INTRODUCTION
Due to the aggressive nature of pancreatic ductal adenocarcinoma (PDAC), it is one of the most lethal malignancies in the world [1, 2]. Recently developed targeted molecular strategies have not contributed to improved therapies for PDAC, and the 5-year survival rate after diagnosis is only 5% [2, 3]. More than 50% of patients develop local recurrence or distant metastasis after curative resection. PDAC frequently metastasizes to the liver, peritoneum and lung [4, 5]. We hypothesized that current genomic approaches might be used to elucidate the molecular mechanisms underlying PDAC metastasis and suggest improved treatments for this disease.
microRNA (miRNA) belongs to a family of small non-coding RNAs that fine-tune the expression of protein coding/noncoding RNAs by repressing translation or cleaving RNA transcripts in a sequence-dependent manner [6]. A unique characteristic of human miRNAs is that a single miRNA species can regulate a large number of RNA transcripts [7, 8]. Therefore, dysregulated miRNAs can disrupt tightly controlled RNA networks and promote cancer cell metastasis. At present, numerous studies have indicated that miRNAs are aberrantly expressed in several cancers, including PDAC [9–11]. The discovery of miRNAs and subsequent studies have deepened our understanding of the roles of miRNA in human cancer pathogenesis [12, 13].
Using miRNA expression signatures, we have identified antitumor miRNAs that modulate novel cancer networks in several types of cancer [14–16]. More recently, we reported the anti-tumor function of miR-375, and the observation that it regulated oncogenic zinc finger protein 36 ring finger protein-like 2 (ZFP36L2) in PDAC cells [17]. Moreover, ZFP36L2 was overexpressed in PDAC specimens and Kaplan–Meier survival curves showed that high expression of ZFP36L2 predicted shorter survival in PDAC patients [17]. We have also constructed an miRNA expression signature of PDAC clinical specimens based on RNA-sequencing. Our data showed that miR-217 was significantly downregulated in PDAC tissues, suggesting that it functions as an antitumor miRNA that targeted several oncogenic genes in PDAC cells. Past studies demonstrated that miR-217 acts as an antitumor miRNA in several types of cancer [18–20]. miR-217 expression was observed to be negatively correlated with KRAS protein expression in PDAC cell lines and miR-217 directly regulated KRAS [21, 22]. However, the RNA networks mediated by miR-217 in PDAC are still obscure.
The aim of this study was to investigate the antitumor roles of miR-217 in PDAC cells and to identify miR-217-mediated molecular pathways involved in PDAC aggressiveness. Our present data showed that the actin-binding protein anillin (ANLN) (coded by the ANLN gene) was directly regulated by antitumor miR-217 in PDAC cells. Kaplan–Meier survival curves showed that high expression of ANLN predicted shorter survival in patients. Moreover, we showed that “Focal adhesion” and “Regulation of actin binding protein” were downstream pathways modulated by ANLN protein in PDAC cells. Elucidation of antitumor miR-217-mediated molecular networks in PDAC may provide new insights into the potential mechanisms of PDAC aggressiveness.
RESULTS
Expression levels of miR-217 in PDAC specimens and cell lines
We evaluated expression levels of miR-217 in PDAC tissues (n = 27), normal pancreatic tissues (n = 14) and two PDAC cell lines (PANC-1 and SW1990). The clinical samples’ backgrounds and clinicopathological characteristics are summarized in Table 1A. Normal pancreatic tissues are summarized in Table 1B. The expression levels of miR-217 were significantly lower in tumor tissues compared with normal pancreatic tissues (P < 0.0001, Figure 1A, Supplementary Figure 1). However, there were no significant relationships between any of the clinicopathological parameters, (i.e., neoadjuvant chemotherapy, metastasis or recurrence) and the expression of miR-217 (data not shown).
Table 1A: Patient characteristics
Pancreatic ductal adenocarcinoma (PDAC) |
(%) |
||
|---|---|---|---|
Total number |
27 |
||
Average age (range), years |
67.1 (42-85) |
||
Gender |
Male |
12 |
(44.4) |
Female |
15 |
(55.6) |
|
T category |
pTis |
1 |
(3.7) |
pT1 |
2 |
(7.4) |
|
pT2 |
0 |
(0) |
|
pT3 |
22 |
(81.5) |
|
pT4 |
2 |
(7.4) |
|
N category |
0 |
15 |
(55.6) |
1 |
12 |
(44.4) |
|
M category |
0 |
25 |
(92.6) |
1 |
2 |
(7.4) |
|
Neoadjuvant chemotherapy |
(-) |
12 |
(44.4) |
(+) |
15 |
(56.6) |
|
Recurrence |
(-) |
7 |
(25.9) |
(+) |
20 |
(74.1) |
|
Table 1B: Patient characteristics
Normal pancreatic tissue |
|||
|---|---|---|---|
Total number |
14 |
||
Average age (range), years |
63.8 (42-85) |
||
Gender |
Male |
6 |
(42.8) |
Female |
8 |
(57.2) |
|
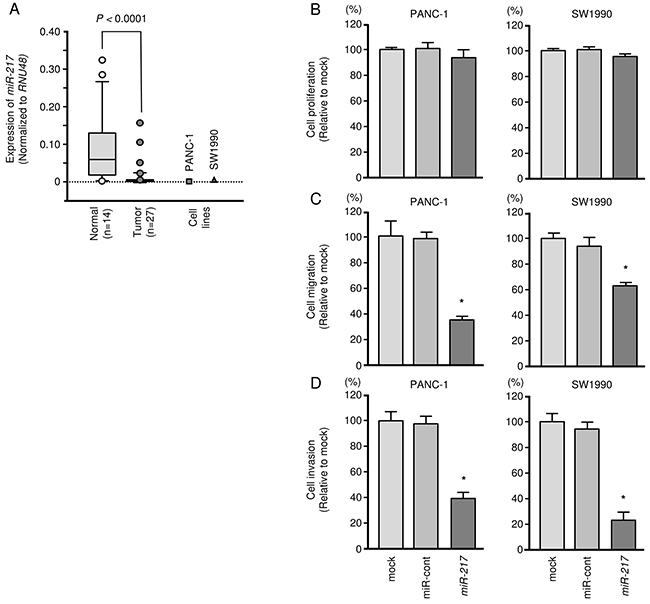
Figure 1: Antitumor functions of miR-217 in PDAC cell lines (PANC-1 and SW1990). (A) Expression levels of miR-217 in PDAC clinical specimens and cell lines were determined by qRT-PCR. Data were normalized to RNU48 expression. (B) Cell proliferation was determined by XTT assays 72 h after transfection with 10 nM miR-217. *, P < 0.0001. (C) Cell migration activity was determined by migration assays. *, P < 0.0001. (D) Cell invasion activity was determined using Matrigel invasion assays. *, P < 0.0001.
Effect of miR-217 expression on cell growth, migration and invasiveness in PDAC cell lines
To investigate the functional roles of miR-217, we performed gain-of-function studies using transfection of PANC-1 and SW1990 cells. The expression levels of miR-217 were markedly lower in two cell lines (Figure 1A). To elucidate molecular mechanisms of low expression of miR-217 in PDAC cells, PANC-1 and SW1990 cells were treated with the demethylating agent [5-aza-2’-deoxycytidine (5-aza-dC)]. Expression levels of miR-217 in PDAC cells were significantly elevated by 5-aza-dc treatment (Supplementary Figure 2). These data suggested that DNA methylation might cause silencing of miR-217 in PDAC cells.
XTT assays revealed no significant inhibition of cell proliferation in PANC-1 or in SW1990 cells transfected with miR-217 in comparison with mock or control transfectants (Figure 1B). In vitro assays demonstrated that migration and invasion were significantly inhibited in miR-217 transfectants compared with mock or miR-control transfectants (each, P < 0.0001, Figure 1C and 1D, Supplementary Figure 5). These results suggested that miR-217 could have an antitumor function in PDAC cells.
Identification of candidate genes regulated by miR-217 in PDAC cells
To gain further insight into the molecular mechanisms and pathways regulated by tumor-suppressive miR-217 in PDAC cells, we used in silico analyses. The strategy for narrowing down the genes targeted by miR-217 is shown in Figure 2. The TargetScan database showed that 3,970 genes have putative target sites for miR-217 in their 3’-UTRs. Gene expression data showed that 996 genes were upregulated (fold-change log2 > 1.5) in cancer tissues by GEO database analyses (GEO accession number; GSE15471). We identified 167 genes that were putative targets of miR-217 and were upregulated in PDAC specimens. Finally, we found that 19 genes had conserved sequences that were putatively targeted by miR-217 (Table 2).
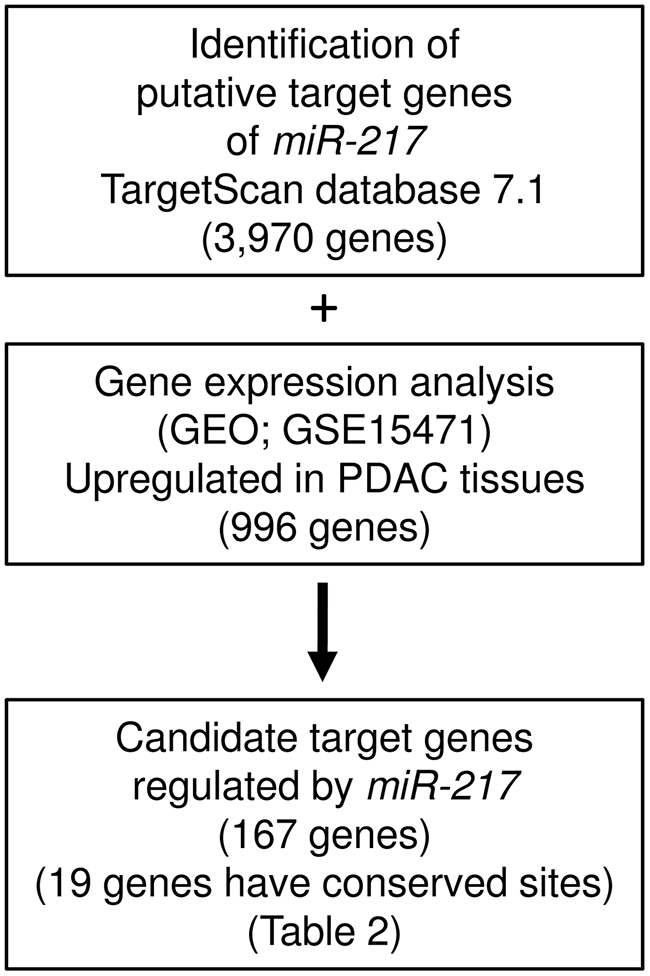
Figure 2: The strategy for analysis of miR-217 target genes. Flow chart illustrates the strategy for analysis of miR-217 target genes in PDAC cells.
Table 2: Candidate target genes regulated by miR-217
Entrez gene |
Gene symbol |
Gene name |
Conserved sites |
GEO data |
*TCGA_OncoLnc |
|||
|---|---|---|---|---|---|---|---|---|
8mer sites |
7mer-m8 sites |
7mer-A1 sites |
Total |
FC(log2) |
P-value |
|||
9644 |
SH3PXD2A |
SH3 and PX domains 2A |
1 |
1 |
1 |
3 |
1.631 |
0.3620 |
54443 |
ANLN |
anillin, actin binding protein |
1 |
1 |
0 |
2 |
1.729 |
0.0014 |
3624 |
INHBA |
inhibin, beta A |
0 |
1 |
0 |
1 |
5.190 |
0.0414 |
1295 |
COL8A1 |
collagen, type VIII, alpha 1 |
1 |
0 |
0 |
1 |
4.623 |
0.3640 |
2335 |
FN1 |
fibronectin 1 |
0 |
1 |
0 |
1 |
3.727 |
0.0950 |
860 |
RUNX2 |
runt-related transcription factor 2 |
1 |
0 |
0 |
1 |
2.932 |
0.4790 |
10085 |
EDIL3 |
EGF-like repeats and discoidin I-like domains 3 |
1 |
0 |
0 |
1 |
2.560 |
0.5300 |
6400 |
SEL1L |
sel-1 suppressor of lin-12-like (C. elegans) |
1 |
0 |
0 |
1 |
2.375 |
0.0950 |
22943 |
DKK1 |
dickkopf WNT signaling pathway inhibitor 1 |
0 |
0 |
1 |
1 |
2.303 |
0.0110 |
800 |
CALD1 |
caldesmon 1 |
0 |
0 |
1 |
1 |
2.067 |
0.5440 |
10630 |
PDPN |
podoplanin |
0 |
1 |
0 |
1 |
1.957 |
0.3500 |
79070 |
KDELC1 |
KDEL (Lys-Asp-Glu-Leu) containing 1 |
1 |
0 |
0 |
1 |
1.870 |
0.0969 |
55454 |
CSGALNACT2 |
chondroitin sulfateN-acetylgalactosaminyltransferase 2 |
0 |
1 |
0 |
1 |
1.781 |
0.6930 |
57181 |
SLC39A10 |
solute carrier family 39 (zinc transporter),member 10 |
0 |
1 |
0 |
1 |
1.770 |
0.0007 |
6091 |
ROBO1 |
roundabout, axon guidance receptor, homolog 1 (Drosophila) |
0 |
0 |
1 |
1 |
1.769 |
0.8790 |
22856 |
CHSY1 |
chondroitin sulfate synthase 1 |
0 |
0 |
1 |
1 |
1.652 |
0.3330 |
23603 |
CORO1C |
coronin, actin binding protein, 1C |
0 |
0 |
1 |
1 |
1.566 |
0.0056 |
50515 |
CHST11 |
carbohydrate (chondroitin 4) sulfotransferase 11 |
1 |
0 |
0 |
1 |
1.559 |
0.0024 |
9444 |
QKI |
QKI, KH domain containing, RNA binding |
0 |
0 |
1 |
1 |
1.554 |
0.9170 |
*Kaplan-Meier analysis log-rank P-value < 0.05
Poor prognosis with a high expression
We also checked the expression status of those 19 genes and the clinical significance of PDAC by using the OncoLnc database (http://www.oncolnc.org/). Kaplan-Meier survival curves showed that high expression of 6 genes was associated with poor prognosis in PDAC by TCGA database searching (Table 2, Supplementary Figure 3). In this study, we focused on the actin binding protein anillin (ANLN) because past studies had indicated that actin binding proteins were involved in metastasis [23–26].
ANLN is a direct target of miR-217 in PDAC cells
We performed qRT-PCR to validate miR-217 repression of ANLN mRNA expression in PDAC cell lines. Our studies revealed that ANLN mRNA was significantly reduced in miR-217 transfectants in comparison with mock or miR-control transfectants (P < 0.0001 and P < 0.0001, Figure 3A). Expression of ANLN protein was also repressed in the miR-217 transfectants (Figure 3B, Supplementary Figure 6).
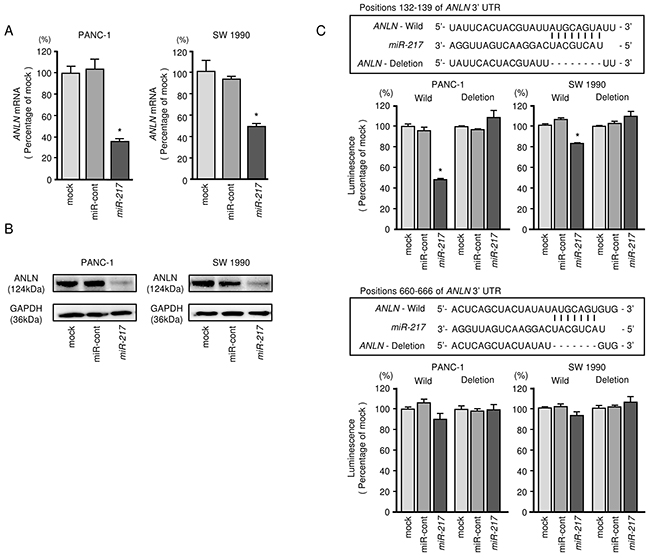
Figure 3: Direct regulation of ANLN by miR-217 in PDAC cell lines. (A) ANLN mRNA expression in PDAC cell lines was evaluated by qRT-PCR 72 h after transfection with miR-217. GUSB was used as an internal control. *, P < 0.0001. (B) ANLN protein expression in PDAC cell lines was evaluated by Western blot analyses 96 h after transfection with miR-217. GAPDH was used as a loading control. (C) miR-217 binding sites in the 3′-UTR of ANLN mRNA. Dual luciferase reporter assays using vectors encoding putative miR-217 (positions 132 - 139 or 660 - 666) target sites of the ANLN 3′-UTR for both wild-type and deleted regions. Normalized data were calculated as ratios of Renilla/firefly luciferase activities. *, P < 0.005.
Target prediction databases indicated two putative target sites in the 3’-UTR of ANLN (Figure 3C). To determine whether ANLN mRNA had a functional target site, we performed a dual luciferase reporter assay. We used vectors encoding the partial wild-type sequence of the3’-UTR of the mRNA, including the predicted miR-217 target sites. We found that the luminescence intensity was significantly reduced by co-transfection with miR-217 and the vector carrying the wild-type 3’-UTR (position 132 – 139), whereas transfection with the deletion vector (binding site had been removed) blocked the decrease in luminescence (P < 0.005, Figure 3C). In contrast, the luminescence intensity was not decreased in co-transfection with miR-217 and the vector carrying the wild-type 3’-UTR (position 660 – 666) (Figure 3C).
Effects of silencing ANLN on PDAC cell lines
To investigate the functional role of ANLN in PDAC cells, we carried out loss-of-function studies using si-ANLN transfectants. First, we evaluated the knockdown efficiency of si-ANLN transfection in PDAC cell lines. In the present study, we used two types of si-ANLN (si-ANLN-1 and si-ANLN-2). Based on qRT-PCR assessment and Western blot analyses, both siRNAs effectively downregulated ANLN expression in both cell lines (P < 0.0001, Figure 4A and 4B). XTT, cell migration and cell invasion assays demonstrated that cell proliferation, migration and invasion were inhibited in si-ANLN transfectants compared with mock- or siRNA-control-transfected cells (Figure 4C, 4D and 4E).
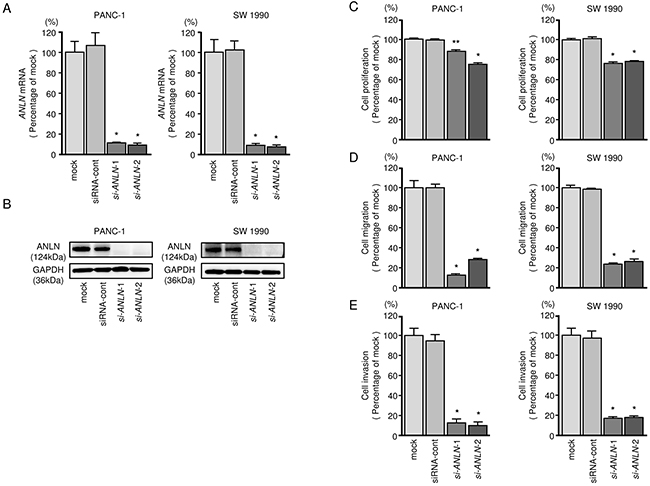
Figure 4: ANLN mRNA and ANLN protein expression after si-ANLN transfection and effects of ANLN silencing in PDAC cell lines. (A) ANLN mRNA expression in PDAC cell lines was evaluated by qRT-PCR 72 h after transfection with si-ANLN-1 or si-ANLN-2. GUSB was used as an internal control. (B) ANLN protein expression in PDAC cell lines was evaluated by Western blot analysis 96 h after transfection with miR-217. GAPDH was used as a loading control. (C) Cell proliferation was determined with the XTT assays 72 h after transfection with 10 nM si-ANLN-1 or si-ANLN-2. *,P < 0.0001, **, P < 0.05. (D) Cell migration activity was determined by migration assays. *,P < 0.0001. (E) Cell invasion activity was determined using Matrigel invasion assays. *, P < 0.0001.
Effects of cotransfection of ANLN and miR-217 in PDAC cell line
To confirm the antitumor effect of miR-217 by targeting ANLN in PDAC cells, we performed rescue experiments by ANLN overexpression in PANC-1 cells with miR-217 restoration. The rescue studies indicated that cell migration and invasion properties were rescued by ANLN transfectants compared with cells with restored miR-217 only (Supplementary Figure 4). These data indicated that ANLN expression was regulated by miR-217 and expression of miR-217 was induced antitumor effects of migration and invasion in PDAC cells.
Expression of ANLN in PDAC clinical specimens and the TCGA database
ANLN was upregulated in clinical PDAC samples and cell lines, PANC-1 and SW1990 (P < 0.0001, Figure 5A). Negative correlations between miR-217 expression and ANLN mRNA expression were found using Spearman’s rank test (R = -0.841, P < 0.0001, Figure 5B).
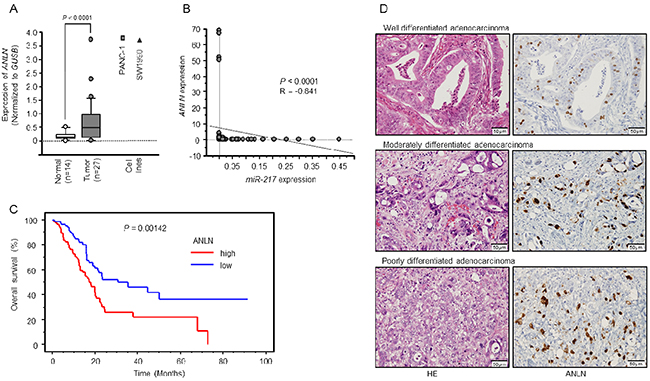
Figure 5: Expression levels of ANLN mRNA and immunohistochemical staining of ANLN protein in PDAC specimens. (A) Expression levels of ANLN mRNA in PDAC or normal pancreatic tissues and PDAC cell lines. (B) The expression levels of miR-217 and ANLN were negatively correlated (R = -0.841, P < 0.0001). (C) Kaplan–Meier curve analysis of overall patient survival between those with high ANLN expression and low ANLN expression in the PDAC TCGA database. (D) Immunohistochemical staining of ANLN in PDAC specimens. All differentiated types of PDAC (well, moderately, poorly) showed strong nuclear immunoreactivity (Left panel: hematoxylin-eosin staining, Right panel: ANLN staining, original magnification, X400).
Using a PDAC TCGA database, we found that the mRNA expression levels of ANLN were significantly upregulated. The ANLN high expression group had a significantly poorer overall survival by Kaplan-Meier analysis (Log-rank P-value = 0.00142, Figure 5C).
We confirmed the expression of ANLN protein in PDAC clinical specimens using immunohistochemistry. A total of 27 specimens were evaluated. All tumors were of the differentiated adenocarcinoma type (well, moderately, poorly) and showed strong nuclear immunoreactivity (Figure 5D).
Investigation of downstream genes regulated by ANLN in PDAC cells
To identify the downstream genes regulated by ANLN, genome-wide gene expression and in silico analyses were performed in a PDAC cell line (PANC-1) transfected with si-ANLN. Our selection strategy is shown in Figure 6. A total of 937 genes were commonly downregulated (log2 ratio < - 1.0) in si-ANLN-transfected PANC-1 cells. We also assessed the downregulated genes using KEGG pathways and the GENECODIS program. With that approach, we identified 8 pathways and 15 genes that were significantly enriched (Table 3A and 3B). Our microarray expression data was deposited in Gene Expression Omnibus (GEO) database (accession number; GSE93290).
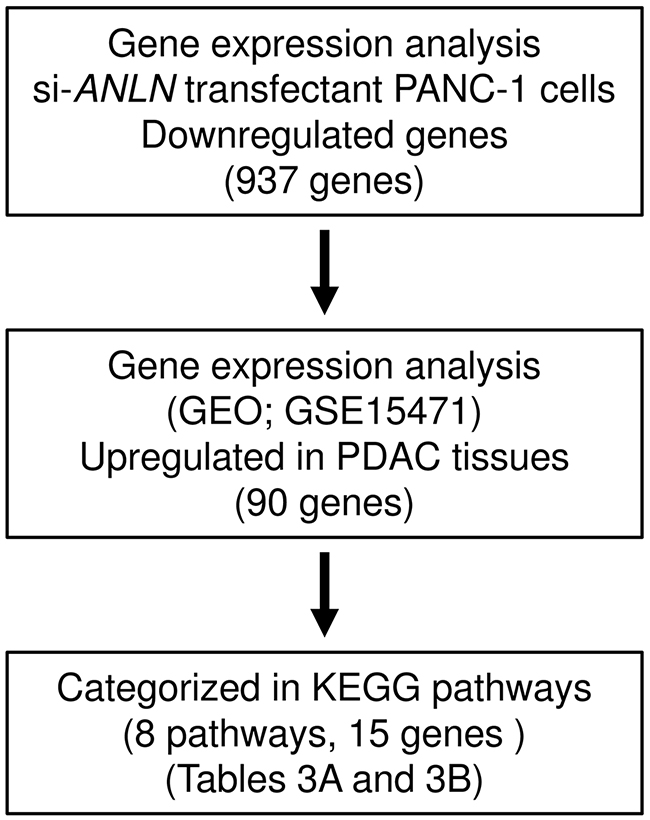
Figure 6: The strategy for analysis of ANLN downstream genes. Flow chart illustrates the strategy for analysis of ANLN-mediated downstream pathways and genes in PDAC cells.
Table 3A: Enriched KEGG pathways in si-ANLN transfectant
KEGG ID |
Pathways |
P-value |
No.of genes |
Genes |
|---|---|---|---|---|
Kegg:04510 |
Focal adhesion |
0.00011 |
5 |
ITGA3,RAC2,FLNA,LAMC2,MYL9 |
Kegg:05200 |
Pathways in cancer |
0.00105 |
5 |
ITGA3,RAC2,FZD1,FZD2,LAMC2 |
Kegg:04512 |
ECM-receptor interaction |
0.00005 |
4 |
ITGA3,HSPG2,LAMC2,CD47 |
Kegg:04810 |
Regulation of actin cytoskeleton |
0.00156 |
4 |
ITGA3,RAC2,MSN,MYL9 |
Kegg:04670 |
Leukocyte transendothelial migration |
0.00013 |
3 |
RAC2,MSN,MYL9 |
Kegg:04610 |
Complement and coagulation cascades |
0.00050 |
3 |
F8,SERPINE1,PLAU |
Kegg:04310 |
Wnt signaling pathway |
0.00050 |
3 |
RAC2,FZD1,FZD2 |
Kegg:05146 |
Amoebiasis |
0.00186 |
3 |
SERPINB9,LAMC2,GNA15 |
Table 3B: Downregulated genes by si-ANLN transfectant in enrichied KEGG pathways
Entrez gene ID |
Gene symbol |
Gene name |
miR-217 target sites |
Expression FC(log2) |
*TCGA_OncoLnc |
|
|---|---|---|---|---|---|---|
mock vs si-ANLN |
GEO |
P-value |
||||
8321 |
FZD1 |
frizzled class receptor 1 |
0 |
-1.603 |
1.186 |
0.2750 |
10398 |
MYL9 |
myosin, light chain 9, regulatory |
0 |
-1.536 |
2.181 |
0.9090 |
961 |
CD47 |
CD47 molecule |
0 |
-1.511 |
1.236 |
0.0705 |
5880 |
RAC2 |
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
0 |
-1.444 |
1.323 |
0.7490 |
3918 |
LAMC2 |
laminin, gamma 2 |
0 |
-1.436 |
2.761 |
0.0595 |
3675 |
ITGA3 |
integrin, alpha 3 (antigen CD49C, alpha 3 subunit of VLA-3 receptor) |
0 |
-1.363 |
1.329 |
0.0004 |
2316 |
FLNA |
filamin A, alpha |
0 |
-1.332 |
1.426 |
0.3290 |
2769 |
GNA15 |
guanine nucleotide binding protein (G protein), alpha 15 (Gq class) |
0 |
-1.309 |
1.089 |
0.0070 |
5328 |
PLAU |
plasminogen activator, urokinase |
1 |
-1.252 |
2.341 |
0.0052 |
2535 |
FZD2 |
frizzled class receptor 2 |
0 |
-1.243 |
1.295 |
0.0349 |
3339 |
HSPG2 |
heparan sulfate proteoglycan 2 |
0 |
-1.177 |
1.080 |
0.3290 |
5272 |
SERPINB9 |
serpin peptidase inhibitor, clade B (ovalbumin), member 9 |
1 |
-1.111 |
1.670 |
0.3210 |
4478 |
MSN |
moesin |
1 |
-1.068 |
1.740 |
0.0389 |
5054 |
SERPINE1 |
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 |
0 |
-1.060 |
1.966 |
0.0066 |
2157 |
F8 |
coagulation factor VIII, procoagulant component |
0 |
-1.021 |
1.110 |
0.3080 |
*Kaplan-Meier analysis log-rank P-value < 0.05
Poor prognosis with a high expression
Furthermore, we checked the expression status of those genes and their pathologic relations to PDAC by using the TCGA-based large cohort study data. Kaplan-Meier analysis showed that high expression group of 6 genes (ITGA3, GNA15, PLAU, FZD2, MSM and SERPINE1) had a significantly poorer overall survival for patients with PDAC (Table 3B and Figure 7).
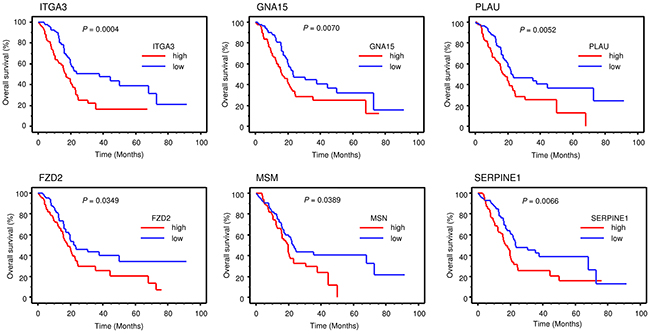
Figure 7: TCGA database analysis of ANLN downstream genes. Kaplan–Meier plots overall survival with log-rank tests between those with high and low candidacy ANLN downstream 6 genes expression in the PDAC TCGA database.
DISCUSSION
In this study, we confirmed the downregulation and antitumor function of miR-217 in PDAC cells, showing that miR-217 inhibited cancer cell migration and invasion. Based on those findings, we hypothesized that miR-217 suppressed novel metastatic pathways in PDAC cells. In human genome, miR-217 is located on the human chromosome 2q16.1 region and miR-217 forms miRNA cluster with miR-216s (miR-216a,miR-216b). Expression of these clustered miRNAs were significantly downregulated in PDAC cells (data not shown). Past studies demonstrated that miR-217 were downregulated in esophageal adenocarcinoma cells and leukemia cells [27, 28]. Moreover, our data showed that these miRNAs acted as antitumor miRNAs in PDAC cells (data not shown). First, we combined genome-wide gene expression analysis and in silico database search to identify antitumor miR-217 targets in PDAC cells. Nineteen genes were identified as putative miR-217 targets. Among them, we selected the actin-binding protein gene ANLN because our past studies showed that several actin-binding protein genes (CORO1C, FSCN1,LASP1 and MSN) were overexpressed in cancer tissues and were deeply involved in promoting human cancer cell migration and invasion [29–32]. Interestingly, we showed that expression of these actin-binding proteins was regulated by several antitumor miRNAs (miR-1,miR-133a, miR-145 and miR-218) [33–36]. Our present data demonstrated that ANLN was directly regulated by antitumor miR-217 and knockdown of ANLN significantly inhibited cancer cell aggressiveness in PDAC cells.
The human ANLN gene was cloned as a human homologue of the Anln gene in Drosophila melanogaster that encodes a 1,125–amino acid actin-binding protein. ANLN has a unique multi-domain structure: an actin-binding and a myosin-binding domain at the N-terminus and a pleckstrin homology domain at the C-terminus [37]. ANLN is a conserved protein implicated in cytoskeletal dynamics, and it is a ubiquitously expressed protein required for cytokinesis. ANLN is primarily located in the nucleus in interphase, whereas in telophase it relocates to the cytoplasm where it accumulates in the contractile ring and cleavage furrow [38]. A small guanosine triphosphatase, RHOA, interacts with ANLN and stabilizes actin-related proteins (F-actin, myosin, septins and CD2AP) in the cleavage furrow [39].
Overexpressed ANLN was reported in several cancers and elevated expression appears to be involved in the metastatic potential of human cancers [23–25]. In non-small cell lung cancer (NSCLC), nuclear localization of ANLN was associated with poor survival of patients with NSCLC [40]. Likewise, detection of nuclear ANLN was significantly associated with decreased breast cancer survival and recurrence-free survival [41]. Our present immunohistochemical assessment of ANLN protein expression showed that ANLN was localized in cell nuclei in PDAC cells. Overexpression of ANLN was confirmed in PDAC cells, and Kaplan–Meier survival curves showed that high expression of ANLN predicted shorter survival in patients with PDAC by TCGA dataset analysis. Moreover, our data showed that reduced expression of ANLN in PDAC cells suppressed cancer cell migration and invasion. Past studies showed that knockdown of ANLN expression in breast cancer cells arrested the cell cycle in S/G2 or G2/M transition [26]. These findings suggest that (1) ANLN has multiple functions, (2) its expression affects several oncogenic pathways and (3) overexpression of ANLN enhances cancer cell aggressiveness.
To investigate the molecular pathways regulated by ANLN in PDAC cells, we applied a genome-wide gene expression analysis using si-ANLN transfectants. Our data showed that several pathways were downstream from ANLN, such as the “focal adhesion”, “pathways in cancer”, “ECM-receptor interaction” and “regulation of the actin cytoskeleton”. Finally, we identified 15 genes that were upregulated in PDAC specimens and involved in those pathways. Among the 15 genes, high expression of ITGA3, GNA15, PLAU, FZD2, MSM and SERPINE1 predicted shorter survival in patients with PDAC as determined by TCGA database analysis. Our previous study showed that overexpression of ITGA3 enhanced cancer cell migration and invasion and directly regulated antitumor expression of miR-223 in prostate cancer [42].
Integrins play an important role in cell adhesion by linking the cytoskeleton of cells to components in the extracellular matrix (ECM) by integrin-mediated signaling pathways [43]. Several studies show that dysregulated ECM-integrin signaling is implicated in the development and progression of cancer [44, 45]. A complex of PLAUR and its ligand PLAU is an important regulator of ECM proteolysis, ECM interactions and cell signaling [46]. In cancer, aberrant expression of PLAU and PLAUR has been linked to tumor progression, metastasis, and shortened survival in cancer [47–49]. These findings suggest that miR-217/ANLN-mediated genes are deeply involved in PDAC pathogenesis. However, the detailed molecular mechanism how downstream genes of ANLN are regulated is still unclear. Future analysis is needed. Exploration of novel antitumor miR-217-mediated pathways may lead to the development of new treatment protocols for this disease.
In conclusion, downregulation of miR-217 was detected in PDAC clinical specimens. miR-217 acts as an antitumor miRNA through its targeting of ANLN expression in PDAC cells. Expression of ANLN enhanced cancer cell aggressiveness and its high expression predicts poorer survival of PDAC patients. Elucidation of miR-217/ANLN-mediated molecular networks may lead to a better understanding of PDAC pathogenesis and the development of new treatment protocols.
MATERIALS AND METHODS
Clinical specimens and cell lines
Clinical tissues specimens (n = 27) were collected from patients with PDAC who underwent curative surgical resection at Kagoshima University Hospital between 1997 and 2015. Normal pancreatic tissue specimens (n = 14) were obtained from noncancerous tumor-adjacent tissue. Each surgical specimen was histologically classified according to the TNM classification system [50]. All patients in this study provided informed consent and the study protocol was approved by the Institutional Review Board of Kagoshima University. Two human PDAC cell lines were investigated in this study. PANC-1 cells were obtained from RIKEN Cell Bank (Tsukuba, Ibaraki, Japan) and SW 1990 cells were obtained from the ATCC (Manassas, Virginia, USA).
Total RNA, including miRNA, was isolated using ISOGEN (NIPPON GENE, Toyama, Japan) according to the manufacturer’s protocol.
Quantitative real-time PCR (qRT-PCR)
Quantification of miRNA was performed using qRT-PCR as previously described [12, 13, 17, 51, 52]. Briefly, miRNAs were quantified using stem-loop RT-PCR, TaqMan MicroRNA Assays and Assay-on-Demand Gene Expression TaqMan probes and primers as directed by the manufacturer. Probes and primers for miR-217 (product ID: 002337; Thermo Fisher Scientific, Waltham, MA, USA), ANLN (product ID: Hs01122612_m1; Thermo Fisher Scientif), human GUSB (product ID: Hs99999908_m1; Thermo Fisher Scientific) and RNU48 (product ID: 001006; Thermo Fisher Scientific) were used as internal controls. Expression fold-changes were determined using the ΔΔCt method.
Transfection of miRNA mimic and small interfering RNA (siRNA) into PDAC cell lines
PDAC cell lines were transfected with a miRNA mimic for gain-of-function experiments and siRNA for loss-of-function experiments. Pre-miRTM miRNA precursors for miR-217 (product ID: PM 12774), negative control miRNA (product ID: AM 17111), two ANLN siRNAs (product IDs: HSS122893 and HSS122895) and negative control siRNA (product ID: D-001810-10) were purchased from Thermo Fisher Scientific. The transfection efficiencies of miRNA in PANC-1 and SW 1990 cells were calculated as described in previous studies [12, 13, 17, 51, 52].
Cell proliferation, migration and invasion assays
XTT assays were used to assess cell proliferation (Cell Proliferation Kit II, Roche Applied Sciences, Penzberg, Germany). Cell migration assays were performed with BD Falcon Cell Culture Inserts (BD Biosciences, Franklin Lakes, NJ, USA). Modified Boyden chambers containing Transwell-precoated Matrigel membrane filter inserts were used to quantitate cellular invasion. The protcols of these assays were described as previously [12, 13, 17, 51, 52].
Western blot analyses
Protein lysates were collected 96 h after transfection and 20 μg of protein were separated using gel electrophoresis on e-PAGEL 5-20% gels (ATTO, Tokyo, Japan) before transfer to polyvinylidene fluoride membranes. Solutions of mouse anti-ANLN antibodies (product ID: AMAb90662; Atlas Antibodies AB, Stockholm, Sweden) were diluted 1:750 for immunoblotting. Anti-GAPDH antibodies at a 1:1000 dilution (product ID: SAF6698; Wako, Osaka, Japan) were used as an internal loading control. A detailed description of the Western blotting procedure was published elsewhere [12, 13, 17, 51, 52].
Immunohistochemistry
Tissue sections were incubated overnight at room temperature with anti-ANLN antibodies diluted 1:200 (product ID: AMA90662; Atlas Antibodies AB, Stockholm, Sweden). Overnight, antibodies were visualized using an avidin-biotin complex (ABC) detection kit (Vector Laboratories, Burlingame, CA, USA) and a diaminobenzidine substrate system according to the manufacturer’s protocol. The expression of ANLN was evaluated in cancer-rich fields using high-power microscopy (400×).
Genome-wide gene expression and in silico analyses
To identify miR-217 target genes, we used in silico analyses conducted as described previously [12, 13, 17, 51, 52]. The microarray data were deposited into the GEO database (accession number GSE15471). Next, we selected putative miRNA target genes using the TargetScanHuman 7.1(June, 2016 release, http://www.targetscan.org/vert_71) database and TCGA (The Cancer Genome Atlas, https://cancergenome.nih.gov/). Figure 2 shows the strategy for selecting target genes.
TCGA database analysis of PDAC specimens
The clinical significance of ANLN in PDAC was assessed by RNA sequencing in TCGA OncoLnc (http://www.oncolnc.org/) [53, 54]. We linked TCGA survival data to mRNA expression levels. We obtained clinical information on 174 PDAC samples from the National Cancer Institute CDC Data Portal (https://gdc-portal.nci.nih.gov/projects/TCGA-PAAD). We selected high and low ANLN expression groups defined by the median value, and data were analyzed by Kaplan-Meier survival curves and log-rank statistics.
Plasmid construction and dual-luciferase reporter assay
The partial wild-type sequence of the ANLN 3’-untranslated region (UTR) was inserted between the XhoI–PmeI restriction sites in the 3’-UTR of the hRluc gene in the psiCHECK-2 vector (C8021; Promega, Madison, WI, USA). Alternatively, we used sequences that were missing the miR-217 target sites (position 132 - 139 or position 660 - 666). The synthesized DNA was cloned into the psiCHECK-2 vector. PANC-1 and SW1990 cells were transfected with 20 ng of the vector, 20 nM microRNAs, and 1 μL Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) in 100 μL Opti-MEM (Invitrogen). The procedure of dual-luciferase reporter assay was described previously [12, 13, 17, 51, 52].
Identification of downstream targets regulated by ANLN in PDAC
We used genome-wide gene expression analysis in a PDAC cell line (PANC-1) transfected with si-ANLN. Genes downregulated by ANLN were categorized by KEGG pathways using the GENECODIS program.
Statistical analysis
The relationships between 2 groups and expression values obtained by RT-qPCR were analyzed using the Mann-Whitney U-test. The correlation between expression of miR-217 and ANLN was evaluated using Spearman’s rank test. The relationships among more than 3 variables and numerical values were analyzed using the Bonferroni-adjusted Mann-Whitney U-test. Overall survival (OS) after surgery was gauged using Kaplan–Meier curves. Patients were divided into two groups based on ANLN expression, and differences in survival were estimated using the log-rank test. We used Expert StatView software (version 5.0 SAS Institute Inc., Cary, NC, USA) for these analyses [12, 13, 17].
ACKNOWLEDGMENTS
We wish to thank the Joint Research Laboratory, Kagoshima University Graduate School of Medical and Dental Sciences, for the use of their facilities. The present study was supported by KAKENHI(C) grant 15K10801, 15K15501 and 26293306.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Chrystoja CC, Diamandis EP, Brand R, Ruckert F, Haun R, Molina R. Pancreatic cancer. Clin Chem. 2013; 59:41-46.
2. Kamisawa T, Wood LD, Itoi T, Takaori K. Pancreatic cancer. Lancet. 2016; 388:73-85.
3. Corbo V, Tortora G, Scarpa A. Molecular pathology of pancreatic cancer: from bench-to-bedside translation. Curr Drug Targets. 2012; 13:744-752.
4. Xu Z, Pothula SP, Wilson JS, Apte MV. Pancreatic cancer and its stroma: a conspiracy theory. World J Gastroenterol. 2014; 20:11216-11229.
5. Das S, Batra SK. Pancreatic cancer metastasis: are we being pre-EMTed? Curr Pharm Des. 2015; 21:1249-1255.
6. Zhang Y, Wang Z, Gemeinhart RA. Progress in microRNA delivery. J Control Release. 2013; 172:962-974.
7. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009; 136:215-233.
8. Salmanidis M, Pillman K, Goodall G, Bracken C. Direct transcriptional regulation by nuclear microRNAs. Int J Biochem Cell Biol. 2014; 54:304-311.
9. Su Y, Li X, Ji W, Sun B, Xu C, Li Z, Qian G, Su C. Small molecule with big role: microRNAs in cancer metastatic microenvironments. Cancer Lett. 2014; 344:147-156.
10. Kita Y, Yonemori K, Osako Y, Baba K, Mori S, Maemura K, Natsugoe S. Noncoding RNA and colorectal cancer: its epigenetic role. J Hum Genet. 2017; 62:41-47.
11. Yonemori K, Kurahara H, Maemura K, Natsugoe S. MicroRNA in pancreatic cancer. J Hum Genet. 2016; 62:33-40.
12. Mataki H, Seki N, Mizuno K, Nohata N, Kamikawaji K, Kumamoto T, Koshizuka K, Goto Y, Inoue H. Dual-strand tumor-suppressor microRNA-145 (miR-145-5p and miR-145-3p) coordinately targeted MTDH in lung squamous cell carcinoma. Oncotarget. 2016; 7:72084-72098. doi: 10.18632/oncotarget.12290.
13. Osako Y, Seki N, Kita Y, Yonemori K, Koshizuka K, Kurozumi A, Omoto I, Sasaki K, Uchikado Y, Kurahara H, Maemura K, Natsugoe S. Regulation of MMP13 by antitumor microRNA-375 markedly inhibits cancer cell migration and invasion in esophageal squamous cell carcinoma. Int J Oncol. 2016; 49:2255-2264.
14. Goto Y, Kojima S, Nishikawa R, Kurozumi A, Kato M, Enokida H, Matsushita R, Yamazaki K, Ishida Y, Nakagawa M, Naya Y, Ichikawa T, Seki N. MicroRNA expression signature of castration-resistant prostate cancer: the microRNA-221/222 cluster functions as a tumour suppressor and disease progression marker. Br J Cancer. 2015; 113:1055-1065.
15. Goto Y, Kurozumi A, Nohata N, Kojima S, Matsushita R, Yoshino H, Yamazaki K, Ishida Y, Ichikawa T, Naya Y, Seki N. The microRNA signature of patients with sunitinib failure: regulation of UHRF1 pathways by microRNA-101 in renal cell carcinoma. Oncotarget. 2016; 7:59070-59086. doi: 10.18632/oncotarget.10887.
16. Koshizuka K, Nohata N, Hanazawa T, Kikkawa N, Arai T, Okato A, Fukumoto I, Katada K, Okamoto Y, Seki N. Deep sequencing-based microRNA expression signatures in head and neck squamous cell carcinoma: dual strands of pre-miR-150 as antitumor miRNAs. Oncotarget. 2017; 8:30288-30304. doi: 10.18632/oncotarget.16327.
17. Yonemori K, Seki N, Kurahara H, Osako Y, Idichi T, Arai T, Koshizuka K, Kita Y, Maemura K, Natsugoe S. ZFP36L2 promotes cancer cell aggressiveness and is regulated by antitumor microRNA-375 in pancreatic ductal adenocarcinoma. Cancer Sci. 2017; 108:124-135.
18. Guo J, Feng Z, Huang Z, Wang H, Lu W. MicroRNA-217 functions as a tumour suppressor gene and correlates with cell resistance to cisplatin in lung cancer. Mol Cells. 2014; 37:664-671.
19. Chen DL, Zhang DS, Lu YX, Chen LZ, Zeng ZL, He MM, Wang FH, Li YH, Zhang HZ, Pelicano H, Zhang W, Xu RH. microRNA-217 inhibits tumor progression and metastasis by downregulating EZH2 and predicts favorable prognosis in gastric cancer. Oncotarget. 2015; 6:10868-10879. doi: 10.18632/oncotarget.3451.
20. Zhang Q, Yuan Y, Cui J, Xiao T, Jiang D. MiR-217 promotes tumor proliferation in breast cancer via targeting DACH1. J Cancer. 2015; 6:184-191.
21. Zhao WG, Yu SN, Lu ZH, Ma YH, Gu YM, Chen J. The miR-217 microRNA functions as a potential tumor suppressor in pancreatic ductal adenocarcinoma by targeting KRAS. Carcinogenesis. 2010; 31:1726-1733.
22. Azevedo-Pouly AC, Sutaria DS, Jiang J, Elgamal OA, Amari F, Allard D, Grippo PJ, Coppola V, Schmittgen TD. miR-216 and miR-217 expression is reduced in transgenic mouse models of pancreatic adenocarcinoma, knockout of miR-216/miR-217 host gene is embryonic lethal. Funct Integr Genomics. 2016; 17:203-212.
23. Hall PA, Todd CB, Hyland PL, McDade SS, Grabsch H, Dattani M, Hillan KJ, Russell SE. The septin-binding protein anillin is overexpressed in diverse human tumors. Clin Cancer Res. 2005; 11:6780-6786.
24. Liang PI, Chen WT, Li CF, Li CC, Li WM, Huang CN, Yeh HC, Ke HL, Wu WJ, Chai CY. Subcellular localisation of anillin is associated with different survival outcomes in upper urinary tract urothelial carcinoma. J Clin Pathol. 2015; 68:1026-1032.
25. Wang S, Mo Y, Midorikawa K, Zhang Z, Huang G, Ma N, Zhao W, Hiraku Y, Oikawa S, Murata M. The potent tumor suppressor miR-497 inhibits cancer phenotypes in nasopharyngeal carcinoma by targeting ANLN and HSPA4L. Oncotarget. 2015; 6:35893-35907. doi: 10.18632/oncotarget.5651.
26. Zhou W, Wang Z, Shen N, Pi W, Jiang W, Huang J, Hu Y, Li X, Sun L. Knockdown of ANLN by lentivirus inhibits cell growth and migration in human breast cancer. Mol Cell Biochem. 2015; 398:11-19.
27. Nishioka C, Ikezoe T, Yang J, Nobumoto A, Tsuda M, Yokoyama A. Downregulation of miR-217 correlates with resistance of Ph(+) leukemia cells to ABL tyrosine kinase inhibitors. Cancer Sci. 2014; 105:297-307.
28. Xi S, Inchauste S, Guo H, Shan J, Xiao Z, Xu H, Miettenen M, Zhang MR, Hong JA, Raiji MT, Altorki NK, Casson AG, Beer DG, et al. Cigarette smoke mediates epigenetic repression of miR-217 during esophageal adenocarcinogenesis. Oncogene. 2015; 34:5548-5559.
29. Carmeci C, Thompson DA, Kuang WW, Lightdale N, Furthmayr H, Weigel RJ. Moesin expression is associated with the estrogen receptor-negative breast cancer phenotype. Surgery. 1998; 124:211-217.
30. Darnel AD, Behmoaram E, Vollmer RT, Corcos J, Bijian K, Sircar K, Su J, Jiao J, Alaoui-Jamali MA, Bismar TA. Fascin regulates prostate cancer cell invasion and is associated with metastasis and biochemical failure in prostate cancer. Clin Cancer Res. 2009; 15:1376-1383.
31. Chiyomaru T, Enokida H, Kawakami K, Tatarano S, Uchida Y, Kawahara K, Nishiyama K, Seki N, Nakagawa M. Functional role of LASP1 in cell viability and its regulation by microRNAs in bladder cancer. Urol Oncol. 2012; 30:434-443.
32. Ren G, Tian Q, An Y, Feng B, Lu Y, Liang J, Li K, Shang Y, Nie Y, Wang X, Fan D. Coronin 3 promotes gastric cancer metastasis via the up-regulation of MMP-9 and cathepsin K. Mol Cancer. 2012; 11:67.
33. Chiyomaru T, Enokida H, Tatarano S, Kawahara K, Uchida Y, Nishiyama K, Fujimura L, Kikkawa N, Seki N, Nakagawa M. miR-145 and miR-133a function as tumour suppressors and directly regulate FSCN1 expression in bladder cancer. Br J Cancer. 2010; 102:883-891.
34. Kinoshita T, Nohata N, Fuse M, Hanazawa T, Kikkawa N, Fujimura L, Watanabe-Takano H, Yamada Y, Yoshino H, Enokida H, Nakagawa M, Okamoto Y, Seki N. Tumor suppressive microRNA-133a regulates novel targets: moesin contributes to cancer cell proliferation and invasion in head and neck squamous cell carcinoma. Biochem Biophys Res Commun. 2012; 418:378-383.
35. Nishikawa R, Goto Y, Sakamoto S, Chiyomaru T, Enokida H, Kojima S, Kinoshita T, Yamamoto N, Nakagawa M, Naya Y, Ichikawa T, Seki N. Tumor-suppressive microRNA-218 inhibits cancer cell migration and invasion via targeting of LASP1 in prostate cancer. Cancer Sci. 2014; 105:802-811.
36. Mataki H, Enokida H, Chiyomaru T, Mizuno K, Matsushita R, Goto Y, Nishikawa R, Higashimoto I, Samukawa T, Nakagawa M, Inoue H, Seki N. Downregulation of the microRNA-1/133a cluster enhances cancer cell migration and invasion in lung-squamous cell carcinoma via regulation of Coronin1C. J Hum Genet. 2015; 60:53-61.
37. Oegema K, Savoian MS, Mitchison TJ, Field CM. Functional analysis of a human homologue of the Drosophila actin binding protein anillin suggests a role in cytokinesis. J Cell Biol. 2000; 150:539-552.
38. Giansanti MG, Bonaccorsi S, Gatti M. The role of anillin in meiotic cytokinesis of Drosophila males. J Cell Sci. 1999; 112:2323-2334.
39. Gbadegesin RA, Hall G, Adeyemo A, Hanke N, Tossidou I, Burchette J, Wu G, Homstad A, Sparks MA, Gomez J, Jiang R, Alonso A, Lavin P, et al. Mutations in the gene that encodes the F-actin binding protein anillin cause FSGS. J Am Soc Nephrol. 2014; 25:1991-2002.
40. Suzuki C, Daigo Y, Ishikawa N, Kato T, Hayama S, Ito T, Tsuchiya E, Nakamura Y. ANLN plays a critical role in human lung carcinogenesis through the activation of RHOA and by involvement in the phosphoinositide 3-kinase/AKT pathway. Cancer Res. 2005; 65:11314-11325.
41. Magnusson K, Gremel G, Ryden L, Ponten V, Uhlen M, Dimberg A, Jirstrom K, Ponten F. ANLN is a prognostic biomarker independent of Ki-67 and essential for cell cycle progression in primary breast cancer. BMC Cancer. 2016; 16:904.
42. Kurozumi A, Goto Y, Matsushita R, Fukumoto I, Kato M, Nishikawa R, Sakamoto S, Enokida H, Nakagawa M, Ichikawa T, Seki N. Tumor-suppressive microRNA-223 inhibits cancer cell migration and invasion by targeting ITGA3/ITGB1 signaling in prostate cancer. Cancer Sci. 2016; 107:84-94.
43. Gilcrease MZ. Integrin signaling in epithelial cells. Cancer Lett. 2007; 247:1-25.
44. Kinoshita T, Nohata N, Hanazawa T, Kikkawa N, Yamamoto N, Yoshino H, Itesako T, Enokida H, Nakagawa M, Okamoto Y, Seki N. Tumour-suppressive microRNA-29s inhibit cancer cell migration and invasion by targeting laminin-integrin signalling in head and neck squamous cell carcinoma. Br J Cancer. 2013; 109:2636-2645.
45. Hamidi H, Pietila M, Ivaska J. The complexity of integrins in cancer and new scopes for therapeutic targeting. Br J Cancer. 2016; 115:1017-1023.
46. Smith HW, Marshall CJ. Regulation of cell signalling by uPAR. Nat Rev Mol Cell Biol. 2010; 11:23-36.
47. Gorantla B, Asuthkar S, Rao JS, Patel J, Gondi CS. Suppression of the uPAR-uPA system retards angiogenesis, invasion, and in vivo tumor development in pancreatic cancer cells. Mol Cancer Res. 2011; 9:377-389.
48. Margheri F, Luciani C, Taddei ML, Giannoni E, Laurenzana A, Biagioni A, Chilla A, Chiarugi P, Fibbi G, Del Rosso M. The receptor for urokinase-plasminogen activator (uPAR) controls plasticity of cancer cell movement in mesenchymal and amoeboid migration style. Oncotarget. 2014; 5:1538-1553. doi: 10.18632/oncotarget.1754.
49. Pavon MA, Arroyo-Solera I, Cespedes MV, Casanova I, Leon X, Mangues R. uPA/uPAR and SERPINE1 in head and neck cancer: role in tumor resistance, metastasis, prognosis and therapy. Oncotarget. 2016; 7:57351-57366. doi: 10.18632/oncotarget.10344.
50. Brierley JD, Gospodarowicz MK, Wittekind C. TNM Classification of Malignant Tumours, 7th Edition. Chichester, West Sussex, United Kingdom: Wiley-Blackwell; 2010.
51. Fukumoto I, Kinoshita T, Hanazawa T, Kikkawa N, Chiyomaru T, Enokida H, Yamamoto N, Goto Y, Nishikawa R, Nakagawa M, Okamoto Y, Seki N. Identification of tumour suppressive microRNA-451a in hypopharyngeal squamous cell carcinoma based on microRNA expression signature. Br J Cancer. 2014; 111:386-394.
52. Itesako T, Seki N, Yoshino H, Chiyomaru T, Yamasaki T, Hidaka H, Yonezawa T, Nohata N, Kinoshita T, Nakagawa M, Enokida H. The microRNA expression signature of bladder cancer by deep sequencing: the functional significance of the miR-195/497 cluster. PLoS One. 2014; 9:e84311.
53. Anaya J. OncoRank: a pan-cancer method of combining survival correlations and its application to mRNAs, miRNAs, and lncRNAs. PeerJ Prepr. 2016; 1:e2574.
54. Anaya J. OncoLnc: linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. PeerJ Comput Sci. 2016; 2:e67.

