INTRODUCTION
Sarcomas are a group of heterogeneous malignant tumors that arise from mesenchymal origin. These malignancies are characterized by more than 50 subtypes with distinct clinical presentation and histology [1]. Although relatively rare, sarcomas account for nearly 21% of all pediatric solid cancers and are the third leading cause of cancer-related deaths in patients under the age of 20 [2, 3]. Despite the significant development of novel multidisciplinary treatments, including surgery, chemotherapy and radiation therapy, the 5-year survival rate has reached to a plateau in recent years [4]. This phenomenon is seen particularly in patients with metastatic or recurrent advanced disease, where the overall median survival is around 15 months, and only about 10% of these cases are still alive at five years [5].
Since the introduction of adjuvant and neoadjuvant chemotherapy in the 1970s, chemotherapy with multimodality therapeutic strategies have made considerable progress in improving the survival rate and quality of life for patients with certain types of sarcomas, especially for osteosarcoma, Ewing’s sarcoma and rhabdomyosarcoma [6–12]. However, progress has remained stagnant over the last three decades with conventional chemotherapy approach [13, 14]. The development of drug resistance to chemotherapy is one of the major reasons for treatment failure, but the mechanisms of drug resistance have not been completely understood. Multiple subtypes make drug resistance mechanisms more diverse and controversial in sarcoma. To date, none of the ongoing attempts to address current mechanisms have been applied in the clinic with the aim of restoring drug efficacy in sarcoma. Therefore, identification of novel mechanisms of drug resistance in sarcoma that can be directly translated to clinic is required.
With the rapid development of next-generation sequencing technology to whole genomes and transcriptomes, it has been shown that only 2% of the human genome encodes mRNAs, whereas 98% of transcriptional products are noncoding RNAs (ncRNAs) [15]. ncRNAs have emerged as a class of cellular regulators involved in biological processes such as proliferation, differentiation, apoptosis and cell cycle, whereas mutations and dysregulation of ncRNAs have been linked to diverse human diseases, including sarcomas [16, 17]. Recent evidence has highlighted ncRNAs as being associated with drug resistance in sarcoma, mainly in two distinct subtype forms: microRNAs (miRs) and long noncoding RNAs (lncRNAs). miRs have moved to the forefront of ncRNA research in the past decade, while the role of lncRNAs is also emerging in different areas of cancer biology [18]. In this review, we summarize recent discoveries of the molecular functions of ncRNAs in cellular pathways related to drug resistance, and their potential as novel therapeutic strategies to overcome treatment failure of chemotherapies in sarcoma.
NCRNAS AND DRUG RESISTANCE IN SARCOMA
ncRNAs are extensively transcribed within the human genome [19, 20]. They lack valid open reading frames and have no protein-coding capacity [21–23]; however, ncRNAs represent a broad class of structurally and functionally distinct RNAs (Table 1). These include long and short ncRNAs that represent around 60% of the RNAs distributed throughout the human genome [24, 25]. Short ncRNAs are no more than 200 nucleotides (nt) [26]. This group is composed of miR, PIWI interacting RNA, small nucleolar RNA, and others [26]. In contrast, transcripts longer than 200 nt are regarded as lncRNAs [27]. This group is diverse and highly abundant [28], and contains long intergenic ncRNAs, long intronic ncRNAs, pseudogene RNAs, to name a few [26]. Novel classes of ncRNAs are constantly being discovered, including circular RNAs [29] and enhancer RNAs [30].
Table 1: Classes of discovered human ncRNAs
Noncoding RNAs |
Class |
Symbol |
Length (nt) |
Functions |
Significance |
|---|---|---|---|---|---|
House-keeping ncRNAs |
Transfer RNAs |
tRNAs |
73–94 |
Connect amino acids with mRNA |
Translation |
Ribosomal RNAs |
rRNAs |
121–5070 |
Component of ribosome |
Translation |
|
Vault RNAs |
vRNAs |
86–141 |
Component of vault |
Expulsion of xenobiotics, such as chemotherapeutic compounds |
|
Small ncRNAs |
MicroRNAs |
miRNAs |
18–25 |
Modulate protein-coding genes, guide suppression of translation, Drosha and Dicer dependent small ncRNAs |
Regulation of proliferation, |
Piwi-interacting RNAs |
piRNAs |
26–30 |
Bind to Piwi proteins, principally restricted to the germline |
Involved in germ cell development, stem self-renewal, and retrotransposon silencing |
|
Small nuclear RNAs |
snRNAs |
150~ |
Assemble with proteins into spliceosomes to remove introns during mRNA processing |
Aid in the regulation of transcription factors or RNA polymerase II, maintaining the telomeres |
|
Small nucleolar RNAs |
snoRNAs |
60–200 |
Guide modifications of other noncoding RNAs, function as miRNA to regulate mRNAs, alternative splicing |
Associated with the development of some cancers |
|
Promoter-associated small RNAs |
paRNAs |
20–200 |
Involved in the regulation of the transcription of protein-coding genes by targeting epigenetic silencing complexes |
Relation to diseases has not yet been discovered |
|
Transcription initiation RNAs |
tiRNAs |
~18 |
Involved in the regulation of the transcription of protein-coding genes by targeting epigenetic silencing complexes |
Relation to diseases has not yet been discovered |
|
Long ncRNAs |
Long intergenic noncoding RNAs |
lincRNAs |
> 200 |
Involved in diverse biological processes, such as mRNA splicing and miRNA silencing |
Involved in tumorigenesis and cancer metastasis |
Long intronic noncoding RNAs |
> 200 |
Likely to be involved in post-transcriptional gene silencing |
Aberrantly expressed in human cancers |
||
Telomere associated noncoding RNAs |
TERRAs |
> 200 |
Negative regulation of telomere length and activity through inhibition of telomerase |
Possible impact on telomere-associated diseases, including many cancers |
|
Long noncoding RNAs with dual functions |
> 200 |
Both protein-coding and functionally regulatory RNA capacity |
Deregulation has been described in cancers |
||
Transcribed pseudogenes |
> 200 |
Regulation of tumor suppressors and oncogenes by acting as microRNA decoys |
Often deregulated during tumorigenesis and cancer progression |
||
Transcribed-ultraconserved regions |
T-UCRs |
> 200 |
Antisense inhibitors for protein-coding genes or other ncRNAs |
Expression is often altered in some cancers, possible involvement in tumorigenesis |
|
Circular RNAs |
circRNAs |
> 100 |
Do not have 5′ or 3′ ends, act as miRNA sponge, transport miRNAs, regulate mRNA through limited base pairing |
May be used to study pathogenesis and devise therapeutic interventions |
|
Enhancer RNAs |
eRNAs |
50–2000 |
Important components in enhancer activity, have effect on the trans-criptional regulation in cis and in trans |
May provide promising regulatory routes in tumor suppression |
ncRNAs were considered to be junk or debris derived from the DNA transcriptional process initially [25]. However, recent studies have revealed that ncRNAs could be deregulated or mutated in many human cancers and they play an important role in pathological processes, including in tumorigenesis, metastasis and treatment resistance [31–34]. The development of drug resistance in sarcoma is affected by many factors, most of which involve a change in expression of certain genes. Mutations, amplifications or deletions within oncogenes and/or tumor suppressors were considered to be associated with the development of drug resistance [35–37]. ncRNAs have emerged as major regulators of epigenetic, transcriptional, and post-transcriptional gene expression and alternative splicing, which provides cells with yet another mode to fine-tune their transcriptome and adjust their proteome in response to stimuli. ncRNAs, mainly in the form of miRs and lncRNAs, have been linked to sarcoma drug resistance through abnormal gene regulation related to the activity of drug efflux transporters; the activation of drug target mutations or changes; DNA repair and cell cycle arrest; apoptosis and cell-survival pathways; cancer stem cells (CSCs) and autophagy involved signaling pathways [38–42].
ROLE OF MIRS IN DRUG RESISTANT SARCOMA
miRs are a large family of conserved endogenous single-stranded small ncRNAs derived from the human genome [43]. The initial discovery of miR was in 1993 [44], and to date, over 1,400 human miRs have been identified [45]. Over one-third of the genes in human genome are regulated by miRs [46]. miRs bind to the 3′-untranslated region (3′-UTR) of its mRNA target, based on a partial base-pairing complementarity [47]. Translation of the mRNA is ultimately prevented either by transcript degradation, inhibition of translation, or mRNA decay, leading to a reduced level of protein [48].
Mutation patterns or aberrant expressions of miRs have been detected in sarcoma and have been associated with clinical features such as progression, metastasis and drug resistance [49]. Upregulation of miRs induces variations in drug metabolism and disposition through the regulation of target mRNAs (Table 2). The products of these target genes can be drug transporters, nuclear receptors, transcription factors or drug-metabolizing enzymes [50]. This leads to altered expression of proteins that are involved in the signal transduction network of drug resistance, and ultimately changing the chemosensitivity of cancer cells [51].
Table 2: Summary of miRs involved in drug resistant sarcoma
miRs |
Sarcomas |
Alteration |
Drugs |
Resistance mechanisms |
Major targets |
References |
|---|---|---|---|---|---|---|
miR-143 |
Liposarcoma Osteosarcoma |
↓ |
Doxorubicin |
Alterations in drug targets Cell cycle Evasion of apoptosis Cancer stem cells Autophagy |
Top2A PRC1 PLK1 Bcl-2 ALDH1 CD133 ATG2B LC3-II |
|
miR-124 |
Osteosarcoma |
↓ |
Camptothecin Etoposide Doxorubicin |
DNA repair |
ATMIN PARP1 |
[55] |
miR-708 |
Ewing’s sarcoma |
↓ |
Doxorubicin Etoposide |
DNA repair |
EYA3 |
[56] |
miR-17 |
Synovial sarcoma |
↑ |
Doxorubicin |
Cell cycle |
p21 |
[57] |
miR-140 |
Osteosarcoma |
↓ |
Methotrexate |
Cell cycle |
HDAC4 |
[58] |
miRNA-215 |
Osteosarcoma |
↑ |
Methotrexate |
Cell cycle |
DTL |
[59] |
miR-301a |
Osteosarcoma |
↑ |
Doxorubicin |
Evasion of apoptosis |
AMPKα1 |
[63] |
miR-382 |
Osteosarcoma |
↓ |
Doxorubicin Cisplatin Methotrexate |
Evasion of apoptosis |
HIPK3 |
[64] |
Let-7d |
Osteosarcoma |
↑ |
Doxorubicin Cisplatin Paclitaxel Etoposide |
Evasion of apoptosis Cancer stem cells |
Bcl-2 Caspase-3 Oct3/4 Sox2 Nanog Lin28B HMGA2 |
[65] |
miR-138 |
Osteosarcoma |
↓ |
Cisplatin |
Evasion of apoptosis |
EZH2 |
[66] |
miR-21 |
Osteosarcoma |
↑ |
Cisplatin |
Evasion of apoptosis |
Bcl-2 |
|
miR-34a |
Ewing’s sarcoma |
↓ |
Doxorubicin Vincristine |
Evasion of apoptosis |
Not reported |
[69] |
miR-125b |
Ewing’s sarcoma |
↑ |
Doxorubicin Etoposide Vincristine |
Evasion of apoptosis |
P53 Bak |
[74] |
miRNA-193a-5p |
Osteosarcoma Ewing’s sarcoma |
↑ |
Cisplatin |
Evasion of apoptosis Activation of cell-survival pathways |
TAp73β Wnt/β-catenin pathway |
|
miR-202 |
Osteosarcoma |
↑ |
Doxorubicin |
Activation of cell-survival pathways |
PDCD4 |
[76] |
miR-221 |
Osteosarcoma |
↑ |
Cisplatin |
Activation of cell-survival pathways |
PTEN |
[82] |
miR-217 |
Osteosarcoma |
↓ |
Cisplatin |
Activation of cell-survival pathways |
KRAS |
[86] |
miR-100 |
Chondrosarcoma |
↓ |
Cisplatin |
Activation of cell-survival pathways |
mTOR |
[87] |
miR-497 |
Osteosarcoma |
↓ |
Cisplatin |
Activation of cell-survival pathways |
VEGFA |
[89] |
miR-92a miR-99b miR-422a |
Osteosarcoma |
↑ |
Ifosfamide |
Wnt/β-catenin pathway |
[90] |
|
miR-132 |
Osteosarcoma |
↓ |
Ifosfamide |
Activation of cell-survival pathways |
Wnt/β-catenin pathway |
[90] |
miR-33a |
Osteosarcoma |
↑ |
Cisplatin |
Activation of cell-survival pathways |
TWIST |
[91] |
miR-34c |
Osteosarcoma |
↓ |
Doxorubicin Cisplatin Methotrexate |
Activation of cell-survival pathways |
North1 LEF1 |
[93] |
miR-146b-5p |
Osteosarcoma |
↑ |
Doxorubicin Cisplatin Methotrexate |
Activation of cell-survival pathways |
ZNRF3 |
[97] |
miR-199a-3p |
Osteosarcoma |
↓ |
Doxorubicin |
Cancer stem cells |
CD44 |
[102] |
miR-29b-1 |
Osteosarcoma |
↓ |
Doxorubicin Cisplatin Etoposide |
Cancer stem cells |
Oct3/4 Sox2 Nanog CD133 N-Myc CCND2 |
[105] |
miR-199-5p |
Osteosarcoma |
↓ |
Cisplatin |
Autophagy |
LC3-II Beclin-1 |
[114] |
miR-30a |
Osteosarcoma |
↓ |
Doxorubicin |
Autophagy |
Beclin-1 |
[115] |
miR-22 |
Osteosarcoma |
↑ |
Doxorubicin Cisplatin |
Autophagy |
HMGB1 |
|
miR-101 |
Osteosarcoma |
↓ |
Doxorubicin |
Autophagy |
LC3 AVOs |
[119] |
miR-488 |
Osteosarcoma |
↑ |
Doxorubicin |
Evasion of apoptosis |
Bim |
[145] |
miR-184 |
Osteosarcoma |
↑ |
Doxorubicin |
Evasion of apoptosis |
Bcl-2 |
[146] |
miR-23b |
Chondrosarcoma |
↓ |
Cisplatin |
Evasion of apoptosis |
Src-Akt pathway |
[149] |
miR-141 |
Neuroblastoma |
↓ |
Cisplatin |
Not reported |
FUS gene |
[150] |
miRs are associated with changes to drug targets in drug resistant sarcoma
Quantitative or qualitative alterations to drug targets can be a factor contributing to drug resistance. miRs that inhibit DNA topoisomerase 2A (Top2A), such as miR-143, are implicated in drug resistance in dedifferentiated liposarcoma (Figure 1). The miR expression profile showed a decreased miR-143 in dedifferentiated liposarcoma tissues. Restored miR-143 decreased the expression of Top2A, inhibited proliferation and induced apoptosis in dedifferentiated liposarcoma cells. Top2A encoded by the Top2A gene is a nuclear enzyme. It catalyzes the transient breaking and rejoining of two strands of duplex DNA, thus altering the topology of DNA for transcription and replication [52]. The gene encoding Top2A functions as the target for several chemotherapeutic agents used for sarcoma treatment, such as doxorubicin (DOX) and etoposide [53]. Amplification or mutations in this gene have been associated with the development of drug resistance. In dedifferentiated liposarcoma, reduced expression of miR-143 increased nuclear expression of Top2A and cell proliferation, and this led to the dedifferentiated liposarcoma cells acquiring resistance to Top2A-targeted drugs [54].
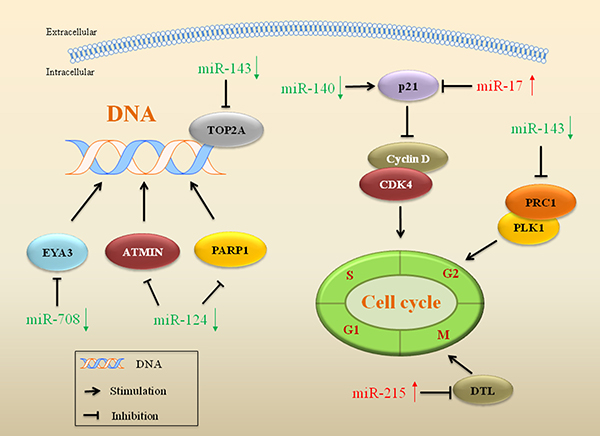
Figure 1: miRs involved in drug target alterations, DNA repair and cell cycle in drug resistant sarcoma. miRs that regulate gene expression of crucial proteins in drug targets, DNA repair and cell cycle are implicated in drug resistance in sarcoma. Red arrows indicate upregulation of miRs and green arrows indicate downregulation.
miRs influence DNA repair in drug resistant sarcoma
miRs that have an impact on DNA repair, such as miR-124 and miR-708, are implicated in resistance to drugs aimed at damage tumor cell DNA (Figure 1). The damaged DNA leads to genetic instability and in turn activates apoptosis, when DNA repair proves inefficient. Overexpression of miR-124 reduced poly ADP-ribose polymerase 1 (PARP1) and ataxia telangiectasia mutated interactor (ATMIN) proteins expression in osteosarcoma cells, which reduced DNA repair capacity and increased sensitivity to DNA-damaging drugs, such as DOX [55]. Overexpression of miR-124 induced the DNA repair defect through binding to the 3′-UTR of the mRNAs of PARP1 and ATMIN, thus reversed drug resistance in osteosarcoma [55]. Decreased expression of another miR, miR-708, induced EYA3 expression and led to drug resistance in Ewing’s sarcoma. EYA3 is a highly expressed transcriptional cofactor that enhances DNA repair in response to DNA damage. Restored miR-708 directly targeted EYA3 and reduced its expression, thus sensitized Ewing’s sarcoma cells to the DNA-damaging drugs etoposide and DOX. This confirmed that miR-708 could be a promising target for sensitization to DNA-damaging drugs applied in the treatment of Ewing’s sarcoma [56].
miRs regulate cell cycle-associated genes in drug resistant sarcoma
miRs associated with drug resistance have been shown to regulate genes related to cell cycle in sarcoma, include miR-17, miR-140, miR-215 and miR-143 (Figure 1). Overexpression of miR-17 suppressed DOX-evoked higher expression of p21, a tumor suppressor, and conferred the drug resistance in synovial sarcoma. Treatment of the synovial sarcoma cells with anti-miR-17 rescued expression of p21, which may be a therapeutic target to reverse drug resistance in sarcoma [57]. miR-140 is involved in chemosensitivity with reduced cell proliferation via G1 and G2 phase arrest mediated in part, by the suppression of histone deacetylase 4 (HDAC4) expression. HDAC4 is related to chemoresistance via repression of p21 and regulating cell-cycle progression and differentiation. Transfection with miR-140 reversed methotrexate-resistance through induction of the p21 gene accompanied by G1 and G2 phase arrest only in cell lines containing wild type of p53 [58]. Another miR, miR-215, inhibited expression of denticleless protein homolog (DTL) and led to an increased resistance to methotrexate and Tomudex in osteosarcoma. DTL is a cell cycle G2/M checkpoint regulatory protein. Inhibition of DTL induced G2-arrest and decreased cell proliferation. miR-215 inhibited expression of the DTL gene accompanied by upregulation of p21 resulting in G2-arrest, and thus drug resistance emerged in osteosarcoma [59]. Protein Regulator of cytokinesis 1 (PRC1) is a protein that is highly expressed during S and G2/M phases of the cell cycle [60]. Polo-like kinase 1 (PLK1) or serine/threonine-protein kinase 13 (STPK13), is an early trigger enzyme for G2/M transition [61]. miR-143 was downregulated in dedifferentiated liposarcoma. Restoring miR-143 expression inhibited cytokinesis in dedifferentiated liposarcoma cells, with decreased expression of PRC1 and PLK1. Treatment with a PLK1 inhibitor potently induced G2-M cell cycle arrest and apoptosis in liposarcoma cells, indicating that miR-143 could be a therapeutic target for the reversal of drug resistance [54].
miRs affect evasion of apoptosis in drug resistant sarcoma
Dysfunctions of miRs, including miR-301a, miR-382, Led-7, miR-21, miR-34a and miR-125b, are associated with defective apoptosis-induced drug resistance in sarcoma (Figure 2) [62]. DOX treatment promoted the cleavage of caspase-3, which was blocked by miR-301a in osteosarcoma, suggesting that miR-301a enhanced the resistance of osteosarcoma cells to DOX by inhibiting apoptosis [63]. miR-382, increased cleavage of caspase-3 and enhanced drug-induced cell apoptosis in osteosarcoma. Downregulation of miR-382 was associated with a poor chemoresponse to DOX, cisplatin (CDDP), and methotrexate in osteosarcoma [64]. High expression of Let-7d also decreased caspase-3 cleavage induced by chemotherapeutic agents, in combination with reduced apoptosis and drug resistance in osteosarcoma [65]. In contrast, increased miR-138 expression enhanced caspase-3 activation and CCDP-induced apoptosis by targeting EZH2, resulting in reinforcing chemosensitivity to CCDP in osteosarcoma [66].
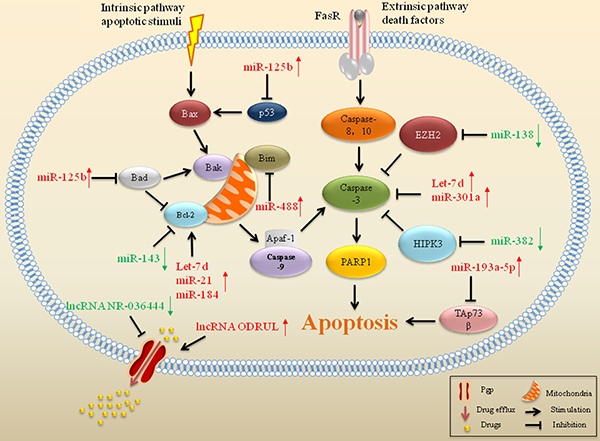
Figure 2: ncRNAs associated with apoptosis and drug efflux in drug resistant sarcoma. miRs and lncRNAs that interfere with gene expression of significant proteins in apoptosis and drug efflux have an impact on drug resistance in sarcoma. Red arrows indicate upregulation of miRs and green arrows indicate downregulation.
miR-21 is implicated in the modulation of drug resistance in sarcoma and serum miR-21 can serve as a potential biomarker for prediction of chemotherapeutic sensitivity in patients with osteosarcoma [67]. Transfection with miR-21 mimics depleted the sensitivity of osteosarcoma cells to CDDP, while miR-21 suppression led to enhanced CDDP cytotoxicity. Changes in chemoresponse induced by miR-21 upregulation were ameliorated by downregulation of apoptotic inhibitor Bcl-2 [68]. In addition to miR-21, miR Let-7d also reduced cell sensitivity to DOX, CDDP, etoposide and paclitaxel, consistent with increased expression of Bcl-2 in osteosarcoma [65].
miR-34a was associated with good prognosis in Ewing’s sarcoma, and reinforced that miR-34a could sensitize Ewing’s sarcoma cells to DOX and vincristine [69]. miR-34a is a component of the p53 tumor suppressor network [70]. Loss of miR-34a is mainly caused by inactivating mutations of p53, genetic alterations of the region Chr1p36 (which contains the coding region for miR-34a), or epigenetic changes [71–73]. miR-125b could induce drug resistance through evasion of p53-dependent apoptosis in DOX-resistant Ewing’s sarcoma cells. Knockdown of miR-125b enhanced the sensitivity of Ewing’s sarcoma cells to DOX, which was associated with an increased level of p53. In addition to DOX, overexpression of miR-125b also resulted in enhanced resistance to etoposide and vincristine in Ewing’s sarcoma [74]. Other members of the p53 tumor suppressor network also influence drug-induced apoptosis in sarcoma. High levels of miR-193a-5p blocked CDDP-induced apoptosis via inhibition of TAp73β, a p53-family protein. The role of the miR-193a-5p/TAp73β axis in drug resistance was confirmed in sarcomas, including in osteosarcoma and Ewing’s sarcoma [75].
miRs interfere with cell-survival pathways in drug resistant sarcoma
Expressions of miRs interfere with cell growth and survival mediated by the PI3K/Akt pathway, which plays a role in drug resistance in sarcoma (Figure 3). To identify PI3K/Akt-related miRs, a group of miRs was shown to have elevated expression upon treatment of drugs, such as miR-202 and miR-221, while other miRs were downregulated, including miR-217, miR-100 and miR-497. Overexpression of miR-202 promoted drug resistance by targeting programmed cell death 4 (PDCD4) in osteosarcoma [76]. PDCD4 is a tumor suppressor that inhibits the PI3K/Akt pathway and enhances apoptosis [77]. Activation of the PI3K/Akt pathway leads to drug resistance, while suppression of PI3K/Akt signaling in turn restores drug sensitivity in cancers, including in sarcoma [78–81]. High levels of miR-221 in osteosarcoma also induced resistance to CDDP and increased cell survival by suppressing PTEN expression [82]. miR-221 directly targeted the 3′-UTR of PTEN and regulated CDDP resistance through the PI3K/Akt pathway with loss of PTEN [82]. As a tumor suppressor, miR-217 increases sensitivity to CDDP through inhibiting the PI3K/Akt signaling pathway in various cancers [83–85]. miR-217 enhanced sensitivity to CDDP via targeting KRAS and inhibiting the PI3K/Akt pathway in osteosarcoma [86]. In chondrosarcoma, reduced expression of miR-100 was associated with resistance to CDDP with activation of the Akt-mTOR pathway. Upregulated expression of miR-100 sensitized chondrosarcoma cells to CDDP by inhibiting the Akt/mTOR pathway [87]. Loss of miR-497 induced drug resistance through targeting vascular endothelial growth factor A (VEGFA) in osteosarcoma. VEGFA is a member of the vascular endothelial growth factor (VEGF) family, which is a positive modulator of the PI3K/Akt pathway [88]. With depletive VEGFA, decreased miR-497 induced Akt activation, resulting in resistance to CDDP. In contrast, increased miR-497 inhibited cell survival and promoted sensitivity to CDDP with inhibition of the PI3K/Akt pathway [89].
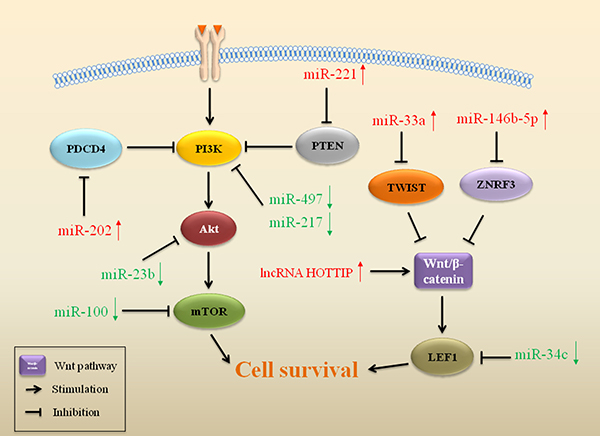
Figure 3: ncRNAs affect cell-survival pathways in drug resistant sarcoma. ncRNAs that regulate gene expression of important regulators in cell-survival pathways play a role in drug resistance in sarcoma. Red arrows indicate upregulation and green arrows indicate downregulation of miRs.
The Wnt pathway is another cell-survival signaling pathway associated with drug resistance in sarcoma (Figure 3). miR profiles identified five discrete miRs (miR-92a, miR-99b, miR-132, miR-193a-5p and miR-422a) in osteosarcoma, which have an impact on drug response through targeting the Wnt, TGFβ and MAP kinase pathways [90]. Overexpression of miR-33a also resulted in drug resistance in osteosarcoma, which was mediated by downregulation of TWIST [91]. TWIST is a negative regulator of CDDP resistance by suppressing the Wnt/β-catenin pathway in osteosarcoma [92]. Low expression of miR-34c contributed to drug resistance development in osteosarcoma [93]. miR-34c may target and inhibit Notch1 and LEF1 expression. Notch1 and LEF1 activated the Notch and the Wnt signaling pathway, promoting drug resistance in cancers, including in osteosarcoma [94–96]. Overexpression of miR-146b-5p promoted resistance to DOX, CDDP and methotrexate by promoting the Wnt/β-catenin pathway in osteosarcoma. Zinc and ring finger 3 (ZNRF3) is a direct target of miR-146b-5p. Overexpression of miR-146b-5p downregulated ZNRF3 , thus activate Wnt/β-catenin pathway and induced drug resistance in osteosarcoma [97].
miRs are involved in CSC-related pathways in drug resistant sarcoma
The CSC hypothesis postulates that within a tumor, a minor subpopulation of cells possess the capacity to self-renew and drive the heterogeneous lineages of cancer cells that account for the initiation, proliferation, recurrence, metastasis and therapeutic resistance in cancer [98]. CSCs may be involved in resistance to drugs and toxins through the expression of drug efflux transporters, activation of DNA repair, and evasion of apoptosis [99]. For example, CSCs express more anti-apoptotic proteins, including Bcl-2, FLIP, IAP-1, IAP-2 and survivin than normal tumor cells to resist apoptosis in sarcoma [100]. Even though the molecular pathways modulating CSCs remain unclear, growing evidence highlights the roles of miRs in the regulation of drug resistance related to CSCs (Figure 4).
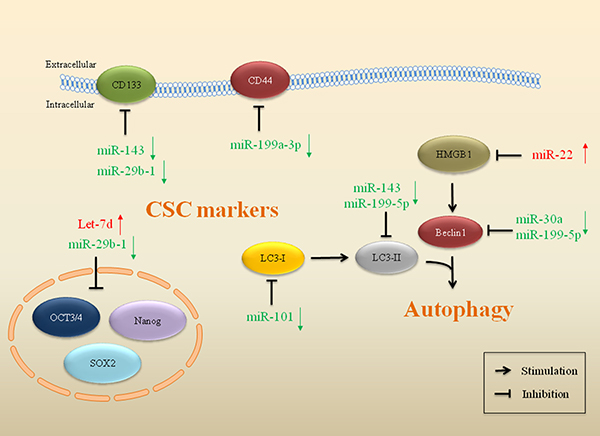
Figure 4: miRs impact on CSCs and autophagy in drug resistant sarcoma. miRs that mediate gene expression of markers and regulators in CSCs and autophagy are implicated in drug resistance in sarcoma. Red arrows indicate upregulation of miRs and green arrows indicate downregulation.
Cluster of differentiation 44 (CD44) is a cell surface marker commonly used to identify and enrich CSCs [101]. Higher expression of this cell-surface glycoprotein was associated with response to chemotherapy in osteosarcoma. miR-199a-3p is one of the most dramatically decreased miRs in osteosarcoma. Transfection of miR-199a-3p enhanced drug sensitivity, consistent with downregulation of CD44 in osteosarcoma. The 3′-UTR of CD44 mRNA was the direct target of miR-199a-3p [102]. In chemoresistant osteosarcoma cell lines, it was also observed that loss of miR-143 expression was accompanied by a high level of ALDH1+CD133+ cells. Forced miR-143 expression could reverse drug resistance in DOX-resistant osteosarcoma CSCs through activation of apoptosis [103]. CSCs in sarcoma can also induce evasion of apoptosis through p53 and Rb pathways to resist chemotherapeutic drugs [104]. The AB-OS cell line is a CSC cell line selected from human osteosarcoma MG63 cells. Overexpression of miR-29b-1 reduced the expression of stem cell markers (Oct3/4, Sox2, Nanog, CD133, and N-Myc) and anti-apoptotic markers (Bcl-2 and IAP-2) in 3AB-OS CSCs, suggesting that miR-29b-1 can inhibit stemness properties of 3AB-OS CSCs and sensitize 3AB-OS cells to chemotherapy drugs (DOX, CDDP and etoposide) in osteosarcoma [105]. On the contrary, upregulation of Let-7d reduced cell sensitivity to apoptosis and induced drug resistance in 3AB-OS cells, in combination with decreased caspase-3 and increased Bcl-2 in osteosarcoma [65].
miRs impact autophagy in drug resistant sarcoma
Recent findings have suggested that dysregulated miRs are closely connected to autophagy modulation and cancer drug resistance (Figure 4). Autophagy is an evolutionary conserved catabolic process involving in the degradation of proteins and organelles, cellular remodeling and survival, promoting the controlled degradation of cytoplasmic material both at steady state and during nutrient deprivation [106]. Tumor cell autophagy is activated during chemotherapy and contributes to chemotherapy resistance [107, 108]. Thus, inhibition of autophagy can re-sensitize resistant cancer cells and enhance cytotoxicity of chemotherapeutic agents [109, 110]. Certain miRs have been revealed as potent inhibitors of drug resistance-related autophagy [111], indicating their potential role in cancer therapy [112, 113].
It has been shown that miR143 and miR-199-5p suppress autophagy and reverse drug resistance in osteosarcoma (Figure 4). Upregulation of miR-143 was correlated with the sensitization of osteosarcoma cells to DOX, which is accompanied by downregulation of LC3-II and Bcl-2 [103]. CDDP treatment upregulated protein levels of LC3-II and Beclin1 in osteosarcoma cells, indicating that autophagy was activated. Restoration of miR-199a-5p inhibited CDDP-induced autophagy, indicating that miR-199a-5p promotes the cytotoxicity of CDDP in osteosarcoma through inhibition of autophagy [114]. Reduced miR-30a expression activated Beclin-1-dependent autophagy, which influenced drug resistance in osteosarcoma cells to DOX. miR-30a directly binds to the 3′-UTR of Beclin-1 gene, resulting in the suppression of Beclin-1 mediated autophagy. Elevated miR-30a expression promoted drug-induced apoptosis and inhibited autophagy activity in osteosarcoma [115, 116]. The expression of miR-22 was induced by chemotherapeutic drugs in osteosarcoma. High-mobility group box 1 (HMGB1) is a direct target of miR-22, which could facilitate autophagy and promote drug resistance. miR-22 blocked the HMGB1-mediated autophagy and reversed drug resistance in osteosarcoma [117, 118]. Autophagy-characteristic acidic vesicular organelles (AVOs) are characteristic in autophagy. miR-101 could also reduce the DOX-induced AVOs and decrease autophagy-related proteins in osteosarcoma. Upregulated miR-101 enhanced chemosensitivity to DOX through inhibiting autophagy, thus reversing autophagy-mediated drug resistance in osteosarcoma [119].
ROLE OF LNCRNAS IN DRUG RESISTANT SARCOMA
LncRNAs are transcriptional polyadenylated RNAs that were first reported in 1990 [120]. They consist of exons and introns in structure, but lack valid ORFs [22]. LncRNAs regulate gene expression at epigenetic, transcriptional and post-transcriptional levels, through epigenetic regulation, splicing, imprinting, transcriptional regulation and subcellular transport [121–124]. They function in both cis, to act on the same chromosome, and trans, to affect different chromosomes [125, 126]. LncRNAs play a role in a number of biological processes, including chromatin modification, telomere biology and subcellular structural organization [127]. Current estimates of unique lncRNAs present in humans that range from 7000–23,000, with a growing cohort being validated as having a role in human disease processes, including in sarcoma [128, 129]. The misregulated lncRNAs can function as decoys, scaffolds, signals or guides for specific regulatory modules, resulting in a gene expression profile in favor of cancer drug resistance development (Table 3) [24, 130].
Table 3: Summary of lncRNAs involved in drug resistant sarcoma
lncRNAs |
Sarcomas |
Alteration |
Drugs |
Resistance mechanisms |
Major targets |
References |
|---|---|---|---|---|---|---|
ENST00000563280 (ODRUL) |
Osteosarcoma |
↑ |
Doxorubicin |
Evasion of apoptosis |
ABCB1 |
|
NR-036444 |
Osteosarcoma |
↓ |
Doxorubicin |
Evasion of apoptosis |
ABCB1 |
[131] |
HOTTIP |
Osteosarcoma |
↑ |
Cisplatin |
Activation of cell-survival pathways |
Wnt/β-catenin pathways |
[139] |
lncRNAs are associated with drug efflux and accumulation in drug resistant sarcoma
A human lncRNA-mRNA combined microarray revealed that 3,465 lncRNAs (1,761 up and 1,704 down) were aberrantly expressed in DOX-resistant osteosarcoma cells. This lncRNA-mRNA co-expression network identified that upregulated expression of lncRNA EST00000563280 and downregulated expression of lncRNA NR-036444 could stimulate the expression of Pgp to transport the drugs out of the cell, leading to drug resistance to DOX in osteosarcoma (Figure 2) [131]. Pgp, also known as MDR1, is a 170 kDa cell membrane glycoprotein encoded by the ATP-binding cassette subfamily B member 1 (ABCB1) gene. ABCB1 is a member of the ubiquitous ATP-binding cassette (ABC) superfamily, which contains 49 members divided into seven subclasses ranging from ABCA to ABCG [132]. As an ATP-dependent drug transporter, Pgp can extrude cytostatic agents against a drug concentration gradient at the expense of ATP hydrolysis. The action of Pgp triggers drug efflux and promotes resistance to drugs in cancer [133, 134]. In sarcoma, the acute induced expression of Pgp has been demonstrated as a function of time in response to in vivo exposure to chemotherapy [135, 136]. The novel lncRNA EST00000563280, termed osteosarcoma DOX-resistance related upregulated lncRNA (ODRUL), was the most upregulated lncRNA in DOX-resistant osteosarcoma cells. ODRUL plays a role in the emergence of resistance to DOX by inducing Pgp expression in osteosarcoma [137]. This indicates that lncRNAs may be a novel target for reversing drug resistance in sarcoma.
lncRNAs regulate cell-survival pathways in drug resistant sarcoma
In addition to promoting cancer cell survival, activation of Wnt/β-catenin pathway can also induce drug resistance through affecting cell cycle in osteosarcoma (Figure 3) [138]. HOTTIP is another lncRNA upregulated in osteosarcoma. Overexpression of HOTTIP promoted cellular resistance to CDDP, accompanied by an activation of Wnt/β-catenin pathway. Decreasing HOTTIP induced cell cycle arrest in G1 phase through inhibiting the Wnt/β-catenin pathway, thus reversing drug resistance in osteosarcoma [139].
The therapeutic potential of ncRNAs in drug resistant sarcoma
Identification of chemotherapy-resistant tumors at diagnosis is a first priority for treating sarcoma, where more individualized, effective, and less toxic treatments are highly desirable. ncRNA therapiesappear to be a novel field in which ncRNA activity is the major target of the intervention [140].
miRs are believed to be relatively safe and more effective in cancer treatment in early preclinical studies [141]. Recent evidence has confirmed that the selective modulation of miR activity could improve responses to chemotherapy. The rationale for developing miR therapies is based on the premise that aberrant expression of miRs play key roles in the development of resistance to chemotherapy and that correcting these miR deficiencies by either antagonizing or restoring miR function may provide a therapeutic benefit [142].
Overexpressions of some miRs are involved in drug-resistant human sarcoma. Inhibition of these miRs may increase chemoresponse to antitumor therapy. For example, miR-21 promotes drug resistance in sarcoma, and inhibition of miR-21 may function to reverse drug resistance in osteosarcoma cells [68]. Further understanding of miR-21-mediated signaling pathways will help to promote the clinical use of miR-21 in cancer treatment [143]. DOX is a widely used chemotherapy drug, but it has a narrow therapeutic window with toxic side-effects. The antitumor activity of DOX is achieved by induction of cell apoptosis through interacting with DNA [144]. The miR-488 is overexpressed in osteosarcoma and is induced by hypoxia through binding to the hypoxia response element. Overexpression of miR-488 decreased the sensitivity to DOX via direct targeting of the mediator of apoptosis Bim. Transfection of the miR-488 inhibitor resulted in an increase in apoptosis and enhanced the sensitivity of osteosarcoma cells to DOX treatment [145]. Overexpression of Let-7d also reduced osteosarcoma cell sensitivity to apoptosis induced by chemotherapy agents DOX, CDDP, etoposide and paclitaxel, concomitant with a decrease in caspase-3 and increase in BCL2 expression [65]. miR-184 also contributes to chemoresistance in osteosarcoma. The expression of miR-184 was upregulated in DOX-treated osteosarcoma cells. BCL2L1 served as the direct target gene of miR-184. Overexpression of miR-184 reduced DOX-induced cell apoptosis and led to poor response to drug therapy by targeting BCL2L1 [146]. These observations indicate that miR-488, Let-7d and miR-184 may serve as predictors of response to chemotherapy and as potential therapeutic targets for the treatment of drug resistant osteosarcoma.
Some miRs are downregulated and related to drug resistance in sarcoma. Exogenous upregulation of these may promote sensitivity to chemotherapy. In Ewing’s sarcoma, miRNA expression was investigated in patients with different clinical outcomes. Microarray analysis identified a signature of miR-34a as a prognostic indicator for Ewing’s sarcoma [147]. Functional analysis indicated that miR-34a increased tumor sensitivity to the effects of DOX and vincristine in Ewing’s sarcoma cells, and that restoration of miR-34a activity may be useful to decrease malignancy. The result suggested a potential role of miR-34a in the treatment of sarcoma and may spare excessive long-term toxicity to Ewing’s sarcoma patients [69]. Chondrosarcoma is the second most common type of primary bone malignancy in the United States. The expression of miR-125b was decreased in both parental and DOX-resistant chondrosarcoma cells. Overexpression of miR-125 enhanced the sensitivity of both parental and DOX-resistant cells to DOX by inhibiting glycolysis through directly targeting the oncogene, ErbB2 [148]. CDDP is another widely used antitumor agent via effective DNA-damaging. However, chondrosarcoma is resistant to conventional chemotherapies, including CDDP. Compared with the parental cells, miR-23b was significantly downregulated in CDDP-resistant cells. Src kinase is a direct target of miR-23b in chondrosarcoma cells. The chemo-sensitization effect induced by miR-23b-mediated suppression of the Src-Akt pathway suggests that miR-23b could be a potential therapeutic target to reverse the acquirement of resistance to CDDP in chondrosarcoma [149]. These findings provide a novel therapy to overcoming chemoresistance in human chondrosarcoma cells. In neuroblastoma, miR-141 upregulation promoted chemosensitivity to CDDP, which was associated with the FUS gene, which may also help in the development of therapeutic strategies for the treatments of patients [150].
Moreover, lncRNAs have become a new frontier for drug development. The structural and functional novelty of lncRNAs offers promise as anticancer therapeutics that may avoid the emergence of drug resistance commonly seen with the currently used agents. For example, HOTAIR is a novel factor involved in drug resistance and metastasis in various types of solid tumors. Overexpression of HOTAIR was involved in drug resistance in primary sarcoma and decreased levels of HOTAIR were associated with good chemoresponse, suggesting that clinical application of HOTAIR may be a potential predictor of treatment efficacy [151].
Although there are still several obstacles to overcome before clinical testing of miR therapeutics, such as delivery and chemical modification of ncRNA modulators, it can be expected that ncRNAs and ncRNA-targeting oligonucleotides may become promising tools in the fight against drug resistant sarcoma in the near future [152].
CONCLUSIONS AND FUTURE PERSPECTIVES
The current rapid expansion of research on ncRNAs evidently shows that ncRNAs have many implications in drug resistant sarcoma. This review brings together information on the roles of ncRNAs in drug resistant sarcomas, including miRs and lncRNAs. Although the exact mechanisms of ncRNAs in drug resistance in sarcoma are still unclear, these studies highlight the increasing interest and development of knowledge of ncRNAs in drug resistant sarcoma.
Although recent findings present a positive trend in development, the research progress of ncRNAs in drug resistant sarcoma is still in its infancy. There is a tough route from discovering the function of ncRNAs to its clinical use. Despite extensive investigation and distinct progress in this field, there are still many challenges. Most of the current studies focus on the role of ncRNA in chemoresistance with a single agent, such as either DOX or CDDP. However, sarcoma patients commonly receive combined chemotherapy in the clinic. Thus, it is necessary to evaluate the role of ncRNA in the regulation of multidrug resistance. With regard to ncRNA-based therapeutic agent, the field will benefit from the development of RNA delivery technology and chemical modifications of ncRNAs. Novel RNA delivery technology may protect ncRNA from degradation and enable ncRNA to reach tumors efficiently and specifically. The chemical modifications of ncRNAs can enhance the stability of ncRNAs and reduce toxicity, minimizing side effects. Furthermore, discovery of more miR targets will be useful to better identify new and more effective drug targets [153].
A rising strategy to solve gene specificity limitations is the technology of genome editing by clustered regulatory interspaced short palindromic repeats-associated endonuclease 9 (CRISPR-Cas9), which is a rapid and efficient way to generate total or partial downregulation of specific genes, including ncRNAs, by the targeted interruption of the promoter and the chosen sequence through insertion of polyadenylation signals. Furthermore, CRISPR-Cas9 can be applied to achieve ncRNA overexpression from its endogenous locus by inserting a strong promoter upstream of the ncRNA sequence or by targeting transcriptional activator complexes to the promoter [154, 155]. In the future, after gaining insight into the ncRNAs involved in drug resistance mechanisms in sarcoma, ncRNA-based approaches may provide important advances in overcoming drug resistance and improving chemotherapy responses of sarcoma patients.
ACKNOWLEDGMENTS
This work was supported by the National Natural Science Foundation of China (NO.: 81372871, 81502332 and 81672176). Dr. Li is supported by a scholarship from the China Scholarship Council (201606370194). Dr. Duan is supported, in part, through a Grant from Sarcoma Foundation of America (SFA) (222433), a Grant from National Cancer Institute (NCI)/National Institutes of Health (NIH), UO1, CA151452-01.
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
REFERENCES
1. Fletcher CD HP, Mertens F, Bridge JA. World Health Organization Classification of Tumors. Pathology and Genetics of Tumors of Soft Tissue and Bone. 4th ed Lyon, France: IARC Press. 2013.
2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016; 66:7–30. https://doi.org/10.3322/caac.21332.
3. Burningham Z, Hashibe M, Spector L, Schiffman JD. The epidemiology of sarcoma. Clin Sarcoma Res. 2012; 2:14. https://doi.org/10.1186/2045-3329-2-14.
4. Allemani C, Weir HK, Carreira H, Harewood R, Spika D, Wang XS, Bannon F, Ahn JV, Johnson CJ, Bonaventure A, Marcos-Gragera R, Stiller C, Azevedo e Silva G, et al. Global surveillance of cancer survival 1995-2009: analysis of individual data for 25,676,887 patients from 279 population-based registries in 67 countries (CONCORD-2). Lancet. 2015; 385:977–1010. https://doi.org/10.1016/S0140-6736(14)62038-9.
5. Italiano A, Mathoulin-Pelissier S, Cesne AL, Terrier P, Bonvalot S, Collin F, Michels JJ, Blay JY, Coindre JM, Bui B. Trends in survival for patients with metastatic soft-tissue sarcoma. Cancer. 2011; 117:1049–54. https://doi.org/10.1002/cncr.25538.
6. Bielack SS, Kempf-Bielack B, Delling G, Exner GU, Flege S, Helmke K, Kotz R, Salzer-Kuntschik M, Werner M, Winkelmann W, Zoubek A, Jurgens H, Winkler K. Prognostic factors in high-grade osteosarcoma of the extremities or trunk: an analysis of 1,702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. J Clin Oncol. 2002; 20:776–90.
7. Goorin AM, Schwartzentruber DJ, Devidas M, Gebhardt MC, Ayala AG, Harris MB, Helman LJ, Grier HE, Link MP, Pediatric Oncology Group. Presurgical chemotherapy compared with immediate surgery and adjuvant chemotherapy for nonmetastatic osteosarcoma: Pediatric Oncology Group Study POG-8651. J Clin Oncol. 2003; 21:1574–80. https://doi.org/10.1200/JCO.2003.08.165.
8. Ferrari S, Sundby Hall K, Luksch R, Tienghi A, Wiebe T, Fagioli F, Alvegard TA, Brach Del Prever A, Tamburini A, Alberghini M, Gandola L, Mercuri M, Capanna R, et al. Nonmetastatic Ewing family tumors: high-dose chemotherapy with stem cell rescue in poor responder patients. Results of the Italian Sarcoma Group/Scandinavian Sarcoma Group III protocol. Ann Oncol. 2011; 22:1221–7. https://doi.org/10.1093/annonc/mdq573.
9. Grier HE, Krailo MD, Tarbell NJ, Link MP, Fryer CJ, Pritchard DJ, Gebhardt MC, Dickman PS, Perlman EJ, Meyers PA, Donaldson SS, Moore S, Rausen AR, et al. Addition of ifosfamide and etoposide to standard chemotherapy for Ewing’s sarcoma and primitive neuroectodermal tumor of bone. N Engl J Med. 2003; 348:694–701. https://doi.org/10.1056/NEJMoa020890.
10. Ferrari S, Smeland S, Mercuri M, Bertoni F, Longhi A, Ruggieri P, Alvegard TA, Picci P, Capanna R, Bernini G, Muller C, Tienghi A, Wiebe T, et al. Neoadjuvant chemotherapy with high-dose Ifosfamide, high-dose methotrexate, cisplatin, and doxorubicin for patients with localized osteosarcoma of the extremity: a joint study by the Italian and Scandinavian Sarcoma Groups. J Clin Oncol. 2005; 23:8845–52. https://doi.org/10.1200/JCO.2004.00.5785.
11. Harwood JL, Alexander JH, Mayerson JL, Scharschmidt TJ. Targeted Chemotherapy in Bone and Soft-Tissue Sarcoma. Orthop Clin North Am. 2015; 46:587–608. https://doi.org/10.1016/j.ocl.2015.06.011.
12. Hiniker SM, Donaldson SS. Recent advances in understanding and managing rhabdomyosarcoma. F1000Prime Rep. 2015; 7:59. https://doi.org/10.12703/P7-59.
13. Miller KD, Siegel RL, Lin CC, Mariotto AB, Kramer JL, Rowland JH, Stein KD, Alteri R, Jemal A. Cancer treatment and survivorship statistics, 2016. CA Cancer J Clin. 2016; 66:271–89. https://doi.org/10.3322/caac.21349.
14. Howlader N, Noone AM, Krapcho M, Garshell J, Miller D, Altekruse SF, Kosary CL, Yu M, Ruhl J, Tatalovich Z, Mariotto A, Lewis DR, Chen HS, et al, eds. SEER Cancer Statistics Review, 1975-2012, National Cancer Institute. Bethesda, MD, http://seer.cancer.gov/csr/1975_2012/, based on November 2014 SEER data submission, posted to the SEER web site, April 2015.
15. International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature. 2004; 431:931–45. https://doi.org/10.1038/nature03001.
16. Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011; 12:861–74. https://doi.org/10.1038/nrg3074.
17. Taft RJ, Pang KC, Mercer TR, Dinger M, Mattick JS. Non-coding RNAs: regulators of disease. J Pathol. 2010; 220:126–39. https://doi.org/10.1002/path.2638.
18. Dey BK, Mueller AC, Dutta A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription. 2014; 5:e944014. https://doi.org/10.4161/21541272.2014.944014.
19. Ulitsky I, Bartel DP. lincRNAs: genomics, evolution, and mechanisms. Cell. 2013; 154:26–46. https://doi.org/10.1016/j.cell.2013.06.020.
20. Bentwich I, Avniel A, Karov Y, Aharonov R, Gilad S, Barad O, Barzilai A, Einat P, Einav U, Meiri E, Sharon E, Spector Y, Bentwich Z. Identification of hundreds of conserved and nonconserved human microRNAs. Nat Genet. 2005; 37:766–70. https://doi.org/10.1038/ng1590.
21. ENCODE Project Consortium, Birney E, Stamatoyannopoulos JA, Dutta A, Guigo R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE, Kuehn MS, Taylor CM, et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007; 447:799–816. https://doi.org/10.1038/nature05874.
22. Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, et al. The transcriptional landscape of the mammalian genome. Science. 2005; 309:1559–63. https://doi.org/10.1126/science.1112014.
23. Ponting CP, Belgard TG. Transcribed dark matter: meaning or myth? Hum Mol Genet. 2010; 19:R162–8. https://doi.org/10.1093/hmg/ddq362.
24. Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011; 43:904–14. https://doi.org/10.1016/j.molcel.2011.08.018.
25. Wright MW, Bruford EA. Naming ‘junk’: human non-protein coding RNA (ncRNA) gene nomenclature. Hum Genomics. 2011; 5:90–8.
26. Sana J, Faltejskova P, Svoboda M, Slaby O. Novel classes of non-coding RNAs and cancer. J Transl Med. 2012; 10:103. https://doi.org/10.1186/1479-5876-10-103.
27. Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, Bell I, Cheung E, Drenkow J, et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007; 316:1484–8. https://doi.org/10.1126/science.1138341.
28. Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009; 458:223–7. https://doi.org/10.1038/nature07672.
29. Lasda E, Parker R. Circular RNAs: diversity of form and function. RNA. 2014; 20:1829–42. https://doi.org/10.1261/rna.047126.114.
30. Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, Harmin DA, Laptewicz M, Barbara-Haley K, Kuersten S, Markenscoff-Papadimitriou E, Kuhl D, Bito H, et al. Widespread transcription at neuronal activity-regulated enhancers. Nature. 2010; 465:182–7. https://doi.org/10.1038/nature09033.
31. Kapranov P, Drenkow J, Cheng J, Long J, Helt G, Dike S, Gingeras TR. Examples of the complex architecture of the human transcriptome revealed by RACE and high-density tiling arrays. Genome Res. 2005; 15:987–97. https://doi.org/10.1101/gr.3455305.
32. Yang C, Wu D, Gao L, Liu X, Jin Y, Wang D, Wang T, Li X. Competing endogenous RNA networks in human cancer: hypothesis, validation, and perspectives. Oncotarget. 2016; 7:13479–90. https://doi.org/10.18632/oncotarget.7266.
33. Nagano T, Fraser P. No-nonsense functions for long noncoding RNAs. Cell. 2011; 145:178–81. https://doi.org/10.1016/j.cell.2011.03.014.
34. Alsidawi S, Malek E, Driscoll JJ. MicroRNAs in brain metastases: potential role as diagnostics and therapeutics. Int J Mol Sci. 2014; 15:10508–26. https://doi.org/10.3390/ijms150610508.
35. Bonasio R, Tu S, Reinberg D. Molecular signals of epigenetic states. Science. 2010; 330:612–6. https://doi.org/10.1126/science.1191078.
36. Croce CM. Oncogenes and cancer. N Engl J Med. 2008; 358:502–11. https://doi.org/10.1056/NEJMra072367.
37. Sherr CJ. Principles of tumor suppression. Cell. 2004; 116:235–46.
38. Goldman B. Multidrug resistance: can new drugs help chemotherapy score against cancer? J Natl Cancer Inst. 2003; 95:255–7.
39. Lipovich L, Johnson R, Lin CY. MacroRNA underdogs in a microRNA world: evolutionary, regulatory, and biomedical significance of mammalian long non-protein-coding RNA. Biochim Biophys Acta. 2010; 1799:597–615. https://doi.org/10.1016/j.bbagrm.2010.10.001.
40. Haenisch S, Cascorbi I. miRNAs as mediators of drug resistance. Epigenomics. 2012; 4:369–81. https://doi.org/10.2217/epi.12.39.
41. Harries LW. Long non-coding RNAs and human disease. Biochem Soc Trans. 2012; 40:902–6. https://doi.org/10.1042/BST20120020.
42. d’Adda di Fagagna F. Living on a break: cellular senescence as a DNA-damage response. Nat Rev Cancer. 2008; 8:512–22. https://doi.org/10.1038/nrc2440.
43. Carthew RW, Sontheimer EJ. Origins and Mechanisms of miRNAs and siRNAs. Cell. 2009; 136:642–55. https://doi.org/10.1016/j.cell.2009.01.035.
44. Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993; 75:843–54.
45. Kozomara A, Griffiths-Jones S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 2011; 39:D152–7. https://doi.org/10.1093/nar/gkq1027.
46. Iorio MV, Croce CM. MicroRNAs in cancer: small molecules with a huge impact. J Clin Oncol. 2009; 27:5848–56. https://doi.org/10.1200/JCO.2009.24.0317.
47. Lytle JR, Yario TA, Steitz JA. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′ UTR as in the 3′ UTR. Proc Natl Acad Sci U S A. 2007; 104:9667–72. https://doi.org/10.1073/pnas.0703820104.
48. Winter J, Diederichs S. MicroRNA biogenesis and cancer. Methods Mol Biol. 2011; 676:3–22. https://doi.org/10.1007/978-1-60761-863-8_1.
49. Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014; 20:460–9. https://doi.org/10.1016/j.molmed.2014.06.005.
50. Zheng T, Wang J, Chen X, Liu L. Role of microRNA in anticancer drug resistance. Int J Cancer. 2010; 126:2–10. https://doi.org/10.1002/ijc.24782.
51. Hong L, Yang Z, Ma J, Fan D. Function of miRNA in controlling drug resistance of human cancers. Curr Drug Targets. 2013; 14:1118–27.
52. Azarova AM, Lyu YL, Lin CP, Tsai YC, Lau JY, Wang JC, Liu LF. Roles of DNA topoisomerase II isozymes in chemotherapy and secondary malignancies. Proc Natl Acad Sci USA. 2007; 104:11014–9. https://doi.org/10.1073/pnas.0704002104.
53. Pommier Y, Leo E, Zhang H, Marchand C. DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem Biol. 2010; 17:421–33. https://doi.org/10.1016/j.chembiol.2010.04.012.
54. Ugras S, Brill E, Jacobsen A, Hafner M, Socci ND, Decarolis PL, Khanin R, O’Connor R, Mihailovic A, Taylor BS, Sheridan R, Gimble JM, Viale A, et al. Small RNA sequencing and functional characterization reveals MicroRNA-143 tumor suppressor activity in liposarcoma. Cancer Res. 2011; 71:5659–69. https://doi.org/10.1158/0008-5472.CAN-11-0890.
55. Chen SM, Chou WC, Hu LY, Hsiung CN, Chu HW, Huang YL, Hsu HM, Yu JC, Shen CY. The Effect of MicroRNA-124 Overexpression on Anti-Tumor Drug Sensitivity. PLoS One. 2015; 10:e0128472. https://doi.org/10.1371/journal.pone.0128472.
56. Robin TP, Smith A, McKinsey E, Reaves L, Jedlicka P, Ford HL. EWS/FLI1 regulates EYA3 in Ewing sarcoma via modulation of miRNA-708, resulting in increased cell survival and chemoresistance. Mol Cancer Res. 2012; 10:1098–108. https://doi.org/10.1158/1541-7786.MCR-12-0086.
57. Minami Y, Kohsaka S, Tsuda M, Yachi K, Hatori N, Tanino M, Kimura T, Nishihara H, Minami A, Iwasaki N, Tanaka S. SS18-SSX-regulated miR-17 promotes tumor growth of synovial sarcoma by inhibiting p21WAF1/CIP1. Cancer Sci. 2014; 105:1152–9. https://doi.org/10.1111/cas.12479.
58. Song B, Wang Y, Xi Y, Kudo K, Bruheim S, Botchkina GI, Gavin E, Wan Y, Formentini A, Kornmann M, Fodstad O, Ju J. Mechanism of chemoresistance mediated by miR-140 in human osteosarcoma and colon cancer cells. Oncogene. 2009; 28:4065–74. https://doi.org/10.1038/onc.2009.274.
59. Song B, Wang Y, Titmus MA, Botchkina G, Formentini A, Kornmann M, Ju J. Molecular mechanism of chemoresistance by miR-215 in osteosarcoma and colon cancer cells. Mol Cancer. 2010; 9:96. https://doi.org/10.1186/1476-4598-9-96.
60. Jiang W, Jimenez G, Wells NJ, Hope TJ, Wahl GM, Hunter T, Fukunaga R. PRC1: a human mitotic spindle-associated CDK substrate protein required for cytokinesis. Mol Cell. 1998; 2:877–85.
61. Holtrich U, Wolf G, Brauninger A, Karn T, Bohme B, Rubsamen-Waigmann H, Strebhardt K. Induction and down-regulation of PLK, a human serine/threonine kinase expressed in proliferating cells and tumors. Proc Natl Acad Sci USA. 1994; 91:1736–40.
62. Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000; 100:57–70.
63. Zhang Y, Duan G, Feng S. MicroRNA-301a modulates doxorubicin resistance in osteosarcoma cells by targeting AMP-activated protein kinase alpha 1. Biochem Biophys Res Commun. 2015; 459:367–73. https://doi.org/10.1016/j.bbrc.2015.02.101.
64. Xu M, Jin H, Xu CX, Sun B, Mao Z, Bi WZ, Wang Y. miR-382 inhibits tumor growth and enhance chemosensitivity in osteosarcoma. Oncotarget. 2014; 5:9472–83. https://doi.org/10.18632/oncotarget.2418.
65. Di Fiore R, Drago-Ferrante R, Pentimalli F, Di Marzo D, Forte IM, Carlisi D, De Blasio A, Tesoriere G, Giordano A, Vento R. Let-7d miRNA Shows Both Antioncogenic and Oncogenic Functions in Osteosarcoma-Derived 3AB-OS Cancer Stem Cells. J Cell Physiol. 2016; 231:1832–41. https://doi.org/10.1002/jcp.25291.
66. Zhu Z, Tang J, Wang J, Duan G, Zhou L, Zhou X. MiR-138 Acts as a Tumor Suppressor by Targeting EZH2 and Enhances Cisplatin-Induced Apoptosis in Osteosarcoma Cells. PLoS One. 2016; 11:e0150026. https://doi.org/10.1371/journal.pone.0150026.
67. Yuan J, Chen L, Chen X, Sun W, Zhou X. Identification of serum microRNA-21 as a biomarker for chemosensitivity and prognosis in human osteosarcoma. J Int Med Res. 2012; 40:2090–7.
68. Ziyan W, Yang L. MicroRNA-21 regulates the sensitivity to cisplatin in a human osteosarcoma cell line. Ir J Med Sci. 2016; 185:85–91. https://doi.org/10.1007/s11845-014-1225-x.
69. Nakatani F, Ferracin M, Manara MC, Ventura S, Del Monaco V, Ferrari S, Alberghini M, Grilli A, Knuutila S, Schaefer KL, Mattia G, Negrini M, Picci P, et al. miR-34a predicts survival of Ewing’s sarcoma patients and directly influences cell chemo-sensitivity and malignancy. J Pathol. 2012; 226:796–805. https://doi.org/10.1002/path.3007.
70. He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y, Xue W, Zender L, Magnus J, Ridzon D, Jackson AL, Linsley PS, Chen C, et al. A microRNA component of the p53 tumour suppressor network. Nature. 2007; 447:1130–4. https://doi.org/10.1038/nature05939.
71. Lodygin D, Tarasov V, Epanchintsev A, Berking C, Knyazeva T, Korner H, Knyazev P, Diebold J, Hermeking H. Inactivation of miR-34a by aberrant CpG methylation in multiple types of cancer. Cell Cycle. 2008; 7:2591–600. https://doi.org/10.4161/cc.7.16.6533.
72. He X, He L, Hannon GJ. The guardian’s little helper: microRNAs in the p53 tumor suppressor network. Cancer Res. 2007; 67:11099–101. https://doi.org/10.1158/0008-5472.CAN-07-2672.
73. Hermeking H. p53 enters the microRNA world. Cancer Cell. 2007; 12:414–8. https://doi.org/10.1016/j.ccr.2007.10.028.
74. Iida K, Fukushi J, Matsumoto Y, Oda Y, Takahashi Y, Fujiwara T, Fujiwara-Okada Y, Hatano M, Nabashima A, Kamura S, Iwamoto Y. miR-125b develops chemoresistance in Ewing sarcoma/primitive neuroectodermal tumor. Cancer Cell Int. 2013; 13:21. https://doi.org/10.1186/1475-2867-13-21.
75. Jacques C, Calleja LR, Baud’huin M, Quillard T, Heymann D, Lamoureux F, Ory B. miRNA-193a-5p repression of p73 controls Cisplatin chemoresistance in primary bone tumors. Oncotarget. 2016; 7:54503–14. https://doi.org/10.18632/oncotarget.10950.
76. Lin Z, Song D, Wei H, Yang X, Liu T, Yan W, Xiao J. TGF-beta1-induced miR-202 mediates drug resistance by inhibiting apoptosis in human osteosarcoma. J Cancer Res Clin Oncol. 2016; 142:239–46. https://doi.org/10.1007/s00432-015-2028-9.
77. Wang WQ, Zhang H, Wang HB, Sun YG, Peng ZH, Zhou G, Yang SM, Wang RQ, Fang DC. Programmed cell death 4 (PDCD4) enhances the sensitivity of gastric cancer cells to TRAIL-induced apoptosis by inhibiting the PI3K/Akt signaling pathway. Mol Diagn Ther. 2010; 14:155–61. https://doi.org/10.2165/11536340-000000000-00000.
78. Diaz-Montero CM, Wygant JN, McIntyre BW. PI3-K/Akt-mediated anoikis resistance of human osteosarcoma cells requires Src activation. Eur J Cancer. 2006; 42:1491–500. https://doi.org/10.1016/j.ejca.2006.03.007.
79. Cenni V, Maraldi NM, Ruggeri A, Secchiero P, Del Coco R, De Pol A, Cocco L, Marmiroli S. Sensitization of multidrug resistant human ostesarcoma cells to Apo2 Ligand/TRAIL-induced apoptosis by inhibition of the Akt/PKB kinase. Int J Oncol. 2004; 25:1599–608.
80. Inoue R, Matsuki NA, Jing G, Kanematsu T, Abe K, Hirata M. The inhibitory effect of alendronate, a nitrogen-containing bisphosphonate on the PI3K-Akt-NFkappaB pathway in osteosarcoma cells. Br J Pharmacol. 2005; 146:633–41. https://doi.org/10.1038/sj.bjp.0706373.
81. Wang H, Luo QF, Peng AF, Long XH, Wang TF, Liu ZL, Zhang GM, Zhou RP, Gao S, Zhou Y, Chen WZ. Positive feedback regulation between Akt phosphorylation and fatty acid synthase expression in osteosarcoma. Int J Mol Med. 2014; 33:633–9. https://doi.org/10.3892/ijmm.2013.1602.
82. Zhao G, Cai C, Yang T, Qiu X, Liao B, Li W, Ji Z, Zhao J, Zhao H, Guo M, Ma Q, Xiao C, Fan Q, et al. MicroRNA-221 induces cell survival and cisplatin resistance through PI3K/Akt pathway in human osteosarcoma. PLoS One. 2013; 8:e53906. https://doi.org/10.1371/journal.pone.0053906.
83. Zhao WG, Yu SN, Lu ZH, Ma YH, Gu YM, Chen J. The miR-217 microRNA functions as a potential tumor suppressor in pancreatic ductal adenocarcinoma by targeting KRAS. Carcinogenesis. 2010; 31:1726–33. https://doi.org/10.1093/carcin/bgq160.
84. Guo J, Feng Z, Huang Z, Wang H, Lu W. MicroRNA-217 functions as a tumour suppressor gene and correlates with cell resistance to cisplatin in lung cancer. Mol Cells. 2014; 37:664–71. https://doi.org/10.14348/molcells.2014.0121.
85. Wang H, Dong X, Gu X, Qin R, Jia H, Gao J. The MicroRNA-217 Functions as a Potential Tumor Suppressor in Gastric Cancer by Targeting GPC5. PLoS One. 2015; 10:e0125474. https://doi.org/10.1371/journal.pone.0125474.
86. Zhang X, Guo Q, Chen J, Chen Z. Quercetin Enhances Cisplatin Sensitivity of Human Osteosarcoma Cells by Modulating microRNA-217-KRAS Axis. Mol Cells. 2015; 38:638–42. https://doi.org/10.14348/molcells.2015.0037.
87. Zhu Z, Wang CP, Zhang YF, Nie L. MicroRNA-100 resensitizes resistant chondrosarcoma cells to cisplatin through direct targeting of mTOR. Asian Pac J Cancer Prev. 2014; 15:917–23.
88. Wu J, Wu X, Zhong D, Zhai W, Ding Z, Zhou Y. Short Hairpin RNA (shRNA) Ether a go-go 1 (Eag1) inhibition of human osteosarcoma angiogenesis via VEGF/PI3K/AKT signaling. Int J Mol Sci. 2012; 13:12573–83. https://doi.org/10.3390/ijms131012573.
89. Shao XJ, Miao MH, Xue J, Xue J, Ji XQ, Zhu H. The Down-Regulation of MicroRNA-497 Contributes to Cell Growth and Cisplatin Resistance Through PI3K/Akt Pathway in Osteosarcoma. Cell Physiol Biochem. 2015; 36:2051–62. https://doi.org/10.1159/000430172.
90. Gougelet A, Pissaloux D, Besse A, Perez J, Duc A, Dutour A, Blay JY, Alberti L. Micro-RNA profiles in osteosarcoma as a predictive tool for ifosfamide response. Int J Cancer. 2011; 129:680–90. https://doi.org/10.1002/ijc.25715.
91. Zhou Y, Huang Z, Wu S, Zang X, Liu M, Shi J. miR-33a is up-regulated in chemoresistant osteosarcoma and promotes osteosarcoma cell resistance to cisplatin by down-regulating TWIST. J Exp Clin Cancer Res. 2014; 33:12. https://doi.org/10.1186/1756-9966-33-12.
92. Wu J, Liao Q, He H, Zhong D, Yin K. TWIST interacts with beta-catenin signaling on osteosarcoma cell survival against cisplatin. Mol Carcinog. 2014; 53:440–6. https://doi.org/10.1002/mc.21991.
93. Xu M, Jin H, Xu CX, Bi WZ, Wang Y. MiR-34c inhibits osteosarcoma metastasis and chemoresistance. Med Oncol. 2014; 31:972. https://doi.org/10.1007/s12032-014-0972-x.
94. Hsu KW, Hsieh RH, Huang KH, Fen-Yau Li A, Chi CW, Wang TY, Tseng MJ, Wu KJ, Yeh TS. Activation of the Notch1/STAT3/Twist signaling axis promotes gastric cancer progression. Carcinogenesis. 2012; 33:1459–67. https://doi.org/10.1093/carcin/bgs165.
95. Devy L, Huang L, Naa L, Yanamandra N, Pieters H, Frans N, Chang E, Tao Q, Vanhove M, Lejeune A, van Gool R, Sexton DJ, Kuang G, et al. Selective inhibition of matrix metalloproteinase-14 blocks tumor growth, invasion, and angiogenesis. Cancer Res. 2009; 69:1517–26. https://doi.org/10.1158/0008-5472.CAN-08-3255.
96. Mu X, Isaac C, Greco N, Huard J, Weiss K. Notch Signaling is Associated with ALDH Activity and an Aggressive Metastatic Phenotype in Murine Osteosarcoma Cells. Front Oncol. 2013; 3:143. https://doi.org/10.3389/fonc.2013.00143.
97. Xu E, Zhao J, Ma J, Wang C, Zhang C, Jiang H, Cheng J, Gao R, Zhou X. miR-146b-5p promotes invasion and metastasis contributing to chemoresistance in osteosarcoma by targeting zinc and ring finger 3. Oncol Rep. 2016; 35:275–83. https://doi.org/10.3892/or.2015.4393.
98. Clarke MF, Dick JE, Dirks PB, Eaves CJ, Jamieson CH, Jones DL, Visvader J, Weissman IL, Wahl GM. Cancer stem cells--perspectives on current status and future directions: AACR Workshop on cancer stem cells. Cancer Res. 2006; 66:9339–44. https://doi.org/10.1158/0008-5472.CAN-06-3126.
99. Dean M, Fojo T, Bates S. Tumour stem cells and drug resistance. Nat Rev Cancer. 2005; 5:275–84. https://doi.org/10.1038/nrc1590.
100. Di Fiore R, Santulli A, Ferrante RD, Giuliano M, De Blasio A, Messina C, Pirozzi G, Tirino V, Tesoriere G, Vento R. Identification and expansion of human osteosarcoma-cancer-stem cells by long-term 3-aminobenzamide treatment. J Cell Physiol. 2009; 219:301–13. https://doi.org/10.1002/jcp.21667.
101. Yu Z, Pestell TG, Lisanti MP, Pestell RG. Cancer stem cells. Int J Biochem Cell Biol. 2012; 44:2144–51. https://doi.org/ 10.1016/j.biocel.2012.08.022.
102. Gao Y, Feng Y, Shen JK, Lin M, Choy E, Cote GM, Harmon DC, Mankin HJ, Hornicek FJ, Duan Z. CD44 is a direct target of miR-199a-3p and contributes to aggressive progression in osteosarcoma. Sci Rep. 2015; 5:11365. https://doi.org/10.1038/srep11365.
103. Zhou J, Wu S, Chen Y, Zhao J, Zhang K, Wang J, Chen S. microRNA-143 is associated with the survival of ALDH1+CD133+ osteosarcoma cells and the chemoresistance of osteosarcoma. Exp Biol Med (Maywood). 2015; 240:867–75. https://doi.org/10.1177/1535370214563893.
104. Li S, Sun W, Wang H, Zuo D, Hua Y, Cai Z. Research progress on the multidrug resistance mechanisms of osteosarcoma chemotherapy and reversal. Tumour Biol. 2015; 36:1329–38. https://doi.org/10.1007/s13277-015-3181-0.
105. Di Fiore R, Drago-Ferrante R, Pentimalli F, Di Marzo D, Forte IM, D’Anneo A, Carlisi D, De Blasio A, Giuliano M, Tesoriere G, Giordano A, Vento R. MicroRNA-29b-1 impairs in vitro cell proliferation, selfrenewal and chemoresistance of human osteosarcoma 3AB-OS cancer stem cells. Int J Oncol. 2014; 45:2013–23. https://doi.org/10.3892/ijo.2014.2618.
106. Gibbings D, Mostowy S, Jay F, Schwab Y, Cossart P, Voinnet O. Selective autophagy degrades DICER and AGO2 and regulates miRNA activity. Nat Cell Biol. 2012; 14:1314–21. https://doi.org/10.1038/ncb2611.
107. Yang ZJ, Chee CE, Huang S, Sinicrope FA. The role of autophagy in cancer: therapeutic implications. Mol Cancer Ther. 2011; 10:1533–41. https://doi.org/10.1158/1535-7163.MCT-11-0047.
108. Maes H, Rubio N, Garg AD, Agostinis P. Autophagy: shaping the tumor microenvironment and therapeutic response. Trends Mol Med. 2013; 19:428–46. https://doi.org/10.1016/j.molmed.2013.04.005.
109. Sui X, Chen R, Wang Z, Huang Z, Kong N, Zhang M, Han W, Lou F, Yang J, Zhang Q, Wang X, He C, Pan H. Autophagy and chemotherapy resistance: a promising therapeutic target for cancer treatment. Cell Death Dis. 2013; 4:e838. https://doi.org/10.1038/cddis.2013.350.
110. Wang C, Hu Q, Shen HM. Pharmacological inhibitors of autophagy as novel cancer therapeutic agents. Pharmacol Res. 2016; 105:164–75. https://doi.org/10.1016/j.phrs.2016.01.028.
111. Frankel LB, Lund AH. MicroRNA regulation of autophagy. Carcinogenesis. 2012; 33:2018–25. https://doi.org/10.1093/carcin/bgs266.
112. Bader AG, Brown D, Stoudemire J, Lammers P. Developing therapeutic microRNAs for cancer. Gene Ther. 2011; 18:1121–6. https://doi.org/10.1038/gt.2011.79.
113. Thorsen SB, Obad S, Jensen NF, Stenvang J, Kauppinen S. The therapeutic potential of microRNAs in cancer. Cancer J. 2012; 18:275–84. https://doi.org/10.1097/PPO.0b013e318258b5d6.
114. Li Y, Jiang W, Hu Y, Da Z, Zeng C, Tu M, Deng Z, Xiao W. MicroRNA-199a-5p inhibits cisplatin-induced drug resistance via inhibition of autophagy in osteosarcoma cells. Oncol Lett. 2016; 12:4203–8. https://doi.org/10.3892/ol.2016.5172.
115. Xu R, Liu S, Chen H, Lao L. MicroRNA-30a downregulation contributes to chemoresistance of osteosarcoma cells through activating Beclin-1-mediated autophagy. Oncol Rep. 2016; 35:1757–63. https://doi.org/10.3892/or.2015.4497.
116. Zou Z, Wu L, Ding H, Wang Y, Zhang Y, Chen X, Chen X, Zhang CY, Zhang Q, Zen K. MicroRNA-30a sensitizes tumor cells to cis-platinum via suppressing beclin 1-mediated autophagy. J Biol Chem. 2012; 287:4148–56. https://doi.org/10.1074/jbc.M111.307405.
117. Guo S, Bai R, Liu W, Zhao A, Zhao Z, Wang Y, Wang Y, Zhao W, Wang W. miR-22 inhibits osteosarcoma cell proliferation and migration by targeting HMGB1 and inhibiting HMGB1-mediated autophagy. Tumour Biol. 2014; 35:7025–34. https://doi.org/10.1007/s13277-014-1965-2.
118. Li X, Wang S, Chen Y, Liu G, Yang X. miR-22 targets the 3′ UTR of HMGB1 and inhibits the HMGB1-associated autophagy in osteosarcoma cells during chemotherapy. Tumour Biol. 2014; 35:6021–8. https://doi.org/10.1007/s13277-014-1797-0.
119. Chang Z, Huo L, Li K, Wu Y, Hu Z. Blocked autophagy by miR-101 enhances osteosarcoma cell chemosensitivity in vitro. ScientificWorldJournal. 2014; 2014:794756. https://doi.org/10.1155/2014/794756.
120. Brannan CI, Dees EC, Ingram RS, Tilghman SM. The product of the H19 gene may function as an RNA. Mol Cell Biol. 1990; 10:28–36.
121. Hung T, Chang HY. Long noncoding RNA in genome regulation: prospects and mechanisms. RNA Biol. 2010; 7:582–5.
122. Wapinski O, Chang HY. Long noncoding RNAs and human disease. Trends Cell Biol. 2011; 21:354–61. https://doi.org/10.1016/j.tcb.2011.04.001.
123. Gong C, Maquat LE. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature. 2011; 470:284–8. https://doi.org/10.1038/nature09701.
124. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009; 10:155–9. https://doi.org/10.1038/nrg2521.
125. Kotake Y, Nakagawa T, Kitagawa K, Suzuki S, Liu N, Kitagawa M, Xiong Y. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene. 2011; 30:1956–62. https://doi.org/10.1038/onc.2010.568.
126. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010; 464:1071–6. https://doi.org/10.1038/nature08975.
127. Amaral PP, Mattick JS. Noncoding RNA in development. Mamm Genome. 2008; 19:454–92. https://doi.org/10.1007/s00335-008-9136-7.
128. Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012; 22:1775–89. https://doi.org/10.1101/gr.132159.111.
129. Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011; 25:1915–27. https://doi.org/10.1101/gad.17446611.
130. Bartonicek N, Maag JL, Dinger ME. Long noncoding RNAs in cancer: mechanisms of action and technological advancements. Mol Cancer. 2016; 15:43. https://doi.org/10.1186/s12943-016-0530-6.
131. Zhu KP, Zhang CL, Shen GQ, Zhu ZS. Long noncoding RNA expression profiles of the doxorubicin-resistant human osteosarcoma cell line MG63/DXR and its parental cell line MG63 as ascertained by microarray analysis. Int J Clin Exp Pathol. 2015; 8:8754–73.
132. Dean M, Rzhetsky A, Allikmets R. The human ATP-binding cassette (ABC) transporter superfamily. Genome Res. 2001; 11:1156–66. https://doi.org/10.1101/gr.184901.
133. Holohan C, Van Schaeybroeck S, Longley DB, Johnston PG. Cancer drug resistance: an evolving paradigm. Nat Rev Cancer. 2013; 13:714–26. https://doi.org/10.1038/nrc3599.
134. Cascorbi I. Role of pharmacogenetics of ATP-binding cassette transporters in the pharmacokinetics of drugs. Pharmacol Ther. 2006; 112:457–73. https://doi.org/10.1016/j.pharmthera.2006.04.009.
135. Abolhoda A, Wilson AE, Ross H, Danenberg PV, Burt M, Scotto KW. Rapid activation of MDR1 gene expression in human metastatic sarcoma after in vivo exposure to doxorubicin. Clin Cancer Res. 1999; 5:3352–6.
136. Wu Q, Yang Z, Nie Y, Shi Y, Fan D. Multi-drug resistance in cancer chemotherapeutics: mechanisms and lab approaches. Cancer Lett. 2014; 347:159–66. https://doi.org/10.1016/j.canlet.2014.03.013.
137. Zhang CL, Zhu KP, Shen GQ, Zhu ZS. A long non-coding RNA contributes to doxorubicin resistance of osteosarcoma. Tumour Biol. 2016; 37:2737–48. https://doi.org/10.1007/s13277-015-4130-7.
138. Dieudonne FX, Marion A, Marie PJ, Modrowski D. Targeted inhibition of T-cell factor activity promotes syndecan-2 expression and sensitization to doxorubicin in osteosarcoma cells and bone tumors in mice. J Bone Miner Res. 2012; 27:2118–29. https://doi.org/10.1002/jbmr.1650.
139. Li Z, Zhao L, Wang Q. Overexpression of long non-coding RNA HOTTIP increases chemoresistance of osteosarcoma cell by activating the Wnt/beta-catenin pathway. Am J Transl Res. 2016; 8:2385–93.
140. van Rooij E, Purcell AL, Levin AA. Developing microRNA therapeutics. Circ Res. 2012; 110:496–507. https://doi.org/10.1161/CIRCRESAHA.111.247916.
141. Kim M, Kasinski AL, Slack FJ. MicroRNA therapeutics in preclinical cancer models. Lancet Oncol. 2011; 12:319–21. https://doi.org/10.1016/S1470-2045(11)70067-5.
142. Bader AG, Brown D, Winkler M. The promise of microRNA replacement therapy. Cancer Res. 2010; 70:7027–30. https://doi.org/10.1158/0008-5472.CAN-10-2010.
143. Hong L, Han Y, Zhang Y, Zhang H, Zhao Q, Wu K, Fan D. MicroRNA-21: a therapeutic target for reversing drug resistance in cancer. Expert Opin Ther Targets. 2013; 17:1073–80. https://doi.org/10.1517/14728222.2013.819853.
144. Tacar O, Sriamornsak P, Dass CR. Doxorubicin: an update on anticancer molecular action, toxicity and novel drug delivery systems. J Pharm Pharmacol. 2013; 65:157–70. https://doi.org/10.1111/j.2042-7158.2012.01567.x.
145. Zhou C, Tan W, Lv H, Gao F, Sun J. Hypoxia-inducible microRNA-488 regulates apoptosis by targeting Bim in osteosarcoma. Cell Oncol (Dordr). 2016; 39:463–71. https://doi.org/10.1007/s13402-016-0288-2.
146. Lin BC, Huang D, Yu CQ, Mou Y, Liu YH, Zhang DW, Shi FJ. MicroRNA-184 Modulates Doxorubicin Resistance in Osteosarcoma Cells by Targeting BCL2L1. Med Sci Monit. 2016; 22:1761–5.
147. Marino MT, Grilli A, Baricordi C, Manara MC, Ventura S, Pinca RS, Bellenghi M, Calvaruso M, Mattia G, Donati D, Tripodo C, Picci P, Ferrari S, et al. Prognostic significance of miR-34a in Ewing sarcoma is associated with cyclin D1 and ki-67 expression. Ann Oncol. 2014; 25:2080–6. https://doi.org/10.1093/annonc/mdu249.
148. Tang XY, Zheng W, Ding M, Guo KJ, Yuan F, Feng H, Deng B, Sun W, Hou Y, Gao L. miR-125b acts as a tumor suppressor in chondrosarcoma cells by the sensitization to doxorubicin through direct targeting the ErbB2-regulated glucose metabolism. Drug Des Devel Ther. 2016; 10:571–83. https://doi.org/10.2147/DDDT.S90530.
149. Huang K, Chen J, Yang MS, Tang YJ, Pan F. Inhibition of Src by microRNA-23b increases the cisplatin sensitivity of chondrosarcoma cells. Cancer Biomark. 2017; 18:231–9. https://doi.org/10.3233/CBM-160102.
150. Wang Z, Lei H, Sun Q. MicroRNA-141 and its associated gene FUS modulate proliferation, migration and cisplatin chemosensitivity in neuroblastoma cell lines. Oncol Rep. 2016; 35:2943–51. https://doi.org/10.3892/or.2016.4640.
151. Milhem MM, Knutson T, Yang S, Zhu D, Wang X, Leslie KK, Meng X. Correlation of MTDH/AEG-1 and HOTAIR Expression with Metastasis and Response to Treatment in Sarcoma Patients. J Cancer Sci Ther. 2011; S5.
152. Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017; 16:203–22. https://doi.org/10.1038/nrd.2016.246.
153. Christopher AF, Kaur RP, Kaur G, Kaur A, Gupta V, Bansal P. MicroRNA therapeutics: Discovering novel targets and developing specific therapy. Perspect Clin Res. 2016; 7:68–74. https://doi.org/10.4103/2229-3485.179431.
154. Zhao Y, Dai Z, Liang Y, Yin M, Ma K, He M, Ouyang H, Teng CB. Sequence-specific inhibition of microRNA via CRISPR/CRISPRi system. Sci Rep. 2014; 4:3943. https://doi.org/10.1038/srep03943.
155. Ho TT, Zhou N, Huang J, Koirala P, Xu M, Fung R, Wu F, Mo YY. Targeting non-coding RNAs with the CRISPR/Cas9 system in human cell lines. Nucleic Acids Res. 2015; 43:e17. https://doi.org/10.1093/nar/gku1198.

