INTRODUCTION
Breast cancer is the most common cancer in women worldwide [1]. There are different therapeutic options for patients with invasive breast cancer, depending on the presented subtype. Systemic treatment for breast cancer consists of chemotherapy, endocrine or targeted therapy, guided by hormone receptor or HER2/neu status and other clinico-pathological features. For HER2/neu positive disease, targeted therapies are available, including monoclonal antibodies (trastuzumab or pertuzumab), the antibody-drug conjugate trastuzumab-emtansine or the tyrosine kinase inhibitor lapatinib. However, during the course of treatment, a number of patients gain resistance to the current therapy. Therefore, the development of innovative biomarker concepts and additional therapeutic strategies for breast cancer patients is of high clinical importance.
Prostate stem cell antigen (PSCA) is located on chromosome 8q24.2, encodes for a 123 amino acid glycosylphosphatidylinositol (GPI)-anchored cell surface protein and belongs to the Thy-1/Ly-6 family. It was originally defined as an upregulated gene in a human prostate cancer LAPC-4 xenograft model [2, 3]. Therefore, subsequent studies on its potential clinical application primarily focused on prostate cancer. It was shown that PSCA is expressed in 94% of all primary prostate cancers and that its expression correlates with advanced clinical stage, invasion to seminal vesicles, capsular invasion and prostate cancer progression to an androgen independent state [2, 4–6]. Moreover, PSCA mRNA detection in the peripheral blood of prostate cancer patients was shown to be of prognostic relevance [7]. Interestingly, besides its diagnostic potential, PSCA was also evaluated as therapeutic target. In this context, immunotherapies, such as the PSCA-mediated re-direction of T-lymphocytes towards prostate tumor cells or PSCA-mediated vaccination strategies have been proposed [8–10].
PSCA shares 30% homology with the stem cell antigen type 2 (SCA-2), which is an immature lymphocyte cell surface marker. However, considering this rather weak homology to SCA-2, PSCA was inaccurately named, since it is actually neither a marker for a stem cell population, nor is PSCA exclusively expressed in prostatic tissue [3]. The function of PSCA is not yet fully understood. It is believed that this protein is associated with the IFNα/β mediated immune response [11]. Subsequent reports showed that PSCA is likewise up-regulated in other cancer entities, such as gallbladder, urinary bladder cancer, renal cell carcinoma, pancreatic cancer or glioma [12–16], while it is down-regulated in others, such as esophageal and gastric cancers [17–20]. The clinical utility of PSCA as a diagnostic marker or therapeutic target has been demonstrated in prostrate, pancreatic and bladder cancer. Hitherto, PSCA-protein expression was described in only a few normal tissues including prostate epithelium, epithelial layers of the urinary bladder, neuroendocrine cells of the stomach and colon, collecting ducts of the kidney and trophoblasts of the placenta, with conflicting reports about its expression in the normal pancreas [21–23]. PSCA mRNA expression is predominantly found in prostate, placenta, kidney and urogenital tissues [21, 22]. This selective expression in normal tissue makes PSCA a suitable target for immunotherapy.
Currently, for breast cancer patients, there is limited data on PSCA. Some of these investigations have an epidemiological focus and report on genetic variation of the PSCA gene and its relation to breast cancer development. In this context, it was reported, for instance, that PSCA single nucleotide polymorphims (e.g. rs2294008 or rs2978974) are associated with increased risk of developing breast cancer [24]. Moreover, there is some evidence by two preliminary studies suggesting that PSCA-protein might be expressed in at least a subpopulation of breast cancer patients [25, 26]. However one of these studies only focusses on a limited set of patients with micropapillary carcinoma of the breast. The other study investigated PSCA-protein in only 20 fresh frozen breast cancer tissues and suggested that PSCA is rather downregulated in malignant compared to healthy breast tissue. Therefore, considering that clinical relevance of PSCA-protein expression in breast cancer patients is still an open and important question, the objective of the current study was to i) analyze expression pattern and compartmental distribution of the PSCA-protein in normal breast tissue and in breast cancer tissue from a comprehensive cohort of 405 clinically documented breast cancer patients ii) to correlate PSCA-expression to the patient’s clinicopathological data and, finally, iii) to investigate prognostic significance of PSCA-expression for breast cancer.
RESULTS
Immunohistochemical PSCA-staining in breast cancer tissue and normal breast
In order to exclude a heterogeneous PSCA-expression as well as to assess expression pattern and compartmental distribution of PSCA-protein in breast cancer patients, we performed immunohistochemical PSCA-staining in conventional tissue sections from primary breast tumors. PSCA-staining, which was positive in 4 of 18 samples (22%), appeared homogeneously and was observed with membranous and occasionally cytoplasmic PSCA-reactivity (Figure 1A). PSCA-staining was virtually absent in the vasculature (comprising endothelial cells and smooth muscle cells of the vascular wall), in lymph endothelial cells and in the tumor stroma (comprising fibroblasts, inflammatory cells and fibro-muscular cells, Figure 1B, Figure 2A and 2B). Healthy breast tissue, adjacent to the analyzed tumors (n=18), or breast tissue in specimens from breast reduction surgery (n=2), showed no PSCA-expression (Figure 1B).
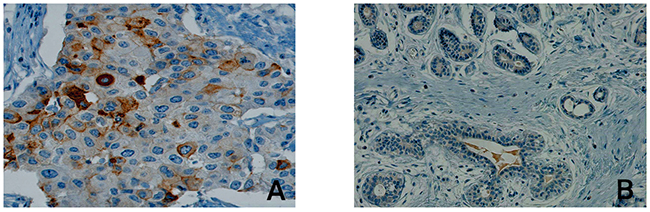
Figure 1: Compartmental distribution of PSCA-expression in breast cancer tissue. The Figure shows representative immunohistological PSCA-staining results in (A) malignant breast cancer tissue including tumor stroma and (B) in non-neoplastic breast tissue. Besides membranous and cytoplasmatic expression of PSCA in invasive breast cancer cells, PSCA-expression is completely absent in stromal cells and in non-neoplastic breast tissue.
PSCA-expression in breast cancer and its correlation with clinico-pathological features
Given the spatially homogeneous staining in PSCA-positive breast cancer tissues, we subsequently performed PSCA-tissue microarray analysis in a comprehensive set of clinically documented breast cancer patients (n=405). PSCA-expression was observed with a positivity rate of 23% (94/405 patients). Low PSCA-expression was detected in 16% (66/405 patients) and a high expression was observed in 7% (28/405 patients, Figure 2 and 3A). Subsequently, we correlated PSCA-expression with the patients’ clinico-pathological data (Table 1). PSCA-protein expression was significantly associated with unfavorable histopathological grade (p=0.011) and higher Ki67 proliferation index (p=0.006). No association with age, histological subtype, tumor stage, nodal status, lymphatic-invasion or angio-invasion was observed.
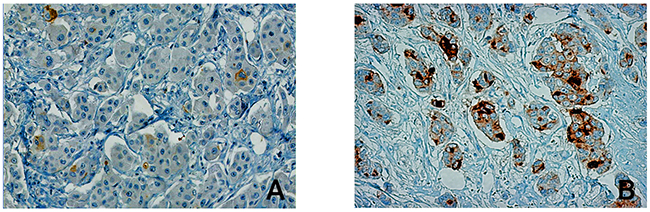
Figure 2: PSCA-expression levels in breast cancer tissue. The Figure shows a representative immunohistological PSCA-staining in malignant breast cancer tissue. Among the studied breast cancer patients, PSCA-expression was detected (A) in only a few disseminated tumor cells with weak intensity, referred to as “low expression” or (B) in a high number of invasive tumor cells, referred to as “high expression”.
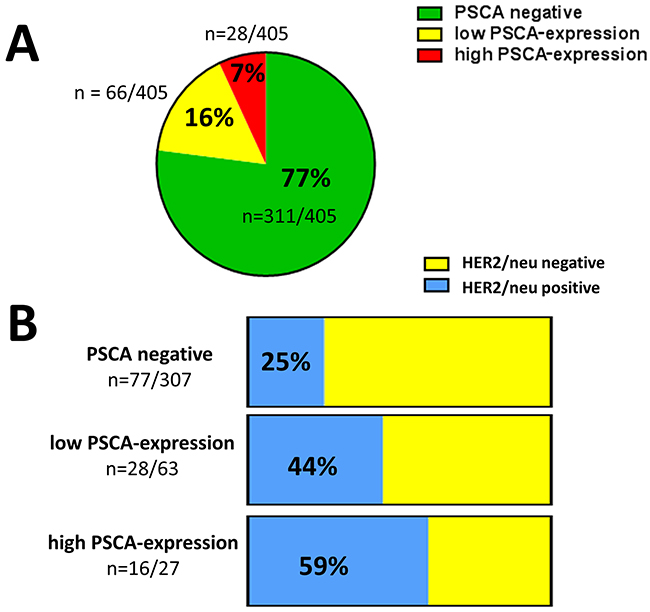
Figure 3: PSCA-expression in breast cancer and its association with HER2/neu status. (A) The pie chart shows the proportion of low and high PSCA-expression among the studied breast cancer patients according to TMA-analysis. (B) The stacked columns show the relative distribution of HER2/neu positive and HER2/neu negative patients with regard to the PSCA-status. Absolute patient numbers of each subgroup are indicated.
Table 1: Correlation of PSCA-expression with clinico-pathological parameter
PSCA negative |
PSCA low |
PSCA high |
p-value |
|
|---|---|---|---|---|
Her-2/neu status |
||||
HER2-positive |
77 (25.1) |
28 (44.4) |
16 (59.3) |
< 0.001 |
HER2-negative |
230 (74.9) |
35 (55.6) |
11 (40.7) |
|
histopathological grade |
||||
G1 |
41 (13.2) |
5 (7.6) |
0.011 |
|
G2 |
124 (39.9) |
25 (37.9) |
10 (35.7) |
|
G3 |
146 (46.9) |
36 (54.5) |
18 (64.3) |
|
Ki67 proliferation index |
||||
0-9% |
126 (41.3) |
21 (31.8) |
4 (14.3) |
0.006 |
10-20% |
81 (26.6) |
18 (27.3) |
11 (39.3) |
|
>20% |
98 (32.1) |
27 (40.9) |
13 (46.4) |
|
Age, n (%) |
||||
≤ 50 years |
89 (28.6) |
11 (16.7) |
6 (21.4) |
0.112 |
>50 years |
222 (71.4) |
55 (83.3) |
22 (78.6) |
|
ER Hormone receptor status |
||||
ER negative |
115 (37) |
22 (33.3) |
16 (57.1) |
0.078 |
ER positive |
196 (63) |
44 (66.7) |
12 (42.9) |
|
PR Hormone receptor status |
||||
PR negative |
131 (42.1) |
23 (34.8) |
15 (53.6) |
0.232 |
PR positive |
180 (57.9) |
43 (65.2) |
13 (46.4) |
|
Combination of hormone receptor and HER-2/neu status |
||||
triple negative |
75 (24.1) |
8 (12.3) |
7 (25) |
0.108 |
no triple negative |
236 (75.9) |
57 (87.7) |
21 (75) |
|
Histological type |
||||
no special type |
261 (83.9) |
61 (92.4) |
25 (89.3) |
0.172 |
invasive lobular |
25 (8) |
2 (3) |
||
other types |
25 (8) |
3 (4.5) |
3 (10.7) |
|
pT-stage |
||||
pT1 |
130 (41.8) |
31 (47) |
13 (46.4) |
0.690 |
pT2/pT3/pT4 |
181 (58.2) |
35 (53) |
15 (53.6) |
|
nodal status |
||||
pN0 |
161 (56.1) |
28 (45.2) |
12 (46.2) |
0.215 |
pN1 - pN3 |
126 (40.5) |
34 (51.5) |
14 (50) |
To sum up, PSCA-expression in breast cancer is associated with unfavorable histopathological grade and increased proliferative activity, however, it does not correlate with other clinico-pathological features.
PSCA-expression and ER, PR, HER2/neu receptor status
We inquired, whether there is an association between PSCA-expression and receptor status in breast cancer patients. No correlation between estrogen receptor status and progesterone receptor status with PSCA-protein expression was observed. Interestingly, there was a very strong correlation between PSCA-protein expression and HER2/neu receptor status (p<0.001, Table 1). Therefore, in the subgroup of patients with low PSCA-expression, we observed a considerably higher proportion of HER2/neu positivity than in patients with non-detectable PSCA (44% vs. 25%). Interestingly, for patients with high PSCA-expression, the proportion of HER2/neu positivity further increased up to 59% (Figure 3B). Of note, the distribution of PSCA-expression among triple negative patients was comparable to the total population.
Taken together, we demonstrate that PSCA-positive breast cancer patients, especially those with high PSCA-expression, are more likely to overexpress HER2/neu.
Prognostic significance of PSCA-expression in breast cancer
In order to assess prognostic significance of PSCA in breast cancer patients, we correlated PSCA-expression status with the patient’s survival data. No association was observed between PSCA-protein expression and PFS (p=0.44) or OS (p=0.89; Figure 4).
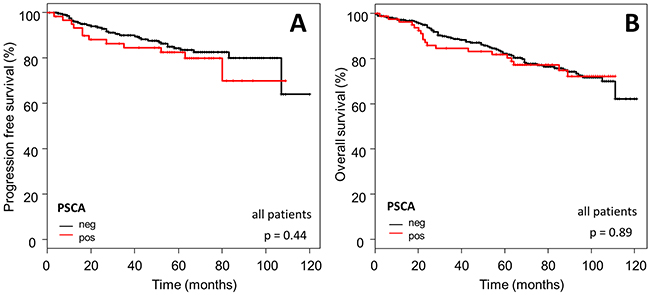
Figure 4: Prognostic relevance of PSCA-expression in the total breast cancer population. The Kaplan-Meier plots show (A) progression-free survival and (B) overall survival of the total study population with PSCA-positive (red curves) vs. PSCA-negative tumors (black curves).
Subsequently, we confined our survival analysis, with additional stratification into clinically relevant subgroups of breast cancer. In accordance to the overall Kaplan-Meier analysis described above, PSCA-protein did not provide prognostic information in the subgroup of i) HER2/neu overexpressing patients (n=110 for OS, p=0.87; n=74 for PFS, p=0.61) or ii) patients with intermediate risk, according to the criteria of St. Gallen 2007 [27] (n=236 for OS, p=0.39; n=182 for PFS, p=0.46, Figure 5 and 6).
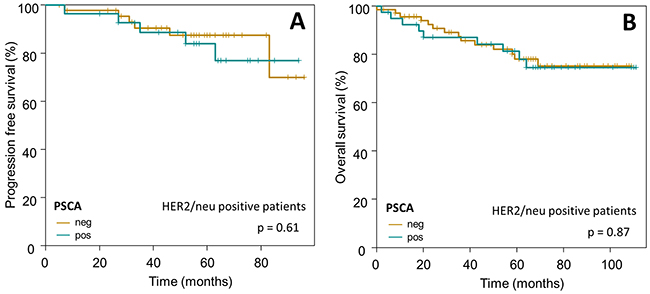
Figure 5: Prognostic relevance of PSCA-expression in HER2/neu positive breast cancer patients. The Kaplan-Meier plots show (A) progression-free survival and (B) overall survival of HER2/neu positive breast cancer patients with PSCA-positive (dark-green curves) vs. PSCA-negative tumors (olive-green curves).
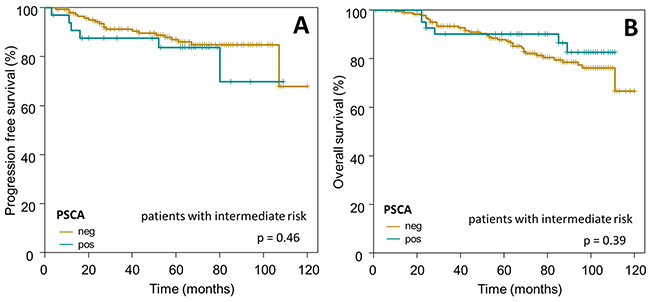
Figure 6: Prognostic relevance of PSCA-expression in breast cancer patients with intermediate risk. The Kaplan-Meier plots show (A) progression-free survival and (B) overall survival of intermediate risk breast cancer patients with PSCA-positive (dark-green curves) vs. PSCA-negative tumors (olive-green curves).
In summary, PSCA does not provide prognostic significance in our breast cancer cohort, neither in the overall study population nor in clinically relevant breast cancer subgroups.
DISCUSSION
In the present study, we demonstrate clinical relevance of PSCA-expression in breast cancer patients. We observed homogenous distribution of PSCA-expressing tumor cells in breast cancer tissue and expected that a TMA-analysis of PSCA-expression will allow conclusions on PSCA-expression across a large number of patients. This assumption was corroborated by the fact that PSCA-positivity rate in conventional breast cancer tissue sections almost exactly corresponded to our TMA-analysis (22% vs. 23%). Although this positivity rate is smaller than in other cancer entities, such as prostate or pancreatic cancer, in which PSCA is up-regulated in 60- 80% [4, 13], we were able to define a subgroup of PSCA-positive breast cancer patients, potentially considered for a PSCA-directed therapy. We observed mostly membranous PSCA-staining of breast cancer cells, which is in accordance with the fact that PSCA is a cell surface antigen [2, 3]. Virtual absence of PSCA-immunoreactivity in the tumor stroma is an interesting finding, since e.g. in prostate cancer, PSCA-protein expression occurs not only in epithelial but also in stromal compartments [28]. Moreover, outside the prostate, immunohistochemical studies in normal human tissues reported on PSCA-staining in the bladder, stomach, colon and kidney [4]. We did not detect PSCA-expression in healthy breast tissues, which is in contrast to a recent study on breast cancer, reporting that PSCA is strongly expressed in normal adjacent tissue but not in the tumor [26]. However, these findings are not directly comparable to our results, since this study included only a very limited number of fresh frozen breast cancer tissues (n=20) and a different staining protocol was applied. Our results indicate that PSCA-expression in breast cancer is restricted to tumor cells and is rather induced as a “neo-antigen” or at least up-regulated in a subgroup of invasive breast cancers.
We did not find an association of PSCA-expression and the patients’ survival, not even in consideration of clinically relevant breast cancer subgroups. This is in accordance to the aforementioned study on breast cancer, which also failed to show any statistically significant associations between PSCA mRNA expression and the patient’s clinico-pathological data [26] and could be explained by the heterogeneity of breast cancer disease. Therefore, we conclude that PSCA is not directly suitable as a prognostic biomarker for breast cancer.
Little is known about the function of PSCA. PSCA could possibly interact with other membranous proteins in order to activate downstream signalling pathways. Alternatively, PSCA could be shed from the cellular membrane through the cleavage of its GPI-anchor by phospholipase C and may act as a secreted ligand, which activates other receptor-mediated pathways [29]. Recent functional studies suggested that PSCA could suppress tumor growth and metastasis by inhibiting epithelial-mesenchymal transition (EMT) or by immunomodulation [30–32]. However, since context dependent tumor-suppressive as well as oncogenic functions have been reported, the exact role of PSCA in tumorigenesis is not fully understood. This is in contrast to a similar protein, prostate specific membrane antigen (PSMA), which is highly expressed in the prostate and which has well characterized enzymatic functions (so called “NAALADase” and folate hydrolase activity). PSMA is not only strongly expressed in prostate cancer, it is also upregulated in the neovasculature of solid tumors [33]. Nevertheless, our observations corroborate the assumption, that PSCA-expression parallels a more aggressive and proliferative tumor phenotype and may therefore be rather of oncogenic than tumor-suppressive character in breast cancer. We observed a very strong correlation between PSCA-positivity and HER2/neu overexpression. HER2/neu is a 185-kDa transmembrane oncoprotein, which is overexpressed in about 15-20% of breast cancer patients, who benefit from a HER2/neu directed therapy [34–37]. This finding is of significant clinical interest, since we showed that PSCA-expression status parallels a clinically relevant subgroup of breast cancer patients, eligible for HER2/neu directed therapy. This finding encourages further investigation, to determine whether PSCA-expression could be a response predictor for HER2/neu directed therapy. However, this question could not be addressed in the present study, because a part of our patients were diagnosed before trastuzumab administration became clinical standard for primary breast cancer [36]. Thus, the number of patients, which were HER2/neu overexpressing and actually treated with trastuzumab in our cohort, was too small for reasonable comparison between PSCA-status in trastuzumab responders and non-responders in a statistically substantiated manner.
PSCA has attracted some attention as a therapeutic target. Early clinical trials using full-length antibodies against PSCA in prostate and pancreatic cancer have failed to show signs of efficacy [38, 39]. Consequently, more potent targeting approaches have been developed in a preclinical setting, such as bispecific antibodies for re-direction of T-lymphocytes towards PSCA-positive tumor cells [8, 9, 40, 41]] or chimeric antigen receptor (CAR)-based strategies [42–47]. Of note, a first anti-PSCA CAR trial in pancreatic carcinoma has recently been initiated (NCT02744287). Moreover, radiolabeled antibody derivatives for tumor targeting [48] and PSCA-mediated vaccination strategies have been proposed [49]. Meantime, a variety of approved HER2/neu directed therapies are available, however patients often develop resistance to this kind of treatment [50]. The strong correlation between HER2/neu and PSCA-expression points towards the possibility of targeting PSCA in case of resistance against HER2/neu directed therapy.
Conclusively, we provide the first comprehensive analysis of PSCA-expression and its clinical relevance in breast cancer. Our results suggest that PSCA-based immunotherapeutic and theragnostic approaches may become valuable additions to breast cancer therapy, especially for patients resistant to therapies targeting HER2/neu and to the subset of triple negative patients with PSCA-positive tumors.
MATERIALS AND METHODS
Patient characteristics
We analyzed formalin fixed paraffin embedded (FFPE) cancer tissue of 405 primary breast cancers patients, treated according to uniform guidelines in the Regional Breast Cancer Center Dresden between 2003 and 2011. This study was approved by the ethics committee of the University Hospital of Dresden, Germany (EK59032007) and was performed, according to the Declaration of Helsinki. Since this study was performed retrospectively on long term archived FFPE breast cancer tissue without interventions, no patient’s consent was required by the guidelines of ethics committee. From each breast cancer, at least two cores were included in the tissue microarrays, resulting in a total number of five microarrays. In order to increase statistical power of comparison of breast cancers with different biological features, two of the tissue microarrays were enriched with triple negative and HER2/neu positive breast cancers cases. In order to exclude heterogeneous expression as well as to assess expression pattern and compartmental distribution of PSCA-expression in breast cancer patients, we performed immunohistochemical PSCA-staining in conventional tissue sections from primary breast tumors (n=18). To check the PSCA-expression in normal breast tissue, the tumor free cancer surrounding tissue in the conventional sections and two breast reduction specimens were analyzed. Table 2 summarizes the patient’s clinico-pathological characteristics. Follow up data were available for 366 patients. The follow up time ranged from 1 to 121 months (median follow up 84 months). Twenty four patients died of breast cancer between 10 and 105 month after first surgery (5-year overall survival 82%, 95%-CI [78% - 87%]).
Table 2: Patient characteristics at primary diagnosis of breast cancer
n (%) |
|
|---|---|
Histological subtype |
|
no special type |
347 (85.7) |
invasive lobular |
27 (6.7) |
other subtypes |
31 (7.7) |
pT-stage |
|
pT1 |
174 (43) |
pT2 |
181 (44.7) |
pT3 |
24 (5.9) |
pT4 |
26 (6.4) |
nodal status |
|
pN0 |
201 (49.6) |
pN1 - pN3 |
174 (43) |
unknown |
30 (7.4) |
histopathological grade |
|
G1 |
46 (11.4) |
G2 |
159 (39.3) |
G3 |
200 (49.4) |
hormone receptor status |
|
ER positive |
252 (62.2) |
ER negative |
153 (37.8) |
PR positive |
236 (58.3) |
PR negative |
169 (41.7) |
Her-2/neu status |
|
HER2-positive |
121 (29.9) |
HER2-negative |
276 (68.1) |
HER2/neu stage unknown |
8 (2) |
triple negative |
90 (22.2) |
PSCA-expression |
|
negative |
311 (76.8) |
low |
66 (16.3) |
high |
28 (6.9) |
Immunohistochemistry
The expression of PSCA, estrogen and progesterone receptor, HER2/neu and Ki67 was analysed by immunohistochemistry. All staining results were scored semi-quantitatively considering the percentage of positive cells and the staining intensity. The staining result for PSCA was subdivided in negative (no positive cells), low expression (1 to 9% of tumor cells positive) and high expression (>9% positive tumor cells). Table 3 gives an overview of the immunohistochemical analyses, including the antibodies used and the stratification for the staining results. For a more detailed analysis of the outcome, the cases were stratified in three prognosis groups, according to the criteria of St. Gallen 2007 [27] and HER2/neu expression.
Table 3: Used antibodies and positivity thresholds for immunohistochemistry
Antigen |
Antibody |
Dilution |
Stratification of staining results |
|---|---|---|---|
PSCA |
MB1[4] |
5 ng/μl |
1-9% weak positiv; >9% strong positiv |
Ki-67 |
MIB-1 (DAKO) |
1:50 |
0-9%, 10-20%, >20% |
estrogen receptor |
SP1 (Ventana) |
prediluted by manufacturer |
threshold for positivity: >1% |
progesterone receptor |
1E2 (Ventana) |
prediluted by manufacturer |
threshold for positivity: >1% |
HER-2/neu |
A0485 (DAKO) |
1:800 |
negative: IRS 0, 1+, 2+ without Her-2/neu |
Statistical analysis
Statistical analysis was performed by Chi-squared test for comparing the categorical clinico-pathological markers and PSCA-expression. The Cochrane-Armitage-trend test was performed for comparing ordered categorical variables and Kaplan-Meier estimation (log rank test) was used for survival analyses. Overall survival (OS) was defined from day of surgery until death. Progression-free survival (PFS) was defined from day of surgery until disease progression. OS and PFS for patients who did not die (for OS), respective suffer from progression/death (for PFS) during the observational period, were censored at the end of follow-up. Median follow up time was estimated using the reverse Kaplan-Meier-method. The significance threshold was p<0.05 for all tests. Because of the exploratory character of the analyses, no adjustment for multiple testing was performed.
Ethical approval and consent to participate
This study was approved by the ethics committee of the University Hospital of Dresden, Germany (EK59032007) and was performed, according to the Declaration of Helsinki. Since this study was performed retrospectively on long term archived FFPE breast cancer tissue, no patient’s consent was required.
Abbreviations
CAR, chimeric antigen receptor; EMT, epithelial-to-mesenchymal-transition; FFPE, formalin fixed paraffin embedded; GPI, glycosylphosphatidylinositol; OS, overall survival; PFS, progression free survival; PSCA, prostate stem cell antigen; SCA-2, stem cell antigen type 2 PSMA: Prostate-Specific Membrane Antigen.
Authors contributions
TL, FK, AE, JDK, MK, AW, AG, BR, GE, GB, MB, PW and KF made substantial contributions to the conception and design of the study, to the experimental work or to the acquisition of data and to the analysis/interpretation of the results. JDK, KF, TL and AE were involved in drafting the manuscript or revising it. All authors read and approved the manuscript in its final version.
ACKNOWLEDGMENTS
The authors want to thank Dr. Dora Tang for proof reading our manuscript as a native speaker.
CONFLICTS OF INTERESTS
AE, GE and MB hold patents related to an anti-PSCA antibody. The remaining authors declare that no conflict of interest exists.
FUNDING
The present study was supported by an internal budget from the Technical University of Dresden, Germany.
REFERENCES
1. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. International journal of cancer. 2015; 136:E359-386.
2. Reiter RE, Gu Z, Watabe T, Thomas G, Szigeti K, Davis E, Wahl M, Nisitani S, Yamashiro J, Le Beau MM, Loda M, Witte ON. Prostate stem cell antigen: a cell surface marker overexpressed in prostate cancer. Proc Natl Acad Sci USA. 1998; 95:1735–40.
3. Raff AB, Gray A, Kast WM. Prostate stem cell antigen: a prospective therapeutic and diagnostic target. Cancer Lett. 2009; 277:126–32.
4. Gu Z, Thomas G, Yamashiro J, Shintaku IP, Dorey F, Raitano A, Witte ON, Said JW, Loda M, Reiter RE. Prostate stem cell antigen (PSCA) expression increases with high gleason score, advanced stage and bone metastasis in prostate cancer. Oncogene. 2000; 19:1288–96.
5. Han KR, Seligson DB, Liu X, Horvath S, Shintaku PI, Thomas GV, Said JW, Reiter RE. Prostate stem cell antigen expression is associated with gleason score, seminal vesicle invasion and capsular invasion in prostate cancer. J Urol. 2004; 171:1117–21.
6. Zhigang Z, Wenlv S. Prostate stem cell antigen (PSCA) expression in human prostate cancer tissues and its potential role in prostate carcinogenesis and progression of prostate cancer. World J Surg Oncol. 2004; 2:13.
7. Hara N, Kasahara T, Kawasaki T, Bilim V, Obara K, Takahashi K, Tomita Y. Reverse transcription-polymerase chain reaction detection of prostate-specific antigen, prostate-specific membrane antigen, and prostate stem cell antigen in one milliliter of peripheral blood: value for the staging of prostate cancer. Clinical cancer research. 2002; 8:1794–1799.
8. Feldmann A, Arndt C, Töpfer K, Stamova S, Krone F, Cartellieri M, Koristka S, Michalk I, Lindemann D, Schmitz M, Temme A, Bornhäuser M, Ehninger G, Bachmann M. Novel humanized and highly efficient bispecific antibodies mediate killing of prostate stem cell antigen-expressing tumor cells by CD8+ and CD4+ T cells. J Immunol. 2012; 189:3249–59.
9. Arndt C, Feldmann A, Töpfer K, Koristka S, Cartellieri M, Temme A, Ehninger A, Ehninger G, Bachmann M. Redirection of CD4+ and CD8+ T lymphocytes via a novel antibody-based modular targeting system triggers efficient killing of PSCA+ prostate tumor cells. Prostate. 2014; 74:1347–58.
10. Xiao L, Joo KI, Lim M, Wang P. Dendritic cell-directed vaccination with a lentivector encoding PSCA for prostate cancer in mice. PLoS One. 2012; 7:e48866.
11. Marra E, Uva P, Viti V, Simonelli V, Dogliotti E, De Rinaldis E, Lahm A, La Monica N, Nicosia A, Ciliberto G, Palombo F. Growth delay of human bladder cancer cells by Prostate Stem Cell Antigen downregulation is associated with activation of immune signaling pathways. BMC Cancer. 2010; 10:129.
12. Elsamman EM, Fukumori T, Tanimoto S, Nakanishi R, Takahashi M, Toida K, Kanayama HO. The expression of prostate stem cell antigen in human clear cell renal cell carcinoma: a quantitative reverse transcriptase-polymerase chain reaction analysis. BJU Int. 2006; 98:668–73.
13. Argani P, Rosty C, Reiter RE, Wilentz RE, Murugesan SR, Leach SD, Ryu B, Skinner HG, Goggins M, Jaffee EM, Yeo CJ, Cameron JL, Kern SE, Hruban RH. Discovery of new markers of cancer through serial analysis of gene expression: prostate stem cell antigen is overexpressed in pancreatic adenocarcinoma. Cancer Res. 2001; 61:4320–24.
14. Geiger KD, Hendruschk S, Rieber EP, Morgenroth A, Weigle B, Juratli T, Senner V, Schackert G, Temme A. The prostate stem cell antigen represents a novel glioma-associated antigen. Oncol Rep. 2011; 26:13–21.
15. Amara N, Palapattu GS, Schrage M, Gu Z, Thomas GV, Dorey F, Said J, Reiter RE. Prostate stem cell antigen is overexpressed in human transitional cell carcinoma. Cancer Res. 2001; 61:4660–65.
16. Zou Q, Yang L, Yang Z, Huang J, Fu X. PSCA and Oct-4 expression in the benign and malignant lesions of gallbladder: implication for carcinogenesis, progression, and prognosis of gallbladder adenocarcinoma. Biomed Res Int. 2013; 2013:648420.
17. Bahrenberg G, Brauers A, Joost HG, Jakse G. Reduced expression of PSCA, a member of the LY-6 family of cell surface antigens, in bladder, esophagus, and stomach tumors. Biochem Biophys Res Commun. 2000; 275:783–88.
18. Sakamoto H, Yoshimura K, Saeki N, Katai H, Shimoda T, Matsuno Y, Saito D, Sugimura H, Tanioka F, Kato S, Matsukura N, Matsuda N, Nakamura T, et al. Genetic variation in PSCA is associated with susceptibility to diffuse-type gastric cancer. Nat Genet. 2008; 40:730–40.
19. Zhao X, Wang F, Hou M. Expression of stem cell markers nanog and PSCA in gastric cancer and its significance. Oncol Lett. 2016; 11:442–48.
20. Ono H, Hiraoka N, Lee YS, Woo SM, Lee WJ, Choi IJ, Saito A, Yanagihara K, Kanai Y, Ohnami S, Chiwaki F, Sasaki H, Sakamoto H, et al. Prostate stem cell antigen, a presumable organ-dependent tumor suppressor gene, is down-regulated in gallbladder carcinogenesis. Genes Chromosomes Cancer. 2012; 51:30–41.
21. Cunha AC, Weigle B, Kiessling A, Bachmann M, Rieber EP. Tissue-specificity of prostate specific antigens: comparative analysis of transcript levels in prostate and non-prostatic tissues. Cancer Lett. 2006; 236:229–38.
22. Ross S, Spencer SD, Holcomb I, Tan C, Hongo J, Devaux B, Rangell L, Keller GA, Schow P, Steeves RM, Lutz RJ, Frantz G, Hillan K, et al. Prostate stem cell antigen as therapy target: tissue expression and in vivo efficacy of an immunoconjugate. Cancer Res. 2002; 62:2546–53.
23. Ono H, Yanagihara K, Sakamoto H, Yoshida T, Saeki N. Prostate stem cell antigen gene is expressed in islets of pancreas. Anat Cell Biol. 2012; 45:149–54.
24. Wang M, Wang X, Fu SW, Liu X, Jin T, Kang H, Ma X, Lin S, Guan H, Zhang S, Liu K, Dai C, Zhu Y, Dai Z. Single-nucleotide polymorphisms in PSCA and the risk of breast cancer in a Chinese population. Oncotarget. 2016; 7:27665–75. doi: 10.18632/oncotarget.8491.
25. Hao JY, Yang YL, Li S, Qian XL, Liu FF, Fu L. [PSCA expression in invasive micropapillary carcinoma of breast]. [Article in Chinese]. Zhonghua Bing Li Xue Za Zhi. 2011; 40:382–6.
26. Martin TA, Jiang WG. Evaluation of the expression of stem cell markers in human breast cancer reveals a correlation with clinical progression and metastatic disease in ductal carcinoma. Oncol Rep. 2014; 31:262–72.
27. Goldhirsch A, Wood WC, Gelber RD, Coates AS, Thürlimann B, Senn HJ; 10th St. Gallen conference. Progress and promise: highlights of the international expert consensus on the primary therapy of early breast cancer 2007. Ann Oncol. 2007; 18:1133–44.
28. da Silva MM Jr, Matheus WE, Garcia PV, Stopiglia RM, Billis A, Ferreira U, Favaro WJ. Characterization of reactive stroma in prostate cancer: involvement of growth factors, metalloproteinase matrix, sexual hormones receptors and prostatic stem cells. Int Braz J Urol. 2015; 41:849–58.
29. Saeki N, Gu J, Yoshida T, Wu X. Prostate stem cell antigen: a Jekyll and Hyde molecule? Clin Cancer Res. 2010; 16:3533–38.
30. Wang L, Sang Y, Tang J, Zhang RH, Luo D, Chen M, Deng WG, Kang T. Down-regulation of prostate stem cell antigen (PSCA) by Slug promotes metastasis in nasopharyngeal carcinoma. J Pathol. 2015; 237:411–22.
31. Saeki N, Ono H, Sakamoto H, Yoshida T. Down-regulation of Immune-related Genes by PSCA in Gallbladder Cancer Cells Implanted into Mice. Anticancer Res. 2015; 35:2619–25.
32. Kang R, Zhao S, Liu L, Li F, Li E, Luo L, Xu L, Wan S, Zhao Z. Knockdown of PSCA induces EMT and decreases metastatic potentials of the human prostate cancer DU145 cells. Cancer Cell Int. 2016; 16:20.
33. Ghosh A, Heston WD. Tumor target prostate specific membrane antigen (PSMA) and its regulation in prostate cancer. J Cell Biochem. 2004; 91:528–39.
34. Schechter AL, Stern DF, Vaidyanathan L, Decker SJ, Drebin JA, Greene MI, Weinberg RA. The neu oncogene: an erb-B-related gene encoding a 185,000-Mr tumour antigen. Nature. 1984; 312:513–16.
35. Slamon DJ, Leyland-Jones B, Shak S, Fuchs H, Paton V, Bajamonde A, Fleming T, Eiermann W, Wolter J, Pegram M, Baselga J, Norton L. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med. 2001; 344:783–92.
36. Piccart-Gebhart MJ, Procter M, Leyland-Jones B, Goldhirsch A, Untch M, Smith I, Gianni L, Baselga J, Bell R, Jackisch C, Cameron D, Dowsett M, Barrios CH, et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med. 2005; 353:1659–72.
37. Varga Z, Noske A, Ramach C, Padberg B, Moch H. Assessment of HER2 status in breast cancer: overall positivity rate and accuracy by fluorescence in situ hybridization and immunohistochemistry in a single institution over 12 years: a quality control study. BMC Cancer. 2013; 13:615.
38. Morris MJ, Eisenberger MA, Pili R, Denmeade SR, Rathkopf D, Slovin SF, Farrelly J, Chudow JJ, Vincent M, Scher HI, Carducci MA. A phase I/IIA study of AGS-PSCA for castration-resistant prostate cancer. Annals of oncology. 2012; 23:2714–2719.
39. Wolpin BM, O’Reilly EM, Ko YJ, Blaszkowsky LS, Rarick M, Rocha-Lima CM, Ritch P, Chan E, Spratlin J, Macarulla T, McWhirter E, Pezet D, Lichinitser M, et al. Global, multicenter, randomized, phase II trial of gemcitabine and gemcitabine plus AGS-1C4D4 in patients with previously untreated, metastatic pancreatic cancer. Annals of oncology. 2013; 24:1792–1801.
40. Feldmann A, Stamova S, Bippes CC, Bartsch H, Wehner R, Schmitz M, Temme A, Cartellieri M, Bachmann M. Retargeting of T cells to prostate stem cell antigen expressing tumor cells: comparison of different antibody formats. Prostate. 2011; 71:998–1011.
41. Arndt C, Feldmann A, Koristka S, Cartellieri M, Dimmel M, Ehninger A, Ehninger G, Bachmann M. Simultaneous targeting of prostate stem cell antigen and prostate-specific membrane antigen improves the killing of prostate cancer cells using a novel modular T cell-retargeting system. Prostate. 2014; 74:1335–46.
42. Morgenroth A, Cartellieri M, Schmitz M, Günes S, Weigle B, Bachmann M, Abken H, Rieber EP, Temme A. Targeting of tumor cells expressing the prostate stem cell antigen (PSCA) using genetically engineered T-cells. Prostate. 2007; 67:1121–31.
43. Katari UL, Keirnan JM, Worth AC, Hodges SE, Leen AM, Fisher WE, Vera JF. Engineered T cells for pancreatic cancer treatment. HPB. 2011; 13:643–650.
44. Kloss CC, Condomines M, Cartellieri M, Bachmann M, Sadelain M. Combinatorial antigen recognition with balanced signaling promotes selective tumor eradication by engineered T cells. Nat Biotechnol. 2013; 31:71–75.
45. Abate-Daga D, Lagisetty KH, Tran E, Zheng Z, Gattinoni L, Yu Z, Burns WR, Miermont AM, Teper Y, Rudloff U, Restifo NP, Feldman SA, Rosenberg SA, Morgan RA. A novel chimeric antigen receptor against prostate stem cell antigen mediates tumor destruction in a humanized mouse model of pancreatic cancer. Hum Gene Ther. 2014; 25:1003–12.
46. Cartellieri M, Koristka S, Arndt C, Feldmann A, Stamova S, von Bonin M, Töpfer K, Krüger T, Geib M, Michalk I, Temme A, Bornhäuser M, Lindemann D, et al. A novel ex vivo isolation and expansion procedure for chimeric antigen receptor engrafted human T cells. PLoS One. 2014; 9:e93745.
47. Hillerdal V, Ramachandran M, Leja J, Essand M. Systemic treatment with CAR-engineered T cells against PSCA delays subcutaneous tumor growth and prolongs survival of mice. BMC Cancer. 2014; 14:30.
48. Olafsen T, Gu Z, Sherman MA, Leyton JV, Witkosky ME, Shively JE, Raubitschek AA, Morrison SL, Wu AM, Reiter RE. Targeting, imaging, and therapy using a humanized antiprostate stem cell antigen (PSCA) antibody. J Immunother. 2007; 30:396–405.
49. Ahmad S, Casey G, Sweeney P, Tangney M, O’Sullivan GC. Prostate stem cell antigen DNA vaccination breaks tolerance to self-antigen and inhibits prostate cancer growth. Molecular therapy. 2009; 17:1101–1108.
50. Pohlmann PR, Mayer IA, Mernaugh R. Resistance to Trastuzumab in Breast Cancer. Clin Cancer Res. 2009; 15:7479–91.
51. Wolff AC, Hammond ME, Schwartz JN, Hagerty KL, Allred DC, Cote RJ, Dowsett M, Fitzgibbons PL, Hanna WM, Langer A, McShane LM, Paik S, Pegram MD, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. Arch Pathol Lab Med. 2007; 131:18–43.

