INTRODUCTION
Urothelial carcinoma of the bladder is one of most common malignancy all over the world and the most common urologic tumors in China [1, 2]. The incidence and mortality of bladder cancer have been significantly increased in the past decade [3–6]. Despite improvements in current clinical treatment such as surgery, adjuvant chemoradiotherapies, and immunological therapy, the prognosis of patients has not been significantly improved [7–11]. Patient’s prognosis is closely related to the stage of the disease, but there are no efficient methods for diagnosis at early stage [12, 13]. Therefore, more sensitive and specific markers for diagnosis at early stage and more efficient and safer treatments are urgently needed [14].
LncRNAs are a class of noncoding RNAs which are greater than 200 nucleotides in length and limited coding potential [15–17]. The rapid development of human genomics has highlighted the important role of non-coding RNAs in diverse biological processes of cancer [18–20]. Recent emerging evidences have shown that lncRNAs play key roles in development and progression of bladder cancer, such as UCA-1, PVT-1, MALAT1, SPRY4-IT1, PANDAR and etc [21–30]. CCEPR (cervical carcinoma expressed PCNA regulatory lncRNA, GenBank number AK055418) is a novel identified lncRNA with 2504 nucleotides in length and localized at the chromosome 10 [20]. Recently, CCEPR originally was identified as a powerful tumor biomarker for cervical cancer [31]. However, its biological function in bladder cancer development is still completely unknown.
Although CCERP has been shown to serve as an oncogene, the underlying molecular mechanism of CCERP in tumorigenesis remains to be clarified. The function of lncRNAs usually relies on the proteins that they interact with. For example, the lncRNA HOTTIP has been reported to directly interact with the WDR5 protein and target WDR5/MLL complexes [32]. The lncRNA PANDAR has also been reported to interact with the NF-YA to repress gene regulation [33]. And the lncRNA HNF1A-AS1 has also been reported to mediate the binding of DNMT1 to E-cadherin [26, 34]. Yang and colleagues have provided evidence that CCEPR associates with PCNA (proliferating cell nuclear antigen) mRNA, consequently increases the expression level of PCNA [31]. We speculated that CCERP may play a similar role in bladder cancer cell.
In this study, we found that the expression of CCEPR is significantly increased in bladder cancer tissues and cell lines, moreover, CCEPR promotes proliferation and suppresses apoptosis of bladder cancer cells in vitro. Our results suggested that CCEPR plays an key role and may serve as a promising diagnostic and therapeutic target for bladder cancer.
RESULTS
The expression of CCEPR is increased in bladder cancer
The expression of CCEPR in bladder cancer tissues and cell lines was detected by qRT-PCR. CCEPR mRNA levels were signifcantly up-regulated in bladder cancer tissues compared to corresponding non-tumor tissues (Figure 1A, 1B, 1C). Furthermore, elevated expression of CCEPR was positively associated with advanced TNM stage(Figure 1D). CCEPR mRNA levels was up-regulated in bladder cancer cell lines (Figure 1E). Statistical results and clinicopathological features of 55 patients are shown in Table 1 and Supplementary Table 1, respectively.
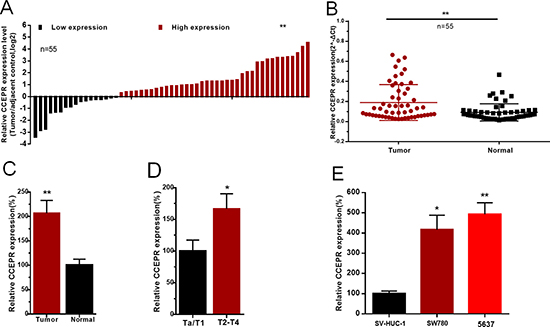
Figure 1: The relative expression levels of CCEPR in bladder cancer. (A) The heights of the columns in the chart represent the log2-transformed fold changes (bladder cancer tissue/normal bladder tissue) in CCEPR expression in 55 patients with bladder cancer. (B and C) CCEPR is up-regulated in bladder cancer tissues compared with corresponding non-tumor tissues. (D) CCEPR was up-regulated in patients with advanced TNM stage. (E) CCEPR was up-regulated in bladder cancer cell lines. Data are shown as mean ± SD. *p < 0.05; **p < 0.01.
Table 1: Correlation between CCEPR expression and clinicopathological features of UCB patients
Parameters Total |
Group |
Total |
CCEPR expression |
P value |
|
|---|---|---|---|---|---|
High |
Low |
||||
Gender |
Male |
40 (73%) |
27 (49%) |
13 (24%) |
0.677 |
Female |
15 (27%) |
11 (20%) |
4 (7%) |
||
Age (years) |
< 60 |
20 (36%) |
15 (27%) |
5 (9%) |
0.473 |
≥ 60 |
35 (64%) |
23 (42%) |
12 (22%) |
||
Tumor size (cm) |
< 3 cm |
21 (38%) |
15 (27%) |
6 (11%) |
0.768 |
≥ 3 cm |
34 (62%) |
23 (42%) |
11 (20%) |
||
Multiplicity |
Single |
32 (58%) |
25 (45%) |
7 (13%) |
0.087 |
Multiple |
23 (42%) |
13 (24%) |
10 (18%) |
||
Histological grade |
L |
23 (42%) |
12 (22%) |
11 (20%) |
0.021* |
H |
32 (58%) |
26 (47%) |
6 (11%) |
||
Tumor stage T |
Ta, T1 |
14 (26%) |
6 (11%) |
8 (15%) |
0.014* |
T2–T4 |
41 (74%) |
32 (58%) |
9 (16%) |
||
*P < 0.05 was considered significant (Chi-square test between 2 groups).
Corresponding specific siRNA/pcDNA3.1 down/up-regulated expression level of CCEPR
5637 and SW780 cells were cultured and then transfected with CCEPR specific siRNA or expression vector. The results of qRT-PCR showed that CCEPR mRNA levels in 5637 and SW780 cells was significantly decreased by siRNA-CCEPR (Figure 2A) and increased by pcDNA3.1-CCEPR (Figure 2B).
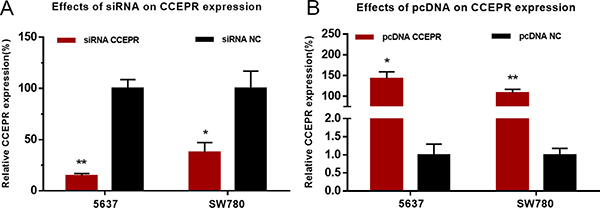
Figure 2: Effects of corresponding siRNA or pcDNA on CCEPR expression level. (A) The CCEPR specific siRNA significantly down-regulated the expression level of CCEPR in 5637 and SW780 cells. (B) The CCEPR specific pcDNA3.1 significantly up-regulated the expression level of CCEPR in 5637 and SW780 cells. Data are indicated as mean ± SD. *p < 0.05; **p < 0.01.
CCEPR promotes the proliferation of bladder cancer cells
The cell proliferative capacity were evaluated by Cell Counting Kit-8 assay and Ethynyl-2-deoxyuridine incorporation assay. Silencing CCEPR inhibited cell proliferation in 5637 and SW780 cells (Figure 3A, 3C, 3E). Overexpressing CCEPR promoted cell proliferation in 5637 and SW780 cells (Figure 3B, 3D, 3F).
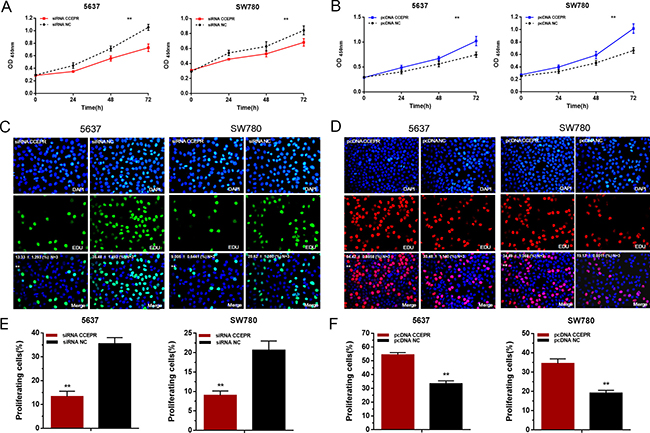
Figure 3: The effect of CCEPR on bladder cancer cell proliferation. (A, C and E) Silencing CCEPR inhibited cell proliferation in 5637 and SW780 cells. (B, D and F) Overexpressing CCEPR promoted cell proliferation in 5637 and SW780 cells. Data are shown as mean ± SD. *p < 0.05; **p < 0.01.
CCEPR promotes cell cycle and increases the expression of PCNA
The cell cycle was evaluated by Flow cytometry. Silencing CCEPR inhibited cell cycle in 5637 and SW780 cells (Figure 4A). Overexpressing CCEPR promoted cell cycle in 5637 and SW780 cells (Figure 4B). To investigate the underlying mechanisms of CCEPR-mediated biological processes, we performed bio-information analysis, qRT-PCR and western blotting. As shown in Figure 4C and 4D, down-regulation of CCEPR significantly decreased PCNA mRNA and protein levels and up-regulation of CCEPR significantly increased PCNA mRNA and protein levels in 5637 and SW780 cells.
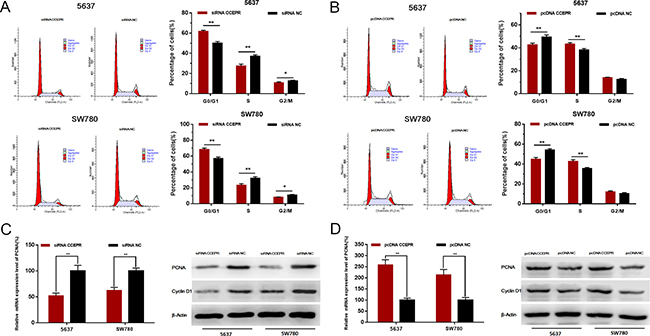
Figure 4: The effect of CCEPR on bladder cancer cell cycle and the expression of PCNA. (A) Silencing CCEPR inhibited cell cycle in 5637 and SW780 cells. (B) Overexpressing CCEPR promoted cell cycle in 5637 and SW780 cells. (C) Down-regulation of CCEPR decreased the mRNA and protein level of PCNA. (D) Up-regulation of CCEPR increased the mRNA and protein level of PCNA. Data are shown as mean ± SD. *p < 0.05; **p < 0.01.
CCEPR suppresses the apoptosis of bladder cancer cells
We further evaluated the apoptosis of bladder cancer cells by ELISA assay and Flow cytometry. Silencing CCEPR induced cell apoptosis in 5637 and SW780 cells (Figure 5A, 5C, 5E). Overexpressing CCEPR suppressed cell apoptosis in 5637 and SW780 cells (Figure 5B, 5D, 5F). As shown in Figure 5G, 5H, Down-/Up-regulation of CCEPR decreased /increasedthe anti-apoptosis associated markers (Bcl-2/Bcl-XL).
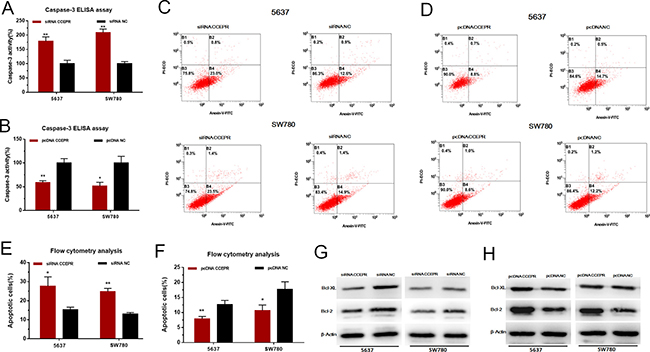
Figure 5: The effect of CCEPR on bladder cancer cell apoptosis. (A, C and E) Silencing CCEPR induced cell apoptosis in 5637 and SW780 cells. (B, D and F) Overexpressing CCEPR suppressed cell apoptosis in 5637 and SW780 cells. (G) Down-regulation of CCEPR decreased the anti-apoptosis associated markers (Bcl-2/Bcl-XL). (H) Up-regulation of CCEPR increased the anti-apoptosis associated markers (Bcl-2/Bcl-XL). Data are shown as mean ± SD. *p < 0.05; **p < 0.01.
CCEPR does not regulate cell migration and invasion in bladder cancer
We further evaluated whether CCEPR regulates the migration and invasion of bladder cancer cells. Regrettably, there was no significant difference in the migratory and invasive abilities of cells transfected with CCEPR specific siRNA (Figure 6A, 6C) or pcDNA3.1 (Figure 6B, 6D). The results indicated that CCEPR does not regulate cell migration and invasion in bladder cancer.
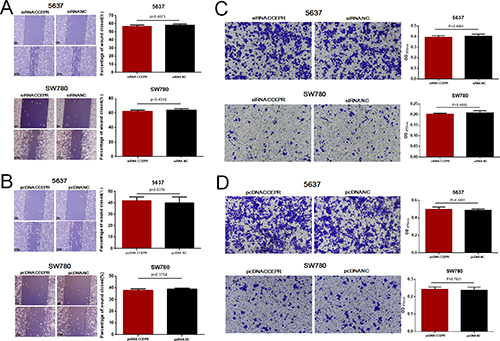
Figure 6: The effect of CCEPR on bladder cancer cell metastasis. (A and B) There was no significant difference in the migratory ability of bladder cancer cells transfected with corresponding specific siRNA/pcDNA3.1. (C and D) There was no significant difference in the invasive ability of bladder cancer cells transfected with corresponding specific siRNA/pcDNA3.1. Data are shown as mean ± SD. *p < 0.05; **p < 0.01.
DISCUSSION
Urothelial carcinoma of the bladder is the most common genitourinary malignancies and a major cause of morbidity and mortality in China [35, 36]. Because the molecular mechanisms of tumorigenesis in bladder cancer are still unknown, the therapeutic outcomes for patients with bladder cancer remain unsatisfactory [37, 38]. Therefore, to explore detailed molecular mechanisms of bladder cancer development and progression is essential in improving the clinical strategies and outcomes of bladder cancer [39, 40].
The lncRNAs which are longer than 200 nucleotides are a class of noncoding RNAs [41]. Recently, an increasing number of evidences indicated that lncRNAs play important roles in cancer development and progression [42–44]. CCEPR was previously reported to interact with PCNA and regulate cervical carcinoma occurrence and progression [31]. However, the biological functions and underlying mechanisms of CCEPR in bladder cancer are still unknown.
In this study, we found the expression level of CCEPR is significantly increased in bladder cancer and CCEPR could promote proliferation and suppresse apoptosis in bladder cancer cells. Mechanistically, the function of lncRNAs usually relies on the proteins that they interact with. We found CCEPR upregulates the expression of PCNA and serves as a key regulator in bladder cancer development and progression. In conclusion, these findings demonstrated that CCEPR plays an important regulatory role in bladder cancer and may provide a promising diagnostic and therapeutic target in bladder cancer.
MATERIALS AND METHODS
Patients and clinical samples collection
55 patients with urothelial carcinoma of bladder who received radical or partial cystectomy were included in this study. After radical or partial cystectomy bladder cancer tissues and normal bladder tissues from each patient were snap-frozen in liquid nitrogen immediately. All patients included in this study signed informed consent and this study was approved by the Institutional Review Board of Peking University First Hospital and Shenzhen University First Hospital.
Bladder cancer cell lines and cell culture
Bladder cancer cell lines used in this study were purchased from the Institute of Cell Research, Chinese Academy of Sciences, Shanghai, China. The SV-HUC-1 cells were cultured in DMEM Medium (Invitrogen, Carlsbad, CA, USA). The 5637 and SW780 cells were cultured in RPMI-1640 Medium (Invitrogen, Carlsbad, CA, USA).
siRNA and pcDNA transfection
The specific small interfering RNA and expression vector used in this study were purchased from GenePharma, Shanghai, China. The target sequence of siRNA-CCEPR was 5′- CGAGGGCGAGCATGTTTGTTGTTTA -3′ [31]. The cells were transiently transfected with corresponding siRNA or pcDNA using Lipofectamine 3000 Transfection Reagent (Invitrogen, Carlsbad, CA, USA).
RNA extraction and quantitative real-time PCR
The total RNA of the tissue samples and cells were extracted using the Trizol reagent (Invitrogen, Carlsbad, CA, USA). The detailed primer sequences included in this study are shown in Supplementary Table 2. Quantitative real-time PCR was performed using the ABI PRISM 7000 Fluorescent Quantitative PCR System according to the manufacturer’s instructions.
Cell counting Kit-8 assay
Cell proliferation was evaluated by Cell Counting Kit-8 (Beyotime Inst Biotech, China) following the Beyotime’s instructions. Briefly, 5 × 103 cells per well were seeded in a 96-well plate, then transfected with corresponding siRNA or pcDNA. Finally, the absorbance was finally evaluated at a wavelength of 450 nm using a microplate reader.
Ethynyl-2-deoxyuridine (EdU) incorporation assay
Cell proliferation was also evaluated by Ethynyl-2-deoxyuridine incorporation assay (Ribo Bio, Guangzhou, China) according to the Ribo Bio’s instructions. Briefly, after transfected with corresponding siRNA or pcDNA cells were incubated with 100 μl of 50 μM EdU per well for 2 h at 37°C, respectively. Finally, the cells were visualized under a fluorescence microscopy.
Cleaved caspase-3 ELISA assay
Cell apoptosis was evaluated by ELISA assay. Briefly, 5 × 105 cells per well were seeded in a 6-well plate, then transfected with corresponding siRNA or pcDNA, respectively. At 48 h after transfection, Cell cleaved caspase-3 activity was measured using the Caspase-3 Colorimetric Assay kit (Abcam, Cambridge, UK) following the Abcam’s instructions.
Flow cytometry analysis
Cell apoptosis and cell cycle were evaluated by Flow cytometry. Cells were collected after transfection for 48 h. Cell apoptosis was determined using PE Annexin V apoptosis detection kits (BD Pharmingen, San Diego, CA, USA) following the instructions. Cell cycle analysis was determined using propidium iodide cell cycle detected kits (BD Pharmingen) according to the instructions. Finally, cell apoptosis and cell cycle were determined using flow cytometry.
Western blotting analysis
Total protein was separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred onto nitrocellulose membranes. Then the membrane was blocked with 5% non-fat milk and incubated with primary antibodies at 4°C overnight. After incubation with specific antibodies (Abcam, Hong Kong, China), the blots were incubated with goat anti-rabbit secondary antibody (Abcam, Hong Kong, China) and visualized with enhanced chemiluminescence.
Wound healing assay
Cell motility was evaluated by wound healing assay. At 24 h after transfection, a wound field was created using a sterile 200 μl pipette tip. The cells were incubated for 24 h at 37°C, and then the migration of cells were visualized under a digital camera system.
Transwell assay
The invasion of bladder cancer cells was evaluated by a transwell insert (8 μm, Corning). At 24 h after transfection, 5 × 104 cells were starved in 200 ml serum free medium and placed in the dishes. The lower chamber was filled with 500 ml of complete medium. Then the cells that had migrated to the bottom surface of the filter membrane were stained with 0.5% crystal violet solution and photographed. Finally, the absorbance were evaluated using an microplate reader.
Statistical analyses
All experimental data from three independent experiments were analyzed by Student’s t-test or χ2 test with SPSS version 19.0 software (SPSS Inc. Chicago, IL, USA). Results were expressed as mean ± standard deviation and P-values of less than 0.05 were considered to be statistically significant.
ACKNOWLEDGMENTS AND FUNDING
This work was supported by the National Key Basic Research Program of China (973 Program) (2014CB745201), National Natural Science Foundation of China 81672546, 81602253, 81372746, Natural Science Foundation of Beijing 71772219, 7152146, the Shenzhen Municipal Government of China (ZDSYS201504301722174, JCYJ20150330102720130, GJHZ20150316154912494).
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest with this study.
REFERENCES
1. Resnick MJ. Variation in Bladder Cancer Spending: A Global Call to Action. European urology. 2016; 69:448–449.
2. Antoni S, Ferlay J, Soerjomataram I, Znaor A, Jemal A, Bray F. Bladder Cancer Incidence and Mortality: A Global Overview and Recent Trends. European urology. 2017; 71:96–108.
3. Knowles MA, Hurst CD. Molecular biology of bladder cancer: new insights into pathogenesis and clinical diversity. Nature reviews Cancer. 2015; 15:25–41.
4. Chamie K, Litwin MS, Bassett JC, Daskivich TJ, Lai J, Hanley JM, Konety BR, Saigal CS, Urologic Diseases in America P. Recurrence of high-risk bladder cancer: a population-based analysis. Cancer. 2013; 119:3219–3227.
5. Burger M, Catto JW, Dalbagni G, Grossman HB, Herr H, Karakiewicz P, Kassouf W, Kiemeney LA, La Vecchia C, Shariat S, Lotan Y. Epidemiology and risk factors of urothelial bladder cancer. European urology. 2013; 63:234–241.
6. Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature. 2014; 507:315–322.
7. Ingelmo-Torres M, Lozano JJ, Izquierdo L, Carrion A, Costa M, Gomez L, Ribal MJ, Alcaraz A, Mengual L. Urinary cell microRNA-based prognostic classifier for non-muscle invasive bladder cancer. Oncotarget. 2017; 8:18238–18247. doi: 10.18632/oncotarget.15315.
8. Powles T, Eder JP, Fine GD, Braiteh FS, Loriot Y, Cruz C, Bellmunt J, Burris HA, Petrylak DP, Teng SL, Shen X, Boyd Z, Hegde PS, et al. MPDL3280A (anti-PD-L1) treatment leads to clinical activity in metastatic bladder cancer. Nature. 2014; 515:558–562.
9. Izumi K, Taguri M, Miyamoto H, Hara Y, Kishida T, Chiba K, Murai T, Hirai K, Suzuki K, Fujinami K, Ueki T, Udagawa K, Kitami K, et al. Androgen deprivation therapy prevents bladder cancer recurrence. Oncotarget. 2014; 5:12665–12674. doi: 10.18632/oncotarget.2851.
10. Alfred Witjes J, Lebret T, Comperat EM, Cowan NC, De Santis M, Bruins HM, Hernandez V, Espinos EL, Dunn J, Rouanne M, Neuzillet Y, Veskimae E, van der Heijden AG, et al. Updated 2016 EAU Guidelines on Muscle-invasive and Metastatic Bladder Cancer. European urology. 2017; 71:462–475.
11. Pichler R, Fritz J, Zavadil C, Schafer G, Culig Z, Brunner A. Tumor-infiltrating immune cell subpopulations influence the oncologic outcome after intravesical Bacillus Calmette-Guerin therapy in bladder cancer. Oncotarget. 2016; 7:39916–39930. doi: 10.18632/oncotarget.9537.
12. Gui Y, Guo G, Huang Y, Hu X, Tang A, Gao S, Wu R, Chen C, Li X, Zhou L, He M, Li Z, Sun X, et al. Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nature genetics. 2011; 43:875–878.
13. Delto JC, Kobayashi T, Benson M, McKiernan J, Abate-Shen C. Preclinical analyses of intravesical chemotherapy for prevention of bladder cancer progression. Oncotarget. 2013; 4:269–276. doi: 10.18632/oncotarget.852.
14. Morales EE, Grill S, Svatek RS, Kaushik D, Thompson IM Jr, Ankerst DP, Liss MA. Finasteride Reduces Risk of Bladder Cancer in a Large Prospective Screening Study. European urology. 2016; 69:407–410.
15. Necsulea A, Soumillon M, Warnefors M, Liechti A, Daish T, Zeller U, Baker JC, Grutzner F, Kaessmann H. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature. 2014; 505:635–640.
16. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nature reviews Genetics. 2009; 10:155–159.
17. Di Gesualdo F, Capaccioli S, Lulli M. A pathophysiological view of the long non-coding RNA world. Oncotarget. 2014; 5:10976–10996. doi: 10.18632/oncotarget.2770.
18. Deng L, Yang SB, Xu FF, Zhang JH. Long noncoding RNA CCAT1 promotes hepatocellular carcinoma progression by functioning as let-7 sponge. Journal of experimental & clinical cancer research. 2015; 34:18.
19. Olivieri M, Ferro M, Terreri S, Durso M, Romanelli A, Avitabile C, De Cobelli O, Messere A, Bruzzese D, Vannini I, Marinelli L, Novellino E, Zhang W, et al. Long non-coding RNA containing ultraconserved genomic region 8 promotes bladder cancer tumorigenesis. Oncotarget. 2016; 7:20636–20654. doi: 10.18632/oncotarget.7833.
20. Ota T, Suzuki Y, Nishikawa T, Otsuki T, Sugiyama T, Irie R, Wakamatsu A, Hayashi K, Sato H, Nagai K, Kimura K, Makita H, Sekine M, et al. Complete sequencing and characterization of 21,243 full-length human cDNAs. Nature genetics. 2004; 36:40–45.
21. Yan TH, Lu SW, Huang YQ, Que GB, Chen JH, Chen YP, Zhang HB, Liang XL, Jiang JH. Upregulation of the long noncoding RNA HOTAIR predicts recurrence in stage Ta/T1 bladder cancer. Tumour biology. 2014; 35:10249–10257.
22. Zhan Y, Lin J, Liu Y, Chen M, Chen X, Zhuang C, Liu L, Xu W, Chen Z, He A, Zhang Q, Sun X, Zhao G, et al. Up-regulation of long non-coding RNA PANDAR is associated with poor prognosis and promotes tumorigenesis in bladder cancer. Journal of experimental & clinical cancer research. 2016; 35:83.
23. Hong HH, Hou LK, Pan X, Wu CY, Huang H, Li B, Nie W. Long non-coding RNA UCA1 is a predictive biomarker of cancer. Oncotarget. 2016; 7:44442–44447. doi: 10.18632/oncotarget.10142.
24. Luo M, Li Z, Wang W, Zeng Y, Liu Z, Qiu J. Long non-coding RNA H19 increases bladder cancer metastasis by associating with EZH2 and inhibiting E-cadherin expression. Cancer letters. 2013; 333:213–221.
25. Zhuang C, Li J, Liu Y, Chen M, Yuan J, Fu X, Zhan Y, Liu L, Lin J, Zhou Q, Xu W, Zhao G, Cai Z, et al. Tetracycline-inducible shRNA targeting long non-coding RNA PVT1 inhibits cell growth and induces apoptosis in bladder cancer cells. Oncotarget. 2015; 6:41194–41203. doi: 10.18632/oncotarget.5880.
26. Wu Y, Liu H, Shi X, Yao Y, Yang W, Song Y. The long non-coding RNA HNF1A-AS1 regulates proliferation and metastasis in lung adenocarcinoma. Oncotarget. 2015; 6:9160–9172. doi: 10.18632/oncotarget.3247.
27. Zhen S, Hua L, Liu YH, Sun XM, Jiang MM, Chen W, Zhao L, Li X. Inhibition of long non-coding RNA UCA1 by CRISPR/Cas9 attenuated malignant phenotypes of bladder cancer. Oncotarget. 2017; 8:9634–9646. doi: 10.18632/oncotarget.14176.
28. Liu D, Li Y, Luo G, Xiao X, Tao D, Wu X, Wang M, Huang C, Wang L, Zeng F, Jiang G. LncRNA SPRY4-IT1 sponges miR-101–3p to promote proliferation and metastasis of bladder cancer cells through up-regulating EZH2. Cancer letters. 2016; 388:281–291.
29. Zhan Y, Liu Y, Wang C, Lin J, Chen M, Chen X, Zhuang C, Liu L, Xu W, Zhou Q, Sun X, Zhang Q, Zhao G, et al. Increased expression of SUMO1P3 predicts poor prognosis and promotes tumor growth and metastasis in bladder cancer. Oncotarget. 2016; 7:16038–16048. doi: 10.18632/oncotarget.6946.
30. Wang T, Yuan J, Feng N, Li Y, Lin Z, Jiang Z, Gui Y. Hsa-miR-1 downregulates long non-coding RNA urothelial cancer associated 1 in bladder cancer. Tumour biology. 2014; 35:10075–10084.
31. Yang M, Zhai X, Xia B, Wang Y, Lou G. Long noncoding RNA CCHE1 promotes cervical cancer cell proliferation via upregulating PCNA. Tumour biology. 2015; 36:7615–7622.
32. Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta RA, Wysocka J, Lei M, Dekker J, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011; 472:120–124.
33. Puvvula PK, Desetty RD, Pineau P, Marchio A, Moon A, Dejean A, Bischof O. Long noncoding RNA PANDA and scaffold-attachment-factor SAFA control senescence entry and exit. Nature communications. 2014; 5:5323.
34. Yang X, Song JH, Cheng Y, Wu W, Bhagat T, Yu Y, Abraham JM, Ibrahim S, Ravich W, Roland BC, Khashab M, Singh VK, Shin EJ, et al. Long non-coding RNA HNF1A-AS1 regulates proliferation and migration in oesophageal adenocarcinoma cells. Gut. 2014; 63:881–890.
35. Nielsen ME, Smith AB, Meyer AM, Kuo TM, Tyree S, Kim WY, Milowsky MI, Pruthi RS, Millikan RC. Trends in stage-specific incidence rates for urothelial carcinoma of the bladder in the United States: 1988 to 2006. Cancer. 2014; 120:86–95.
36. Babjuk M. Trends in Bladder Cancer Incidence and Mortality: Success or Disappointment? European urology. 2017; 71:109–110.
37. Masson-Lecomte A, Rava M, Real FX, Hartmann A, Allory Y, Malats N. Inflammatory biomarkers and bladder cancer prognosis: a systematic review. European urology. 2014; 66:1078–1091.
38. Nguyen DP, Al Hussein Al Awamlh B, Wu X, O’Malley P, Inoyatov IM, Ayangbesan A, Faltas BM, Christos PJ, Scherr DS. Recurrence patterns after open and robot-assisted radical cystectomy for bladder cancer. European urology. 2015; 68:399–405.
39. Mostafid AH, Palou Redorta J, Sylvester R, Witjes JA. Therapeutic options in high-risk non-muscle-invasive bladder cancer during the current worldwide shortage of bacille Calmette-Guerin. European urology. 2015; 67:359–360.
40. Shiota M, Yokomizo A, Takeuchi A, Imada K, Kiyoshima K, Inokuchi J, Tatsugami K, Ohga S, Nakamura K, Honda H, Naito S. Secondary bladder cancer after anticancer therapy for prostate cancer: reduced comorbidity after androgen-deprivation therapy. Oncotarget. 2015; 6:14710–14719. doi: 10.18632/oncotarget.3817.
41. Rion N, Ruegg MA. LncRNA-encoded peptides: More than translational noise? Cell research. 2017; 27:604–605.
42. Engreitz JM, Pandya-Jones A, McDonel P, Shishkin A, Sirokman K, Surka C, Kadri S, Xing J, Goren A, Lander ES, Plath K, Guttman M. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science. 2013; 341:1237973.
43. Malek E, Jagannathan S, Driscoll JJ. Correlation of long non-coding RNA expression with metastasis, drug resistance and clinical outcome in cancer. Oncotarget. 2014; 5:8027–8038. doi: 10.18632/oncotarget.2469.
44. Martens-Uzunova ES, Bottcher R, Croce CM, Jenster G, Visakorpi T, Calin GA. Long noncoding RNA in prostate, bladder, and kidney cancer. European urology. 2014; 65:1140–1151.

