INTRODUCTION
MicroRNAs (miRNAs) are class of non-coding small RNAs which approximately range from 18-25 nucleotides in length. Mature miRNAs regulate a wide variety of target genes in post-transcriptional level by binding to the 3’-untranslated complementary sequence of messenger RNA (mRNA) [1, 2]. Thus, miRNAs play pivotal roles in gene expression and diverse biological processes, such as cell proliferation, cycle, apoptosis and differentiation [3–5]. Emerging studies have declared that miRNAs were proposed as predictive indicators for multiple human neoplasms due to the aberrant expression discrepancy between tumor tissues and normal tissues [6–8]. In 2002 Calin et al first reported the biological role of miR-15 and miR-16 in chronic lymphocytic leukemia, this was the first time to investigate the relationship between microRNA and cancer [9]. Additionally, miRNAs are generally classified into two categories: hazardous miRNAs which correlated with poor prognosis and protective miRNAs which are known as favorable survival predictors [10–12]. As a member of miRNAs, miR-148/152 family have been reported to develop prognostic role in multiple carcinomas.
The miR-148/152 family consists of three highly homologous members (miR-148a, miR-148b, and miR-152), of which ectopic expression was observed in multiple diseases such as: atherosclerosis, diabetes, and cancers [13–15]. The majority of studies have considered miR-148/152 family as a tumor suppressor and exerted anti-tumor effect in human neoplasms [16–18]. For instance, Qiu et al reported that miR-148a expression was down-regulated in gastric tumor tissues compared with non-tumor tissues. Evaluated expression of miR-148a significantly predicted favorable overall survival of patients with gastric cancer [19]. Ma et al detected the decreased miR-148a level in bladder carcinoma specimens and reduced miR-148a expression correlated with shorter survival time and increased recurrence risk [20]. In addition, downregulation of miR-148b was found to associate with poorer outcomes in patients with hepatocellular carcinoma [21], and high miR-152 expression developed a negative impact on recrudescence in NSCLC [22]. However, there still exists a series of investigations presenting an adverse function of miR-148/152 family, indicating that the correlation between them remains controversial. Kjersem et al have declared that evaluated miR-148a predicted pernicious progression and shoter overall survival of colorectal cancer patients [23]. Furthermore, Wang et al reported the onceogenic value of miR-152 in colorectal carcinoma, but failed to demonstrate a significant impact on prognosis [24]. Therefore, consensus has not been reached to the reliability of miR-148/152 family as prognostic indicators in various human neoplasms.
Considering the limitation of study scale, we sought to carry out this meta-analysis to summarize available findings and clarify the predictive significance of miR-148/152 family in malignancies prognoses.
RESULTS
Overview of eligible studies
A total of 267 studies from published database PubMed, EMBASE, and the Web of Science were identified to focus on the association between miR-148/152 family expression and multiple human malignancies. After a manual screening of titles and abstracts, 220 studies were excluded on account of the following reasons: review articles or letters, not human studies, unrelated to prognosis or outcomes, no relationship between miR-148/152 family and malignancies. For further quality evaluation of remaining candidates, 23 potential studies were excluded due to insufficient survival data, indirectly related to specific prognosis, incomprehensive or reduplicative data. Finally, 24 studies were considered to be included in the meta-analysis. The selection process of candidate studies are presented in detail in Figure 1.
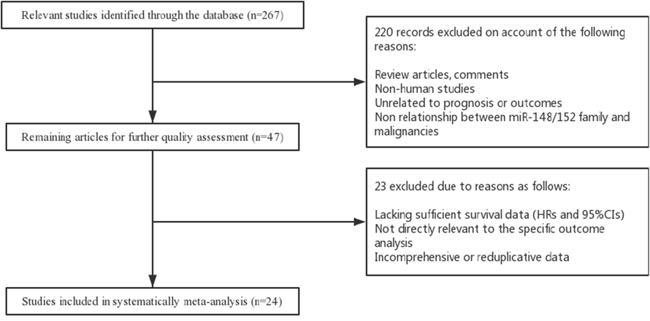
Figure 1: Flow diagram of literature search and selection process.
Dominant characteristics of included investigations were summarized in Table 1 and Table 2. For data extracted from the 24 eligible studies, 13 evaluated the relationship between miR-148a and human carcinomas, while 9 reported miR-148b and other 5 focused on miR-152. For survival analyses, 25 focused on patients OS/CSS and 8 reported DFS/RFS/PFS. Of these eligible studies, 20 evaluated Asian population while other 4 studies concentrated on Caucasian. In addition, the malignant neoplasms consisted of hepatocellular carcinoma (HCC), non-small cell lung cancer (NSCLC), colorectal cancer (CRC), and bladder cancer, ovarian cancer, gastric cancer and pancreatic cancer, skin cancer, endometrial serous adenocarcinoma and Osteosarcoma. All of the analyzed studies were retrospective except for one prospective. 14 studies focused on pathological type of adenocarcinoma (AdenoCA), 2 evaluated squamous carcinoma (SqCa), 3 reported (SqCa/AdenoCA) and 2 assessed AdenoCA/Mucinous, other 3 studies focused on epithelial carcinoma, transitional cell carcinoma and sarcoma, respectively. Quantitative real-time PCR (qRT-PCR) was widely used in all eligible studies to calculate miRNA-148/152 family expression.
Table 1: Main characteristics of eligible studies in the meta-analysis
First author, publication year |
MicroRNA type |
Case nationality |
Median or mean age |
Dominant ethnicity |
Study design |
Malignant disease |
Main type of pathology |
Detected sample |
Survival analysis |
Source of HR |
Maximum months of follow-up |
|---|---|---|---|---|---|---|---|---|---|---|---|
Qiu,2016 |
miR-148a |
China |
60 |
Asian |
R |
Gastric cancer |
AdenoCA |
Tissue |
OS |
Reported |
60 |
NG,2016 |
miR-148a |
China |
55 |
Asian |
R |
HCC |
AdenoCA |
Plasma |
OS/DFS |
Reported |
138 |
Wang,2016 |
miR-152 |
China |
60 |
Asian |
R |
CRC |
AdenoCA |
Tissue |
OS |
Reported |
54 |
Wang,2016 |
miR-148b |
China |
60 |
Asian |
R |
NSCLC |
SqCa/AdenoCA |
Tissue |
OS |
Reported |
60 |
Gong,2016 |
miR-148a |
China |
50 |
Asian |
R |
Ovarian cancer |
Epithelial carcinoma |
Plasma |
OS/RFS |
Reported |
60 |
Ma,2016 |
miR-148a |
China |
60 |
Asian |
R |
Bladder cancer |
Transitional cell carcinoma |
Tissue |
OS/RFS |
Reported/SC |
120 |
Ziari,2016 |
miR-148b |
Iran |
60 |
Asian |
R |
HCC |
AdenoCA |
Tissue |
OS |
Reported |
80 |
Wang,2016 |
miR-148a |
China |
51 |
Asian |
R |
HCC |
AdenoCA |
Serum |
OS |
Reported |
35 |
Wang,2016 |
miR-148b |
China |
51 |
Asian |
R |
HCC |
AdenoCA |
Serum |
OS |
Reported |
35 |
Wang,2016 |
miR-152 |
China |
51 |
Asian |
R |
HCC |
AdenoCA |
Serum |
OS |
Reported |
35 |
Tian,2015 |
miR-148a |
China |
60 |
Asian |
R |
Skin cancer |
SqCa |
Tissue |
OS |
Reported |
60 |
Wang,2015 |
miR-152 |
China |
19 |
Asian |
P |
Osteosarcoma |
Sarcoma |
Tissue |
OS |
Reported |
60 |
Hibino,2015 |
miR-148a |
Japan |
63 |
Asian |
R |
CRC |
AdenoCA |
Tissue |
CSS |
Reported |
60 |
Ghasemkhani, 2015 |
miR-148b |
Iran |
50 |
Asian |
R |
NSCLC |
SqCa/AdenoCA |
Tissue |
OS |
Reported |
60 |
Sadeghian,2015 |
miR-148b |
Iran |
60 |
Asian |
R |
HCC |
AdenoCA |
Tissue |
OS |
Reported |
60 |
Ge,2015 |
miR-148b |
China |
60 |
Asian |
R |
NSCLC |
SqCa/AdenoCA |
Tissue |
OS |
Reported |
60 |
Zhang,2015 |
miR-148b |
China |
50 |
Asian |
R |
HCC |
AdenoCA |
Tissue |
OS |
SC |
48 |
Zhang,2014 |
miR-148b |
China |
50 |
Asian |
R |
HCC |
AdenoCA |
Tissue |
OS |
Reported |
80 |
Sakamoto,2014 |
miR-148a |
Japan |
60 |
Asian |
R |
Gastric cancer |
AdenoCA |
Tissue |
OS |
Reported |
66 |
Kjersem,2014 |
miR-148a |
Norway |
60 |
Caucasian |
R |
CRC |
AdenoCA |
Plasma |
OS/PFS |
Reported |
NM |
Sanfiorenzo, 2013 |
miR-152 |
France |
65.1 |
Caucasian |
R |
NSCLC |
SqCa |
Plasma |
DFS |
Reported |
NM/SC |
Tsai,2013 |
miR-148a |
China |
65 |
Asian |
R |
CRC |
AdenoCA/Mucinous |
Serum |
OS/DFS |
SC |
67 |
Christensen,2013 |
miR-148a |
Denmark |
66 |
Caucasian |
R |
CRC |
AdenoCA/Mucinous |
Tissue |
DFS |
SC |
96 |
Zhao,2013 |
miR-148b |
China |
60 |
Asian |
R |
Pancreatic Cancer |
AdenoCA |
Tissue |
OS |
SC |
40 |
Schultz,2012 |
miR-148a |
Denmark |
64 |
Caucasian |
R |
PDAC |
AdenoCA |
Tissue |
OS |
Reported |
300 |
Schultz,2012 |
miR-148a |
Denmark |
64 |
Caucasian |
R |
A-AC |
AdenoCA |
Tissue |
OS |
Reported |
300 |
Hiroki,2009 |
miR-152 |
Japan |
64.9 |
Asian |
R |
Endometrial serous adenocarcinoma |
AdenoCA |
Tissue |
OS/DFS |
Reported |
64 |
Study design is described as prospective (P) or retrospective (R). miR-148a, microRNA-148a; miR-148b, microRNA-148b; miR-152, microRNA-152.
AdenoCA, adenocarcinoma; SqCa, squamous carcinoma; OS, overall survival; CSS, cause-specific survival; DFS, disease-free survival; RFS, recurrence-free survival; PFS, progression-free survival.
HCC, hepatocellular carcinoma; NSCLC, non-small cell lung cancer; CRC, colorectal cancer; PDAC, pancreatic ductal adenocarcinoma; A-AC, ampullary adenocarcinomas.
NM, not mentioned; SC, survival curve.
Table 2: HRs and 95% CIs of patient survival or cancer progression relating to MMPs expression in eligible studies
First author, publication year |
MicroRNA type |
Main assay method |
Cut-off value |
Case number |
OS/CSS |
DFS/RFS/PFS |
|||
|---|---|---|---|---|---|---|---|---|---|
High expression |
Low expression |
HR(95%CI)(U/M) |
P Value |
HR(95%CI)(U/M) |
P Value |
||||
Qiu,2016 |
miR-148a |
qRT-PCR |
median |
39 |
55 |
0.80(0.65-0.99)U |
0.045 |
NM |
NM |
NG,2016 |
miR-148a |
qRT-PCR |
NM |
31 |
31 |
0.45(0.03-6.25)M |
0.548 |
0.43 (0.02-7.57)M |
0.561 |
Wang,2016 |
miR-152 |
qRT-PCR |
median |
101 |
101 |
1.171(0.479-2.864)M |
0.73 |
NM |
NM |
Wang,2016 |
miR-148b |
qRT-PCR |
NM |
NM |
NM |
0.48(0.32-0.93)M |
0.003 |
NM |
NM |
Gong,2016 |
miR-148a |
qRT-PCR |
median |
55 |
47 |
0.589(0.41-0.85) M |
0.005 |
0.70 (0.49-1.O1)M |
0.058 |
Ma,2016 |
miR-148a |
qRT-PCR |
median |
56 |
70 |
0.60(0.38-0.96)U* |
0.005 |
0.07 (0.045-0.148)M |
<0.001 |
Ziari,2016 |
miR-148b |
qRT-PCR |
median |
NM |
NM |
0.452(0.110-0.699)M |
0.012 |
NM |
NM |
Wang,2016 |
miR-148a |
qRT-PCR |
median |
38 |
38 |
0.442(0.212-0.923)M |
0.03 |
NM |
NM |
Wang,2016 |
miR-148b |
qRT-PCR |
median |
38 |
38 |
0.709(0.343-1.462)M |
0.352 |
NM |
NM |
Wang,2016 |
miR-152 |
qRT-PCR |
median |
38 |
38 |
0.578(0.278-1.2)M |
0.141 |
NM |
NM |
Tian,2015 |
miR-148a |
qRT-PCR |
NM |
50 |
55 |
0.053(0.005-0.548)M |
0.014 |
NM |
NM |
Wang,2015 |
miR-152 |
qRT-PCR |
mean |
38 |
42 |
0.126(0.023-0.7010)M |
0.004 |
NM |
NM |
Hibino,2015 |
miR-148a |
qRT-PCR |
median |
16 |
33 |
0.226(0.054-0.679)M |
0.006 |
NM |
NM |
Ghasemkhani, 2015 |
miR-148b |
qRT-PCR |
median |
58 |
46 |
0.32(0.09-0.65)M |
0.021 |
NM |
NM |
Sadeghian, 2015 |
miR-148b |
qRT-PCR |
median |
NM |
NM |
0.378(0.19-0.57)M |
0.01 |
NM |
NM |
Ge,2015 |
miR-148b |
qRT-PCR |
median |
74 |
77 |
0.424(0.109-0.62)M |
0.011 |
NM |
NM |
Zhang,2015 |
miR-148b |
qRT-PCR |
median |
20 |
20 |
0.64(0.16-2.56)U* |
<0.01 |
NM |
NM |
Zhang,2014 |
miR-148b |
qRT-PCR |
median |
58 |
98 |
0.54(0.34-0.81)M |
0.002 |
NM |
NM |
Sakamoto,2014 |
miR-148a |
qRT-PCR |
median |
50 |
52 |
0.325(0.114-0.822)M |
0.0169 |
NM |
NM |
Kjersem,2014 |
miR-148a |
qRT-PCR |
median |
NM |
NM |
1.179(0.96-1.45)M |
0.118 |
1.290 (1.07-1.55)M |
0.007 |
Sanfiorenzo, 2013 |
miR-152 |
qRT-PCR |
mean |
NM |
NM |
NM |
NM |
0.333 (0.125-0.892)M |
0.029 |
Tsai,2013 |
miR-148a |
qRT-PCR |
mean |
55 |
55 |
0.58(0.29-1.17)U* |
0.0156 |
0.43(0.20-0.90)U* |
0.0006 |
Christensen, 2013 |
miR-148a |
qRT-PCR |
median |
20 |
26 |
NM |
NM |
0.78 (0.22-2.76)U* |
0.0236 |
Zhao,2013 |
miR-148b |
qRT-PCR |
median |
24 |
24 |
0.56(0.25-1.25)U* |
<0.05 |
NM |
NM |
Schultz,2012 |
miR-148a |
qRT-PCR |
median |
NM |
NM |
0.97(0.91-1.04)M |
0.54 |
NM |
NM |
Schultz,2012 |
miR-148a |
qRT-PCR |
median |
NM |
NM |
0.82(0.74-0.91)M |
<0.001 |
NM |
NM |
Hiroki,2009 |
miR-152 |
qRT-PCR |
median |
NM |
NM |
0.005(4.77E-5-0.440)M |
0.021 |
0.003 (4.37E-5-0.250)M |
0.01 |
The source of HR and 95% CI was extracted from survival curves or article reports. miR-148a, microRNA-148a; miR-148b, microRNA-148b; miR-152, microRNA-152.
*HR calculated from survival curves; SC, survival curve; U, univariate analysis; M, multivariate analysis; NM, not mentioned.
OS, overall survival; CSS, cause-specific survival; DFS, disease-free survival; RFS, recurrence-free survival; PFS, progression-free survival.
qRT-qPCR, reverse transcriptase-quantitative PCR.
Patients survival associated with miR-148/152 expression
For studies evaluating OS/CSS analysis, a random-effects model was performed due to significant heterogeneity (P<0.001, I2=71.9%). Our analyses indicated that high expression of miR-148/152 family could significantly predict a favorable OS/CSS for various human carcinomas, with a combined HR of 0.63 (95% CI: 0.54-0.74, Figure 2A). Furthermore, we carried out stratified analyses by classifying studies into subgroups. Results from subgroups suggested that miR-148a and miR-148b exerted enhanced OS/CSS, with a pooled HR of 0.76 (95% CI: 0.69-0.90) and 0.49 (95% CI: 0.39-0.61), while abnormal miR-152 expression developed no statistical impact (HR=0.40, 95% CI: 0.12-1.29; Figure 3A). In stratified analyses with cancer types, 6 studies reporting HCC and 3 reporting NSCLC indicated that miR-148/152 family were particularly associated with favorable OS/CSS (HCC: HR=0.5, 95% CI: 0.39-0.65; NSCLC: HR=0.43, 95% CI: 0.29-0.66; Figure 3B). Other 4 studies demonstrated that miR-148/152 family exerted no significant function on OS/CSS in CRC patients (HR=0.77, 95% CI: 0.42-1.41; Figure 3B), and 2 studies with gastric cancer obtained a similar result (HR=0.58, 95% CI: 0.25-1.35; Figure 3B). In addition, up-regulated miR-148/152 family correlated with superior OS/CSS in Asian (HR=0.53, 95% CI: 0.44-0.64) than that in Caucasian population (HR=0.96, 95% CI: 0.82-1.13; Figure 3C). Considering the limitation of investigation quantity, the prognostic significance of miR-148/152 family in other caner types still needed further confirmation.
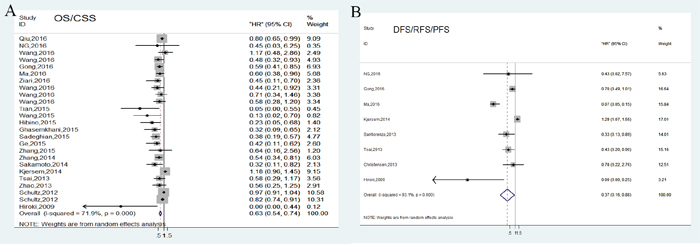
Figure 2: Forest plots of pooled analyses associated with miR-148/152 family expression. Survival data are reported as overall survival/cause-disease survival (OS/CSS) (A) and disease-free survival/recurrence-free survival/progression-free survival (DFS/RFS/PFS) (B).

Figure 3: Forest plots of subgroup analysis of the OS/CSS. (A) stratified by MicroRNA subgroup; (B) stratified by malignancy subgroup; (C) stratified by ethnic subgroup; (D) stratified by sample subgroup.
Tumor progression associated with miR-148/152 expression
In general, tumor progression was assessed by combining disease recurrence and metastasis. A total of 8 independent studies reported DFS/RFS/PFS analysis and revealed a protective significance of upregulated miR-148/152 family expression in multiple human neoplasms (HR=0.37, 95% CI: 0.16-0.88; Figure 2B). A random-effects model was applied to estimation due to a significant heterogeneity between studies (P<0.001, I2=93.1%). Stratified analyses indicated that miR-148/152 overexpression was a significant prediction for tumor recurrence and progression in tissues (HR=0.11, 95% CI: 0.01-0.98) but not in plasma/serum (HR=0.67, 95% CI: 0.38-1.18; Figure 4D). In ethnic subgroups, our analysis suggested that high miR148/152 expression correlated with favorable DFS/RFS/PFS in Asian population (HR=0.21, 95% CI: 0.06-0.81), but failed to obtain a significant consequence in Caucasian (HR=0.76, 95% CI: 0.31-1.87; Figure 4C). Results from other stratified analysis were presented in Figure 4A and 4B.
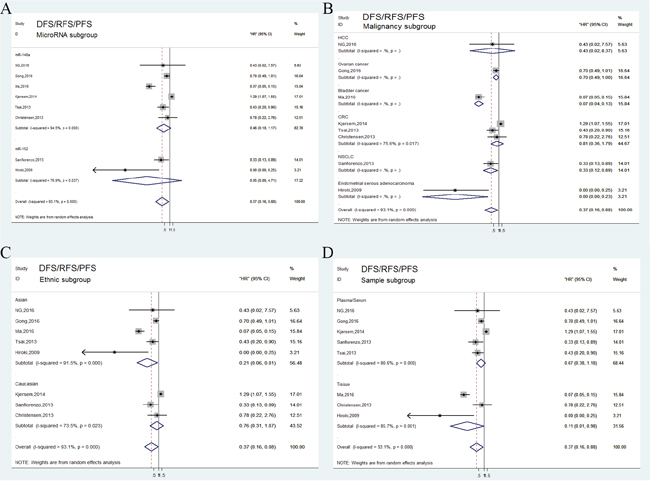
Figure 4: Forest plots of subgroup analysis of the DFS/RFS/PFS. (A) stratified by MicroRNA subgroup; (B) stratified by malignancy subgroup; (C) stratified by ethnic subgroup; (D) stratified by sample subgroup.
Sensitivity analyses
We also conducted sensitivity analyses by sequentially omitting individual studies to evaluate whether exclusion of any individual study indicated alterations in the results. The sensitive analyses from a random-effect model indicated the analyzed results was stable (Figure 5A, 5B).
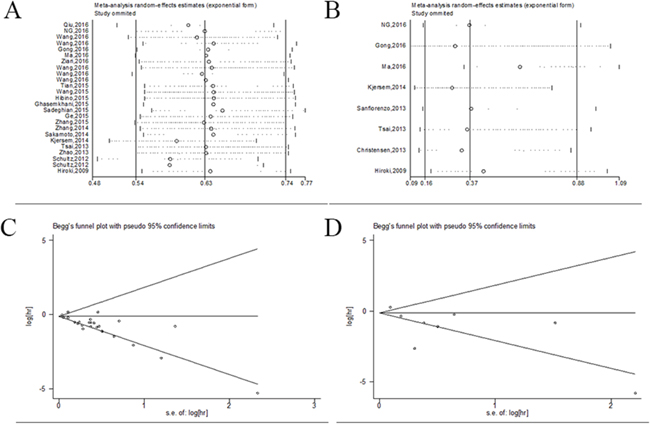
Figure 5: Sensitivity and bias analysis under specific model. (A) effect of individual studies on the pooled HR for OS/CSS; (B) effect of individual studies on the pooled HR for DFS/RFS/PFS; (C) Begg’s funnel plots of publication bias for OS/CSS. (D) Begg’s funnel plots of publication bias for DFS/RFS/PFS.
Publication bias
Publication bias of the included investigations was performed by funnel plots and Begg’s tests. In the combined prognostic analysis as determined, the funnel plots were symmetric and P values were 0.118 for OS/CSS and 0.536 for DFS/RFS/PFS (P>0.05), respectively (Figure 5C, 5D). Therefore, no significant publication bias was observed in the meta-analysis.
DISCUSSION
Recently, exhaustive efforts have been invested in identifying prognostic biomarkers for patients with multiple malignancies. Mounting evidence has indicated that microRNAs play crucial roles in carcinogenesis and cancer progression, which are closely associated with various biological activities such as cell proliferation, cycle, invasion, and metastasis [25]. MicroRNAs are more stably expressed in multiple specimen samples compared with mRNAs and proteins, which can be accurately quantified by qRT-PCR [26, 27]. Therefore, an increasing number of investigations have verified microRNAs as potential targets for clinical treatment, as well as promising biomarkers for cancer prognoses [28–30].
Currently, miR-148/152 family members include miR-148a, miR-148b, and miR-152, of which the three share a common seeds sequence in domains [31]. Previous studies have confirmed that down-regulation of miR-148/152 family is associated with unfavorable survival and prognostic outcomes of patients with malignancies [32–34]. Downregulated miR-148a expression was found in gastric tumor compared with non-neoplastic mucosa, and this was correlated with advanced tumor invasiveness and poor prognosis by targeting MMP7 [35]. Zhang et al found that decreased miR-148b expression in HCC predicted poor prognosis, and downregulated miR-148b significantly enhanced cancer progression with advanced vein invasion and TNM stage [36]. In addition, Wang et al detected the circulating miR-148a, miR-148b, and miR-152 and revealed that loss of miR-148a expression independently predicted a shorter overall time in patients with HCC than miR-148b and miR-152 [37]. Even though, certain investigations have presented contradictory results. For instance, loss level of miR-148 was found to predict longer period of recurrence and favorable overall survival in esophageal adenocarcinoma patients [38]. Ma et al confirmed that miR-148a high expression was an independent indicator for unfavorable overall survival and disease-specific survival, respectively [39]. The observation of these oncogenic role of miR-148/152 family in multiple cancers might cast doubt on its dominant anti-tumor effects. Despite these controversial results, miR-148/152 family was still an attractive biomarker for considerable prognostic significance.
The prognostic role of miR-148/152 family in human neoplasms may partly attributed to its underlying molecular mechanism, as well as dissimilar biological function. In HCC patients, miR-148a overexpression was found to suppress cell invasion and affects prognosis by directly targeting sphingosine-1-phosphate receptor 1(S1PR1) [40, 41]. He et al verified that miR-148a inhibited NSCLC cell proliferation and invasion activity through silencing signal transducer and activator of transcription 3 (STAT3), which highlighted miR-148a/STAT3 axis as a potential target for clinical treatment with NSCLC patients [42]. Kim et al also found that miR-148a acted as a tumor suppressor and holds vital potential for renal carcinoma therapy by directly targeting Rab14 [43]. Furthermore, miR-148/152 family was found to involved into DNA methylation by interacting with DNA methyltransferase enzyme 1 (DNMT1) in many malignancies types. Zhu et al found that DNMT1 overexpression inactivated miR-148a by hypermethylation of DNA in gastric cancer. Inhibition of miR-148a might promote DNA hypermethylation in case of the overexpression of DNMT1 [44]. In a breast cancer study, Xu et al that high DNMT1 expression was responsible for hypermethylation of miR-148a and miR-152 promoters. Besides, DNMT1 was conversely associated with miR-148a/152 expression, which highlighted a potential miR-148a/152-DNMT1 regulatory framework might exist in breast cancer [45]. Based on these underlying mechanism, we concluded that miR-148/152 family in specific cancer category might induce particular biological behaviors.
In this meta-analysis, we first collected available data from published studies to assess the prognostic significance of miR-148/152 family in multiple human malignancies. Subgroup, sensitivity, and heterogeneity analysis were conducted to explore the effects of main characteristics in relevant studies. Results from OS/CSS analysis indicated that up-regulated miR-148/152 family could predict favorable outcomes with a pooled HR of 0.63 (95% CI: 0.54-0.74). Additionally, the pooled outcome in the DFS/RFS/PFS analysis indicated that increased miRNA-148/152 expression is predictive of slower cancer progression (HR=0.37, 95% CI: 0.16-0.88). In stratified analysis, we found that upregulated miR-148/152 family predicted superior OS/CSS in Asian (HR=0.53, 95% CI: 0.44-0.64), but analysis in Caucasian population failed to obtain the significance (HR=0.96, 95% CI: 0.82-1.13). Similar outcomes of DFS/RFS/PFS analysis in ethnic subgroups was observed that aberrant miR-148/152 expression contributed to favorable disease progression in Asian population (HR=0.21, 95% CI: 0.06-0.81), but not in Caucasian (HR=0.76, 95% CI: 0.31-1.87). Previous investigations have confirmed that specifc miRNAs emerged diverse expression levels and develop particular survival impact in multiple ethnic groups [46, 47]. The occurrence of these discrepancies might be caused by the difference in environmental exposures and genetic backgrounds.
Admittedly, miR-148a, miR-148b, and miR-152 are the three members of the miR-148/152 family with the same seed sequence, of which are pivotal for binding to target mRNAs. In analysis of microRNA subgroups, our results demonstrated that miR-148a and miR-148b promoted favorable OS/CSS (HR=0.76, 95% CI: 0.69-0.90) and (HR=0.49, 95% CI: 0.39-0.61), nevertheless abnormal miR-152 expression exerted no statistical significance (HR=0.40, 95% CI: 0.12-1.29). Diverse prognostic values between miR-148a/b and miR-152 may attributed to different domains of the three, even though they possessed the same seed sequence. In addition, the deficiencies of studies focusing on miR-152 and cancer outcomes also accounted to some extent. Furthermore, malignancies species also had a considerable impact on the prognostic role of miR-148/152 family. Seven survival data from OS/CSS analysis indicated that miR-148/152 family play a vital role in overall survival for patients with HCC (HR=0.51, 95% CI: 0.39-0.65), revealing the independent value of miR-148/152 family in HCC [21, 34, 36, 48-50]. Although analyses of other neoplasms also obtained a statistical outcome, these results need to be further confirmed due to the deficiency of studies. Interesting, stratification analysis of different detected specimens suggested that miR-148/152 served as a significant indicator for tumor recurrence and progression in tissues (HR=0.11, 95% CI: 0.01-0.98) other than plasma/serum (HR=0.67, 95% CI: 0.38-1.18). These might origin from the specificity of diverse samples in various neoplasm and limitations of lacking more researches. To summarize, our meta-analysis indicated that detection of abnormal miR-148/152 family levels is of great significance in predicting prognosis of various human malignancies.
What can not be overlooked is the existence of heterogeneity when accounting for the results of this meta-analysis [51]. As is determined by Stata software, heterogeneity and publication bias of eligible subjects might affect the stability of our demonstration. In this meta-analysis, a significant heterogeneity was observed when we carried out OS/CSS analysis of miR-148/152 family, as well as comparison for DFS/RFS/PFS. All of these might weaken the pooled results of meta-analysis and cannot explicitly states the prognostic status of miR-148/152 family. Based on respective results above, we performed stratified analysis to minimize the impact of heterogeneity by classifying studies into subgroups of microRNA types, dominant ethnicity, malignant diseases, detected samples and pathological categories. Partly decreasing heterogeneity was presented in some subgroups, even though they still existed in the results. Furthermore, sensitivity analysis was also carried out to strengthen the conclusion of the meta-analysis. We found that exclusion of individual studies brought about quite tiny change of estimated pooled HRs. No evidence of significant publication bias was noted in this meta-analysis, indicating our analyzed results were credible.
Although elaborate check was conducted along with statistical analysis, our conclusion still needs further refinement for the following accounts. Firstly, all investigations included were published in English except one Chinese article, which might induced English language bias in pooled results [52, 53]. Secondly, the number of eligible studies reporting DFS/RFS/PFS was not sufficiently enough for a comprehensive analysis. Thirdly, a recognized miR-148/152 expression level could hardly to achieve even the majority of articles regarded the median/mean points as the cut-off value. What’s more, only Asian and Caucasian population were analyzed in this meta-analysis, which might weaken the meta-analytic worth to some extent. Considering these limitations, the significance of miR-148/152 family as a prognostic indicator in multiple human malignancies might be overestimated. Our results should be interpreted minutely and need further confirmation.
In summary, this meta-analysis demonstrates that miR-148/152 overexpression can significantly predict favorable prognostic outcomes in diverse human neoplasms, particularly in Asian population and tissues specimens. Besides, miR-148a/b are promising biomarkers for predicting patients overall outcomes than miR-152. Taking insufficient evidence into account, in order to get a better evaluation of the prognostic role in patients with malignancies, further large-scale researches and clinical studies are needed for further convince.
MATERIALS AND METHODS
Search strategy
We conducted this meta-analysis in accordance with the standard guidelines of the Meta-analysis of Observational Studies in Epidemiology group (MOOSE) [54]. A literature search through online databases such as PubMed, Embase, and Web of Science were performed up to March 2017, using the following keywords (“microRNA-148a” or “miR-148a” or “microRNA-148b” or “miR-148b” or “microRNA-152” or “miR-152”) and (“cancer” or “carcinoma” or “Neoplasm” or “Tumor”) and (“prognostic” or “prognosis” or “survival” or “outcome” or “recurrence” or “relapse”). Eligible studies to be included in this analysis should meet the following criteria: (1) studies exploring various human malignancies; (2) a relationship between miR-148/152 family and cancer prognosis. With the aim to supplement our literature search, reference lists of eligible studies were screened for additional publications.
Quality assessment
In order to evaluate the quality of all included studies, we used a critical review checklist of the Dutch Cochrane Centre proposed by MOOSE. The key points of the quality assessment included the following: (1) origin of country and definition of study population, (2) clear microRNA subtypes and carcinoma classifications, (3) the study design and cut-off value of miR-148/152 family, (4) detected samples and pathology, (5) description of outcomes and follow-up period of patients. Studies without specifying the points mentioned above were excluded to maintain the quality of the meta-analysis. A flow diagram of the study selection process is presented in Figure 1.
Data extraction
Two investigators (Chen-kui Miao and Jian-zhong Zhang) independently identified all eligible studies and extracted relevant data to rule out any discrepancy. Following data elements were included and recorded: (1) the first authors’ names, publication year, and nationality of study population, (2) microRNA type, (3) dominant ethnicity and malignant types, (4) detected samples and pathology, (5) main assay method and cut-off definition, (6) following up duration and (7) HRs associated with evaluated miR-148/152 for OS/CSS and DFS/RFS/PFS along with 95% CIs and P values. If HRs and 95% CIs were only available in Kaplan-Meier curves, data were extracted from graphical survival plots using Engauge Digitizer version 4.1 [55, 56].
Statistical analysis
Cochran’s Q-test and Higgins I2 statistics (I2) were carried out to test the heterogeneity of pooled HRs. P<0.05 was considered statistically significant. The fixed-effects model (Mantel-Haenszel method) or the random-effects model (DerSimonian-Laird method) was performed for analysis according to the heterogeneity of all eligible investigations. If the heterogeneity was considered significantly at P<0.05 or the percentage of I2 was greater than 50%, a random-effects model was applied to calculate the pooled HR, otherwise a fixed-effects model was conducted. In addition, we also executed stratified analyses upon similar characteristics to minimize the sources of heterogeneity. Publication bias was assessed by using Begg’s test and Egger’s test [57, 58]. All above statistical calculations were conducted by using Stata version 12.0 (Stata Corporation, College Station, TX, USA).
ACKNOWLEDGMENTS AND FUNDING
The study was supported by National Natural Science Foundation of China (grant numbers 81270685, 81672531 and 81372757), Six talent peak project in jiangsu province (WSN-011) and Project of Nanjing Science and Technology Committee (201605001).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Carthew RW, Sontheimer EJ. Origins and Mechanisms of miRNAs and siRNAs. Cell. 2009; 136:642-655.
2. Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993; 75:843-854.
3. Yang W, Lee DY, Ben-David Y. The roles of microRNAs in tumorigenesis and angiogenesis. Int J Physiol Pathophysiol Pharmacol. 2011; 3:140-155.
4. Hulf T, Sibbritt T, Wiklund ED, Patterson K, Song JZ, Stirzaker C, Qu W, Nair S, Horvath LG, Armstrong NJ, Kench JG, Sutherland RL, Clark SJ. Epigenetic-induced repression of microRNA-205 is associated with MED1 activation and a poorer prognosis in localized prostate cancer. Oncogene. 2013; 32:2891-2899.
5. Markou A, Yousef GM, Stathopoulos E, Georgoulias V, Lianidou E. Prognostic significance of metastasis-related microRNAs in early breast cancer patients with a long follow-up. Clin Chem. 2014; 60:197-205.
6. Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, et al. MicroRNA expression profiles classify human cancers. Nature. 2005; 435:834-838.
7. Karakatsanis A, Papaconstantinou I, Gazouli M, Lyberopoulou A, Polymeneas G, Voros D. Expression of microRNAs, miR-21, miR-31, miR-122, miR-145, miR-146a, miR-200c, miR-221, miR-222, and miR-223 in patients with hepatocellular carcinoma or intrahepatic cholangiocarcinoma and its prognostic significance. Mol Carcinog. 2013; 52:297-303.
8. Wang QZ, Xu W, Habib N, Xu R. Potential uses of microRNA in lung cancer diagnosis, prognosis, and therapy. Curr Cancer Drug Targets. 2009; 9:572-594.
9. Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K, Rassenti L, Kipps T, Negrini M, et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA. 2002; 99:15524-15529.
10. Haapa-Paananen S, Chen P, Hellstrom K, Kohonen P, Hautaniemi S, Kallioniemi O, Perala M. Functional profiling of precursor MicroRNAs identifies MicroRNAs essential for glioma proliferation. Plos One. 2013; 8:e60930.
11. Campayo M, Navarro A, Vinolas N, Diaz T, Tejero R, Gimferrer JM, Molins L, Cabanas ML, Ramirez J, Monzo M, Marrades R. Low miR-145 and high miR-367 are associated with unfavourable prognosis in resected nonsmall cell lung cancer. Eur Respir J. 2013; 41:1172-1178.
12. Lee HK, Bier A, Cazacu S, Finniss S, Xiang C, Twito H, Poisson LM, Mikkelsen T, Slavin S, Jacoby E, Yalon M, Toren A, Rempel SA, et al. MicroRNA-145 is downregulated in glial tumors and regulates glioma cell migration by targeting connective tissue growth factor. Plos One. 2013; 8:e54652.
13. Bidzhekov K, Gan L, Denecke B, Rostalsky A, Hristov M, Koeppel TA, Zernecke A, Weber C. microRNA expression signatures and parallels between monocyte subsets and atherosclerotic plaque in humans. Thromb Haemost. 2012; 107:619-625.
14. Nielsen LB, Wang C, Sorensen K, Bang-Berthelsen CH, Hansen L, Andersen ML, Hougaard P, Juul A, Zhang CY, Pociot F, Mortensen HB. Circulating levels of microRNA from children with newly diagnosed type 1 diabetes and healthy controls: evidence that miR-25 associates to residual beta-cell function and glycaemic control during disease progression. Exp Diabetes Res. 2012; 2012:896362.
15. Zhou X, Zhao F, Wang ZN, Song YX, Chang H, Chiang Y, Xu HM. Altered expression of miR-152 and miR-148a in ovarian cancer is related to cell proliferation. Oncol Rep. 2012; 27:447-454.
16. Gong L, Wang C, Gao Y, Wang J. Decreased expression of microRNA-148a predicts poor prognosis in ovarian cancer and associates with tumor growth and metastasis. Biomed Pharmacother. 2016; 83:58-63.
17. Tian Y, Wei W, Li L, Yang R. Down-Regulation of miR-148a Promotes Metastasis by DNA Methylation and is Associated with Prognosis of Skin Cancer by Targeting TGIF2. Med Sci Monit. 2015; 21:3798-3805.
18. Tsai HL, Yang IP, Huang CW, Ma CJ, Kuo CH, Lu CY, Juo SH, Wang JY. Clinical significance of microRNA-148a in patients with early relapse of stage II stage and III colorectal cancer after curative resection. Transl Res. 2013; 162:258-268.
19. Qiu X, Zhu H, Liu S, Tao G, Jin J, Chu H, Wang M, Tong N, Gong W, Zhao Q, Qiang F, Zhang Z. Expression and prognostic value of microRNA-26a and microRNA-148a in gastric cancer. J Gastroenterol Hepatol. 2016; 32:819-827.
20. Ma L, Xu Z, Xu C, Jiang X. MicroRNA-148a represents an independent prognostic marker in bladder cancer. Tumour Biol. 2016; 37:7915-7920.
21. Ziari K, Zarea M, Gity M, Fayyaz AF, Yahaghi E, Darian EK, Hashemian AM. Downregulation of miR-148b as biomarker for early detection of hepatocellular carcinoma and may serve as a prognostic marker. Tumour Biol. 2016; 37:5765-5768.
22. Sanfiorenzo C, Ilie MI, Belaid A, Barlesi F, Mouroux J, Marquette CH, Brest P, Hofman P. Two panels of plasma microRNAs as non-invasive biomarkers for prediction of recurrence in resectable NSCLC. Plos One. 2013; 8:e54596.
23. Kjersem JB, Ikdahl T, Lingjaerde OC, Guren T, Tveit KM, Kure EH. Plasma microRNAs predicting clinical outcome in metastatic colorectal cancer patients receiving first-line oxaliplatin-based treatment. Mol Oncol. 2014; 8:59-67.
24. Wang Y, Yuan W, Ma X, Ma J. [Expression of microRNA-152 in colorectal cancer and its relationship with prognosis]. [Article in Chinese]. Zhonghua Zhong Liu Za Zhi. 2016; 38:763-766.
25. Hong L, Han Y, Li S, Yang J, Zheng J, Zhang H, Zhao Q, Wu K, Fan D. The malignant phenotype-associated microRNA in gastroenteric, hepatobiliary and pancreatic carcinomas. Expert Opin Biol Ther. 2010; 10:1693-1701.
26. Ferracin M, Negrini M. Micromarkers 2.0: an update on the role of microRNAs in cancer diagnosis and prognosis. Expert Rev Mol Diagn. 2015; 15:1369-1381.
27. Kim DJ, Linnstaedt S, Palma J, Park JC, Ntrivalas E, Kwak-Kim JY, Gilman-Sachs A, Beaman K, Hastings ML, Martin JN, Duelli DM. Plasma components affect accuracy of circulating cancer-related microRNA quantitation. J Mol Diagn. 2012; 14:71-80.
28. Ge YZ, Wu R, Xin H, Zhu M, Lu TZ, Liu H, Xu Z, Yu P, Zhao YC, Li MH, Hu ZK, Zhao Y, Zhong B, et al. A tumor-specific microRNA signature predicts survival in clear cell renal cell carcinoma. J Cancer Res Clin Oncol. 2015; 141:1291-1299.
29. Yu SL, Chen HY, Chang GC, Chen CY, Chen HW, Singh S, Cheng CL, Yu CJ, Lee YC, Chen HS, Su TJ, Chiang CC, Li HN, et al. MicroRNA signature predicts survival and relapse in lung cancer. Cancer Cell. 2008; 13:48-57.
30. Landi MT, Zhao Y, Rotunno M, Koshiol J, Liu H, Bergen AW, Rubagotti M, Goldstein AM, Linnoila I, Marincola FM, Tucker MA, Bertazzi PA, Pesatori AC, et al. MicroRNA expression differentiates histology and predicts survival of lung cancer. Clin Cancer Res. 2010; 16:430-441.
31. Yang JS, Li BJ, Lu HW, Chen Y, Lu C, Zhu RX, Liu SH, Yi QT, Li J, Song CH. Serum miR-152, miR-148a, miR-148b, and miR-21 as novel biomarkers in non-small cell lung cancer screening. Tumour Biol. 2015; 36:3035-3042.
32. Schultz NA, Andersen KK, Roslind A, Willenbrock H, Wojdemann M, Johansen JS. Prognostic microRNAs in cancer tissue from patients operated for pancreatic cancer--five microRNAs in a prognostic index. World J Surg. 2012; 36:2699-2707.
33. Hiroki E, Akahira J, Suzuki F, Nagase S, Ito K, Suzuki T, Sasano H, Yaegashi N. Changes in microRNA expression levels correlate with clinicopathological features and prognoses in endometrial serous adenocarcinomas. Cancer Sci. 2010; 101:241-249.
34. Ng KT, Lo CM, Wong N, Li CX, Qi X, Liu XB, Geng W, Yeung OW, Ma YY, Chan SC, Man K. Early-phase circulating miRNAs predict tumor recurrence and survival of hepatocellular carcinoma patients after liver transplantation. Oncotarget. 2016; 7:19824-19839. doi: 10.18632/oncotarget.7627.
35. Sakamoto N, Naito Y, Oue N, Sentani K, Uraoka N, Zarni OH, Yanagihara K, Aoyagi K, Sasaki H, Yasui W. MicroRNA-148a is downregulated in gastric cancer, targets MMP7, and indicates tumor invasiveness and poor prognosis. Cancer Sci. 2014; 105:236-243.
36. Zhang Z, Zheng W, Hai J. MicroRNA-148b expression is decreased in hepatocellular carcinoma and associated with prognosis. Med Oncol. 2014; 31:984.
37. Wang F, Ying H, He B, Pan Y, Sun H, Wang S. Circulating miR-148/152 family as potential biomarkers in hepatocellular carcinoma. Tumour Biol. 2016; 37:4945-4953.
38. Hezova R, Kovarikova A, Srovnal J, Zemanova M, Harustiak T, Ehrmann J, Hajduch M, Svoboda M, Sachlova M, Slaby O. Diagnostic and prognostic potential of miR-21, miR-29c, miR-148 and miR-203 in adenocarcinoma and squamous cell carcinoma of esophagus. Diagn Pathol. 2015; 10:42.
39. Ma L, Xu Z, Xu C, Jiang X. MicroRNA-148a represents an independent prognostic marker in bladder cancer. Tumour Biol. 2016; 37:7915-7920.
40. Zhang SL, Liu L. microRNA-148a inhibits hepatocellular carcinoma cell invasion by targeting sphingosine-1-phosphate receptor 1. Exp Ther Med. 2015; 9:579-584.
41. Heo MJ, Kim YM, Koo JH, Yang YM, An J, Lee SK, Lee SJ, Kim KM, Park JW, Kim SG. microRNA-148a dysregulation discriminates poor prognosis of hepatocellular carcinoma in association with USP4 overexpression. Oncotarget. 2014; 5:2792-2806. doi: 10.18632/oncotarget.1920.
42. He M, Xue Y. MicroRNA-148a suppresses proliferation and invasion potential of non-small cell lung carcinomas via regulation of STAT3. Onco Targets Ther. 2017; 10:1353-1361.
43. Kim EA, Kim TG, Sung EG, Song IH, Kim JY, Doh KO, Lee TJ. miR-148a increases the sensitivity to cisplatin by targeting Rab14 in renal cancer cells. Int J Oncol. 2017; 50:984-992.
44. Zhu A, Xia J, Zuo J, Jin S, Zhou H, Yao L, Huang H, Han Z. MicroRNA-148a is silenced by hypermethylation and interacts with DNA methyltransferase 1 in gastric cancer. Med Oncol. 2012; 29:2701-2709.
45. Xu Q, Jiang Y, Yin Y, Li Q, He J, Jing Y, Qi YT, Xu Q, Li W, Lu B, Peiper SS, Jiang BH, Liu LZ. A regulatory circuit of miR-148a/152 and DNMT1 in modulating cell transformation and tumor angiogenesis through IGF-IR and IRS1. J Mol Cell Biol. 2013; 5:3-13.
46. Bovell LC, Shanmugam C, Putcha BD, Katkoori VR, Zhang B, Bae S, Singh KP, Grizzle WE, Manne U. The prognostic value of microRNAs varies with patient race/ethnicity and stage of colorectal cancer. Clin Cancer Res. 2013; 19:3955-3965.
47. Huang RS, Gamazon ER, Ziliak D, Wen Y, Im HK, Zhang W, Wing C, Duan S, Bleibel WK, Cox NJ, Dolan ME. Population differences in microRNA expression and biological implications. RNA Biol. 2011; 8:692-701.
48. Wang F, Ying H, He B, Pan Y, Sun H, Wang S. Circulating miR-148/152 family as potential biomarkers in hepatocellular carcinoma. Tumour Biol. 2016; 37:4945-4953.
49. Sadeghian Y, Kamyabi-Moghaddam Z, Nodushan SM, Khoshbakht S, Pedram B, Yahaghi E, Mokarizadeh A, Mohebbi M. RETRACTED ARTICLE: Profiles of tissue microRNAs; miR-148b and miR-25 serve as potential prognostic biomarkers for hepatocellular carcinoma. Tumour Biol. 2015 Jul 25. [Epub ahead of print].
50. Zhang JG, Shi Y, Hong DF, Song M, Huang D, Wang CY, Zhao G. MiR-148b suppresses cell proliferation and invasion in hepatocellular carcinoma by targeting WNT1/beta-catenin pathway. Sci Rep. 2015; 5:8087.
51. Boccia S, De Feo E, Galli P, Gianfagna F, Amore R, Ricciardi G. A systematic review evaluating the methodological aspects of meta-analyses of genetic association studies in cancer research. Eur J Epidemiol. 2010; 25:765-775.
52. Gregoire G, Derderian F, Le Lorier J. Selecting the language of the publications included in a meta-analysis: is there a Tower of Babel bias? J Clin Epidemiol. 1995; 48:159-163.
53. Moher D, Fortin P, Jadad AR, Juni P, Klassen T, Le Lorier J, Liberati A, Linde K, Penna A. Completeness of reporting of trials published in languages other than English: implications for conduct and reporting of systematic reviews. Lancet. 1996; 347:363-366.
54. Stroup DF, Berlin JA, Morton SC, Olkin I, Williamson GD, Rennie D, Moher D, Becker BJ, Sipe TA, Thacker SB. Meta-analysis of observational studies in epidemiology: a proposal for reporting. Meta-analysis Of Observational Studies in Epidemiology (MOOSE) group. Jama. 2000; 283:2008-2012.
55. Williamson PR, Smith CT, Hutton JL, Marson AG. Aggregate data meta-analysis with time-to-event outcomes. Stat Med. 2002; 21:3337-3351.
56. DerSimonian R, Laird N. Meta-analysis in clinical trials revisited. Contemp Clin Trials. 2015; 45:139-145.
57. Egger M, Smith GD. Bias in location and selection of studies. BMJ. 1998; 316:61-66.
58. Egger M, Davey SG, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629-634.

