INTRODUCTION
Prostate Cancer (PCa) is worldwide one of the most common male tumors. About 95% of prostate cancers are adenocarcinoma. PCa is characterized by an evident clinical, histological and biological heterogeneity. Clinically PCa goes from “latent” to an “aggressive” form. The “latent” form, a slow-growing tumor, appears histologically differentiated and indolent, without any symptoms. On the contrary, the aggressive form is a fast-growing tumor with a lethal progression [1, 2]. The identification of the PCa type is an important step for the optimal treatment and for the decrease of the morbidity and mortality incidence. Recently, the early stage detection is based mostly on the evaluation of the Prostate Specific Antigen (PSA) serum levels in patients. However, the most important limitation of PSA is its poor specificity. PSA concentration can be elevated also in non-malignant conditions such as prostatitis and benign prostatic hypertrophy [3]. Actually, all the parameters used in urology, PSA levels, Gleason Score classification and TNM system, are not sufficient to predict the tumor progression types [4]. Therefore, the clinicians and researchers are interested in finding more precise and sensitive biomarkers suitable for PCa diagnosis as well as prognosis and therapy. In this scenario, many data suggest a crucial role of cytokines and growth factor pathways in prostate cancer progression and in the development of therapy resistance [5–8]. A key central player of these signaling pathways is the protein Signal Transducer and Activator of Transcription 3 (STAT3) which was found constitutively activated in many cancer types including PCa [9].
STAT3 is ubiquitously expressed and plays multiple functions in normal cell [10]. In the canonical way, the latent cytoplasmic STAT3 is activated following phosphorylation of the tyrosine 705 (pY705-STAT3), in the C-terminal domain. This modification causes a conformational change in the dimers from anti-parallel to parallel and the activated STAT3 dimers translocate to the nuclear compartment [11, 12, 13]. Once in the nucleus, STAT3 regulate transcription through the binding to specific DNA response elements in the promoter regions of many target genes and moreover the nucleus can be considered to be its inactivation compartment, since STAT3 dephosphorylation at pY705 occurs mainly here [14, 15, 16]. However, recently a plethora of evidences on non-canonical STAT3 activities has emerged [17]. Severals studies suggested the presence of Stat3 in mitochondria (mitoStat3), where it acts as regulator of mitochondrial respiration and transcriptional process [18, 19] by altering ROS production. Another STAT3 non canonical activity involves unphosphorylated Stat3 (uStat3), which can also act as a nuclear transcription factor, broadening in this way the potential mechanisms involved in Stat3 signaling. These studies suggested that non-canonical Stat3 signaling, together with the well characterized, canonical Stat3 signaling pathways, may play an important role in tumor development and progression.
In addition to tyrosine phosphorylation, the STAT3 activity can be modulated by other important post-translational modifications (PTMs) [20]. The phosphorylation of Ser727 (pS727-STAT3) increases STAT3 transcriptional activity [21, 22] and it is necessary for her mitochondrial activities [23]. The Acetylation of Lys685 (acK685-STAT3), usually presents during inflammatory processes, has been reported to stabilize STAT3 dimers [24, 25]. The S-glutathionylation has been observed in response to oxidative stress [26]. Therefore, STAT3 PTMs are specific and reflect particular cell conditions and they may be implicated in PCa development and progression. Our efforts are focused on understanding the correlation between STAT3 PTMs, as well as STAT3 protein interactors and gene expression at different clinical tumor stages [27]. The protein interactors analyzed are: CBP/p300 (Histone Acetyltransferase p300 interactor), directly responsible for STAT3 acetylation [28], PDIA3/ERp57 [29] and APE1/Ref-1 [30] modulators of oxidative stress conditions. Studies were carried out using Formalin Fixed and Paraffin Embedded (FFPE) tissues, derived from radical prostatectomy of 65 patients ranging from Gleason Score 6 and Gleason Score 9. FFPE tissues are widely used as an “archival FFPE tissue banks”, since can be easily stored for years due to their inherent stability at room temperature and they have the advantage of a known patient outcome history. Therefore, these samples are invaluable source of biological information to elucidate pathological pathways or to retrieve disease-associated biomarkers.
RESULTS
Classification of analyzed PCa tissues
All human tissue samples were obtained from PCa patients in the “Policlinico Umberto I” Hospital of Sapienza University. The patient profiles including age, PSA value, Gleason Score and TNM system, were summarized in Table 1. Histological diagnosis was performed by independent certified pathologists of the hospital. Cancer tissues of each Gleason pattern and adjacent normal counterparts were collected separately from the formalin-fixed paraffin embedded (FFPE) sections using microtome. As a reference haematoxylin/eosin stained [31] sections of the tissues were histological verified by an experienced pathologist.
Table 1: Characteristic of patients classified according to Gleason Score
Groups |
Gleason6 |
Gleason7 |
Gleason8 |
Gleason9 |
|---|---|---|---|---|
Age, median |
68, 66 |
62, 5 |
64 |
62, 5 |
Preoperative PSA |
5, 55 |
11, 36 |
12, 4 |
8, 2 |
pTNM |
||||
pT3 |
9 |
3 |
8 |
1 |
pT4 |
6 |
12 |
14 |
12 |
Total |
15 |
15 |
22 |
13 |
Note: Patient profiles including median age, median PSA value, Gleason Score and TNM system.
Immunofluorescence analysis of pY705-STAT3 expression in prostate cancer tissues
Sections of sixty-five FFPE of non-neoplastic and neoplastic tissue specimens were processed for immunofluorescence staining to detect STAT3 and pY705-STAT3, according to previously established protocols [32]. This analysis revealed a similar level of pY705-STAT3 modification in all tumor samples as showed in a representative image of different specimens corresponding to a Gleason score ranging from 6 to 9 (Figure 1). In 6–7 Gleason the nuclear compartment was enriched of pY705-STAT3. However into 8–9 Gleason we observed also an evident cytoplasmatic presence of pY705-STAT3; in particular this result was directly related to the persistently activated STAT3 which plays an important role in tumor progression and malignancy [16]. As negative control, sections were stained with only secondary antibodies or with primary antibodies preincubated with different concentration of a synthetic phosphopeptide corresponding to residues flanking Tyr705 of STAT3 [33]. Results were reported in Supplementary Figure 3.
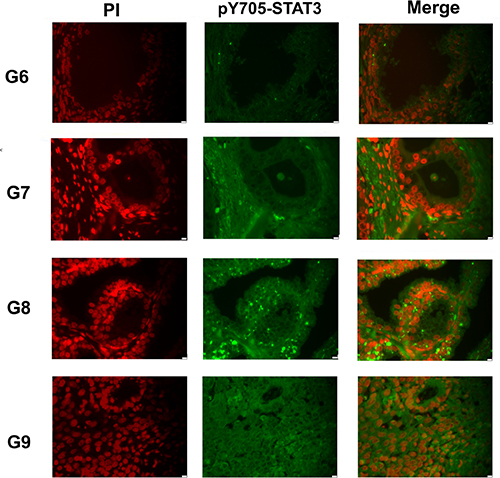
Figure 1: Analysis of pY705-STAT3 distribution in prostate cancer FFPE tissues Immunofluorescence staining of representative FFPE tissue sections with different Gleason score. pY705-STAT3: phosphorylated at tyrosine 705, PI: Propidium Iodide, G: Gleason Score, FFPE: Formalin Fixed and Paraffin Embedded.
STAT3 PTMs evaluation in PCa tissues by Western blotting analysis
The analyzed FFPE tissues had different Gleason score values. Each tumor section was compared to the normal tissue adjacent to the tumor. The number of specimens for each Gleason grade is summarized in Table 1. The levels of STAT3 and its main PTMs, were detected by immunoblotting analysis with anti-STAT3, anti-pY705-STAT3, anti-pS727-STAT3 and anti-acK685-STAT3 antibodies. As described by Robertson et al. [32], for the glutathionylation we performed a double staining with a monoclonal antibody against protein-bound GSH and anti-STAT3 antibody. The results showed a constitutively activated STAT3 (pY705-STAT3) in all tumor grades compared to the matched normal sections. A significant change between the levels of the acetylated and glutathionylated forms was present in the different clinical stages. In samples with lower Gleason Score (Gleason 6 and 7), the acK685-STAT3 is overexpressed compared to the more malignant scores. On the other hand, the pS727-STAT3 and the glutathionylated forms were relevant in higher Gleason grades. Intensity of the immunostained bands was normalized to β-actin or to the total amount of proteins in the gel revealed by coomassie staining using the ImageJ program (Figure 2). The analysis of STAT3 PTM’s profile was also performed on fresh tumoral and normal tissues samples from biopsies. Immunoblotting releaved similar STAT3 PTM’s profiles to those obtained from FFPE samples (Supplementary Figure 2), supporting the efficiency and accuracy of the data obtained from the FFPE sections.
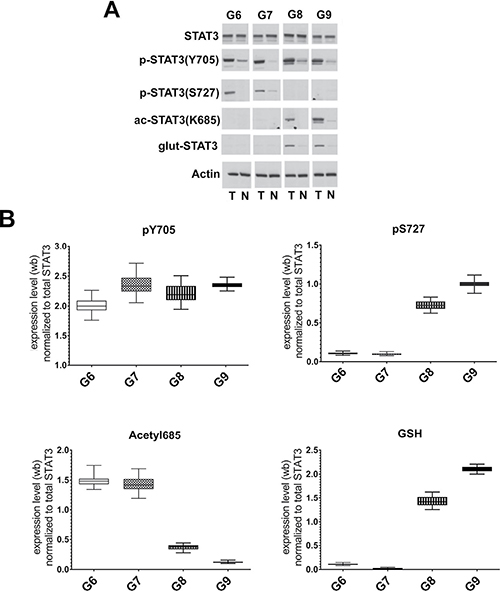
Figure 2: Analysis of STAT3 PTMs in FFPE samples with different Gleason score. (A) Representative western blot analysis of protein extracted from FFPE corresponding to Gleason Scores (G) 6, 7, 8 and 9. T: prostate carcinomas; N: normal tissue, adjacent to tumor. (B) Cumulative results of STAT3 PTMs levels in tumor prostate tissues from 65 matched prostate carcinomas. Values are presented as the means, standard deviations (boxes) and min/max values (bars). Data were analyzed by One Way Anova test and all differences between Gleason score groups in each data set were statistically significant (p < 0,01%) with the exception of pS727 G6 vs G7. pY705-STAT3: phosphorylated at tyrosine 705, pS727-STAT3: phosphorylated at serine 727, acK685-STAT3: acetylated at lysine 685, glut-STAT3 or GSH: glutathionylated STAT3.
Immunoblotting analysis of STAT3 interactors in PCA FFPE tissues: PDIA3/ERp57, CBP/p300 and APE1/REF-1
We investigated the presence and relative amount of selected STAT3 interactors, CBP/p300, APE1/Ref-1 and PDIA3/ERp57 analyzing, by immunoblotting, the protein extracts derived from patients presented in Table 1. The results showed a significant expression of the protein CBP/p300 in the lower Gleason Score (Gleason 6); on the contrary, the protein APE1/Ref-1 and PDIA3/ERp57 were overexpressed in the more malignant scores (Gleason 8 and 9 (Figure 3)). Alterations in the amount of STAT3 interactors in the analyzed samples suggest a different behaviour of STAT3 which activity could correlate with the cancer progression in the tissue.
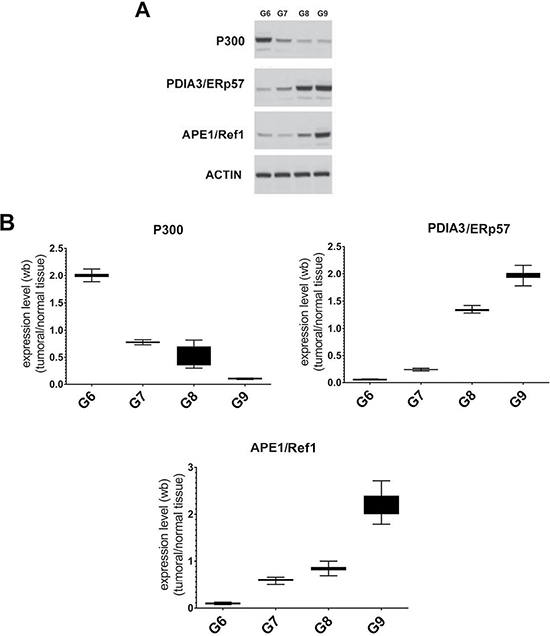
Figure 3: Analysis of STAT3 interactors in FFPE samples with different Gleason score. (A) Representative Immunoblotting of selected STAT3 protein interactors expressed in each Gleason Score. (B) Average level of expression of the selected STAT3 protein interactors matching to the same Gleason Score normalized to the corresponding normal samples. Values are presented as the means, standard deviations (boxes) and min/max values (bars). Data were analyzed by One Way Anova test and all differences between Gleason score groups in each data set were statistically significant (p < 0,01%). CBP/p300: CREB-binding protein/Histone Acetyltransferase p300 interactor. PDIA3/ERp57: Protein Disulfide-isomerase A3/Endoplasmic Reticulum Protein 57, APE1/Ref-1: Apurinic/apyrimidinic endonuclease 1/redox factor-1.
Expression of BIRC5, SOD2, SRD5A2 MMP2, PSA and CRP genes correlated to Gleason Score
We evaluated the expression of several STAT3 target genes by Quantitative Real Time-PCR in all samples. In particular, we focused attention to the levels of BIRC5, PSA, SRD5A2, MMP2 CRP and SOD2 genes, known to be involved in PCa initiation and progression [38–41], and associated with inflammatory [42] and oxidative stress conditions. The results showed an increase of the MMP2, BIRC5, CRP and SOD2 expression in samples characterized by high Gleason Score. On the contrary, as described in literature [43, 44], we saw a sharp decrease of the SRD5A2 expression in Gleason Score 9. The trend of PSA expression levels was variable among the different tissues (Figure 4).
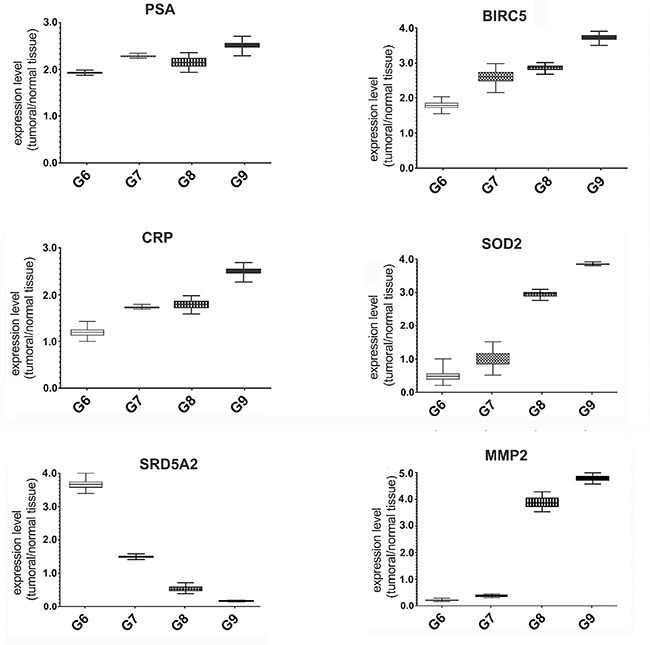
Figure 4: Expression analysis of STAT3 dependent genes in FFPE samples with different Gleason score. Quantitative RealTime-PCR for PSA, BIRC5, CRP, SOD2, SRD5A2 and MMP2 genes in FFPE tissues with different Gleason Score. Each gene expression value was normalized to housekeeping gene (β-actin) and to a control sample (normal matching tissue). Values are presented as the means, standard deviations (boxes) and min/max values (bars). Data were analyzed by One Way Anova test and all differences between Gleason score groups in each data set were statistically significant (p < 0,01%). PSA: Prostate Specific Antigen, BIRC5: Baculoviral Inhibitor of Apoptosis Repeat-containing 5, CRP: C-reactive Protein, SOD2: Superoxide Dismutase 2, SRD5A2: Steroid-5-Alpha-Reductase, Alpha Polypeptide 2, MMP2: Matrix Metalloproteinase-2.
DISCUSSION
Emerging evidences revealed that cancer progression is largely regulated by protein PTMs [45, 46]. In particular, in molecular hub proteins for signaling pathways, like STAT3, PTMs may influence gene regulation, cellular functions, tissue development, diseases, malignant progression and drug resistance [47, 48]. Here, we investigated the specificity of STAT3 PTMs in different PCa Gleason Score tissues and their influences in gene expressions. STAT3, activated by phosphorylation on tyrosine 705, was shown to be associated with the malignant transformation of prostatic epithelial cells [49]. However, beside tyrosine 705 phosphorylation, other post-translational modifications have been proven to be essential for the pleiotropic functions of STAT3 [50]. Among these, acetylation of Lysine 685 by the mammalian histone acetyltransferase complex CBP/p300, has been reported to increase the transcriptional activity stabilizing STAT3 homodimers. In addition this PTM is known to be involved in inflammatory processes activated by cytokines [24, 51]. pS727-STAT3 is known to influence the protein through canonical and non canonical mechanisms of activation [52]. In the canonical way, which first requires tyrosine phosphorylation, pS727-STAT3 induces the maximal transcriptional activity of the protein [53]. In the non canonical way, STAT3 phosphorylated only on Serine 727 residue regulates its mithocondrial translocation [23]. Mithocondrial STAT3 interacts with respiratory complexes I, II, and V of the electron transport chain and influences ROS production, thus promoting tumor cell survival and invasion [54]. Several reports have highlighted that STAT3 is also a redox-sensitive protein and may be controlled by S-glutathionylation, a post-translational protein modification regulated by intracellular redox state [26, 55]. Literature data demonstrated that an increase in glutathionylated proteins is always associated with oxidative stress in vitro and in vivo [56]. The identification of PTMs involved in STAT3 regulation will also include the characterization of proteins correlated with this cellular processes, such as writers, readers, modifiers and erasers. In particular, during oxidative stress conditions, some STAT3 protein partners, such as PDIA3/ERp57 and APE1/Ref-1, are highly expressed. Other researchers confirmed the presence of these two proteins in prostate cancer [57, 58] and it has been reported an interaction between them [59]. The protein ERp57, also known as GRP58/PDIA3, is a stress-responsive protein and is a member of the protein disulfide isomerase family. It has been recognized as a multifunctional chaperone that regulates proper folding and the quality control of glycoproteins. Although PDIA3/ERp57 has been characterized by its functions in the endoplasmic reticulum (ER), many evidences indicate that PDIA3/ERp57 is also involved in a variety of functions in the cytosol and nucleus. Several reports suggested that PDIA3/ERp57 is associated with tumor progression and the modulation of STAT3 activity [60, 61]. Apurinic/apyrimidinic endonuclease/redox factor-1 (APE1/Ref-1) is a multifunctional protein that, in addition to its base excision DNA repair activity, exerts a redox control on cancer-associated transcription factors, including STAT3 [62]. Our studies, carried out on sixty-five FFPE tissues, confirmed the constitutively activation of STAT3 protein (pY705-STAT3) in tumor samples, without any relevant variation in expression levels between the Gleason grades. On the contrary, there is a significant increase of the acK685-STAT3 in the low Gleason Scores. The result was correlated with the higher presence of the histone acetyltransferase CBP/p300 interactor in these tissues. On the other hand, pS727-STAT3 and the glutathionylated-STAT3 forms were highly expressed in the most malignant tumor (Gleason 8 and 9) together with an increase in the APE1/Ref-1 and PDIA3/ERp57 levels which are known to be present under oxidative stress [63]. Quantitative Real Time-PCR in FFPE tissues confirmed our hypothesis. In this experiment we looked at the expression of specific STAT3 target genes involved in PCa. The PSA gene, as expected, was always overexpressed between the different tissue grades and its values were a reflection of the pY705-STAT3 levels, sustaining the fact that its expression doesn’t follow an universal value, but it is variable and differs among patients. This observation, according to the data reported in the Table 1, highlights the absence of correlation between PSA values and the Gleason Score. SOD2 gene expression gradually increased from low to high Gleason Scores, responding to an oxidative state [64]. Similarly, BIRC5, an anti-apoptotic protein, and CRP gene, known to be involved in response to inflammation [36], rise its expression with increasing Gleason score, as reflection of clinical-pathological patient conditions. These findings are correlated with the overexpression of the pS727-STAT3 in the aggressive forms, as responsible modification for the mitochondrial translocation and for the equilibrium of the ROS species. The SRD5A2 gene expression profile showed a downregulation in Gleason Score 9, related to the progression of an aggressive hormone-independent form. The MMP2 gene expression followed exactly the opposite trend of SRD5A2, and this was consistent with data described also by other researches, where exogenous expression of SRD5A2 reduces the cell migration/invasion in cancer cells. In conclusion, and in agreement with the literature, tumor microenvironment can modify different cellular events and promote tumor progression. STAT3, as a signal mediator protein, reflects these particular tumor stages through its PTMs. Here, we demonstrated that inflammatory processes are predominant in low Gleason grades as confirmed by the presence of STAT3 acetylated and CBP/p300 interactor [65]. This modification seems related to the regulation of MMP2 and SRD5A2 gene expression; Sestito et al. [66] demonstrated that SIRT1 was responsible for STAT3 deacetylation and, according with our results, Lovaas et al. [61] showed that SIRT1 induced an increase of MMP2 gene expression, thus promoting tumor invasion. In parallel, other evidences, described the control of SRD5A2 expression by inflammatory mediators (IL-6 and TNF-a) which upregulates DNMT1. The latter is responsible for the SRD5A2 promoter methylation. This epigenetic event, that cause the inhibition of SRD5A2 expression, is typically associated with the development of the hormone resistant form in prostate cancer. On the other hand, the occurrence of the pS727-STAT3 and the glutathionylated STAT3 forms [56, 68] as well as the increase of ERp57 protein suggests an oxidative stress environment in the higher Gleason Score tissues. In this scenario, the STAT3 PTMs profile and the presence and relative amount of its interactors in PCa might enable clinicians to make rational decisions concerning prognosis: the currently diagnosic elements, PSA levels, Gleason Score classification and TNM system, are not sufficient to predict which tumor will develop the aggressive form or stay indolent. For many years prostatectomy was the standard of care for PCa patients followed by a lower life quality. However the era of personalized medicine has arrived and is shaping therapy in several solid tumors. The need for individual treatment in prostate cancer is particularly notable because the disease is biologically and clinically heterogeneous. The correlation between STAT3 PTMs and tumor grades would be an additional biomarker for the exactly evaluation of the PCa stages and the optimal treatment options: Watchful Waiting for the indolent tumor, prostatectomy or hormone therapy for the aggressive ones [69]. In addition, because of their important role in cellular regulation, STAT3 PTMs might be important targets in cancer as adjuvants in chemiotherapy.
MATERIALS AND METHODS
Patients and tumor samples
Formalin fixed and paraffin-embedded (FFPE) blocks corresponding to PCa patients were retrieved from the archives of Pathology Laboratory “Department gynecological-obstetric and urological sciences” University “Sapienza” Rome, according to the following criteria: radical prostatectomy specimens with no previous treatment for PCa (including androgen deprivation therapy or chemotherapy prior to surgery). We obtained sixty-five PCa specimens during the period between 2008 and 2014, stratified in four groups by Gleason score. Patients and tissue data are summarized in Table 1. All patients gave written informed consent, and the study was approved by the Ethics Committee of our institution. Immunohistochemistry and Immunofluorescence staining
FFPE sections (5 μm), cut from each prostatectomy tissue block, were stained with hematoxylin/eosin and examined to localize normal and tumor sites (Supplementary Figure 1) [31]. Sections were subjected to immunofluorescence analysis [33]. Briefly, FFPE tumor and normal sections were deparaffinized in xylene and rehydrated through graded ethanol. The slides were subjected to heat-induced antigen retrieval in citrate buffer for 15 minutes. The sections were incubated with primary rabbit polyclonal antibody overnight at 4°C (p-Y705-STAT3 Cell Signaling Technology, diluited 1:200). After PBS washings, the incubation with secondary antibody FITC conjugated goat-anti-rabbit-IgG (Jackson Immunoresearch diluited 1:500) was performed for 1h at 25°C. The sections also were counterstained with propidium iodide (0, 3 μg/ml) for 10 minutes and mounted with glass coverslips. As negative controls, the sections, pretreated with 1% BSA, were incubated with secondary antibodies for 1h at room temperature. Additionally, primary antibodies were preincubated with a synthetic phosphopeptide (PpYLKTK, Sigma Aldrich) corresponding to residues flanking Tyr705 of STAT3 [33]. Stained sections were analyzed and photographed using a fluorescence microscope (Leica AF6000 Modular System) with 63× oil immersion objective. Scale bars 10 μm in magnifications.
Protein extraction from FFPE sections
Normal and tumor FFPE sections (10 μm) were weighed (3 sections approximately equals to 30 mg) [34, 35]. The deparaffinization was carried out incubating twice with xylene for 5 minutes at room temperature before rehydration in a graded series of ethanol (100%, 90% and 70%) for 10 min each one. Vials containing the deparaffinized sections were put in a vacuum dryer for 15 minutes. Afterwards, samples were mixed with FFPE extraction buffer (150 μl per sample), sonicated about 7–8 times for 30 seconds, incubated at 100°C for 20 minutes and 80°C for 2 hours, and then centrifuged for 30 min at 14,000 rpm. The supernatant was supplemented with thiourea 100 mM and stored at −20°C. Protein concentrations were determined by the BCA assay (Bio-Rad, Munich, Germany).
Extraction of total RNA from FFPE sections
Total RNA was extracted from three 10 μm thick FFPE sections (corresponding about 30 mg of tissue). RNA was purified with WAXFREE™ paraffin kit (TrimGen) according to the manufacturer’s instructions. RNA was quantified spectrophotometrically, and its quality was assessed by 1.5% agarose gel electrophoresis and staining with ethidium bromide. To rule out the possibility of DNA contamination, the extracted RNA was incubated with 10 μm/ml RNase-free DNase at 37°C for 30 min.
Reverse transcription and quantitative real time-PCR
The reverse transcription was carried out with PrimeScript™ RT reagent Kit (Takara). Gene expression was evaluated with specific primers for PSA, BIRC5, SRD5A2, MMP2, CRP and SOD2 (Quanti-Tect QIAGEN) by Real-Time PCR using a MJ MiniOpticon Detection System (BioRad Laboratories) with SYBR green fluorophore using Brilliant SYBR Green QPCR Master Mix (Stratagene).
Western blot analysis
Western blot analysis was carried out on proteins extracted from tumor and control regions of FFPE tissues (15–20 μg protein) resolved by 10% SDS-PAGE and transferred to nitrocellulose membranes [30]. The membranes were blocked with 3% nonfat dried milk in Tris-buffered saline containing 0.05% Tween-20 (TBS-T) and incubated with the desired primary antibody for 1h. Subsequently, the membranes were washed three times in T-TBS, and bound antibodies were detected using the appropriate horseradish peroxidase-conjugated secondary antibody (1 h) [37], followed by an ECL PlusWestern blotting Detection System (GE Healthcare Bio-Sciences). ECL was detected using a Molecular ImagerR ChemiDoc™ mod. MP System (Bio-Rad Laboratories), acquired by ImageLab Software ver. 4.1, and quantified using ImageJ analysis software (http://rsbweb.nih.gov/ij/). Measured protein abundances were normalized to actin amounts and the tumor/control abundance ratio for each protein was used for analysis. The ratios of tumor to control tissue values were used to correct for possible inter-individual differences in protein abundance not related to PCa. Immunodetection was carried out using polyclonal antibodies (Cell Signaling Technology; antibody dilution 1:1000) against STAT3 (cat. n. 9132), pY705-STAT3 (cat. n. 9131), pS727-STAT3 (cat. n. 9134) and acK685-STAT3 (cat. n. 2523). At least three experimental replicates were performed for each biological sample.
Statistical analysis
The repeatability of results was confirmed by performing all experiments at least three times, and values are presented as the means, standard deviations (boxes) and min/max values (bars). Statistical analyses were performed with GraphPad software using a one-way ANOVA test followed by Bonferroni post-hoc test to analyze differences between Gleason groups.
Abbreviations
PCa : Prostate Cancer; PSA: Prostate Specific Antigen; FFPE: Formalin Fixed and Paraffin Embedded; STAT3: Signal Transducer and Activator of Transcription; PTM: Post Translational Modification; pY705-STAT3: phosphorylation of tyrosine 705; pS727-STAT3: phosphorylation of serine 727; acK685-STAT3: acetylation of lysine 685; CBP/p300: CREB-binding protein/Histone Acetyltransferase p300 interactor; PDIA3/ERp57: Protein Disulfide-isomerase A3/Endoplasmic Reticulum Protein 57; APE1/Ref-1: Apurinic/apyrimidinic endonuclease 1/redox factor-1; FITC: Fluorescein isothiocyanate; TRITC: Tetramethylrhodamine; PI: Propidium Iodide; PBS: Phosphate Buffered Saline; BIRC5: Baculoviral Inhibitor of Apoptosis Repeat-containing 5; SRD5A2: Steroid-5-Alpha-Reductase, Alpha Polypeptide 2; MMP2: Matrix Metalloproteinase-2; CRP: C-reactive Protein; SOD2: Superoxide Dismutase 2; ROS: Reactive Oxygen Species; SIRT1: Sirtuin 1; DNMT1: DNA (cytosine-5)-methyltransferase 1; qRT-PCR: Quantitative Real Time - Polymerase Chain Reaction; SDS-PAGE: Sodium Dodecyl Sulphate - PolyAcrylamide Gel Electrophoresis; TBS: Tris-buffered Saline; ECL: Enhanced Chemiluminescence.
ACKNOWLEDGMENTS
This paper is dedicated to the memory of our beloved student, Dott.ssa Stefania Gasbarrone, for unforgettable passion and commitment in study and research. We are grateful to Professor Carlo Turano (Sapienza University of Rome, Italy) for helpful discussions.
CONFLICTS OF INTEREST
The authors have declared that no competing interests exist
GRANT SUPPORT
This work has been partly supported by Istituto Pasteur Italia - Fondazione Cenci Bolognetti, Credito Cooperativo Cassa Rurale ed Artigiana di Paliano (to ME), Deutsche Bank Frosinone, Italy (to ME), Fondazione Enrico ed Enrica Sovena and Regione Lazio prog.FILAS-RU-2015. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
REFERENCES
1. Silberstein JL, Pal SK, Lewis B, Sartor O. Current clinical challenges in prostate cancer. Transl Androl Urol. 2013; 122–36.
2. Ziaee S, Chu GC, Huang JM, Sieh S, Chung LW. Prostate cancer metastasis: roles of recruitment and reprogramming, cell signal network and three-dimensional growth characteristics. Transl Androl Urol. 2015; 4:438–54.
3. Chandrasekar T, Yang JC, Gao AC, Evans CP. Mechanisms of resistance in castration-resistant prostate cancer (CRPC). Transl Androl Urol. 2015; 4:365–80.
4. Tabayoyong W, Abouassaly R. Prostate Cancer Screening and the Associated Controversy. Surg Clin North Am. 2015; 95:1023–39.
5. Pinsky PF, Andriole G, Crawford ED, Chia D, Kramer BS, Grubb R, Greenlee R, Gohagan JK. Prostate-specific antigen velocity and prostate cancer gleason grade and stage. Cancer. 2007; 109:1689–95.
6. Ramsay AK, Leung HY. Signalling pathways in prostate carcinogenesis: potentials for molecular-targeted therapy. Clin Sci (Lond). 2009; 117:209–28.
7. Tindall EA, Severi G, Hoang HN, Southey MC, English DR, Hopper JL, Giles GG, Hayes VM, Australian Prostate Cancer BioResource. Interleukin-6 promoter variants, prostate cancer risk, and survival. Prostate. 2012; 72:1701–7.
8. Rybak AP, Bristow RG, Kapoor A. Prostate cancer stem cells: deciphering the origins and pathways involved in prostate tumorigenesis and aggression. Oncotarget. 2015; 6:1900–19. doi: 10.18632/oncotarget.2953.
9. Yuan J, Zhang F, Niu R. Multiple regulation pathways and pivotal biological functions of STAT3 in cancer. Sci Rep. 2015; 5:17663.
10. Bishop JL, Thaper D, Zoubeidi A. The Multifaceted Roles of STAT3 Signaling in the Progression of Prostate Cancer. Cancers (Basel). 2014; 6:829–59.
11. Zhong M, Henriksen MA, Takeuchi K, Schaefer O, Liu B, ten Hoeve J, Ren Z, Mao X, Chen X, Shuai K, Darnell JE Jr. Implications of an antiparallel dimeric structure of nonphosphorylated STAT1 for the activation-inactivation cycle. Proc Natl Acad Sci USA. 2005; 102:3966–71.
12. Mao X, Ren Z, Parker GN, Sondermann H, Pastorello MA, Wang W, McMurray JS, Demeler B, Darnell JE Jr, Chen X. Structural bases of unphosphorylated STAT1 association and receptor binding. Mol Cell. 2005; 17:761–71.
13. Domoszlai T, Martincuks A, Fahrenkamp D, Schmitz-Van de Leur H, Küster A, Müller-Newen G. Consequences of the disease-related L78R mutation for dimerization and activity of STAT3. J Cell Sci. 2014; 127:1899–910.
14. Carpenter RL, Lo HW. STAT3 Target Genes Relevant to Human Cancers. Cancers (Basel). 2014; 6:897–925.
15. Fagard R, Metelev V, Souissi I, Baran-Marszak F. STAT3 inhibitors for cancer therapy: Have all roads been explored? JAKSTAT. 2013; 2:e22882.
16. Herrmann A, Vogt M, Mönnigmann M, Clahsen T, Sommer U, Haan S, Poli V, Heinrich PC, Müller-Newen G. Nucleocytoplasmic shuttling of persistentlyactivated STAT3. J Cell Sci. 2007; 120:3249–61.
17. Srivastava J, DiGiovanni J. Non-canonical Stat3 signaling in cancer. Mol Carcinog. 2015; 55:1889–1898. doi: 10.1002/mc.22438.
18. Demaria M, Giorgi C, Lebiedzinska M, Esposito G, D’Angeli L, Bartoli A, Gough DJ, Turkson J, Levy DE, Watson CJ, Wieckowski MR, Provero P, Pinton P, et al. A STAT3-mediated metabolic switch is involved in tumour transformation and STAT3 addiction. Aging (Albany NY). 2010; 2:823–42. doi: 10.18632/aging.100232.
19. Macias E, Rao D, Carbajal S, Kiguchi K, DiGiovanni J. Stat3 binds to mtDNA and regulates mitochondrial gene expression in keratinocytes. J Invest Dermatol. 2014; 134:1971–80.
20. Qi QR, Yang ZM. Regulation and function of signal transducer and activator of transcription 3. World J Biol Chem. 2014; 5:231–9.
21. Wakahara R, Kunimoto H, Tanino K, Kojima H, Inoue A, Shintaku H, Nakajima K. Phospho-Ser727 of STAT3 regulates STAT3 activity by enhancing dephosphorylation of phospho-Tyr705 largely through TC45. Genes Cells. 2012; 17:132–45.
22. Waitkus MS, Chandrasekharan UM, Willard B, Tee TL, Hsieh JK, Przybycin CG, Rini BI, Dicorleto PE. Signal integration and gene induction by a functionally distinct STAT3 phosphoform. Mol Cell Biol. 2014; 34:1800–11.
23. Zhang Q, Raje V, Yakovlev VA, Yacoub A, Szczepanek K, Meier J, Derecka M, Chen Q, Hu Y, Sisler J, Hamed H, Lesnefsky EJ, Valerie K, et al. Mitochondrial localized Stat3 promotes breast cancer growth via phosphorylation of serine 727. J Biol Chem. 2013; 288:31280–8.
24. Ray S, Boldogh I, Brasier AR. STAT3 NH2-terminal acetylation is activated by the hepatic acute-phase response and required for IL-6 induction of angiotensinogen. Gastroenterology. 2005; 129:1616–32.
25. Zhuang S. Regulation of STAT signaling by acetylation. Cell Signal. 2013; 25:1924–31.
26. Butturini E, Carcereri de Prati A, Chiavegato G, Rigo A, Cavalieri E, Darra E, Mariotto S. Mild oxidative stress induces S-glutathionylation of STAT3 and enhances chemosensitivity of tumoural cells to chemotherapeutic drugs. Free Radic Biol Med. 2013; 65:1322–30.
27. Yeh JE, Frank DA. STAT3-Interacting Proteins as Modulators of Transcription Factor Function: Implications to Targeted Cancer Therapy. Chem Med Chem. 2015; 11:795–801. doi: 10.1002/cmdc.201500482.
28. Hou T, Ray S, Lee C, Brasier AR. The STAT3 NH2-terminal domain stabilizes enhanceosome assembly by interacting with the p300 bromodomain. J Biol Chem. 2008; 283:30725–34.
29. Chichiarelli S, Gaucci E, Ferraro A, Grillo C, Altieri F, Cocchiola R, Arcangeli V, Turano C, Eufemi M. Role of ERp57 in the signaling and transcriptional activity of STAT3 in a melanoma cell line. Arch Biochem Biophys. 2010; 494:178–83.
30. Ray S, Lee C, Hou T, Bhakat KK, Brasier AR. Regulation of signal transducer and activator of transcription 3 enhanceosome formation by apurinic/apyrimidinic endonuclease 1 in hepatic acute phase response. Mol Endocrinol. 2010; 24:391–401.
31. Feldman AT, Wolfe D. Tissue processing and hematoxylin and eosin staining. Methods Mol Biol. 2014; 1180:31–43.
32. Robertson D, Isacke CM. Multiple immunofluorescence labeling of formalin-fixed paraffin-embedded tissue. Methods Mol Biol. 2011; 724:69–77.
33. Turkson J, Ryan D, Kim JS, Zhang Y, Chen Z, Haura E, Laudano A, Sebti S, Hamilton AD, Jove R. Phosphotyrosyl peptides block Stat3-mediated DNA binding activity, gene regulation, and cell transformation. J Biol Chem. 2001; 276: 45443–45455.
34. Gräntzdörffer I, Yumlu S, Gioeva Z, von Wasielewski R, Ebert MP, Röcken C. Comparison of different tissue sampling methods for protein extraction from formalin-fixed and paraffin-embedded tissue specimens. Exp Mol Pathol. 2010; 88:190–6.
35. Gustafsson OJ, Arentz G, Hoffmann P. Proteomic developments in the analysis of formalin-fixed tissue. Biochim Biophys Acta. 2015; 1854:559–80.
36. Chapdelaine P, Vignola K, Fortier MA. Protein estimation directly from SDS-PAGE loading buffer for standardization of samples from cell lysates or tissue homogenates before Western blot analysis. Biotechniques. 2001; 31:478, 480, 482.
37. Gallagher S, Winston SE, Fuller SA, Hurrell JG. Immunoblotting and immunodetection. Curr Protoc Cell Biol. 2011; Chapter 6:Unit6.2.
38. De Cicco C, Ravasi L, Zorzino L, Sandri MT, Botteri E, Verweij F, Granchi D, de Cobelli O, Paganelli G. Circulating levels of VCAM and MMP-2 may help identify patients with more aggressive prostate cancer. Curr Cancer Drug Targets. 2008; 8:199–206.
39. Danilewicz M, Stasikowska-Kanicka O, Wągrowska-Danilewicz M. Augmented immunoexpression of survivin correlates with parameters of aggressiveness in prostate cancer. Pol J Pathol. 2015; 66:44–8.
40. Kryza T, Silva ML, Loessner D, Heuzé-Vourc'h N, Clements JA. The kallikrein-related peptidase family: Dysregulation and functions during cancer progression. JA Biochimie. 2016; 122:283–99.
41. Miar A, Hevia D, Muñoz-Cimadevilla H, Astudillo A, Velasco J, Sainz RM, Mayo JC. Manganese superoxide dismutase (SOD2/MnSOD)/catalase and SOD2/GPx1 ratios as biomarkers for tumor progression and metastasis in prostate, colon, and lung cancer. Free Radic Biol Med. 2015; 85:45–55.
42. Sevcenco S, Mathieu R, Baltzer P, Klatte T, Fajkovic H, Seitz C, Karakiewicz PI, Rouprêt M, Rink M, Kluth L, Trinh QD, Loidl W, Briganti A, et al. The prognostic role of preoperative serum C-reactive protein in predicting the biochemical recurrence in patients treated with radical prostatectomy. Prostate Cancer Prostatic Dis. 2016; 19:163–7. doi: 10.1038/pcan.2015.60.
43. Oliveira OL, Koff WJ, Muraro F, Santos EB, Gomes Soares DF, Trindade VM. Steroid 5-alpha reductase type 2 activity in biopsies from malignant and normal prostatic tissues. Clin Chim Acta. 2008; 391:36–40.
44. Aggarwal S, Singh M, Kumar A, Mukhopadhyay T. SRD5A2 gene expression inhibits cell migration and invasion in prostate cancer cell line via F-actin reorganization. Mol Cell Biochem. 2015; 408:15–23.
45. Shen F, Kirmani KZ, Xiao Z, Thirlby BH, Hickey RJ, Malkas LH. Nuclear protein isoforms: implications for cancer diagnosis and therapy. J Cell Biochem. 2011; 112:756–60.
46. Basak S, Lu C, Basak A. Post-translational Protein Modifications of Rare and Unconventional Types: Implications in Functions and Diseases. Curr Med Chem. 2016; 23:714–45.
47. Siveen KS, Sikka S, Surana R, Dai X, Zhang J, Kumar AP, Tan BK, Sethi G, Bishayee A. Targeting the STAT3 signaling pathway in cancer: role of synthetic and natural inhibitors. Biochim Biophys Acta. 2014; 1845:136–54.
48. Zhao C, Li H, Lin HJ, Yang S, Lin J, Liang G. Feedback Activation of STAT3 as a Cancer Drug-Resistance Mechanism. Trends Pharmacol Sci. 2016; 37:47–61.
49. Han G, Yu JY, Chen YD, Cao XL, Zhu J, Wang W, Wang XX, Zhang X, Yan JQ, Gao JP. The usefulness of phosphorylated-signal transduction and activators of transcription 3 in detecting prostate cancer from negative biopsies. Eur J Surg Oncol. 2012; 38:367–73.
50. Yu H, Lee H, Herrmann A, Buettner R, Jove R. Revisiting STAT3 signalling in cancer: new and unexpected biological functions. Nat Rev Cancer. 2014; 14:736–46.
51. Ohbayashi N, Ikeda O, Taira N, Yamamoto Y, Muromoto R, Sekine Y, Sugiyama K, Honjoh T, Matsuda T. LIF- and IL-6-induced acetylation of STAT3 at Lys-685 through PI3K/Akt activation. Biol Pharm Bull. 2007; 30:1860–4.
52. Qin HR, Kim HJ, Kim JY, Hurt EM, Klarmann GJ, Kawasaki BT, Duhagon Serrat MA, Farrar WL. Activation of signal transducer and activator of transcription 3 through a phosphomimetic serine 727 promotes prostate tumorigenesis independent of tyrosine 705 phosphorylation. Cancer Res. 2008; 68:7736–41.
53. Yokogami K, Wakisaka S, Avruch J, Reeves SA. Serine phosphorylation and maximal activation of STAT3 during CNTF signaling is mediated by the rapamycin target mTOR. Curr Biol. 2000; 10:47–50.
54. Demaria M, Camporeale A, Poli V. STAT3 and metabolism: how many ways to use a single molecule? Int J Cancer. 2014; 135:1997–2003.
55. Butturini E, Darra E, Chiavegato G, Cellini B, Cozzolino F, Monti M, Pucci P, Dell’Orco D, Mariotto S. S-Glutathionylation at Cys328 and Cys542 impairs STAT3 phosphorylation. ACS Chem Biol. 2014; 9:1885–93.
56. Gallogly MM, Mieyal JJ. Mechanisms of reversible protein glutathionylation in redox signaling and oxidative stress. Curr Opin Pharmacol. 2007; 7:381–91.
57. Basu A, Cajigas-Du Ross CK, Rios-Colon L, Mediavilla-Varela M, Daniels-Wells TR, Leoh LS, Rojas H, Banerjee H, Martinez SR, Acevedo-Martinez S, Casiano CA. LEDGF/p75 Overexpression Attenuates Oxidative Stress-Induced Necrosis and Upregulates the Oxidoreductase ERP57/PDIA3/GRP58 in Prostate Cancer. PLoS One. 2016; 11:e0146549.
58. Kelley MR, Cheng L, Foster R, Tritt R, Jiang J, Broshears J, Koch M. Elevated and altered expression of the multifunctional DNA base excision repair and redox enzyme Ape1/ref-1 in prostate cancer. Clin Cancer Res. 2001; 7:824–30.
59. Grillo C, D’Ambrosio C, Scaloni A, Maceroni M, Merluzzi S, Turano C, Altieri F. Cooperative activity of Ref-1/APE and ERp57 in reductive activation of transcription factors. Free Radic Biol Med. 2006; 41:1113–23.
60. Turano C, Gaucci E, Grillo C, Chichiarelli S. ERp57/GRP58: a protein with multiple functions. Cell Mol Biol Lett. 2011; 16:539–63.
61. Choe MH, Min JW, Jeon HB, Cho DH, Oh JS, Lee HG, Hwang SG, An S, Han YH, Kim JS. ERp57 modulates STAT3 activity in radioresistant laryngeal cancer cells and serves as a prognostic marker for laryngeal cancer. Oncotarget. 2015; 6:2654–66. doi: 10.18632/oncotarget.3042.
62. Thakur S, Dhiman M, Tell G, Mantha AK. A review on protein-protein interaction network of APE1/Ref-1 and its associated biological functions. Cell Biochem Funct. 2015; 33:101–12.
63. Hsu FN, Chen MC, Lin KC, Peng YT, Li PC, Lin E, Chiang MC, Hsieh JT, Lin H. Cyclin-dependent kinase 5 modulates STAT3 and androgen receptor activation through phosphorylation of Ser727 on STAT3 in prostate cancer cells. Am J Physiol Endocrinol Metab. 2013; 305:E975–86.
64. Che M, Wang R, Li X, Wang HY, Zheng XF. Expanding roles of superoxide dismutases in cell regulation and cancer. Drug Discov Today. 2016; 21:143–9.
65. Lee JL, Wang MJ, Chen JY. Acetylation and activation of STAT3 mediated by nuclear translocation of CD44. J Cell Biol. 2009; 185:949–57.
66. Sestito R, Madonna S, Scarponi C, Cianfarani F, Failla CM, Cavani A, Girolomoni G, Albanesi C. STAT3-dependent effects of IL-22 in human keratinocytes are counterregulated by sirtuin 1 through a direct inhibition of STAT3 acetylation. FASEB J. 2011; 25:916–27.
67. Lovaas JD, Zhu L, Chiao CY, Byles V, Faller DV, Dai Y. SIRT1 enhances matrix metalloproteinase-2 expression and tumor cell invasion in prostate cancer cells. Prostate. 2013; 73:522–30.
68. Capron C, Jondeau K, Casetti L, Jalbert V, Costa C, Verhoeyen E, Massé JM, Coppo P, Béné MC, Bourdoncle P, Cramer-Bordé E, Dusanter-Fourt I. Viability and stress protection of chronic lymphoid leukemia cells involves overactivation of mitochondrial phosphoSTAT3Ser727. Cell Death Dis. 2014; 5:e1451.
69. Swanson GP, Epstein JI, Ha CS, Kryvenko ON. Pathological characteristics of low risk prostate cancer based on totally embedded prostatectomy specimens. Prostate. 2015; 75:424–9.

