INTRODUCTION
Well-established poor prognostic factors in chronic lymphocytic leukemia (CLL) include deletions in 11q, which involve ATM locus (delATM), and losses affecting TP53 gene located in 17p13 (delTP53) detected by fluorescence in situ hybridization (FISH) [1]. While the adverse prognosis of delATM has been partially overcome by chemoimmunotherapy-based treatments, patients showing TP53 deletions or mutations are typically resistant to these treatments [2, 3]. New agents acting independently of the p53 pathway have changed the treatment response rate for CLL patients with TP53 abnormalities [4, 5]. However, recent studies have revealed that current FISH analyses (using Dohner’s hierarchical model probes) underestimate the true genomic complexity revealed by chromosome banding analysis (CBA) which predicts a worse clinical outcome in patients treated either with standard treatments or new agents [6-11]. The aims of the present study were: (i) to describe and compare the characteristics and clinical course of CLL patients according to karyotype complexity detected by CBA, (ii) to analyze the impact of complex karyotype (CK) and high-risk FISH deletions (ATM and/or TP53, HR-FISH) on time to the first therapy and overall survival.
RESULTS AND DISCUSSION
In the whole series, 435/1043 (41.6%) patients displayed abnormal karyotypes, being 99 (9.5%) complex. While most patients with abnormal but non-CK carried a single aberration (267/336, 80%), cases with CK displayed a median of 4 abnormalities (range: 3-18). Of note, structural aberrations were slightly predominant in both groups (63% and 57% in CK and non-CK, respectively). Notably, abnormal karyotypes from non-CK were mainly restricted to known recurrent CLL aberrations, namely trisomy 12 (37%), 13q14 structural anomalies (17% deletions and 4% translocations), del(11q) (12%), and translocations involving 14q (6%). On the contrary, CK patients displayed a wide variety of abnormalities distributed along the genome, which reflected the genomic instability of this group. Among them, the most frequent were 17p structural rearrangements (deletions, unbalanced translocations or isochromosome 17q) leading to TP53 loss (26%), trisomy 12 (26%), del(13q) (19%), del(11q) (16%), trisomy 18 (11%) and del(6q) (10%). In addition, clonal evolution by CBA was identified in 33/99 (33%) patients with CK. Detailed karyotypes from those patients from CK group are displayed in Supplementary Table 3.
The CK group included patients with more advanced disease at diagnosis, including a higher proportion of patients diagnosed at Binet stage B/C (22% vs 11%; P = 0.001) and higher β2-microglobulin levels (2.6 mg/L vs 2 mg/L; P < 0.001). Moreover, features of aggressive disease were also observed: higher CD38 (33% vs 20%; P = 0.007) and ZAP-70 (46% vs 30%; P = 0.018) positivity rates, delATM (25% vs 7%; P < 0.001) and delTP53 (40% vs 5%; P < 0.001). Despite the limited availability of IGHV data in this retrospective cohort (N = 98/1045), CK group also displayed a significant higher proportion of unmutated IGHV genes [71% (15/21) vs 23% (18/77); P < 0.001]. Indeed, CK group showed a significant higher two-year cumulative incidence of treatment (48%, 95%CI 36-58%) than non-CK (20%, 95%CI 18-23%; P < 0.001), as well as a shorter overall survival (OS) (79 mo vs not reached; P < 0.001). The delTP53 enrichment observed in CK group was in accordance with the previously described frequencies (ranging from 28-50%) and could be underlying genomic instability leading to CK [6, 9, 12–13]. However, only 44.4% (40/90) patients with delTP53 showed a CK. CK negatively impacted on OS even within delTP53 cases (38 mo vs 133 mo; P < 0.001). In this regard, when patients were categorized regarding both CK and HR-FISH, no relevant differences in clinical characteristics at diagnosis were observed among those patients with at least one high-risk cytogenetic marker (CK and/or HR-FISH) (Table 1). Nevertheless, patients could be stratified in terms of time to first treatment (Figure 1). Of note, while patients with both HR-FISH and CK showed the worst median OS (52 mo) being clearly distinct from those non-CK and non-HR-FISH (median not reached), no significant differences were detected between cases with only CK or HR-FISH (Figure 1). Similarly, if only CK and delTP53 were considered, no differences in OS for CK patients lacking delTP53 and those with delTP53 but non-CK were observed (132 mo vs 89 mo; P = 0.269). In the multivariate analysis for OS, both CK and delTP53 remained statistically significant. Of note, delTP53 was the predictor showing higher hazard ratio in the multivariate analysis (Table 2). Unfortunately, as this is a retrospective and multicentric study, TP53 mutational status was not routinely assessed and availability of DNA samples was limited. Despite this limitation, TP53 mutations could be analyzed in 8/42 cases from CK and non-HR-FISH subgroup and all of them were negative. In accordance, although it has been described that nearly 20% of patients with TP53 abnormalities show mutations in absence of 17p deletion [14, 15], this percentage is much lower in general untreated CLL cohorts (4.5%) [16]. In this regard, despite the limited availability of information regarding TP53 mutational status, only a minority of CK and non-HR-FISH patients could harbor TP53 mutations and therefore could be misclassified. So that, it could be assumed that other factors than abnormal TP53 function are underlying the global poor outcome observed in this subgroup. On the other hand, despite the detection of unmutated IGHV was significantly associated with shorter OS in the global cohort (55 mo vs 133 mo; P < 0.001), the limited availability of IGHV data did not allow the stratification by both IGHV and CK in the OS assessment.
Table 1: Baseline characteristics of patients with unfavourable cytogenetic markers (CK and/or HR-FISH) at diagnosis
Patients characteristics |
CK and non-HR-FISH (N = 42) |
non-CK and HR-FISH (N = 113) |
CK and HR-FISH (N = 57) |
P-value |
|---|---|---|---|---|
Median age at diagnosis (range) |
73 (46–87) |
67 (27–90) |
69 (45–88) |
0.322 |
Male |
31 (73.9%) |
78 (69.0%) |
35 (61.4%) |
0.284 |
Diagnosis |
||||
MBL |
10 (23.8%) |
13 (11.5) |
4 (7.0%) |
0.039 |
CLL |
32 (76.2%) |
100 (88.5%) |
53 (93.0%) |
|
Advanced Binet stage * |
9/42 (21.4%) |
33/109 (30.3%) |
11/56 (19.6%) |
0.262 |
Lymphadenopathy |
16/42 (38.1%) |
56/110 (50.1%) |
28/55 (50.9%) |
0.323 |
Splenomegaly |
3/40 (7.5%) |
15/109 (13.8%) |
8/53 (15.1%) |
0.512 |
Hepatomegaly |
2/42 (4.8%) |
11/112 (9.8%) |
2/56 (3.6%) |
0.266 |
Absolute white blood cell count (x109/L) |
16 (6–164) |
17 (4–168) |
21 (5–89) |
0.122 |
Absolut lymphocyte count (x109/L) |
11 (2–105) |
11 (1–155) |
14 (2–82) |
0.091 |
Hemoglobin (g/dL) |
14 (9–17) |
14 (6–18) |
14 (8–17) |
0.219 |
Platelets (x109/L) |
190 (42–473) |
191 (10–413) |
173 (34–331) |
0.470 |
Lactate dehydrogenase (IU/L) |
332 (160–1279) |
340 (139–1094) |
383 (156–1082) |
0.237 |
Beta-2 Microglobulin (mg/L) |
2.4 (1.0–10.0) |
2.2 (1.0–13.4) |
2.6 (1.1–8.4) |
0.305 |
ZAP-70 positive** |
7/20 (35.0%) |
17/62 (27.4%) |
16/31 (51.6%) |
0.071 |
CD38 positive* |
11/31 (35.5%) |
33/96 (34.4%) |
14/42 (33.3%) |
0.982 |
Unmutated IGHV |
1/5 (20.0%) |
2/2 (100%) |
13/14 (92.9%) |
0.003 |
FISH |
||||
13q deletion |
16/42 (38.1%) |
54/110 (49.1%) |
34/55 (61.8%) |
0.064 |
Trisomy 12 |
24/42 (57.1%) |
15/109 (13.8%) |
7/54 (13.02%) |
< 0.001 |
11q deletion (ATM) |
– |
68/113 (60.2%) |
25/57 (43.9%) |
N.A. |
17p deletion (TP53) |
– |
50/113 (44.2%) |
40/57 (70.2%) |
N.A. |
Values are given as median (range) or number (%). Hemoglobin is expressed as mean (range). N.A. Not applicable.
*Binet stage was not available in five patients (information regarding physical examination and/or analytical parameters not provided).
**Positivity was considered when ZAP-70 > 20% and CD38 > 30%.
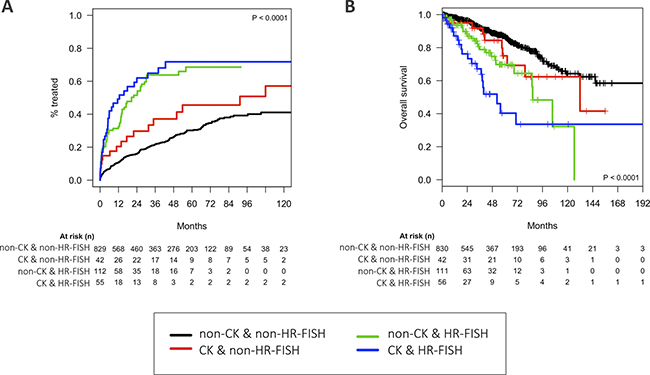
Figure 1: (A) Cumulative incidence of treatment (TFT) for the presence of CK and/or HR-FISH and (B) Kaplan–Meier plot for overall survival (OS) and the presence of CK and/or HR-FISH.
Table 2: Univariate and multivariate analysis on median overall survival (OS)
Variable |
Univariate |
Multivariate |
|||
|---|---|---|---|---|---|
Median OS in months (95% CI) |
P-value |
Hazard ratio (95% CI) |
P-value |
||
Complex karyotype (CK) |
Positive |
79 (23–135) |
< 0.001 |
1.66 (1.06–2.59) |
0.027 |
Negative |
NR |
||||
ATM deletion |
Positive |
87 (64–108) |
0.003 |
1.51 (0.93–2.43) |
0.093 |
Negative |
NR |
||||
TP53 deletion |
Positive |
56 (34–77) |
< 0.001 |
3.03 (1.93–4.77) |
< 0.001 |
Negative |
197 (107–287) |
||||
NR: Not reached.
Mounting evidence suggests a relevant role of CBA in risk assessment of CLL patients. Indeed, Rigolin et al. demonstrated that 25-37% of patients showing no aberrations by FISH carried chromosomal abnormalities not covered by the standard FISH panel, which strongly correlated with worse prognosis [17]. Moreover, Jaglowski et al. described that both the number of karyotypic abnormalities as a continuous variable and the CK detection (defined as ≥ 5 abnormalities) could predict a worse clinical outcome in patients undergoing reduced-intensity conditioning allogeneic stem cell transplantation; remarkably, both variables retained their predictive prognostic value even within patients with HR-FISH [6]. In this regard, a recent study described the impact of CK as an independent unfavorable prognostic factor in a prospective trial using chlorambucil-based regimens in CLL patients with relevant comorbidity. In accordance to our results, an important proportion of patients with CK lacking TP53 abnormalities (73% cases with CK) showed a dismal survival, equivalent to those patients with sole TP53 lesions [11]. Novel evidences suggesting the value of CBA have arisen with the recent development of new treatment agents in CLL. Initially, Thompson et al. demonstrated that CK showed stronger impact on outcome than delTP53 in CLL patients treated with ibrutinib [10]. Furthermore, this negative impact on the outcome of ibrutinib-treated patients has been subsequently confirmed after a longer 5-year follow-up [18], as well as in a large “real-world” multicentric analysis of patients treated with novel agents in CLL [19]. It is noteworthy that a lack of any adverse prognostic effect of CK on idelalisib-treated have been recently reported [20]. However, this finding should be confirmed in a larger cohort of CLL undergoing idelalisib-based regimens. All these findings point out CK as a potent prognostic factor in the new therapies era, especially considering that ibrutinib treatment increased survival in treatment-naïve older CLL patients without TP53 deletion irrespectively of other historical poor prognostic factors, such as delATM or unmutated IGHV [5]. Nevertheless, patients with CK are indeed a heterogeneous group: patients harboring multiple trisomies, namely +12+18+19, and non-HR-FISH show a particularly benign clinical evolution [21]. Herein, we detected nine such cases presenting a better outcome compared with the remaining CK cases (data not shown). This observation suggests that criteria for CK definition in CLL should be revised; and that cytogenetic complexity defined by solely numerical aberrations should not be unquestionably considered as an unfavorable prognostic marker in this entity [21]. Even though previous publications evaluate cytogenetic complexity by the number of abnormalities detected by CBA, as well as the number or total size of altered regions by genomic microarrays, standard criteria focusing on both the number and the nature of the abnormalities are highly needed to provide a more accurate prognostic marker in CLL.
In conclusion, we report the existence of a subgroup of CLL patients with CK lacking HR-FISH abnormalities (delATM/delTP53) that show an equivalent impaired clinical evolution as those with HR-FISH and non-CK. Despite CK group is globally associated with advanced disease and poor prognostic markers, genetic basis other than TP53/ATM dysfunction underlying the heterogeneity of CK remain to be elucidated. In order to better determine the patients’ prognosis according to genomic complexity detected by CBA, and make possible to select tailored therapies, further investigation in larger patient cohorts with CBA data is required.
MATERIALS AND METHODS
Patient selection
We selected 1045 CLL and monoclonal B-cell lymphocytosis (MBL) patients (821 and 224, respectively) with available simultaneous karyotype and FISH at diagnosis or prior to therapy from the CLL/MBL database of the Grupo Cooperativo Español de Citogenética Hematológica (GCECGH) and Grupo Español de Leucemia Linfática Crónica (GELLC). Of note, MBL individuals were also included, as MBL represents an initial phase in the disease evolution, some of them harbor poor prognosis cytogenetics and could evolve to CLL requiring treatment during follow-up. The median time from diagnosis to CBA was 0.5 months (range, 0-229). The study was approved by the ethical committee of Hospital del Mar, Barcelona (ref. 2016/6861/I).
Clinical information collected at diagnosis included demographics (age and gender), Binet stage, physical examination, and analytical information regarding absolute white blood cell and lymphocyte counts, platelet count, hemoglobin level, as well as lactate dehydrogenase (LDH) and serum beta2-microglobulin (B2M) concentrations. When available, CD38 and ZAP-70 expression and mutational status of immunoglobulin heavy chain genes (IGHV) were also compiled. Evolutive data on treatment administration and last follow-up were recorded (Supplementary Table 1). Treatment administration and response evaluation were performed according to current guidelines [22–24]. At the end of the study, 30.2% of patients had required therapy. The most frequently used therapies in first-line treatments are detailed in Supplementary Table 2. Of note, only four of the included patients received novel agents acting independently of TP53 pathway (ibrutinib) during follow-up.
Chromosome banding analyses and fluorescence in situ hybridization
Chromosome banding analyses were mainly assessed on peripheral blood samples (n = 881, 84%), whereas bone marrow and lymph node samples were only analyzed in 159 and five cases, respectively. Cell cultures were set for 72 hours with TPA (12-O-tetradecanoyl-phorbol-13-acetate) according to standard procedures. A minimum of 20 metaphases were analyzed when possible. Karyotypes were described following the International System for Human Cytogenetic Nomenclature (2013) [25]. Only those abnormalities detected in at least two metaphases or those detected in a single metaphase but further confirmed by FISH or in a subsequent CBA were considered. CK was defined as the presence of three or more chromosomal abnormalities in a single clone. Data from routine FISH panel for TP53 (17p13), ATM (11q22), D13S319 (13q14) and chromosome 12 centromeric region were obtained simultaneously to CBA.
Statistical analysis
Firstly, patients were classified according to the presence of CK independently of FISH results (CK vs non-CK). Discrete variables were compared between groups by Chi-square or Fisher exact tests, while Mann-Whitney test was used for continuous variables. Time to first treatment (TFT) was defined as time from sampling to start of treatment or last follow-up; overall survival (OS) was defined as time from sampling to death or last follow-up. Both TFT and OS were assessed using cumulative incidence and Kaplan–Meier plots, respectively and the effect of different covariates was evaluated using the Gray’s and log-rank test, respectively. The maintenance of the independent predictive value on OS was assessed in multivariate analyses using Cox proportional hazards regression models. To further clarify the impact of high-risk FISH deletions (ATM and/or TP53, HR-FISH) in CK prognosis, a second classification including CK and HR-FISH was performed. TFT and OS from four groups were compared: (1) CK and HR-FISH; (2) CK and non-HR-FISH; (3) non-CK and HR-FISH; and (4) non-CK and non-HR-FISH. Statistical analyses were performed using SPSS v.22 software (SPSS Inc, Chicago, IL, USA) and R v.3.2.2. Results were considered statistically significant for P-values < 0.05.
Author contributions
AP and BE designed the research study, provided patients data, analyzed the data and wrote the manuscript. JD provided patients information, performed data analysis and critical discussion of the results. RC, MJC, MO, NRX, ARD, EL, TG, BN, MDGM, AV, JAH, MTA, MAP, MLB, MHS, RS, AB, MS, AF, PA, EG, EA, CF, MJT, FJO, DC, CM , FC and FB provided patient data. All authors read the last version of the manuscript.
ACKNOWLEDGMENTS
This work has been supported by the following grants: PI11/01621, PI15/00437, RD12/0036/0044, RD12/0036/0069 and PT13/0010/0005 FEDER, Instituto de Salud Carlos III, Spanish Ministry of Economy and Competitiveness; 2014/SGR585 from Generalitat de Catalunya.
CONFLICTS OF INTEREST
The authors made no disclosure of conflicts of interest.
REFERENCES
1. Döhner H, Stilgenbauer S, Benner A, Leupolt E, Kröber A, Bullinger L, Döhner K, Bentz M, Lichter P. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med. 2000; 343:1910–16.
2. Tsimberidou AM, Tam C, Abruzzo LV, O’Brien S, Wierda WG, Lerner S, Kantarjian HM, Keating MJ. Chemoimmunotherapy may overcome the adverse prognostic significance of 11q deletion in previously untreated patients with chronic lymphocytic leukemia. Cancer. 2009; 115:373–80.
3. Stilgenbauer S, Schnaiter A, Paschka P, Zenz T, Rossi M, Döhner K, Bühler A, Böttcher S, Ritgen M, Kneba M, Winkler D, Tausch E, Hoth P, et al. Gene mutations and treatment outcome in chronic lymphocytic leukemia: results from the CLL8 trial. Blood. 2014; 123:3247–54.
4. Furman RR, Sharman JP, Coutre SE, Cheson BD, Pagel JM, Hillmen P, Barrientos JC, Zelenetz AD, Kipps TJ, Flinn I, Ghia P, Eradat H, Ervin T, et al. Idelalisib and rituximab in relapsed chronic lymphocytic leukemia. N Engl J Med. 2014; 370:997–1007.
5. Burger JA, Tedeschi A, Barr PM, Robak T, Owen C, Ghia P, Bairey O, Hillmen P, Bartlett NL, Li J, Simpson D, Grosicki S, Devereux S, et al, and RESONATE-2 Investigators. Ibrutinib as initial therapy for patients with chronic lymphocytic leukemia. N Engl J Med. 2015; 373:2425–37.
6. Jaglowski SM, Ruppert AS, Heerema NA, Bingman A, Flynn JM, Grever MR, Jones JA, Elder P, Devine SM, Byrd JC, Andritsos LA. Complex karyotype predicts for inferior outcomes following reduced-intensity conditioning allogeneic transplant for chronic lymphocytic leukaemia. Br J Haematol. 2012; 159:82–87.
7. Woyach JA, Lozanski G, Ruppert AS, Lozanski A, Blum KA, Jones JA, Flynn JM, Johnson AJ, Grever MR, Heerema NA, Byrd JC. Outcome of patients with relapsed or refractory chronic lymphocytic leukemia treated with flavopiridol: impact of genetic features. Leukemia. 2012; 26:1442–44.
8. Travella A, Ripollés L, Aventin A, Rodríguez A, Bezares RF, Caballín MR, Slavutsky I. Structural alterations in chronic lymphocytic leukaemia. Cytogenetic and FISH analysis. Hematol Oncol. 2013; 31:79–87.
9. Baliakas P, Iskas M, Gardiner A, Davis Z, Plevova K, Nguyen-Khac F, Malcikova J, Anagnostopoulos A, Glide S, Mould S, Stepanovska K, Brejcha M, Belessi C, et al. Chromosomal translocations and karyotype complexity in chronic lymphocytic leukemia: a systematic reappraisal of classic cytogenetic data. Am J Hematol. 2014; 89:249–55.
10. Thompson PA, O’Brien SM, Wierda WG, Ferrajoli A, Stingo F, Smith SC, Burger JA, Estrov Z, Jain N, Kantarjian HM, Keating MJ. Complex Karyotype is a Stronger Predictor Than Del(17p) for an Inferior Outcome in Relapsed or Refractory Chronic Lymphocytic Leukemia Patients Treated With Ibrutinib-Based Regimens. Cancer. 2016; 122:2505–11.
11. Herling CD, Klaumünzer M, Rocha CK, Altmüller J, Thiele H, Bahlo J, Kluth S, Crispatzu G, Herling M, Schiller J, Engelke A, Tausch E, Döhner H, et al. Complex karyotypes and KRAS and POT1 mutations impact outcome in CLL after chlorambucil-based chemotherapy or chemoimmunotherapy. Blood. 2016; 128:395–404.
12. Haferlach C, Dicker F, Schnittger S, Kern W, Haferlach T. Comprehensive genetic characterization of CLL: a study on 506 cases analysed with chromosome banding analysis, interphase FISH, IgV(H) status and immunophenotyping. Leukemia. 2007; 21:2442–51.
13. Yu L, Kim HT, Kasar SN, Benien P, Du W, Hoang K, Aw A, Tesar B, Improgo R, Fernandes SM, Radhakrishnan S, Klitgaard JL, Lee C, et al. Survival of Del17p CLL Depends on Genomic Complexity and Somatic Mutation. Clin Cancer Res. 2016. [Epub ahead of print].
14. Rossi D, Cerri M, Deambrogi C, Sozzi E, Cresta S, Rasi S, De Paoli L, Spina V, Gattei V, Capello D, Forconi F, Lauria F, Gaidano G. The prognostic value of TP53 mutations in chronic lymphocytic leukemia is independent of Del17p13: implications for overall survival and chemorefractoriness. Clin Cancer Res. 2009; 15:995–1004.
15. Pospisilova S, Sutton LA, Malcikova J, Tausch E, Rossi D, Montserrat E, Moreno C, Stamatopoulos K, Gaidano G, Rosenquist R, Ghia P, European Research Initiative on CLL (ERIC). Innovation in the prognostication of chronic lymphocytic leukemia: how far beyond TP53 gene analysis can we go? Haematologica. 2016; 101:263–65.
16. Zenz T, Kröber A, Scherer K, Häbe S, Bühler A, Benner A, Denzel T, Winkler D, Edelmann J, Schwänen C, Döhner H, Stilgenbauer S. Monoallelic TP53 inactivation is associated with poor prognosis in chronic lymphocytic leukemia: results from a detailed genetic characterization with long-term follow-up. Blood. 2008; 112:3322–29.
17. Rigolin GM, Cibien F, Martinelli S, Formigaro L, Rizzotto L, Tammiso E, Saccenti E, Bardi A, Cavazzini F, Ciccone M, Nichele I, Pizzolo G, Zaja F, et al. Chromosome aberrations detected by conventional karyotyping using novel mitogens in chronic lymphocytic leukemia with “normal” FISH: correlations with clinicobiologic parameters. Blood. 2012; 119:2310–13.
18. O’Brien S, Furman R, Coutre S, Flinn IM, Burger JM, Blum K, Sharman J, Wierda W, Jones J, Zhao W, Heerema N, et al. Five-Year Experience with Single-Agent Ibrutinib in Patients with Previously Untreated and Relapsed/Refractory Chronic Lymphocytic. American Society of Hematology Anunal Meeting 2016. https://ash.confex.com/ash/2016/webprogram/Paper89757.html.
19. Mato AR, Hill BT, Lamanna N, Barr PM, Ujjani CS, Brander DM, Howlett C, Skarbnik AP, Cheson BD, Zent CS, Pu JJ, Kiselev P, Foon K, et al. Optimal Sequencing of Ibrutinib, Idelalisib, and Venetoclax in Chronic Lymphocytic Leukemia: Results from a Multi-Center Study of 683 Patients. Ann Oncol. 2017mdx031; Epub ahead of print.
20. Kreuzer KA, Furman R, Stilgenbauer S, Dubowy RL, Kim Y, Munugalavadla V, Lilienweiss E, Reinhardt H, Pettitt A, Hallek M. Outcome of Patients with Complex Karyotype in a Phase 3 Randomized Study of Idelalisib Plus Rituximab for Relapsed Chronic Lymphocytic Leukemia. Blood. 2016; 128:192.
21. Baliakas P, Puiggros A, Xochelli A, Sutton LA, Nguyen-Khac F, Gardiner A, Plevova K, Minga E, Hadzidimitriou A, Walewska R, McCarthy H, Ortega M, Collado R, et al. Additional trisomies amongst patients with chronic lymphocytic leukemia carrying trisomy 12: the accompanying chromosome makes a difference. Haematologica. 2016; 101:e299–302.
22. Cheson BD, Bennett JM, Rai KR, Grever MR, Kay NE, Schiffer CA, Oken MM, Keating MJ, Boldt DH, Kempin SJ, Foon KA. Guidelines for clinical protocols for chronic lymphocytic leukemia: recommendations of the National Cancer Institute-sponsored working group. Am J Hematol. 1988; 29:152–63.
23. Cheson BD, Bennett JM, Grever M, Kay N, Keating MJ, O’Brien S, Rai KR. National Cancer Institute-sponsored Working Group guidelines for chronic lymphocytic leukemia: revised guidelines for diagnosis and treatment. Blood. 1996; 87:4990–97.
24. Hallek M, Cheson BD, Catovsky D, Caligaris-Cappio F, Dighiero G, Döhner H, Hillmen P, Keating MJ, Montserrat E, Rai KR, Kipps TJ, International Workshop on Chronic Lymphocytic Leukemia. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia: a report from the International Workshop on Chronic Lymphocytic Leukemia updating the National Cancer Institute-Working Group 1996 guidelines. Blood. 2008; 111:5446–56.
25. Shaffer L, McGowan-Jordan J, Schmid M. 2013. ISCN 2013: An International System for Human Cytogenetic Nomenclature. Karger Medical and Scientific Publishers; 2013.

