INTRODUCTION
Chromatin is organized in arrays of nucleosomes in which the core histones, H2A, H2B, H3 and H4, are arranged as an octameric core around which DNA is wrapped. The linker histones H1 bind to the linker DNA connecting adjacent nucleosomes [1]. In addition to the above major histone types, many histone variants have been found, including H2A variants H2A.Z, MacroH2A, H2A-Bbd, H2AvD, and H2A.X; H3 variants H3.3 and centromeric H3 (CenH3) [1]. The similarity between the major histone subtypes and the variants range from almost no amino acid differences to extremely divergent changes [2]. Histones are functionally conserved as indicated by their high degree of structural conservation [3]. Each histone contains a conserved C-terminal histone fold domain (HFD) and a less structured and unique amino-terminus, commonly referred to as an ‘N-tail’. Histones are highly basic proteins and the ‘N-tail’ provides the majority of the basic amino acids Arg (R) and Lys (K) to the protein. The amino-terminus of histones is subjected to a wide variety of post-translational modifications, most of which occur on the Arg and Lys residues. The combinations of modifications on Arg or Lys and the Ser/Thr residues at the amino-terminus are thought to constitute a code directing distinct structural states of chromatin [4, 5].
The minor histone variant forms can replace the corresponding major histone in the nucleosome and carry out specific functions [6, 7]. The incorporation of different species of histone variants into nucleosomes provides further differentiation and epigenetic chromatin diversity [1, 8]. The differentiation and specific functions of chromatin directed by a histone variant is especially conspicuous at centromeres, where the H3 variant, CENP-A, is assembled into specialized nucleosomes that form the foundation for the kinetochore assembly [9–11]. The crystal structure of CENP-A revealed that it is quite similar to the structure of histone H3 variants and consists of an unstructured amino terminal αN-helix, α1 helix, Loop1, β1-sheet, β2-sheet, α2 helix, and α3-helix [12]. The specific localization of CENP-A at centromeres plays a central role in proper chromosome segregation and has been linked to cell cycle timing regulation, genome stability and cancer development [13–20]. The precise and spatiotemporal localization of CENP-A in centromeric nucleosomes is mediated by a system distinct from that used for the core histone [21–25], and the HJURP (Holliday junction recognition protein) is the dedicated deposition factor [17, 26, 27]. A domain found in the HFD of CENP-A, called the CENP-A targeting domain (CATD), mediates the specific interaction of CENP-A with HJURP and to date is the only domain identified essential for CENP-A centromeric localization [28]. Some modifications in the amino-terminus of CENP-A have recently been identified as regulatory signals for its centromeric targeting and chromosome segregation [9, 13, 29, 30]. However, the role of the amino-terminus of CENP-A is poorly understood.
Histones are synthesized in the cytoplasm, and the first step of their assembly into new chromatin is their import into the nucleus from the cytoplasm. Core histones contain a nuclear localization sequence (NLS) in the amino-terminal tail and are imported into the nucleus by members of the karyopherin (Kap)/importin family [31]. It has been determined that the positive charge of the basic acid residues in the N-tail promotes NLS function [32]. CENP-A is a unique variant that possesses a distinct protein sequence at its amino-terminus, compared with the core histones, and the molecular mechanism for the nuclear import of CENP-A has not yet been deciphered.
In this study, we examined the role of the amino-terminal tail of CENP-A. We identified two motifs to be critical for CENP-A nuclear accumulation and a third motif essential for centromeric accumulation.
RESULTS
A region, R29-K53, of the amino-terminus of CENP-A is required for its nuclear accumulation
CENP-A possesses an HFD quite similar to that of the canonical histone H3, but it varies at the amino-terminus. To examine the role of the amino-terminus of CENP-A, we first generated two deletion mutants of CENP-A, Del2-29 and Del2-53, which contained deletions of G2-R29 and G2-K53, respectively (Figure 1A). The wild type and mutant proteins were fused to mCherry at the C-terminal end to facilitate visualization of their cellular localization in both 293T and HeLa cells [17, 26]. The expression of wild-type and mutant CENP-A-mCherry in 293T cells was confirmed by both CENP-A and mCherry antibody because the deletion mutant cannot be recognized by CENP-A antibody (Figure 1F). Compared to the exclusive nuclear localization of wild-type CENP-A (Figure 1B and 1D, first rows), the mutant Del2-29 localized to nucleus (Figure 1B and 1D, second rows), and the mutant Del2-53 mainly localized to the cytoplasm (Figure 1B and 1D, third rows). The results suggested that the region R29-K53 is required for CENP-A nuclear import. To further confirm this, a mutant with the region R29-K53 deleted, Del29-53, was generated (Figure 1A). The Del29-53 mutant localized to both the cytoplasm and the nucleus (Figure 1B and 1D, bottom rows). More than 100 mCherry-positive 293T or HeLa cells were examined for each group, and the localization results were plotted and compared (Figure 1C, 1E).
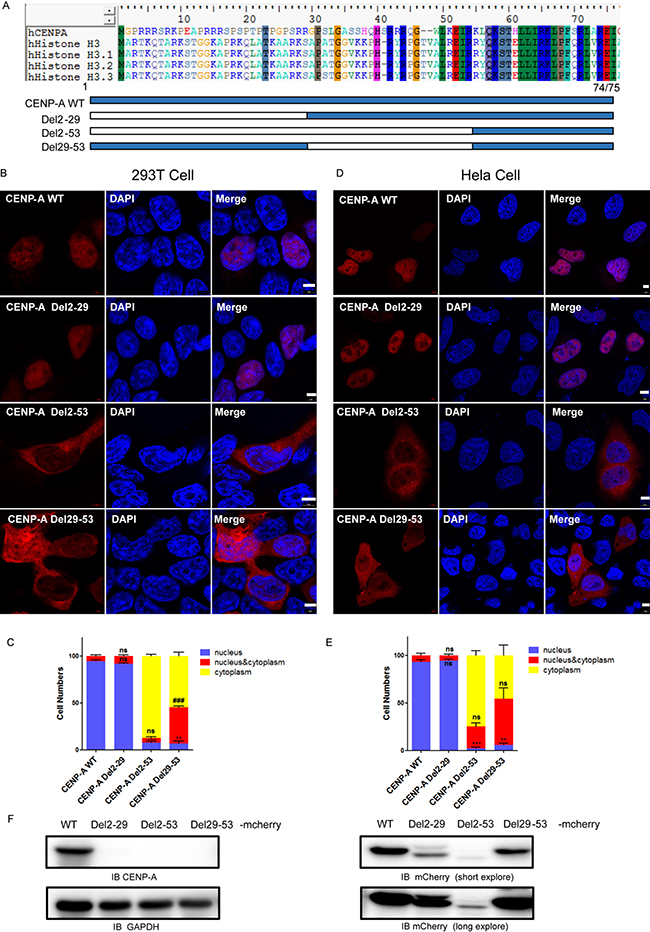
Figure 1: The nuclear localization of CENP-A requires its amino-terminus. (A) Schematic drawing showing the N-tail deletion mutants of CENP-A that we generated. (B, D) The localization of wild-type CENP-A and the mutants in 293T (B) and HeLa (D) cells. Scale bar is 5 μm. C, D, Statistical results from 100 of the 293T (C) and HeLa (E) mCherry-positive cells. Data are presented as the mean ± SEM. Nuclear localization (compared to wild type), **p < 0.01, ***p < 0.001. nucleus/cytoplasm localization (compared to wild type), ##p < 0.01, ###p < 0.001. ns, no significance. (F) Western blotting confirmed the expression of mCherry-tagged CENP-A protein in 293T cells.
Two motifs in the region R29-K53, specifically R42R43R44 and K49R52K53K56, are required for CENP-A nuclear accumulation
We examined the amino residues of the region R29-K53 (Figure 2A). There are two motifs in the region, R42R43R44 and K49R52K53K56, that contain tandem repeats of polar basic residues, typical of a protein nuclear localization signal [33]. We noticed that there was a leucine-rich motif, L60L61I62R63K64L65, which is typical of a protein nuclear export signal (NES), close to the R29-K53 region [34]. We generated three mutants, R42A R43A R44A (3A), K49A R52A K53A K56A (4A) and Del60-64, in which L60-K64 was deleted (Figure 2A). The expression of the 3A, 4A and Del60-64 mutants in 293T cells was confirmed by both CENP-A and mCherry antibodies (Figure 2F). In contrast to the exclusive nuclear localization of wild type CENP-A (Figure 2B and 2D, first rows), the mutants 3A (Figure 2B and 2D, second rows) and 4A (Figure 2B and 2D, third rows) were distributed in both the nucleus and the cytoplasm in both 293T and HeLa cells. The Del60-64 mutant (Figure 2B and 2D, bottom rows) was localized to the nucleus as efficiently as the wild-type protein. More than 100 cells for each group were examined to confirm the observation (Figure 2C and 2E). The results suggested that the motifs R42R43R44 and K49R52K53K56 are required for CENP-A nuclear accumulation, and the motif L60L61I62R63K64 is not involved in its cytoplasm/nuclear localization.
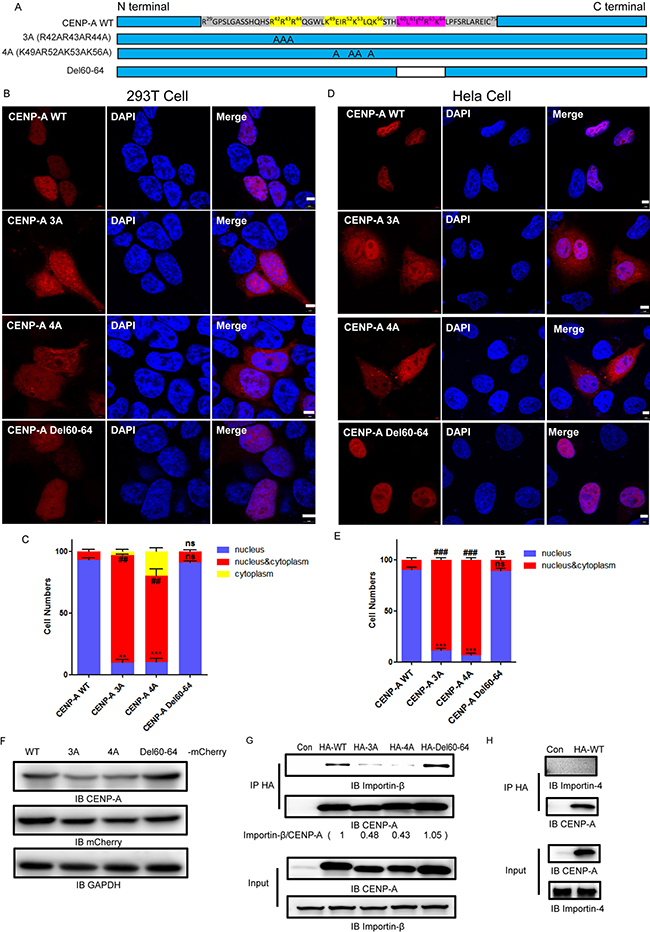
Figure 2: Two motifs, R42R43R44 and K49R52K53K56, are required for nuclear accumulation of CENP-A. (A) A schematic drawing showing the amino acid constituents of R29-C75. Three mutants, R42A R43A R44A (3A), K49A R52A K53A K56A (4A) and Del60-64, were generated. (B, D), The localization of wild-type CENP-A and mutants 3A, 4A and Del60-64 in 293T (B) and HeLa cells (D). Scale bar is 5 μm. C, (E) Statistical results from 100 mCherry-positive 293T (C) and HeLa (D) cells. Nuclear localization (compared to wild type), **p < 0.01, ***p < 0.001. nucleus/cytoplasm localization (compared to wild type), ##p < 0.01, ###p < 0.001. ns, no significance. (F) Western blotting confirmed the expression of mCherry-tagged CENP-A and mutant proteins in 293T cells. (G) CENP-A interacts with endogenous Importin-β. (H) CENP-A does not interact with Importin 4. The western blotting images were quantitated using FluorChem FC3 software.
Previous reports have suggested that the core histones are imported into the nucleus by members of the importin family [35–37]. We explored whether importins might be responsible for the nuclear import of CENP-A. Protein co-immunoprecipitation assays suggested that endogenous Importin-β (Figure 2G), but not Importin-4 (Figure 2H), co-immunoprecipitated with CENP-A. More importantly, the results showed that the interactions of mutants 3A or 4A with Importin-β were significantly reduced (Figure 2G). The mutant Del60-64 interacted with Importin-β as efficiently as the wild type protein. These results suggested that Importin-β interacts with the two motifs, R42R43R44 and K49R52K53K56, and potentially mediates the nuclear import of CENP-A.
To clarify whether the amino-tail of CENP-A is sufficient for nuclear targeting, we fused the amino-tail of CENP-A to mCherry (Figure 3A) and expressed the fusion protein in both 293T and HeLa cells (Figure 3B and 3D). More than 100 mCherry-positive 293T or HeLa cells were examined for each group, and the localization results were plotted and compared (Figure 3C and 3E). The data revealed that the amino-tail alone is not sufficient for targeting CENP-A to the nucleus and suggested that there are other elements beyond the amino-tail that are also required for CENP-A nuclear localization.
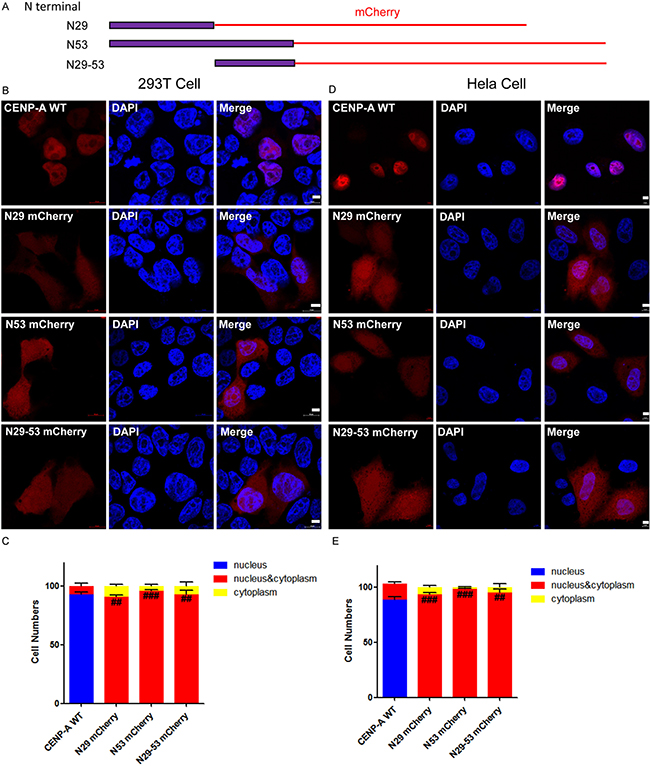
Figure 3: Evaluation of the ability of the amino-terminus of CENP-A to target CENP-A to the nucleus. (A) Different regions, the first 29, the first 53 or the 29th-53rd amino acid residues, of the amino-terminus of CENP-A were fused to mCherry (N29-mCherry, N53-mCherry or N29-53-mCherry). (B, D) Fused mCherry proteins were expressed in both 293T and HeLa cells. Scale bar is 5 μm. (C, E) Statistical results from 100 of the 293T (C) and HeLa (E) mCherry-positive cells. ##p < 0.01, ###p < 0.001 (nucleus/cytoplasm localization compared to wild-type CENP-A-mCherry).
The two motifs R42R43R44 and K49R52K53K56 of CENP-A are not involved in CENP-A localization to the centromere
We noticed that mutants Del29-53, 3A and 4A were not exclusively cytoplasmic, and there was a significant nuclear distribution of these mutated proteins. Given that CENP-A is a structural and functional component of the centromere, we asked whether these mutants are also defective in centromere targeting. To do this, we generated lentiviruses for mCherry-fused wild-type CENP-A and its mutants and infected HeLa cells to obtain stable expression of these proteins. We examined whether these mutated proteins were targeted to the centromere with both the ImageStream system and microscopy. In the imaging flow cytometer, cells in each group were classified into one of two categories, either high-spot or low-spot. In the high-spot population, the mCherry signal presented as many discrete dots, which is the typical localization pattern of a centromeric protein (Figure 4A, left panel), and in the low-spot group, less discrete dots were presented (Figure 4A, right panel). The results obtained from imaging flow cytometer suggested that G2-R29 and the two motifs R42R43R44 and K49R52K53K56 are not involved in centromeric accumulation because they behaved similarly to wild type (Figure 4B). The localization to the centromere of Del2-53 and Del29-53 was greatly impaired, primarily because they are defective in nuclear import. The cellular localization of each mutant under microscopy (Figure 4C) was consistent with that obtained from the ImageStream system. The data suggested that G2-R29, R42R43R44 and K49R52K53K56 of CENP-A are not involved in CENP-A localization to the centromere.
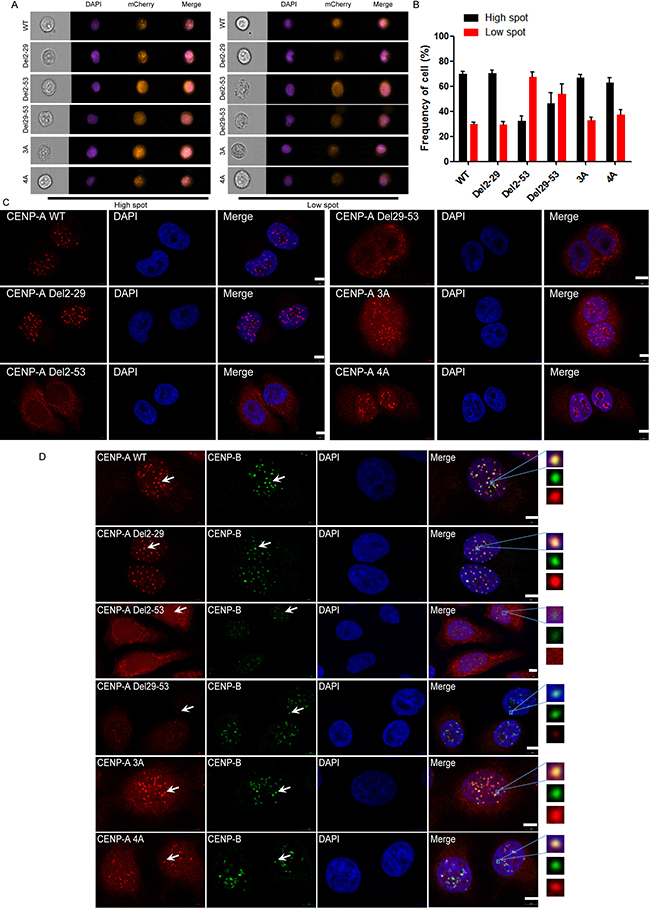
Figure 4: R42R43R44 and K49R52K53K56 were not involved in the centromeric localization of CENP-A. (A) Representative pictures of the high-spot and low-spot population of HeLa cells expressing mCherry-fused CENP-A and its mutants acquired with Amnis® ImageStreamx Mark II flow cytometry. (B) Statistical analysis of the high-spot and low-spot population of CENP-A and its mutants. (C) Centromeric localization of wild-type CENP-A and Del2-29, 3A and 4A mutant proteins. Scale bar is 5 μm. (D) Localization of the endogenous centromeric marker protein CENP-B and wild-type CENP-A, Del2-29, 3A and 4A mutant proteins. Scale bar is 5 μm.
To further confirm whether the dots were localized to the centromere, we visualized an endogenous centromeric marker, CENP-B, using immunofluorescence (Figure 4D). The dots in the mutants Del2-29, 3A and 4A co-localized very well with CENP-B, as did wild-type CENP-A (Figure 4D). There was no signal overlap with CENP-B for the mutants Del2-53 and very weak signal for the mutant Del29-53 (Figure 4D). The data suggested that the mutants Del2-29, 3A and 4A are functionally intact in regard to targeting to the centromere, and the motifs involved in CENP-A nuclear accumulation are not involved in its centromeric accumulation.
A new motif, L60-I62, is involved in CENP-A centromeric accumulation and H4 association
We found that the mutant Del60-64, in which L60L61I62R63K64 was deleted, localized to the nucleus as efficiently (Figure 2B and 2D, bottom panels) as wild-type CENP-A. A protein sequence alignment of the centromere-specific histone variants of histone H3 from different species suggested that this region is relatively conserved (Figure 5A). The mutant Del60-64 was expressed in HeLa cells to a similar level as wild-type CENP-A and a CATD mutant, subCATD, in which the CATD of CENP-A is substituted with the corresponding region of the core Histone H3 (Figure 5B). The CATD mutant, subCATD, is known to be defective in centromeric localization (Figure 5C–5E) [38]. We did detect very weak centromeric localization with the mutant Del60-64 (Figure 5C–5E). The centromeric localization of the mutant Del60-64 was greatly impaired compared to wild type. The CENP-B co-localization assay yielded the same results (Figure 5F). The data suggested that the motif L60L61I62R63K64 is required for CENP-A centromeric accumulation.
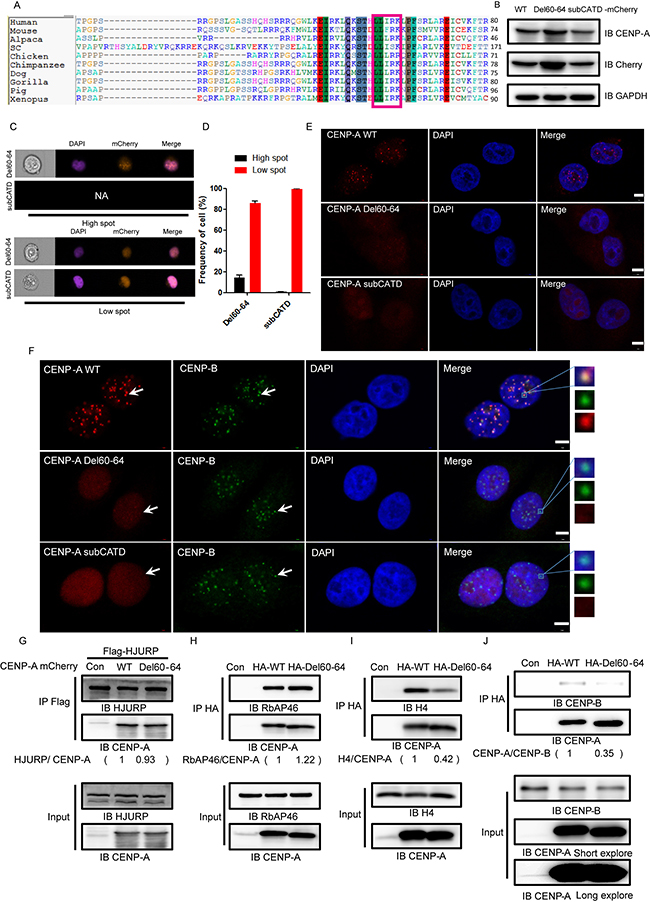
Figure 5: A new motif, L60L61I62R63K64, is required for CENP-A centromeric localization. (A) The protein sequence alignment of the centromere-specific histone variants of histone H3 from different species showing conservation of the L60L61I62R63K64 motif. (B) The expression of mCherry-tagged wild-type CENP-A and the mutant proteins Del60-64 and subCATD in HeLa cells. (C) Representative pictures of the high-spot and low-spot populations of HeLa cells expressing mCherry-fused CENP-A and mutant proteins acquired with ImageStream cytometry. (D) The statistical analysis results of high-spot and low-spot populations of CENP-A and mutants. (E) Del60-64 partially lost centromeric localization,. (F) Localization of the endogenous centromeric marker protein CENP-B and mCherry-tagged CENP-A, Del60-64, and subCATD. Scale bar is 5 μm. (G–J) The association of CENP-A or Del60-64 with HJURP (G), RbAp46 (H), Histone H4 (I) and CENP-B (J).
To investigate why the mutant Del60-64 lost its ability to localize to the centromere, we examined its interaction with HJURP and RbAp46, both of which are involved in targeting CENP-A to the centromere. We found that the interaction of Del60-64 with HJURP (Figure 5G) and RbAp46 (Figure 5H) was not affected. Interestingly, we found that its interaction with Histone H4 or with CENP-B was significantly reduced (Figure 5I and 5J). The results suggested that the motif L60L61I62R63K64 is potentially involved in the association of CENP-A with the core Histone H4.
There are two types of amino acid residues within the L60L61I62R63K64 sequence, the nonpolar and hydrophobic residues Leu/Ile and the polar basic residues Arg/Lys. We evaluated the role of these Leu/Ile and Arg/Lys residues in the centromeric localization of CENP-A. In total, 5 mutants were generated: L60A L61A I62A R63A K64A (5A), K64R, RK2QQ, RK2AA, and DelLLI (in which L60L61I62 were deleted) (Figure 6A). Each of these mutants expressed intact protein that was recognized by CENP-A and mCherry antibodies (Figure 6F). Mutating five of the residues to Ala impaired but did not abrogate the CENP-A centromeric accumulation. The polar basic residues R63 and K64 played a minor role in CENP-A centromeric accumulation because the mutation of these two residues had no effect on the centromeric localization of CENP-A (Figure 6B–6D). However, deletion of L60L61I62 (DelLLI in Figure 6B–6D) abrogated the centromeric accumulation of CENP-A. The CENP-B co-localization assay yielded the same results (Figure 6E). These results underscored the critical role of the nonpolar and hydrophobic residues L60L61I62 in this motif in CENP-A centromeric accumulation.
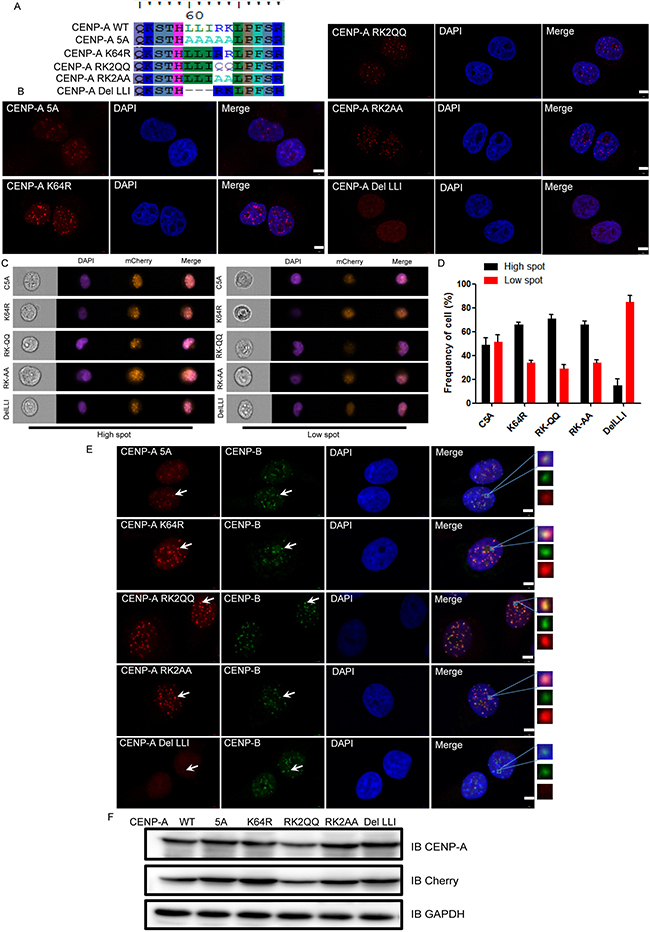
Figure 6: L60L61I62 are the most important residues for CENP-A centromeric localization within the L60L61I62R63K64 region. (A) A protein sequence alignment showing the mutants we generated within the motif L60L61I62R63K64. (B) The centromeric localization of the mutant DelLLI was greatly impaired when compared to other mutants with mutation within the region L60-K64. Scale bar is 5 μm. (C) Representative pictures of high-spot and low-spot populations of HeLa cells expressing mCherry-fused CENP-A and its mutants acquired with ImageStream cytometry. (D) The statistical analysis results of high-spot and low-spot populations of CENP-A and its mutants. (E) Localization of CENP-B and CENP-A mutants. Scale bar is 5 μm. (F) The expression of mCherry-tagged CENP-A proteins in HeLa cells.
The effect of CENP-A mutants on cell cycle and mitotic arrest
We were interested in whether mutant CENP-A has any physiological impact on cellular function. We expressed the mutant in HeLa cells and examined cell cycle progression and mitotic arrest upon the mitotic insult. Western blotting confirmed that the expression levels of HA-tagged CENP-A wild-type and mutant proteins in HeLa cells were similar (Figure 7A). The cell cycle progression was not altered upon the expression of these CENP-A mutants (Figure 7B). The cells with the expression of mutant and wild-type CENP-A responded similarly to the mitotic insult induced by the microtubule depolymerizer nocodazole (Figure 7C and 7D). Overall, the effects of the mutant CENP-A on cell cycle progression and mitotic arrest were minor in the current experimental setting. Expression of the mutant in the context of endogenous CENP-A depletion could be helpful in evaluating the effect of these mutants.
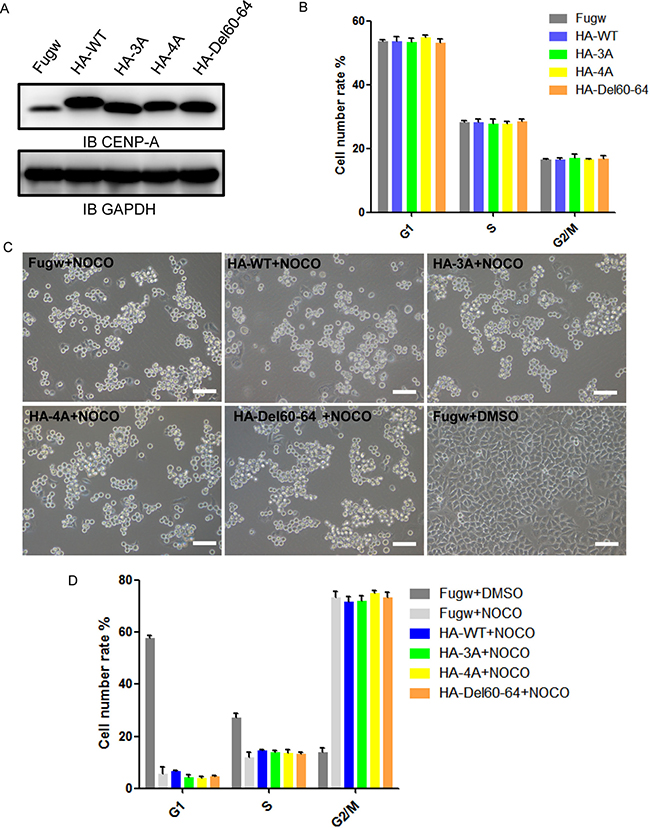
Figure 7: The effect of CENP-A mutants on cell cycle and mitotic arrest. (A) Western blotting confirmed the expression of HA-tagged wild-type CENP-A and mutant proteins in HeLa cells. (B) Cell cycle analysis of HeLa cells with the expression of wild-type CENP-A and mutant proteins for 72 hours. (C) Cells in mitotic arrest expressing wild-type CENP-A and mutants in HeLa cells. Cells were observed using microscopy after treatment with 100 ng/ml nocodazole for 18 hours. Scale bar is 100 μm. (D) The percentages of the G1, S and G2/M populations after 100 ng/ml nocodazole treatment for 18 hours were determined using PI staining and FACS analysis.
DISCUSSION
Histones and histone variants are synthesized in the cytoplasm, and nuclear import of histones is a prerequisite for the downstream deposition to form chromatin, which is important for the efficient progression of the cell cycle [39]. In yeast, it has been demonstrated that each core histone contains an NLS located at their amino terminus [35, 36]. The minimal NLS domains of H3 and H4 have been mapped to the initial residues, residues 1–28 for Histone H3 and residues 1–21 for Histone H4 [36]. The NLS of CENP-A has never been reported. We identified two motifs in CENP-A, R42R43R44 and K49R52K53K56, both of which consist of basic amino acid residues, to be critical for CENP-A accumulation in the nucleus. Distinct from the NLSs of Histone H3/H4, which are located at the very beginning of their amino-termini, the two motifs essential for CENP-A nuclear accumulation are located in the region R29-K53. The initial residues 1–28 of CENP-A are not involved in its nuclear import. To our surprise, the two motifs are required but not sufficient for CENP-A nuclear accumulation (data not shown). There are other elements beyond these two motifs that participate in the nuclear import of CENP-A. In the current structural model of the CENP-A nucleosome, these two motifs are located preceding to or in the αN helix of CENP-A, which functions in contacting DNA and stabilizing the conventional nucleosomal DNA ends in the nucleus [9, 10]. Our findings suggest that in the cytoplasm, this region of CENP-A mediates the interaction with importin for nuclear targeting [40]. This highlights the dual functions of the CENP-A amino terminus, participating in nuclear targeting in the cytoplasm and stabilizing DNA binding in the centromere nucleosome.
The motif L60L61I62 is located in the region between the αN-helix and the α1-helix, and its function has not been defined yet [10, 11, 41]. We found that this motif is critical for centromeric accumulation of CENP-A. Deletion of this motif greatly impaired the centromeric localization of CENP-A. The CATD of CENP-A is the only unique region identified to date that is required for CENP-A localization. The motif L60L61I62 is a newly discovered site allowing efficient incorporation of CENP-A into centromeric chromatin. The CATD mediates the specific interaction of CENP-A and HJURP [17, 26, 49] for centromeric targeting. However, we found that this motif does not mediate the interaction of CENP-A with its centromeric targeting molecules HJURP and RbAp46 [17, 26]. The results suggest that this motif is involved in CENP-A association with the core histone H4, which is mediated by the α2-L2-α3 segment of CENP-A. The L60L61I62 motif is a novel region at the amino-terminus of CENP-A that is potentially required for assembly of the CENP-A-H4 heterodimer. This observation is consistent with the report that the deposition of CENP-A requires formation of the CENP-A-H4 heterodimer to provide a specific recognition site for HJURP binding [11, 41].
Our study characterized the key region and residues at the amino-terminus of CENP-A that are critical for nuclear accumulation, CENP-A/H4 assembly and centromere accumulation. These findings underscore the multiple functions and importance of the flexible amino-terminus of CENP-A [9]. Our study facilitates improved understanding of the behavior of CENP-A in cells.
MATERIALS AND METHODS
Cell culture and chemical treatment
The 293T and HeLa cell lines were cultured in Dulbecco’s modified Eagle’s medium (DMEM; GIBCO, Grand Island, USA) supplemented with 10% fetal bovine serum (ExCell Bio, Shanghai, China), 100 U/ml streptomycin and 100 U/ml penicillin. Cells were cultured at 37°C in a humidified incubator under 5% CO2. For drug treatment, cells were treat with Paclitaxel or nocodazole (S1150, S2775; Selleckchem, Houston, USA) for 18 hours.
Plasmids, transfections and virus generation
The human CENP-A (Gene ID: 1058) coding sequence was amplified using PCR (E003-01A, Novoprotein) and substituted for the H2B coding sequencing in the PGK-H2BmCherry plasmid (Addgene Plasmid #21217) with the recombinase NR001A (Novoprotein). This generated an mCherry-fused CENP-A overexpression plasmid. CENP-A mutants were generated using a QuikChange kit from Agilent and verified by DNA sequencing. Transient transfections of the DNA were performed using a transfection reagent from Exelgene (USA) as described in [43]. Virus generation and the infection procedure were conducted as previously described [44–46].
Western blotting, immunofluorescence staining, immunoprecipitation and antibodies
Western blotting, immunoprecipitation and immunofluorescence were performed as previously described [44–47]. For immunoprecipitation, cell lysates were obtained by sonication in lysis buffer (50 mM Tris (pH 7.4), 150 mM NaCl, 1 mM EDTA, 1% (v/v) NP-40 alternative, and 15% (v/v) glycerol). Prior to use, the buffer was supplemented with complete protease inhibitor cocktail (Roche) and 2 mM sodium fluoride and 1 mM sodium orthovanadate (Na3 VO4, Sigma) as phosphatase inhibitors. The cell lysates were incubated with antibody and Protein A/G beads (Sigma) for 4 hours. The beads were washed with lysis buffer three times and the immunoprecipitates were analyzed by western blotting. For immunofluorescence staining, cells plated on a glass coverslip were fixed with 4% paraformaldehyde for 15 min, rinsed in PBS, treated with 0.2% Triton X-100 for 10 min, washed with PBS, blocked with 10% FBS for 1 hour, then incubated with primary antibody, and washed with PBS. Then, the cells were incubated with dye-conjugated secondary antibody and were finally mounted using mounting medium with DAPI. All the western blotting, immunoprecipitations and immunofluorescence staining were repeated three times independently, and the following antibodies were used: anti-CENP-A (Cell Signaling Technology, CST2186), anti-CENP-B, (ab25734), anti-Importin4 (ab181037), anti-Importin-β/NTF-97 (ab45938), anti-RbAP46 (Abcam, ab109285), anti-HJURP (Santa Cruz, sc-168091), anti-RbAp46 (ab28500), anti-GAPDH (ABclonal), anti-HA (Bioworld; Nanjing, China; AP0005M), anti-Flag (Sigma, F1804), and horseradish peroxidase (HRP)-conjugated secondary antibodies (Cell Signaling Technology). In addition, Alexa Fluor-conjugated secondary antibodies were obtained from Invitrogen (Eugene, OR. USA). A Leica TSC SP5 microscope was used for imaging (Leica Microsystems, Bensheim, Germany). The western blotting images were quantitated with FluorChem FC3 software. Analysis was performed using a Student’s t-test or One-way analysis of variance (ANOVA). Values of P < 0.05 were considered to be statistically significant.
Imagestream
Amnis® ImageStreamx Mark II is a multispectral flow cytometer that combines standard microscopy with flow cytometry. It acquires up to 100 cells/sec, with simultaneous acquisition of six images of each cell including bright field, scatter, and multiple fluorescence images. Cells were fixed with 70% cold-ethanol, stained with propidium iodide and subjected to Amnis® ImageStreamx Mark II flow cytometry for analysis of nuclear/cytoplasm and centromeric localization [48, 49].
Authors’ contributions
RQJ performed the experiments and wrote the paper. JJX analyzed some of the experimental results and provided technical assistance. YL, WC, GYW and WWJ contributed to the preparation of the figures. SCZ conceived and designed the study and wrote the paper. JHK coordinated the work. All authors reviewed the results and approved the final version of the manuscript.
ACKNOWLEDGMENTS AND FUNDING
This work was supported by grants obtained from the National Natural Science Foundation of China and the Ministry of Science and Technology (grant numbers 31571390, 2016YFA0101300, 2013CB967600, 2013CB967401, 81530042, 31210103905, 31371510, 31571529, 31571519, and 31471250), the Ministry of Education Grants IRT_15R51, the Science and Technology Commission of Shanghai Municipality (grant numbers 15JC1403200 and 15JC1403201), and Fundamental Research Funds for the Central Universities (2000219136, 1500219106, and 20002310002).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Kamakaka RT, Biggins S. Histone variants: deviants? Genes Dev. 2005; 19:295–310. doi: 10.1101/gad.1272805.
2. Malik HS, Henikoff S. Phylogenomics of the nucleosome. Nat Struct Biol. 2003; 10:882–91. doi: 10.1038/nsb996.
3. Ausio J. Histone variants—the structure behind the function. Brief Funct Genomic Proteomic. 2006; 5:228–43. doi: 10.1093/bfgp/ell020.
4. Turner BM. Cellular memory and the histone code. Cell. 2002; 111:285–91.
5. Cosgrove MS, Wolberger C. How does the histone code work? Biochem Cell Biol. 2005; 83:468–76. doi: 10.1139/o05-137.
6. Biterge B, Schneider R. Histone variants: key players of chromatin. Cell Tissue Res. 2014; 356:457–66. doi: 10.1007/s00441-014-1862-4.
7. Smith MM. Histone variants and nucleosome deposition pathways. Mol Cell. 2002; 9:1158–60.
8. Jin J, Cai Y, Li B, Conaway RC, Workman JL, Conaway JW, Kusch T. In and out: histone variant exchange in chromatin. Trends Biochem Sci. 2005; 30:680–7. doi: 10.1016/j.tibs.2005.10.003.
9. Roulland Y, Ouararhni K, Naidenov M, Ramos L, Shuaib M, Syed SH, Lone IN, Boopathi R, Fontaine E, Papai G, Tachiwana H, Gautier T, Skoufias D, et al. The Flexible Ends of CENP-A Nucleosome Are Required for Mitotic Fidelity. Mol Cell. 2016; 63:674–85. doi: 10.1016/j.molcel.2016.06.023.
10. Tachiwana H, Kagawa W, Shiga T, Osakabe A, Miya Y, Saito K, Hayashi-Takanaka Y, Oda T, Sato M, Park SY, Kimura H, Kurumizaka H. Crystal structure of the human centromeric nucleosome containing CENP-A. Nature. 2011; 476:232–5. doi: 10.1038/nature10258.
11. Sekulic N, Bassett EA, Rogers DJ, Black BE. The structure of (CENP-A-H4)(2) reveals physical features that mark centromeres. Nature. 2010; 467:347–51. doi: 10.1038/nature09323.
12. Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature. 1997; 389:251–60. doi: 10.1038/38444.
13. Yu Z, Zhou X, Wang W, Deng W, Fang J, Hu H, Wang Z, Li S, Cui L, Shen J, Zhai L, Peng S, Wong J, et al. Dynamic phosphorylation of CENP-A at Ser68 orchestrates its cell-cycle-dependent deposition at centromeres. Dev Cell. 2015; 32:68–81. doi: 10.1016/j.devcel.2014.11.030.
14. Niikura Y, Kitagawa R, Ogi H, Abdulle R, Pagala V, Kitagawa K. CENP-A K124 Ubiquitylation Is Required for CENP-A Deposition at the Centromere. Dev Cell. 2015; 32:589–603. doi: 10.1016/j.devcel.2015.01.024.
15. McKinley KL, Cheeseman IM. Polo-like kinase 1 licenses CENP-A deposition at centromeres. Cell. 2014; 158:397–411. doi: 10.1016/j.cell.2014.06.016.
16. Shuaib M, Ouararhni K, Dimitrov S, Hamiche A. HJURP binds CENP-A via a highly conserved N-terminal domain and mediates its deposition at centromeres. Proc Natl Acad Sci U S A. 2010; 107:1349–54. doi: 10.1073/pnas.0913709107.
17. Dunleavy EM, Roche D, Tagami H, Lacoste N, Ray-Gallet D, Nakamura Y, Daigo Y, Nakatani Y, Almouzni-Pettinotti G. HJURP is a cell-cycle-dependent maintenance and deposition factor of CENP-A at centromeres. Cell. 2009; 137:485–97. doi: 10.1016/j.cell.2009.02.040.
18. Lacoste N, Woolfe A, Tachiwana H, Garea AV, Barth T, Cantaloube S, Kurumizaka H, Imhof A, Almouzni G. Mislocalization of the centromeric histone variant CenH3/CENP-A in human cells depends on the chaperone DAXX. Mol Cell. 2014; 53:631–44. doi: 10.1016/j.molcel.2014.01.018.
19. Regnier V, Vagnarelli P, Fukagawa T, Zerjal T, Burns E, Trouche D, Earnshaw W, Brown W. CENP-A is required for accurate chromosome segregation and sustained kinetochore association of BubR1. Mol Cell Biol. 2005; 25:3967–81. doi: 10.1128/MCB.25.10.3967-3981.2005.
20. Goutte-Gattat D, Shuaib M, Ouararhni K, Gautier T, Skoufias DA, Hamiche A, Dimitrov S. Phosphorylation of the CENP-A amino-terminus in mitotic centromeric chromatin is required for kinetochore function. Proc Natl Acad Sci U S A. 2013; 110:8579–84. doi: 10.1073/pnas.1302955110.
21. Verreault A. Histone deposition at the replication fork: a matter of urgency. Mol Cell. 2003; 11:283–4.
22. Jackson V, Granner D, Chalkley R. Deposition of histone onto the replicating chromosome: newly synthesized histone is not found near the replication fork. Proc Natl Acad Sci U S A. 1976; 73:2266–9.
23. Jackson V, Chalkley R. A new method for the isolation of replicative chromatin: selective deposition of histone on both new and old DNA. Cell. 1981; 23:121–34.
24. Franco AA, Kaufman PD. Histone deposition proteins: links between the DNA replication machinery and epigenetic gene silencing. Cold Spring Harb Symp Quant Biol. 2004; 69:201–8. doi: 10.1101/sqb.2004.69.201.
25. Li Q, Burgess R, Zhang Z. All roads lead to chromatin: Multiple pathways for histone deposition. Biochim Biophys Acta. 2012; 1819:238–46. doi: 10.1016/j.bbagrm.2011.06.013.
26. Foltz DR, Jansen LE, Bailey AO, Yates JR 3rd, Bassett EA, Wood S, Black BE, Cleveland DW. Centromere-specific assembly of CENP-a nucleosomes is mediated by HJURP. Cell. 2009; 137:472–84. doi: 10.1016/j.cell.2009.02.039.
27. Sanchez-Pulido L, Pidoux AL, Ponting CP, Allshire RC. Common ancestry of the CENP-A chaperones Scm3 and HJURP. Cell. 2009; 137:1173–4. doi: 10.1016/j.cell.2009.06.010.
28. Zasadzinska E, Barnhart-Dailey MC, Kuich PH, Foltz DR. Dimerization of the CENP-A assembly factor HJURP is required for centromeric nucleosome deposition. EMBO J. 2013; 32:2113–24. doi: 10.1038/emboj.2013.142.
29. Liu RM, Tian XY, Huang XT, Zhou H. Transfer of Ser7 phosphorylated CENP-A from centromere to midbody during mitosis in MCF-7 cells. Folia Biol (Praha). 2013; 59:105–9.
30. Bailey AO, Panchenko T, Sathyan KM, Petkowski JJ, Pai PJ, Bai DL, Russell DH, Macara IG, Shabanowitz J, Hunt DF, Black BE, Foltz DR. Posttranslational modification of CENP-A influences the conformation of centromeric chromatin. Proc Natl Acad Sci U S A. 2013; 110:11827–32. doi: 10.1073/pnas.1300325110.
31. Keck KM, Pemberton LF. Histone chaperones link histone nuclear import and chromatin assembly. Biochim Biophys Acta. 2012; 1819:277–89. doi: 10.1016/j.bbagrm.2011.09.007.
32. Blackwell JS Jr, Wilkinson ST, Mosammaparast N, Pemberton LF. Mutational analysis of H3 and H4 N termini reveals distinct roles in nuclear import. J Biol Chem. 2007; 282:20142–50. doi: 10.1074/jbc.M701989200.
33. Christophe D, Christophe-Hobertus C, Pichon B. Nuclear targeting of proteins: how many different signals? Cell Signal. 2000; 12:337–41.
34. Nigg EA. Nucleocytoplasmic transport: signals, mechanisms and regulation. Nature. 1997; 386:779–87. doi: 10.1038/386779a0.
35. Mosammaparast N, Jackson KR, Guo Y, Brame CJ, Shabanowitz J, Hunt DF, Pemberton LF. Nuclear import of histone H2A and H2B is mediated by a network of karyopherins. J Cell Biol. 2001; 153:251–62.
36. Mosammaparast N, Guo Y, Shabanowitz J, Hunt DF, Pemberton LF. Pathways mediating the nuclear import of histones H3 and H4 in yeast. J Biol Chem. 2002; 277:862–8. doi: 10.1074/jbc.M106845200.
37. Muhlhausser P, Muller EC, Otto A, Kutay U. Multiple pathways contribute to nuclear import of core histones. EMBO Rep. 2001; 2:690–6. doi: 10.1093/embo-reports/kve168.
38. Black BE, Foltz DR, Chakravarthy S, Luger K, Woods VL Jr, Cleveland DW. Structural determinants for generating centromeric chromatin. Nature. 2004; 430:578–82. doi: 10.1038/nature02766.
39. Marzluff WF, Duronio RJ. Histone mRNA expression: multiple levels of cell cycle regulation and important developmental consequences. Curr Opin Cell Biol. 2002; 14:692–9.
40. Jakel S, Albig W, Kutay U, Bischoff FR, Schwamborn K, Doenecke D, Gorlich D. The importin beta/importin 7 heterodimer is a functional nuclear import receptor for histone H1. EMBO J. 1999; 18:2411–23. doi: 10.1093/emboj/18.9.2411.
41. Hu H, Liu Y, Wang M, Fang J, Huang H, Yang N, Li Y, Wang J, Yao X, Shi Y, Li G, Xu RM. Structure of a CENP-A-histone H4 heterodimer in complex with chaperone HJURP. Genes Dev. 2011; 25:901–6. doi: 10.1101/gad.2045111.
42. Bassett EA, DeNizio J, Barnhart-Dailey MC, Panchenko T, Sekulic N, Rogers DJ, Foltz DR, Black BE. HJURP uses distinct CENP-A surfaces to recognize and to stabilize CENP-A/histone H4 for centromere assembly. Dev Cell. 2012; 22:749–62. doi: 10.1016/j.devcel.2012.02.001.
43. Song C, Zhu S, Wu C, Kang J. Histone deacetylase (HDAC) 10 suppresses cervical cancer metastasis through inhibition of matrix metalloproteinase (MMP) 2 and 9 expression. J Biol Chem. 2013; 288:28021–33. doi: 10.1074/jbc.M113.498758.
44. Yan H, Zhu S, Song C, Liu N, Kang J. Bone morphogenetic protein (BMP) signaling regulates mitotic checkpoint protein levels in human breast cancer cells. Cell Signal. 2012; 24:961–8. doi: 10.1016/j.cellsig.2011.12.019.
45. Zhu S, Jing R, Yang Y, Huang Y, Wang X, Leng Y, Xi J, Wang G, Jia W, Kang J. A motif from Lys216 to Lys222 in human BUB3 protein is a nuclear localization signal and critical for BUB3 function in mitotic checkpoint. J Biol Chem. 2015; 290:11282–92. doi: 10.1074/jbc.M114.598029.
46. Yang Y, Huang Y, Wang Z, Wang HT, Duan B, Ye D, Wang C, Jing R, Leng Y, Xi J, Chen W, Wang G, Jia W, et al. HDAC10 promotes lung cancer proliferation via AKT phosphorylation. Oncotarget. 2016; 7:59388–59401. doi: 10.18632/oncotarget.10673.
47. Chen L, Wei T, Si X, Wang Q, Li Y, Leng Y, Deng A, Chen J, Wang G, Zhu S, Kang J. Lysine acetyltransferase GCN5 potentiates the growth of non-small cell lung cancer via promotion of E2F1, cyclin D1, and cyclin E1 expression. J Biol Chem. 2013; 288:14510–21. doi: 10.1074/jbc.M113.458737.
48. Scheffel MJ, Scurti G, Simms P, Garrett-Mayer E, Mehrotra S, Nishimura MI, Voelkel-Johnson C. Efficacy of Adoptive T-cell Therapy Is Improved by Treatment with the Antioxidant N-Acetyl Cysteine, Which Limits Activation-Induced T-cell Death. Cancer Res. 2016; 76:6006–16. doi: 10.1158/0008-5472.CAN-16-0587.
49. Chang SY, Song JH, Guleng B, Cotoner CA, Arihiro S, Zhao Y, Chiang HS, O'Keeffe M, Liao G, Karp CL, Kweon MN, Sharpe AH, Bhan A, et al. Circulatory antigen processing by mucosal dendritic cells controls CD8(+) T cell activation. Immunity. 2013; 38:153–65. doi: 10.1016/j.immuni.2012.09.018.

