Recognition for the diversity of lncRNAs and their involvement IN cancer
The first lncRNA was discovered decades ago during the characterization of X-chromosome inactivation [1]. Following that early discovery, several lncRNAs were inadvertently uncovered and characterized as anomalous molecules before the community recognized that lncRNAs represent a distinct class of regulatory RNAs. With the completion of the human genome project in 2003 and subsequent characterization the genomic landscape, attempts at bioinformatics prediction of mRNA genes was found to be cluttered with many fold higher predicted transcripts than were experimentally verified as the precursors of proteins [2-5]. What these programs revealed was a glut of predicted transcripts, genes with hallmarks of transcription but no discernable protein coding function. In addition, these were thought to have no practical biological function because they had little evolutionary conservation [6]. Initially, these unverified genes were dismissed as programming artifacts to be eliminated [7-9]. However, it was quickly realized the lack of evolutionary conservation did not rule out function [10]. It is now accepted that the human genome contains many thousands of lncRNA transcripts. Functional implications of this discovery have yet to be fully elucidated. To date lncRNAs have been detected throughout development and in every cell type tested thus far.
One field that has been particularly active in lncRNA discovery is cancer biology. Due to the pressing need for development of novel therapeutics and diagnostics, many newly emergent fields have been focused on cancer research. These include the discovery of microRNAs, targeted immunotherapy, and most recently circulating tumor cells. Added to this ever-growing list are lncRNAs. Their implication in a diverse array of regulatory roles has heightened interest in these molecules as functional players in the development and heterogeneity of cancer [11]. Recently, the pace of discovery and functional validation for lncRNAs has been increasing exponentially with the advent of sequencing technologies (Figure 1). But due to the rapid pace of discovery little headway has been made in functionally characterizing the bulk of these newly-discovered genes.
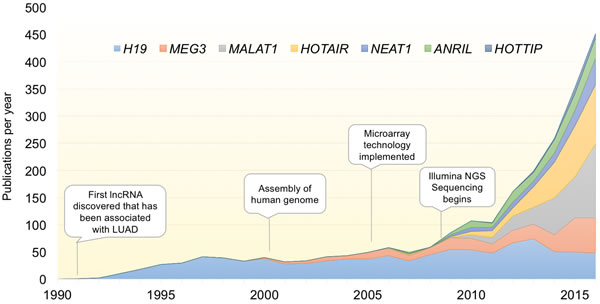
Figure 1: Exponential discovery of lncRNAs with the adoption of transcriptome-wide gene expression technologies. Graph indicates the total number of publications per year for select lncRNAs with known involvement in LUAD. With the advent of transcriptomic profiling, the pace of lncRNA discovery and papers characterizing their function has increased exponentially over the last decade. lncRNAs included are those described in Table 1.
One cancer type with particularly high mortality is lung cancer. The overall mortality rate (all stages) remains at ~85% [12], which is comparable to stage IV breast cancer. The landmark National Lung cancer Screening Trial (NLST) study identified Spiral CT as an effective detection tool that reduced overall mortality, however the study was only applicable to lifelong smokers and has a false positive rate of 92% [13]. One issue complicating the development of early detection strategies is that lung cancer is composed of several distinct subtypes, each with their own etiology and clinical outcomes. Lung cancer is loosely broken into two subcategories, small cell (SCLC, ~13% of cases) and non-small cell lung cancer (NSCLC, ~87% of cases). Lung adenocarcinoma (LUAD) is the most common subtype of NSCLC, and arises from the distal alveolar epithelium (Figure 2). This cancer has been linked to mutations in EGFR for never-smokers and mutations in kRAS for smokers [14]. In addition, dozens of other oncogenic mutations, copy number variations, and epigenetic alterations have been described in LUAD [15, 16]. Several oncogenic mutations in protein coding genes have been exploited for the development of targeted therapeutics. Notably among them are Erlotinib and Gefintinib, both EGFR inhibitors, and Crizotinib, an ALK/ROS1/MET kinase inhibitor [17-20]. While Erlotinib and Gefitinib are in use clinically, each is associated with a high rate of relapse in patients due to further molecular alterations that develop, such as the 790M mutation to EGFR, which renders the cancer resistant [21]. Therefore, there is a pressing need to both define molecular hallmarks that distinguish LUAD from other lung cancers and normal tissues, and to specifically target those cancerous cells while leaving lung function intact.
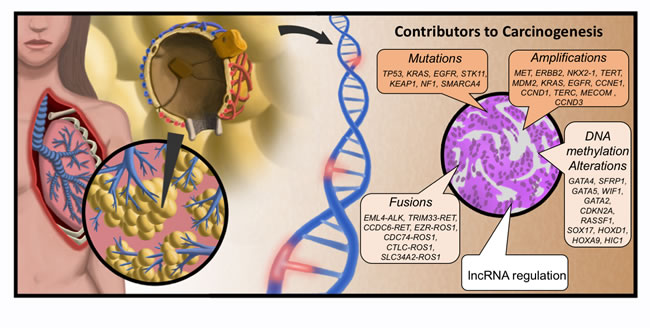
Figure 2: Molecular origins of LUAD. Lung adenocarcinoma (LUAD) arises in the distal alveolar epithelium from progenitor alveolar epithelial cells. LUAD develops from these precursor cells though oncogenic activation (and deactivation of tumor suppressors) by induced mutations to the DNA, amplification and fusion events, as well as epigenomic alterations. Genes listed were taken from TCGA analysis of LUAD (15). Added to this is the newly-emergent appreciation for altered lncRNA regulation of cellular processes as an oncogenic event.
In this review, we focus on lncRNAs with characteristics indicating they could be exploited in improved efficacy of LUAD detection, clinical management, and outcome prediction. We first outline the current state of molecular characterization for lncRNAs with known involvement in LUAD etiology. Then, we utilize multiple high-throughput analysis recently made publicly available to define a subset of high-interest candidate lncRNAs. Of these, we provide a synopsis on what is currently known about the predicted candidates. We end with discussion of ways in which knowledge of dysregulated lncRNAs in LUAD can be leveraged in the clinic.
Known lncRNAs involved in LUAD
The biological significance of lncRNAs is under intense investigation. Because lncRNAs were grouped into a broad category of any non-coding RNA longer than 200 nucleotides, this class of RNAs represents a heterogeneous group in terms of mechanism and function. lncRNAs are implicated in transcriptional regulation, cellular signaling, chromatin remodeling, splicing, and a host of other processes [22-25]. Mechanistically, lncRNAs can regulate transcriptional activity at the endogenous locus through antisense activity and in trans through the regulation of epigenetic structure [26, 27]. At the post-transcriptional level, lncRNAs regulate splicing, micro-RNA targeting, and through RNA-protein interactions, can influence their binding partner function, localization, and activity [28-30]. In terms of biological processes, lncRNAs are involved in regulation of the cell cycle, apoptosis, differentiation, and immunological response [31-34]. Despite the large repertoire of lncRNAs expressed in lung, only a handful have been functionally linked to LUAD development. Some exhibit hallmarks of tumor suppression, such as MEG3 [35], while others, such as HOTAIR, behave as oncogenes through increased proliferation and reduced survival [36]. Table 1 highlights some of the known lncRNAs involved in LUAD and their cellular mechanism of action. However, for most lncRNAs, a defined molecular mechanism has yet to be discovered.
Table 1: Known lncRNAs involved in LUAD
GENE |
Locus |
Nearby factor implicated in cancer |
Mechanism(s) of action |
Additional cancer association(s) |
|
ANRIL |
9p21 |
CDKN2B (INK4-ARF) tumor suppressor |
Oncogene. Antagonizes the CDKN2A and CDKN2B tumor suppressors via recruitment of PRC2 and PRC1 [57,58]. |
basal cell, breast, cervical, esophageal, gallbladder, gastric, liver, melanoma, ovarian. |
|
H19 |
11p15 |
IGF2 growth factor |
Oncogene. Targets multiple tumor suppressive miRNAs [29,45]; parent transcript of miRNAs involved in regulation of tight junction dynamics [44,46]. |
adrenal, bladder, cervical, colorectal, gallbladder, gastric, esophageal, laryngeal, nasopharyngeal, ovarian, pancreatic, thyroid. |
HOTAIR |
12q13 |
HOXC transcription factors |
Oncogene. Long-range epigenetic action via recruitment of PRC2 and LSD1 [65-67]; serves as a miRNA sponge to block miR-331-3p mediated destruction of HER2 transcripts [68]. |
bladder, colorectal, ER (+) breast, liver, nasopharyngeal, oral, ovarian, pancreatic, pituitary, small cell lung. |
|
HOTTIP |
7p15 |
HOXA transcription factors |
Oncogene. Regulates chromatin structure at the HOXA transcription factor locus [98]. |
colorectal, pancreatic, osteosarcoma, tongue. |
|
MALAT1 |
11q13 |
NEAT1 lncRNA |
Oncogene. Suppresses E-cadherin via suz12 recruitment, leading to metastasis [112]. |
glioma, multiple myeloma, pituitary, renal clear cell, tongue. |
|
MEG3 |
14q32 |
DLK1 growth factor receptor |
Tumor Suppressor. Long-range epigenetic action, leading to suppression of TFG β [90]; blocks oncogenic activity of miR-21 [86]. |
AML, cervical, colorectal, gastric, meningioma, ovarian, pancreatic, pituitary, prostate, thyroid. |
|
NEAT1 |
11q13 |
MALAT1 lncRNA |
Oncogene. Promotes survival when DNA damage present via paraspeckle formation [122,123]. |
colorectal, esophageal, gastric, glioma, leukemia, ovarian, prostate. |
The RefSeq gene name annotation, alongside the hg19 chromosomal location are listed. In addition, the established mechanism of action is listed, as well as other cancers where the lncRNA has demonstrated effects on tumor initiation, promotion, progression, and/or patient survival outcomes.
Pan-cancer lncRNAs
It is important to note that, while the lncRNAs in Table 1 play a role in LUAD development, they are all implicated in the development of multiple cancer types, and therefore do not confer specificity to any given cancer. Because multiple types of cancers depend on similar pathways for sustained growth, it is not surprising that a subset of lncRNAs have been linked to suppression of p53, Wnt signaling activation, epithelial to mesenchymal transition (EMT), and similar early steps in the process of oncogenesis. Here, we discuss examples of lncRNAs that not only promote LUAD but are also involved in tumorigenesis in a variety of cancers.
H19
The maternally expressed and imprinted gene H19 is elevated in numerous cancers. Overexpression occurs through the loss of epigenetic repression at the paternal allele [37-39]. More recently, H19 was found to be upregulated in NSCLC tissue and correlated with poor prognosis [40]. Many genes involved in embryonic growth and implicated in cancer lie within the H19 locus and are cis regulated by H19 [41, 42]. In addition, H19 upregulation has been linked to MYC oncogene activation [43]. Therefore, disrupted paternal imprinting on H19 acts as an oncogenic driver in several cancers, including NSCLC.
Mechanistically, H19 serves as the precursor miR-675, which is processed from the first exon of H19 and in its mature miRNA form mediates degradation of ZO-1 and E-Cadherin mRNA, disrupting tight junction formation, which in turn disrupts epithelial architecture and leads to increased invasion [44, 45]. In addition to being a precursor to miRNAs, H19 can suppress several miRNAs, including let-7 [46], miR-138, and miR-200a [29] by serving as a competing endogenous RNA (ceRNA). Suppression of both miR-138 and miR-200a via H19 was shown to re-activate expression of the mesenchymal marker genes ZEB1, ZEB2, and Vimentin, resulting in EMT progression in bladder cancer [29]. In addition, H19 was recently shown to suppress miR-141 and miR-22, both of which function as antagonists of Wnt signaling [47, 48]. This H19-mediated suppression lead to the activation of the Wnt/β-catenin pathway during osteoblast differentiation [49]. Their role in Wnt/B-catenin signaling might suggest an alternative means by which H19 can promote tumorigenesis. In addition, H19 has been linked with Wnt-mediated tumorigenesis via PRC2/EZH2 recruitment to the Wnt antagonist gene NKD1 [50]. The convergent activity of H19 on different parts of the Wnt signaling pathway is interesting from both a mechanistic and evolutionary standpoint, revealing that individual lncRNAs can perform multiple functional roles simultaneously to effect intracellular signaling cascades.
H19 has also been shown to negatively regulate p53 signaling. Ectopic expression of H19 can cause increased cell growth and decreased p53 transcriptional activity [51]. This was attributed to a physical interaction between H19 and the p53 protein. However, the mechanism by which this interaction mediates the inactivation of p53 remains ambiguous and more investigation is needed to fully evaluate the effect of H19 on p53-mediated cellular arrest and apoptosis.
ANRIL (antisense non-coding RNA in the INK4 locus)
This gene lies within the 9p21.3 gene cluster, consisting of the p14ARF, p15INK4b, and p16INK4a tumor suppressor genes. Within this locus, ANRIL is the natural antisense transcript of the p16INK4a gene. p14ARF is involved in stabilizing p53 levels by negatively regulating MDM2 [52], whereas both p15INK4b and p16INK4a are critical regulators of the cell cycle [53]. Their deactivation promotes an increase in cellular proliferation and is seen in several cancers [54, 55]. In addition, the deactivation of the 9p21.3 gene cluster often occurs in conjunction with LUAD driven by mutationally-activated kRAS [56]. The proximity of ANRIL within the gene cluster allows for a cis-mediated suppression of p16INK4a that occurs through recruitment of PRC2 complex which compacts chromatin and subsequently deactivates gene expression [57, 58]. A more recent study demonstrated that ANRIL is overexpressed in NSCLC, correlating with poor prognosis [59].
HOTAIR (HOX transcript antisense RNA)
This well-characterized lncRNA has been the sole subject of previous reviews [36, 60-63], and acts by binding and promoting chromatin compaction through association with GA-rich DNA sequence motifs that subsequently recruit PRC2 [64-66]. This results in genome-wide epigenetic regulation of differentiation and cancer development [67]. HOTAIR can also regulate miRNA by acting as a competing endogenous RNA (ceRNA) to deplete cells of miR-331-3p, enhancing expression of the HER2 receptor tyrosine kinase and thereby promoting oncogenesis [68]. Indeed, HOTAIR displays all the canonical behaviors of an oncogene, including poor prognosis when present [69, 70], chemoresistance [71], reduced overall survival [72-75], and increased metastasis [76-80]. This occurs in a number of cancers, including both SCLC [81] and NSCLC [82, 83].
MEG3 (maternally expressed gene 3)
One of many maternally imprinted lncRNAs [84], MEG3 exhibits the hallmarks of a tumor suppressor, namely inhibition of proliferation and induction of apoptosis [35, 85-87] in numerous cancers. Multiple functions for MEG3 in cancer have been described [87, 88], Locally, expression is inversely correlated with the nearby tumor suppressor DLK1, which it may regulate [89]. MEG3 can also act throughout the genome as an epigenomic regulator of TGFβ-responsive distal regulatory elements. It does so by forming RNA:DNA triplex helix structures at GA-rich sequence recognition sites, which bring EZH2 to target loci, effectively condensing local chromatin regions to disrupt enhancer activity and block TGFβ-induced proliferation [90]. Additionally, MEG3 can mediate the destruction of miR-21, blocking this microRNAs oncogenic potential [86].
It is interesting to note that both HOTAIR and MEG3 bind GA-rich sequence elements that facilitate recruitment of PRC2 complex and condense the local chromatin environment, yet HOTAIR functions as an oncogene while MEG3 functions as a tumor suppressor. Considering that HOTAIR oncogenic activity is seen in multiple cancers and MEG3 tumor suppressor activity is also observed across cancers, the simple explanation of differing mechanisms in differing tumors does not seem applicable. Instead, follow up studies on the genomic distribution of the two lncRNAs, their relative expression to each other, and any mechanistic interactions they may have are warranted to address this question.
HOTTIP (HOXA transcript at the distal TIP)
HOTTIP is another lncRNA transcribed from the HOXA locus which exhibits the oncogenic properties of increased proliferation, expression in advanced pathological stages alongside distant metastasis, inhibition of apoptosis, and association with overall poor prognosis in multiple cancers [91-96]. While there are observed correlations between HOTTIP and vitamin D receptor signaling [97] as well as p21 silencing [91], the main role of HOTTIP described in cancer progression is its ability to utilize three-dimensional chromatin looping structures. These allow HOTTIP to regulate cis-members of the HOXA cluster by recruitment of WDR5 to drive H3K4me3 deposition into chromatin, activating target gene expression [98].
The MEN (MALAT1-NEAT1) locus
The MEN locus is located on chromosome 11 at p13.1 and harbors both the MALAT1 and NEAT1 lncRNA genes. NEAT1 is about 53kb upstream of the 5’ end of MALAT1, and both transcripts are deregulated in LUAD [99, 100].
MALAT1 (metastasis-associated lung adenocarcinoma transcript-1; multiple endocrine neoplasia-alpha)
This is a single exon gene originally identified as expressed specifically in lung cancer. Because it is associated with poor prognosis and distant metastasis in NSCLC [101-103], along with other cancers [102, 104-107], much of the emphasis in studying this gene is to utilize it as a prognostic biomarker [108, 109]. Identified over two decades ago, this functional RNA has been the sole subject of previous reviews [110, 111] and was initially implicated in RNA splicing through extensive studies in vitro.
Functionally, MALAT1 interacts with Suz12 resulting in decreased expression of E-cadherin, a cell adhesion molecule essential in maintaining epithelial architecture. The loss of E-cadherin is a commonly observed phenomenon in cancers of epithelial origin, and co-occurs with upregulation of N-cadherin and fibronectin. This ultimately leads to metastasis, as reported in bladder cancer [112]. In addition, knockout studies of MALAT1 have demonstrated that in lung cancer, MALAT1 can also directly regulate the expression of pro-metastatic genes [99]. These observations tie together the poor prognosis and increased metastatic behaviors observed when MALAT1 is over expressed in tumors.
NEAT1 (nuclear paraspeckle assembly transcript 1; Nuclear Enriched Abundant Transcript-1)
This is another single exon gene that is transcribed from the same locus as MALAT1. It exhibits many similar characteristics with MALAT1, including tumor recurrence [113], poor prognosis [114], and metastasis [115, 116]. Of special note, the close proximity of NEAT1 to MALAT1, and their similar roles as oncogenes in multiple cancers suggests that the entire locus may be subject to aberrant regulation in cancer [117]. Indeed, several studies have demonstrated a correlation between MALAT1 and NEAT1 expression [115]. While NEAT1 acts as part of chromatin remodeling complexes [118], less is understood about a direct functional role for NEAT1 in carcinogenesis. NEAT1 serves as a scaffold for nuclear paraspeckle formation [119-121], which accumulate in response to DNA-damaged induced genotoxic stress [122, 123]. It is possible that NEAT1 acts to promote carcinogenesis directly by abrogating the stressors placed on the genomes of cancer cells. Further research is needed to determine if NEAT1 plays a direct role in oncogenesis, or if instead the MALAT1/NEAT1 locus is under mutual regulation, with NEAT1 upregulation in cancer being a byproduct of its genomic proximity to MALAT1.
Unclear mechanism(s) affecting cancer progression
Highlighting the need for further research, there are still several lncRNAs which are deregulated across cancer types, yet have not undergone in-depth functional characterization. This lack of mechanistic understanding hinders further investigation into the application of targeted therapeutics toward these deregulated lncRNAs, for concerns regarding off-target effects. One such transcript is CCAT2. It was originally identified as having LUAD-specific expression [124], but is now implicated in a host of cancers [125, 126] and associated with smoking [127]. It contains rs6983267, a single nucleotide polymorphism (SNP) identified through genome-wide association studies (GWAS) as conferring an increased risk of prostate and colorectal cancer [128-130]. This gene lies within the 8q24 ‘gene desert’ hotspot that is home to the MYC oncogene and is associated with numerous cancers [131-134], highlighting the significance of lncRNAs in genetic predisposition to cancer. However, rs6983267 has not been associated with LUAD risk in the numerous LUAD GWAS studies performed to date. Instead, it appears that copy number alterations in 8q24 occur frequently in lung cancers, suggesting an alternate mechanism other than SNP regulation of the CCAT2 transcript in lung cancer pathogenesis. Mechanistically, CCAT2 can alter cancer metabolism depending on the allele transcribed through altered binding affinity to pre-mRNA cleavage (CFIm) splicing factors [135]. However, little is known regarding how this altered splicing affects other cellular processes, or whether the differing alleles of CCAT2 target the splicing complex to different chromatin locations to affect cancer development. This is but one example of the many lncRNAs that have been identified as deregulated in LUAD. Below, we highlight recent methods that have taken a more systematic approach to identifying the extent of lncRNA deregulation in LUAD, and what, if anything, is known about these genes.
Genome-wide analysis of lncRNAs in LUAD
Original attempts to characterize the lncRNA landscape in LUAD were performed using microarray technology. While these arrays were designed to target mRNAs, many unaccounted-for exons that were later classified as lncRNAs were included in several platforms, notably the Affymetrix Human Exon 1.0 ST Array. Illumina-developed arrays contained less information on lncRNAs due to their design emphasis on 3’UTR targeting, however several lncRNAs were included under the ‘LOC_” definition. Using this probe-based approach several studies were able to identify lncRNA expression profiles in lung cancer and perform preliminary analysis [136-139], the results of which have been conveniently collated by lnc2cancer [140]. However, the discernable drawbacks of such techniques include the lack of discovery and low-expression levels of lncRNAs, thwarting detection efforts. With the advent and widespread adoption of RNA sequencing technology, the ability to detect novel transcripts had increased exponentially. Indeed, the rate at which the non-coding RNA transcriptome expanded has rapidly outpaced the identification of mRNA genes over the last five years.
Adding to the discovery landscape was The Cancer Genome Atlas (TCGA). LUAD samples from TCGA [15] underwent whole section RNA-seq analysis using Illumina TruSeq technology. This allows for polyA selection to minimize genomic contamination; however, it eliminates any non-polyadenyated signal from the final sequence alignments, therefore expression of only the poly adenylated lncRNAs were captured with this method. While the main purpose of RNA-seq analysis performed by TCGA was to quantify mRNA expression, several groups have utilized this dataset for dual interrogation of lncRNA transcriptome changes. LncRNAtor [141], MiTranscriptome [142], and The Atlas of Noncoding RNA In Cancer [TANRIC] [143] have each performed re-analysis of RNA-seq data from TCGA data to detect lncRNAs, with differing results based on the reference genomes utilized, filtering criteria, lncRNA references databases, and incorporation of secondary data sets. Their differing results highlight how alternate bioinformatic approaches can vastly affect the results of an analysis.
LncRNAtor showcases a re-analysis of several NGS datasets. They constructed a reference lncRNA library that included sequence from the EMSEMBL, lncRNAdb, HGNC and MBI datasets. They then analyzed each transcript for phylogenetic conservation and filtered transcripts against protein coding potential to arrive at a consensus reference set. Against this pipeline they re-aligned over 200 large-scale NGS datasets from 23 different cancers, including from GEO, ENCODE, and TCGA. Of these, only TCGA LUAD and LUSC dataset were specific for lung cancers. Their reanalysis identified 860 lncRNAs significantly differentially expressed between LUAD tumors and non-paired normal lung (adjusted p-value <0.01). However, many of the transcripts annotated as lncRNAs included known protein coding genes, such as MMP12 and UHRF1, suggesting that inadequate filtering for coding potential may have inadvertently included protein coding genes. By comparison, lncRNAtor computed that 15,331 mRNAs were differentially expressed between LUAD tumors and normal tissue. The vast difference in the magnitude of changes (10-fold greater number of differentially expressed mRNAs) indicates that their analysis showed more variability in mRNAs then in lncRNAs. However, follow up work will be needed to determine if this is a reproducible phenomenon across cancers and sample sets.
MiTranscriptome was a re-analysis of data generated by TCGA, the Michigan Center for Translational Pathology, and ENCODE, developed to discover novel lncRNAs involved in cancer. Utilizing these sample sets, 58,674 lncRNA genes were identified, 1,150 of which were differentially expressed in LUAD (and obtainable through their portal linked to the UCSC genome browser). While their dataset has been made accessible via the BETA portal, as of July 2016 it does not contain the entire statistical analysis presented in their paper.
TANRIC contains the systematic re-analysis of TCGA data from over 20 cancer types, one of which was LUAD. Included in their analysis were 488 LUAD tumors and 58 unmatched normal lung sections from TCGA, along with re-analysis of RNA-seq data derived from the SEO study [16] of Korean LUAD cancer patients (83 LUAD and 77 matched normal samples). However, TANRIC focused on tumor heterogeneity and correlation with clinical covariants rather than simple tumor/normal comparisons, which can be influenced by issues such as tumor purity, degree of necrosis and other confounders.
Another study by the Mather group took this analysis a step further [144]. After initially performing tumor-normal differential analysis and finding 592 altered lncRNAs, they then subtracted differentially expressed lncRNAs in multiple cancers to arrive at a lung cancer specific deregulated lncRNA class (including LUAD and LUSC). Their analysis also included re-analysis of the SEO dataset and a smaller subset of TCGA LUAD samples as performed by TANRIC (55 LUAD tumors and matched normals, with another 243 unmatched tumors). For the purposes of this review, the differentially expressed lncRNAs released by miTranscriptome, the Mather group, and microarray data [136] were compared (Figure 3). In this way, we referenced the newest technologies as well as compared them to the older microarray data, to provide a more refined review of the current state of LUAD-associated lncRNA discovery. Forty transcripts were differentially expressed between tumor and normal in all three data sets. We have segregated these forty differentially-expressed lncRNAs into stand-alone genes, heretofore labeled as intergenic lncRNAs (Table 2), and those that occur antisense to a protein-coding gene (Table 3). We have highlighted for further discussion several of these with a high degree of validation and prior mechanistic work, but all are potential candidate driver genes for LUAD.
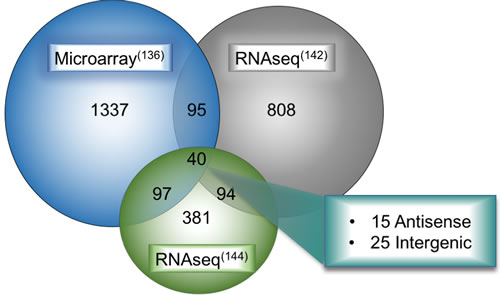
Figure 3: Overlap of deregulated lncRNAs in LUAD between multiple large-scale bioinformatic studies. Results from microarray analysis, de novo RNA-seq transcriptome assembly of TCGA LUAD datasets from the Maher study on LCALs (supplementary data file 5 from their study which includes LUAD-specific lncRNAs), and robust statistical analysis of multiple lung cancer datasets were overlapped in the hg19 UCSC genome browser to determine a unifying set of lncRNAs deregulated in all three studies. These forty deregulated lncRNAs fall broadly into two categories, 15 were antisense transcripts and the remaining 25 were intergenic genes.
Table 2: Differentially expressed intergenic lncRNAs
Level of Validation |
Genecode ID |
Alternate ID |
Position (GRCh37/hg19) |
Known Genes tsl1 |
LINC00880 |
NR_034007 |
chr3: 156799455-156840793 |
MIR3945HG |
TCONS_00008359 |
chr4: 185748077-185776905 |
|
LUCAT1 |
TCONS_00010402 |
chr5: 90597873-90621000 |
|
PVT1 |
ENST00000504719 |
chr8: 128806778-129113503 |
|
SFTA1P |
TCONS_00018434 |
chr10: 10825866-10837007 |
|
LINC00460 |
ENST00000439790 |
chr13: 107028910-107030941 |
|
LINC00197 |
TCONS_00023799 |
chr15: 95752084-96051089 |
|
MSN |
TCONS_00016990 |
chrX: 64808260-64845760 |
|
Validated Genes tsl2-tsl3 |
MIR4435-2HG |
TCONS_l2_00015916 |
chr2: 112186885-112268567 |
CTB-43E15.1 |
ENST00000523242 |
chr5: 173069521-173085197 |
|
HCG15 |
TCONS_l2_00024132 |
chr6: 28953517-28959134 |
|
LINC00968 |
ENST00000522511 |
chr8: 57401656-57472382 |
|
LINC01290 |
ENST00000566787 |
chr16: 10608698-10622059 |
|
LINC00511 |
ENST00000579631 |
chr17: 70319263-70599647 |
|
RP11-353N14.2 |
TCONS_l2_00011006 |
chr17: 77796264-77801616 |
|
LINC00665 |
ENST00000586345 |
chr19: 36795481-36822667 |
|
LINC00478 |
ENST00000428669 |
chr21: 17553910-18013444 |
|
Predicted Genes tsl4-tsl5 |
RP11-815M8.1 |
TCONS_00000738 |
chr1: 222054322-222158306 |
LINC00152 |
ENST00000331944 |
chr2: 87754947-87821037 |
|
RP11-83M16.6 |
ENST00000510621 |
chr5: 66995256-67198428 |
|
RP11-359M6.1 |
ENST00000548359 |
chr12: 79933982-79944315 |
|
RP11-1008C21.2 |
TCONS_00023630 |
chr15: 38360990-38365188 |
|
Predicted Genes tslNA |
TCONS_00007953 |
chr4: 1546984-1555291 |
|
TCONS_00025436 |
chr17: 53690342-53725799 |
||
TCONS_00029745 |
chr22: 50981205-50983413 |
LncRNAs that were deregulated in three studies (from Figure 3) that also occupy chromosomal regions in between mRNAs. LncRNAs were segregated by their Gencode transcript confidence level (tsl1 = highest quality, full transcript is validated; tsl2-3, one or many spliced ESTs are validated; tsl4-5 = one or none ESTs support the validity of the transcript, and those ESTs are suspect.) Those without a tsl ranking do not have a representative transcript in Gencode. All coordinates span the entire transcript length and are hg19 genome-based. The MSN transcript overlaps in the same orientation the MSN mRNA, therefore there is a high probability that this is not truly a lncRNA (and thus is greyed out in figure).
Table 3: Differentially expressed anti-sense lncRNAs
Genecode |
Alternate IDs |
Anti-Sense Gene |
Multi-exonic |
Position (GRCh37/hg19) |
LOC101928370 |
RP4-575N6.1 |
S1PR1 |
YES |
chr1: 101701238-101702084 |
LINC00883 |
DUBR |
LINC00882 |
YES |
chr3: 106959538-107045811 |
LINC00312 |
LINC00312 |
LMCD1 |
NO |
chr3: 8613467-8634810 |
LHFPL3-AS2 |
RP11-325F22.5 |
LHFPL3 |
YES |
chr7: 104558006-104567077 |
FEZF1-AS1 |
FEZF1-AS1 |
FEZF1 |
YES |
chr7: 121945003-121945871 |
HSPC324 |
RP11-251M1.1 |
EGFL7 |
YES |
chr9: 139541826-139554873 |
LOC105369340 |
RP11-783K16.5 |
PPP1R14B |
YES |
chr11: 64014525-64015649 |
LOC101929340 |
RP11-677M14.3 |
ESAM |
YES |
chr11: 124632326-124635257 |
SBK1-AS1 |
RP11-57A19.2 |
SBK1 |
YES |
chr16: 28270020-28303385 |
FENDRR |
FENDRR |
FOXF1 |
YES |
chr16: 86508050-86542705 |
TBX2-AS1 |
RP11-332H18.5 |
TBX2 |
YES |
chr17: 59470732-59477096 |
RP11-720L2.4 |
COLEC2 |
YES |
chr18: 314886-319165 |
|
GATA6-AS1 |
GATA6-AS1 |
GATA6 |
YES |
chr18: 19746858-19748929 |
LINC01271 |
RP11-290F20.2 |
LINC01270 |
YES |
chr20: 48909256-48937879 |
LINC00649 |
LINC00649 |
ATP5O |
YES |
chr21: 35295736-35351160 |
The Gencode annotation for each lncRNA is indicated, along with the genomic coordinates of the lncRNA and the mRNA that is transcribed in the antisense orientation. All but LINC00312 are multi-exonic, indicative of splicing. The hg19-based lncRNA coordinates are listed.
Intergenic discovery-based lncRNAs and their functional implications
Intergenic lncRNAs are herein defined as those lncRNA genes located in the space between protein coding genes. Prior evidence has shown that lncRNAs occurring in cis with protein coding genes can loop back and affect the nearby mRNA [98]. However, for intergenic lncRNA genes, their distal location makes functional predictions difficult. Instead, intergenic lncRNA can mediate their function in trans through a variety of mechanisms, such as their involvement with chromatin remodeling complexes [145]. Here, we focus on four lncRNAs that came from our review of LUAD transcriptomic profiling (Table 2).
LUCAT1: (lung cancer associated transcript-1, also known as smoke and cancer-associated lncRNA-1, SCAL1)
is a multi-exonic lncRNA located deep within the gene desert of chromosome 5q14. This lncRNA is upregulated by cigarette smoke in vivo and in vitro through activation of the NRF-2 transcription factor [146]. NRF-2 (also known as NFE2L2) protects cells from oxidative stress and cigarette smoke toxicity [147], but its overexpression in LUAD cell lines results in drug resistance [148]. Consistent with dichotomous nature of the NRF-2 response [149], downregulation of LUCAT1 also results in smoke-mediated cell death [146] suggesting LUCAT1 may in part mediate the response of NRF-2 to oxidative stress. In addition, LUCAT1 is upregulated in cisplatin-resistant ovarian cancer [150]. It remains unclear whether LUCAT1 targets novel downstream genes involved in oxidative stress or whether it aids NRF-2 in activating NRF-2-dependent genes. Further mechanistic research on LUCAT1 can elucidate if the oxidative stress response is related to or independent from LUCAT1 upregulation by chemotherapy.
PVT1 (plasmacytoma variant transcript 1)
is transcribed ~60kb downstream of the MYC oncogene, and both reside within the 8q24 locus which undergoes copy number amplification in several cancers [151, 152]. Although MYC is an established oncogene, PVT1 is also emerging as a prominent player in cancer. A recent study illustrated that some MYC-driven cancers are dependent on PVT1 activity, as PVT1 could stabilize MYC protein levels by preventing MYC phosphorylation [153]. In addition, silencing of PVT1 in PVT1/MYC amplified cancers resulted in apoptosis, whereas MYC silencing had no effect, implying PVT1 has a MYC-independent role in blocking apoptosis. The inhibition of apoptosis due to PVT1 overexpression may be partially due to its role in silencing the LATS2 gene via recruitment of EZH2 to its locus, inducing chromatin remodeling and gene silencing [154]. The LATS2 tumor suppressor is involved in a variety of functions, including induction of apoptosis and cell cycle control [155, 156]. Inhibition of LATS2 was previously observed in NSCLC, and PVT1 overexpression was found to correlate with poorer overall prognosis [154].
The PVT1 locus also contains multiple miRNA genes, including miR-1204, miR-1205, miR-1206, miR-1207-3p, miR-1207-5p, and miR-1208 [157]. Of interest, both miR-1204 and miR-1207-5p have demonstrated tumor suppressive properties [158, 159]. Surprisingly, p53 mediates the transcriptional expression of both PVT1 and miR-1204 [158]. In addition, ectopic expression of miR-1204 induced p53-mediated growth inhibition in HCT116 cells. Therefore, induction of transcripts from the 8q24 locus results in lncRNAs that promote oncogenesis, and paradoxically, miRNAs that inhibit tumor promotion via p53. This may seem contradictory, but there have been reports of p53 mediating pro-survival pathways during DNA repair [160], such as p53 activation of p21/NRF2 signaling [161]. How this tight balance between pro-survival during DNA repair and apoptosis/cell death is disrupted in cancer will require further research in the downstream targets of the PVT1 locus transcripts. The significance of other miRNAs inhabiting the 8q24 locus in p53-mediated signaling is unknown at the time of this writing.
Targeted therapy against 8q24 amplified cancers has remained challenging due to MYC being essential and in high abundance across normal tissue [162]. Because PVT1 is less abundant in across normal tissue and possesses a protective role for MYC protein, PVT1 appears to be a promising target for 8q24 amplified cancers.
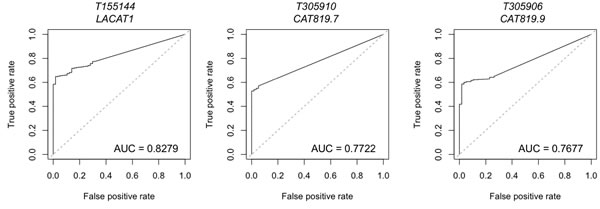
Figure 4: Lineage-specific lncRNAs identified in LUAD by miTranscriptome. Data on lineage and cancer-specific LUAD lncRNAs was obtained from miTranscriptome. The top three lncRNAs with specificity to LUAD are shown. AUC = Area under the curve. False positive rate and false negative rate were generated using miTranscriptome-calculated expression levels for primary LUAD and normal lung tissue samples generated by TCGA. ROC curves were generated using the ROCR package in R.
SFTA1P (surfactant associated 1 pseudogene)
Surfactant signaling is the distinguishing hallmark of alveolar epithelial type 2 (AT2) cells, a purported cell of origin for LUAD [163]. SFTA1P expression is correlated with other components of the surfactant machinery [164], and elevated SFTA1P levels indicate a better prognostic outcome for LUAD cancer patients (cox p-value = 0.009)[143]. This indicates that SFTA1P may hold potential as a biomarker of outcome prediction. However, because this gene is co-expressed with markers of differentiated AT2 cells, the loss of SFTA1P in a subset of LUAD cancers may be reflective of the overall differentiation state of the tumors. Moreover, the SFTA1P pseudogene is not located within genomic proximity to any of the other surfactant-protein producing genes, and the mechanisms (if any) by which SFTA1P functions remains unknown.
LINC00460
This lncRNA is a multi-exonic, intergenic lncRNA over 100kb from the nearest mRNA gene, EFNB2. In addition to being found overexpressed in LUAD in the above studies, LINC00460 is upregulated in head and neck squamous cell carcinoma, kidney carcinoma, and pancreatic cancer [165, 166]. While expression of LINC00460 is correlated with EFNB2 in LUAD (R=0.54, p=4.35e-28(143)), little research into the function or application of this lncRNA has been performed. The neighboring gene, EFNB2, encodes for EphrinB, one of many ligands for the Ephrin tyrosine kinase receptor. Much has been done implicating EFNB2 and the EphrinB receptor in development and progression of lung cancer [167-169]. However, the role LINC00460 plays in this process, if any, has yet to be determined.
LUAD antisense transcripts and their relation to nearby protein coding genes
Antisense transcription has been observed at the transcription start site of numerous protein coding genes [170, 171]. This class of antisense transcripts range from siRNA [172] to antisense lncRNAs, such as HOTAIR [173]. Many have documented antagonistic activity, from epigenetic regulation [174] to direct disruption of the transcriptional machinery [175]. Here, we highlight a few antisense lncRNAs identified through bioinformatics analysis to be involved in LUAD, while the entire list is summarized in Table 3.
FENDRR (FoxF1 adjacent non-coding developmental RNA)
This gene is transcribed in the antisense direction from the adjacent FOXF1 transcription factor. As expected, expression of FENDRR is highly correlated to FOXF1 (R=0.816, p value =1.52e-85 [143]). The FOXF1 transcription factor is implicated in mesoderm development, and similarly FENDRR is implicated in embryogenic mesoderm formation, specifically heart development [176, 177]. FOXF1 is overexpressed in LUAD and plays a central role in regulating epithelial-to-mesenchymal transition by promoting tumorigenesis of adenomas toward adenocarcinomas [178, 179]. Mechanistically, FENDRR is proposed to affect the extracellular matrix due to its inverse correlation with fibronectin1 expression in gastric cancer cell lines [180]. Disrupting fibronectin1 is associated with tumor migration and metastasis [181]. Adding extra weight to the argument that FENDRR may promote EMT and metastasis, Xu et al., found that lower FENDRR expression correlates with higher metastatic potential and poorer outcomes in LUAD patients [180]. Similar to HOTAIR and MEG3, FENDRR appears to form RNA:DNA triplexes to recruit PRC2 complex during embryonic mesoderm patterning, which when disrupted leads to deformation of the heart and embryo death [182].
FEZF1-AS1 (FEZ family zinc finger 1- antisense 1)
FEZF1 (also known as ZNF312b) is a zinc finger transcriptional repressor that is an epigenetically-regulated oncogene in gastric cancer [183]. This protein promotes proliferation via kRAS-oncogene activation [184]. FEZF1-AS1 positively regulates expression of FEZF1 mRNA expression in vitro as well as in the TANRIC analysis of TCGA LUAD data (p=2.85e-104). FEZF1-AS1 is upregulated in human primary colorectal carcinoma, and affects colorectal cancer cell proliferation, metastasis, and invasion [185]. However, it remains to be determined how FEZF1-AS1 and FEZF1 interact mechanistically.
SBK1-AS1 (SH3 domain binding kinase 1 antisense-1; RP11-57A19.2)
SBK1 is a serine/threonine kinase family member implicated in ovarian serous adenocarcinoma cell survival [186]. Several serine threonine kinase family members exhibit oncogenic behavior, such as PIM1 and BRAF. These are attractive therapeutic targets, as small molecule inhibitors have proven effective in halting cancer progression [187, 188]. RP11-57A19.2 is transcribed in an antisense direction from the SBK1 promoter, and the two have correlated expression in multiple cancer types (TCGA-LUAD R=0.597, TCGA-OVR=0.701, TCGA-BRCA=0.671, p=3.95e-108 [143]), however little to nothing is known about the expression, function, and regulation of RP11-57A19.2.
GATA6-AS1 (GATA-binding protein-6 antisense-1)
GATA6 is an important regulatory transcription factor in alveolar epithelial cell biology [189, 190]. The antisense GATA6-AS1 is correlated to GATA6 expression during lung development, albeit with more cell-type specific restriction than the GATA6 transcription factor [191], as well as in TCGA LUAD datasets (R=0.772, p=3.53e-71 [143]). Overexpression of BM742401, an expressed sequence tag which corresponds to GATA6-AS1, reduced cancer metastasis and decreased secretion levels of MMP9, though the mechanism by which GATA6-AS1 mediated these effects remains unknown. Interestingly, these authors investigated whether GATA6-AS1 overexpression affected the expression of GATA6, but found no change (this data not reported) [192]. Expression of GATA6 in LUAD is associated with a more differentiated state, and reflective of that, with better overall patient survival [193]. Whether the expression of GATA6-AS1 functions to maintain the differentiated state, or is merely a passive reflection of differentiation, remains to be determined.
Clinical Ramifications
Lung cancer remains the leading cause of cancer-related death in the United States. Although improvements in surgical treatment and chemotherapies have shown some progress, the 5-year survival rate lingers at ~15% [12]. In addition, NSCLC is composed of several differing subtypes, each with their own set of heterogenic factors that result in cancer. The array of molecular mechanisms implicated in the genesis of this disease underscores the need more accurate prognostic markers, to implement therapies targeted to the specific pathways disrupted in each disease subtype and inform clinicians on predicted patient outcomes.
Non-coding RNAs hold promise as biomarkers for a variety of cancers. For instance, increased miR-486 levels showed efficacy as a blood-based biomarker for early detection of NSCLC, and lower levels of miR-486 post-surgical resection was an effective predictor of recurrence-free survival of NSCLC patients [194]. As mentioned above, MALAT1 and LUCAT1 also hold promise as prognostic biomarkers as their elevated expression is linked with poorer overall survival [195]. However, enthusiasm surrounding MALAT1’s applicability to early detection was tempered by research describing its overall sensitivity of detection at only 56%, meaning almost half of true positive LUAD cases would be missed using this biomarker alone [196]. To circumvent this issue, MALAT1 was included in a panel of lncRNAs to improve the detection sensitivity while maintaining the specificity for LUAD. The other lncRNAs included in the panel were: ENST00000540136, NR034174, uc001gzl.3, uc004bbl.1, and ENST00000434223. When combined, this panel outperformed any individual lncRNA in the training set. The testing set reached an AUC of 0.978 for tumor identification, with 92% sensitivity and 98% specificity [138], reinforcing the notion that the combining the inherent cell-type specificity of many lncRNAs with the sensitivity of others can aide in the development of early detection tools.
Separate from this panel of biomarkers, miTranscriptome has also undertaken the task of calculating the specificity of expression within each individual cancer for the entire lncRNA transcriptome. They identified 25 lncRNAs that demonstrated statistically robust specificity for LUAD [142], only five of which were previously annotated. These genes included PVT1 (mentioned above as adjacent to the MYC oncogene), DYPD-AS1, LACAT24, and LACAT8 (Table 4). It is interesting to note that this analysis included 23 cancerous tissues alongside 12 normal tissues, a step not typically undertaken when assessing the feasibility of early detection tools. As no single study can yet address the wide diversity of cell types present in the human body [197], including as many cell types as is available can nevertheless decrease the risk of investing heavily in the development of a biomarker, only to see it fail in clinical trials due to off-target effects.
Table 4: miTranscriptome-defined LUAD lineage-specific lncRNAs
miTranscriptome |
Alternate IDs |
Position (GRCh37/hg19) |
Genomic Context |
Patient Outcome |
CAT1100.3 |
PVT1 |
chr8:128996355-129130070 (-) |
Adjacent to MYC oncogene |
Correlated to MYC expression and tumor progression. PVT1 presence indicates poorer prognosis for patients |
DPYD-AS1.2 |
NR_046590.1 |
chr1:97720954-97751573 (+) |
Antisense to DYPD |
DYPD implicated in increased toxicity to patients treated with 5FU. |
LACAT24 |
LOC101927132 |
chr16:47936832-47961855 (+) |
Intergenic |
Unknown |
LACAT8.1 |
LACAT8 |
chr12:131471968-131478539 (-) |
Antisense to GPR133 (ADGRD1) |
GPR133 upregulation correlated to poorer survival in GBM |
miTranscriptome-calcuated lncRNAs with lineage specificity to LUAD.
In the future, testing for lncRNA expression could also yield gains in personalized medicine. For instance, overexpression of HOTAIR in LUAD results in chemo-resistance towards cisplatin [198]. HOTAIR-induced drug resistance was attributed in part to downregulation of p21. Although the mechanism by which HOTAIR regulates p21 remains unknown, previous studies have reported EZH2 is involved in p21 suppression [199]. Considering the known involvement of HOTAIR with the PRC2/EZH2 complex, HOTAIRs role in mediating p21 suppression and cisplatin resistance might be due to epigenic modifications. Therefore, testing for HOTAIR expression after resection may insulate a subset of the patient population from having to undergo chemotherapy and instead direct them toward promising EZH2 inhibitors, such as E7438, currently under evaluation in clinical trials [200].
A lncRNA with oncogenic activity and expression restricted to a specific cancer would be an ideal therapeutic target. To that end, anti-RNA treatments are currently being developed to diversify the options available to clinicians. However, many obstacles have arisen, including a lack of efficient delivery methods, RNA degradation, and aberrant immune system activation [201]. In addition, some lncRNAs have high turnover and low transcriptional expression, making them difficult to target effectively. In such cases, knowledge of pathway involvement is needed to develop effective treatments that target downstream signaling molecules affected by aberrant lncRNA activity.
Conclusions
Over the last decade, lncRNAs have been recognized as a diverse class of macromolecules in terms of function and mechanism. Many lncRNAs have demonstrated functional activity in a wide range of cancers. While the lncRNA transcriptome is far from complete, we now have an appreciation of their diversity due to cell-type specificity. This characteristic can aid in the development of early detection methods and targeted therapies for multiple cancer types. Alongside these immediate applications, understanding the mechanism(s) by which these transcripts are regulated will shed light on the etiology of cancer development, allowing clinicians to implement better treatment strategies and improve overall survival rates.
While research in this field is still in preliminary stages, multiple large-cohort patient studies such as TCGA, ENCODE, and SEO alongside previous microarray studies have delineated a host of lncRNAs with potential for novel therapeutic strategies. As such, they have provided framework for the interrogation of molecular mechanisms that lncRNAs utilize in LUAD initiation, promotion, and progression. While in this review we focused on lncRNAs validated by three different studies, it is by no means exhaustive. It remains to be seen if other case-controlled, multi-ethnic cohorts will unveil an entirely new set of lncRNAs with implicit utility in the diagnosis and treatment of LUAD. It will be interesting to follow what discoveries that work will yield in the development of novel therapeutics and early detection strategies in the years to come.
Acknowledgments
The authors acknowledge input from Ite A. Laird-Offringa on the topics discussed, and artwork generated by Pat Marconett. Funding was provided by the Department of Surgery, Keck School of Medicine, USC.
Conflicts of Interest
There is no conflict of interest.
References
1. Brown CJ, Ballabio A, Rupert JL, Lafreniere RG, Grompe M, Tonlorenzi R, Willard HF. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature. 1991;349:38-44.
2. Borodovsky M, Lomsadze A, Ivanov N, Mills R. Eukaryotic gene prediction using GeneMark.hmm. Curr Protoc Bioinformatics. 2003;Chapter 4:Unit4.6.
3. Besemer J, Borodovsky M. GeneMark: web software for gene finding in prokaryotes, eukaryotes and viruses. Nucleic Acids Res. 2005;33:W451-4.
4. Lukashin AV, Borodovsky M. GeneMark.hmm: new solutions for gene finding. Nucleic Acids Res. 1998;26:1107-15.
5. van Baren MJ, Koebbe BC, Brent MR. Using N-SCAN or TWINSCAN to predict gene structures in genomic DNA sequences. Curr Protoc Bioinformatics. 2007;Chapter 4:Unit 4.8.
6. Wang J, Zhang J, Zheng H, Li J, Liu D, Li H, Samudrala R, Yu J, Wong GK. Mouse transcriptome: neutral evolution of ‘non-coding’ complementary DNAs. Nature. 2004;431:1 p following 757; discussion following
7. Gross SS, Brent MR. Using multiple alignments to improve gene prediction. J Comput Biol. 2006;13:379-93.
8. Korf I, Flicek P, Duan D, Brent MR. Integrating genomic homology into gene structure prediction. Bioinformatics. 2001 (Suppl 1); 17:S140–48.
9. van Baren MJ, Brent MR. Iterative gene prediction and pseudogene removal improves genome annotation. Genome Res. 2006;16:678-85.
10. Pang KC, Frith MC, Mattick JS. Rapid evolution of noncoding RNAs: lack of conservation does not mean lack of function. Trends Genet. 2006;22:1-5.
11. Chen Z, Fillmore CM, Hammerman PS, Kim CF, Wong KK. Non-small-cell lung cancers: a heterogeneous set of diseases. Nat Rev Cancer. 2014;14:535-46.
12. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7-30.
13. Aberle DR, Adams AM, Berg CD, Black WC, Clapp JD, Fagerstrom RM, Gareen IF, Gatsonis C, Marcus PM, Sicks JD, and National Lung Screening Trial Research Team. Reduced lung-cancer mortality with low-dose computed tomographic screening. N Engl J Med. 2011;365:395-409.
14. Sun S, Schiller JH, Gazdar AF. Lung cancer in never smokers—a different disease. Nat Rev Cancer. 2007;7:778-90.
15. Network CG, and Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511:543-50.
16. Seo JS, Ju YS, Lee WC, Shin JY, Lee JK, Bleazard T, Lee J, Jung YJ, Kim JO, Shin JY, Yu SB, Kim J, Lee ER, et al. The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res. 2012;22:2109-19.
17. Pao W, Miller V, Zakowski M, Doherty J, Politi K, Sarkaria I, Singh B, Heelan R, Rusch V, Fulton L, Mardis E, Kupfer D, Wilson R, et al. EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci USA. 2004;101:13306-11.
18. Camidge DR, Bang YJ, Kwak EL, Iafrate AJ, Varella-Garcia M, Fox SB, Riely GJ, Solomon B, Ou SH, Kim DW, Salgia R, Fidias P, Engelman JA, et al. Activity and safety of crizotinib in patients with ALK-positive non-small-cell lung cancer: updated results from a phase 1 study. Lancet Oncol. 2012;13:1011-9.
19. Solomon BJ, Mok T, Kim DW, Wu YL, Nakagawa K, Mekhail T, Felip E, Cappuzzo F, Paolini J, Usari T, Iyer S, Reisman A, Wilner KD, et al, and PROFILE 1014 Investigators. First-line crizotinib versus chemotherapy in ALK-positive lung cancer. N Engl J Med. 2014;371:2167-77.
20. Christensen JG, Zou HY, Arango ME, Li Q, Lee JH, McDonnell SR, Yamazaki S, Alton GR, Mroczkowski B, Los G. Cytoreductive antitumor activity of PF-2341066, a novel inhibitor of anaplastic lymphoma kinase and c-Met, in experimental models of anaplastic large-cell lymphoma. Mol Cancer Ther. 2007;6:3314-22.
21. Pao W, Miller VA, Politi KA, Riely GJ, Somwar R, Zakowski MF, Kris MG, Varmus H. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med. 2005;2:e73.
22. Sun M, Kraus WL. Minireview: Long noncoding RNAs: new “links” between gene expression and cellular outcomes in endocrinology. Mol Endocrinol. 2013;27:1390-402.
23. Fu X, Li H, Liu C, Hu B, Li T, Wang Y. Long noncoding RNA AK126698 inhibits proliferation and migration of non-small cell lung cancer cells by targeting Frizzled-8 and suppressing Wnt/-catenin signaling pathway. Onco Targets Ther. 2016;9:3815-27.
24. Bernstein E, Allis CD. RNA meets chromatin. Genes Dev. 2005;19:1635-55.
25. Shahryari A, Jazi MS, Samaei NM, Mowla SJ. Long non-coding RNA SOX2OT: expression signature, splicing patterns, and emerging roles in pluripotency and tumorigenesis. Front Genet. 2015;6:196.
26. Guil S, Esteller M. Cis-acting noncoding RNAs: friends and foes. Nat Struct Mol Biol. 2012;19:1068-75.
27. Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, Thomas K, Presser A, Bernstein BE, van Oudenaarden A, Regev A, Lander ES, Rinn JL. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA. 2009;106:11667-72.
28. Wilusz JE. Long noncoding RNAs: Re-writing dogmas of RNA processing and stability. Biochim Biophys Acta. 2016;1859:128-38.
29. Liang WC, Fu WM, Wong CW, Wang Y, Wang WM, Hu GX, Zhang L, Xiao LJ, Wan DC, Zhang JF, Waye MM. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget. 2015; 6:22513–25. https://doi.org/10.18632/oncotarget.4154.
30. Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007;129:1311-23.
31. Kitagawa M, Kitagawa K, Kotake Y, Niida H, Ohhata T. Cell cycle regulation by long non-coding RNAs. Cell Mol Life Sci. 2013;70:4785-94.
32. Rossi MN, Antonangeli F. LncRNAs: New Players in Apoptosis Control. Int J Cell Biol. 2014;2014:473857.
33. Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014;15:7-21.
34. Heward JA, Lindsay MA. Long non-coding RNAs in the regulation of the immune response. Trends Immunol. 2014;35:408-19.
35. Luo G, Wang M, Wu X, Tao D, Xiao X, Wang L, Min F, Zeng F, Jiang G. Long Non-Coding RNA MEG3 Inhibits Cell Proliferation and Induces Apoptosis in Prostate Cancer. Cell Physiol Biochem. 2015;37:2209-20.
36. Botti G, Marra L, Malzone MG, Anniciello A, Botti C, Franco R, Cantile M. LncRNA HOTAIR as prognostic circulating marker and potential therapeutic target in patients with tumor diseases. Curr Drug Targets. 2015.
37. Hibi K, Nakamura H, Hirai A, Fujikake Y, Kasai Y, Akiyama S, Ito K, Takagi H. Loss of H19 imprinting in esophageal cancer. Cancer Res. 1996;56:480-2.
38. Douc-Rasy S, Barrois M, Fogel S, Ahomadegbe JC, Stéhelin D, Coll J, Riou G. High incidence of loss of heterozygosity and abnormal imprinting of H19 and IGF2 genes in invasive cervical carcinomas. Uncoupling of H19 and IGF2 expression and biallelic hypomethylation of H19. Oncogene. 1996;12:423-30.
39. Kondo M, Takahashi T. Altered genomic imprinting in the IGF2 and H19 genes in human lung cancer. [Article in Japanese] Nihon Rinsho. 1996; 54:492–96. https://www.ncbi.nlm.nih.gov/pubmed/8838103.
40. Zhang E, Li W, Yin D, De W, Zhu L, Sun S, Han L. c-Myc-regulated long non-coding RNA H19 indicates a poor prognosis and affects cell proliferation in non-small-cell lung cancer. Tumour Biol. 2016;37:4007-15.
41. Raveh E, Matouk IJ, Gilon M, Hochberg A. The H19 Long non-coding RNA in cancer initiation, progression and metastasis - a proposed unifying theory. Mol Cancer. 2015;14:184.
42. Court F, Baniol M, Hagege H, Petit JS, Lelay-Taha MN, Carbonell F, Weber M, Cathala G, Forne T. Long-range chromatin interactions at the mouse Igf2/H19 locus reveal a novel paternally expressed long non-coding RNA. Nucleic Acids Res. 2011;39:5893-906.
43. Barsyte-Lovejoy D, Lau SK, Boutros PC, Khosravi F, Jurisica I, Andrulis IL, Tsao MS, Penn LZ. The c-Myc oncogene directly induces the H19 noncoding RNA by allele-specific binding to potentiate tumorigenesis. Cancer Res. 2006;66:5330-7.
44. Cai X, Cullen BR. The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA. 2007;13:313-6.
45. Zou T, Jaladanki SK, Liu L, Xiao L, Chung HK, Wang JY, Xu Y, Gorospe M, Wang JY. LncRNA H19 regulates intestinal epithelial barrier function via miR-675 by interacting with RNA-binding protein HuR. Mol Cell Biol. 2016.
46. Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L, Lu L, Liu C, Yi JS, Zhang H, Min W, Bennett AM, Gregory RI, et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell. 2013;52:101-12.
47. Abedi N, Mohammadi-Yeganeh S, Koochaki A, Karami F, Paryan M. miR-141 as potential suppressor of -catenin in breast cancer. Tumour Biol. 2015;36:9895-901.
48. Tang H, Kong Y, Guo J, Tang Y, Xie X, Yang L, Su Q, Xie X. Diallyl disulfide suppresses proliferation and induces apoptosis in human gastric cancer through Wnt-1 signaling pathway by up-regulation of miR-200b and miR-22. Cancer Lett. 2013;340:72-81.
49. Liang WC, Fu WM, Wang YB, Sun YX, Xu LL, Wong CW, Chan KM, Li G, Waye MM, Zhang JF. H19 activates Wnt signaling and promotes osteoblast differentiation by functioning as a competing endogenous RNA. Sci Rep. 2016;6:20121.
50. Luo M, Li Z, Wang W, Zeng Y, Liu Z, Qiu J. Long non-coding RNA H19 increases bladder cancer metastasis by associating with EZH2 and inhibiting E-cadherin expression. Cancer Lett. 2013;333:213-21.
51. Yang F, Bi J, Xue X, Zheng L, Zhi K, Hua J, Fang G. Up-regulated long non-coding RNA H19 contributes to proliferation of gastric cancer cells. FEBS J. 2012;279:3159-65.
52. Tao W, Levine AJ. P19(ARF) stabilizes p53 by blocking nucleo-cytoplasmic shuttling of Mdm2. Proc Natl Acad Sci USA. 1999;96:6937-41.
53. Mullighan CG, Williams RT, Downing JR, Sherr CJ. Failure of CDKN2A/B (INK4A/B-ARF)-mediated tumor suppression and resistance to targeted therapy in acute lymphoblastic leukemia induced by BCR-ABL. Genes Dev. 2008;22:1411-5.
54. Roussel MF. The INK4 family of cell cycle inhibitors in cancer. Oncogene. 1999;18:5311-7.
55. Gil J, Peters G. Regulation of the INK4b-ARF-INK4a tumour suppressor locus: all for one or one for all. Nat Rev Mol Cell Biol. 2006;7:667-77.
56. Schuster K, Venkateswaran N, Rabellino A, Girard L, Peña-Llopis S, Scaglioni PP. Nullifying the CDKN2AB locus promotes mutant K-ras lung tumorigenesis. Mol Cancer Res. 2014;12:912-23.
57. Yap KL, Li S, Muñoz-Cabello AM, Raguz S, Zeng L, Mujtaba S, Gil J, Walsh MJ, Zhou MM. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell. 2010;38:662-74.
58. Kotake Y, Nakagawa T, Kitagawa K, Suzuki S, Liu N, Kitagawa M, Xiong Y. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene. 2011;30:1956-62.
59. Nie FQ, Sun M, Yang JS, Xie M, Xu TP, Xia R, Liu YW, Liu XH, Zhang EB, Lu KH, Shu YQ. Long noncoding RNA ANRIL promotes non-small cell lung cancer cell proliferation and inhibits apoptosis by silencing KLF2 and P21 expression. Mol Cancer Ther. 2015;14:268-77.
60. Yu X, Li Z. Long non-coding RNA HOTAIR: A novel oncogene (Review). Mol Med Rep. 2015; 12:5611–18. Review.
61. Loewen G, Zhuo Y, Zhuang Y, Jayawickramarajah J, Shan B. lincRNA HOTAIR as a novel promoter of cancer progression. J Cancer Res Updates. 2014;3:134-40.
62. Loewen G, Jayawickramarajah J, Zhuo Y, Shan B. Functions of lncRNA HOTAIR in lung cancer. J Hematol Oncol. 2014;7:90.
63. Zhou X, Chen J, Tang W. The molecular mechanism of HOTAIR in tumorigenesis, metastasis, and drug resistance. Acta Biochim Biophys Sin (Shanghai). 2014;46:1011-5.
64. Cifuentes-Rojas C, Hernandez AJ, Sarma K, Lee JT. Regulatory interactions between RNA and polycomb repressive complex 2. Mol Cell. 2014;55:171-85.
65. Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329:689-93.
66. Chu C, Qu K, Zhong FL, Artandi SE, Chang HY. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol Cell. 2011;44:667-78.
67. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010;464:1071-6.
68. Liu XH, Sun M, Nie FQ, Ge YB, Zhang EB, Yin DD, Kong R, Xia R, Lu KH, Li JH, De W, Wang KM, Wang ZX. Lnc RNA HOTAIR functions as a competing endogenous RNA to regulate HER2 expression by sponging miR-331-3p in gastric cancer. Mol Cancer. 2014;13:92.
69. Geng YJ, Xie SL, Li Q, Ma J, Wang GY. Large intervening non-coding RNA HOTAIR is associated with hepatocellular carcinoma progression. J Int Med Res. 2011;39:2119-28.
70. Wu S, Zheng C, Chen S, Cai X, Shi Y, Lin B, Chen Y. Overexpression of long non-coding RNA HOTAIR predicts a poor prognosis in patients with acute myeloid leukemia. Oncol Lett. 2015;10:2410-4.
71. Li J, Yang S, Su N, Wang Y, Yu J, Qiu H, He X. Overexpression of long non-coding RNA HOTAIR leads to chemoresistance by activating the Wnt/-catenin pathway in human ovarian cancer. Tumour Biol. 2015; 36:9093–94.
72. Huang L, Liao LM, Liu AW, Wu JB, Cheng XL, Lin JX, Zheng M. Overexpression of long noncoding RNA HOTAIR predicts a poor prognosis in patients with cervical cancer. Arch Gynecol Obstet. 2014;290:717-23.
73. Nie Y, Liu X, Qu S, Song E, Zou H, Gong C. Long non-coding RNA HOTAIR is an independent prognostic marker for nasopharyngeal carcinoma progression and survival. Cancer Sci. 2013;104:458-64.
74. Ishibashi M, Kogo R, Shibata K, Sawada G, Takahashi Y, Kurashige J, Akiyoshi S, Sasaki S, Iwaya T, Sudo T, Sugimachi K, Mimori K, Wakabayashi G, et al. Clinical significance of the expression of long non-coding RNA HOTAIR in primary hepatocellular carcinoma. Oncol Rep. 2013;29:946-50.
75. Berrondo C, Flax J, Kucherov V, Siebert A, Osinski T, Rosenberg A, Fucile C, Richheimer S, Beckham CJ. Expression of the Long Non-Coding RNA HOTAIR Correlates with Disease Progression in Bladder Cancer and Is Contained in Bladder Cancer Patient Urinary Exosomes. PLoS One. 2016;11:e0147236.
76. Cai B, Wu Z, Liao K, Zhang S. Long noncoding RNA HOTAIR can serve as a common molecular marker for lymph node metastasis: a meta-analysis. Tumour Biol. 2014;35:8445-50.
77. Sørensen KP, Thomassen M, Tan Q, Bak M, Cold S, Burton M, Larsen MJ, Kruse TA. Long non-coding RNA HOTAIR is an independent prognostic marker of metastasis in estrogen receptor-positive primary breast cancer. Breast Cancer Res Treat. 2013;142:529-36.
78. Chen FJ, Sun M, Li SQ, Wu QQ, Ji L, Liu ZL, Zhou GZ, Cao G, Jin L, Xie HW, Wang CM, Lv J, De W, et al. Upregulation of the long non-coding RNA HOTAIR promotes esophageal squamous cell carcinoma metastasis and poor prognosis. Mol Carcinog. 2013;52:908-15.
79. Wu Y, Zhang L, Wang Y, Li H, Ren X, Wei F, Yu W, Liu T, Wang X, Zhou X, Yu J, Hao X. Long non-coding RNA HOTAIR promotes tumor cell invasion and metastasis by recruiting EZH2 and repressing E-cadherin in oral squamous cell carcinoma. Int J Oncol. 2015;46:2586-94.
80. Wu ZH, Wang XL, Tang HM, Jiang T, Chen J, Lu S, Qiu GQ, Peng ZH, Yan DW. Long non-coding RNA HOTAIR is a powerful predictor of metastasis and poor prognosis and is associated with epithelial-mesenchymal transition in colon cancer. Oncol Rep. 2014;32:395-402.
81. Ono H, Motoi N, Nagano H, Miyauchi E, Ushijima M, Matsuura M, Okumura S, Nishio M, Hirose T, Inase N, Ishikawa Y. Long noncoding RNA HOTAIR is relevant to cellular proliferation, invasiveness, and clinical relapse in small-cell lung cancer. Cancer Med. 2014;3:632-42.
82. Liu XH, Liu ZL, Sun M, Liu J, Wang ZX, De W. The long non-coding RNA HOTAIR indicates a poor prognosis and promotes metastasis in non-small cell lung cancer. BMC Cancer. 2013;13:464.
83. Nakagawa T, Endo H, Yokoyama M, Abe J, Tamai K, Tanaka N, Sato I, Takahashi S, Kondo T, Satoh K. Large noncoding RNA HOTAIR enhances aggressive biological behavior and is associated with short disease-free survival in human non-small cell lung cancer. Biochem Biophys Res Commun. 2013;436:319-24.
84. Miyoshi N, Wagatsuma H, Wakana S, Shiroishi T, Nomura M, Aisaka K, Kohda T, Surani MA, Kaneko-Ishino T, Ishino F. Identification of an imprinted gene, Meg3/Gtl2 and its human homologue MEG3, first mapped on mouse distal chromosome 12 and human chromosome 14q. Genes Cells. 2000;5:211-20.
85. Liu B, Shen ED, Liao MM, Hu YB, Wu K, Yang P, Zhou L, Chen WD. Expression and mechanisms of long non-coding RNA genes MEG3 and ANRIL in gallbladder cancer. Tumour Biol. 2016.
86. Zhang J, Yao T, Wang Y, Yu J, Liu Y, Lin Z. Long noncoding RNA MEG3 is downregulated in cervical cancer and affects cell proliferation and apoptosis by regulating miR-21. Cancer Biol Ther. 2016;17:104-13.
87. Peng W, Si S, Zhang Q, Li C, Zhao F, Wang F, Yu J, Ma R. Long non-coding RNA MEG3 functions as a competing endogenous RNA to regulate gastric cancer progression. J Exp Clin Cancer Res. 2015;34:79.
88. Su L, Han D, Wu J, Huo X. Skp2 regulates non-small cell lung cancer cell growth by Meg3 and miR-3163. Tumour Biol. 2015.
89. Kawakami T, Chano T, Minami K, Okabe H, Okada Y, Okamoto K. Imprinted DLK1 is a putative tumor suppressor gene and inactivated by epimutation at the region upstream of GTL2 in human renal cell carcinoma. Hum Mol Genet. 2006;15:821-30.
90. Mondal T, Subhash S, Vaid R, Enroth S, Uday S, Reinius B, Mitra S, Mohammed A, James AR, Hoberg E, Moustakas A, Gyllensten U, Jones SJ, et al. MEG3 long noncoding RNA regulates the TGF- pathway genes through formation of RNA-DNA triplex structures. Nat Commun. 2015;6:7743.
91. Lian Y, Ding J, Zhang Z, Shi Y, Zhu Y, Li J, Peng P, Wang J, Fan Y, De W, Wang K. The long noncoding RNA HOXA transcript at the distal tip promotes colorectal cancer growth partially via silencing of p21 expression. Tumour Biol. 2015.
92. Ren YK, Xiao Y, Wan XB, Zhao YZ, Li J, Li Y, Han GS, Chen XB, Zou QY, Wang GC, Lu CM, Xu YC, Wang YC. Association of long non-coding RNA HOTTIP with progression and prognosis in colorectal cancer. Int J Clin Exp Pathol. 2015;8:11458-63.
93. Li F, Cao L, Hang D, Wang F, Wang Q. Long non-coding RNA HOTTIP is up-regulated and associated with poor prognosis in patients with osteosarcoma. Int J Clin Exp Pathol. 2015;8:11414-20.
94. Deng HP, Chen L, Fan T, Zhang B, Xu Y, Geng Q. Long non-coding RNA HOTTIP promotes tumor growth and inhibits cell apoptosis in lung cancer. Cell Mol Biol (Noisy-le-grand). 2015;61:34-40.
95. Cheng Y, Jutooru I, Chadalapaka G, Corton JC, Safe S. The long non-coding RNA HOTTIP enhances pancreatic cancer cell proliferation, survival and migration. Oncotarget. 2015; 6:10840–52. https://doi.org/10.18632/oncotarget.3450.
96. Zhang H, Zhao L, Wang YX, Xi M, Liu SL, Luo LL. Long non-coding RNA HOTTIP is correlated with progression and prognosis in tongue squamous cell carcinoma. Tumour Biol. 2015;36:8805-9.
97. Jiang YJ, Bikle DD. LncRNA profiling reveals new mechanism for VDR protection against skin cancer formation. J Steroid Biochem Mol Biol. 2014;144 Pt A:87-90.
98. Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta RA, Wysocka J, Lei M, Dekker J, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011;472:120-4.
99. Gutschner T, Hämmerle M, Eissmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, Zörnig M, MacLeod AR, Spector DL, et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013;73:1180-9.
100. Sun C, Li S, Zhang F, Xi Y, Wang L, Bi Y, Li D. Long non-coding RNA NEAT1 promotes non-small cell lung cancer progression through regulation of miR-377-3p-E2F3 pathway. Oncotarget. 2016; 7:51784–814. https://doi.org/10.18632/oncotarget.10108.
101. Shi XS, Li J, Yang RH, Zhao GR, Zhou HP, Zeng WX, Zhou M. Correlation of increased MALAT1 expression with pathological features and prognosis in cancer patients: a meta-analysis. Genet Mol Res. 2015;14:18808-19.
102. Zhang J, Zhang B, Wang T, Wang H. LncRNA MALAT1 overexpression is an unfavorable prognostic factor in human cancer: evidence from a meta-analysis. Int J Clin Exp Med. 2015;8:5499-505.
103. Schmidt LH, Spieker T, Koschmieder S, Schäffers S, Humberg J, Jungen D, Bulk E, Hascher A, Wittmer D, Marra A, Hillejan L, Wiebe K, Berdel WE, et al. The long noncoding MALAT-1 RNA indicates a poor prognosis in non-small cell lung cancer and induces migration and tumor growth. J Thorac Oncol. 2011;6:1984-92.
104. Ma KX, Wang HJ, Li XR, Li T, Su G, Yang P, Wu JW. Long noncoding RNA MALAT1 associates with the malignant status and poor prognosis in glioma. Tumour Biol. 2015;36:3355-9.
105. Zhang HM, Yang FQ, Chen SJ, Che J, Zheng JH. Upregulation of long non-coding RNA MALAT1 correlates with tumor progression and poor prognosis in clear cell renal cell carcinoma. Tumour Biol. 2015;36:2947-55.
106. Wu XS, Wang XA, Wu WG, Hu YP, Li ML, Ding Q, Weng H, Shu YJ, Liu TY, Jiang L, Cao Y, Bao RF, Mu JS, et al. MALAT1 promotes the proliferation and metastasis of gallbladder cancer cells by activating the ERK/MAPK pathway. Cancer Biol Ther. 2014;15:806-14.
107. Wei Y, Niu B. Role of MALAT1 as a Prognostic Factor for Survival in Various Cancers: A Systematic Review of the Literature with Meta-Analysis. Dis Markers. 2015;2015:164635.
108. Cho SF, Chang YC, Chang CS, Lin SF, Liu YC, Hsiao HH, Chang JG, Liu TC. MALAT1 long non-coding RNA is overexpressed in multiple myeloma and may serve as a marker to predict disease progression. BMC Cancer. 2014;14:809.
109. Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003;22:8031-41.
110. Yoshimoto R, Mayeda A, Yoshida M, Nakagawa S. MALAT1 long non-coding RNA in cancer. Biochim Biophys Acta. 2016;1859:192-9.
111. Gutschner T, Hämmerle M, Diederichs S. MALAT1 — a paradigm for long noncoding RNA function in cancer. J Mol Med (Berl). 2013;91:791-801.
112. Fan Y, Shen B, Tan M, Mu X, Qin Y, Zhang F, Liu Y. TGF--induced upregulation of malat1 promotes bladder cancer metastasis by associating with suz12. Clin Cancer Res. 2014;20:1531-41.
113. Li Y, Li Y, Chen W, He F, Tan Z, Zheng J, Wang W, Zhao Q, Li J. NEAT expression is associated with tumor recurrence and unfavorable prognosis in colorectal cancer. Oncotarget. 2015; 6:27641–50. https://doi.org/10.18632/oncotarget.4737.
114. Chen X, Kong J, Ma Z, Gao S, Feng X. Up regulation of the long non-coding RNA NEAT1 promotes esophageal squamous cell carcinoma cell progression and correlates with poor prognosis. Am J Cancer Res. 2015;5:2808-15.
115. Guo S, Chen W, Luo Y, Ren F, Zhong T, Rong M, Dang Y, Feng Z, Chen G. Clinical implication of long non-coding RNA NEAT1 expression in hepatocellular carcinoma patients. Int J Clin Exp Pathol. 2015;8:5395-402.
116. Pan LJ, Zhong TF, Tang RX, Li P, Dang YW, Huang SN, Chen G. Upregulation and clinicopathological significance of long non-coding NEAT1 RNA in NSCLC tissues. Asian Pac J Cancer Prev. 2015;16:2851-5.
117. West JA, Davis CP, Sunwoo H, Simon MD, Sadreyev RI, Wang PI, Tolstorukov MY, Kingston RE. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol Cell. 2014;55:791-802.
118. Kawaguchi T, Tanigawa A, Naganuma T, Ohkawa Y, Souquere S, Pierron G, Hirose T. SWI/SNF chromatin-remodeling complexes function in noncoding RNA-dependent assembly of nuclear bodies. Proc Natl Acad Sci USA. 2015;112:4304-9.
119. Clemson CM, Hutchinson JN, Sara SA, Ensminger AW, Fox AH, Chess A, Lawrence JB. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell. 2009;33:717-26.
120. Sasaki YT, Ideue T, Sano M, Mituyama T, Hirose T. MENepsilon/beta noncoding RNAs are essential for structural integrity of nuclear paraspeckles. Proc Natl Acad Sci USA. 2009;106:2525-30.
121. Sunwoo H, Dinger ME, Wilusz JE, Amaral PP, Mattick JS, Spector DL. MEN epsilon/beta nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res. 2009;19:347-59.
122. Liu Q, Sun S, Yu W, Jiang J, Zhuo F, Qiu G, Xu S, Jiang X. Altered expression of long non-coding RNAs during genotoxic stress-induced cell death in human glioma cells. J Neurooncol. 2015;122:283-92.
123. Choudhry H, Albukhari A, Morotti M, Haider S, Moralli D, Smythies J, Schödel J, Green CM, Camps C, Buffa F, Ratcliffe P, Ragoussis J, Harris AL, et al. Tumor hypoxia induces nuclear paraspeckle formation through HIF-2 dependent transcriptional activation of NEAT1 leading to cancer cell survival. Oncogene. 2015;34:4546.
124. Qiu M, Xu Y, Yang X, Wang J, Hu J, Xu L, Yin R. CCAT2 is a lung adenocarcinoma-specific long non-coding RNA and promotes invasion of non-small cell lung cancer. Tumour Biol. 2014;35:5375-80.
125. Wang CY, Hua L, Yao KH, Chen JT, Zhang JJ, Hu JH. Long non-coding RNA CCAT2 is up-regulated in gastric cancer and associated with poor prognosis. Int J Clin Exp Pathol. 2015;8:779-85.
126. Chen X, Liu L, Zhu W. Up-regulation of long non-coding RNA CCAT2 correlates with tumor metastasis and poor prognosis in cervical squamous cell cancer patients. Int J Clin Exp Pathol. 2015;8:13261-6.
127. Wang J, Qiu M, Xu Y, Li M, Dong G, Mao Q, Yin R, Xu L. Long noncoding RNA CCAT2 correlates with smoking in esophageal squamous cell carcinoma. Tumour Biol. 2015;36:5523-8.
128. Berndt SI, Wang Z, Yeager M, Alavanja MC, Albanes D, Amundadottir L, Andriole G, Beane Freeman L, Campa D, Cancel-Tassin G, Canzian F, Cornu JN, Cussenot O, et al, and African Ancestry Prostate Cancer GWAS Consortium. Two susceptibility loci identified for prostate cancer aggressiveness. Nat Commun. 2015;6:6889.
129. Zhang B, Jia WH, Matsuda K, Kweon SS, Matsuo K, Xiang YB, Shin A, Jee SH, Kim DH, Cai Q, Long J, Shi J, Wen W, et al, and Genetics and Epidemiology of Colorectal Cancer Consortium (GECCO), and Colorectal Transdisciplinary (CORECT) Study, and Colon Cancer Family Registry (CCFR). Large-scale genetic study in East Asians identifies six new loci associated with colorectal cancer risk. Nat Genet. 2014;46:533-42.
130. Yeager M, Orr N, Hayes RB, Jacobs KB, Kraft P, Wacholder S, Minichiello MJ, Fearnhead P, Yu K, Chatterjee N, Wang Z, Welch R, Staats BJ, et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet. 2007;39:645-9.
131. Skibola CF, Berndt SI, Vijai J, Conde L, Wang Z, Yeager M, de Bakker PI, Birmann BM, Vajdic CM, Foo JN, Bracci PM, Vermeulen RC, Slager SL, et al. Genome-wide association study identifies five susceptibility loci for follicular lymphoma outside the HLA region. Am J Hum Genet. 2014;95:462-71.
132. Cerhan JR, Berndt SI, Vijai J, Ghesquières H, McKay J, Wang SS, Wang Z, Yeager M, Conde L, de Bakker PI, Nieters A, Cox D, Burdett L, et al. Genome-wide association study identifies multiple susceptibility loci for diffuse large B cell lymphoma. Nat Genet. 2014;46:1233-8.
133. Wolpin BM, Rizzato C, Kraft P, Kooperberg C, Petersen GM, Wang Z, Arslan AA, Beane-Freeman L, Bracci PM, Buring J, Canzian F, Duell EJ, Gallinger S, et al. Genome-wide association study identifies multiple susceptibility loci for pancreatic cancer. Nat Genet. 2014;46:994-1000.
134. Rajaraman P, Melin BS, Wang Z, McKean-Cowdin R, Michaud DS, Wang SS, Bondy M, Houlston R, Jenkins RB, Wrensch M, Yeager M, Ahlbom A, Albanes D, et al. Genome-wide association study of glioma and meta-analysis. Hum Genet. 2012;131:1877-88.
135. Redis RS, Vela LE, Lu W, Ferreira de Oliveira J, Ivan C, Rodriguez-Aguayo C, Adamoski D, Pasculli B, Taguchi A, Chen Y, Fernandez AF, Valledor L, Van Roosbroeck K, et al. Allele-Specific Reprogramming of Cancer Metabolism by the Long Non-coding RNA CCAT2. Mol Cell. 2016;61:520-34.
136. Xu G, Chen J, Pan Q, Huang K, Pan J, Zhang W, Chen J, Yu F, Zhou T, Wang Y. Long noncoding RNA expression profiles of lung adenocarcinoma ascertained by microarray analysis. PLoS One. 2014;9:e104044.
137. Li J, Li P, Zhao W, Yang R, Chen S, Bai Y, Dun S, Chen X, Du Y, Wang Y, Zang W, Zhao G, Zhang G. Expression of long non-coding RNA DLX6-AS1 in lung adenocarcinoma. Cancer Cell Int. 2015;15:48.
138. Wang P, Lu S, Mao H, Bai Y, Ma T, Cheng Z, Zhang H, Jin Q, Zhao J, Mao H. Identification of biomarkers for the detection of early stage lung adenocarcinoma by microarray profiling of long noncoding RNAs. Lung Cancer. 2015;88:147-53.
139. Wang Y, Chen W, Chen J, Pan Q, Pan J. LncRNA expression profiles of EGFR exon 19 deletions in lung adenocarcinoma ascertained by using microarray analysis. Med Oncol. 2014;31:137.
140. Ning S, Zhang J, Wang P, Zhi H, Wang J, Liu Y, Gao Y, Guo M, Yue M, Wang L, Li X. Lnc2Cancer: a manually curated database of experimentally supported lncRNAs associated with various human cancers. Nucleic Acids Res. 2016;44:D980-5.
141. Park C, Yu N, Choi I, Kim W, Lee S. lncRNAtor: a comprehensive resource for functional investigation of long non-coding RNAs. Bioinformatics. 2014;30:2480-5.
142. Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, Barrette TR, Presner JR, Evans JR, Zhao S, Poliakov A, Cao X, Dhanasekaran SM, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47:199-208.
143. Li J, Han L, Roebuck P, Diao L, Liu L, Yuan Y, Weinstein JN, Liang H. TANRIC: An Interactive Open Platform to Explore the Function of lncRNAs in Cancer. Cancer Res. 2015;75:3728-37.
144. White NM, Cabanski CR, Silva-Fisher JM, Dang HX, Govindan R, Maher CA. Transcriptome sequencing reveals altered long intergenic non-coding RNAs in lung cancer. Genome Biol. 2014;15:429.
145. Saxena A, Carninci P. Long non-coding RNA modifies chromatin: epigenetic silencing by long non-coding RNAs. BioEssays. 2011;33:830-9.
146. Thai P, Statt S, Chen CH, Liang E, Campbell C, Wu R. Characterization of a novel long noncoding RNA, SCAL1, induced by cigarette smoke and elevated in lung cancer cell lines. Am J Respir Cell Mol Biol. 2013;49:204-11.
147. Cho HY, Jedlicka AE, Reddy SP, Kensler TW, Yamamoto M, Zhang LY, Kleeberger SR. Role of NRF2 in protection against hyperoxic lung injury in mice. Am J Respir Cell Mol Biol. 2002;26:175-82.
148. Homma S, Ishii Y, Morishima Y, Yamadori T, Matsuno Y, Haraguchi N, Kikuchi N, Satoh H, Sakamoto T, Hizawa N, Itoh K, Yamamoto M. Nrf2 enhances cell proliferation and resistance to anticancer drugs in human lung cancer. Clin Cancer Res. 2009;15:3423-32.
149. Müller T, Hengstermann A. Nrf2: friend and foe in preventing cigarette smoking-dependent lung disease. Chem Res Toxicol. 2012;25:1805-24.
150. Zheng ZG, Xu H, Suo SS, Xu XL, Ni MW, Gu LH, Chen W, Wang LY, Zhao Y, Tian B, Hua YJ. The Essential Role of H19 Contributing to Cisplatin Resistance by Regulating Glutathione Metabolism in High-Grade Serous Ovarian Cancer. Sci Rep. 2016;6:26093.
151. Guan Y, Kuo WL, Stilwell JL, Takano H, Lapuk AV, Fridlyand J, Mao JH, Yu M, Miller MA, Santos JL, Kalloger SE, Carlson JW, Ginzinger DG, et al. Amplification of PVT1 contributes to the pathophysiology of ovarian and breast cancer. Clin Cancer Res. 2007;13:5745-55.
152. Riquelme E, Suraokar MB, Rodriguez J, Mino B, Lin HY, Rice DC, Tsao A, Wistuba II. Frequent coamplification and cooperation between C-MYC and PVT1 oncogenes promote malignant pleural mesothelioma. J Thorac Oncol. 2014;9:998-1007.
153. Tseng YY, Moriarity BS, Gong W, Akiyama R, Tiwari A, Kawakami H, Ronning P, Reuland B, Guenther K, Beadnell TC, Essig J, Otto GM, O’Sullivan MG, et al. PVT1 dependence in cancer with MYC copy-number increase. Nature. 2014;512:82-6.
154. Wan L, Sun M, Liu GJ, Wei CC, Zhang EB, Kong R, Xu TP, Huang MD, Wang ZX. Long Noncoding RNA PVT1 Promotes Non-Small Cell Lung Cancer Cell Proliferation through Epigenetically Regulating LATS2 Expression. Mol Cancer Ther. 2016;15:1082-94.
155. Li Y, Pei J, Xia H, Ke H, Wang H, Tao W. Lats2, a putative tumor suppressor, inhibits G1/S transition. Oncogene. 2003;22:4398-405.
156. Ke H, Pei J, Ni Z, Xia H, Qi H, Woods T, Kelekar A, Tao W. Putative tumor suppressor Lats2 induces apoptosis through downregulation of Bcl-2 and Bcl-x(L). Exp Cell Res. 2004;298:329-38.
157. Huppi K, Volfovsky N, Runfola T, Jones TL, Mackiewicz M, Martin SE, Mushinski JF, Stephens R, Caplen NJ. The identification of microRNAs in a genomically unstable region of human chromosome 8q24. Mol Cancer Res. 2008;6:212-21.
158. Barsotti AM, Beckerman R, Laptenko O, Huppi K, Caplen NJ, Prives C. p53-Dependent induction of PVT1 and miR-1204. J Biol Chem. 2012;287:2509-19.
159. Dang W, Qin Z, Fan S, Wen Q, Lu Y, Wang J, Zhang X, Wei L, He W, Ye Q, Yan Q, Li G, Ma J. miR-1207-5p suppresses lung cancer growth and metastasis by targeting CSF1. Oncotarget. 2016; 7:32421–32. https://doi.org/10.18632/oncotarget.8718.
160. Kruiswijk F, Labuschagne CF, Vousden KH. p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol. 2015;16:393-405.
161. Chen W, Jiang T, Wang H, Tao S, Lau A, Fang D, Zhang DD. Does Nrf2 contribute to p53-mediated control of cell survival and death? Antioxid Redox Signal. 2012;17:1670-5.
162. Carramusa L, Contino F, Ferro A, Minafra L, Perconti G, Giallongo A, Feo S. The PVT-1 oncogene is a Myc protein target that is overexpressed in transformed cells. J Cell Physiol. 2007;213:511-8.
163. Xu X, Rock JR, Lu Y, Futtner C, Schwab B, Guinney J, Hogan BL, Onaitis MW. Evidence for type II cells as cells of origin of K-Ras-induced distal lung adenocarcinoma. Proc Natl Acad Sci USA. 2012;109:4910-5.
164. Zhao W, Luo J, Jiao S. Comprehensive characterization of cancer subtype associated long non-coding RNAs and their clinical implications. Sci Rep. 2014;4:6591.
165. Wang ZL, Li B, Piccolo SR, Zhang XQ, Li JH, Zhou H, Yang JH, Qu LH. Integrative analysis reveals clinical phenotypes and oncogenic potentials of long non-coding RNAs across 15 cancer types. Oncotarget. 2016; 7:35044–55. https://doi.org/10.18632/oncotarget.9037.
166. Wang Y, Li Z, Zheng S, Zhou Y, Zhao L, Ye H, Zhao X, Gao W, Fu Z, Zhou Q, Liu Y, Chen R. Expression profile of long non-coding RNAs in pancreatic cancer and their clinical significance as biomarkers. Oncotarget. 2015; 6:35684–98. https://doi.org/10.18632/oncotarget.5533.
167. Ferguson BD, Liu R, Rolle CE, Tan YH, Krasnoperov V, Kanteti R, Tretiakova MS, Cervantes GM, Hasina R, Hseu RD, Iafrate AJ, Karrison T, Ferguson MK, et al. The EphB4 receptor tyrosine kinase promotes lung cancer growth: a potential novel therapeutic target. PLoS One. 2013;8:e67668.
168. Zheng MF, Ji Y, Wu XB, Ye SG, Chen JY. EphB4 gene polymorphism and protein expression in non-small-cell lung cancer. Mol Med Rep. 2012;6:405-8.
169. Xiong Z, Liu JK, Zhou ML, Hu CP, Zhou H, Chen W. Relationship between multi-slice spiral CT pulmonary perfusion imaging and the expression of EphB4 and ephrinB2 in non-small cell lung cancer. [Article in Chinese] Zhonghua Zhong Liu Za Zhi. 2011; 33:192–96.
170. Chen J, Sun M, Kent WJ, Huang X, Xie H, Wang W, Zhou G, Shi RZ, Rowley JD, Shi RZ, Rowley JD. Over 20% of human transcripts might form sense-antisense pairs. Nucleic Acids Res. 2004;32:4812-20.
171. Katayama S, Tomaru Y, Kasukawa T, Waki K, Nakanishi M, Nakamura M, Nishida H, Yap CC, Suzuki M, Kawai J, Suzuki H, Carninci P, Hayashizaki Y, et al, and FANTOM Consortium. Antisense transcription in the mammalian transcriptome. Science. 2005;309:1564-6.
172. Werner A, Cockell S, Falconer J, Carlile M, Alnumeir S, Robinson J. Contribution of natural antisense transcription to an endogenous siRNA signature in human cells. BMC Genomics. 2014;15:19.
173. Wu J, Xie H. Expression of long noncoding RNA-HOX transcript antisense intergenic RNA in oral squamous cell carcinoma and effect on cell growth. Tumour Biol. 2015;36:8573-8.
174. Magistri M, Faghihi MA, St Laurent G, Wahlestedt C. Regulation of chromatin structure by long noncoding RNAs: focus on natural antisense transcripts. Trends Genet. 2012;28:389-96.
175. Chen LL, Carmichael GG. Decoding the function of nuclear long non-coding RNAs. Curr Opin Cell Biol. 2010;22:357-64.
176. Mahlapuu M, Ormestad M, Enerbäck S, Carlsson P. The forkhead transcription factor Foxf1 is required for differentiation of extra-embryonic and lateral plate mesoderm. Development. 2001;128:155-66.
177. Grote P, Wittler L, Hendrix D, Koch F, Währisch S, Beisaw A, Macura K, Bläss G, Kellis M, Werber M, Herrmann BG. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell. 2013;24:206-14.
178. Wang X, Balli D, Le T, Kalinichenko VV, Kalin TV. Abstract A60: Foxf1 transcription factor drives progression of benign lung adenomas to malignant adenocarcinomas. Cancer Res. 2013.
179. Saito RA, Micke P, Paulsson J, Augsten M, Peña C, Jönsson P, Botling J, Edlund K, Johansson L, Carlsson P, Jirström K, Miyazono K, Ostman A. Forkhead box F1 regulates tumor-promoting properties of cancer-associated fibroblasts in lung cancer. Cancer Res. 2010;70:2644-54.
180. Xu TP, Huang MD, Xia R, Liu XX, Sun M, Yin L, Chen WM, Han L, Zhang EB, Kong R, De W, Shu YQ. Decreased expression of the long non-coding RNA FENDRR is associated with poor prognosis in gastric cancer and FENDRR regulates gastric cancer cell metastasis by affecting fibronectin1 expression. J Hematol Oncol. 2014;7:63.
181. Bae YK, Kim A, Kim MK, Choi JE, Kang SH, Lee SJ. Fibronectin expression in carcinoma cells correlates with tumor aggressiveness and poor clinical outcome in patients with invasive breast cancer. Hum Pathol. 2013;44:2028-37.
182. Grote P, Herrmann BG. The long non-coding RNA Fendrr links epigenetic control mechanisms to gene regulatory networks in mammalian embryogenesis. RNA Biol. 2013;10:1579-85.
183. Song IS, Ha GH, Kim JM, Jeong SY, Lee HC, Kim YS, Kim YJ, Kwon TK, Kim NS. Human ZNF312b oncogene is regulated by Sp1 binding to its promoter region through DNA demethylation and histone acetylation in gastric cancer. Int J Cancer. 2011;129:2124-33.
184. Song IS, Oh NS, Kim HT, Ha GH, Jeong SY, Kim JM, Kim DI, Yoo HS, Kim CH, Kim NS. Human ZNF312b promotes the progression of gastric cancer by transcriptional activation of the K-ras gene. Cancer Res. 2009;69:3131-9.
185. Chen N, Guo D, Xu Q, Yang M, Wang D, Peng M, Ding Y, Wang S, Zhou J. Long non-coding RNA FEZF1-AS1 facilitates cell proliferation and migration in colorectal carcinoma. Oncotarget. 2016; 7:11271–83. https://doi.org/10.18632/oncotarget.7168.
186. Wang P, Guo J, Wang F, Shi T, Ma D. Human SBK1 is dysregulated in multiple cancers and promotes survival of ovary cancer SK-OV-3 cells. Mol Biol Rep. 2011;38:3551-9.
187. Shannan B, Watters A, Chen Q, Mollin S, Dörr M, Meggers E, Xu X, Gimotty PA, Perego M, Li L, Benci J, Krepler C, Brafford P, et al. PIM kinases as therapeutic targets against advanced melanoma. Oncotarget. 2016; 7:54897–912. https://doi.org/10.18632/oncotarget.10703.
188. Fiskus W, Mitsiades N. B-Raf Inhibition in the Clinic: Present and Future. Annu Rev Med. 2016;67:29-43.
189. Zhou B, Francis TA, Yang H, Tseng W, Zhong Q, Frenkel B, Morrisey EE, Ann DK, Minoo P, Crandall ED, Borok Z. GATA-6 mediates transcriptional activation of aquaporin-5 through interactions with Sp1. Am J Physiol Cell Physiol. 2008;295:C1141-50.
190. Zhang Y, Goss AM, Cohen ED, Kadzik R, Lepore JJ, Muthukumaraswamy K, Yang J, DeMayo FJ, Whitsett JA, Parmacek MS, Morrisey EE. A Gata6-Wnt pathway required for epithelial stem cell development and airway regeneration. Nat Genet. 2008;40:862-70.
191. Herriges MJ, Swarr DT, Morley MP, Rathi KS, Peng T, Stewart KM, Morrisey EE. Long noncoding RNAs are spatially correlated with transcription factors and regulate lung development. Genes Dev. 2014;28:1363-79.
192. Park SM, Park SJ, Kim HJ, Kwon OH, Kang TW, Sohn HA, Kim SK, Moo Noh S, Song KS, Jang SJ, Sung Kim Y, Kim SY. A known expressed sequence tag, BM742401, is a potent lincRNA inhibiting cancer metastasis. Exp Mol Med. 2013;45:e31.
193. Cheung WK, Zhao M, Liu Z, Stevens LE, Cao PD, Fang JE, Westbrook TF, Nguyen DX. Control of alveolar differentiation by the lineage transcription factors GATA6 and HOPX inhibits lung adenocarcinoma metastasis. Cancer Cell. 2013;23:725-38.
194. Li W, Wang Y, Zhang Q, Tang L, Liu X, Dai Y, Xiao L, Huang S, Chen L, Guo Z, Lu J, Yuan K. MicroRNA-486 as a Biomarker for Early Diagnosis and Recurrence of Non-Small Cell Lung Cancer. PLoS One. 2015;10:e0134220.
195. Tian X, Xu G. Clinical value of lncRNA MALAT1 as a prognostic marker in human cancer: systematic review and meta-analysis. BMJ Open. 2015;5:e008653.
196. Weber DG, Johnen G, Casjens S, Bryk O, Pesch B, Jöckel KH, Kollmeier J, Brüning T. Evaluation of long noncoding RNA MALAT1 as a candidate blood-based biomarker for the diagnosis of non-small cell lung cancer. BMC Res Notes. 2013;6:518.
197. Stueve TR, Marconett CN, Zhou B, Borok Z, Laird-Offringa IA. The importance of detailed epigenomic profiling of different cell types within organs. Epigenomics. 2016;8:817-29.
198. Liu Z, Sun M, Lu K, Liu J, Zhang M, Wu W, De W, Wang Z, Wang R. The long noncoding RNA HOTAIR contributes to cisplatin resistance of human lung adenocarcinoma cells via downregualtion of p21(WAF1/CIP1) expression. PLoS One. 2013;8:e77293.
199. Fan T, Jiang S, Chung N, Alikhan A, Ni C, Lee CC, Hornyak TJ. EZH2-dependent suppression of a cellular senescence phenotype in melanoma cells by inhibition of p21/CDKN1A expression. Mol Cancer Res. 2011;9:418-29.
200. Momparler RL, Côté S. Targeting of cancer stem cells by inhibitors of DNA and histone methylation. Expert Opin Investig Drugs. 2015;24:1031-43.
201. Kole R, Krainer AR, Altman S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat Rev Drug Discov. 2012;11:125-40.

