyperactivated in a variety of tumors and is involved in maintaining the stemness of CSCs [43]. In melanomas, Notch4 interacts with CSCs which express CD133 and ABCG2 to promote melanomagenesis [44]. In medulloblastoma, sustained high expression of Notch2 significantly increases the number of medulloblastoma stem cells [45-48], whereas the inhibition of Notch signaling decreases the medulloblastoma stem cells and suppresses carcinogenesis [49].
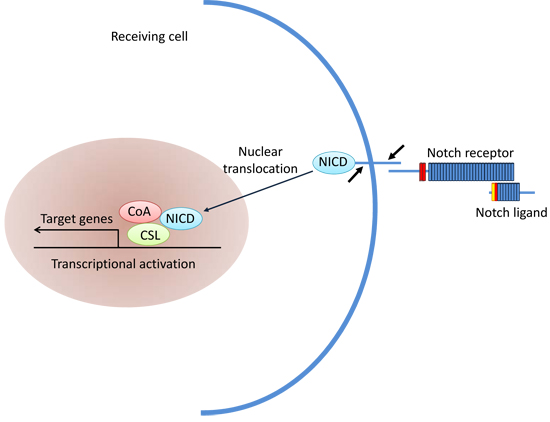
Figure 3: Notch signaling pathway.
Wnt signaling pathway
The Wnt signaling pathway (Figure 4) is another important signaling pathway involved in embryogenesis and cell proliferation, survival and development [50, 51]. It is involved in stemness and maintenance of CSCs [52]. Wnt signaling maintains the stem cell pool and prevents their differentiation by stabilizing cytoplasmic β-catenin levels [53]. The main components of the Wnt signaling pathway include the extracellular factor Wnt, the transmembrane receptor, β-catenin, the “destruction complex”, and the transcription factor referred to as T cell factor (TCF). Wnt signaling is generally activated when both frizzled (FZD) and lipoprotein receptor-related protein (LRP) bind to Wnt.
After activation, the Wnt protein bound to the cell surface receptor FZD and LRP5/6 triggers intracellular signal transduction resulting in activation of the Dsh protein [54]. Activated Dsh then binds to Axin and Frat1 to form a complex with GSK-3β and APC. Then, Frat1 mediates the release of GSK-3β from Axin resulting in accumulation of dephosphorylated β-catenin. Dephosphorylated β-catenin is transported to the nucleus to activate the target genes including c-myc, stromelysin, fibroblast growth factor, epithelial cell growth factor and cyclin D1. Expression of these genes promotes tumorigenesis. Thus, tumorigenic β-catenin mutation results in aberrant Wnt signaling in the CSCs, thereby inducing tumorigenic proliferation [55].
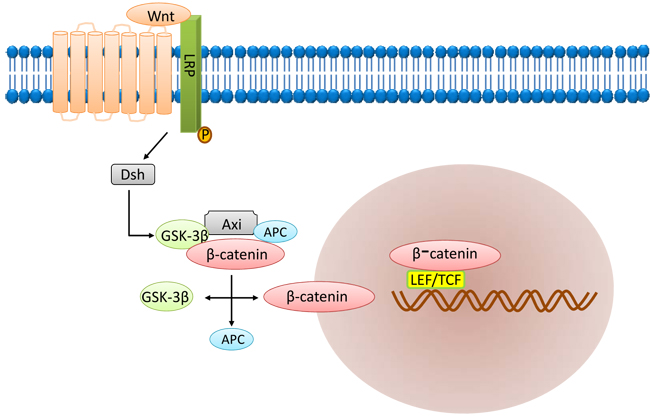
Figure 4: Wnt signaling pathway.
Other signaling pathways
Aberrant activation of the JAK-STAT signaling pathway has been shown to induce tumorigenesis [56, 57]. STAT3 activation is required for the tumorigenic functions of multiple CSCs [57-60]. IL-6 cytokine that is critical for the stemness of normal stem cells that also regulates CSCs by modulating OCT2, CD44 and SOX2 gene expression. The IL6/JAK/STAT3 signaling pathway helps maintain plasticity of breast CSCs and also activates mTORC1-STAT3 signaling pathway to maintain BCSC stemness [37, 41]. The tumorigenic tyrosine kinase receptor also contributes to stemness maintenance and chemotherapy resistance of non-small cell lung cancer stem cells [61].
Drug resistant mechanism of BCSCs
The high recurrence rates of papillary bladder cancer as well as high invasiness and metastasis of myometrial invasive bladder cancer are attributed to BCSCs [12, 15, 62-64]. Also, recurrent bladder cancers are usually chemoresistant, although the primary cancer is sensitive to chemotherapy [65]. There are two possible reasons for the chemoresistance. First, the chemotherapeutic drugs are unable to access the core of the tumor [64]; second, the BCSCs are chemoresistant [65, 66]. In fact, chemotherapy in combination with stem cell targeted therapy is the most promising therapy currently available for bladder cancer [67]. Therefore, it is important to identify the mechanisms that contribute to chemoresistance of BCSCs and deciphere ways to suppress or eliminate it to improve the prognosis of bladder cancer patients. The mechanisms that contribute to chemoresistance mechanisms of bladder cancer stem cells are: (1) active pumping out of chemotherapy drugs by the BCSCs; (2) enzymatic breakdown of drugs and their metabolites rendering them ineffective; (3) inhibition of programmed cell death (apoptosis) and (4) regulation of cytokines and other immune substances that inhibit the anti-tumor immune response (Figure 5).
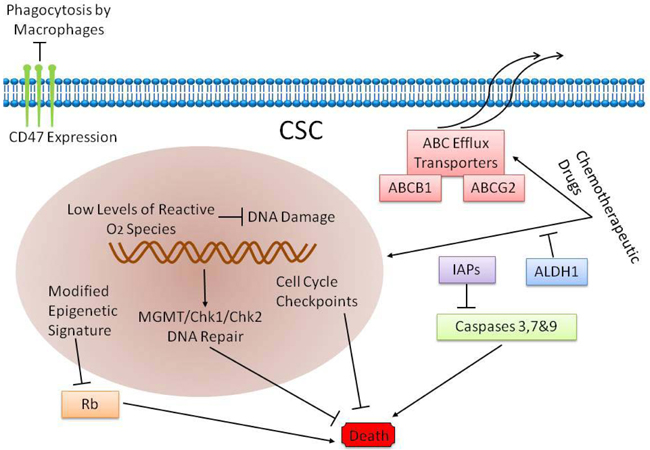
Figure 5: Drug resistant mechanism of BCSCs.
ABC transport proteins
The ABC transport proteins in stem cells, especially the Vallazapine-sensitive ABCG2, pump out intracellular metabolites, drugs and toxic substances [68]. This was previously used as the principle for isolating the side population (SP) CSCs [69]. The SP cells sorted from T24 human bladder cancer cell lines showed significant colony-forming ability upon subculturing and were tolerant to radiotherapy and chemotherapy [36]. This suggested that the ABC transport proteins contributed to the chemoresistance of BCSCs.
Acetaldehyde dehydrogenase (ALDH)
Acetaldehyde dehydrogenase (ALDH) is significantly overexpressed in BCSCs [70]. In xenograft studies of mouse colon cancer, the cyclophosphamide and its metabolites were oxidized and inactivated by ALDH, thereby helping tumor cells to survive chemotherapeutic treatment [71, 72]. Since ALDH is overexpressed in BCSCs, inactivation of chemotherapeutic agents and their metabolites by ALDH is a likely resistance mechanism for bladder cancer.
Antioxidant enzymes
During radiotherapy and chemotherapy, reactive oxygen species (ROS) that are generated in tumor cells, irreversibly damage the DNA, thereby resulting in tumor cell death [73, 74]. In epithelial CSCs, ROS levels were significantly lower as a result of robust free radical scavenging system that protected the tumor cell DNA from endogenous or exogenous ROS and oxidative damage [75, 76]. Also, urinary epithelial CSCs upregulate antioxidant enzymes like superoxide dismutase (SOD2) and Hemeoxygenase, thereby decreasing oxidative DNA damage [77]. The presence of these antioxidant enzymes leads to a significant increase in the viability of CSCs during chemotherapeutic treatment.
Apoptosis resistance
Urinary epithelial CSCs express high levels of IL11, IL18 and IL23 mRNA [70]. These interleukins promote in vitro survival and growth of tumor cells by activation of anti-apoptotic genes including cFLIP / FLAME-1 and Bcl-xL [78]. High interleukin levels have also been reported in prostate, breast, bladder and colorectal cancers [78]. IL23 is involved in the activation of the STAT-3 pathway, which promotes tumor survival, proliferation, invasion and angiogenesis [79].
Therefore, cytokines produced by the BCSCs enable them to survive chemotherapy and mediate recurrence. Therefore, to improve therapeutic outcomes, it is important to uncover the molecular mechanisms relevant for chemoresistance of BCSCs.
Targeted therapy for bladder cancer CSCs
Drug resistance is a barrier to the effectual remedy of most cancers. Although CSCs-targeting treatment is still far away from clinical use, it is still widely believed to be a promising approach to drug resistance and may create better curative effects on cancers [80]. However, we need to identify CSCs-associated genes with a comprehensive study on the genomic profiles of CSCs in order to establish intervention targets. By single cell sequencing, Yang et al identified 21 key mutation genes in BCSCs including 6 genes,ETS1, GPRC5A, MKL1, AWR, PITX2 and RGS9BP that were reported for the first time in bladder cancer. Besides, co-mutagenesis of ARID1A, GPRC5A and MLL2 by CRISPR/Cas9 technology significantly enhanced self-renewal and tumor initiation properties of BCSCs [19]. This study explored the genetic basis of human BCSCs and demonstrated their phylogeny. Besides, cancer stem cell function also interacts with epigenetics. Histone modification and chromatin rearrangement mutations usually exist in CSCs [81], promoting tumour initiation in different mechanisms [82]such as influencing the genome integrity [83]or leading to epigenetic reprogramming in iPS [84].
Analysis of the heterogeneity of different CD44+ subsets in bladder transitional cell carcinoma demonstrated many proteins that are involved in self-renewal such as nuclear Bmi-1, Stat3, and β-Catenin. Also, Activated Gli1 was found in 85% of CD44+ tumor cells suggesting that Gli1 was a potential molecular target [85].
The above mutations and abnormally expressed genes in CSCs are all potential therapeutic targets.
The monoclonal antibody therapy targeting tumor cell antigens is most effective among cancer treatments. Immunofluorescence and FACS analyses showed that CD47 is widely expressed in bladder urothelial carcinoma cells [32]. Compared to CD44- tumor cell subsets, higher CD47 expression was observed in CD44+ tumor stem cells. CD47 is immunosuppressive and inhibits macrophage phagocytosis by interacting with SIRPα, a plasma membrane protein that is mainly expressed on bone marrow cells including macrophages, neutrophils, eosinophils and dendritic cells [86]. Treatment with monoclonal anti-CD47 antibody can induce phagocytosis of macrophages in bladder cancer cells in vitro and significantly reduce tumor growth in bladder cancer xenografts in a dose-dependent manner in the recipient mice [32, 86].
The CSC-targeted chemo-radiotherapy, which is different from conventional chemo-radiotherapy, combined with small molecule inhibitors that target CSC-specific signaling pathways are expected to be effective therapeutics in the future.
Conclusions and future prospects
The CSCs, which were first identified in human malignant blood tumors [87], have been subsequently identified in many solid tumors [10, 38, 88-90]. In vivo and in vitro experiments with tumor cells in genetically engineered mice has greatly advanced our understanding regarding cancer stem cell role, including insights regarding their relevance for clinical applications. However, clinical success from the point of view of CSCs needs further understanding and clinical development. Compared to CSCs in hematological malignant tumors, the solid tumor CSCs interact with a large number of endothelial cells, fibroblasts, and inflammatory cells. To some extent, CSCs isolated from solid tumors with one or several specific markers is not equal to the whole tumor mass [91]. Therefore, study and clinical applications regarding solid tumor CSCs need to consider cell-cell and cell-stroma interactions, which are highly complex and sometimes difficult to simulate. Also, the field of solid tumor CSCs is still developing and there is no consensus yet regarding the existence, origin and development mechanism of CSCs. These areas need robust scientific exploration.
In the 21st century, a variety of malignant tumors including bladder cancer have extremely poor prognosis. The CSCs that have been discovered in the past decade or two provide a novel perspective to anticancer therapy that needs to be pursued strongly. In regard to BCSCs, it is important to determine specific bladder cancer stem cells markers, isolate and identify the origin and role of different categories of BCSCs with self-renewal and differentiation ability. Greater understanding of the biology of BCSCs will provide important information regarding disease classification, prognosis, treatment and early intervention, which will all improve clinical outcomes. We believe that with the deep understanding of BCSCs, the diagnosis and treatment of bladder cancer will make great progress in the near future.
ACKNOWLEDGMENTS
This work was supported by the National Natural Science Foundation of China (81472413, 81672956, 81602644).
conflicts of Interests
No potential conflicts of interest were disclosed.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016; 66: 7-30. http://doi.org/10.3322/caac.21332.
2. Witjes JA, Comperat E, Cowan NC, De Santis M, Gakis G, Lebret T, Ribal MJ, Van der Heijden AG, Sherif A. EAU guidelines on muscle-invasive and metastatic bladder cancer: summary of the 2013 guidelines. Eur Urol. 2014; 65: 778-92. http://doi.org/10.1016/j.eururo.2013.11.046.
3. Khandelwal P, Abraham SN, Apodaca G. Cell biology and physiology of the uroepithelium. Am J Physiol Renal Physiol. 2009; 297: F1477-501. http://doi.org/10.1152/ajprenal.00327.2009.
4. Easwaran H, Tsai HC, Baylin SB. Cancer epigenetics: tumor heterogeneity, plasticity of stem-like states, and drug resistance. Mol Cell. 2014; 54: 716-27. http://doi.org/10.1016/j.molcel.2014.05.015.
5. Bonnet D, Dick JE. Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med. 1997; 3: 730-7. http://doi.org/org/10.1038/nm0797-730
6. Reya T, Morrison SJ, Clarke MF, Weissman IL. Stem cells, cancer, and cancer stem cells. Nature. 2001; 414: 105-11. http://doi.org/10.1038/35102167.
7. Ricci-Vitiani L, Lombardi DG, Pilozzi E, Biffoni M, Todaro M, Peschle C, De Maria R. Identification and expansion of human colon-cancer-initiating cells. Nature. 2007; 445: 111-5. http://doi.org/10.1038/nature05384.
8. Li J, Condello S, Thomes-Pepin J, Ma X, Xia Y, Hurley TD, Matei D, Cheng JX. Lipid Desaturation Is a Metabolic Marker and Therapeutic Target of Ovarian Cancer Stem Cells. Cell Stem Cell. 2016. http://doi.org/10.1016/j.stem.2016.11.004.
9. Ma S, Chan KW, Hu L, Lee TK, Wo JY, Ng IO, Zheng BJ, Guan XY. Identification and characterization of tumorigenic liver cancer stem/progenitor cells. Gastroenterology. 2007; 132: 2542-56. http://doi.org/10.1053/j.gastro.2007.04.025.
10. Visvader JE, Lindeman GJ. Cancer stem cells: current status and evolving complexities. Cell Stem Cell. 2012; 10: 717-28. http://doi.org/10.1016/j.stem.2012.05.007.
11. Chen C, Hu L, Chen Y, Hou J. The prognostic value of histological subtype in patients with metastatic bladder cancer. Oncotarget. 2017; 8: 28408-17. http://doi.org/10.18632/oncotarget.16083.
12. Brandt WD, Matsui W, Rosenberg JE, He X, Ling S, Schaeffer EM, Berman DM. Urothelial carcinoma: stem cells on the edge. Cancer Metastasis Rev. 2009; 28: 291-304. http://doi.org/10.1007/s10555-009-9187-6.
13. Dinney CP, McConkey DJ, Millikan RE, Wu X, Bar-Eli M, Adam L, Kamat AM, Siefker-Radtke AO, Tuziak T, Sabichi AL, Grossman HB, Benedict WF, Czerniak B. Focus on bladder cancer. Cancer Cell. 2004; 6: 111-6. http://doi.org/10.1016/j.ccr.2004.08.002.
14. Cordon-Cardo C. Molecular alterations associated with bladder cancer initiation and progression. Scand J Urol Nephrol Suppl. 2008: 154-65. http://doi.org/10.1080/03008880802291915.
15. Zhang ZT, Pak J, Huang HY, Shapiro E, Sun TT, Pellicer A, Wu XR. Role of Ha-ras activation in superficial papillary pathway of urothelial tumor formation. Oncogene. 2001; 20: 1973-80. http://doi.org/10.1038/sj.onc.1204315.
16. Bjerkvig R, Tysnes BB, Aboody KS, Najbauer J, Terzis AJ. Opinion: the origin of the cancer stem cell: current controversies and new insights. Nat Rev Cancer. 2005; 5: 899-904. http://doi.org/10.1038/nrc1740.
17. Jordan CT. Cancer stem cells: controversial or just misunderstood? Cell Stem Cell. 2009; 4: 203-5. http://doi.org/10.1016/j.stem.2009.02.003.
18. Passegue E, Jamieson CH, Ailles LE, Weissman IL. Normal and leukemic hematopoiesis: are leukemias a stem cell disorder or a reacquisition of stem cell characteristics? Proc Natl Acad Sci U S A. 2003; 100 Suppl 1: 11842-9. http://doi.org/10.1073/pnas.2034201100.
19. Yang Z, Li C, Fan Z, Liu H, Zhang X, Cai Z, Xu L, Luo J, Huang Y, He L, Liu C, Wu S. Single-cell Sequencing Reveals Variants in ARID1A, GPRC5A and MLL2 Driving Self-renewal of Human Bladder Cancer Stem Cells. Eur Urol. 2017; 71: 8-12. http://doi.org/10.1016/j.eururo.2016.06.025.
20. Yang YM, Chang JW. Bladder cancer initiating cells (BCICs) are among EMA-CD44v6+ subset: novel methods for isolating undetermined cancer stem (initiating) cells. Cancer Invest. 2008; 26: 725-33. http://doi.org/10.1080/07357900801941845.
21. Li C, Yang Z, Du Y, Tang H, Chen J, Hu D, Fan Z. BCMab1, A Monoclonal Antibody against Aberrantly Glycosylated Integrin a3b1, Has Potent Antitumor Activity of Bladder Cancer In Vivo. Clinical Cancer Research. 2014; 20: 4001-13. http://doi.org/10.1158/1078-0432.ccr-13-3397.
22. Barker N, Ridgway RA, van Es JH, van de Wetering M, Begthel H, van den Born M, Danenberg E, Clarke AR, Sansom OJ, Clevers H. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature. 2009; 457: 608-11. http://doi.org/10.1038/nature07602.
23. Kurzrock EA, Lieu DK, Degraffenried LA, Chan CW, Isseroff RR. Label-retaining cells of the bladder: candidate urothelial stem cells. Am J Physiol Renal Physiol. 2008; 294: F1415-21. http://doi.org/10.1152/ajprenal.00533.2007.
24. Gaisa NT, Graham TA, McDonald SA, Canadillas-Lopez S, Poulsom R, Heidenreich A, Jakse G, Tadrous PJ, Knuechel R, Wright NA. The human urothelium consists of multiple clonal units, each maintained by a stem cell. J Pathol. 2011; 225: 163-71. http://doi.org/10.1002/path.2945.
25. Van Batavia J, Yamany T, Molotkov A, Dan H, Mansukhani M, Batourina E, Schneider K, Oyon D, Dunlop M, Wu XR, Cordon-Cardo C, Mendelsohn C. Bladder cancers arise from distinct urothelial sub-populations. Nat Cell Biol. 2014; 16: 982-91, 1-5. http://doi.org/10.1038/ncb3038.
26. McConkey DJ, Lee S, Choi W, Tran M, Majewski T, Lee S, Siefker-Radtke A, Dinney C, Czerniak B. Molecular genetics of bladder cancer: Emerging mechanisms of tumor initiation and progression. Urol Oncol. 2010; 28: 429-40. http://doi.org/10.1016/j.urolonc.2010.04.008.
27. Houghton J, Stoicov C, Nomura S, Rogers AB, Carlson J, Li H, Cai X, Fox JG, Goldenring JR, Wang TC. Gastric cancer originating from bone marrow-derived cells. Science. 2004; 306: 1568-71. http://doi.org/10.1126/science.1099513.
28. Lin H, Hu L, Chen L, Yu H, Wang Q, Chen P, Hu XT, Cai XJ, Guan XY. Chemically-induced cancers do not originate from bone marrow-derived cells. PLoS One. 2012; 7: e30493. http://doi.org/10.1371/journal.pone.0030493.
29. Shin K, Lim A, Odegaard JI, Honeycutt JD, Kawano S, Hsieh MH, Beachy PA. Cellular origin of bladder neoplasia and tissue dynamics of its progression to invasive carcinoma. Nat Cell Biol. 2014; 16: 469-78. http://doi.org/10.1038/ncb2956.
30. Kreso A, Dick JE. Evolution of the cancer stem cell model. Cell Stem Cell. 2014; 14: 275-91. http://doi.org/10.1016/j.stem.2014.02.006.
31. Bryan RT. Bladder cancer and cancer stem cells: basic science and implications for therapy. ScientificWorldJournal. 2011; 11: 1187-94. http://doi.org/10.1100/tsw.2011.117.
32. Chan KS, Espinosa I, Chao M, Wong D, Ailles L, Diehn M, Gill H, Presti J Jr, Chang HY, van de Rijn M, Shortliffe L, Weissman IL. Identification, molecular characterization, clinical prognosis, and therapeutic targeting of human bladder tumor-initiating cells. Proc Natl Acad Sci U S A. 2009; 106: 14016-21. http://doi.org/10.1073/pnas.0906549106.
33. Edris B, Weiskopf K, Volkmer AK, Volkmer JP, Willingham SB, Contreras-Trujillo H, Liu J, Majeti R, West RB, Fletcher JA, Beck AH, Weissman IL, van de Rijn M. Antibody therapy targeting the CD47 protein is effective in a model of aggressive metastatic leiomyosarcoma. Proc Natl Acad Sci U S A. 2012; 109: 6656-61. http://doi.org/10.1073/pnas.1121629109.
34. Su Y, Qiu Q, Zhang X, Jiang Z, Leng Q, Liu Z, Stass SA, Jiang F. Aldehyde dehydrogenase 1 A1-positive cell population is enriched in tumor-initiating cells and associated with progression of bladder cancer. Cancer Epidemiol Biomarkers Prev. 2010; 19: 327-37. http://doi.org/10.1158/1055-9965.epi-09-0865.
35. Li C, Du Y, Yang Z, He L, Wang Y, Hao L, Ding M, Yan R, Wang J, Fan Z. GALNT1-Mediated Glycosylation and Activation of Sonic Hedgehog Signaling Maintains the Self-Renewal and Tumor-Initiating Capacity of Bladder Cancer Stem Cells. Cancer Res. 2016; 76: 1273-83. http://doi.org/10.1158/0008-5472.can-15-2309.
36. Ning ZF, Huang YJ, Lin TX, Zhou YX, Jiang C, Xu KW, Huang H, Yin XB, Huang J. Subpopulations of stem-like cells in side population cells from the human bladder transitional cell cancer cell line T24. J Int Med Res. 2009; 37: 621-30. http://doi.org/10.1177/147323000903700304.
37. Zhou J, Wulfkuhle J, Zhang H, Gu P, Yang Y, Deng J, Margolick JB, Liotta LA, Petricoin E 3rd, Zhang Y. Activation of the PTEN/mTOR/STAT3 pathway in breast cancer stem-like cells is required for viability and maintenance. Proc Natl Acad Sci U S A. 2007; 104: 16158-63. http://doi.org/10.1073/pnas.0702596104.
38. Dalerba P, Cho RW, Clarke MF. Cancer stem cells: models and concepts. Annu Rev Med. 2007; 58: 267-84. http://doi.org/10.1146/annurev.med.58.062105.204854.
39. Monteiro J, Fodde R. Cancer stemness and metastasis: therapeutic consequences and perspectives. Eur J Cancer. 2010; 46: 1198-203. http://doi.org/10.1016/j.ejca.2010.02.030.
40. Conner SD. Regulation of Notch Signaling Through Intracellular Transport. Int Rev Cell Mol Biol. 2016; 323: 107-27. http://doi.org/10.1016/bs.ircmb.2015.12.002.
41. Wang Z, Li Y, Banerjee S, Sarkar FH. Exploitation of the Notch signaling pathway as a novel target for cancer therapy. Anticancer Res. 2008; 28: 3621-30.
42. Nam Y, Sliz P, Song L, Aster JC, Blacklow SC. Structural basis for cooperativity in recruitment of MAML coactivators to Notch transcription complexes. Cell. 2006; 124: 973-83. http://doi.org/10.1016/j.cell.2005.12.037.
43. Pannuti A, Foreman K, Rizzo P, Osipo C, Golde T, Osborne B, Miele L. Targeting Notch to target cancer stem cells. Clin Cancer Res. 2010; 16: 3141-52. http://doi.org/10.1158/1078-0432.ccr-09-2823.
44. Speigl L, Janssen N, Weide B, Pawelec G, Shipp C. Prognostic impact of the putative cancer stem cell markers ABCG2, CD133, ALDH1A1 and CD44V7/8 in metastatic melanoma. Br J Dermatol. 2016. http://doi.org/10.1111/bjd.15194.
45. Cordeiro BM, Oliveira ID, Alves MT, Saba-Silva N, Capellano AM, Cavalheiro S, Dastoli P, Toledo SR. SHH, WNT, and NOTCH pathways in medulloblastoma: when cancer stem cells maintain self-renewal and differentiation properties. Childs Nerv Syst. 2014; 30: 1165-72. http://doi.org/10.1007/s00381-014-2403-x.
46. Fan X, Eberhart CG. Medulloblastoma stem cells. J Clin Oncol. 2008; 26: 2821-7. http://doi.org/10.1200/jco.2007.15.2264.
47. Keith B, Simon MC. Hypoxia-inducible factors, stem cells, and cancer. Cell. 2007; 129: 465-72. http://doi.org/10.1016/j.cell.2007.04.019.
48. Gustafsson MV, Zheng X, Pereira T, Gradin K, Jin S, Lundkvist J, Ruas JL, Poellinger L, Lendahl U, Bondesson M. Hypoxia requires notch signaling to maintain the undifferentiated cell state. Dev Cell. 2005; 9: 617-28. http://doi.org/10.1016/j.devcel.2005.09.010.
49. Fan X, Matsui W, Khaki L, Stearns D, Chun J, Li YM, Eberhart CG. Notch pathway inhibition depletes stem-like cells and blocks engraftment in embryonal brain tumors. Cancer Res. 2006; 66: 7445-52. http://doi.org/10.1158/0008-5472.can-06-0858.
50. Klaus A, Birchmeier W. Wnt signalling and its impact on development and cancer. Nat Rev Cancer. 2008; 8: 387-98. http://doi.org/10.1038/nrc2389.
51. Nusse R. Wnt signaling and stem cell control. Cell Res. 2008; 18: 523-7. http://doi.org/10.1038/cr.2008.47.
52. de Sousa EMF, Vermeulen L. Wnt Signaling in Cancer Stem Cell Biology. Cancers (Basel). 2016; 8. http://doi.org/10.3390/cancers8070060.
53. Van Camp JK, Beckers S, Zegers D, Van Hul W. Wnt signaling and the control of human stem cell fate. Stem Cell Rev. 2014; 10: 207-29. http://doi.org/10.1007/s12015-013-9486-8.
54. Daniels DL, Weis WI. Beta-catenin directly displaces Groucho/TLE repressors from Tcf/Lef in Wnt-mediated transcription activation. Nat Struct Mol Biol. 2005; 12: 364-71. http://doi.org/10.1038/nsmb912.
55. Teng Y, Wang X, Wang Y, Ma D. Wnt/beta-catenin signaling regulates cancer stem cells in lung cancer A549 cells. Biochem Biophys Res Commun. 2010; 392: 373-9. http://doi.org/10.1016/j.bbrc.2010.01.028.
56. Lu R, Zhang YG, Sun J. STAT3 activation in infection and infection-associated cancer. Mol Cell Endocrinol. 2017. http://doi.org/10.1016/j.mce.2017.02.023.
57. Ojha R, Singh SK, Bhattacharyya S. JAK-mediated autophagy regulates stemness and cell survival in cisplatin resistant bladder cancer cells. Biochim Biophys Acta. 2016; 1860: 2484-97. http://doi.org/10.1016/j.bbagen.2016.07.021.
58. Luo Y, Cui Y, Cao X, Li X, Chen A, Zhang J, Chen X, Cao J. 8-Bromo-7-methoxychrysin-blocked STAT3/Twist axis inhibits the stemness of cancer stem cell-like cell originated from SMMC-7721 cells. Acta Biochim Biophys Sin (Shanghai). 2017; 49: 458-64. http://doi.org/10.1093/abbs/gmx025.
59. Junk DJ, Bryson BL, Smigiel JM, Parameswaran N, Bartel CA, Jackson MW. Oncostatin M promotes cancer cell plasticity through cooperative STAT3-SMAD3 signaling. Oncogene. 2017. http://doi.org/10.1038/onc.2017.33.
60. Ghoshal S, Fuchs BC, Tanabe KK. STAT3 is a key transcriptional regulator of cancer stem cell marker CD133 in HCC. Hepatobiliary Surg Nutr. 2016; 5: 201-3. http://doi.org/10.21037/hbsn.2016.03.02.
61. Singh AK, Arya RK, Maheshwari S, Singh A, Meena S, Pandey P, Dormond O, Datta D. Tumor heterogeneity and cancer stem cell paradigm: updates in concept, controversies and clinical relevance. Int J Cancer. 2015; 136: 1991-2000. http://doi.org/10.1002/ijc.28804.
62. Zhang ZT, Pak J, Shapiro E, Sun TT, Wu XR. Urothelium-specific expression of an oncogene in transgenic mice induced the formation of carcinoma in situ and invasive transitional cell carcinoma. Cancer Res. 1999; 59: 3512-7. http://doi.org/Published July 1999
63. Cicalese A, Bonizzi G, Pasi CE, Faretta M, Ronzoni S, Giulini B, Brisken C, Minucci S, Di Fiore PP, Pelicci PG. The tumor suppressor p53 regulates polarity of self-renewing divisions in mammary stem cells. Cell. 2009; 138: 1083-95. http://doi.org/10.1016/j.cell.2009.06.048.
64. Tran MN, Goodwin Jinesh G, McConkey DJ, Kamat AM. Bladder cancer stem cells. Curr Stem Cell Res Ther. 2010; 5: 387-95. http://doi.org/HTTP://DOI.ORG/10.2174/157488810793351640
65. Ishii H, Iwatsuki M, Ieta K, Ohta D, Haraguchi N, Mimori K, Mori M. Cancer stem cells and chemoradiation resistance. Cancer Sci. 2008; 99: 1871-7. http://doi.org/10.1111/j.1349-7006.2008.00914.x.
66. Rich JN. Cancer stem cells in radiation resistance. Cancer Res. 2007; 67: 8980-4. http://doi.org/10.1158/0008-5472.CAN-07-0895.
67. Tharakan ST, Inamoto T, Sung B, Aggarwal BB, Kamat AM. Curcumin potentiates the antitumor effects of gemcitabine in an orthotopic model of human bladder cancer through suppression of proliferative and angiogenic biomarkers. Biochem Pharmacol. 2010; 79: 218-28. http://doi.org/10.1016/j.bcp.2009.08.007.
68. Schinkel AH, Jonker JW. Mammalian drug efflux transporters of the ATP binding cassette (ABC) family: an overview. Adv Drug Deliv Rev. 2003; 55: 3-29. http://doi.org/org/10.1016/s0169-409x(02)00169-2
69. Zhou S, Schuetz JD, Bunting KD, Colapietro AM, Sampath J, Morris JJ, Lagutina I, Grosveld GC, Osawa M, Nakauchi H, Sorrentino BP. The ABC transporter Bcrp1/ABCG2 is expressed in a wide variety of stem cells and is a molecular determinant of the side-population phenotype. Nat Med. 2001; 7: 1028-34. http://doi.org/10.1038/nm0901-1028.
70. He X, Marchionni L, Hansel DE, Yu W, Sood A, Yang J, Parmigiani G, Matsui W, Berman DM. Differentiation of a highly tumorigenic basal cell compartment in urothelial carcinoma. Stem Cells. 2009; 27: 1487-95. http://doi.org/10.1002/stem.92.
71. Matsui W, Wang Q, Barber JP, Brennan S, Smith BD, Borrello I, McNiece I, Lin L, Ambinder RF, Peacock C, Watkins DN, Huff CA, Jones RJ. Clonogenic multiple myeloma progenitors, stem cell properties, and drug resistance. Cancer Res. 2008; 68: 190-7. http://doi.org/10.1158/0008-5472.can-07-3096.
72. Kastan MB, Schlaffer E, Russo JE, Colvin OM, Civin CI, Hilton J. Direct demonstration of elevated aldehyde dehydrogenase in human hematopoietic progenitor cells. Blood. 1990; 75: 1947-50.
73. Idelchik M, Begley U, Begley TJ, Melendez JA. Mitochondrial ROS control of cancer. Semin Cancer Biol. 2017. http://doi.org/10.1016/j.semcancer.2017.04.005.
74. Chio IIC, Tuveson DA. ROS in Cancer: The Burning Question. Trends Mol Med. 2017; 23: 411-29. http://doi.org/10.1016/j.molmed.2017.03.004.
75. Diehn M, Cho RW, Lobo NA, Kalisky T, Dorie MJ, Kulp AN, Qian D, Lam JS, Ailles LE, Wong M, Joshua B, Kaplan MJ, Wapnir I, et al. Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature. 2009; 458: 780-3. http://doi.org/10.1038/nature07733.
76. Vitale I, Manic G, De Maria R, Kroemer G, Galluzzi L. DNA Damage in Stem Cells. Mol Cell. 2017; 66: 306-19. http://doi.org/10.1016/j.molcel.2017.04.006.
77. Eisele L, Klein-Hitpass L, Chatzimanolis N, Opalka B, Boes T, Seeber S, Moritz T, Flasshove M. Differential expression of drug-resistance-related genes between sensitive and resistant blasts in acute myeloid leukemia. Acta Haematol. 2007; 117: 8-15. http://doi.org/10.1159/000096854.
78. Todaro M, Alea MP, Di Stefano AB, Cammareri P, Vermeulen L, Iovino F, Tripodo C, Russo A, Gulotta G, Medema JP, Stassi G. Colon cancer stem cells dictate tumor growth and resist cell death by production of interleukin-4. Cell Stem Cell. 2007; 1: 389-402. http://doi.org/10.1016/j.stem.2007.08.001.
79. Aggarwal BB, Kunnumakkara AB, Harikumar KB, Gupta SR, Tharakan ST, Koca C, Dey S, Sung B. Signal transducer and activator of transcription-3, inflammation, and cancer: how intimate is the relationship? Ann N Y Acad Sci. 2009; 1171: 59-76. http://doi.org/10.1111/j.1749-6632.2009.04911.x.
80. Mashima T. Cancer Stem Cells (CSCs) as a Rational Therapeutic Cancer Target, and Screening for CSC-targeting Drugs. Yakugaku Zasshi. 2017; 137: 129-32. http://doi.org/10.1248/yakushi.16-00229-1.
81. Yamazaki J, Estecio MR, Lu Y, Long H, Malouf GG, Graber D, Huo Y, Ramagli L, Liang S, Kornblau SM, Jelinek J, Issa JP. The epigenome of AML stem and progenitor cells. Epigenetics. 2013; 8: 92-104. http://doi.org/10.4161/epi.23243.
82. Avgustinova A, Benitah SA. The epigenetics of tumour initiation: cancer stem cells and their chromatin. Curr Opin Genet Dev. 2016; 36: 8-15. http://doi.org/10.1016/j.gde.2016.01.003.
83. Polak P, Karlic R, Koren A, Thurman R, Sandstrom R, Lawrence MS, Reynolds A, Rynes E, Vlahovicek K, Stamatoyannopoulos JA, Sunyaev SR. Cell-of-origin chromatin organization shapes the mutational landscape of cancer. Nature. 2015; 518: 360-4. http://doi.org/10.1038/nature14221.
84. Christophorou MA, Castelo-Branco G, Halley-Stott RP, Oliveira CS, Loos R, Radzisheuskaya A, Mowen KA, Bertone P, Silva JC, Zernicka-Goetz M, Nielsen ML, Gurdon JB, Kouzarides T. Citrullination regulates pluripotency and histone H1 binding to chromatin. Nature. 2014; 507: 104-8. http://doi.org/10.1038/nature12942.
85. Jamieson CH, Ailles LE, Dylla SJ, Muijtjens M, Jones C, Zehnder JL, Gotlib J, Li K, Manz MG, Keating A, Sawyers CL, Weissman IL. Granulocyte-macrophage progenitors as candidate leukemic stem cells in blast-crisis CML. N Engl J Med. 2004; 351: 657-67. http://doi.org/10.1056/NEJMoa040258.
86. Willingham SB, Volkmer JP, Gentles AJ, Sahoo D, Dalerba P, Mitra SS, Wang J, Contreras-Trujillo H, Martin R, Cohen JD, Lovelace P, Scheeren FA, Chao MP, et al. The CD47-signal regulatory protein alpha (SIRPa) interaction is a therapeutic target for human solid tumors. Proc Natl Acad Sci U S A. 2012; 109: 6662-7. http://doi.org/10.1073/pnas.1121623109.
87. Fialkow PJ, Jacobson RJ, Papayannopoulou T. Chronic myelocytic leukemia: clonal origin in a stem cell common to the granulocyte, erythrocyte, platelet and monocyte/macrophage. Am J Med. 1977; 63: 125-30. http://doi.org/org/10.1016/0002-9343(77)90148-6
88. Haraguchi N, Ishii H, Mimori K, Tanaka F, Ohkuma M, Kim HM, Akita H, Takiuchi D, Hatano H, Nagano H, Barnard GF, Doki Y, Mori M. CD13 is a therapeutic target in human liver cancer stem cells. J Clin Invest. 2010; 120: 3326-39. http://doi.org/10.1172/jci42550.
89. O’Brien CA, Pollett A, Gallinger S, Dick JE. A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature. 2007; 445: 106-10. http://doi.org/10.1038/nature05372.
90. Schatton T, Murphy GF, Frank NY, Yamaura K, Waaga-Gasser AM, Gasser M, Zhan Q, Jordan S, Duncan LM, Weishaupt C, Fuhlbrigge RC, Kupper TS, Sayegh MH, et al. Identification of cells initiating human melanomas. Nature. 2008; 451: 345-9. http://doi.org/10.1038/nature06489.
91. Merlo LM, Pepper JW, Reid BJ, Maley CC. Cancer as an evolutionary and ecological process. Nat Rev Cancer. 2006; 6: 924-35. http://doi.org/10.1038/nrc2013.

