INTRODUCTION
With the soaring prevalence in obesity and diabetes, understanding the molecular mechanisms of adipogenesis has become an area of immense interest, and much of our current knowledge has emphasized on the transcriptional cascade that governs adipocyte differentiation [1]. Specifically, the peroxisome proliferator-activated receptor γ (PPARγ) is an essential adipogenic transcription factor, which cooperates with CCAAT/enhancer-binding proteins (C/EBPs), mainly C/EBPα, to induce the expression of genes important for differentiation [1].
The Src-associated in mitosis of 68kDa (Sam68) is an RNA binding protein belonging to the signal transduction and activation of RNA (STAR) family [2–4]. We have previously identified Sam68 as a critical regulator of adipogenesis, where whole-body Sam68 knockout (Sam68-/-) mice are lean due to increased energy expenditure, decreased commitment to early adipocyte progenitors, and defects in adipogenic differentiation [5]. Consistently, mouse embryonic fibroblasts (MEFs) isolated from Sam68-/- mice are compromised in their ability to differentiate into adipocytes [6]. More importantly, similar to the phenotype found in adipose tissue-specific PPARγ knockout mice [7], Sam68-/- mice are protected against dietary-induced obesity, as well as insulin resistance and glucose intolerance associated with diabetes [5]. Molecular evidence to date demonstrates that Sam68 can promote adipogenesis by regulating the alternative splicing of mTOR and S6K1 transcripts [5, 8]. Specifically, Sam68-depletion leads to decreased intron 5 retention of the mTOR pre-mRNA, which reduces mTOR protein levels, and causes defects in insulin-stimulated S6 and Akt phosphorylation [5]. In contrast, Sam68 prevents the production of adipogenic repressor p31S6K1, which is encoded by the short isoform of S6K1 [8]. However, it is unclear whether Sam68 has functions beyond that of a splicing regulator during adipogenesis. Moreover, although Sam68-deficient preadipocytes are defective in differentiation, little is known about the exact role of Sam68 in these cells.
A substantial proportion of the genome is transcribed into long noncoding RNAs (lncRNAs), which are RNA transcripts longer than 200 nucleotides, often polyadenylated, and lack evident open reading frames [9]. Despite their biochemical resemblance to messenger RNAs (mRNAs), lncRNAs are poorly conserved, lower in abundance, and have greater tissue specificity [9, 10]. Nevertheless, mounting evidence suggests that specific lncRNAs have regulatory roles in numerous cellular processes, including cell differentiation, development, and disease pathogenesis [9–12]. Previous large-scale studies have shown that lncRNAs can influence adipogenic differentiation, most of which are expressed in a tissue-specific manner, and are strongly induced during adipogenesis [13, 14]. Furthermore, results obtained from gain and loss-of-function screens suggest that some lncRNAs can serve as key regulators of adipocyte-specific genes [13, 15]. Indeed, a few lncRNAs have been characterized with important roles in adipocyte development, including ADINR [15], HOTAIR [16], NEAT1 [17, 18], as well as lnc-BATE-1 specifically for brown adipocyte differentiation [14]; however, the function and regulation for many existing lncRNAs remain poorly understood.
In the present study, we performed deep RNA sequencing (RNA-seq) to profile the transcriptome influenced by the loss of Sam68 in mouse preadipocytes. We found a small subset of lncRNAs that are differentially regulated by Sam68, which contained consensus Sam68 binding sites, and are modulated by Sam68 via transcript stability. Furthermore, we confirmed by RNA interference (RNAi) that these lncRNAs play an important role in preadipocyte differentiation. Hence, we have characterized a new role for Sam68 as a regulator of lncRNAs during adipogenesis.
RESULTS
Transcriptional profiling of Sam68 wild type and deficient preadipocytes
To identify the transcripts whose expression is regulated by Sam68, we first performed RNA-seq using undifferentiated Sam68 wild type (pRetroSuper) and Sam68-deficient (Sam68sh) 3T3-L1 cells. These Sam68sh cells were previously characterized for their differentiation defects [5]. We identified 652 protein-coding genes and 9 noncoding RNAs (ncRNAs) that were significantly up or downregulated (≥ 2 fold change, adjusted p < 0.05) by Sam68 (Figures 1A and 1B, Supplemental Tables 1 and 2). Indeed, the expression of pro-adipogenic factors such as C/EBP δ (Cebpd) (adjusted p = 1.52 x 10-77) was among the most significantly downregulated genes in Sam68sh 3T3-L1 cells, while the gene expression of repressors, for instance GATA-3 (Gata3) (adjusted p = 1.45 x 10-5), was significantly upregulated with Sam68 depletion. And interestingly, the lncRNA Hotair (adjusted p = 6.14 x 10-3) was one of the most downregulated ncRNAs in Sam68-deficient cells. As expected, global gene ontology (GO) analysis revealed that downregulated genes in Sam68-deficient cells were significantly associated with “regulation of cell migration”, “regulation of cell motility”, “fat cell differentiation”, “response to lipid”, “extracellular matrix organization”, and other known Sam68-regulated biological processes (q < 0.05) (Figure 1C). Conversely, upregulated genes in Sam68-deficient cells were significantly associated with GO terms linked to “regulation of neurogenesis”, “negative regulation of fat cell differentiation”, and “negative regulation of nucleic acid-templated transcription” (q < 0.05, Figure 1C), all of which are consistent with previous reports of Sam68 functions [19–21]. Since Sam68 is known to regulate alternative splicing during adipogenesis [5, 8], we also analyzed the data for genome-wide alternation in transcript usage. Indeed, we detected a total of 725 differential alternative splicing events across 631 genes between wild type and Sam68-deficient 3T3-L1 preadipocytes (Figures 1D and 1E, Supplemental Table 3). We did not detect the previously identified mTOR and Rps6kb1 alternative splicing events, likely because of the limited sequence coverage. Further analysis will be required to confirm the new alternative spliced events identified.
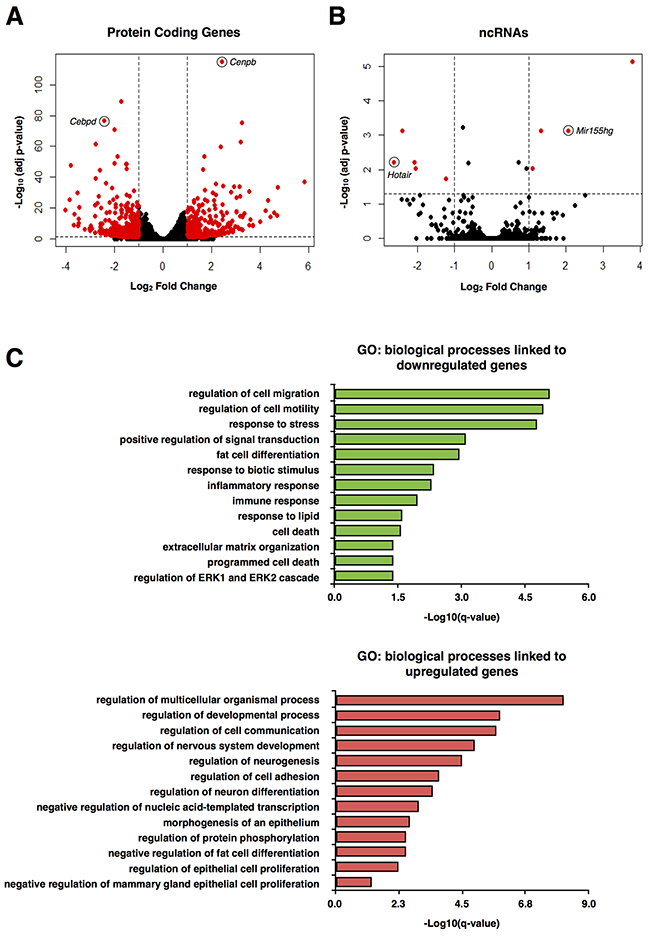
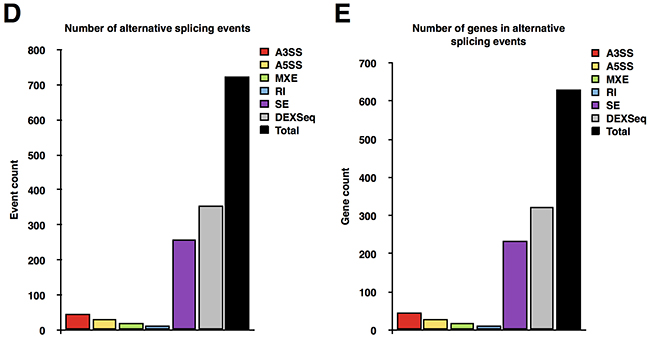
Figure 1: RNA-seq analysis of protein-coding genes and ncRNAs in mouse preadipocytes. (A and B) Volcano plot of protein-coding genes (A) and noncoding RNAs (ncRNAs) (B) differentially expressed between pRetroSuper and Sam68sh 3T3-L1 cells (red). (C) Most significant gene ontology (GO) terms associated with downregulated (green) and upregulated (red) genes in Sam68sh 3T3-L1 cells. (D and E) Differential alternative splicing events detected by rMATS and DEXSeq between pRetroSuper and Sam68sh 3T3-L1 preadipocytes. Bar plots represent the number of each type of alternative splice event (D) and the number of different genes found in each event (E). A3SS, alternative 3’ splice site; A5SS, alternative 5’ splice site; MXE, mutually exclusive exons; RI, retained intron; SE, skipped exon.
Genome-wide identification of Sam68-regulated lncRNAs in preadipocytes
Detailed analysis of the 9 ncRNAs identified from our RNA-seq data revealed lncRNAs, microRNAs (miRNAs), small cytoplasmic RNAs (scRNAs), and uncharacterized RIKEN sequences that were differentially regulated by Sam68. To further investigate the role of Sam68 in lncRNA regulation, we performed RT-qPCR in pRetroSuper and Sam68sh 3T3-L1 cells. We confirmed the depletion of Sam68 mRNA (~80%), and we validated 4 lncRNAs that were differentially regulated by Sam68 (Figure 2). These lncRNAs include 2 known transcripts, Hotair and Mir155hg, as well as 2 previously uncharacterized lncRNAs herein referred to as Sam68-regulated lncRNAs (SR-lncRNAs), SR-lncRNA-1 (2610035D17Rik) and SR-lncRNA-2 (4930461G14Rik). Our results showed that Sam68-deficiency significantly decreased Hotair (~50-fold) and SR-lncRNA-1 (~1.7-fold) levels, while the expression of Mir155hg and SR-lncRNA-2 were substantially increased by ~1.7-fold and ~13-fold, respectively, compared to Sam68 wild type cells (Figure 2). Of note, ectopic expression of Hotair has been shown to promote preadipocyte differentiation [16], whereas the mature product of Mir155hg, miR-155, is a known inhibitor of adipogenesis [22]. Together, we identify lncRNA targets for Sam68 in preadipocytes that are both positive and negative regulators of adipogenesis.
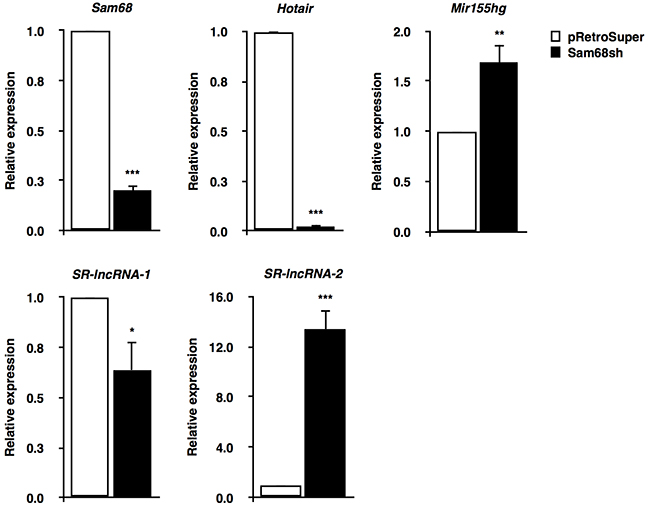
Figure 2: Validation of Sam68 lncRNA targets. RT-qPCR confirmation of Sam68 mRNA expression and Sam68-regulated lncRNAs (Hotair, Mir155hg, SR-lncRNA-1, SR-lncRNA-2) in pRetroSuper (white) and Sam68sh (black) 3T3-L1 cells. Expressions were normalized to 18S rRNA levels. Data are represented as mean ± S.D. from 3 independent experiments done in biological triplicates (*p ≤0.05, **p ≤0.005, ***p ≤0.0005).
Sam68 controls lncRNA expression by regulating transcript stability
To understand how Sam68 regulates the expression of lncRNAs in preadipocytes, we first searched for potential Sam68 binding sites in the available sequences for Hotair, Mir155hg, SR-lncRNA-1, and SR-lncRNA-2, given that Sam68 binds to UU/AAA bipartite elements [23, 24]. While we found potential Sam68 binding sites on all 4 lncRNAs (Table 1), only Hotair, Mir155hg, and SR-lncRNA-2 contained perfect consensus Sam68 binding sites.
Table 1: Consensus Sam68 binding sites on target lncRNAs.
Target |
RefSeq |
Consensus Sam68 binding sites |
Nucleotides (bp) |
|---|---|---|---|
Hotair |
NR_047528.1 |
…UUAAGCTGUAAA…UUUAUUUUUAAA… |
26-37, 1916-1927 |
Mir155hg |
NR_132106.1 |
…UAAAUAUUUAAUUUAAUUAAUAUUUAA… |
1032-1058 |
SR-lncRNA-1 |
NR_015556.1 |
…UUUAUAUAAA…UAAAUUCCUUUA… |
1605-1614, 1834-1845 |
SR-lncRNA-2 |
NR_040736.1 |
…UUAAAAUUAA… |
731-740 |
We next performed actinomycin D experiments to monitor the half-life of Hotair, Mir155hg, SR-lncRNA-1, and SR-lncRNA-2 in pRetroSuper and Sam68sh 3T3-L1 cells. Our results demonstrate that Sam68 status affects the transcript half-life of 3 out of the 4 lncRNAs (Figure 3). Specifically, while depletion of Sam68 did not significantly influence the half-life of SR-lncRNA-1 (1.98 h vs. 1.91 h), which does not contain a perfect Sam68 binding site, the half-life of Hotair was dramatically reduced from 3.43 h to 2.28 h (Figure 3). In contrast, Sam68-deficiency significantly increased the transcript half-life of Mir155hg (1.85 h vs. 2.13 h) and SR-lncRNA-2 (2.35 h vs. 4.19 h) (Figure 3). Altogether, these data suggest that Sam68 mainly modulates target lncRNA expression by differentially regulating their transcript stability. However, given the evidence of Sam68 as a transcriptional regulator [25–27], we do not exclude the possibility that Sam68 may also contribute to lncRNA expression in preadipocytes at the transcription level.
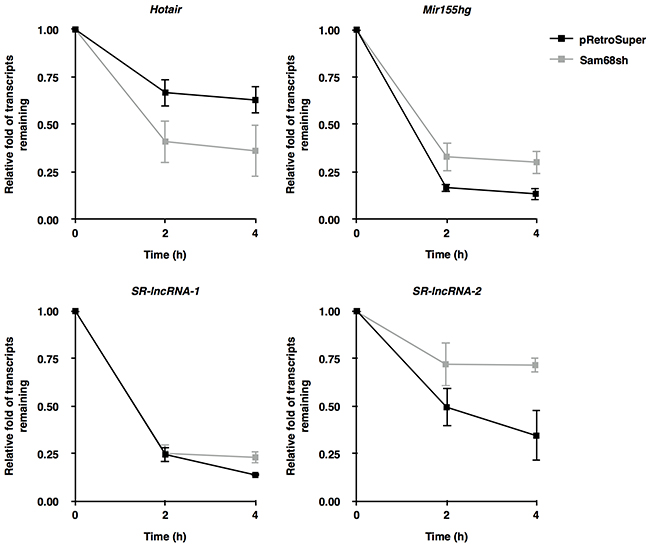
Figure 3: Sam68 regulates lncRNA expression by modulating transcript stability. Transcript stability was assessed in pRetroSuper (black) or Sam68sh (grey) 3T3-L1 cells treated with actinomycin D (5μg /mL). Total RNA was extracted at the indicated time points and transcripts were quantified by RT-qPCR using primers specific for Hotair, Mir155hg, SR-lncRNA-1, and SR-lncRNA-2, and were normalized to 18S rRNA levels. Data are represented as mean ± S.D. from 3 independent experiments done in biological triplicates.
Sam68-regulated lncRNAs in preadipocytes are important for adipogenic differentiation
To assess the functional role of Sam68-regulated lncRNAs in adipogenesis, we next performed loss-of-function studies using small interfering RNAs (siRNAs). Our earlier data showed that Hotair and SR-lncRNA-1 had a higher expression in wild type 3T3-L1 cells compared to Sam68sh cells, whereas it was the contrary for SR-lncRNA-2. Thus, we decided to deplete Hotair and SR-lncRNA-1 transcripts from wild type 3T3-L1 cells, and SR-lncRNA-2 from Sam68sh 3T3-L1 cells (Figure 4A).
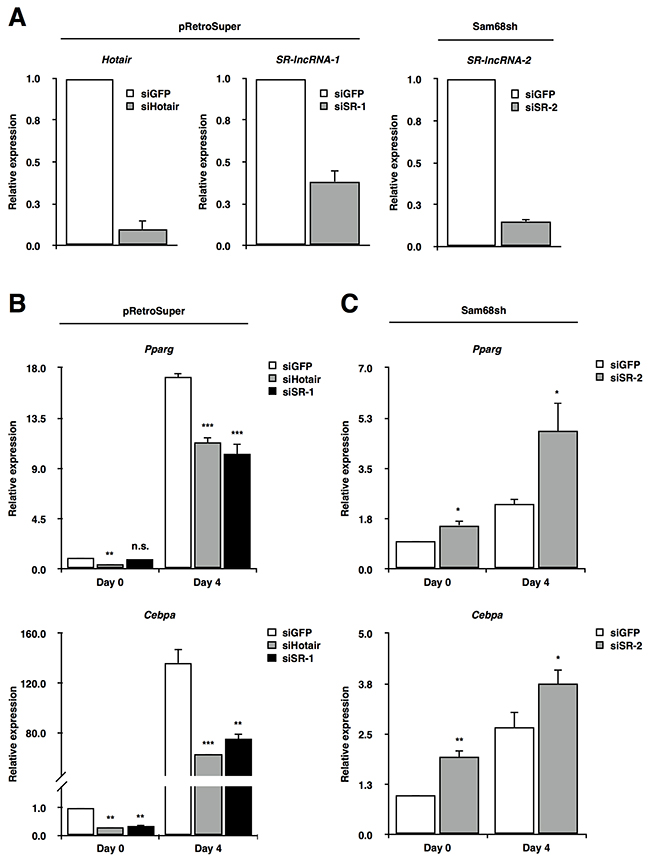
Figure 4: Sam68-regulated lncRNAs are required for adipogenic differentiation. (A) RT-qPCR confirmation of lncRNA knockdown in pRetroSuper (Hotair, SR-lncRNA-1) and Sam68sh (SR-lncRNA-2) 3T3-L1 cells. Expressions were normalized to 18S rRNA levels. (B and C) RT-qPCR analysis of Pparg and Cebpa on day 0 and day 4 of adipogenic differentiation after knockdown of the indicated lncRNAs in pRetroSuper (B) and Sam68sh 3T3-L1 cells (C). Expressions were normalized to 18S rRNA levels. Data, replicated in 3 independent experiments, are represented as mean ± S.D. of biological triplicates (*p ≤0.05, **p ≤0.005, ***p ≤0.0005).
Consistent with previous studies demonstrating Hotair as a positive regulator of adipogenesis [16], knockdown of Hotair markedly reduced the expression of adipogenic marker genes Pparg and Cebpa on both day 0 and day 4 of differentiation (Figure 4B). Similarly, although knockdown of SR-lncRNA-1 did not significantly alter Pparg mRNA levels on day 0, Pparg induction was compromised on day 4 compared to control cells, while Cebpa expression was dramatically decreased on both day 0 and day 4 with SR-lncRNA-1 knockdown compared to control cells (Figure 4B). Unlike Hotair and SR-lncRNA-1, Sam68 depletion increased SR-lncRNA-2 expression (Figure 2), suggesting that SR-lncRNA-2 may be an early inhibitor of adipogenic differentiation. Indeed, although not comparable to Sam68 wild type cells, depletion of SR-lncRNA-2 in Sam68sh cells significantly increased Pparg and Cebpa mRNA levels on day 0 and day 4 of differentiation compared to Sam68sh control cells (Figure 4C).
To confirm our gene expression data, we performed oil red O (ORO) staining for lncRNA-depleted 3T3-L1 cells on day 0 and day 4 of differentiation to monitor lipid accumulation. Consistently, the number of ORO+ cells was dramatically reduced when Hotair or SR-lncRNA-1 was depleted, as a result of defective adipogenic differentiation (Figure 5A). On the other hand, knockdown of SR-lncRNA-2 visibly increased the number of ORO+ Sam68sh cells on day 4 of differentiation, suggesting a slight rescue in adipogenic differentiation (Figure 5B). Collectively, these results demonstrate that Sam68-regulated lncRNAs have critical functions in adipogenesis.
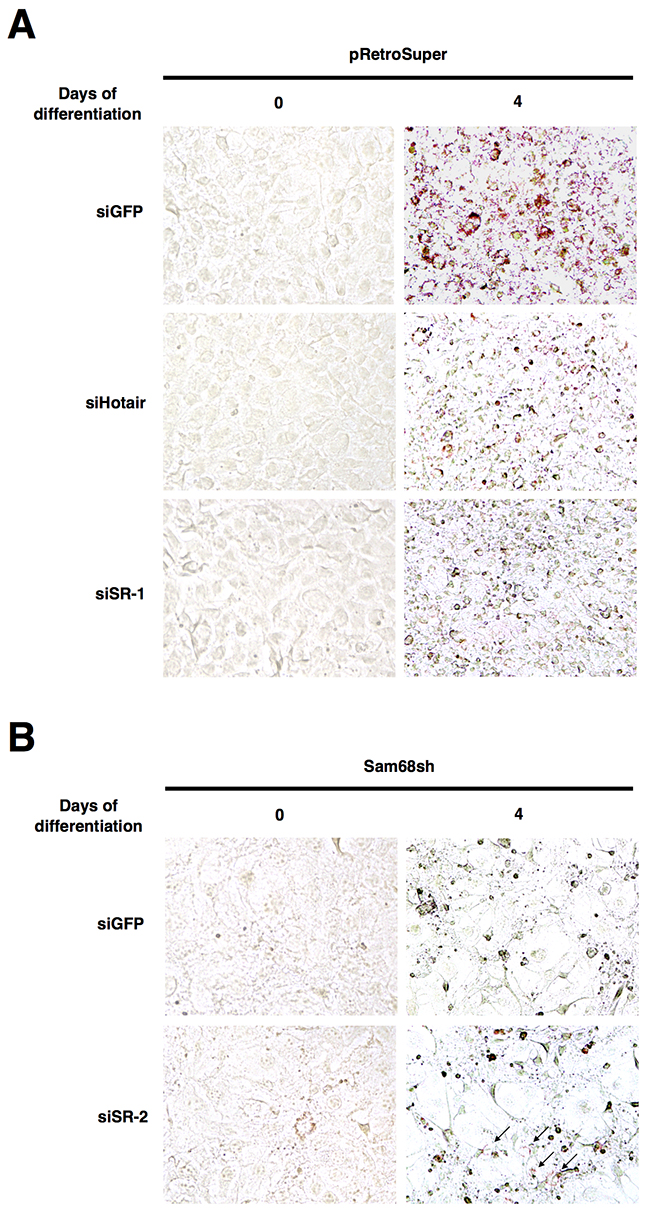
Figure 5: Sam68-regulated lncRNAs are required for lipid accumulation. (A and B) Oil red O staining (40X) on day 0 and day 4 of adipogenic differentiation after knockdown of the indicated lncRNAs in pRetroSuper (A) and Sam68sh (B) 3T3-L1 cells.
DISCUSSION
Although the study of adipogenesis has traditionally focused on nuclear transcription factors, lncRNAs are recently emerging as key players in adipogenic differentiation [11, 13, 14]. In the present manuscript, we unveiled a new role for Sam68 as a regulator of lncRNAs in mouse preadipocytes to promote adipogenesis. We identified Sam68-regulated lncRNAs that have important functions in adipocyte differentiation including Hotair, Mir155hg, SR-lncRNA-1, and SR-lncRNA-2, where the latter two are new lncRNAs that have never been previously associated with adipogenesis. These lncRNAs contain consensus Sam68 binding sites, specifically with the stability of Hotair, Mir155hg, and SR-lncRNA-2 modulated by Sam68. These findings define a new role for Sam68 during adipogenesis, namely in its regulation on transcript stability.
LncRNAs are known to regulate adipogenesis through lncRNA-protein interactions [11]; however few studies have linked lncRNAs with RNA binding proteins. Given that Sam68 is required for adipogenesis [5, 8], we now demonstrate its importance in cell differentiation as a regulator of specific lncRNAs using mouse preadipocytes. Interestingly, Sam68 has also been described to modulate lncRNA functions in human cancer cells. Specifically, INXS requires the interaction of Sam68-containing ribonucleoprotein complexes for efficient BCL-XS splicing, leading to increased apoptosis and tumor regression [28]. Furthermore, Sam68 associates with the p53 lncRNA target, PR-lncRNA-1, to synergistically enhance p53 transcriptional activities and effector functions [25]. Taken together, these studies suggest that the role of Sam68 as lncRNA regulator is conserved across different species, cell types, and cellular networks.
Much of the previous studies have focused on identifying differentially expressed lncRNAs during adipocyte differentiation [13, 14]. We now add to these studies by defining new functional lncRNAs in preadipocytes. Among our validated Sam68 lncRNA targets, Hotair is one of the few lncRNAs with well-characterized molecular functions. In humans, increased HOTAIR expression is associated with promotion of cancer metastasis, where it has been shown to reprogram the cellular chromatin state by re-targeting Polycomb Repressive Complex 2 (PRC2) [29]. Additionally to cancer, HOTAIR expression is also detected in gluteal adipose tissues, with enrichment in adipocyte fractions, suggesting its potential role in adipogenic differentiation (Supplemental Figure 1) [16, 30]. Indeed, ectopic expression of HOTAIR promotes adipogenesis [16]. We now extend these findings, and show that Sam68 increases Hotair stability to promote adipogenesis.
Moreover, we also identified Mir155hg, which is the primary sequence of miR-155, a known inhibitor of adipogenesis [22]. Although the relation between Sam68 and miRNAs was not investigated in our current study, many publications have shown that lncRNAs can block miRNA maturation by complementing with their primary sequences, or reduce miRNA levels by acting as sponges [10, 11]. In addition to the two annotated targets, we discovered two previously uncharacterized Sam68-regulated lncRNAs, SR-lncRNA-1 and SR-lncRNA-2, with positive and negative roles in adipocyte differentiation, respectively. Given the versatility and ubiquitous nature of Sam68 [3, 4], it would be of great interest to further explore their possible functions in other cellular processes and diseases.
One aspect of Sam68 that we have yet to investigate is its possible transcriptional role in preadipocytes. Indeed, previous studies have identified Sam68 as a co-regulator for a number of transcription factors, including the androgen receptor (AR) [27], NF-κB [26], and p53 [25]. These evidences suggest that Sam68 may contribute to the preadipocyte transcriptome by modulating both the transcription and transcript stability of its target genes. Moreover, it is interesting to note that Sam68 can differentially regulate transcript stability. Specifically, while Sam68-deficiency decreased the half-life of Hotair, it increased the half-lives of Mir155hg and SR-lncRNA-2. Given that spliced lncRNAs are more stable than single exon transcripts [31], it can be speculated that Sam68 may influence stability by regulating the splicing of its target lncRNAs.
To date, several mechanisms of action have been elucidated for lncRNAs in gene regulation. Most lncRNAs are nuclear, and function by modifying chromatin structures [32, 33], where they act as molecular scaffolds to recruit DNA methyltransferases [34], or histone modification complexes that ultimately influence transcription [35]. On the other hand, cytoplasmic lncRNAs can modulate translation, regulate mRNA degradation, or act as sponges for miRNAs [36].
In summary, our study demonstrates a previously unappreciated role for Sam68 as a regulator of lncRNAs during adipogenesis. This feature of Sam68 can be potentially harnessed as a therapeutic target for treating obesity and its related metabolic disorders, possibly by inhibiting the activity of Sam68 [37]. However, much remains to be determined on deciphering the detailed mechanisms by which Sam68 acts through lncRNAs for regulating gene expression.
MATERIALS AND METHODS
RNA-seq, GO, and alternative splicing event analysis
Total RNA was isolated from undifferentiated Sam68 wild type (pRetroSuper) and Sam68-deficient (Sam68sh) 3T3-L1 preadipocytes using TRIzol reagent according to the manufacturer’s instructions. RNA integrity and quality were checked as previously described [5], and biological triplicates of each sample were sent for sequencing. For analysis, raw reads were trimmed using Trimmomatic v0.32 [38], removing low-quality bases at the end of the reads (phred33 < 30), clipping the first 4 bases and clipping the Illumina adaptor sequences using palindrome mode. A sliding window quality trimming was performed, cutting once the average quality of a window of 4 bases that fell below 30. Reads shorter than 30 base pairs after trimming were discarded. Trimmed reads were aligned to the reference genome mm10 using STAR v2.3.0e [39]. Read counts for each gene at the exonic level were calculated by featureCounts v1.4.4 using Gencode VM7 annotation, normalized using DESeq2 R library [40], and were annotated using org.Mm.eg.db R bioconductor library. Differential expression analysis was performed using DESeq2 for all Gencode VM7 genes at the exonic level [40]. Genes were considered differentially expressed if they had a |fold change| ≥ 2 and an adjusted p-value < 0.05. Gene ontology (GO) analysis was performed with GOrilla tools [41]. Differential alternative splicing events were detected using rMATS (FDR < 0.05) and DEXSeq (adjusted p < 0.05) [42, 43].
Preadipocyte differentiation and ORO staining
Sam68 wild type (pRetroSuper) and deficient (Sam68sh) 3T3-L1 cells were previously described [5]. Adipogenic differentiation of 3T3-L1 cells was performed as previously described [5]. For oil red O (ORO) staining, cells were fixed in 3% formaldehyde and 0.025% glutaraldehyde. After fixation, cells were washed with PBS and were stained with freshly prepared ORO solution (Sigma-Aldrich).
RNA extraction and RT-qPCR
Total RNA was extracted at the indicated time points as described above. Reverse transcription (RT) was performed with the M-MLV reverse transcriptase (Promega). Real-time PCR (qPCR), including primer design and efficiency tests, was performed according to the Minimum Information for Publication of Quantitative Real-Time PCR Experiments (MIQE) guidelines [44]. qPCR experiments were performed using PowerUp SYBR Green Master Mix (Thermo Fisher Scientific) following the manufacturer’s instructions. Data were normalized by the ΔΔCt method. Sequences of primers used for qPCR are listed in Table 2.
Table 2: List of primers used for qPCR.
Target (mouse) |
Forward Primer (5’-3’) |
Reverse Primer (5’-3’) |
|---|---|---|
Sam68 |
GTGGAGACCCCAAATATGCCCA |
AAACTGCTCCTGACAGATATCA |
Pparg |
GAACGTGAAGCCCATCGAGGAC |
CTGGAGCACCTTGGCGAACA |
Cebpa |
CGCAAGAGCCGAGATAAAGC |
GCGGTCATTGTCACTGGTCA |
Hotair |
AAGGCTGAAATGGAGGACCG |
TACCGATGTTGGGGACCTCT |
Mir155hg |
GGGGTTTTGGCCTCTGACTG |
TTGGACTTGTCATCCTCCCAC |
SR-lncRNA-1 |
TCCTTGCCCGACTACAAACC |
TACAGCTCTGCGCCTTTCTT |
SR-lncRNA-2 |
GCATCACTGTGGTCTCTACCC |
TAACCCTTCTGCCAAGCTCTC |
18S |
GTAACCCGTTGAACCCCATT |
CCATCCAATCGGTAGTAGCG |
Hprt |
TTGGGCTTACCTCACTGCTT |
TCGCTAATCACGACGCTGG |
Arp |
GAGGAATCAGATGAGGATATGGGA |
AAGCAGGCTGACTTGGTTGC |
LncRNA knockdown
The following siGENOME SMARTpool small interfering RNAs (siRNAs) were purchased from Dharmacon: Hotair, SR-lncRNA-1 (2610035D17Rik), and SR-lncRNA-2 (4930461G14Rik). Transfection of siRNAs was performed with RNAiMAX reagents (Invitrogen) according to the manufacturer’s instructions. Cells were recovered and grown to confluence before induction of differentiation. Total RNA was harvested as described above.
ACKNOWLEDGMENTS
We thank Zhenbao Yu for helpful discussions and for critically reading the manuscript.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
GRANT SUPPORT
This work was supported by a grant from the Canadian Institute of Health Research (MOP-123531) to SR and a grant from the Fonds de Recherche du Québec – Santé (FRQS) to CLK. The computing infrastructure was provided by Calcul Québec and Compute Canada.
REFERENCES
1. Rosen ED, MacDougald OA. Adipocyte differentiation from the inside out. Nat Rev Mol Cell Biol. 2006; 7:885–96.
2. Lukong KE, Richard S. Sam68, the KH domain-containing superSTAR. Biochim Biophys Acta. 2003; 1653:73–86.
3. Richard S. Reaching for the stars: linking RNA binding proteins to diseases. Adv Exp Med Biol. 2010; 693:142–57.
4. Bielli P, Busà R, Paronetto MP, Sette C. The RNA-binding protein Sam68 is a multifunctional player in human cancer. Endocr Relat Cancer. 2011; 18:R91–102.
5. Huot ME, Vogel G, Zabarauskas A, Ngo CT, Coulombe-Huntington J, Majewski J, Richard S. The Sam68 STAR RNA-binding protein regulates mTOR alternative splicing during adipogenesis. Mol Cell. 2012; 46:187–99.
6. Richard S, Torabi N, Franco GV, Tremblay GA, Chen T, Vogel G, Morel M, Cléroux P, Forget-Richard A, Komarova S, Tremblay ML, Li W, Li A, et al. Ablation of the Sam68 RNA binding protein protects mice from age-related bone loss. PLoS Genet. 2005; 1:e74.
7. Jones JR, Barrick C, Kim KA, Lindner J, Blondeau B, Fujimoto Y, Shiota M, Kesterson RA, Kahn BB, Magnuson MA. Deletion of PPARgamma in adipose tissues of mice protects against high fat diet-induced obesity and insulin resistance. Proc Natl Acad Sci USA. 2005; 102:6207–12.
8. Song J, Richard S. Sam68 Regulates S6K1 Alternative Splicing during Adipogenesis. Mol Cell Biol. 2015; 35:1926–39.
9. Rinn JL, Chang HY. Genome regulation by long noncoding RNAs. Annu Rev Biochem. 2012; 81:145–66.
10. Knoll M, Lodish HF, Sun L. Long non-coding RNAs as regulators of the endocrine system. Nat Rev Endocrinol. 2015; 11:151–60.
11. Chen Z. Progress and prospects of long noncoding RNAs in lipid homeostasis. Mol Metab. 2015; 5:164–70.
12. Audas TE, Lee S. Stressing out over long noncoding RNA. Biochimica et biophysica acta. 2016; 1859:184–191. doi: 10.1016/j.bbagrm.2015.06.010.
13. Sun L, Goff LA, Trapnell C, Alexander R, Lo KA, Hacisuleyman E, Sauvageau M, Tazon-Vega B, Kelley DR, Hendrickson DG, Yuan B, Kellis M, Lodish HF, Rinn JL. Long noncoding RNAs regulate adipogenesis. Proc Natl Acad Sci USA. 2013; 110:3387–92.
14. Alvarez-Dominguez JR, Bai Z, Xu D, Yuan B, Lo KA, Yoon MJ, Lim YC, Knoll M, Slavov N, Chen S, Chen P, Lodish HF, Sun L. De Novo Reconstruction of Adipose Tissue Transcriptomes Reveals Long Non-coding RNA Regulators of Brown Adipocyte Development. Cell Metab. 2015; 21:764–76.
15. Xiao T, Liu L, Li H, Sun Y, Luo H, Li T, Wang S, Dalton S, Zhao RC, Chen R. Long Noncoding RNA ADINR Regulates Adipogenesis by Transcriptionally Activating C/EBPα. Stem Cell Reports. 2015; 5:856–65.
16. Divoux A, Karastergiou K, Xie H, Guo W, Perera RJ, Fried SK, Smith SR. Identification of a novel lncRNA in gluteal adipose tissue and evidence for its positive effect on preadipocyte differentiation. Obesity (Silver Spring). 2014; 22:1781–85.
17. Cooper DR, Carter G, Li P, Patel R, Watson JE, Patel NA. Long Non-Coding RNA NEAT1 Associates with SRp40 to Temporally Regulate PPARγ2 Splicing during Adipogenesis in 3T3-L1 Cells. Genes (Basel). 2014; 5:1050–63.
18. Gernapudi R, Wolfson B, Zhang Y, Yao Y, Yang P, Asahara H, Zhou Q. MicroRNA 140 Promotes Expression of Long Noncoding RNA NEAT1 in Adipogenesis. Mol Cell Biol. 2015; 36:30–38.
19. Huot ME, Brown CM, Lamarche-Vane N, Richard S. An adaptor role for cytoplasmic Sam68 in modulating Src activity during cell polarization. Mol Cell Biol. 2009; 29:1933–43.
20. Iijima T, Wu K, Witte H, Hanno-Iijima Y, Glatter T, Richard S, Scheiffele P. SAM68 regulates neuronal activity-dependent alternative splicing of neurexin-1. Cell. 2011; 147:1601–14.
21. La Rosa P, Bielli P, Compagnucci C, Cesari E, Volpe E, Farioli Vecchioli S, Sette C. Sam68 promotes self-renewal and glycolytic metabolism in mouse neural progenitor cells by modulating Aldh1a3 pre-mRNA 3’-end processing. eLife. 2016; 5:e20750.
22. Liu S, Yang Y, Wu J. TNFα-induced up-regulation of miR-155 inhibits adipogenesis by down-regulating early adipogenic transcription factors. Biochem Biophys Res Commun. 2011; 414:618–24.
23. Galarneau A, Richard S. The STAR RNA binding proteins GLD-1, QKI, SAM68 and SLM-2 bind bipartite RNA motifs. BMC Mol Biol. 2009; 10:47.
24. Lin Q, Taylor SJ, Shalloway D. Specificity and determinants of Sam68 RNA binding. Implications for the biological function of K homology domains. J Biol Chem. 1997; 272:27274–80.
25. Li N, Richard S. Sam68 functions as a transcriptional coactivator of the p53 tumor suppressor. Nucleic Acids Res. 2016; 44:8726–41.
26. Fu K, Sun X, Zheng W, Wier EM, Hodgson A, Tran DQ, Richard S, Wan F. Sam68 modulates the promoter specificity of NF-κB and mediates expression of CD25 in activated T cells. Nat Commun. 2013; 4:1909.
27. Rajan P, Gaughan L, Dalgliesh C, El-Sherif A, Robson CN, Leung HY, Elliott DJ. The RNA-binding and adaptor protein Sam68 modulates signal-dependent splicing and transcriptional activity of the androgen receptor. J Pathol. 2008; 215:67–77.
28. DeOcesano-Pereira C, Amaral MS, Parreira KS, Ayupe AC, Jacysyn JF, Amarante-Mendes GP, Reis EM, Verjovski-Almeida S. Long non-coding RNA INXS is a critical mediator of BCL-XS induced apoptosis. Nucleic Acids Res. 2014; 42:8343–55.
29. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010; 464:1071–76.
30. Consortium G, and GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013; 45:580–85.
31. Clark MB, Johnston RL, Inostroza-Ponta M, Fox AH, Fortini E, Moscato P, Dinger ME, Mattick JS. Genome-wide analysis of long noncoding RNA stability. Genome Res. 2012; 22:885–98.
32. Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta RA, Wysocka J, Lei M, Dekker J, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011; 472:120–24.
33. Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007; 129:1311–23.
34. Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R, Fraser P. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science. 2008; 322:1717–20.
35. Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010; 329:689–93.
36. Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014; 15:7–21.
37. Wang H, Chen Y, Lu XA, Liu G, Fu Y, Luo Y. Endostatin Prevents Dietary-Induced Obesity by Inhibiting Adipogenesis and Angiogenesis. Diabetes. 2015; 64:2442–56.
38. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014; 30:2114–20.
39. Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013; 29:15–21.
40. Anders S, McCarthy DJ, Chen Y, Okoniewski M, Smyth GK, Huber W, Robinson MD. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nat Protoc. 2013; 8:1765–86.
41. Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics. 2009; 10:48.
42. Shen S, Park JW, Lu ZX, Lin L, Henry MD, Wu YN, Zhou Q, Xing Y. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci USA. 2014; 111:E5593–601.
43. Anders S, Reyes A, Huber W. Detecting differential usage of exons from RNA-seq data. Genome Res. 2012; 22:2008–17.
44. Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009; 55:611–22.

