INTRODUCTION
A recent comprehensive analysis of RNA sequencing data from human tissues identified nearly 60,000 long non-coding RNAs (lncRNAs), 3 times the number of protein-coding mRNAs [1]. LncRNAs represent a large family of evolutionarily conserved non-coding RNAs, about 200 nucleotides in length [2]. Recently, some studies demonstrated that some lncRNAs may encode small peptides, probably because minimum cutoff previously used for a classic open reading frame prediction was 100 codons [3–6]. LncRNAs play crucial roles in diverse biological pathways and cellular processes including embryonic development, cell growth and carcinogenesis by affecting gene expression at the transcriptional, post-transcriptional and translational levels [7]. LncRNAs function in either nuclei and/or cytoplasm. In the nucleus, lncRNAs are involved in chromatin remodeling and modification, transcriptional modulation, or RNA processing; in the cytoplasm, lncRNAs interact with mature RNAs and/or proteins [8]. In the last decade, many studies have shown that lncRNAs play important roles as oncogenes or tumor suppressors [9]. Several lncRNAs such as HOTAIR, MALAT1, GAS5, PC3, and H19 are aberrantly regulated in various malignant tumors and associated with carcinogenesis, metastasis, and prognosis [10–14]. The Food and Drug Administration (FDA) approved urine biomarker lncRNA PCA3 is a more specific and sensitive biomarker for prostate cancer compared to serum prostate-specific antigen [13, 15]. Also, phase I/II clinical trials are underway in patients with bladder, pancreatic and ovarian cancers for lncRNA H19 targeted therapy based on in vitro experiments demonstrating tumor cell growth arrest without affecting normal cells with H19 promoter cloned into BC-819 plasmid [16–18]. Hence, lncRNAs have shown immense potential as diagnostic and prognostic biomarkers and therapeutic targets in malignant tumors.
The lncRNA UCA1 was first cloned and identified from bladder cancer cell line BLZ-211 [19]. It is located on human chromosome 19p13.12 positive strand and has three exons and two introns with multiple stop codons without any conserved long open reading frames (ORFs) [19]. The three transcriptional isoforms of UCA1 are 1.4 kb, 2.2 kb and 2.7 kb long, generated by splicing and polyadenylated. The 1.4 kb isoform is denoted as lncRNA UCA1, while the 2.2 kb isoform is denoted lncRNA UCA1a or lncRNA CUDR; the biological role of the 2.7 kb isoform is not known [20]. LncRNA UCA1 is the most abundant isoform in various malignant tumors such as bladder cancer, breast cancer and hepatocellular carcinoma [19, 21, 22, 23]. Hence, most studies have mainly focused on the expression, regulation and functional significance of lncRNA UCA1.
Ectopic expression of lncRNA UCA1 in bladder cancer cell line BLS-211 promoted cancer progression demonstrating that lncRNA UCA1 was oncogenic [21]. Several recent studies have also identified oncogenic functions of lncRNA UCA1 in various cancers such as bladder cancer, breast cancer, colorectal cancer, esophageal squamous cell carcinoma, gastric cancer, hepatocellular carcinoma, melanoma, ovarian cancer, and tongue squamous cell carcinoma [19–29]. Besides the oncogenic function, lncRNA UCA1 also regulates drug resistance in multiple types of malignant tumors. In this article, we review the pivotal roles of the lncRNA UCA1 in drug resistance and discuss the potential future clinical applications.
LncRNA UCA1 expression in various cancers
LncRNA UCA1 is upregulated at 5–10 weeks of gestation; after 28 weeks of gestation, it is highly expressed in bladder, heart and uterus compared to cervix, kidney, liver, lung, intestine, skin, spleen and stomach; after birth, it is turned off in most tissues except heart and spleen [19, 21]. Moreover, lncRNA UCA1 is reactivated in various malignant tumors including bladder cancer, breast cancer, cervical cancer, colorectal cancer, esophageal squamous cell carcinoma, hepatocellular carcinoma, gastric cancer, lung cancer, melanoma, ovarian cancer, thyroid cancer, tongue squamous cell carcinoma, prostate cancer, acute myeloid leukemia, pancreatic cancer, glioma and osteosarcoma [19, 21–33].
The subcellular localization data indicates that lncRNA UCA1 is mainly located in cytoplasm suggesting that it interacts with mature RNAs and/or proteins and regulates them [34]. Interestingly, lncRNA UCA1 is also detected in blood and urine samples from bladder cancer patients and is a useful circulating biomarker [35]. Its detection in urine samples is an effective and noninvasive mode of diagnosing bladder cancer with high sensitivity (100%) and specificity (67%) in combination with the standard cytology method [36]. However, the underlying mechanisms regarding its release from cancer cells into circulation remain to be elucidated. Extracellular vesicles such as exosomes and microvesicles carry coding and non-coding RNAs, proteins and lipids through plasma membranes from various cell types [37]. A recent in vitro study reported that incubation of exosomes with lncRNA UCA1 from the tamoxifen-resistant breast cancer cell line LCC2 with the tamoxifen-sensitive breast cancer cell line MCF-7 resulted in acquired drug resistance [38].
LncRNA UCA1 and regulation
LncRNAs are transcriptionally regulated similar to mRNAs [39, 40]. LncRNA UCA1 expression is regulated by transcription factors, Ets-2 [41], C/EBPα [42], HIF-1α [43], SATB1 [44] and transcriptional complexes, TAZ/YAP/TEAD/SMAD2/3 [45], CAPER/TBX3 [46]. Transcription factors Ets-2 [41] and C/EBPα [42] upregulate lncRNA expression in bladder cancer cells by directly binding to the core promoters and enhancing UCA1 promoter activity (Table 1). Transcription factor HIF-1α enhances lncRNA UCA1 overexpression in bladder and ovarian cancer cells in a hypoxia-dependent manner by directly binding to the hypoxia response elements (HREs) of UCA1 promoter [43]. The transcriptional complex is composed of Hippo pathway effectors (TAZ/YAP/TEAD) and members of the transforming growth factor-β (TGF-β) pathway (SMAD2/3) that are recruited to the UCA1 promoter and synergistically induce its expression in breast cancer cells after TGF-β treatment [45, 47]. Conversely, transcription factor SATB1 represses lncRNA UCA1 expression in breast cancer cells by directly binding to the promoter and 3.0 kb upstream regions of UCA1 and closing the chromatin structure, thereby blocking transcription [44]. The transcriptional complex composed of coactivator of activating protein-1 and estrogen receptors (CAPERα)/T-box 3 (TBX3) inhibits lncRNA UCA1 by binding to its promoter in cultured primary human foreskin fibroblasts (HFFs) [46].
Table 1: Regulators of lncRNA UCA1 expression
Regulators |
Roles on lncRNA UCA1 |
regulation level |
References |
|---|---|---|---|
Ets-2 |
upregulate expression |
transcriptional |
Wu w, et al. [41] |
C/EBPα |
upregulate expression |
transcriptional |
Xue M, et al. [42] |
TAZ/YAP/TEAD |
upregulate expression |
transcriptional |
Hiemer SE, et al. [45] |
HIF-1α |
upregulate expression |
transcriptional |
Xue M, et al. [43] |
SATB1 |
downregulate expression |
transcriptional |
Lee JJ, et al. [44] |
CAPERα/TBX3 |
downregulate expression |
transcriptional |
Kumar PP, et al. [46] |
miR-1 |
downregulate expression |
post-transcriptional |
Wang T, et al. [48] |
The expression of lncRNA UCA1 is also regulated at post-transcriptional level. A recent study reported that knockdown of miR-1 resulted in Ago2-slicer-dependent lncRNA UCA1 overexpression, whereas miR-1 overexpression decreased UCA1 levels in bladder cancer cells [48]. Wang et al. postulated that miR-1 targets and cleaves lncRNA UCA1 in an Ago2-dependent manner similar to degradation of protein-coding mRNAs by miRNAs [48].However, this hypothesis needs to be experimentally proven.
LncRNA UCA1 and prognosis
The prognostic value of lncRNA UCA1 in various human malignant tumors has been evaluated by many studies. Liu et al. conducted a meta-analysis of 1,240 patients from 15 articles on colorectal cancer, esophageal squamous cell carcinoma, prostate cancer, hepatocellular carcinoma, non-small cell lung cancer, gastric cancer, ovarian cancer and concluded that lncRNA UCA1 was an independent prognostic factor for OS in cancer patients [49]. The meta-analysis showed that higher expression of lncRNA UCA1 (HR = 1.71, 95% CI: 1.43–1.99) was associated with poor OS based on cancer type, cut-off value, analysis type and sample size. Furthermore, the pooled odds ratios (ORs) showed that elevated lncRNA UCA1 was associated with lymph node metastasis (OR = 2.98, 95% CI: 2.06–4.30), distant metastasis (OR = 3.14, 95% CI: 1.77–5.58) and poor clinical stage (OR = 2.76, 95% CI: 2.08–3.68) [49]. However, larger-size, multi-center and higher-quality studies are necessary to validate the results in this study. Higher lncRNA UCA1 expression was also reported as an independent prognostic biomarker for pancreatic cancer [32] and glioma [33]. The poor prognosis in cancer therapy is mainly due to metastasis and drug resistance.
Direct binding targets of LncRNA UCA1
The molecular mechanisms for most lncRNAs remain to be fully elucidated. They directly bind to DNA, RNA, or proteins, and many techniques have been developed to identify each of these interactions. LncRNA-DNA interactions can be identified by Capture Hybridization Analysis of RNA Targets (CHART) [50] and Chromatin Isolation by RNA Purification (ChIRP) [51] that involve hybridization of complementary oligonucleotides to the lncRNA of interest and serve as an affinity handle to recruit bound DNA. Similarly, radioimmunoassay sequencing (RIA-seq) is used to assay lncRNA-RNA interactions [52]. Techniques such as RNA immunoprecipitation (RIP) or its variants, cross-linking immunoprecipitation (CLIP) [53, 54], and photoactivatable ribonucleoside-enhanced cross-linking immunoprecipitation (PAR-CLIP) [55] utilize antibodies to reveal lncRNA-protein interactions. Furthermore, high-throughput methods such as in vivo click selective 2′ hydroxyl acylation and profiling experiment (icSHAPE) [56] or selective 2′-hydroxyl acylation analyzed by primer extension sequencing (SHAPE-seq) [57], have been developed in recent years to explore RNA secondary structure.
LncRNA UCA1 has also been reported to bind to several miRNAs in different cancer cells. These include miR-216b in hepatocellular cancer [22], miR-193a in non-small cell lung cancer [58], miR-145 in bladder cancer [59], miR-204 in colorectal cancer [34] and esophageal cancer [60], miR-1 in bladder cancer [48], miR-18a in breast cancer cells [61], miR-16 in CML [62] (Table 2).
Table 2: Direct binding targets of lncRNA UCA1
Direct binding targets |
Tumors |
Drug resistance |
References |
Published journals and years |
|---|---|---|---|---|
miR-216b |
HCC |
N/A |
Wang F, et al. [22] |
Oncotarget.2015 |
miR-193a |
NSCLC |
N/A |
Nie W, et al. [58] |
Cancer Lett.2016 |
miR-145 |
bladder cancer |
N/A |
Xue M, et al. [59] |
Cancer Sci.2015 |
miR-204 |
colorectal cancer |
YES |
Bian Z, et al. [34] |
Sci Rep.2016 |
miR-204 |
esophageal cancer |
N/A |
Jiao C, et al. [60] |
Oncol Rep.2016 |
miR-18a |
breast cancer |
YES |
Li X, et al. [61] |
Tumour Biol.2016 |
miR-16 |
CML |
YES |
Xiao Y, et al. [62] |
DNA AND CELL BIOLOGY.2016 |
miR-1 |
bladder cancer |
N/A |
Wang T, et al. [48] |
Tumour Biol.2014 |
LncRNAs play the role of a signal, decoy, guide or scaffold by binding to DNAs, RNAs and proteins [63]. These mechanisms are not mutually exclusive and lncRNAs may play one or more of the roles. For example, lncRNA UCA1 plays the role of a decoy by acting as a sponge and sink for miRNAs, which results in promoting carcinogenesis or drug resistance.
LncRNA UCA1 in cancer drug resistance
Systemic or local chemotherapy is widely used in cancer therapy. However, intrinsic and acquired drug resistance reduces the efficacy of cancer therapies. Although remarkable progress has been made in understanding the mechanisms underlying anti-cancer drug resistance, many questions still remain unanswered. The cancer cells employ different mechanisms to effectively escape chemotherapy [64]. These include aberrant expression of glutathione transferase and topoisomerase II, reduced uptake of water-soluble drugs, enhanced DNA damage repair, enhanced drug metabolism, decreased apoptosis, and increased energy-dependent efflux of chemotherapeutic drugs, all of which reduce the effects of the cancer therapeutics [65]. In recent years, the role of lncRNAs in non-mutational regulation of gene function has been described [66]. LncRNAs bind to various writers, erasers, readers of histone modifications, and other chromatin regulatory factors [67]. LncRNA UCA1 is a good example of the role of lncRNAs in drug resistance.
LncRNA UCA1 and bladder cancer drug resistance
Bladder cancer is ranked fourth among males and tenth among females in regard to leading causes of deaths worldwide [68]. Cisplatin and gemcitabine are the preferred drugs for chemotherapy after surgery for muscle-invasive bladder cancer patients and drug resistance greatly impedes long-term survival [69]. The molecular mechanisms promoting resistance to cisplatin/gemcitabine-based chemotherapy are only recently being identified.
In 2008, Wang et al. showed that transfection of a 1.4 kb full-length lncRNA UCA1 cDNA strongly promoted the proliferation, motility and invasiveness of the bladder TCC cell line, BLS-211 [21]. Stable UCA1 overexpression in BLS-211 resulted in drug resistance to cisplatin and mitomycin with their IC50 increased by 2.4 and 1.9 fold, respectively. Microarray analysis identified 16 upregulated and 26 downregulated genes. The genes were related to tumorigenesis and/or embryonic development with WNT6, CYP1A1, and AURKC among the upregulated genes, and MBD3 and SRPK1 among the downregulated genes. Fan et al. established a cisplatin-resistant T24-resistance bladder cancer cell line by continuous exposure of the T24 bladder cancer cell line to cisplatin (2.0 μg/ml) over 12 months [70]. The expression of lncRNA UCA1 was higher in the T24-cisplatin resistant cells than T24 cells [70]. Forced expression of lncRNA UCA1 in T24 cells decreased cisplatin-induced cell death, whereas knockdown of lncRNA UCA1 in T24-resistance cells increased cell death [70]. In vitro analysis revealed that lncRNA UCA1 increased Wnt6 expression and subsequent activation of the Wnt signaling pathway, consistent with previous findings [70]. Higher lncRNA UCA1 levels were observed in blood samples of advanced bladder cancer patients after cisplatin-based chemotherapy with a positive correlation between UCA1 and Wnt6 mRNA expression levels in the bladder cancer tissues [70]. Recently, Pan et al. demonstrated that forced expression of lncRNA UCA1 in the bladder cancer cell line UMUC-2 reduced apoptosis upon cisplatin/gemcitabine treatment, whereas silencing of lncRNA UCA1 in the bladder cancer cell line 5637 increased cellular apoptosis [71]. Both in vitro and in vivo experiments confirmed that lncRNA UCA1 induced cisplatin/gemcitabine resistance through activation of CREB by p-AKT and subsequent upregulation of miR-196a-5p [71]. Moreover, expression of lncRNA UCA1, miR-196a-5p and p-CREB was positively correlated in bladder cancer tissues [71].
LncRNA UCA1 and ovarian cancer drug resistance
Ovarian cancer is ranked first in developed countries and second in developing countries as the cause of gynecological cancer-related deaths worldwide [72]. Although platinum-based chemotherapy is effective in ovarian cancer treatment, primary or secondary resistance to cisplatin severely affects therapeutic efficacy [73]. The overall 5-year survival rate of ovarian cancer is less than 50% [72]. Hence, there is a great need for discovering predictive biomarkers for ovarian cancer.
In 2015, Wang et al. found that lncRNA UCA1 and SRPK1 mRNA was substantially upregulated in ovarian cancer tissues compared to normal ovarian tissues [24]. Stable transfection of pcDNA-UCA1 in SKOV3 cells contributes to cisplatin resistance with a 2.41 fold increase in IC50, increased SRPK1 and Bcl-2 and decreased Bax, Caspase-3 and Caspase-9 expression. However, knockdown of SRPK1 overcomes cisplatin resistance in SKOV3-pcDNA-UCA1 cells and increased Bcl-2 and decreased Bax, Caspase-3 and Caspase-9 expression is observed. These results revealed that SRPK1 and apoptosis related proteins played an important role in the lncRNA UCA1 induced cisplatin resistance [24]. Furthermore, Zhang et al. observed that lncRNA UCA1 levels were significantly higher in ovarian cancer tissues compared to normal ovarian tissues, and high UCA1 expression was associated with more lymph node metastasis, advanced FIGO stage, and bad response to platinum-based chemotherapy [74].
LncRNA UCA1 and breast cancer drug resistance
Breast cancer is the most commonly diagnosed malignant tumor among women worldwide with nearly 70% of it being estrogen receptor positive [75]. Tamoxifen is one of the major hormone therapies for ER positive breast cancer in clinical practice. However, tamoxifen resistance limits the long-term effects of the treatment of ER positive breast cancer. Hence, it is necessary to identify the molecular mechanisms underlying tamoxifen resistance.
Wu et al. showed that knockdown of lncRNA UCA1 in tamoxifen resistant LCC2 and LCC9 cells increased apoptosis upon tamoxifen treatment accompanied by significant reduction in p-AKT and p-mTOR [76]. However, forced expression of lncRNA UCA1 in tamoxifen-sensitive MCF-7 cells substantially reduced tamoxifen induced apoptosis, which could be abrogated by the mTOR specific inhibitor rapamycin [76]. These results revealed that lncRNA UCA1 induced tamoxifen resistance in breast cancer cells partly through activation of the mTOR signal pathway [76]. Further, Li et al. showed that ectopic expression of lncRNA UCA1 in MCF-7 induced acquired tamoxifen resistance [61]. In addition, functional analysis demonstrated that lncRNA UCA1 acted as a sponge for miR-18a, which is an important modulator of tamoxifen resistance by regulating cell cycle proteins. Interestingly, tamoxifen treatment upregulated lncRNA UCA1 in MCF-7 cells through a miR-18a-HIF1α feedback loop [61]. Moreover, Liu et al. showed that lncRNA UCA1 expression was upregulated in the tamoxifen resistant breast cancer cells compared to their parental MCF-7 and T47D cells [77]. Stable transfection of lncRNA UCA1 in MCF-7 and T47D cells resulted in drug resistance to tamoxifen, whereas siRNA targeting lncRNA UCA1 enhanced tamoxifen sensitivity. Also, knockdown of lncRNA UCA1 prevented the nuclear translocation of β-catenin, thereby inhibiting the Wnt signaling pathway [77]. This suggested that lncRNA UCA1 induced tamoxifen resistance by increasing the activity of Wnt/β-Catenin signaling [77]. Further, Xu et al. demonstrated that treatment of MCF-7 cells with exosomes released from tamoxifen resistant LCC2 cells with lncRNA UCA1 resulted in enhanced cell viability accompanied by decreased expression of cleaved caspase-3 after tamoxifen treatment [38]. Further, LCC2 exosomes with impaired UCA1 could not induce tamoxifen resistance in MCF-7 cells. These results showed that exosome-mediated transfer of lncRNA UCA1 induced acquired tamoxifen resistance in MCF-7 cells [38].
LncRNA UCA1 and lung cancer drug resistance
Currently, EGFR-TKIs like gefitinib or erlotinib are the first-line treatments against advanced non-small cell lung cancer (NSCLC) harboring EGFR-activating mutations [78, 79, 80]. However, the majority of EGFR-mutated NSCLCs that respond to initial EGFR-TKI therapy develop acquired resistance in about 12 months [81, 82]. Functional analysis indicated that a secondary T790M mutation is the major cause of acquired resistance ; c-MET amplification, PIK3CA mutations (~5%), BRAF mutations and small-cell lung cancer transformation were also associated with acquired resistance [83–85].
Cheng et al. performed microarray expression profiling of lncRNAs/mRNA and demonstrated high lncRNA UCA1 expression in acquired gefitinib-resistant PC9/R cells compared to the parental gefitibib-sensitive PC9 cells [86]. Further, Cheng et al. demonstrated higher lncRNA UCA1 expression in PC9/R and H1975 cells with acquired resistance compared to A549, H460, H23 and H1299 cells with primary resistance [87]. The lncRNA UCA1 levels were higher in lung cancer tissues from patients with acquired resistance to EGFR-TKIs compared to EGFR-TKI sensitive patients or patients with primary resistance to EGFR-TKIs [87]. Functional analysis showed that knockdown of UCA1 in PC9/R cells without T790M mutation partly restored gefitinib sensitivity with increased expression of caspase 3 and caspase 8, whereas H1975 cells with T790M mutation remained gefitinib resistant. Moreover, both in vitro and in vivo analysis confirmed that lncRNA UCA1 contributed to non-T790M acquired resistance to EGFR-TKIs by activating the AKT/mTOR pathway and EMT [87].
LncRNA UCA1 and gastric cancer drug resistance
Gastric cancer is the most frequently diagnosed cancer of the digestive system with high morbidity and mortality [88]. For advanced gastric cancer, both pre-operative and post-operative chemotherapies are critical for survival [89]. However, the underlying mechanisms are unclear.
In 2016, Fang et al. showed increased lncRNA UCA1 and decreased miR-27b in MDR gastric cancer cells such as SGC-7901/ADR, SGC-7901/DDP, and SGC- 7901/5-FU cells compared to their parental SGC-7901 cells, whereas knockdown of lncRNA UCA1 substantially restored miR-27b expression [90]. Bioinformatic analysis demonstrated two possible binding sites of mirR-27b in lncRNA UCA1 and in vivo analysis of gastric cancer tissues confirmed the negative correlation between lncRNA UCA1 and miR-27b expression [90]. Furthermore, lncRNA UCA1 silencing and miR-27b overexpression overcame the drug resistance by reducing the IC50 of ADR, DDP, and 5-FU in SGC-7901/ADR cells, which was accompanied with enhanced apoptosis due to decreased Bcl-2 and increased cleaved caspase-3 levels [90]. In addition, Shang et al. showed that lncRNA UCA1 levels were higher in SGC7901/ADR cells compared to the parental SGC7901 cells and this resulted in a 4.3 fold increase in IC50 after adriamycin treatment [91]. However, knockdown of lncRNA UCA1 suppressed this resistance to adriamycin in SGC7901/ADR and increased cellular apoptosis upon adriamycin treatment due to increased cleaved PARP and decreased Bcl-2 [91].
LncRNA UCA1 and colorectal cancer drug resistance
Colorectal cancer is the third most common cancer in males and the second most common cancer in females worldwide [72]. The initiation and progression of CRC is a multi-step process controlled by the deregulation of multiple oncogenes and tumor suppressors [92]. However, diagnostic methods and therapies are needed to improve CRC detection and survival.
Bian et al. showed that ectopic expression of lncRNA UCA1 in SW480 cells resulted in chemoresistance with reduction in 5-Fu induced apoptosis, whereas knockdown of lncRNA UCA1 in HCT116 cells enhanced apoptosis [34]. Bioinformatic analysis showed that lncRNA UCA1 has a miR-204-5p recognition sequence. Previous in vitro results showed that miR-204-5p enhanced chemotherapeutic sensitivity of CRC cells. Luciferase and RIP assays demonstrated direct binding between UCA1 and miR-204-5p [34]. Mechanistically, lncRNA UCA1 acted as a sponge of miR-204-5p and upregulated the expression of miR-204-5p target genes (BCL2, RAB22A and CREB1), thereby contributing to 5-Fu resistance. Moreover, positive correlation between CREB1 and UCA1 and inverse correlation between CREB1 and miR-204-5p was observed in CRC tissues, thereby suggesting that UCA1 acted as a ceRNA of miR-204-5p to regulate CREB1 expression in CRC tumors [34].
LncRNA UCA1 and prostate cancer drug resistance
Prostate cancer is the second most common cancer in males in the world with good prognosis [72]. However, the prognosis of some advanced or metastatic patients is poor and drug resistance is one of the major reasons.
Wang et al. showed that lncRNA UCA1 overexpression downregulated miR-204 expression in LNCaP cells and 22RV1 cells, while miR-204 upregulation substantially decreased Sirt1 expression suggesting that miR-204 inversely modulated Sirt1 expression [30]. In addition, lncRNA UCA1 overexpression increased Sirt1 expression in PNT2 cells, while silencing of endogenous lncRNA UCA1 decreased Sirt1 expression in LNCaP and 22RV1 cells [30]. This suggested a UCA1/miR-204/Sirt1 axis in LNCaP and 22RV1 cells. Interestingly, lncRNA UCA1 and Sirt1 levels were significantly upregulated in 22RV1/DR cells compared to the parental 22RV1 cells and miR-204 was downregulated. Transfection with UCA1 and Sirt1 siRNAs or miR-204 mimics in 22RV1/DR cells significantly decreased the expression of P-glycoprotein (P-gp), which is an important membrane pump responsible for drug resistance [30]. Additionally, reduced docetaxel IC50 in 22RV1/DR cells due to increased cleaved caspase-3 expression and enhanced apoptosis was observed [30]. These results revealed that lncRNA UCA1 induced docetaxel resistance in prostate cancer cells via the UCA1/miR-204/Sirt1 axis.
LncRNA UCA1 and CML drug resistance
To date, imatinib has been a revolutionary treatment against chronic myelogenous leukemia (CML) with about 95% of CML patients achieving remission after imatinib therapy [93, 94]. However, some patients in accelerated phase or blast crisis demonstrate drug resistance to imatinib [94, 95]. The underlying mechanisms of imatinib resistance could be a result of point mutations in the breakpoint cluster region-Abelson murine leukemia (BCR-ABL) kinase domain, amplification of the BCR-ABL gene, and overexpression of the multidrug resistance protein 1 (MRP1) gene in tumor cells [96, 97].
In 2017, Xiao et al. showed that K562/IM cells displayed a 10 fold increase in IC50 compared to a 5 fold increase in IM resistance of K562/IMR cells in relation to the parent K562 cells whereas [62]. Further, MDR1 protein expression was higher in K562/IM-R and lower in K562/IM cells compared to K562 cells. Also, lncRNA UCA1 expression was higher in K562/IM-R and lower in K562/IM cells than in K562 cells, which indicated the role of lncRNA UCA1 in IM resistance of CML cells [63]. Stable transfection of lncRNA UCA1 in K562 cells markedly upregulated MDR1 mRNA and protein levels and resulted in IM resistance, whereas silencing of UCA1 in K562/IM cells significantly inhibited MDR1 expression. Thus, lncRNA UCA1 emerged as a competitive endogenous RNA (ceRNA) of MDR1 that induced IM resistance by sequestering miR-16 [62].
Taken together, these studies confirm the pivotal role of lncRNA UCA1 in anti-cancer drug resistance to cisplatin, gemcitabine, 5-FU, tamoxifen, imatinib and EGFR-TKIs. However, the underlying molecular mechanisms still remain to be investigated in greater detail (Table 3). Therefore, based on current knowledge, we postulate that lncRNA UCA1 promotes drug resistance by directly binding to specific miRNAs and acting as a “sponge”. To date, miR-216b, miR-193a, miR-145, miR-204, miR-1, miR-18a and miR-16 (Table 2) have been shown to directly bind to lncRNAs. Among these, miR-204, miR-18a and miR-16 have been verified to bind to lncRNA UCA1 directly and then induce drug resistance. In future, computer prediction, RIA-seq screening and luciferase and RIP assays should be performed to comprehensively identify specific miRNAs that bind to lncRNA UCA1 specifically. Next, the interactions between lncRNA UCA1 and miRNAs could relieve miRNAs mediated target mRNAs repression. Up to now, there is none concrete target mRNAs reported in these literatures. Maybe, SRPK1, AKT, Wnt6 belong to this kind of target mRNAs. And then, the overexpression of the target proteins results in activation of signaling pathways that decrease apoptosis, enhance proliferation or reduce cell cycle arrest. Based on available data, several signaling pathways like AKT/mTOR/HIF-1a, AKT/CREB and Wnt/β-catenin could be involved in the process (Figure 1). It is a systematic and thorough work to elucidate this network in great need of basic and clinical research.
Table 3: LncRNA UCA1 in anti-cancer drug resistance
Cancer types |
Drugs |
Direct targets |
Mechanisms |
References |
Published Journals and years |
|---|---|---|---|---|---|
bladder cancer |
cisplatin/mitomycin |
N/A |
N/A |
Wang F, et al. [21] |
FEBS Lett.2008 |
bladder cancer |
cisplatin |
N/A |
Wnt6/Wnt pathway |
Fan Y, et al. [70] |
FEBS J.2014 |
bladder cancer |
cisplatin/gemcitabine |
N/A |
p-AKT/CREB/miR-196a-5p/p27Kip1 |
Pan J, et al. [71] |
Cancer Lett.2016 |
breast cancer |
tamoxifen |
N/A |
N/A |
Xu CG, et al. [38] |
Eur Rev Med Pharmacol Sci.2016 |
breast cancer |
tamoxifen |
N/A |
mTOR pathway |
Wu C, et al. [76] |
Med Sci Monit.2016 |
breast cancer |
tamoxifen |
miR-18a |
miR-18a, HIF1α |
Li X, et al. [61] |
Tumour Biol.2016 |
breast cancer |
tamoxifen |
N/A |
Wnt/β-Catenin |
Liu H, et al. [77] |
PLoS One.2016 |
CML |
imatinib |
miR-16 |
miR-16/MDR1 |
Xiao Y, et al. [62] |
DNA Cell Biol.2016 |
colorectal cancer |
5-FU |
miR-204-5p |
miR-204-5p/CREB1,Bcl2,RAB22A |
Bian Z, et al. [34] |
Sci Rep.2016 |
gastric cancer |
doxorubicin/cisplatin/5-FU |
N/A |
miR-27b/Bcl-2,caspase-3 |
Fang Q, et al. [90] |
Med Sci Monit.2016 |
gastric cancer |
adriamycin |
N/A |
Bcl-2,PARP |
Shang C,et al. [91] |
Cancer Chemother Pharmacol.2016 |
lung cancer |
EGFR-TKIs |
N/A |
AKT/mTOR/caspase-3,caspase-8 |
Cheng N,et al. [87] |
Oncotarget.2015 |
ovarian cancer |
cisplatin |
N/A |
SRPK1/Bcl-2,Bax,caspase-3,caspase-9 |
Wang F, et al. [24] |
Neoplasma.2015 |
ovarian cancer |
platinum-based chemotherapy |
N/A |
N/A |
Zhang L,et al. [74] |
Cancer Chemother Pharmacol.2016 |
prostate cancer |
docetaxel |
N/A |
miR-204/Sirt1 |
Wang X,et al. [30] |
Cancer Chemother Pharmacol.2016 |
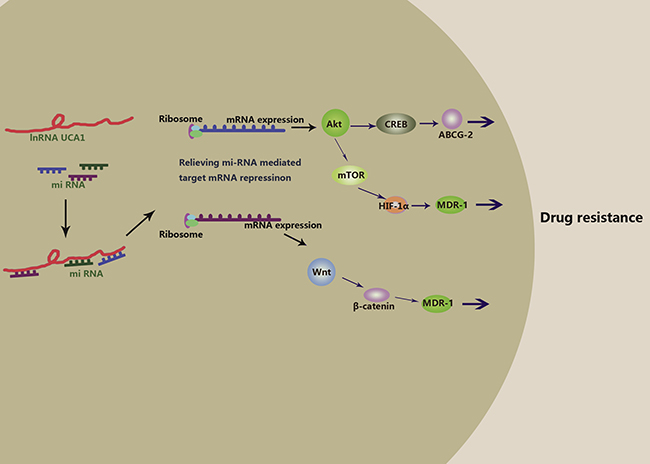
Figure 1: Hypothesis of molecular mechanism of lncRNA UCA1 in anti-cancer drug resistance.
Future clinical applications
LncRNA UCA1 as a cancer biomarker
LncRNAs are promising diagnostic or predictive biomarkers for human cancers because of their specific expression and critical role in cancer initiation and progression. LncRNA UCA1 functions as an oncofetal gene similar to lncRNA H19 [98] and is detected not only in tumor tissue samples but also in blood and urine samples from bladder cancer patients as a circulating biomarker [35]. After birth, lncRNA UCA1 is undetectable in majority of normal tissues [19, 21]. LncRNA UCA1 is reactivated in various malignant tumors including bladder cancer, breast cancer, cervical cancer, colorectal cancer, esophageal squamous cell carcinoma, hepatocellular carcinoma, gastric cancer, lung cancer, melanoma, ovarian cancer, thyroid cancer, tongue squamous cell carcinoma, prostate cancer, acute myeloid leukemia, glioma, pancreatic cancer and osteosarcoma [19–33]. Hence, lncRNA UCA1 in body fluid samples is a potential diagnostic and prognostic biomarker for tumor initiation, metastasis and recurrence of cancers. RT-PCR based lncRNA UCA1 expression detection in urine samples is an effective and noninvasive assay with high sensitivity (100%) and specificity (67%) for the diagnosis of bladder cancer with the standard cytology method [36]. However, the diagnostic value of lncRNA UCA1 in body fluid samples for other types of malignant tumors remains to be confirmed.
The positive correlation between lncRNA UCA1 expression and poor prognosis in a great number of cancer types as mentioned in the above meta-analysis suggests that lncRNA UCA1 could act as an independent prognostic factor for cancer patients [49]. However, larger-size, multi-center and higher-quality studies with a unified criterion for determining lncRNA UCA1 expression are necessary to validate and confirm these results for the different kinds of cancers.
Further, many studies show that lncRNA UCA1 induces drug resistance in various cancers such as bladder cancer, ovarian cancer, breast cancer, lung cancer, gastric cancer, colorectal cancer, prostate cancer and CML [21, 24, 30, 34, 38, 61, 62, 70, 71, 74, 76, 77, 87, 90, 91], thereby greatly reducing the efficacy of cancer therapy. The majority of these results of drug resistance were from in vitro studies, whereas clinical data was available only in ovarian cancers where high lncRNA UCA1 expression was associated with the response to platinum-based chemotherapy [74].
Taken together, lncRNA UCA1 is potentially a good broad spectrum biomarker for cancer diagnosis, prognosis or therapeutic monitoring. However, none of the lncRNA UCA1 products are in clinical trials yet. Therefore, we propose that further animal model studies and prospective clinical studies will highlight and describe the role of lncRNA UCA1 in anti-cancer drug resistance leading to its investigation for various clinical applications in the future.
LncRNA UCA1 related therapies against cancers
Although the role of lncRNAs in cancer drug resistance is only beginning to be discovered, the application potential of lncRNA UCA1 as a candidate to develop novel strategies to reverse drug sensitivity to chemotherapy or molecular targeted therapy cannot be ignored.
LncRNA UCA1 targeted therapeutics (RNAi and ASO)
RNA targeted therapeutics include natural or artificially synthesized oligonucleotides that target mRNA for therapy or elucidate gene functions [99]. Currently, small molecules and antibodies have been used to target proteins, while oligonucleotides target various RNAs [99]. The two main approaches in RNA targeted therapeutics include double stranded RNA-mediated interference (RNAi) and antisense oligonucleotides (ASO). RNAi induces lncRNA degradation, while ASO sterically impedes RNA splicing, formation of functional protein complexes and the formation or function of a mature RNA [100]. Both RNAi and ASO have been verified in clinical trials for many diseases such as cancer and neurodegeneration [99]. Most studies have reported that lncRNA UCA1 is also specifically expressed in various cancer cells and involved in drug resistance [21, 24, 30, 34, 38, 61, 62, 70, 71, 74, 76, 77, 87, 90, 91]. Hence, we postulate that targeting lncRNA UCA1 may partially or completely restore drug sensitivity and improve the efficacy of chemotherapy. In fact, knockdown of lncRNA UCA1 by short interfering RNAs (siRNA) or short hairpin RNA (shRNA) has been shown to reverse drug resistance in various cancer cells such as bladder cancer [70, 71], breast cancer [38, 76, 77], lung cancer [87], gastric cancer [90, 91], colorectal cancer [34], prostate cancer [30], CML [62], ovarian cancer [24]. Also, miR-1 has been shown to suppress lncRNA UCA1 expression in bladder cancer cells [48].
Transcription factor or transcription complex targeted therapeutics
A number of studies have demonstrated that lncRNA UCA1 expression is suppressed by the transcription factor SATB1 [44] and transcriptional complex composed of coactivator of activating protein-1 and estrogen receptors (CAPERα) and T-box 3 (TBX3) [46]. Hence, SATB1 and CAPERα/TBX3 can be potentially exploited to inhibit the expression of lncRNA UCA1 and reverse drug resistance. Apart from these, many studies have indicated that the expression of lncRNA UCA1 is upregulated by transcription factors Ets-2 [41], C/EBPα [42], HIF-1α [43], and transcriptional complexes (TAZ/YAP/TEAD/SMAD2/3) [45]. Thus, Ets-2, C/EBPα, HIF-1α, and transcriptional complexes (TAZ/YAP/TEAD/SMAD2/3) are potential targets to knockdown the expression of lncRNA UCA1.
Signaling pathway and effector molecule targeted therapeutics
Many studies in the last two decades have explored the resistance mechanisms in cancer cells and approaches to reverse drug resistance. These studies have identified the role of proteins such as MDR1, ABCG2, and MRP in cancer drug resistance, and some have been exploited to develop various strategies for cancer therapy. Recently, the role of non-coding RNAs such as microRNAs and lncRNAs has been confirmed in drug resistance [101, 102]. The miRNAs such as miR-204, miR-18a, miR-27b and miR-16 have demonstrated the therapeutic potential to restore drug sensitivity, whereas miRNAs such as miR-196a and proteins such as mTOR, MDR1, P27, BCL2, SRPK1, Sirt1, β-Catenin and Wnt are targets for cancer therapy. Preliminary studies have shown the therapeutic potential of lncRNA UCA1 in cancer therapy, especially drug resistance.
CRISPR/Cas9-based therapeutics
Direct targeting of the UCA1 genomic locus is another method to knockdown UCA1 expression in malignant tumors. Tsui-Ting et al. silenced lncRNA UCA1 expression in HCT-116 human colon cancer cells by using the (CRISPR)/CRISPR-associated (Cas) genome editing system, which involves clustered regularly interspaced short palindromic repeats [103]. Moreover, Zhen et al. designed gRNAs specific to UCA1 and demonstrated significant knockdown of lncRNA UCA1 when transfected into 5637 and T24 bladder cancer cells with CRISPR/Cas9 systems targeting UCA1 [104]. Hence, CRISPR/Cas9 systems are potential tools to inhibit the expression of lncRNA UCA1 at genomic level to attenuate drug resistance in research.
Taken together, potential novel strategies to reverse drug sensitivity to chemotherapy or molecular targeted therapy include lncRNA UCA1 targeted therapeutics, CRISPR/Cas9 system, transcription factor or transcription complex targeted therapeutics, signaling pathway and effector molecule targeted therapeutics.
CONCLUSIONS
LncRNA UCA1 expression is associated with a number of anti-cancer drug resistance tumors, however, the mechanisms remain to be elucidated in greater detail, especially from a clinical standpoint. Yet, lncRNA UCA1 shows great potential as a diagnostic, predictive or prognostic biomarker, and a therapeutic target in malignant tumors.
Abbreviations
lncRNA: long non coding RNA, UCA1: urothelial carcinoma associated 1, UCA1a: urothelial carcinoma associated 1a, ORFs: open reading frames, CUDR: cancer upregulated drug resistant gene, TAZ: Transcriptional coactivator with PDZ-binding motif, YAP: yes-associated protein, TEAD: TEA domain family member, ceRNA: competing endogenous RNA, CAPER/TBX3:coactivator of activating protein-1 and estrogen receptors, C/EBPα: CCAAT/enhancer-binding protein α, HIF-1α: hypoxia-inducible factor 1 α, HREs: hypoxia response elements, TGF-β: transforming growth factor-β, HFFs: human foreskin fibroblasts, RIA-seq: radioimmunoassay sequencing, CHART: capture hybridization analysis of RNA targets, ChIRP: chromatin isolation by RNA purification, RIP: RNA immunoprecipitation, CLIP: cross-linking immunoprecipitation, PAR-CLIP: photoactivatable ribonucleoside–enhanced cross-linking immunoprecipitation, icSHAPE: in vivo click selective 2′ hydroxyl acylation and profiling experiment, SHAPE-seq: selective 2′-hydroxyl acylation analyzed comby primer extension sequencing, BCR-ABL: breakpoint cluster region-Abelson murine leukemia, CML: Chronic Myeloid Leukemia, NSCLC: non-small cell lung cancer, 5-Fu: 5-fluorouracil, ABCB1: ATP binding cassette subfamily B member 1, MRP1: multi-drug resistant associate protein 1, MDR1: multi-drug resistant protein, N/A: not available.
ACKNOWLEDGMENTS
This work was supported by National Natural Science Foundation of China (No.81201089 and No. 81272676) and by grant from National Science and Technology Major Project of the Ministry of Science and Technology of China (No. 2013ZX09506015).
CONFLICTS OF INTEREST
The authors declare no potential conflicts of interest.
REFERENCES
1. Geisler S, Coller J. RNA in unexpected places: long noncoding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol. 2013; 14:699–712.
2. Kogo R, Shimamura T, Mimori K, Kawahara K, Imoto S, Sudo T, Tanaka F, Shibata K, Suzuki A, Komune S, Miyano S, Mori M. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res. 2011; 71:6320–6326.
3. Andrews SJ, Rothnagel JA. Emerging evidence for functional peptides encoded by short open reading frames. Nat Rev Genet. 2014; 15:193–204.
4. Landry CR, Zhong X, Nielly-Thibault L, Roucou X. Found in translation: functions and evolution of a recently discovered alternative proteome. Curr Opin Struct Biol. 2015; 32:74–80.
5. Anderson DM, Anderson KM, Chang CL, Makarewich CA, Nelson BR, McAnally JR, Kasaragod P, Shelton JM, Liou J, Bassel-Duby R, Olson EN. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell. 2015; 160:595–606.
6. Nelson BR, Makarewich CA, Anderson DM, Winders BR, Troupes CD, Wu F, Reese AL, McAnally JR, Chen X, Kavalali ET, Cannon SC, Houser SR, Bassel-Duby R, et al. A peptide encoded by a transcript annotated as long noncoding RNA enhances SERCA activity in muscle. Science. 2016; 351:271–275.
7. Moran VA, Perera RJ, Khalil AM. Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res. 2012; 40:6391–6400.
8. Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016; 29:452–463.
9. Colombo T, Farina L, Macino G, Paci P. PVT1: a rising star among oncogenic long noncoding RNAs. Biomed Res Int. 2015; 2015:304208.
10. Hajjari M, Salavaty A. HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer Biol Med. 2015; 12:1–9.
11. Gutschner T, Hämmerle M, Diederichs S. MALAT1–a paradigm for long noncoding RNA function in cancer. J Mol Med. 2013; 91:791–801.
12. Pickard MR, Williams GT. Molecular and cellular mechanisms of action of tumour suppressor GAS5 lncRNA. Genes. 2015; 6:484–499.
13. Lee GL, Dobi A, Srivastava S. Prostate cancer: diagnostic performance of the PCA3 urine test. Nat Rev Urol. 2011; 8:123–124.
14. Matouk I, Raveh E, Ohana P, Lail RA, Gershtain E, Gilon M, De Groot N, Czerniak A, Hochberg A. The increasing complexity of the oncofetal h19 gene locus: functional dissection and therapeutic intervention. Int J Mol Sci. 2013; 14:4298–4316.
15. Stephan C, Ralla B, Jung K. Prostate-specific antigen and other serum and urine markers in prostate cancer. Biochim Biophys Acta. 2014; 1846:99–112.
16. Smaldone MC, Davies BJ. BC-819, a plasmid comprising the H19 gene regulatory sequences and diphtheria toxin A, for the potential targeted therapy of cancers. Curr Opin Mol Ther. 2010; 12:607–616.
17. Mizrahi A, Czerniak A, Ohana P, Amiur S, Gallula J, Matouk I, Abu-Lail R, Birman T, Hochberg A, Levy T. Treatment of ovarian cancer ascites by intra-peritoneal injection of diphtheria toxin A chain-H19 vector: a case report. J Med Case Rep. 2010; 4:228.
18. Hanna N, Ohana P, Konikoff FM, Leichtmann G, Hubert A, Appelbaum L, Kopelman Y, Czerniak A, Hochberg A. Phase 1/2a, dose-escalation, safety, pharmacokinetic and preliminary efficacy study of intratumoral administration of BC-819 in patients with unresectable pancreatic cancer. Cancer Gene Ther. 2012; 19:374–381.
19. Wang XS, Zhang Z, Wang HC, Cai JL, Xu QW, Li MQ, Chen YC, Qian XP, Lu TJ, Yu LZ, Zhang Y, Xin DQ, Na YQ, et al. Rapid identification of UCA1 as a very sensitive and specific unique marker for human bladder carcinoma. Clin Cancer Res. 2006; 12:4851–4858.
20. Xue M, Chen W, Li X. Urothelial cancer associated 1: a long noncoding RNA with a crucial role in cancer. J Cancer Res Clin Oncol. 2016; 142:1407–1419.
21. Wang F, Li X, Xie X, Zhao L, Chen W. UCA1, a non-protein-coding RNA up-regulated in bladder carcinoma and embryo, influencing cell growth and promoting invasion. FEBS Lett. 2008; 582:1919–1927.
22. Wang F, Ying HQ, He BS, Pan YQ, Deng QW, Sun HL, Chen J, Liu X, Wang SK. Upregulated lncRNA-UCA1 contributes to progression of hepatocellular carcinoma through inhibition of miR-216b and activation of FGFR1/ERK signaling pathway. Oncotarget. 2015; 6:7899–7917. https://doi.org/10.18632/oncotarget.3219.
23. Huang J, Zhou N, Watabe K, Lu Z, Wu F, Xu M, Mo YY. Long non-coding RNA UCA1 promotes breast tumor growth by suppression of p27 (Kip1). Cell Death Dis. 2014; 5:e1008.
24. Wang F, Zhou J, Xie X, Hu J, Chen L, Hu Q, Guo H, Yu C. Involvement of SRPK1 in cisplatin resistance related to long non-coding RNA UCA1 in human ovarian cancer cells. Neoplasma. 2015; 62:432–438.
25. Zheng Q, Wu F, Dai WY, Zheng DC, Zheng C, Ye H, Zhou B, Chen JJ, Chen P. Aberrant expression of UCA1 in gastric cancer and its clinical significance. Clin Transl Oncol. 2015; 17:640–646.
26. Li JY, Ma X, Zhang CB. Overexpression of long non-coding RNA UCA1 predicts a poor prognosis in patients with esophageal squamous cell carcinoma. Int J Clin Exp Pathol. 2014; 7:7938–7944.
27. Tian Y, Zhang X, Hao Y, Fang Z, He Y. Potential roles of abnormally expressed long noncoding RNA UCA1 and Malat-1 in metastasis of melanoma. Melanoma Res. 2014; 24:335–341.
28. Fang Z, Wu L, Wang L, Yang Y, Meng Y, Yang H. Increased expression of the long non-coding RNA UCA1 in tongue squamous cell carcinomas: a possible correlation with cancer metastasis. Oral Surg Oral Med Oral Pathol Oral Radiol. 2014; 117:89–95.
29. Han Y, Yang YN, Yuan HH, Zhang TT, Sui H, Wei XL, Liu L, Huang P, Zhang WJ, Bai YX. UCA1, a long non-coding RNA up-regulated in colorectal cancer influences cell proliferation, apoptosis and cell cycle distribution. Pathology. 2014; 46:396–401.
30. Wang X, Yang B, Ma B. The UCA1/miR-204/Sirt1 axis modulates docetaxel sensitivity of prostate cancer cells. Cancer Chemother Pharmacol. 2016; 78:1025–1031.
31. Hughes JM, Legnini I, Salvatori B, Masciarelli S, Marchioni M, Fazi F, Morlando M, Bozzoni I, Fatica A. C/EBPα-p30 protein induces expression of the oncogenic long non-coding RNA UCA1 in acute myeloid leukemia. Oncotarget. 2015; 6:18534–18544. https://doi.org/10.18632/oncotarget.4069.
32. Chen P, Wan D, Zheng D, Zheng Q, Wu F, Zhi Q. Long non-coding RNA UCA1 promotes the tumorigenesis in pancreatic cancer. Biomed Pharmacother. 2016; 83:1220–1226.
33. Zhao W, Sun C, Cui Z. A long noncoding RNA UCA1 promotes proliferation and predicts poor prognosis in glioma. Clin Transl Oncol. 2017; 19:735–741.
34. Bian Z, Jin L, Zhang J, Yin Y, Quan C, Hu Y, Feng Y, Liu H, Fei B, Mao Y, Zhou L, Qi X, Huang S, et al. LncRNA-UCA1 enhances cell proliferation and 5-fluorouracil resistance in colorectal cancer by inhibiting miR-204-5p. Sci Rep. 2016; 6:23892.
35. Zhang Q, Su M, Lu G, Wang J. The complexity of bladder cancer: long noncoding RNAs are on the stage. Mol Cancer. 2013; 12:101.
36. Rorive S, Sandras F, Biskri L, Fossion C, Catteau X, Roumeguere T, Bossche V, Mijatovic T, Salmon I. RT-PCR-based UCA1 expression detection in urine samples as non-invasive reliable method for urothelial cancer diagnosis. EJC Suppl. 2009; 7:28–29.
37. Février B, Raposo G. Exosomes: endosomal-derived vesicles shipping extracellular messages. Curr Opin Cell Biol. 2004; 16:415–421.
38. Xu CG, Yang MF, Ren YQ, Wu CH, Wang LQ. Exosomes mediated transfer of lncRNA UCA1 results in increased tamoxifen resistance in breast cancer cells. Eur Rev Med Pharmacol Sci. 2016; 20:4362–4368.
39. Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009; 458:223–227.
40. Yang F, Huo XS, Yuan SX, Zhang L, Zhou WP, Wang F, Sun SH. Repression of the long noncoding RNA-LET by histone deacetylase 3 contributes to hypoxia-mediated metastasis. Mol Cell. 2013; 49:1083–1096.
41. Wu W, Zhang S, Li X, Xue M, Cao S, Chen W. Ets-2 regulates cell apoptosis via the Akt pathway, through the regulation of urothelial cancer associated 1, a long non-coding RNA, in bladder cancer cells. PLoS One. 2013; 8:e73920.
42. Xue M, Li X, Wu W, Zhang S, Wu S, Li Z, Chen W. Upregulation of long non-coding RNA urothelial carcinoma associated 1 by CCAAT/enhancer binding protein α contributes to bladder cancer cell growth and reduced apoptosis. Oncol Rep. 2014; 31:1993–2000.
43. Xue M, Li X, Li Z, Chen W. Urothelial carcinoma associated 1 is a hypoxia-inducible factor-1α-targeted long noncoding RNA that enhances hypoxic bladder cancer cell proliferation, migration, and invasion. Tumour Biol. 2014; 35:6901–6912.
44. Lee JJ, Kim M, Kim HP. Epigenetic regulation of long noncoding RNA UCA1 by SATB1 in breast cancer. BMB Rep. 2016; 49:578–583.
45. Hiemer SE, Szymaniak AD, Varelas X. The transcriptional regulators TAZ and YAP direct transforming growth factor β-induced tumorigenic phenotypes in breast cancer cells. J Biol Chem. 2014; 289:13461–13474.
46. Kumar PP, Emechebe U, Smith R, Franklin S, Moore B, Yandell M, Lessnick SL, Moon AM. Coordinated control of senescence by lncRNA and a novel T-box3 co-repressor complex. Elife. 2014; 3.
47. Beyer TA, Weiss A, Khomchuk Y, Huang K, Ogunjimi AA, Varelas X, Wrana JL. Switch enhancers interpret TGF-β and Hippo signaling to control cell fate in human embryonic stem cells. Cell Rep. 2013; 5:1611–1624.
48. Wang T, Yuan J, Feng N, Li Y, Lin Z, Jiang Z, Gui Y. Hsa-miR-1 downregulates long non-coding RNA urothelial cancer associated 1 in bladder cancer. Tumour Biol. 2014; 35:10075–10084.
49. Liu FT, Zhu PQ, Luo HL, Zhang Y, Qiu C. Prognostic value of long non-coding RNA UCA1 in human solid tumors. Oncotarget. 2016; 7:57991–58000. https://doi.org/10.18632/oncotarget.11155.
50. Simon MD, Wang CI, Kharchenko PV, West JA, Chapman BA, Alekseyenko AA, Borowsky ML, Kuroda MI, Kingston RE. The genomic binding sites of a noncoding RNA. Proc Natl Acad Sci USA. 2011; 108:20497–20502.
51. Chu C, Qu K, Zhong FL, Artandi SE, Chang HY. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol Cell. 2011; 44:667–678.
52. Kretz M, Siprashvili Z, Chu C, Webster DE, Zehnder A, Qu K, Lee CS, Flockhart RJ, Groff AF, Chow J, Johnston D, Kim GE, Spitale RC, et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature. 2013; 493:231–235.
53. Ule J, Jensen K, Mele A, Darnell RB. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods. 2005; 37:376–386.
54. Wang Z, Tollervey J, Briese M, Turner D, Ule J. CLIP: construction of cDNA libraries for high-throughput sequencing from RNAs cross-linked to proteins in vivo. Methods. 2009; 48:287–293.
55. Hafner M, Landthaler M, Burger L, Khorshid M, Hausser J, Berninger P, Rothballer A, Ascano M Jr, Jungkamp AC, Munschauer M, Ulrich A, Wardle GS, Dewell S, et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell. 2010; 141:129–141.
56. Spitale RC, Flynn RA, Zhang QC, Crisalli P, Lee B, Jung JW, Kuchelmeister HY, Batista PJ, Torre EA, Kool ET, Chang HY. Structural imprints in vivo decode RNA regulatory mechanisms. Nature. 2015; 519:486–490.
57. Lucks JB, Mortimer SA, Trapnell C, Luo S, Aviran S, Schroth GP, Pachter L, Doudna JA, Arkin AP. Multiplexed RNA structure characterization with selective 2′-hydroxyl acylation analyzed by primer extension sequencing (SHAPE-Seq). Proc Natl Acad Sci USA. 2011; 108:11063–11068.
58. Nie W, Ge HJ, Yang XQ, Sun X, Huang H, Tao X, Chen WS, Li B. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett. 2016; 371:99–106.
59. Xue M, Pang H, Li X, Li H, Pan J, Chen W. Long non-coding RNA urothelial cancer-associated 1 promotes bladder cancer cell migration and invasion by way of the hsa-miR-145-ZEB1/2-FSCN1 pathway. Cancer Sci. 2016; 107:18–27.
60. Jiao C, Song Z, Chen J, Zhong J, Cai W, Tian S, Chen S, Yi Y, Xiao Y. lncRNA-UCA1 enhances cell proliferation through functioning as a ceRNA of Sox4 in esophageal cancer. Oncol Rep. 2016; 36:2960–2966.
61. Li X, Wu Y, Liu A, Tang X. Long non-coding RNA UCA1 enhances tamoxifen resistance in breast cancer cells through a miR-18a-HIF1α feedback regulatory loop. Tumour Biol. 2016; 37:14733–14743.
62. Xiao Y, Jiao C, Lin Y, Chen M, Zhang J, Wang J, Zhang Z. lncRNA UCA1 contributes to imatinib resistance by acting as a ceRNA against miR-16 in chronic myeloid leukemia cells. DNA Cell Biol. 2017; 36:18–25.
63. Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011; 43:904–914.
64. Drinberg V, Bitcover R, Rajchenbach W, Peer D. Modulating cancer multidrug resistance by sertraline in combination with a nanomedicine. Cancer Lett. 2014; 354:290–298.
65. Geng M, Wang L, Chen X, Cao R, Li P. The association between chemosensitivity and Pgp, GST-π and Topo II expression in gastric cancer. Diagn Pathol. 2013; 8:198.
66. Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011; 10:38.
67. Guttman M, Donaghey J, Carey BW, Garber M, Grenier JK, Munson G, Young G, Lucas AB, Ach R, Bruhn L, Yang X, Amit I, Meissner A, et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature. 2011; 477:295–300.
68. Chen Q, Chong T, Yin J, Luo P, Deng A. Molecular events are associated with resistance to vinblastine in bladder cancer. Cell Mol Biol. 2015; 61:33–38.
69. Barocas DA, Clark PE. Bladder cancer. Curr Opin Oncol. 2008; 20:307–314.
70. Fan Y, Shen B, Tan M, Mu X, Qin Y, Zhang F, Liu Y. Long non-coding RNA UCA1 increases chemoresistance of bladder cancer cells by regulating Wnt signaling. FEBS J. 2014; 281:1750–1758.
71. Pan J, Li X, Wu W, Xue M, Hou H, Zhai W, Chen W. Long non-coding RNA UCA1 promotes cisplatin/gemcitabine resistance through CREB modulating miR-196a-5p in bladder cancer cells. Cancer Lett. 2016; 382:64–76.
72. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108.
73. Florea AM, Büsselberg D. Cisplatin as an anti-tumor drug: cellular mechanisms of activity, drug resistance and induced side effects. Cancers. 2011; 3:1351–1371.
74. Zhang L, Cao X, Zhang L, Zhang X, Sheng H, Tao K. UCA1 overexpression predicts clinical outcome of patients with ovarian cancer receiving adjuvant chemotherapy. Cancer Chemother Pharmacol. 2016; 77:629–634.
75. Ojo D, Wei F, Liu Y, Wang E, Zhang H, Lin X, Wong N, Bane A, Tang D. Factors promoting tamoxifen resistance in breast cancer via stimulating breast cancer stem cell expansion. Curr Med Chem. 2015; 22:2360–2374.
76. Wu C, Luo J. Long non-coding rna (lncRNA) urothelial carcinoma-associated 1 (UCA1) enhances tamoxifen resistance in breast cancer cells via inhibiting mTOR signaling pathway. Med Sci Monit. 2016; 22:3860–3867.
77. Liu H, Wang G, Yang L, Qu J, Yang Z, Zhou X. Knockdown of long non-coding RNA UCA1 increases the tamoxifen sensitivity of breast cancer cells through inhibition of Wnt/β-catenin pathway. PLoS One. 2016; 11:e0168406.
78. Zhou C, Wu YL, Chen G, Feng J, Liu XQ, Wang C, Zhang S, Wang J, Zhou S, Ren S, Lu S, Zhang L, Hu C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011; 12:735–742.
79. Keedy VL, Temin S, Somerfield MR, Beasley MB, Johnson DH, McShane LM, Milton DT, Strawn JR, Wakelee HA, Giaccone G. American Society of Clinical Oncology provisional clinical opinion: epidermal growth factor receptor (EGFR) Mutation testing for patients with advanced non-small-cell lung cancer considering first-line EGFR tyrosine kinase inhibitor therapy. J Clin Oncol. 2011; 29:2121–2127.
80. Somaiah N, Fidler MJ, Garrett-Mayer E, Wahlquist A, Shirai K, Buckingham L, Hensing T, Bonomi P, Simon GR. Epidermal growth factor receptor (EGFR) mutations are exceptionally rare in thyroid transcription factor (TTF-1)-negative adenocarcinomas of the lung. Oncoscience. 2014; 1:522–528. doi: 10.18632/oncoscience.72.
81. Gainor JF, Shaw AT. Emerging paradigms in the development of resistance to tyrosine kinase inhibitors in lung cancer. J Clin Oncol. 2013; 31:3987–3996.
82. Wang Y, Xia H, Zhuang Z, Miao L, Chen X, Cai H. Axl-altered microRNAs regulate tumorigenicity and gefitinib resistance in lung cancer. Cell Death Dis. 2014; 5:e1227.
83. Bar J, Onn A. Overcoming molecular mechanisms of resistance to first-generation epidermal growth factor receptor tyrosine kinase inhibitors. Clin Lung Cancer. 2012; 13:267–279.
84. Hammerman PS, Jänne PA, Johnson BE. Resistance to epidermal growth factor receptor tyrosine kinase inhibitors in non-small cell lung cancer. Clin Cancer Res. 2009; 15:7502–7509.
85. Ohashi K, Maruvka YE, Michor F, Pao W. Epidermal growth factor receptor tyrosine kinase inhibitor-resistant disease. J Clin Oncol. 2013; 31:1070–1080.
86. Cheng N, Li X, Zhao C, Ren S, Chen X, Cai W, Zhao M, Zhang Y, Li J, Wang Q, Zhou C. Microarray expression profile of long non-coding RNAs in EGFR-TKIs resistance of human non-small cell lung cancer. Oncol Rep. 2015; 33:833–839.
87. Cheng N, Cai W, Ren S, Li X, Wang Q, Pan H, Zhao M, Li J, Zhang Y, Zhao C, Chen X, Fei K, Zhou C, et al. Long non-coding RNA UCA1 induces non-T790M acquired resistance to EGFR-TKIs by activating the AKT/mTOR pathway in EGFR-mutant non-small cell lung cancer. Oncotarget. 2015; 6:23582–23593. https://doi.org/10.18632/oncotarget.4361.
88. Sugano K. Screening of gastric cancer in Asia. Best Pract Res Clin Gastroenterol. 2015; 29:895–905.
89. Newton AD, Datta J, Loaiza-Bonilla A, Karakousis GC, Roses RE. Neoadjuvant therapy for gastric cancer: current evidence and future directions. J Gastrointest Oncol. 2015; 6:534–543.
90. Fang Q, Chen X, Zhi X. Long non-coding RNA (LncRNA) urothelial carcinoma associated 1 (UCA1) increases multi-drug resistance of gastric cancer via downregulating miR-27b. Med Sci Monit. 2016; 22:3506–3513.
91. Shang C, Guo Y, Zhang J, Huang B. Silence of long noncoding RNA UCA1 inhibits malignant proliferation and chemotherapy resistance to adriamycin in gastric cancer. Cancer Chemother Pharmacol. 2016; 77:1061–1067.
92. Centelles JJ. General aspects of colorectal cancer. ISRN Oncol. 2012; 2012:139268.
93. Walz C, Sattler M. Novel targeted therapies to overcome imatinib mesylate resistance in chronic myeloid leukemia (CML). Crit Rev Oncol Hematol. 2006; 57:145–164.
94. Spolitu S, Uda S, Deligia S, Frau A, Collu M, Angius F, Batetta B. Multidrug resistance P-glycoprotein dampens SR-BI cholesteryl ester uptake from high density lipoproteins in human leukemia cells. Am J Cancer Res. 2016; 6:615–627.
95. Druker BJ, Sawyers CL, Kantarjian H, Resta DJ, Reese SF, Ford JM, Capdeville R, Talpaz M. Activity of a specific inhibitor of the BCR-ABL tyrosine kinase in the blast crisis of chronic myeloid leukemia and acute lymphoblastic leukemia with the Philadelphia chromosome. N Engl J Med. 2001; 344:1038–1042.
96. Huang H, Luo Y, Liang Y, Long XD, Peng Y, Liu Z, Wen X, Jia M, Tian R, Bai C, Li C, Dong X. CD4+CD25+ cells in multiple myeloma related renal impairment. Sci Rep. 2015; 5:16565.
97. Peng XX, Tiwari AK, Wu HC, Chen ZS. Overexpression of P-glycoprotein induces acquired resistance to imatinib in chronic myelogenous leukemia cells. Chin J Cancer. 2012; 31:110–118.
98. Verkerk AJ, Ariel I, Dekker MC, Schneider T, van Gurp RJ, de Groot N, Gillis AJ, Oosterhuis JW, Hochberg AA, Looijenga LH. Unique expression patterns of H19 in human testicular cancers of different etiology. Oncogene. 1997; 14:95–107.
99. Chery J. RNA therapeutics: RNAi and antisense mechanisms and clinical applications. Postdoc J. 2016; 4:35–50.
100. Ulitsky I, Shkumatava A, Jan CH, Sive H, Bartel DP. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell. 2011; 147:1537–1550.
101. Hu Y, Xu K, Yagüe E. miR-218 targets survivin and regulates resistance to chemotherapeutics in breast cancer. Breast Cancer Res Treat. 2015; 151:269–280.
102. Shang Y, Feng B, Zhou L, Ren G, Zhang Z, Fan X, Sun Y, Luo G, Liang J, Wu K, Nie Y, Fan D. The miR27b-CCNG1-P53-miR-508-5p axis regulates multidrug resistance of gastric cancer. Oncotarget. 2016; 7:538–549. https://doi.org/10.18632/oncotarget.6374.
103. Ho TT, Zhou N, Huang J, Koirala P, Xu M, Fung R, Wu F, Mo YY. Targeting non-coding RNAs with the CRISPR/Cas9 system in human cell lines. Nucleic Acids Res. 2015; 43:e17.
104. Zhen S, Hua L, Liu YH, Sun XM, Jiang MM, Chen W, Zhao L, Li X. Inhibition of long non-coding RNA UCA1 by CRISPR/Cas9 attenuated malignant phenotypes of bladder cancer. Oncotarget. 2017; 8:9634–9646. https://doi.org/10.18632/oncotarget.14176.

