INTRODUCTION
A steady decline of gastric carcinoma (GC) incidence has been observed worldwide, primarily as a result of a reduction in distal GC [1]. However, GC is the fourth most common malignancy and is a relatively higher incidence in Eastern Asian (e.g. China, Korea and Japan). Esophagogastric junction adenocarcinoma (EGJA) is one of the most rapidly increasing malignancies in North America and Europe and is thought to have different etiology compared to distal GC [2]. Recently, the increasing incidence of EGJA was also identified in Eastern Asian [3, 4]. EGJA is a highly fatal form of malignancies and is a major public health problem in China. The potential risk factors contributing to EGJA are foods preserved by salting, smoking and obesity et al. In addition, it is reported that individual’s genetic background also plays an important role in pathogenesis of EGJA. Although a number of studies have focused on the etiology of EGJA, it is not well understood. Of late, some studies reported the immune system might be implicated in the etiology of EGJA [5, 6].
Programmed cell death 1 (PD-1) gene was found by Ishida Y in 1992 [7]. It is classified as a member of the immunoglobulin superfamily (IgSF). As other inhibitory costimulatory molecules, PD-1 is expressed on many immunocytes, such as T cells, exhausted T cells, regulatory T cells (Treg), activated monocytes, B cells, natural killer (NK) cells, dendritic cells (DCs) and natural killer T (NKT) cells [8, 9]. PD-1 protein, a transmembrane glycoprotein, consists of an intracellular domain and an extracellular immunoglobulin V domain. PD-1 protein binds two ligands, programmed death ligand-1 (PD-L1) and programmed death ligand-2 (PD-L2). Under common physiological conditions, PD-1 interacts with PD-L1 and PD-L2, and then regulates a immune checkpoint. PD-1 may play a very important role in reducing the function of immune system by inhibiting T-cells and up-regulating Treg [10]. Finally, it decreases autoimmunity and results in self-tolerance.
Most recent studies reported several polymorphisms in PD-1 gene may be associated with susceptibility to some human autoimmune diseases (e.g., systemic lupus erythematosus, rheumatoid arthritis, type 1 diabetes mellitus, and ankylosing spondylitis et al.) [11–16]. Interestingly, accumulating evidences showed that PD-1 single nucleotide polymorphisms (SNPs) were also correlated with susceptibility to human malignancy (e.g., thyroid cancer, breast cancer, cervical cancer, non-small cell lung cancer and gastric cancer et al.) [17–21]. The immune response may be differ greatly among individual tumor hosts, and the potential mechanisms remain unknown. Genetic variations can influence the function of genes and alter the disease phenotypes. Thus, the effect of such functional polymorphisms in immune response genes on cancer risk has attracted our interest. Exploring the relationship of PD-1 SNPs with EGJA risk may be beneficial for providing prevention and personalized diagnosis. Here, we selected PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms and designed a hospital-based case-control study to determine the potential relationship between these functional SNPs in PD-1 gene and EGJA risk.
RESULTS
Baseline characteristics
The demographics (age and sex) and major risk factors (smoking and drinking status) of participants are summarized in Table 1. The mean ± SD of age was not significant in the EGJA patients compared with non-cancer controls (P > 0.05). Our study was well-matched by age and gender. Significant difference was observed on smoking status and alcohol consumption between the EGJA patients and the controls (P < 0.001). For PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms, the success rate of genotyping was more than 99%, respectively (Table 2). The minor allele frequency (MAF) in our controls was similar to the data of Chinese population. The distribution of genotype frequencies in controls accorded with Hardy–Weinberg equilibrium (HWE) (Table 2).
Table 1: Distribution of selected demographic variables and risk factors in EGJA cases and controls
Variable |
Overall Cases (n = 1,063) |
Overall Controls (n = 1,677) |
P a |
||
|---|---|---|---|---|---|
n |
% |
n |
% |
||
Age (years) |
64.19 (±8.63) |
63.91 (±10.22) |
0.451 |
||
Age (years) |
0.165 |
||||
< 64 |
494 |
46.47 |
825 |
49.19 |
|
≥64 |
569 |
53.53 |
852 |
50.81 |
|
Sex |
0.909 |
||||
Male |
759 |
71.40 |
1194 |
71.20 |
|
Female |
304 |
28.60 |
483 |
28.80 |
|
Smoking status |
< 0.001 |
||||
Never |
773 |
72.72 |
1323 |
78.89 |
|
Ever |
290 |
27.28 |
354 |
21.11 |
|
Alcohol use |
< 0.001 |
||||
Never |
908 |
85.42 |
1507 |
89.86 |
|
Ever |
155 |
14.58 |
170 |
10.14 |
|
a Two-sided χ2 test and Student t test.
Table 2: Primary information for PD-1 polymorphisms (PD-1 rs10204525 T>C, rs36084323 T>C, rs7421861 A>G and rs2227982 A>G)
Genotyped polymorphisms |
rs10204525 T>C (PD1.6) |
rs36084323 T>C (PD1.1) |
rs7421861 A>G (PD1.7) |
rs2227982 A>G (PD1.9) |
|---|---|---|---|---|
Chr |
2 |
2 |
2 |
2 |
Position_37 |
242792321 |
242801596 |
242795350 |
242793433 |
Region |
3’UTR |
Promoter |
Intron 1 |
Exon 5 |
MAFa for Chinese in database |
0.302 |
0.490 |
0.165 |
0.488 |
MAF in our controls (n = 1,677) |
0.280 |
0.496 |
0.168 |
0.492 |
P value for HWEb test in our controls |
0.888 |
0.071 |
0.232 |
0.309 |
% Genotyping value |
99.01 |
99.09 |
99.09 |
99.09 |
aMAF: minor allele frequency;
bHWE: Hardy–Weinberg equilibrium.
Association of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms with EGJA
The genotype distributions of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms are summarized in Table 3. The frequencies of PD-1 rs2227982 AA, AG, and GG genotypes were 26.13%, 52.74% and 21.13% in 1,063 EGJA patients and 26.40%, 48.75%, and 24.85% in 1,677 controls, respectively. When compared with the frequency of PD-1 rs2227982 AA genotype, a difference in the frequency of PD-1 rs2227982 GG genotype was found between the EGJA patients and the controls (crude OR = 0.80, 95% CI: 0.64–1.00, P = 0.047). When compared with the frequency of PD-1 rs2227982 AA/AG genotype, there was a difference in the frequency of PD-1 rs2227982 GG genotype between EGJA patients and the controls (crude OR = 0.81, 95% CI: 0.67–0.98, P = 0.026). Adjustment for age, sex, smoking and drinking, there was also difference in recessive genetic model (GG vs. AA/AG: adjusted OR, 0.81; 95% CI, 0.67–0.97; P = 0.024; Table 4).
Table 3: The frequencies of PD-1 rs10204525 T>C, rs36084323 T>C, rs7421861 A>G and rs2227982 A>G polymorphisms in esophagogastric junction adenocarcinoma patients and controls
Genotype |
Overall EGJA case (n = 1,063) |
Overall Controls (n = 1,677) |
||
|---|---|---|---|---|
n |
% |
n |
% |
|
rs36084323 T>C |
||||
TT |
282 |
27.09 |
444 |
26.52 |
TC |
521 |
50.05 |
800 |
47.79 |
CC |
238 |
22.86 |
430 |
25.69 |
CT+CC |
759 |
72.91 |
1,230 |
73.48 |
TT+CT |
803 |
77.14 |
1,244 |
74.31 |
C allele |
997 |
47.89 |
1,660 |
49.58 |
rs10204525 T>C |
||||
TT |
544 |
52.36 |
870 |
51.97 |
TC |
397 |
38.21 |
672 |
40.14 |
CC |
98 |
9.43 |
132 |
7.89 |
TC+CC |
495 |
47.64 |
804 |
48.03 |
TT+TC |
941 |
90.57 |
1,542 |
92.11 |
C allele |
593 |
28.54 |
936 |
27.96 |
rs7421861 A>G |
||||
AA |
642 |
61.67 |
1,166 |
69.65 |
AG |
358 |
34.39 |
454 |
27.12 |
GG |
41 |
3.94 |
54 |
3.23 |
AG+GG |
399 |
38.33 |
508 |
30.35 |
AA+AG |
1,000 |
96.06 |
1,620 |
96.77 |
G allele |
440 |
21.13 |
562 |
16.79 |
rs2227982 A>G |
||||
AA |
272 |
26.13 |
442 |
26.40 |
GA |
549 |
52.74 |
816 |
48.75 |
GG |
220 |
21.13 |
416 |
24.85 |
GG+GA |
769 |
73.87 |
1,232 |
73.60 |
GA+AA |
821 |
78.87 |
1,258 |
75.15 |
G allele |
989 |
47.50 |
1,648 |
49.22 |
Table 4: Overall and stratified analyses of PD-1 rs10204525 T>C, rs36084323 T>C, rs7421861 A>G and rs2227982 A>G polymorphisms with esophagogastric junction adenocarcinoma
Genotype |
Overall (1,063 cases vs. 1,677 controls) |
|||
|---|---|---|---|---|
Crude OR (95%CI) |
P |
Adjusted ORa (95%CI) |
P |
|
rs36084323 T>C |
||||
additive model |
0.96 (0.80–1.15) |
0.642 |
0.97 (0.81–1.17) |
0.766 |
homozygote model |
0.81 (0.66–1.01) |
0.061 |
0.82 (0.66–1.02) |
0.074 |
Dominant model |
0.97 (0.82–1.16) |
0.746 |
0.99 (0.83–1.18) |
0.888 |
Recessive model |
0.86 (0.72–1.03) |
0.097 |
0.86 (0.71–1.03) |
0.097 |
rs10204525 T>C |
||||
additive model |
0.91 (0.77–1.07) |
0.246 |
0.92 (0.78–1.08) |
0.294 |
homozygote model |
1.14 (0.86–1.51) |
0.359 |
1.16 (0.87–1.54) |
0.310 |
Dominant model |
0.99 (0.84–1.15) |
0.845 |
1.00 (0.85–1.16) |
0.956 |
Recessive model |
1.22 (0.93–1.60) |
0.160 |
1.23 (0.94–1.62) |
0.139 |
rs7421861 A>G |
||||
additive model |
1.39 (1.17–1.64) |
< 0.001 |
1.39 (1.18–1.65) |
< 0.001 |
homozygote model |
1.34 (0.88–2.03) |
0.173 |
1.31 (0.86–1.98) |
0.214 |
Dominant model |
1.43 (1.21–1.68) |
< 0.001 |
1.43 (1.21–1.68) |
<0.001 |
Recessive model |
1.23 (0.81–1.86) |
0.327 |
1.20 (0.79–1.82) |
0.394 |
rs2227982 A>G |
||||
additive model |
1.02 (0.85–1.22) |
0.845 |
1.04 (0.86–1.25) |
0.690 |
homozygote model |
0.80 (0.64–1.00) |
0.047 |
0.81 (0.65–1.01) |
0.057 |
Dominant model |
1.01 (0.85–1.21) |
0.874 |
1.03 (0.87–1.23) |
0.716 |
Recessive model |
0.81 (0.67–0.98) |
0.026 |
0.81 (0.67–0.97) |
0.024 |
aAdjusted for age, sex, smoking status and alcohol use in a logistic regression model.
The frequencies of PD-1 rs7421861 AA, AG, and GG genotypes were 61.67%, 34.39% and 3.94% in 1,063 EGJA patients and 69.65%, 27.12%, and 3.23% in 1,677 controls, respectively. When compared with the frequency of PD-1 rs7421861 AA genotype, there was difference in the frequency of PD-1 rs7421861 AG genotype between the EGJA patients and the controls (crude OR = 1.39, 95% CI: 1.17–1.64, P < 0.001). When compared with the frequency of PD-1 rs7421861 AA genotype, there was also difference in the frequency of PD-1 rs7421861 AG/GG genotype between the EGJA patients and the controls (crude OR = 1.43, 95% CI: 1.21–1.68, P < 0.001). Adjustments for age, sex, smoking and drinking, the observed results were not essentially changed (AG vs. AA: adjusted OR, 1.39; 95% CI, 1.18–1.65; P < 0.001; AG/GG vs. AA: adjusted OR, 1.43; 95% CI, 1.21–1.68; P < 0.001; Table 4).
The PD-1 rs36084323 T>C polymorphism conferred a borderline statistically decreased risk to EGJA in homozygote genetic model (crude OR = 0.81, 95% CI = 0.66–1.01, P = 0.061) and recessive genetic model (crude OR = 0.86, 95% CI = 0.72–1.03, P = 0.097). When adjusted for age, sex, smoking and drinking, a borderline statistically decreased risk of EGJA was also found in homozygote genetic model (crude OR = 0.82, 95% CI = 0.66–1.02, P = 0.074) and recessive genetic model (crude OR = 0.86, 95% CI = 0.71–1.03, P = 0.097). However, there was no difference in genotype distribution of PD-1 rs10204525 T>C polymorphism among EGJA patients and the controls (Table 4).
We used the Power and Sample Size Calculator (http://biostat.mc.vanderbilt.edu/twiki/bin/view/Main/PowerSampleSize) to calculate the power value (α = 0.05) [22]. For PD-1 rs7421861 A>G, the power value was 0.967 in the additive model and 0.990 in the dominant model. For PD-1 rs2227982 A>G, the power value was 0.675 in the homozygote model and 0.607 in the recessive model.
Association of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms with EGJA in different stratification groups
Table 5 summarizes the genotype frequencies of PD-1 rs10204525 T>C polymorphism in the stratified analysis by gender, age, alcohol consumption and smoking status. There was no significant difference in genotype distribution of PD-1 rs10204525 T>C polymorphism among EGJA patients and the controls in any subgroup.
Table 5: Stratified analyses between PD-1 rs10204525 T>C polymorphism and EGJA risk by sex, age, smoking status and alcohol consumption
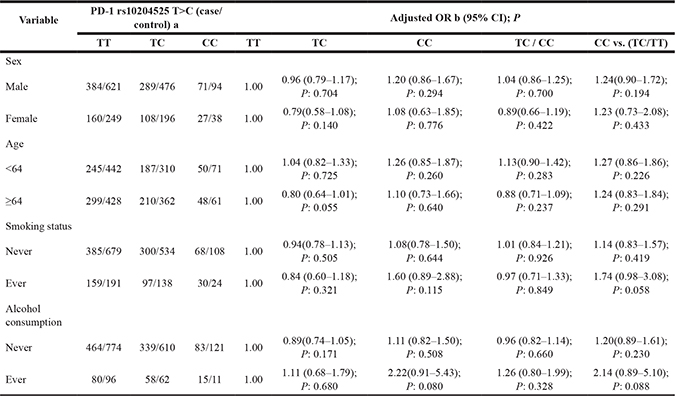
aThe genotyping was successful in 1063 (97.74%) EGJA cases, and 1677 (99.82%) controls for PD-1 rs10204525 T>C;
bAdjusted for age, sex, smoking status and alcohol consumption (besides stratified factors accordingly) in a logistic regression model.
The genotype frequencies of PD-1 rs36084323 T>C polymorphism in the stratified analysis by gender, age, alcohol consumption and smoking status are summarized in Table 6. In never smoking group, after adjustment for gender, age, alcohol consumption and smoking status by logistic regression analysis, the PD-1 rs36084323 CC genotype was associated with a significantly decreased risk of EGJA compared with the TC/TT genotype [CC vs. TC/TT: adjusted OR = 0.80, 95% CI 0.65–0.99, P = 0.043 (Table 6)]. In never drink group, after logistic regression analysis, the CC genotype of PD-1 rs36084323 T>C polymorphism was also associated with a significantly decreased risk of EGJA compared with the TT genotype [CC vs. TT: adjusted OR = 0.78, 95% CI 0.62–0.99, P = 0.037 (Table 6)].
Table 6: Stratified analyses between PD-1 rs36084323 T>C polymorphism and EGJA risk by sex, age, smoking status and alcohol consumption

aThe genotyping was successful in 1063 (97.93%) EGJA cases, and 1677 (99.82%) controls for PD-1 rs36084323 T>C;
bAdjusted for age, sex, smoking status and alcohol consumption (besides stratified factors accordingly) in a logistic regression model.
In the stratified analysis by gender, age, alcohol consumption and smoking status, the genotype frequencies of PD-1 rs7421861 A>G polymorphism are summarized in Table 7. After adjustment by logistic regression analysis, we found that PD-1 rs7421861 A>G polymorphism was associated with a significantly increased risk of EGJA in some subgroups [male group: AG vs. AA: adjusted OR = 1.40, 95% CI 1.15–1.70, P = 0.001 and AG/GG vs. AA: adjusted OR = 1.41, 95% CI 1.16–1.71, P = 0.001; female group: AG/GG vs. AA: adjusted OR = 1.46, 95% CI 1.07–2.00, P = 0.017; <64 years subgroup: AG vs. AA: adjusted OR = 1.62, 95% CI 1.27–2.08, P < 0.001; GG vs. AA: adjusted OR = 2.07, 95% CI 1.12–3.83, P = 0.021 and AG/GG vs. AA: adjusted OR = 1.73, 95% CI 1.36–2.20, P < 0.001; never smoking group: AG vs. AA: adjusted OR = 1.42, 95% CI 1.17–1.73, P < 0.001 and AG/GG vs. AA: adjusted OR = 1.45, 95% CI 1.20–1.75, P < 0.001 and never drinking group: AG vs. AA: adjusted OR = 1.42, 95% CI 1.19–1.70, P < 0.001 and AG/GG vs. AA: adjusted OR = 1.46, 95% CI 1.22–1.73, P < 0.001 (Table 7)].
Table 7: Stratified analyses between PD-1 rs7421861 A>G polymorphism and EGJA risk by sex, age, smoking status and alcohol consumption
Variable |
PD-1 rs7421861 A>G (case/control) a |
Adjusted OR b (95% CI); P |
||||||
|---|---|---|---|---|---|---|---|---|
AA |
AG |
GG |
AA |
AG |
GG |
AG/GG |
GG vs. (AG/AA) |
|
Sex |
||||||||
Male |
455/819 |
262/331 |
29/41 |
1.00 |
1.40 (1.15–1.70); P: 0.001 |
1.20 (0.74–1.97); P: 0.462 |
1.41 (1.16–1.71); P: 0.001 |
1.10 (0.67–1.79); P: 0.712 |
Female |
187/347 |
96/123 |
12/13 |
1.00 |
1.37 (0.99–1.89); P: 0.058 |
1.65 (0.73–3.69); P: 0.228 |
1.46 (1.07–2.00); P: 0.017 |
1.55(0.69–3.46); P: 0.286 |
Age |
||||||||
<64 |
286/590 |
173/212 |
23/21 |
1.00 |
1.62 (1.27–2.08); P: < 0.001 |
2.07 (1.12–3.83); P: 0.021 |
1.73 (1.36–2.20); P: < 0.001 |
1.82 (0.99–3.35); P: 0.054 |
≥64 |
356/576 |
185/242 |
18/33 |
1.00 |
1.21 (0.96–1.53); P: 0.108 |
0.87 (0.48–1.56); P: 0.633 |
1.20 (0.96–1.50); P: 0.113 |
0.83 (0.46–1.49); P: 0.531 |
Smoking status |
||||||||
Never |
465/925 |
263/353 |
26/43 |
1.00 |
1.42 (1.17–1.73); P: < 0.001 |
1.16 (0.70–1.92); P: 0.558 |
1.45 (1.20–1.75); P: < 0.001 |
1.07 (0.65–1.75); P: 0.804 |
Ever |
177/241 |
95/101 |
15/11 |
1.00 |
1.29 (0.91–1.82); P: 0.149 |
1.85 (0.82–4.17); P: 0.139 |
1.36 (0.98–1.90); P: 0.068 |
1.72 (0.77–3.85); P: 0.188 |
Alcohol consumption |
||||||||
Never |
544/1050 |
311/408 |
32/47 |
1.00 |
1.42 (1.19–1.70); P: < 0.001 |
1.26 (0.80–2.00); P: 0.326 |
1.46 (1.22–1.73); P: < 0.001 |
1.15 (0.73–1.82); P: 0.541 |
Ever |
98/116 |
47/46 |
9/7 |
1.00 |
1.14 (0.69–1.89); P: 0.609 |
1.73 (0.59–5.09); P: 0.322 |
1.21 (0.75–1.95); P: 0.441 |
1.66 (0.57–4.83); P: 0.356 |
a The genotyping was successful in 1063 (97.93%) EGJA cases, and 1677 (99.82%) controls for PD-1 rs7421861 A>G;
b Adjusted for age, sex, smoking status and alcohol consumption (besides stratified factors accordingly) in a logistic regression model.
In the stratified analysis by gender, age, alcohol consumption and smoking status, the genotype frequencies of PD-1 rs2227982 A>G polymorphism are summarized in Table 8. After adjustment by logistic regression analysis, the PD-1 rs2227982 GG genotype was associated with a decreased risk of EGJA compared with the AG/AA genotype in three groups [≥ 64 years subgroup: GG vs. AG/AA: adjusted OR = 0.72, 95% CI 0.56–0.93, P = 0.011; never smoking group: GG vs. AG/AA: adjusted OR = 0.74, 95% CI 0.60–0.92, P = 0.008 and never drinking group: GG vs. AG/AA: adjusted OR = 0.76, 95% CI 0.62–0.93, P = 0.008 (Table 8)]. Additionally, compared with the PD-1 rs2227982 AA genotype, the PD-1 rs2227982 GG genotypes were also associated with a significantly decreased risk of EGJA in never drink group [GG vs. AA: adjusted OR = 0.76, 95% CI 0.60–0.97, P = 0.024 (Table 8)].
Table 8: Stratified analyses between PD-1 rs2227982 A>G polymorphism and EGJA risk by sex, age, smoking status and alcohol consumption
Variable |
PD-1 rs2227982 A>G (case/control) a |
Adjusted OR b (95% CI); P |
||||||
|---|---|---|---|---|---|---|---|---|
AA |
AG |
GG |
AA |
AG |
GG |
AG/GG |
GG vs. (AG/AA) |
|
Sex |
||||||||
Male |
188/315 |
393/572 |
165/304 |
1.00 |
1.12(0.89–1.39); P: 0.333 |
0.87 (0.67–1.14); P: 0.310 |
1.10 (0.89–1.35); P: 0.399 |
0.83 (0.67–1.03); P: 0.093 |
Female |
84/127 |
156/244 |
55/112 |
1.00 |
0.87 (0.62–1.23); P: 0.433 |
0.66 (0.43–1.01); P: 0.057 |
0.90 (0.65–1.24); P: 0.513 |
0.75 (0.52–1.08); P: 0.124 |
Age |
||||||||
<64 |
126/212 |
253/426 |
103/185 |
1.00 |
0.93 (0.71–1.22); P: 0.597 |
0.87 (0.63–1.20); P: 0.383 |
0.99 (0.77–1.29); P: 0.948 |
0.93 (0.71–1.23); P: 0.625 |
≥64 |
146/230 |
296/390 |
117/231 |
1.00 |
1.13(0.87–1.45); P: 0.359 |
0.76 (0.56–1.03); P: 0.074 |
1.06 (0.83–1.35); P: 0.666 |
0.72 (0.56–0.93); P: 0.011 |
Smoking status |
||||||||
Never |
188/351 |
415/638 |
151/332 |
1.00 |
1.11 (0.90–1.37); P: 0.339 |
0.77 (0.60–1.00); P: 0.051 |
1.09 (0.89–1.34); P: 0.422 |
0.74(0.60–0.92); P: 0.008 |
Ever |
84/91 |
134/178 |
69/84 |
1.00 |
0.84 (0.58–1.22); P: 0.365 |
0.94 (0.61–1.46); P: 0.788 |
0.90 (0.63–1.28); P: 0.542 |
1.06 (0.73–1.54); P: 0.752 |
Alcohol consumption |
||||||||
Never |
224/387 |
481/739 |
182/379 |
1.00 |
1.04 (0.85–1.26); P: 0.714 |
0.76 (0.60–0.97); P: 0.024 |
1.03 (0.85–1.24); P: 0.783 |
0.76 (0.62–0.93); P: 0.008 |
Ever |
48/55 |
68/77 |
38/37 |
1.00 |
0.99 (0.59–1.67); P: 0.961 |
1.30 (0.70–2.41); P: 0.410 |
1.08 (0.67–1.76); P: 0.753 |
1.30 (0.76–2.24); P: 0.339 |
aThe genotyping was successful in 1063 (97.93%) EGJA cases, and 1677 (99.82%) controls for PD-1 rs2227982 A>G;
bAdjusted for age, sex, smoking status and alcohol consumption (besides stratified factors accordingly) in a logistic regression model.
SNP haplotypes
Using SHESIS software (http://analysis.bio-x.cn/myAnalysis.php) [23], we constructed six haplotypes (Table 9). We found that TGCA haplotypes with the order of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms in gene position significantly decreased the risk of EGJA (OR = 0.83, 95% CI = 0.71–0.96; P = 0.015). However, TGCG and other haplotypes with the same order of PD-1 SNPs in gene position significantly increased the risk of EGJA (OR = 22.19, 95% CI = 5.27–93.56; P < 0.001 and OR = 2.50, 95% CI = 1.73–3.60; P < 0.001).
Table 9: PD-1 haplotype frequencies (%) in cases and controls and risk of esophagogastric junction adenocarcinom
Haplotypes |
Case (n=2,126) |
Control (n=3,354) |
Crude OR (95% CI) |
P |
||
|---|---|---|---|---|---|---|
n |
% |
n |
% |
|||
Trs10204525Ars2227982Trs36084323Ars7421861 |
1000 |
49.24 |
1644 |
49.07 |
1.00 |
|
Crs10204525Grs2227982Crs36084323Grs7421861 |
354 |
17.43 |
544 |
16.24 |
1.07(0.92–1.25) |
0.394 |
Trs10204525Grs2227982Crs36084323Ars7421861 |
347 |
17.09 |
688 |
20.54 |
0.83(0.71–0.96) |
0.015 |
Crs10204525Grs2227982Crs36084323Ars7421861 |
198 |
9.75 |
374 |
11.16 |
0.87(0.72–1.05) |
0.150 |
Trs10204525Ars2227982Crs36084323Ars7421861 |
29 |
1.43 |
48 |
1.43 |
0.99(0.62–1.59) |
0.977 |
Trs10204525Grs2227982Crs36084323Grs7421861 |
27 |
1.33 |
2 |
0.06 |
22.19(5.27–93.56) |
<0.001 |
Others |
76 |
3.74 |
50 |
1.49 |
2.50(1.73–3.60) |
<0.001 |
DISCUSSION
EGJA is considered as a separated carcinoma entirety of upper digestive tract malignancies [24]. Although the incidence of GC is declining, the incidence of EGJA rapidly increases in both western (e.g. North America and Europe) and eastern Asian countries [2–4, 25]. Thus, EGJA is one of the most prevalent malignancies worldwide. The etiology of EGJA is very complicated. Accumulating evidences demonstrated that SNPs in some immune response genes might be associated with risk of cancer [6, 26–28]. In consideration of the vital role of PD-1 in tumor immunology, we chose PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms to examine their potential roles in EGJA. In the present study, we found that PD-1 rs2227982 A>G and rs36084323 T>C polymorphisms might decrease the risk of EGJA. However, PD-1 rs7421861 A>G might be a risk factor for EGJA. In addition, we found Trs10204525Grs2227982Crs36084323Ars7421861 haplotypes significantly decreased the risk of EGJA. On the contrary, Trs10204525Grs2227982Crs36084323Grs7421861 and other haplotypes increased the risk of EGJA.
PD-1 rs7421861 A>G polymorphism locates on intron 1, where a lot of regulatory components and splicing control elements may interact with it [29]. Some epidemiological studies focused on the effect of PD-1 rs7421861 A>G locus on the development of multiple cancers; however, the results remained inconsistent. Several case-control reported that PD-1 rs7421861 A>G polymorphism was not associated with cancer risk [19, 27, 28, 30]. However, Ge et al. found PD-1 rs7421861 A>G polymorphism might increase the risk of overall colorectal cancer [31]. Recently, a meta-analysis indicated that PD-1 rs7421861 A>G was correlated with a borderline statistically increased risk of overall cancer [32]. In the present study, we found PD-1 rs7421861 A>G polymorphism might be associated with the susceptibility of EGJA, which was similar to the findings of the previous studies [31, 32]. This study did not examine the potential effect of this polymorphism on regulating the expression of PD-1 in EGJA patient blood samples. However, in the future, we will conduct a further study to assess whether PD-1 rs7421861 A>G polymorphism is associated with the inhibited activation of T cells in EGJA patients, which may impede the surveillance mechanism of immune system.
PD-1 rs2227982 A>G polymorphism, a SNP in Exon 5, encodes a Val to Ala substitution in the extra-cellular domain of PD-1 receptor during protein synthesis, which influences the sequence, and may alter the function of PD-1 protein. In this study, we found that PD-1 rs2227982 A>G polymorphism was associated with the decreased risk of EGJA risk. Ren et al. reported that PD-1 rs2227982 A>G polymorphism was associated with the development of breast cancer [19]. In addition, several studies reported there was no significant association between PD-1 rs2227982 A>G polymorphism and cancer (e.g. esophageal squamous cell carcinoma, colorectal cancer, breast cancer and non-small cell lung cancer) [27, 28, 30, 31]. A recent meta-analysis suggested that PD-1 rs2227982 A>G polymorphism might be not associated with the risk of overall cancer. However, we found that only five case-control studies with relatively small sample sizes were included in this pooled analysis. The current evidence of the relationship might be very limited. Therefore, whether the Val to Ala substitution in the extra-cellular domain of PD-1 recepter does alter biological activity of PD-1 protein is needed to be further assessed.
Previous report showed that PD-1 rs36084323 CC was associated with a significantly decreased risk of breast cancer compared with the TT genotype [30]. In the present study, we found a borderline statistically decreased risk to EGJA in homozygote genetic model and recessive genetic model. In the stratified analysis by gender, age, alcohol consumption and smoking status, the decreased risk was observed in never smoking group and in never drinking group. To the best of our knowledge, variants in the promoter region of functional gene may influence an initial binding of transcription factors with sequence motifs, and further alter gene expression [33, 34]. Accordingly, as a variant in promoter region of PD-1 gene, PD-1 rs36084323 CC genotype may also affect the activation of transcription, and then regulate the expression of PD-1 gene and increase the risk of EGJA.
In this study, we constructed haplotypes to explore the potential inherited patterns of haplotype. We found that PD-1 Trs10204525Grs2227982C36084323Ars7421861 haplotype significantly decreased the risk of EGJA. However, Trs10204525Grs2227982C36084323Grs7421861 haplotype in PD-1 gene might significantly increase the risk of EGJA. We first studied the relationship of haplotypes in PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms with EGJA susceptibility. Compared PD-1 Trs10204525Grs2227982C36084323Ars7421861 with Trs10204525Grs2227982C36084323Grs7421861 haplotype, we also found that A→G variation in PD-1 rs7421861 locus might significantly inverse the risk of haplotype to EGJA.
In addition, some limitations in this case-control study should be acknowledged. All participants were enrolled in three hospitals from Eastern Chinese Han population. Although the genotype distributions of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms in controls were consistent with HWE and the MAF in the selected controls was very close to the data of Chinese (Table 2), the non-cancer controls might not well-represent the whole Chinese population. As well, only four important SNPs (MAF ≥ 0.05) in PD-1 gene were selected, which might not be enough to determine the total genetic susceptibility in PD-1 gene. Future, a fine-mapping study with larger sample sizes, multiple centers and detailed risk factors is need to confirm these primary findings.
In summary, our study highlights rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms and haplotypes in PD-1 gene, especially within the intron region, may be associated with the risk of EGJA in Eastern Chinese Han population. Our primary findings suggest that PD-1 genetic variation may be beneficial for the exploration of Eastern Chinese subjects genetically susceptible to EGJA.
MATERIALS AND METHODS
Subjects
A total of 2,740 participants were recruited in this case-control study. Among them, 280 EGJA patients were enrolled from the Affiliated Union Hospital of Fujian Medical University and Fujian Medical University Cancer Hospital from January 2014 to May 2016. In addition, 783 EGJA patients were recruited from the Affiliated People’s Hospital of Jiangsu University from January 2008 to November 2016. All cases with histologically confirmed EGJA were enrolled in the present study. EGJA patients who had a history of another malignancy or received prior chemoradiotherapy were excluded. At the same time, 1,677 subjects who underwent health check in these hospitals were recruited as non-cancer controls. Controls were matched with EGJA cases in terms of gender and age. All participants were unrelated Eastern Chinese Han population. Each participant signed the written informed consent. The study was approved by the Review Boards of Jiangsu University (Zhenjiang, China) and Fujian Medical University (Fuzhou, China), in accordance with the Declaration of Helsinki. Two experienced doctors interviewed each individual and collected the relevant risk factors and demographic variables. The corresponding data are summarized in Table 1. Each study participant provided an ethylenediamine tetraacetic acid (EDTA)-anticoagulated intravenous blood sample after an overnight fast. The approved guidelines were used as experimental protocol.
Selection of SNPs
The tagging polymorphisms among the gene region of PD-1 (upstream and downstream of gene extending 5 Kb, respectively) were collected from the CHB population via an internet the HapMap Project (http://hapmap.ncbi.nlm.nih.gov/index.html.en) and were conducted with Haploview 4.2 software with the criterion of pairwise linkage disequilibrium (LD) r2 threshold of 0.8 between polymorphisms (with a minimum LD of r2 > 0.8) [35]. Finally, SNPs with a HWE P ≥ 0.05, MAF ≥ 0.05 and call rate ≥ 95 % in the CHB population were included [36]. PD-1 rs36084323 T>C polymorphism locates on the promoter of PD-1 gene, which region may influence the transcription of PD-1. Thus, in this study, we also included this important SNP. The primary information of PD-1 functional SNPs is summarized in Table 2.
DNA extraction and genotyping
Peripheral blood sample was collected and stored at −20°C. Using the Promega Genomic DNA Purification Kit (Promega, Madison, USA), the genomic DNA was carefully extracted from lymphocytes. The obtained genomic DNA was frozen at −80°C. SNPscanTM genotyping assay (Gnensky Biotechologies Inc., Shanghai, China) was used to analyze the genotyping of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms. For quality control, 110 DNA samples (4%) were randomly selected. The genotypes of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms were reanalyzed by different laboratory technicians. The reproducibility was 100%. The success rate of PD-1 genotyping is shown in Table 2.
Statistical analysis
Age was expressed as the mean ± standard deviation (SD). And we used Student’s t-test to calculate the differences for continuous variables between EGJA patients and controls. An internet-based calculator (http://ihg.gsf.de/cgi-bin/hw/hwa1.pl) was used to determine the deviation of genotype frequencies from HWE. Chi-square test (χ2) was conducted to compare the categorical variables (e.g. age, sex, smoking status, and drinking) and the genotype distributions between EGJA patients and non-cancer controls. Multivariate logistic regression was used to obtain the crude/adjusted odds ratios (OR) and their 95 % confidence intervals (CI) for the relationship of PD-1 rs10204525 T>C, rs2227982 A>G, rs36084323 T>C and rs7421861 A>G polymorphisms with EGJA risk. The SAS software (Version 9.4; SAS Institute Inc., Cary, NC, USA) was used to analyze all data. SHESIS online software [(http://analysis.bio-x.cn/myAnalysis.php), Bio-X Inc., Shanghai, China] was harnessed to construct the haplotypes of PD-1 gene [23]. The criterion of statistical significance was defined as P < 0.05 (two-tailed). We performed the Power and Sample Size Calculator (http://biostat.mc.vanderbilt.edu/twiki/bin/view/Main/PowerSampleSize) to assess the power value of the study (α = 0.05) [22].
ACKNOWLEDGMENTS
We appreciate all subjects who participated in this study. We wish to thank Dr. Yan Liu (Genesky Biotechnologies Inc., Shanghai, China) for technical support.
CONFLICTS OF INTEREST
The authors have no potential financial conflicts of interest.
GRANT SUPPORT
This study was supported in part by Young and Middle-aged Talent Training Project of Health Development Planning Commission in Fujian Province (2016-ZQN-25 and 2014-ZQN-JC-11), Medical Innovation Project of Fujian Province (2014-CX-15 and 2014-CX-18), Nursery Garden Project of Fujian Medical University (2015MP020) and Science and Technology Project of Fujian Province (2060203).
REFERENCES
1. Jemal A, Siegel R, Xu J, Ward E. Cancer statistics, 2010. CA Cancer J Clin. 2010; 60:277–300.
2. Jemal A, Center MM, DeSantis C, Ward EM. Global patterns of cancer incidence and mortality rates and trends. Cancer Epidem Biomar. 2010; 19:1893–1907.
3. Blaser MJ, Saito D. Trends in reported adenocarcinomas of the oesophagus and gastric cardia in Japan. Eur J Gastroen Hepat. 2002; 14:107–113.
4. Zhou Y, Zhang Z, Zhang Z, Wu J, Ren D, Yan X, Wang Q, Wang Y, Wang H, Zhang J, Zhu X, Yang Y, Luo C, et al. A rising trend of gastric cardia cancer in Gansu Province of China. Cancer Lett. 2008; 269:18–25.
5. Sun T, Zhou Y, Yang M, Hu Z, Tan W, Han X, Shi Y, Yao J, Guo Y, Yu D, Tian T, Zhou X, Shen H, et al. Functional genetic variations in cytotoxic T-lymphocyte antigen 4 and susceptibility to multiple types of cancer. Cancer Res. 2008; 68:7025–7034.
6. Tang W, Wang Y, Chen S, Lin J, Chen B, Yu S, Chen Y, Gu H, Kang M. Investigation of Cytotoxic T-lymphocyte antigen 4 Polymorphisms in Gastric Cardia Adenocarcinoma. Scand J Immunol. 2016; 83:212–218.
7. Ishida Y, Agata Y, Shibahara K, Honjo T. Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death. Embo J. 1992; 11:3887–3895.
8. Ceeraz S, Nowak EC, Noelle RJ. B7 family checkpoint regulators in immune regulation and disease. Trends Immunol. 2013; 34:556–563.
9. Li Y, Li F, Jiang F, Lv X, Zhang R, Lu A, Zhang G. A Mini-Review for Cancer Immunotherapy: Molecular Understanding of PD-1/PD-L1 Pathway & Translational Blockade of Immune Checkpoints. Int J Mol Sci. 2016; 17.
10. Francisco LM, Sage PT, Sharpe AH. The PD-1 pathway in tolerance and autoimmunity. Immunol Rev. 2010; 236:219–242.
11. Mahmoudi M, Rezaiemanesh A, Salmaninejad A, Harsini S, Poursani S, Bahrami T, Tahghighi F, Ziaee V, Rezaei N. PDCD1 single nucleotide genes polymorphisms confer susceptibility to juvenile-onset systemic lupus erythematosus. Autoimmunity. 2015; 48:488–493.
12. Kristjansdottir H, Steinsson K, Gunnarsson I, Grondal G, Erlendsson K, Alarcon-Riquelme ME. Lower expression levels of the programmed death 1 receptor on CD4+CD25+ T cells and correlation with the PD-1.3A genotype in patients with systemic lupus erythematosus. Arthritis Rheum. 2010; 62:1702–1711.
13. Huang CH, Wong RH, Wei JC, Tsay MD, Chen WC, Chen HY, Shih WT, Chiou SP, Tu YC, Lee HS. Effects of genetic polymorphisms of programmed cell death 1 and its ligands on the development of ankylosing spondylitis. Rheumatology. 2011; 50:1809–1813.
14. Liu X, Hu LH, Li YR, Chen FH, Ning Y, Yao QF. Programmed cell death 1 gene polymorphisms is associated with ankylosing spondylitis in Chinese Han population. Rheumatol Int. 2011; 31:209–213.
15. Ni R, Ihara K, Miyako K, Kuromaru R, Inuo M, Kohno H, Hara T. PD-1 gene haplotype is associated with the development of type 1 diabetes mellitus in Japanese children. Hum Genet. 2007; 121:223–232.
16. Liu C, Jiang J, Gao L, Hu X, Wang F, Shen Y, Yu G, Zhao Z, Zhang X. A Promoter Region Polymorphism in PDCD-1 Gene Is Associated with Risk of Rheumatoid Arthritis in the Han Chinese Population of Southeastern China. Int J Genomics. 2014; 2014:247637.
17. Yin L, Guo H, Zhao L, Wang J. The programmed death-1 gene polymorphism (PD-1.5 C/T) is associated with non-small cell lung cancer risk in a Chinese Han population. Int J Clin Exp Med. 2014; 7:5832–5836.
18. Haghshenas MR, Dabbaghmanesh MH, Miri A, Ghaderi A, Erfani N. Association of PDCD1 gene markers with susceptibility to thyroid cancer. J Endocrinol Invest. 2016.
19. Ren HT, Li YM, Wang XJ, Kang HF, Jin TB, Ma XB, Liu XH, Wang M, Liu K, Xu P, Yao QL, Dai ZJ. PD-1 rs2227982 Polymorphism Is Associated With the Decreased Risk of Breast Cancer in Northwest Chinese Women: A Hospital-Based Observational Study. Medicine. 2016; 95:e3760.
20. Li XF, Jiang XQ, Zhang JW, Jia YJ. Association of the programmed cell death-1 PD1.5 C>T polymorphism with cervical cancer risk in a Chinese population. Genet Mol Res. 2016; 15.
21. Savabkar S, Azimzadeh P, Chaleshi V, Nazemalhosseini Mojarad E, Aghdaei HA. Programmed death-1 gene polymorphism (PD-1.5 C/T) is associated with gastric cancer. Gastroenterol Hepatol Bed Bench. 2013; 6:178–182.
22. Tang W, Qiu H, Ding H, Sun B, Wang L, Yin J, Gu H. Association between the STK15 F31I polymorphism and cancer susceptibility: a meta-analysis involving 43,626 subjects. PloS one. 2013; 8:e82790.
23. Shi YY, He L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res. 2005; 15:97–98.
24. Kim HI, Cheong JH, Song KJ, An JY, Hyung WJ, Noh SH, Kim CB. Staging of adenocarcinoma of the esophagogastric junction: comparison of AJCC 6th and 7th gastric and 7th esophageal staging systems. Ann Surg Oncol. 2013; 20:2713–2720.
25. Colquhoun A, Arnold M, Ferlay J, Goodman KJ, Forman D, Soerjomataram I. Global patterns of cardia and non-cardia gastric cancer incidence in 2012. Gut. 2015; 64:1881–1888.
26. Tang W, Wang Y, Jiang H, Liu P, Liu C, Gu H, Chen S, Kang M. Programmed death-1 (PD-1) rs2227981 C > T polymorphism is associated with cancer susceptibility: a meta-analysis. Int J Clin Exp Med. 2015; 8:22278–22285.
27. Tang W, Chen Y, Chen S, Sun B, Gu H, Kang M. Programmed death-1 (PD-1) polymorphism is associated with gastric cardia adenocarcinoma. Int J Clin Exp Med. 2015; 8:8086–8093.
28. Qiu H, Zheng L, Tang W, Yin P, Cheng F, Wang L. Programmed death-1 (PD-1) polymorphisms in Chinese patients with esophageal cancer. Clin Biochem. 2014; 47:612–617.
29. Gazave E, Marques-Bonet T, Fernando O, Charlesworth B, Navarro A. Patterns and rates of intron divergence between humans and chimpanzees. Genome Biol. 2007; 8:R21.
30. Hua Z, Li D, Xiang G, Xu F, Jie G, Fu Z, Jie Z, Da P, Li D. PD-1 polymorphisms are associated with sporadic breast cancer in Chinese Han population of Northeast China. Breast Cancer Res Tr. 2011; 129:195–201.
31. Ge J, Zhu L, Zhou J, Li G, Li Y, Li S, Wu Z, Rong J, Yuan H, Liu Y, Chi Q, Piao D, Zhao Y, et al. Association between co-inhibitory molecule gene tagging single nucleotide polymorphisms and the risk of colorectal cancer in Chinese. J Cancer Res Clin. 2015; 141:1533–1544.
32. Zhang J, Zhao T, Xu C, Huang J, Yu H. The association between polymorphisms in the PDCD1 gene and the risk of cancer: A PRISMA-compliant meta-analysis. Medicine. 2016; 95:e4423.
33. de Vooght KM, van Solinge WW. Gene promoter analysis in molecular diagnostics: do or don’t? Expert Rev Mol Diagn. 2009; 9:403–405.
34. de Vooght KM, van Wijk R, van Solinge WW. Management of gene promoter mutations in molecular diagnostics. Clin Chem. 2009; 55:698–708.
35. Tang W, Zhang S, Qiu H, Wang L, Sun B, Yin J, Gu H. Genetic variations in MTHFR and esophageal squamous cell carcinoma susceptibility in Chinese Han population. Med Oncol. 2014; 31:915.
36. Carlson CS, Eberle MA, Kruglyak L, Nickerson DA. Mapping complex disease loci in whole-genome association studies. Nature. 2004; 429:446–452.

