INTRODUCTION
Breast cancer is the most common female cancer, with more than 230,000 new cases estimated to be diagnosed in the United States in 2015 alone [1]. Several studies found that the average rates of recurrence were 11-30% at 5 years and 20-36.8% at 10 years after completion of initial treatments [2–5]. Breast cancer recurrence can be generally categorized into three types: local, regional and distant recurrence, whereas the first two are often combined into loco-regional recurrence [6, 7]. The 5-year PFS and OS were found to be 45% and 71% in the local recurrence group and 34% and 58% in the regional recurrence group [8–10].
To date researchers have identified several factors that are associated with breast cancer recurrence such as age, tumor size, focality, lymph node involvement, grade, estrogen receptor (ER) status, progesterone receptor (PR) status, human epidermal growth factor receptor 2 (HER2) status and Ki67 expression [11–14]. These factors are determined in the primary tumor and are obtained through traditional tumor biopsy [15]. However a tissue sample cannot be continuously monitored during therapy and follow-up of cancer patients [16].
Blood-based biomarkers hold great promises as they are easily accessible and reproducible [17]. A major advantage of blood-based biomarkers, including properties of the cfDNA, in the context of cancer recurrence is the fact that they can be monitored repeatedly, even after the primary tumor has been removed. In recent years blood-based biomarkers such as microRNAs, circulating tumor cells and others have been investigated for the diagnosis and prognosis of breast cancer [18–24]. Circulating DNA is described as cell-free DNA (cfDNA) or circulating tumor DNA present in serum or plasma [25, 26] and elevated cfDNA concentrations have been observed in some types of cancers [17]. The extent of cfDNA fragmentation has also been used in addition to cfDNA concentrations [27]. Generally, the cell free DNA integrity (cfDI) is calculated as the ratio of longer DNA fragments concentration to shorter ones from a specific genetic locus. cfDI has been studied as a biomarker for detection of some types of tumors like head and neck [28], breast cancer [29], renal cancer [30] and acute leukemia [31] et al. As cfDNA integrity can be monitored repeatedly even after the primary tumor has been removed, cfDI might provide the opportunity for an early detection of cancer relapse.
Here in this prospective nested study we aimed to investigate if cfDI can be a biomarker for predicting BC recurrence during the follow-up of BC patients after initial treatment.
RESULTS
Altered cfDI and cfDNA concentrations prior to breast cancer recurrence
The results between the independently measured ALU and LINE1 elements were consistent with high correlation both for log2cfDNA concentration and cfDI (Supplementary Figure 1). As shown in Table 1 and Figure 1, patients with impending recurrence had a significantly lower cfDI (median ALU cfDI = 0.52, median LINE1 cfDI = 0.39) compared to the group of non-recurrent patients (median ALU cfDI =0.62, median LINE1 cfDI = 0.54) (P<0.0001 for each). In contrast, the concentration differences of both ALU and LINE1 were not significant between the two patient groups (ALU: P = 0.16; LINE1: P = 0.17) (Supplementary Figure 2). ROC analysis revealed that cfDI can distinguish patients with impending recurrence from non-recurrent patients with an AUC of 0.710 for ALU and 0.704 for LINE1 (Figure 2A, 2B). When ALU and LINE1 cfDI data were combined, discriminatory power with an AUC of 0.732 was reached (Figure 2C).
Table 1: Mean and median cfDI and cfDNA concentration of recurrent and non-recurrent groups calculated from ALU and LINE1 targets, and P-values of Wilcoxon rank sum tests comparing cfDI and log2cfDNA concentration between recurrent and non-recurrent breast cancer patients
Group |
Index |
Recurrent Patients |
Non-Recurrent Patients |
Comparison |
||
|---|---|---|---|---|---|---|
Mean ± SD |
Median |
Mean ± SD |
Median |
P-value |
||
ALU |
cfDI |
0.51±0.14 |
0.52 |
0.62±0.16 |
0.62 |
7.95E-05 |
|
cfDNA conc (ng/μl) |
1.16±3.42 |
0.21 |
0.55±0.19 |
0.25 |
0.16 |
LINE1 |
cfDI |
0.43±0.15 |
0.39 |
0.56±0.19 |
0.54 |
1.11E-04 |
|
cfDNA conc (ng/μl) |
1.43±4.26 |
0.26 |
0.53±0.40 |
0.33 |
0.17 |
Statistically significant P < 0.05 in the univariate and multivariate analysis is highlighted in bold.
conc concentration, SD standard deviation.
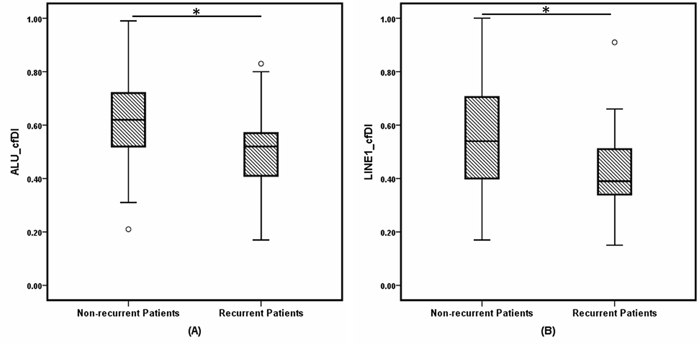
Figure 1: Box and whisker plots of cell-free DNA integrity (cfDI) in non-recurrent breast cancer patients and recurrent BC patients estimated from (A) ALU, (B) LINE1 targets. * indicates P less than 0.001.
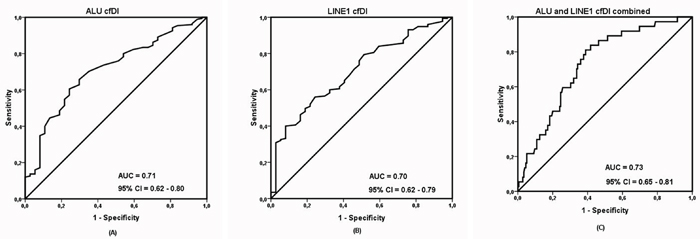
Figure 2: Receiver operating characteristic (ROC) analysis using (A) cell-free DNA integrity (cfDI) calculated from ALU targets, (B) cell-free DNA integrity calculated from LINE1 targets, (C) cfDI from ALU and LINE1 targets combined, to estimate the strength of the model to discriminate two groups, along with area under the curve (AUC) and 95% confidence interval (CI).
Correlation of cfDI and cfDNA concentration with clinical characteristics
To investigate the influence of clinical factors on the detected cfDI and cfDNA concentrations in the samples, the associations between these measures and various clinical characteristics was calculated. Here, a true association was determined only if both ALU and LINE1 elements showed consistent results. As shown in Supplementary Table 1, age was the only factor which showed a consistent association on cfDNA concentration (P = 0.013 for ALU, P = 0.015 for LINE1), whereas it showed no significant association of cfDI. No associations with other factors were observed. Also, primary tumor parameters, including histological type, grading, tumor size, nodal status, ER status, PR status, HER2 status showed no influence on cfDI or cfDNA concentration. To test if cfDI may be affected by the time after therapy until blood withdrawal, we analyzed the correlation between cfDI and the time of the first follow-up date to the time of blood withdrawal which showed no correlation (P = 0.65 for ALU cfDI, P= 0.90 for LINE1 cfDI). What’s more, we also found no significant difference of cfDI and cfDNA concentration of non-recurrent patients between this follow-up time to the follow-up time of the average recurrent patients (P > 0.1).
Univariate and multivariate analysis of factors related to breast cancer recurrence
Univariate analysis demonstrated that ALU and LINE1 cfDI, as well as primary tumor features such as tumor size, ER status, PR status, Ki67 expression level, and type of chemo-therapy were significantly associated with the recurrence status, as shown in Table 2. To evaluate if cfDI can predict breast cancer recurrence independent from the influence of these and other known factors related to recurrence [32], we performed multiple logistic regression analyses. The association of recurrence and cfDI remained significant (P = 0.020 and 0.019 for ALU and LINE1, respectively) with an odds ratio for developing recurrence of 3.69 (95% CI 1.23 – 11.02) for ALU cfDI and 3.74 (95% CI 1.24 – 11.27) for LINE1 cfDI. By using the highest cfDI quartile (Q4) as reference category in the interquartile analysis, it was shown that the risk for patients to develop BC recurrence significantly (P for trend = 0.011 for ALU and P for trend = 0.016 for LINE1) increased for patients in lower cfDI quartiles (Q3, Q2, Q1) compared to patients in the highest cfDI quartile, with an OR between the lowest and highest quartiles of 5.8 (95% CI 1.8 – 18.7) for ALU and10.9 (95% CI 2.4-50.7) for LINE1, as shown in Table 3 and 4. Finally, we constructed different multivariate models to investigate the prognostic ability of cfDI when added with clinical variables. In this way we calculated the corresponding area under the ROC curve was 0.82 (95% CI = 0.73 – 0.91) for clinical variables alone. When combined with cfDI, AUC was increased to 0.84 (95% CI = 0.75 – 0.92) for ALU cfDI and 0.84 (95% CI = 0.76 – 0.92) for LINE1 cfDI (Figure 3). Taken together, these observations confirmed that a decreased cfDI is associated with an increased risk of impending breast cancer recurrence and provide evidence that cfDI is an independent predictor BC recurrence.
Table 2: Univariate and multivariate analysis of factors related to recurrence in breast cancer patients
Variables |
|
ALU |
LINE1 |
||||||
|---|---|---|---|---|---|---|---|---|---|
univariate analysis |
multivariate analysis |
multivariate analysis |
|||||||
P value |
OR |
95%CI |
P value |
OR |
95%CI |
P value |
OR |
95%CI |
|
LINE1 cfDI |
8.45E-04 |
3.96 |
(1.77-8.88) |
|
|
|
0.019 |
3.74 |
(1.24-11.27) |
ALU cfDI |
5.63E-04 |
4.15 |
(1.85-9.31) |
0.020 |
3.69 |
(1.23-11.02) |
|
|
|
Age |
0.58 |
1.01 |
(0.98-1.04) |
0.65 |
0.99 |
(0.94-1.04) |
0.67 |
0.99 |
(0.94-1.04) |
Histotype |
0.46 |
0.84 |
(0.54-1.32) |
0.24 |
0.68 |
(0.35-1.23) |
0.25 |
0.67 |
(0.34-1.32) |
Grading |
0.059 |
0.63 |
(0.39-1.02) |
0.47 |
0.70 |
(0.27-1.84) |
0.49 |
0.72 |
(0.29-1.80) |
Focal |
0.70 |
1.20 |
(0.49-2.94) |
0.080 |
3.52 |
(0.86-14.37) |
0.042 |
4.32 |
(1.06-17.68) |
Tumor Size |
7.21E-3 |
0.49 |
(0.29-0.83) |
0.084 |
0.38 |
(0.13-1.14) |
0.082 |
0.42 |
(0.16-1.12) |
Nodus Stages |
0.039 |
0.47 |
(0.23-0.96) |
0.11 |
0.38 |
(0.12-1.24) |
0.063 |
0.32 |
(0.099-1.06) |
ER |
6.15E-03 |
3.13 |
(1.38-7.08) |
0.056 |
6.38 |
(0.95-42.71) |
0.078 |
5.28 |
(0.83-33.71) |
PR |
0.013 |
1.83 |
(0.84-3.99) |
0.25 |
0.35 |
(0.060-2.07) |
0.28 |
0.39 |
(0.073-2.12) |
HER2 |
0.69 |
1.26 |
(0.41-3.90) |
0.44 |
2.50 |
(0.25-24.92) |
0.38 |
2.88 |
(0.28-29.89) |
p53 |
0.53 |
1.01 |
(0.99-1.02) |
0.044 |
1.02 |
(1.00-1.05) |
0.097 |
1.02 |
(1.00-1.04) |
Ki67 |
0.066 |
0.98 |
(0.97-1.00) |
0.22 |
0.98 |
(0.96-1.01) |
0.13 |
0.98 |
(0.95-1.01) |
Operation |
0.76 |
0.90 |
(0.44-1.84) |
0.36 |
0.56 |
(0.16-1.96) |
0.35 |
0.55 |
(0.16-1.92) |
Radio_therapy |
0.54 |
1.34 |
(0.53-3.36) |
0.32 |
2.23 |
(0.45-10.99) |
0.28 |
2.33 |
(0.50-10.84) |
Chemo_therapy |
0.068 |
0.47 |
(0.21-1.06) |
0.13 |
0.37 |
(0.099-1.34) |
0.24 |
0.46 |
(0.124-1.69) |
Statistically significant P < 0.05 in the univariate and multivariate analysis is highlighted in bold.
Abbreviations: OR - odds ratio, CI - confidence interval.
Table 3: Association of ALU cfDI with recurrence of breast cancer
ALU |
Non-Recurrent Patients |
Recurrent Patients |
OR |
95%CI |
|---|---|---|---|---|
0.17-0.50 (Quartile 1) |
36 |
16 |
5.8 |
(1.8-18.7) |
0.50-0.59 (Quartile 2) |
41 |
11 |
3.5 |
(1.0-11.8) |
0.59-0.71 (Quartile 3) |
46 |
6 |
1.7 |
(0.5-6.4) |
0.71-0.99 (Quartile 4) |
52 |
4 |
1.00 (reference) |
|
P for trend |
0.011 |
|||
Statistically significant P < 0.05 in the univariate and multivariate analysis is highlighted in bold.
Abbreviations: OR - odds ratio, CI - confidence interval.
Table 4: Association of LINE1 cfDI with recurrence of breast cancer
LINE1 |
Non-Recurrent Patients |
Recurrent Patients |
OR |
95%CI |
|---|---|---|---|---|
0.15-0.38 (Quartile 1) |
37 |
15 |
10.9 |
(2.4-50.7) |
0.38-0.52 (Quartile 2) |
41 |
12 |
7.9 |
(1.7-37.3) |
0.52-0.67 (Quartile 3) |
43 |
8 |
5.0 |
(1.0-24.9) |
0.67-0.99 (Quartile 4) |
54 |
2 |
1.00 (reference) |
|
P for trend |
0.016 |
|
||
Statistically significant P < 0.05 in the univariate and multivariate analysis is highlighted in bold.
Abbreviations: OR - odds ratio, CI - confidence interval.
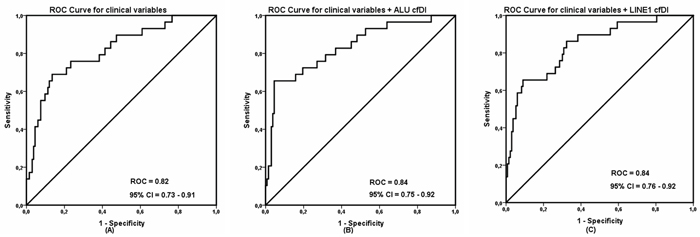
Figure 3: Receiver operating characteristic (ROC) analysis using (A) only clinical variables, (B) clinical variables with ALU cell-free DNA integrity and (C) clinical variables with LINE1 cell-free DNA integrity to estimate the strength of the model to discriminate two groups, along with area under the curve (AUC) and 95% confidence interval (CI).
DISCUSSION
By using a prospective nested study design, we identified that cfDI can be an independent predictor of impending breast cancer recurrence during the follow-up of breast cancer cases. To our knowledge, this is the first study that investigates plasma cfDI as an independent marker for prediction of breast cancer recurrence.
We observed significant decrease of cfDI in the group of patients who had impending recurrence compared to the group of non-recurrent patients (P < 0.001 for ALU and LINE1). The area under the curve (AUC) for ALU and LINE1 is 0.710 and 0.704 indicates that cfDI has moderate ability to differentiate recurrent breast cancer patients from non-recurrent breast cancer subjects. What is more, multivariate analysis confirmed a significant association of recurrence and cfDI (OR = 3.69 for ALU and 3.74 for LINE1). cfDI can also improve the discrimination ability when combined with clinical variables.
The investigation of circulating molecular markers in peripheral blood (“liquid biopsies”) is of great importance because of the advantages such as easily accessible, reproducible and early detectable in cancer [20]. Many biomarkers like microRNA, circulating DNA, circulating tumor cells are now being investigated as markers of diagnosis or prognosis in different types of cancer and these markers show great potential in the clinical utility [33–36]. Among these, cell-free circulating DNA integrity (cfDI) is becoming a potential biomarker for cancer diagnosis and prognosis in recent years.
Several studies have identified an altered integrity of cfDNA in malignant cancer patients compared to healthy controls. However these studies are heterogeneous with some studies showed a reduced cfDI in malignant cancer patients [6, 37–39], while others reported an increased cfDI [27, 28, 40, 41]. Many hypotheses have been applied to explain it. With direct visualization by gel electrophoresis, Giacona et al found that cfDNA from healthy individuals had three to five fold multiples of nucleosome-associated DNA length, and longer fragments compared to cfDNA in pancreatic cancer patients [42]. By massively parallel sequencing, Jiang found elevated amounts of shorter mitochondrial DNA molecules in plasma of carcinoma patients compared to healthy subjects [43]. On the other hand, it was hypothesized that in healthy controls cfDNA fragments are mainly released by apoptotic cells while in cfDNA in cancer patients they are thought to be predominantly released by malignant cells undergoing different pathophysiological processes including necrosis, autophagy, or mitotic catastrophe and thus show increased lengths [44]. However this explanation has not been supported by experimental proof so far, and necrotic DNA has been shown to account for increased cfDI in only a small fraction of cancer patients [45].
As the most abundant classes of repetitive DNA elements, ALU and LINE1 cover approximately 10% and 17% of the genome [46]. While the concentration of cfDNA can vary across several orders of magnitude [47], the cfDI as a ratio of nested long and short cfDNA fragment concentrations obtains standardized values between 0 and 1 which can be observed even in low cfDNA concentration. In addition, these measures reflect a global status of the circulating DNA rather than only one specific genetic locus. In this study, we observed a very good correlation from results between ALU and LINE1. Our concordant observations from independent cfDI measurements of these two elements minimize the possibility of false-positive results.
Many studies have identified the screening or diagnositic relevance of cfDI. However, few focus on the prognostic value of cfDI in cancer. Umetani et al measured serum DNA integrity in 51 healthy women and 83 patients with primary breast cancer by ALU-PCR and observed that cfDI was related to breast cancer AJCC (American Joint Committee on Cancer) stages [41]. It showed that serum DNA integrity was the predictor of lymph nodes metastasis by multivariate logistic analysis. Nevertheless, it only involved in lymph nodes metastasis. In another study about cfDI and its prognostic value in breast cancer, Iqbal et al compared DNA integrity in 25 relapsed and 61 non-relapsed patient samples and observed a significant difference between two groups (P = 0.005) [48]. DNA integrity also has significant association with disease free survival in breast cancer. Here we included all the types of recurrence including local, regional and distant metastases to observe whether cfDI can be a predictable biomarker for recurrence.
As the promising non-invasive biomarkers of clinic utility, it is of great importance to apply a standardized sample processing protocol. Firstly we prefer plasma to serum in cfDI study since studies have found the coagulation process in serum which would induce high variability of the spectrum of circulating nucleic acids [49]. In our study, we conducted a standardized protocol which has been validated in the other study [6]. Furthermore, for a specific gene the correct and validated primer design is essential. Besides its origin, factors for example time between sample collection and processing, plasma purification, the number of freeze-thaw cycles and the employed cfDNA extraction methods can affect cfDNA quality and quantity and thus cfDI measurements [50, 51]. We employed a standardized protocol that included sample processing within 2 hours of blood collection and a two-step centrifugation for plasma purification with a first step at 1300g for 20 min to separate the plasma from blood cells, followed by a second centrifugation step at 15500g for 10 min. This high-speed centrifugation of plasma removes cell debris and prevents interference of contaminating cellular DNA in cfDI measurements [6]. Prior to extraction, only one freeze-thaw cycle was ensured for all plasma samples.
In our study, subjects were included only if the patients has survival at least six months after treatment of the primary breast cancer, to avoid effects of surgery or chemotherapy on the cfDI. We did not observe an influence of the time period between the last time of therapy and blood withdrawal on the cfDI. Characteristics of the primary tumor, for example age, tumor size, focality, nodal status, hormone receptor status are known to be associated with breast cancer recurrence [12]. Likewise, here significant differences were observed in tumor size, ER, PR and p53 between non-recurrent and recurrent patients. Using binary logistic regression, we identified cfDI as an independent biomarker associated with recurrence.
Limitation of this study is the small sample size of the study however it is higher than the size of many other cfDI-related studies. It is not possible for us to compare in different subgroups in this study. Meanwhile the unbalanced number of recurrent and non-recurrent patients impedes the statistical power of this study. Large sample size studies are needed to confirm the results. Due to the lack of prospective studies with standardized processed sample material as mentioned above in other studies, our study is the first study of cfDI in plasma as a predictor in breast cancer recurrence so far. Nevertheless, it would be worthwhile to investigate cfDI combined with other blood-based biomarker in further prospective studies with excellent plasma material. Further it will be interesting to evaluate if cfDI can also contribute to the prediction of recurrence during disease follow-up in other cancers.
In summary, our study shows that cfDI can be an independent predictor of recurrence in breast cancer patients and might be a valuable marker as part of a molecular, blood-based multi-marker assay.
MATERIALS AND METHODS
Study subjects
The GENOM study was approved by the Ethical Committee of the University of Heidelberg (Heidelberg, Germany). Breast cancer patients with a primary tumor diagnosed between November 2008 and July 2015 were included. All subjects were females and Caucasians. For each patient blood samples were collected during follow-up visits in intervals of six months after initial therapy of the primary breast cancer. Recurrent patients were included if collected plasma samples from follow-up time points were available that fulfilled the following criteria: (1) the sample was taken 0-9 months before diagnosis of the recurrence and (2) the recurrence did not occur within the one year after initial therapy. Non-recurrent patients were included for plasma samples at FU3 which had been taken after at least 1 year of follow-up after therapy and at least 18 months of known, recurrence-free follow-up after sample collection. The clinical flow diagram for sample chosen was shown in Supplementary Figure 3. In total, 175 non-recurrent patients and 37 recurrent patients were included in this study. All patients’ demographic and clinical data are presented in Table 5.
Table 5: Distribution of clinical features of patients used in circulating DNA analyses
Characteristics |
Recurrent Patients |
Non-Recurrent Patients |
|||
|---|---|---|---|---|---|
n=37 |
(%) |
n=175 |
(%) |
||
Age |
Mean |
55.5 |
|
56.8 |
|
|
Median |
55 |
|
55 |
|
|
Range |
32-80 |
|
28-81 |
|
Menopausal status |
Pre |
12 |
(32.4%) |
58 |
(33.1%) |
|
Peri |
1 |
(2.7%) |
10 |
(5.7%) |
|
Post |
24 |
(64.8%) |
105 |
(60%) |
|
NA |
0 |
(0.0%) |
2 |
(1.1%) |
Histology |
IDC |
30 |
(81.1%) |
151 |
(86.3%) |
|
ILC |
4 |
(10.8%) |
18 |
(10.3%) |
|
DCIS |
1 |
(2.7%) |
5 |
(2.9%) |
|
NA |
2 |
(5.4%) |
1 |
(2.7%) |
Grading |
1 |
1 |
(2.7%) |
21 |
(14.3%) |
|
2 |
18 |
(48.6%) |
111 |
(60.9%) |
|
3 |
17 |
(45.9%) |
34 |
(18.0%) |
|
NA |
1 |
(2.7%) |
9 |
(6.8%) |
Focality |
Uni |
29 |
(78.4%) |
135 |
(75.2%) |
|
Multi |
7 |
(18.9%) |
39 |
(24.1%) |
|
NA |
1 |
(2.7%) |
1 |
(0.8%) |
Tumor Size |
Tis |
2 |
(5.4%) |
5 |
(2.8%) |
|
T0 |
2 |
(5.4%) |
15 |
(8.6%) |
|
T1 |
11 |
(29.7%) |
76 |
(43.4%) |
|
T2 |
14 |
(37.8%) |
67 |
(38.3%) |
|
T3 |
4 |
(10.8%) |
9 |
(5.1%) |
|
T4 |
4 |
(10.8%) |
2 |
(5.4%) |
Lymph node |
N0 |
21 |
(56.8%) |
121 |
(69.1%) |
|
N1 |
10 |
(27.0%) |
33 |
(18.9%) |
|
N2 |
1 |
(2.7%) |
12 |
(6.9%) |
|
N3 |
5 |
(13.5%) |
9 |
(5.1%) |
ER status |
Positive |
25 |
(67.6%) |
150 |
(87.2%) |
|
Negative |
12 |
(32.4%) |
23 |
(11.3%) |
|
NA |
0 |
(0.0%) |
2 |
(1.5%) |
PR Status |
Positive |
25 |
(67.6%) |
137 |
(82.0%) |
|
Negative |
12 |
(32.4%) |
36 |
(16.5%) |
|
NA |
0 |
(0.0%) |
2 |
(1.5%) |
HER2 Status |
Positive |
4 |
(10.8%) |
23 |
(3.8%) |
|
Negative |
32 |
(86.5%) |
146 |
(91.7%) |
|
NA |
1 |
(2.7%) |
6 |
(4.5%) |
p53 Score |
0-1 |
15 |
(40.5%) |
54 |
(30.9%) |
|
2-10 |
10 |
(27.0%) |
54 |
(30.9%) |
|
> 10 |
6 |
(16.2%) |
34 |
(19.4%) |
|
NA |
6 |
(16.2%) |
33 |
(18.9%) |
Ki67 Score |
1-10 |
6 |
(16.2%) |
79 |
(45.1%) |
|
11-20 |
8 |
(21.6%) |
33 |
(18.9%) |
|
21-50 |
11 |
(30.4%) |
33 |
(18.9%) |
|
> 50 |
10 |
(29.7%) |
22 |
(12.6%) |
|
NA |
2 |
(5.4%) |
8 |
(4.6%) |
Chemo_therapy |
Yes |
28 |
(75.7%) |
104 |
(57.9%) |
|
No |
9 |
(24.3%) |
71 |
(42.1%) |
Radio_therapy |
Yes |
30 |
(81.1%) |
149 |
(86.5%) |
|
No |
7 |
(18.9%) |
26 |
(13.5%) |
Endocrine_therapy |
Yes |
24 |
(64.9%) |
117 |
(63.2%) |
|
No |
13 |
(35.1%) |
58 |
(36.8%) |
Surgical Type |
BCT |
25 |
(67.6%) |
132 |
(78.9%) |
|
Mastectomy |
10 |
(32.4%) |
42 |
(21.1%) |
|
NA |
2 |
(32.4%) |
1 |
(21.1%) |
Distant Recurrence No. |
27 |
lung |
8 |
|
|
|
|
liver |
13 |
|
|
|
|
bone |
20 |
|
|
|
|
other |
15 |
|
|
Local-regional Recurrence No. |
10 |
local |
7 |
|
|
|
|
regional |
3 |
|
|
Abbreviations: ER - oestrogen receptor, PR - progesterone receptor, HER2 - human epidermal growth factor 2, T - tumor size, N - lymph node status, BCT - breast conserving therapy.
Sample collection and cfDNA extraction
Peripheral blood was collected from all patients in 9 ml EDTA tubes (S-Monovette R, Sarstedt, Nümbrecht, Germany). Blood was centrifuged at 1300g for 20 min at 10°C within two hours of blood withdrawal. The supernatant was transferred and centrifuged at 15500g for 10 min at 10°C. This step was done to make sure that the plasma was free of cells or cell debris. The plasma supernatant was snap frozen in liquid nitrogen and stored at -80°C until further use. cfDNA was extracted from 800μl plasma using the QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany) with minor modifications as described before [6]. Extracted cfDNA was eluted in 30 μl of AE elution buffer. The eluate was re-applied onto the column, and the final eluate was collected and stored at -20°C. Blood samples from recurrent and non-recurrent patients were extracted together to avoid batch effects.
Measurement of cfDI and cfDNA concentration
Concentration and integrity of circulating DNA in blood plasma were analyzed by measuring the abundances of short and long fragments from two repetitive DNA elements, ALU (ALU-111bp, ALU-260bp) and LINE1 ( LINE1-97bp, LINE1-266bp) as described before [6]. All primer sequences and amplicon lengths are given in Supplementary Table 2. The fragment concentrations were measured in triplicates by quantitative PCR using ABsolute qPCR SYBR Green Mix (Thermo Scientific, Carlsbad, USA) and the LightCycler480 system (Roche Diagnostics, Mannheim, Germany). The cfDNA eluate was diluted 1:20 before use to achieve optimal PCR efficiency. Concentrations of the long and short fragments were calculated using the absolute quantification method according to the Light Cylcer 480 software instructions. cfDI was calculated as the ratio of long divided by short fragments concentrations for each of the elements: ALU-260/111, LINE1-266/97 as described before [6]. As short fragments were nested within the long fragments, cfDI ranged from 0 to 1. Short fragment concentrations were regarded as overall cfDNA concentrations.
Statistical analysis
All statistical analyses were carried out using the PASW Statistics 18.0 (SPSS, Chicago, IL, USA) package. cfDNA concentrations are not normally distributed and thus were log2-transformed for all data analysis. Differences of cfDNA concentrations and cfDI between the sample groups were evaluated with the Mann-Whitney U test. Influences of clinical parameters on cfDI and cfDNA concentration were studied by Mann-Whitney U test (for categorical and binary data), Spearman correlation permutation tests (for quantitative and continuous data), Jonckheere-Terpstra tests (for ordinal data) and Kruskal-Wallis H tests (for dependent ordinal data). Receiver operating characteristic (ROC) analysis was carried out to assess the discriminatory power of cfDI and cfDNA concentration between non-recurrent and recurrent groups and the corresponding area under the curve (AUC) was calculated. Univariate logistic regression was used to compare different variables between the sample groups. Multivariate logistic regression analyses based on all women grouped by recurrence status were performed to estimate the odds ratio and 95% confidence interval (CI), adjusting for known variables associated with recurrence, such as tumor size, lymph node status, histological grade as well as for significant variables in the univariate analysis. Interquartile analysis of cfDI and recurrence of breast cancer was conducted by logistic regression, with the highest cfDI quartile taken as the reference. P-values less than 0.05 were considered as statistically significant.
CONFLICTS OF INTEREST
All authors disclose no conflicts.
REFERENCES
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015; 65:5–29.
2. Brewster AM, Hortobagyi GN, Broglio KR, Kau SW, Santa-Maria CA, Arun B, Buzdar AU, Booser DJ, Valero V, Bondy M, Esteva FJ. Residual risk of breast cancer recurrence 5 years after adjuvant therapy. J Natl Cancer Inst. 2008; 100:1179–83.
3. Darby S, McGale P, Correa C, Taylor C, Arriagada R, Clarke M, Cutter D, Davies C, Ewertz M, Godwin J, Gray R, Pierce L, Whelan T, et al, and Early Breast Cancer Trialists’ Collaborative Group (EBCTCG). Effect of radiotherapy after breast-conserving surgery on 10-year recurrence and 15-year breast cancer death: meta-analysis of individual patient data for 10,801 women in 17 randomised trials. Lancet. 2011; 378:1707–16.
4. Giuliano AE, Hunt KK, Ballman KV, Beitsch PD, Whitworth PW, Blumencranz PW, Leitch AM, Saha S, McCall LM, Morrow M. Axillary dissection vs no axillary dissection in women with invasive breast cancer and sentinel node metastasis: a randomized clinical trial. JAMA. 2011; 305:569–75.
5. Cheng L, Swartz MD, Zhao H, Kapadia AS, Lai D, Rowan PJ, Buchholz TA, Giordano SH. Hazard of Recurrence among Women after Primary Breast Cancer Treatment-A 10-Year Follow-Up Using Data from SEER-Medicare (vol 21, pg 800, 2012). Cancer Epidem Biomar. 2012; 21:1604–05.
6. Madhavan D, Wallwiener M, Bents K, Zucknick M, Nees J, Schott S, Cuk K, Riethdorf S, Trumpp A, Pantel K, Sohn C, Schneeweiss A, Surowy H, Burwinkel B. Plasma DNA integrity as a biomarker for primary and metastatic breast cancer and potential marker for early diagnosis. Breast Cancer Res Treat. 2014; 146:163–74.
7. Lee Y, Kang E, Lee AS, Baek H, Kim EK, Park SY, Kim JH, Kim YJ, Kim SH, Kim IA, Eom KY, Kim SW. Outcomes and recurrence patterns according to breast cancer subtypes in Korean women. Breast Cancer Res Treat. 2015; 151:183–90.
8. de Boer M, van Dijck JA, Bult P, Borm GF, Tjan-Heijnen VC. Breast cancer prognosis and occult lymph node metastases, isolated tumor cells, and micrometastases. J Natl Cancer Inst. 2010; 102:410–25.
9. Montagna E, Bagnardi V, Rotmensz N, Viale G, Renne G, Cancello G, Balduzzi A, Scarano E, Veronesi P, Luini A, Zurrida S, Monti S, Mastropasqua MG, et al. Breast cancer subtypes and outcome after local and regional relapse. Ann Oncol. 2012; 23:324-331. https://doi.org/10.1093/annonc/mdr129.
10. Harris EE, Hwang WT, Seyednejad F, Solin LJ. Prognosis after regional lymph node recurrence in patients with stage I-II breast carcinoma treated with breast conservation therapy. Cancer. 2003; 98:2144–51.
11. Ahmad A. Pathways to breast cancer recurrence. ISRN Oncol. 2013; 2013:290568. https://doi.org/10.1155/2013/290568.
12. Sestak I, Cuzick J. Markers for the identification of late breast cancer recurrence. Breast Cancer Res. 2015; 17:10.
13. Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004; 351:2817–26.
14. Weigelt B, Peterse JL, van ’t Veer LJ. Breast cancer metastasis: markers and models. Nat Rev Cancer. 2005; 5:591–602.
15. Lyman GH, Giuliano AE, Somerfield MR, Benson AB 3rd, Bodurka DC, Burstein HJ, Cochran AJ, Cody HS 3rd, Edge SB, Galper S, Hayman JA, Kim TY, Perkins CL, et al, and American Society of Clinical Oncology. American Society of Clinical Oncology guideline recommendations for sentinel lymph node biopsy in early-stage breast cancer. J Clin Oncol. 2005; 23:7703–20.
16. Criscitiello C, André F, Thompson AM, De Laurentiis M, Esposito A, Gelao L, Fumagalli L, Locatelli M, Minchella I, Orsi F, Goldhirsch A, Curigliano G. Biopsy confirmation of metastatic sites in breast cancer patients: clinical impact and future perspectives. Breast Cancer Res. 2014; 16:205.
17. Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011; 11:426–37.
18. Molloy TJ, Devriese LA, Helgason HH, Bosma AJ, Hauptmann M, Voest EE, Schellens JH, van’t Veer LJ. A multimarker QPCR-based platform for the detection of circulating tumour cells in patients with early-stage breast cancer. Br J Cancer. 2011; 104:1913–19.
19. Roth C, Rack B, Müller V, Janni W, Pantel K, Schwarzenbach H. Circulating microRNAs as blood-based markers for patients with primary and metastatic breast cancer. Breast Cancer Res. 2010; 12:R90.
20. Cuk K, Zucknick M, Heil J, Madhavan D, Schott S, Turchinovich A, Arlt D, Rath M, Sohn C, Benner A, Junkermann H, Schneeweiss A, Burwinkel B. Circulating microRNAs in plasma as early detection markers for breast cancer. Int J Cancer. 2013; 132:1602–12.
21. Madhavan D, Cuk K, Burwinkel B, Yang R. Cancer diagnosis and prognosis decoded by blood-based circulating microRNA signatures. Front Genet. 2013; 4:116.
22. Cuk K, Zucknick M, Madhavan D, Schott S, Golatta M, Heil J, Marmé F, Turchinovich A, Sinn P, Sohn C, Junkermann H, Schneeweiss A, Burwinkel B. Plasma microRNA panel for minimally invasive detection of breast cancer. PLoS One. 2013; 8:e76729.
23. Madhavan D, Peng C, Wallwiener M, Zucknick M, Nees J, Schott S, Rudolph A, Riethdorf S, Trumpp A, Pantel K, Sohn C, Chang-Claude J, Schneeweiss A, Burwinkel B. Circulating miRNAs with prognostic value in metastatic breast cancer and for early detection of metastasis. Carcinogenesis. 2016; 37:461–70.
24. Madhavan D, Zucknick M, Wallwiener M, Cuk K, Modugno C, Scharpff M, Schott S, Heil J, Turchinovich A, Yang R, Benner A, Riethdorf S, Trumpp A, et al. Circulating miRNAs as surrogate markers for circulating tumor cells and prognostic markers in metastatic breast cancer. Clin Cancer. 2012; 18:5972-5982. https://doi.org/10.1158/1078-0432.CCR-12-1407.
25. Diaz LA Jr, Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014; 32:579–86.
26. Mandel P, Metais P. Les acides nucléiques du plasma sanguin chez l’homme. C R Seances Soc Biol Fil. 1948; 142:241–43.
27. Wang BG, Huang HY, Chen YC, Bristow RE, Kassauei K, Cheng CC, Roden R, Sokoll LJ, Chan DW, Shih IM. Increased plasma DNA integrity in cancer patients. Cancer Res. 2003; 63:3966–68.
28. Jiang WW, Zahurak M, Goldenberg D, Milman Y, Park HL, Westra WH, Koch W, Sidransky D, Califano J. Increased plasma DNA integrity index in head and neck cancer patients. Int J Cancer. 2006; 119:2673–76.
29. Kamel AM, Teama S, Fawzy A, El Deftar M. Plasma DNA integrity index as a potential molecular diagnostic marker for breast cancer. Tumour Biol. 2016; 37:7565–72.
30. Hauser S, Zahalka T, Ellinger J, Fechner G, Heukamp LC, VON Ruecker A, Müller SC, Bastian PJ. Cell-free circulating DNA: diagnostic value in patients with renal cell cancer. Anticancer Res. 2010; 30:2785–89.
31. Gao YJ, He YJ, Yang ZL, Shao HY, Zuo Y, Bai Y, Chen H, Chen XC, Qin FX, Tan S, Wang J, Wang L, Zhang L. Increased integrity of circulating cell-free DNA in plasma of patients with acute leukemia. Clin Chem Lab Med. 2010; 48:1651-1656. https://doi.org/10.1515/CCLM.2010.311.
32. Voduc KD, Cheang MC, Tyldesley S, Gelmon K, Nielsen TO, Kennecke H. Breast cancer subtypes and the risk of local and regional relapse. J Clin Oncol. 2010; 28:1684–91.
33. Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, Dunning MJ, Gale D, Forshew T, Mahler-Araujo B, Rajan S, Humphray S, Becq J, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013; 368:1199–209.
34. Gasparini P, Cascione L, Fassan M, Lovat F, Guler G, Balci S, Irkkan C, Morrison C, Croce CM, Shapiro CL, Huebner K. microRNA expression profiling identifies a four microRNA signature as a novel diagnostic and prognostic biomarker in triple negative breast cancers. Oncotarget. 2014; 5:1174-84. https://doi.org/10.18632/oncotarget.1682.
35. Chang K, Kong YY, Dai B, Ye DW, Qu YY, Wang Y, Jia ZW, Li GX. Combination of circulating tumor cell enumeration and tumor marker detection in predicting prognosis and treatment effect in metastatic castration-resistant prostate cancer. Oncotarget. 2015; 6:41825-36. https://doi.org/10.18632/oncotarget.6167.
36. Brunetti O, Russo A, Scarpa A, Santini D, Reni M, Bittoni A, Azzariti A, Aprile G, Delcuratolo S, Signorile M, Gnoni A, Palermo L, Lorusso V, et al. MicroRNA in pancreatic adenocarcinoma: predictive/prognostic biomarkers or therapeutic targets? Oncotarget. 2015; 6:23323-41. https://doi.org/10.18632/oncotarget.4492.
37. Mead R, Duku M, Bhandari P, Cree IA. Circulating tumour markers can define patients with normal colons, benign polyps, and cancers. Br J Cancer. 2011; 105:239–45.
38. Mouliere F, Robert B, Arnau Peyrotte E, Del Rio M, Ychou M, Molina F, Gongora C, Thierry AR. High fragmentation characterizes tumour-derived circulating DNA. PLoS One. 2011; 6:e23418.
39. Ellinger J, Wittkamp V, Albers P, Perabo FG, Mueller SC, von Ruecker A, Bastian PJ. Cell-free circulating DNA: diagnostic value in patients with testicular germ cell cancer. J Urol. 2009; 181:363–71.
40. Yu J, Gu G, Ju S. Recent advances in clinical applications of circulating cell-free DNA integrity. Lab Med. 2014; 45:6–11.
41. Umetani N, Giuliano AE, Hiramatsu SH, Amersi F, Nakagawa T, Martino S, Hoon DS. Prediction of breast tumor progression by integrity of free circulating DNA in serum. J Clin Oncol. 2006; 24:4270–76.
42. Giacona MB, Ruben GC, Iczkowski KA, Roos TB, Porter DM, Sorenson GD. Cell-free DNA in human blood plasma: length measurements in patients with pancreatic cancer and healthy controls. Pancreas. 1998; 17:89–97.
43. Jiang P, Chan CW, Chan KC, Cheng SH, Wong J, Wong VW, Wong GL, Chan SL, Mok TS, Chan HL, Lai PB, Chiu RW, Lo YM. Lengthening and shortening of plasma DNA in hepatocellular carcinoma patients. Proc Natl Acad Sci USA. 2015; 112:E1317–25.
44. Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, Knippers R. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001; 61:1659–65.
45. Holdenrieder S, Burges A, Reich O, Spelsberg FW, Stieber P. DNA integrity in plasma and serum of patients with malignant and benign diseases. Ann N Y Acad Sci. 2008; 1137:162–70.
46. Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, et al, and International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature. 2001; 409:860–921.
47. Perkins G, Yap TA, Pope L, Cassidy AM, Dukes JP, Riisnaes R, Massard C, Cassier PA, Miranda S, Clark J, Denholm KA, Thway K, Gonzalez De Castro D, et al. Multi-purpose utility of circulating plasma DNA testing in patients with advanced cancers. PLoS One. 2012; 7:e47020.
48. Iqbal S, Vishnubhatla S, Raina V, Sharma S, Gogia A, Deo SS, Mathur S, Shukla NK. Circulating cell-free DNA and its integrity as a prognostic marker for breast cancer. Springerplus. 2015; 4:265.
49. Wang K, Yuan Y, Cho JH, McClarty S, Baxter D, Galas DJ. Comparing the MicroRNA spectrum between serum and plasma. PLoS One. 2012; 7:e41561.
50. El Messaoudi S, Rolet F, Mouliere F, Thierry AR. Circulating cell free DNA: Preanalytical considerations. Clin Chim Acta. 2013; 424:222-230.
51. Devonshire AS, Whale AS, Gutteridge A, Jones G, Cowen S, Foy CA, Huggett JF. Towards standardisation of cell-free DNA measurement in plasma: controls for extraction efficiency, fragment size bias and quantification. Anal Bioanal Chem. 2014; 406:6499–512.

