INTRODUCTION
Lung cancer has the highest morbidity rates of cancer disease, in particular non-small cell lung carcinoma (NSCLC), which accounts for about 80% of lung cancers [1]. While surgery, radiotherapy and chemotherapy have been used to treat NSCLC for several years, the five year survival rate unfortunately remains less than 2% [2]. Tyrosine kinase inhibitors targeting the epidermal growth factor receptor (EGFR-TKIs) have been recently developed and appear to work effectively against several types of cancer, including NSCLC [3]. EGFR is a member of the HER family of transmembrane tyrosine kinase receptors that leads to the production of the proto-oncogene c-erbB1. EGFR plays one of the fundamental roles in the regulation of numerous cellular and physiological processes in homeostasis and during tumorinogenesis, such as cell growth, proliferation, differentiation, angiogenesis and metastasis. EGFR has three domains: an extracellular ligand-binding region, a single transmembrane helix and a cytoplasmic segment [4]. EGF and TGFa(transforming growth factor alpha) ligands, as well as other antigens, bind to the extracellular domain thereby activating the downstream signaling pathways [5]. In NSCLC, an EGFR mutation in the intracellular region has been often identified in correlation with sensitivity to EGFR-TKIs [6]. However, while first generation EGFR-TKIs gefitinib (Iressa,ZD1839; AstraZeneca) and erlotinib (Tarceva, OSI-774; Genentech) have worked well in a subset of patients carrying mutations in the EGFR kinase domain in NSCLC [7], a significant proportion developed primary and secondary drug resistance during treatment [8]. About half of the drug sensitive NSCLC patients harbor a secondary mutation in the EGFR [9] that prevents effective inhibition by EGFR-TKIs due to steric hindrance or increased binding affinity for adenosine triphosphate(ATP) [10]. Thus, detecting the EGFR-activating mutation status prior to treatment is the key for predicting the efficacy of EGFR-TKI therapy in NSCLC patients.
Direct gene sequencing is the standard method used for detection of EGFR mutations in cancer, however, its many limitations, such as low sensitivity and length of procedure (several days), restrict its use in the clinic. Furthermore, gene sequencing cannot reflect EGFR mutation status in the whole tumor. Thus, a noninvasive method to identify patients sensitive to EGFR-targeted therapy and to monitor the EGFR mutation status in real-time during treatment with EGFR-TKIs is urgently needed. Since 1999, when Ralph weissleder from Harvard University first reported the noninvasive imaging technology–Molecular Imaging, it is a useful tool that can measure and characterize a variety of biological processes at the cellular or molecular levels in vivo [11], and which could therefore be utilized to identify EGFR activating mutations and to assess tumor response to EGFR-TKIs therapy in vivo. Notably, some PET(Positron emission tomography) tracers, such as 18F-PEG6-IPQA [12, 13], 11C-erlotinib, 11C-PD153035 and 11C-gefitinib [14–16], and some antibodies [17], have been synthesized and used to evaluate the mutation status of EGFR in NSCLC.
PD153035 is a EGFR tyrosine kinase inhibitor that could bind to activating mutant EGFR(19 exon deletion and L858R point mutation) and wild-type EGFR, but it can not binds to L858R/T790M mutant EGFR, and the capacity of PD153035 binds to activating mutant EGFR is much higher than that in wild-type EGFR. We previously reported a novel SPECT(Single-Photon Emission Computed Tomography) probe, HYNIC-MPG (2-(2-(2-(2-(4-(3-chloro-4- fluorophenylamino)-6-methoxyquinazolin-7-yloxy)ethoxy)ethoxy)ethoxy)ethyl-6-hydrazinylnicotinate hydrochloride), a analogue of PD153035, which could be labeled by 99mTc for targeting mutant (activity) EGFR [18], and we evaluated its chemical and biological properties, including its EGFR binding capacity in vitro and in vivo. In the current study, we further explore the potential of 99mTc-HYNIC-MPG as a novel SPECT radiotracer for the in vivo identification of EGFR exon 19 deletion in NSCLC xenografts.
RESULTS
Cell uptake, efflux and blocking assay of 99mTc-HYNIC-MPG
Cellular uptake and efflux experiments were performed to validate the specificity of 99mTc-HYNIC-MPG in cell levels (Figure 1). PC9 cell lines have a higher uptake ratio than that of H1975, H358 and H520 in all time points examined. At 2h, the accumulation of 99mTc-HYNIC-MPG in PC9 cells reached the highest level at 22.73 ± 1.63% of total input activity, which is about 4.4, 4.6 and 4.8 times higher than H1975, H358 and H520, respectively (P < 0.01). However, the high uptake of 99mTc-HYNIC-MPG in PC9 cells was blocked in the presence of 100 uM PD153035, and the uptake ratio decreased from 22.73 ± 1.63% to 4.89 ± 0.08% of total input activity (P < 0.01), and there is no significant change in the other three NSCLC cell lines. This result suggests that the high uptake rate of 99mTc-HYNIC-MPG in PC9 cells is related to the expression levels of EGFR exon 19 E746-A750 deletion mutant proteins. In the cell efflux assay, 99mTc-HYNIC-MPG was equally well retained in PC9 cells, even at 2h, and the ratio of total input activity in PC9 cells remained at 6.16 ± 0.28%, which was much higher than that of H1975, H358 and H520 cells. These data indicate that 99mTc-HYNIC-MPG might be used for long-term monitoring of the dynamic process of EGFR mutation status at the cellular level.
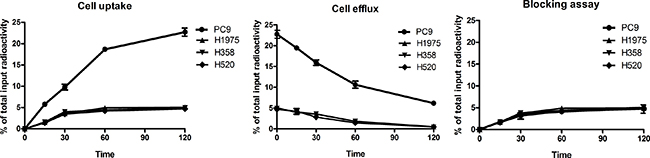
Figure 1: Cell uptake, efflux and blocking assay of 99mTc-HYNIC-MPG in NSCLC cell lines.
SPECT imaging and blocking assay
In vivo SPECT scans were performed in four xenograft groups (PC9, H1975, H358, H520); each group had three tumor-bearing mice. The uptake of 99mTc-HYNIC-MPG in the PC9 tumor was significantly higher than in the other tumors at all time points examined (Figure 2). Tumor to muscle (T/M) ratios were calculated over the region of interest (Table 1). 2 h after the injection of 99mTc-HYNIC-MPG into the tail vein, the T/M ratio in the PC9 group reached 5.47 ± 0.37, versus that of 2.45 ± 0.32 in the H1975 group, 2.35 ± 0.40 in the H358 group and 1.72 ± 0.28 in the H520 group (P < 0.01). The accumulation of 99mTc-HYNIC-MPG in the PC9 tumor was significantly decreased at the presence of 100 mg/Kg PD153035 and the T/M ratio was also decreased to 2.51 ± 0.30 (Figure 3), suggesting that high uptake of 99mTc-HYNIC-MPG in the PC9 tumor is related to the EGFR-activating mutation.
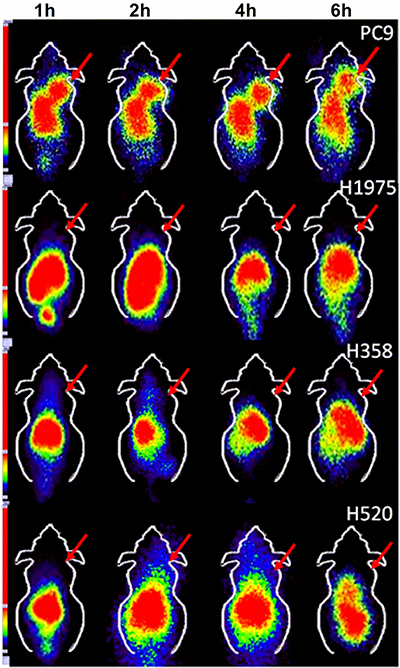
Figure 2: SPECT imaging of 99mTc-HYNIC-MPG in PC9, H1975, H358 and H520 xenografts at 1 h, 2 h, 4 h, and 6 h time points.
Table 1: Tumor to muscle ratio of ROI
1 h |
2 h |
4 h |
6 h |
|
|---|---|---|---|---|
PC9 |
3.63 ± 0.23 |
5.47 ± 0.37 |
4.21 ± 0.35 |
4.11 ± 0.36 |
H1975 |
1.67 ± 0.09 |
2.45 ± 0.32 |
1.81 ± 0.08 |
1.62 ± 0.21 |
H358 |
1.55 ± 0.20 |
2.35 ± 0.40 |
1.66 ± 0.40 |
1.32 ± 0.23 |
H520 |
1.37 ± 0.18 |
1.72 ± 0.28 |
1.64 ± 0.24 |
1.19 ± 0.19 |
Data was analyzed by mean ± SD (n = 3).
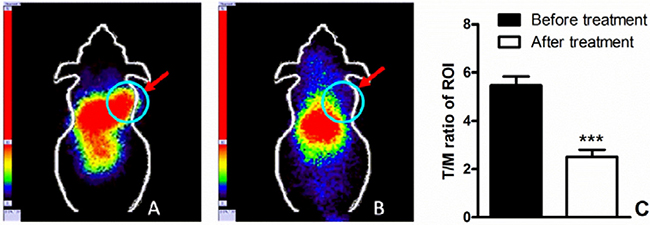
Figure 3: (A) 200 uci of 99mTc-HYNIC-MPG was injected via the tail vein, 2 h later, SPECT imaging was performed on PC9 xenografts; (B) 100 mg/Kg PD153035 was injected before SPECT imaging; T/M ratio before and after treatment was shown in (C).
Ex vivo biodistribution
To quantify the SPECT imaging results of 99mTc-HYNIC-MPG in xenografts, ex vivo biodistribution was monitored at 1 h, 2 h, 4 h and 6 h (Supplementary Tables 1–4). The uptake of 99mTc-HYNIC-MPG in the PC9 tumor reached 7.20 ± 0.27% ID/g at 2h post injection, compared to 2.35 ± 0.14% ID/g in H1975, 2.57 ± 0.20% ID/g in H358 and 1.98 ± 0.13% ID/g in H520 tumors (P < 0.01), shown in Figure 4. The T/M ratio of 99mTc-HYNIC-MPG accumulation in the PC9 group at 2h was 5.14 ± 0.83. In contrast, the T/M ratio was only 2.01 ± 0.19 in H1975, 2.02 ± 0.19 in H358 and 1.42 ± 0.04 in H520 tumors. In the presence of 100 mg/Kg PD153035, the uptake of 99mTc-HYNIC-MPG in PC9 tumors decreased from 7.20 ± 0.27% ID/g to 2.67 ± 0.27% ID/g , and the T/M ratio of 99mTc-HYNIC-MPG accumulation in the PC9 group was reduced to 2.32 ± 0.18. Thus, this ex vivo biodistribution data is in accord with the SPECT imaging results. As expected, the highest accumulation of 99mTc-HYNIC-MPG occurred in the liver, as this is the largest metabolic organ and the pharmacokinetics of these small molecules takes places mainly in the liver. However, adding 4 ethylene glycol to PD153035 decreased the 99mTc-HYNIC-MPG uptake ratio in the liver (16.89 ± 1.56), in agreement with what was previously described [18]. As PEGs(polyethylene glycol) may reduce the liposolubility of the probe, these results suggest that PEGs facilitate the access of 99mTc-HYNIC-MPG to the tumor region by preventing its accumulation in the liver. The uptake of 99mTc-HYNIC-MPG in the brain was very low in all tumor-bearing mice at all time points examined, suggesting that 99mTc-HYNIC-MPG cannot cross the blood-brain barrier.
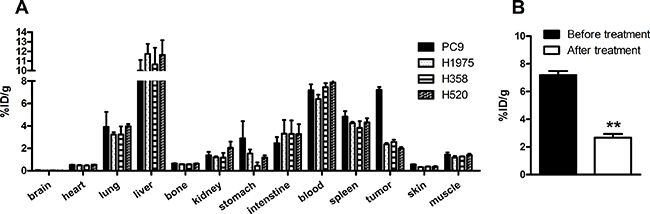
Figure 4: (A) biodistribution of 99mTc-HYNIC-MPG in major organs at 2 h time point; (B) The uptake of 99mTc-HYNIC-MPG in PC9 tumor reached 7.20 ± 0.27% ID/g 2 h post injection through the tail vein, and was blocked at the presence of 100 mg/Kg PD153035.
Western blot analysis and immunofluorescence Staining
Expression levels of wild type EGFR and EGFR exon 19 E746-A750 deletion proteins were quantified in PC9, H1975, H358 and H520 NSCLC tumors by western blot assays and immunofluorescence staining (Figure 5 and Figure 6). PC9, H1975 and H358 all express EGFR protein, but PC9 also expresses EGFR 19 deletion protein, suggesting that the high uptake of 99mTc-HYNIC-MPG in PC9 is likely due to the presence of EGFR exon 19 E746-A750 deletion protein.
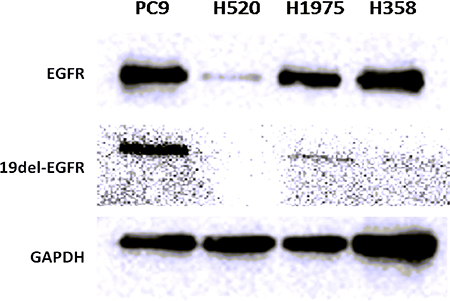
Figure 5: Western blot of EGFR and 19del-EGFR expression level in 4 NSCLC tumors.
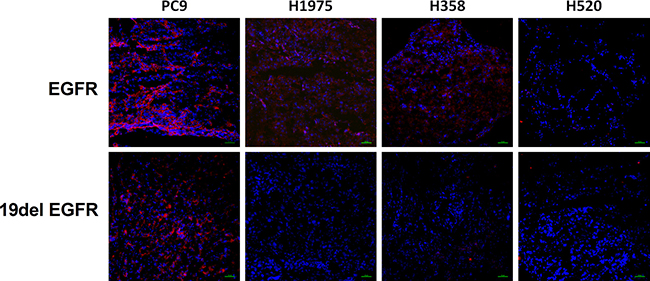
Figure 6: Immunofluorescence staining of EGFR and EGFR exon 19 (19del EGFR) expression level in tumor tissues. Above, red color represents EGFR expression, while blue color represents cell nuclei; blow, red color represents the level of EGFR exon 19 deletion, while blue color represents cell nuclei.
DISCUSSION
Currently, the most widely used methods for detecting EGFR mutations in NSCLC are gene sequencing, fluorescent in situ hybridazation and immunohistochemical assays, however, they present important limitations for EGFR-TKIs therapy. First, analyses from aspiration biopsies cannot reflect the EGFR mutation status of a whole tumor due to its heterogeneity. Second, repeated aspiration biopsies in advanced NSCLC can cause psychological and physical harm, which most of NSCLC patients cannot tolerate. In contrast, noninvasive SPECT imaging offers enormous potential to overcome these hurdles. Indeed, we have shown here that a 99mTc-HYNIC-MPG radiotracer can selectively accumulate in NSCLC as a result of EGFR expressing levels and mutation status in the tumor region. These results demonstrate that noninvasive SPECT imaging could be used to select patients sensitive to EGFR-TKIs therapy.
EGFR mutations are an excellent target for TKIs and occur in 10–35%of NSCLCs [19]. We have recently developed a novel SPECT tracer (99mTc-HYNIC-MPG) that binds specifically and stably to EGFR 19 deletion proteins in vitro[18]. Previously others have utilized alternative radionuclide labeling with EGFR-TKIs, for instance, 18F-gefitinib, 11C-PD 153035 or 11C-gefitinib, among others [14–16]. However, the half-lives of 18F and 11C are 110 min and 20.34 min respectively, which is not sufficient for real-time monitoring of EGFR-TKIs efficacy in vivo [20–22]. In addition, the procedures for synthesizing these probes are highly complex, time-consuming and costly, which greatly limits their availability from commercial manufacturers. In contrast, 99mTc can label many compounds for imaging in nearly every organ in vivo at low cost. Furthermore, 99mTc has a long half-life (6.02 h), a good range of γ ray energy (140 keV) and it is easy to transport, which facilitates its clinical application [23, 24].
While currently available probes targeting EGFR bind to the extracellular region of both wild type and mutant proteins, 99mTc-HYNIC-MPG binds specifically to EGFR 19 E746-A750 deletion proteins. This binding specificity offers the unique advantage of allowing the identification of patients carrying mutations sensitive to EGFR-TKI therapy. Furthermore, with its high sensitivity and stability, 99mTc-HYNIC-MPG has the optimal properties for real-time monitoring of the EGFR mutation status during EGFR-TKI therapy in vivo. However, this promising probe has some limitations that should be addressed in future investigations. For instance, the detailed mechanism for 99mTc-HYNIC-MPG targeting to mutant EGFR remains unclear. Furthermore, although nonspecific uptake of 99mTc-HYNIC-MPG to some organs is reduced when compared to other probes, it is still not low enough for clinical use, and it should therefore be improved by direct modification or other methods. Finally, small animal imaging SPECT equipment could be used to improve the quality of image.
In conclusion, 99mTc-HYNIC-MPG SPECT imaging is a promising approach for selecting NSCLC tumors carrying EGFR exon 19 deletion mutation sensitive to EGFR-TKIs therapy. In view of the capacity of binding to NSCLC EGFR 19 deletion protein, a clinical study is required.
MATERIALS AND METHODS
Radiosynthesis of 99mTc-HYNIC-MPG
99mTc-HYNIC-MPG was synthesized as previously described[18]: a mix of 0.5 ml EDDA solution (10 mg/ml), 10 μl HYNIC-MPG (3 mg/ml), 10 μl SnCl2 (0.2 mg/ml) and Tricine 20 mg with nitrogen protection was heated at 100oC for 10 min (all reagents were purchased from Sigma Company), then purified with C18 Sep-Pak plus cartridges. At the end of synthesis, the radiochemical purity of 99mTc-HYNIC-MPG was > 98%.
Cell culture
Four human NSCLC cell lines were used in this study: PC9 (EGFR 19 exon E746-A750 deletion), H1975 (EGFR L858R /T790M double mutation), H358 (EGFR wild type) and H520 (EGFR negative expression), which were all purchased from American Type Culture Collection (ATCC). Cells were grown in RPMI 1640 medium (HyClone) containing 10% fetal bovine serum (FBS) (HyClone) and then cultured at 37oC in a humidified atmosphere with 5% CO2.
Cell uptake, efflux and blocking assays
Cell uptake: 5 × 106 cells were seeded in 12-well plates 24 h prior to uptake assay. 37 KBq (1.0 μCi/ml) of 99mTc-HYNIC-MPG was added to each well. At 15 min, 30 min, 1 h and 2 h time points, the media was removed and washed twice with 1 ml of ice-cold PBS. 200 μl 0.1 N NaOH was then added for dialysis of the cells. After dialysis the cells were washed twice with 200 μl of ice-cold PBS. The cells were recovered in PBS and counted with CPM (counts per minute) in a gamma counter (PerkinElmer 2480). For the blocking assay, PD153035 (100 μM) was added before 99mTc-HYNIC-MPG in NSCLC cells. 1h later, the cell uptake was performed as mentioned above.
Cell efflux: Efflux assays were performed after 2 h incubation with 99mTc-HYNIC-MPG. The media was then removed and replaced with serum free medium without 99mTc-HYNIC-MPG. 15 min, 30 min, 1 h or 2 h later, the media was removed and the cells were washed twice with ice-cold PBS. 200 μl 0.1 N NaOH was added for dialysis of cells. The well was then washed twice with 200 μl ice-cold PBS. The cells were recovered in PBS and counted with CPM (counts per minute) in a gamma counter (PerkinElmer 2480).
Animal xenograft models
Female BALB/c nude mice (mean weight ± SD, 20 ± 0.5 g; age, 6 week [Department of Animal Center, Shanghai]) were used to establish subcutaneous transplanted human NSCLC xenografts (PC9, H1975, H358, H520) for SPECT imaging and biodistribution. After anesthetization with 2% isoflurane in O2, 5 × 106 cells were injected in the right shoulder. The tumor volume was observed every other day until it reached 250 mm3. The mice were maintained on a standard diet, bedding, and specific pathogen free environments. All animal experiments were approved by the Harbin Medical University Animal Ethics Committee in accordance with Chinese legislation and were performed following their guidelines. The institutional review board approved this study in mice with human tumor xenografts.
SPECT imaging
SPECT scans and image analyses were done using a clinical SPECT scanner (Siemens e.cam Gamma Camera). Each PC9, H1975, H358 and H520 tumor-bearing mouse was injected in a tail vein with 7.4 MBq (200 μCi in 100 uL) of 99mTc-HYNIC-MPG under isoflurane (2% in O2) anesthesia (n = 3 per group). SPECT scan was then performed at 1 h, 2 h, 4 h and 6 h time points. The mice were located in the center field view of the detector. The imaging procedure was completed with a static anterior image (5 min) and with a zoom factor of 1.00 and digitally stored at 1024 × 1024. The energy window setting was 126 keV to 154 keV and nuclide-99mTc. For blocking studies, 100 mg/kg PD153035 was injected in PC9 xenografts 1h before SPECT imaging, and then SPECT scan was performed as described above. The ratio of tumor to muscle (T/M) was calculated over the region of interest.
Ex vivo biodistribution
An average dose of 0.925 MBq (25 μCi) of 99mTc-HYNIC-MPG was injected into xenografts of all NSCLC cell lines via the lateral tail vein. The mice were sacrificed at 1 h, 2 h, 4 h and 6 h time points. Brain, heart, lung, liver, tumor, bone, kidney, stomach, intestine, blood, spleen, skin and muscle were collected immediately. The samples were weighed in wet-weight and the radioactivity of each sample was measured by a gamma-counter (PerkinElmer 2480). After corrected decay of all measurements, the radioactivity was expressed as percentage of injected dose per gram (% ID/g).
Western blot assay
After SPECT imaging, the xenografts were sacrificed immediately and the tumor tissues were harvested. The tumor tissues were extracted in modified RIPA buffer on cold ice and the concentration of each sample was measured with a BCA protein assay kit (Pierce Biotechnology, Inc). Total proteins were loaded and separated by SDS-PAGE (10%), transferred to a nitrocellulose membrane and then incubated with 5% non-fat milk blocking buffer at room temperature for 30min. The membrane was then incubated at 4°C overnight with primary antibodies including EGF Receptor(D38B1) Rabbit mAb antibody or EGF Receptor (E746-A750 Specific) Rabbit mAb antibody, which were purchased from Cell Signaling Technology and this was followed by incubation with anti-rabbit IgG HRP-linked antibody (Cell Signaling Technology) at room temperature for 1 hour. The bands were detected by an ECL western blotting detection system (BD).
Immunofluorescence staining
Immunofluorescence staining was performed to determine the total EGFR and EGFR exon 19 E746-A750 deletion expression levels. Tumor tissues were frozen overnight at −80°C, then cut into 5 μm per slice, fixed with 4% cold paraformaldehyde (PFA) for 30 min and washed three times in PBS, then blocked with 10% normal goat serum for 1 h. The slices were then incubated with EGF Receptor (D38B1) Rabbit mAb antibody (1:50, Cell Signaling Technology) or EGF Receptor (E746-A750 Specific) Rabbit mAb antibody (1:100, Cell Signaling Technology) at 4°C overnight, followed by incubation with Goat anti-Rabbit IgG (H + L) or Alexa Fluor® 594 conjugate (1:1000, ThermoFisher) secondary antibodies and washed five times with PBS buffer. The slices were mounted with 49-6-diamidino-2-phenylindole (DAPI)-containing mounting medium and observed in a confocal microscope (C2Si, Nikon).
Statistical analysis
In our study, all data are expressed as mean ± SD. GraphPad Prism 5 software was used to make statistical analysis. Statistical differences between the groups were compared by using one-way ANOVA and t test. It was considered significant if p <0.05.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
FUNDING
The costs of publication of this article were defrayed in part by the payment of page charges. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC sections 1734. This work was supported, in part, by the National Basic Research Program of China (2015CB931800), National Natural Science Foundation of China (81130028, 31210103913, 81471724, 81101088), Natural Science Foundation of Heilongjiang Province of China (LC2013C26), Innovation Fund Designated of Harbin (2014RFQGJ011), Heilongjiang Province Department of Education Science and Technology Research Projects (12521184), the Youth Science WU LIANDE Foundation of Harbin Medical University (WLD-QN1119), the Fourth Hospital of Harbin Medical University Fund for Distinguished Young Scholars and the Key Laboratory of Molecular Imaging Foundation (College of Heilongjiang Province).
REFERENCES
1. Wender R, Fontham ET, Barrera E Jr, Colditz GA, Church TR, Ettinger DS, Etzioni R, Flowers CR, Gazelle GS, Kelsey DK, LaMonte SJ, Michaelson JS, Oeffinger KC, et al. American Cancer Society lung cancer screening guidelines. CA Cancer J Clin. 2013; 63:107–117.
2. Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014; 64:9–29.
3. Ramalingam SS, Owonikoko TK, Khuri FR. Lung cancer: New biological insights and recent therapeutic advances. CA Cancer J Clin. 2011; 61:91–112.
4. Pao W, Chmielecki J. Rational, biologically based treatment of EGFR-mutant non-small-cell lung cancer. Nat Rev Cancer. 2010; 10:760–774.
5. Rosell R, Bivona TG, Karachaliou N. Genetics and biomarkers in personalisation of lung cancer treatment. Lancet. 2013; 382:720–731.
6. Zhang Z, Lee JC, Lin L, Olivas V, Au V, LaFramboise T, Abdel-Rahman M, Wang X, Levine AD, Rho JK, Choi YJ, Choi CM, Kim SW, et al. Activation of the AXL kinase causes resistance to EGFR-targeted therapy in lung cancer. Nat Genet. 2012; 44:852–860.
7. Dingemans AM, de Langen AJ, van den Boogaart V, Marcus JT, Backes WH, Scholtens HT, van Tinteren H, Hoekstra OS, Pruim J, Brans B, Thunnissen FB, Smit EF, Groen HJ. First-line erlotinib and bevacizumab in patients with locally advanced and/or metastatic non-small-cell lung cancer: a phase II study including molecular imaging. Ann Oncol. 2011; 22:559–566.
8. Tang Z, Du R, Jiang S, Wu C, Barkauskas DS, Richey J, Molter J, Lam M, Flask C, Gerson S, Dowlati A, Liu L, Lee Z, et al. Dual MET-EGFR combinatorial inhibition against T790M-EGFR-mediated erlotinib-resistant lung cancer. Br J Cancer. 2008; 99:911–922.
9. Weaver Z, Difilippantonio S, Carretero J, Martin PL, El Meskini R, Iacovelli AJ, Gumprecht M, Kulaga A, Guerin T, Schlomer J, Baran M, Kozlov S, McCann T, et al. Temporal molecular and biological assessment of an erlotinib-resistant lung adenocarcinoma model reveals markers of tumor progression and treatment response. Cancer Res. 2012; 72:5921–5933.
10. Scheffler M, Kobe C, Zander T, Nogova L, Kahraman D, Thomas R, Neumaier B, Dietlein M, Wolf J. Monitoring reversible and irreversible EGFR inhibition with erlotinib and afatinib in a patient with EGFR-mutated non-small cell lung cancer (NSCLC) using sequential [18F]fluorothymidine (FLT-)PET. Lung Cancer. 2012; 77:617–620.
11. Urano Y, Asanuma D, Hama Y, Koyama Y, Barrett T, Kamiya M, Nagano T, Watanabe T, Hasegawa A, Choyke PL, Kobayashi H. Selective molecular imaging of viable cancer cells with pH-activatable fluorescence probes. Nat Med. 2009; 15:104–109.
12. Yeh HH, Ogawa K, Balatoni J, Mukhapadhyay U, Pal A, Gonzalez-Lepera C, Shavrin A, Soghomonyan S, Flores L 2nd, Young D, Volgin AY, Najjar AM, Krasnykh V, et al. Molecular imaging of active mutant L858R EGF receptor (EGFR) kinase-expressing nonsmall cell lung carcinomas using PET/CT. Proc Natl Acad Sci USA. 2011; 108:1603–1608.
13. Pal A, Balatoni JA, Mukhopadhyay U, Ogawa K, Gonzalez-Lepera C, Shavrin A, Volgin A, Tong W, Alauddin MM, Gelovani JG. Radiosynthesis and initial in vitro evaluation of [18F]F-PEG6-IPQA—a novel PET radiotracer for imaging EGFR expression-activity in lung carcinomas. Mol Imaging Biol. 2011; 13:853–861.
14. Wang JQ, Gao M, Miller KD, Sledge GW, Zheng QH. Synthesis of [11C]Iressa as a new potential PET cancer imaging agent for epidermal growth factor receptor tyrosine kinase. Bioorg Med Chem Lett. 2006; 16:4102–4106.
15. Meng X, Loo BW Jr, Ma L, Murphy JD, Sun X, Yu J. Molecular imaging with 11C-PD153035 PET/CT predicts survival in non-small cell lung cancer treated with EGFR-TKI: a pilot study. J Nucl Med. 2011; 52:1573–1579.
16. Zhang MR, Kumata K, Hatori A, Takai N, Toyohara J, Yamasaki T, Yanamoto K, Yui J, Kawamura K, Koike S, Ando K, Suzuki K. [11C]Gefitinib ([11c]Iressa): radiosynthesis, in vitro uptake, and in vivo imaging of intact murine fibrosarcoma. Mol Imaging Biol. 2010; 12:181–191.
17. Achmad A, Hanaoka H, Yoshioka H, Yamamoto S, Tominaga H, Araki T, Ohshima Y, Oriuchi N, Endo K. Predicting cetuximab accumulation in KRAS wild-type and KRAS mutant colorectal cancer using 64Cu-labeled cetuximab positron emission tomography. Cancer Sci. 2012; 103:600–605.
18. Yan Y, Xiao ZY, Song Y, Kang ZT, Wang P, Sun XL, Shen BZ. (9)(9)mTc-HYNIC-MPG: a novel SPECT probe for targeting mutated EGFR. Bioorg Med Chem Lett. 2015; 25:1647–1652.
19. Punnoose EA, Atwal S, Liu W, Raja R, Fine BM, Hughes BG, Hicks RJ, Hampton GM, Amler LC, Pirzkall A, Lackner MR. Evaluation of circulating tumor cells and circulating tumor DNA in non-small cell lung cancer: association with clinical endpoints in a phase II clinical trial of pertuzumab and erlotinib. Clin Cancer Res. 2012; 18:2391–2401.
20. Su X, Cheng K, Jeon J, Shen B, Venturin GT, Hu X, Rao J, Chin FT, Wu H, Cheng Z. Comparison of two site-specifically (18)F-labeled affibodies for PET imaging of EGFR positive tumors. Mol Pharm. 2014; 11:3947–3956.
21. Burger IA, Zitzmann-Kolbe S, Pruim J, Friebe M, Graham K, Stephens A, Dinkelborg L, Kowal K, Schibli R, Luurtsema G, Maas B, Horn-Tutic M, Haerle SK, et al. First clinical results of (D)-18F-Fluoromethyltyrosine (BAY 86–9596) PET/CT in patients with non-small cell lung cancer and head and neck squamous cell carcinoma. J Nucl Med. 2014; 55:1778–1785.
22. Lin C, Ho CL, Ng SH, Wang PN, Huang Y, Lin YC, Tang TC, Tsai SF, Rahmouni A, Yen TC. (11)C-acetate as a new biomarker for PET/CT in patients with multiple myeloma: initial staging and postinduction response assessment. Eur J Nucl Med Mol Imaging. 2014; 41:41–49.
23. Vallabhajosula S, Nikolopoulou A, Babich JW, Osborne JR, Tagawa ST, Lipai I, Solnes L, Maresca KP, Armor T, Joyal JL, Crummet R, Stubbs JB, Goldsmith SJ. 99mTc-labeled small-molecule inhibitors of prostate-specific membrane antigen: pharmacokinetics and biodistribution studies in healthy subjects and patients with metastatic prostate cancer. J Nucl Med. 2014; 55:1791–1798.
24. Palmedo H, Marx C, Ebert A, Kreft B, Ko Y, Turler A, Vorreuther R, Gohring U, Schild HH, Gerhardt T, Poge U, Ezziddin S, Biersack HJ, et al. Whole-body SPECT/CT for bone scintigraphy: diagnostic value and effect on patient management in oncological patients. Eur J Nucl Med Mol Imaging. 2014; 41:59–67.

