INTRODUCTION
Metastasis is the leading cause of cancer-related death [1]. Despite the development of new effective anticancer treatments, metastases are still difficult to treat. Hence, it is crucial to develop preclinical imaging tools for monitoring the metastatic cascade and for understanding its features in vivo.
Cell tracking or cellular imaging is defined as the “non-invasive and repetitive imaging of targeted cells and cellular processes in living organisms” [2]. Applications of cellular imaging include the monitoring of stem cells in cell-based therapies and the visualization of metastatic cancer cells in small animals [3]. In magnetic resonance imaging (MRI) cell tracking studies, cells are first labeled in vitro with contrast agents prior to their injection in vivo. Superparamagnetic particles of iron oxides (SPIO) are widely used in cell tracking studies because (i) they induce signal voids on T2- and T2*-weighted images (T2(*)-w), (ii) they are biocompatible and (iii) they are easily incorporated in the cell cytoplasm [2, 4, 5].
Previous studies showed that MRI allows single cell detection on high-resolution images [6–8]. For instance, Heyn et al. reported the imaging of isolated breast cancer cells disseminated in the mouse brain and the subsequent development of macrometastases [9]. Since the SPIO contrast is diluted with cell division, previous works performed on melanoma and breast tumor models suggested that the signal void persistence indicated the presence of quiescent cancer cells, whereas regions of contrast loss indicated proliferative tumor cells [10, 11]. For these reasons, MRI offers great promises for monitoring the development of metastases of clonal origin and monitoring dormant metastatic cells [9, 10, 12]. In the clinic, dormant cancer cells are nonresponsive to therapy and are responsible for recurrences [13]. These perspectives are however challenged from other observations. In cell-based therapy approaches, MRI cell tracking experiments failed to characterize the long-term fate of stem cells. Several immunohistochemistry studies performed several weeks after cell delivery showed that macrophages take up SPIO released by dying cells. Our works previously showed that the signal void induced by SPIO-labeled cancer cells on MRI scans quickly decreased at the injection site and at metastatic locations [8, 14]. Moreover, we showed that the iron oxide content in tissues decreased with time [8, 14]. We therefore hypothesized that (i) dilution of SPIO with cell division, (ii) SPIO metabolism by macrophages recruited to the tumor site and (iii) clearance of SPIO from dead cells could explain the loss of contrast and/or the drop of SPIO content in tissues [8, 14]. Hence, the evolution of SPIO contrast may be influenced by the proliferative status but also by the phagocytic activity of tumor macrophages.
Here, we aimed at characterizing the role of macrophages in SPIO uptake and degradation in vivo. It will allow determining if MRI cell tracking can be used for assessing the proliferative status of tumor cells in vivo. MRI was performed to characterize the fate of SPIO-labeled breast cancer cells in vivo. We next questioned the in vitro fate of iron oxides after intracellular incorporation in breast cancer cells and macrophages. We took advantage of the superparamagnetic (SP) properties of these nanoparticles, and used electron paramagnetic resonance (EPR) spectroscopy for measuring superparamagnetic iron. EPR was validated in previous studies for characterizing the SPIO content of cells and tissues [14–22]. Inductively coupled plasma mass spectroscopy (ICP-MS) served for the sensitive quantification of total iron pools (SP + non-SP) [23]. Correlating both ICP-MS and EPR results provided important information on the degradation of iron oxides after SPIO labeling in breast cancer cells and macrophages.
RESULTS
Using MRI (11.7 T), we first tracked green fluorescent protein-tagged 4T1 (4T1-GFP) cells labeled with Modlay Ion Rhodamine B (MIRB) SPIO in vivo. Figure 1 shows that an intramuscular injection of SPIO-labeled 4T1-GFP cells in mice induced a strong signal void on T2*-w images, whereas the injection of control 4T1-GFP cells was not detected on MR scans. In the following days, the hypointense spot progressively disappeared (Figure 1). We also noticed the appearance of a hyperintense area in both groups that was attributed to the development of the tumor (Figure 1).
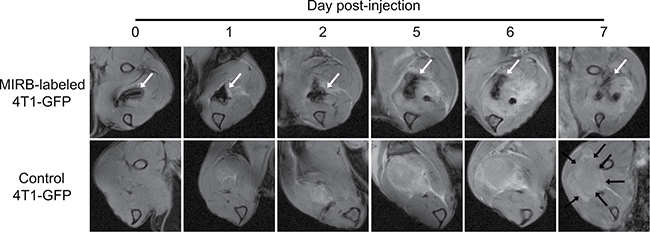
Figure 1: MRI shows that SPIO-induced signal voids progressively disappear after intramuscular injection of iron oxide-labeled 4T1 cells. Mice received an intramuscular injection of 106 MIRB-labeled 4T1-GFP cells (upper images) or 106 SPIO free 4T1-GFP cells (lower images). Longitudinal T2*-w images of MIRB-labeled 4T1-GFP cells (upper images), where the negative contrast due to the presence of SPIO-labeled cells over time is indicated by white arrows. T2*-w images of control 4T1-GFP cells (lower images), highlighting the absence of signal voids due to SPIO and the apparition of a hyperintense area due to tumor growth (black arrows) (n = 4).
We next aimed at characterizing the role of macrophages in the loss of contrast observed on MR scans. For this purpose, we next measured the evolution of SP iron content and total (SP + non-SP) iron content in 4T1-GFP cells and J774 macrophages after SPIO labeling. In the total population of MIRB-labeled 4T1-GFP breast cancer cells, SP iron levels were stable up to five days after labeling (Figure 2A, 0.67 ± 0.03 μg SP iron at day 0 versus 0.64 ± 0.07 μg SP iron at day 5, p = 0.9984). No difference in total iron levels (SP + non-SP) between groups was detected (Figure 2B, 0.70 ± 0.01 μg Fe at day 0 versus 0.51 ± 0.08 μg Fe at day 5, p = 0.53). Conversely, intracellular SP iron oxide content progressively decreased in J774 macrophages after MIRB labeling (Figure 2C, 0.64 ± 0.02 μg SP at day 0 versus 0.20 ± 0.01 μg SP iron at day 5, p < 0.001). Similarly, total (SP + non-SP) iron levels decreased in MIRB-labeled J774 cells after SPIO labeling (Figure 2D, 0.82 ± 0.15 μg iron at day 0 versus 0.26 ± 0.01 μg iron at day 5, p = 0.0031).
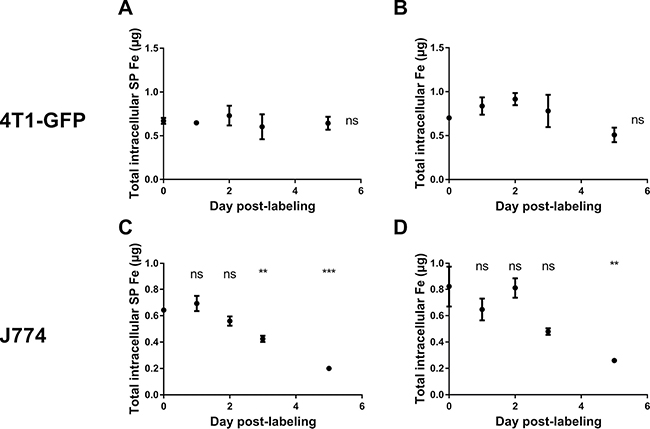
Figure 2: The superparamagnetic iron content remains constant in 4T1-GFP cells after MIRB labeling, whereas it drops in J774 macrophages. (A) The SP iron pool measured by EPR and (B) the total iron (SP + non-SP) pool measured by ICP-MS were quantified in MIRB-labeled 4T1-GFP breast cancer cells. (C) The SP iron pool measured by EPR and (D) the total iron (SP + non-SP) pool measured by ICP-MS were quantified in MIRB-labeled J774 cells. Data are expressed as means ± SEM. **p < 0.01, ***p < 0.001, ns, p > 0.05.
These in vitro experiments showed that the intracellular (SP) iron content dropped in J774 macrophages but not in 4T1-GFP cells after MIRB labeling. It suggested that macrophages in particular metabolize SPIO. Using ICP-MS, we therefore compared iron release by J774 and 4T1-GFP cells after SPIO incubation. Figure 3 shows that J774 macrophages released significant amounts of iron in the culture medium after MIRB-labeling (Figure 3, 0.31 ± 0.01 μg iron in MIRB-labeled J774 cells at day 0 cells versus 0.56 ± 0.01 μg iron at day 5, p < 0,0001). Comparatively, extracellular iron concentration only slightly increased in the 4T1-GFP + MIRB group (Figure 3, 0.52 ± 0.01 μg iron in MIRB-labeled 4T1-GFP cells at day 0 cells versus 0.62 ± 0.05 μg iron at day 5, p = 0.036). Of note, we did not detect any EPR signal coming from extracellular media of cultured 4T1-GFP + MIRB or J774 + MIRB cells (data not shown). When both total intracellular and extracellular iron levels (SP + non-SP) are pooled, no significant variation in iron content was detected in 4T1-GFP and J774 macrophages (Figure 4).
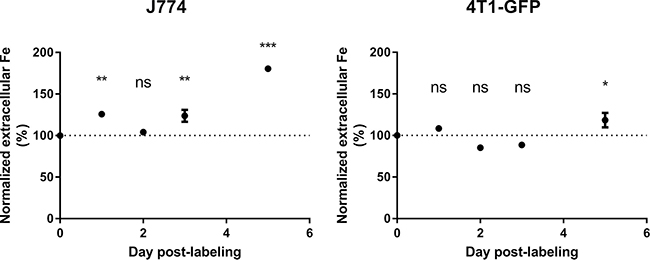
Figure 3: Extracellular iron levels increase in J774 macrophages after MIRB labeling. The extracellular iron levels were measured by ICP-MS in both J774 and 4T1-GFP cells labeled with MIRB. Data are expressed as means ± SEM. n = 3. *p < 0.05, **p < 0.01, ***p < 0.001, ns, p > 0.05.
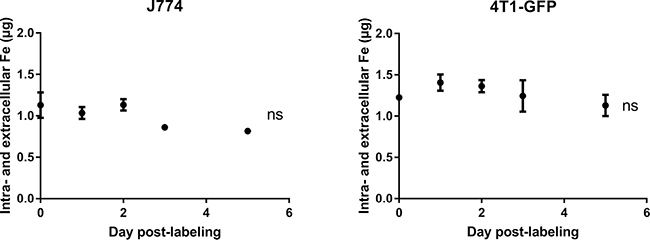
Figure 4: The intra- and extracellular iron pools do not vary with time after SPIO labeling in both 4T1-GFP and J774 cells. Iron values, obtained by ICP-MS, of cell samples and extracellular media were pooled. Data are expressed as means ± SEM. n = 3, ns, p > 0.05.
We addressed the fate of intracellular SPIO when MIRB-labeled 4T1-GFP cells are grown in the presence of control J774 macrophages. For this purpose, SPIO negative/GFP negative J774 macrophages were added to SPIO positive 4T1-GFP cells (overnight incubation with MIRB nanoparticles). Microscopy pictures performed just after incubation with MIRB nanoparticles showed that 4T1-GFP cells incorporated red fluorescent SPIO (Figure 5). At day 2 after the addition of J774 macrophages to MIRB positive 4T1-GFP cells, we observed that the red fluorescence decreased compared to day 0 and co-localized with GFP negative J774 macrophages (Figure 5). Of note, Perls’ Prussian blue staining confirmed that red fluorescence matched with iron clusters (Figure 5). Moreover, Perls’ Prussian blue staining and immunohistochemistry studies performed on 4T1 tumors after MRI cell tracking (See Figure 1) at day +7 showed that remaining iron clusters colocalized with F4/80 positive macrophages (Figure 6).
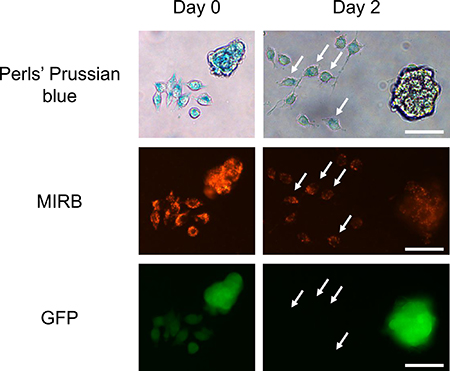
Figure 5: SPIO-free J774 macrophages take up iron oxides released by MIRB-labeled 4T1-GFP cells. Fluorescence imaging of MIRB iron oxides (red fluorescence) and 4T1-GFP cells, combined with the iron detection using Perls’Prussian blue staining were performed just after overnight incubation of 4T1-GFP cells (day 0) with SPIO. GFP/SPIO negative J774 macrophages were added to SPIO-labeled 4T1-GFP cells and allowed to grow for 2 days before imaging (day 2). White arrows indicate the presence of GFP negative J774 macrophages containing red fluorescent SPIO. Scale bar, 50 μm.
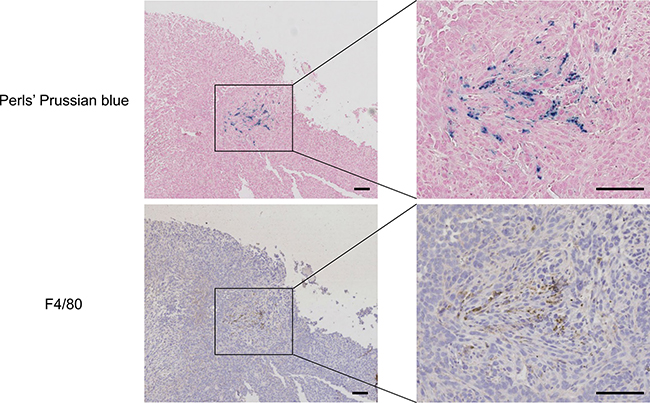
Figure 6: Histology of tumor tissue sample at day 7 post-injection of MIRB-labeled 4T1-GFP cancer cells shows that remaining SPIO are engulfed by macrophages. Mice received an intramuscular injection of 106 MIRB-labeled 4T1-GFP cells and underwent MRI scans for 7 days. After longitudinal MR imaging, mice were sacrificed and tumors were collected for histological analysis. Microscopy pictures show iron positive cells (Perls’ Prussian blue staining) and F4/80-positive macrophages (brown). Scale bar, 100 μm.
DISCUSSION
MRI cell tracking has several potential applications related to cancer. In a series of studies, it was used for characterizing the interplay between immune cells (dendritic cells, T cells and NK cells) and cancer cells for the evaluation of cancer immunotherapy treatments [24–26]. SPIO-based MRI cell tracking can also be used for monitoring the fate of cancer and tumor-associated cells. For instance, cellular imaging techniques were used for visualizing the homing of macrophages into metastasized lymph nodes and the trafficking of endothelial progenitor cells to sites of tumor angiogenesis [27, 28].
We previously used magnetic cell tracking techniques in experimental breast cancer brain metastasis and renal cancer models [8, 14]. These experiments evidenced that the negative MR T2(*) contrast induced by SPIO quickly decreased in tissues after injection [8, 12]. Ex vivo EPR measurements showed that the iron oxide content of tissues quickly dropped after injection of SPIO-labeled cancer cells [8, 14]. Based on these observations, we concluded that reporter gene-based methods (e.g., bioluminescence imaging, MR reporter genes such as ferritin [29]) should be preferred to SPIO-based MR cellular imaging for the long-term tracking of metastatic cells. However, these studies did not document the fate of SPIO in vivo.
Here, we found that macrophages can degrade SPIO after internalization. Two complementary techniques, ICP-MS and EPR, revealed that superparamagnetic and non-superparamagnetic iron decreased in SPIO-labeled macrophages after magnetic labeling, which was associated to a 80% increase in extracellular iron 5 days after SPIO labeling. Comparatively, SPIO-labeled 4T1 breast cancer cells released low amounts of iron in the extracellular medium (17% increase in iron at day 5). The intracellular iron (oxide) content remained unchanged up to 5 days after SPIO labeling in 4T1-GFP cells. A co-culture experiment of SPIO-positive 4T1 breast cancer cells and SPIO-free J774 macrophages further revealed that macrophages engulf SPIO initially present in the cytoplasm of cancer cells. SPIO release by cancer cells is likely due to cell death and/or exocytosis of nanoparticles [30]. Our data evidence that macrophages metabolize superparamagnetic iron (MRI/EPR detectable) into non-superparamagnetic iron (MRI/EPR silent). In agreement with these results, in vivo MRI cell tracking experiments showed that signal voids induced by SPIO on T2*-w images quickly decreased after an intramuscular injection of SPIO-labeled 4T1 cells in mice. Although histology showed that remaining iron oxides were engulfed by macrophages, the use of more sensitive histological methods would perhaps highlight that some SPIO are still present in tumor cells 7 days after injection [31, 32].
We observed in vitro that iron oxide positive macrophages significantly degrade SPIO within 3–5 days after SPIO labeling (Figure 2). Our in vivo results showed that contrast loss occurred from day 2 to day 7 post-injection of SPIO positive 4T1 cancer cells. Hence, Figure 2 (iron quantification in macrophages and Figure 6 (histology) indicated that macrophages were present in the tumor and could quickly degrade SPIO. Based on these pieces of evidence, it appears that the contribution of macrophages to the in vivo signal loss is significant. Although histology showed that remaining iron oxides were engulfed by macrophages, the use of more sensitive histological methods would perhaps highlight that some SPIO are still present in tumor cells 7 days after injection [31, 32].
After in vivo injection of SPIO-labeled breast cancer cells, we can define two concomitant mechanisms to explain the contrast loss observed on T2*-w MR images (Figure 7). First, previous studies showed that the intracellular iron oxide content is shared among daughter cells during cell proliferation [10, 14, 33]. It results in a dilution of the SPIO content in the tumor and a decrease in the darkening effect of SPIO on MR T2-,T2*-w images [14, 33]. Second, the present study showed that SPIO used for cancer cell labeling is eventually taken up by macrophages. The presence of free SPIO in the tumor is likely due to cell death.
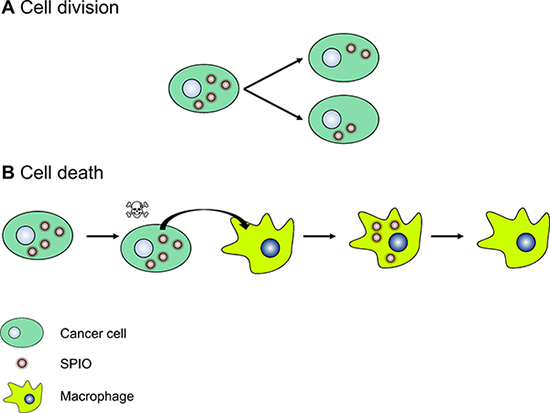
Figure 7: A cartoon representing the mechanisms of the loss of MR T2(*) contrast after injection of SPIO-labeled cancer cells in vivo. (A) Dilution of the iron oxide content following cell division. For the sake of clarity, the iron oxide content is equally shared among daughter cells. (B) Biotransformation of SPIO to non-superparamagnetic iron by macrophages after the release of SPIO by dead cancer cells.
In experimental metastasis models (i.e., the injection of cancer cells in the blood stream), cell tracking techniques were used to ensure the proper delivery of cancer cells to targeted sites and to characterize the long-term fate of cancer cells at the metastatic site [8, 9, 12, 14, 34, 35]. In a previous study, authors evaluated the fate of breast cancer cells in the mouse brain after intracardiac injection [9]. Cells were labeled in vitro with nondegradable micron-sized iron oxides (MPIO) with an inert styrene/divinylbenzene coating [9]. Taking advantage of contrast changes on MR images, authors observed that the vast majority of entrapped iron-containing cancer cells died after injection (loss of negative contrast), whereas remaining cells were either dormant (signal persistence over time) or proliferating (onset of macrometastases [9]). Using a similar approach in a melanoma model, Townson et al. showed that treatment with doxorubicin inhibited tumor growth but did not affect the number of solitary dormant melanoma cells in the mouse liver [12]. Although our study relied on the use of degradable SPIO, we showed that tumor macrophages take up iron oxides a few days after injection of SPIO-labeled cancer cells. Hence, our works challenge the idea that MRI cellular imaging allows the proliferative status of metastatic cells to be determined.
Our results are in line with several studies that attempted to validate MR cellular imaging for tracking stem cells in cell-based therapies. Briefly, in vivo MRI cell tracking experiments showed that SPIO nanoparticles initially incorporated in stem cells finally end up in the cytoplasm of macrophages [36–38]. Thus, a discrepancy was reported between MRI results and the fate of the stem cells after injection in vivo. In our study, we observed a quick loss of the negative contrast induced by SPIO in vivo (< 1 week), whereas above-cited studies reported longer signal persistence after injection (4 weeks) [36]. This can be explained by differences in SPIO formulation, coating and stability.
To conclude, SPIO-based cell tracking suffers from limitations that restricts the follow-up period after injection of cells of interest in vivo. However, this technique is valid for ensuring the proper delivery of cells at a given site. For long-term cell tracking in preclinical models, the use of reporter genes should be preferred because their expression is not diminished after cell division and they are only expressed by living cells.
MATERIALS AND METHODS
Cell culture
Green fluorescent protein-expressing 4T1 murine breast cancer cells cells (4T1-GFP) were generated by lentiviral infection (addgene plasmid pRRLSIN.cPPT.PGK-GFP.WPRE #12252). 4T1-GFP cells were grown in DMEM + GlutaMAX (Invitrogen), supplemented with 10% (v/v) fetal bovine serum (Invitrogen) and 1% (v/v) penicillin/streptomycin (Invitrogen). J774 murine macrophages were grown in RPMI + GlutaMAX (Invitrogen), supplemented with 10% (v/v) fetal bovine serum (Invitrogen).
Cell labeling with SPIO and sample preparation
On day -1, 4T1-GFP or J774 cells were seeded in 6-well plates (7.5 × 104 cells/well). At day 0, the medium was removed and replaced by 2 ml of full medium containing Molday ION™ Rhodamine B red fluorescent SPIO particles (MIRB 25 μg Fe/ml, Biopal, Worcester, USA) for overnight incubation. MIRB are dextran-coated SPIO, have a zeta potential of ~ +31 mV and a colloidal size of 35 nm [39]. After MIRB labeling, cells were washed three times with PBS, harvested using 0.05% trypsin/EDTA (Invitrogen), spun down (180 g for 5 minutes) and resuspended in PBS. Cells were then counted using a hemocytometer, centrifuged (160 g for 3 minutes) and resuspended in PBS for EPR or ICP-MS measurements. Extracellular media were also collected at different time points after labeling and spun down for 5 minutes at 10,000 g, after which supernatants were collected for iron measurements. To characterize SPIO localization after labeling, fluorescence microscopy was performed on days 0, 1, 2, 3 and 5 post-labeling using an AxioVert S100 microscope (Zeiss). Exposure times were kept constant during experiments. All experiments were performed in triplicate.
Iron quantification
For measuring SP iron content, samples were analyzed at room temperature using a Bruker EMX EPR spectrometer operating at 9 GHz (Bruker Biospin GmBh, Germany) with the typical parameters: 3 mT modulation amplitude, 1.724 mW power, 325.1 mT center field, and field 400 mT sweep width. When measurements are performed at room temperature, the EPR signal is specific for superparamagnetic iron [14, 20–22].
For measuring total iron (SP + non-SP) content, samples were analyzed by inductively coupled argon plasma with mass spectrometry using an Agilent 7500ce instrument as previously described [22]. Briefly, samples (50 μl) were diluted quantitatively (1:100) with a HNO3 1%, HCl 0.5% solution containing all internal standards (Sc, Ge, Rh and Ir). Fe was quantified using helium mode (collision cell) and its isotope at m/z 56 with Ge (at m/z 74) as internal standard.
Co-culture of MIRB-labeled 4T1-GFP cells and J774 macrophages and Perls’ Prussian blue staining
On day -2, 4T1-GFP cells were seeded in triplicate in a 6-well plate (3 × 104 cells/well). On day -1, MIRB nanoparticles were added to the culture medium (25 μg/ml, overnight incubation). On day 0, culture medium was removed, cells were washed 3 times with PBS and replenished with full medium. Fluorescence microscopy was performed to confirm the intracellular localization of MIRB particles, and 7.5 × 104 control J774 macrophages were added to MIRB-labeled 4T1-GFP cells. Cells were next imaged for 2 days using fluorescence microscopy (AxioVert S100 microscope, Zeiss). Exposure times were kept constant during experiments. After fluorescence imaging, cells were rinsed 3 times with PBS, fixed with paraformaldehyde 4% (Sigma-Aldrich, Bornem, Belgium), and washed 3 times with PBS containing 0.1% Triton X-100 (Sigma-Aldrich, Bornem, Belgium) for membrane permeabilization. Fixed cells were next incubated with a PBS solution containing 1% potassium ferrocyanide (Sigma-Aldrich, Bornem, Belgium)/1% HCl for 15 minutes. After Perls’ Prussian blue staining of iron, cells were washed with PBS and brightfield pictures were taken (AxioVert S100 microscope, Zeiss).
Ethical statement
All in vivo experiments were performed in compliance with guidelines set by national regulations and were approved by the ethical committee for animal care of the Université catholique de Louvain (UCL).
MRI follow-up of MIRB-labeled 4T1-GFP cells in vivo
Five-week-old female BALB/c mice (Janvier, Le Genest-Saint-Isle, France) were anesthetized with ketamine (80 mg/kg)/xylazine (8 mg/kg). 106 MIRB-labeled 4T1-GFP cells resuspended in 100μl PBS were next injected into the right gastrocnemius muscle. 106 control 4T1-GFP cells were injected into the left gastrocnemius muscle.
In vivo MRI experiments were performed with an 11.7 T Biospec system (Bruker, Ettlingen, Germany) with a volume transmit coil (birdcage, inner diameter = 72 mm) coupled with a receive phase array coil (with four elements) at day 0, 1, 2, 5, 6 and 7 post-injection. Animals (n = 4) were anesthetized by inhalation of 1.5–2% isoflurane (Forene, Abbott, Maidenhead, UK). Mouse temperature was maintained at 37.0°C using a water blanket and respiration was monitored. Coronal T2*-weighted (T2*-w) images were acquired using a multiple gradient-echo (MGE) sequence with the following parameters: TR, 1100 ms; TE, 3.36 ms; number of echoes, 16; excitation pulse angle, 30°; field of view, 3 cm × 2 cm; matrix, 386 × 256; slice thickness, 1 mm; number of slices, 15; in-plane resolution, 78 μm × 78 μm; number of averages, 3; acquisition time, 14 minutes. First-echo images were used as anatomical T2*-w images.
Histological analysis of macrophages recruitment
After MRI experiments, animals were euthanized using an overdose of pentobarbital. Tumors were harvested and cut into 1 mm-thick slices, which were formalin-fixed and paraffin-embedded. Perls’ Prussian blue staining and F4/80 immunostaining were performed on adjacent 5 μm sections to detect iron and macrophages, respectively. For macrophage detection, after manual deparaffinization, endogenous peroxidases were inhibited by a 20 min-treatment with 3% H2O2 in methanol. Sections were then subjected to antigen retrieval in 10 mM citrate buffer pH 5.7. Aspecific antigen binding sites were blocked with a TBS solution containing 5% BSA and 0.05% Triton. Endogenous biotin was blocked with an avidin/biotin blocking kit (Vector SP2001). Slices were stained with anti-F4/80 primary antibodies (AbDSerotec MCA497GA) overnight at 4°C at a 1/500 dilution, followed by incubation with biotinylated goat anti-rat secondary antibodies (Vector BA4001) at a 1/50 dilution for 60 min at room temperature. This reaction was visualized using Streptavidin-HRP (BD Biosciences 554066) and DAB (Dako K3468). After counterstaining with hematoxylin (Dako S3301), slices were dehydrated and coverslipped. Slides were finally digitalized at a 20× magnification with a SCN400 slide scanner (Leica, Wetzlar, Germany) and visualized on the Digital Image Hub (Leica Biosystems, Dublin, Ireland).
Statistical analyses
All results are represented as means ± standard error of the mean (SEM). Groups were compared using One-Way ANOVA followed by Dunnett’s multiple comparisons test; p < 0.05 was considered to be statistically significant. Statistical analyses were performed using GraphPad Prism software (GraphPad Software Inc., La Jolla, USA).
Abbreviations
EPR: Electron paramagnetic resonance; GFP: Green fluorescent protein; ICP-MS: Inductively coupled plasma mass spectroscopy; MIRB: Molday ION™ Rhodamine B red fluorescent iron oxides; MRI: Magnetic resonance imaging; SEM: Standard error of the mean; SP: Superparamagnetic; SPIO: Superparamagnetic particles of iron oxides; T2(*)-w: T2(*)-weighted.
Authors’ contributions
PD, PS and BG designed the experiments, wrote the main manuscript text and prepared figures. PD, GD, NJ, CB, and PL performed the experiments. PD and PL analyzed data. OF supervised fluorescence microscopy experiments. VH supervised ICP-MS experiments. BFJ and BG supervised EPR/MRI experiments. All authors reviewed the manuscript.
ACKNOWLEDGMENTS AND FUNDING
We thank Chantal Fregimilicka and Michèle De Beukelaer for the histological studies. We thank Dr. Fred Miller (Karmanos Cancer Institute) for providing 4T1 cells and Dr. Paolo. E. Porporato for generating 4T1-GFP cells. We thank Prof. Françoise Van Bambeke and Prof. Giulio Muccioli for providing J774 cells. This work was supported by grants from the Belgian National Fund for Scientific Research (F.S.R.-FNRS; PDR T.0107.13), the Fonds Joseph Maisin, the Communauté Française de Belgique (ARC 14/19-058) and the Télévie. Funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. P. Sonveaux and B. F. Jordan are Senior Research Associates and P. Danhier is a Postdoctoral Researcher of the F.R.S-FNRS. The authors have declared that no competing interests exist.
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
REFERENCES
1. Gupta GP, Massague J. Cancer metastasis: building a framework. Cell. 2006; 127:679–95. doi: 10.1016/j.cell.2006.11.001.
2. Bulte JW, Kraitchman DL. Iron oxide MR contrast agents for molecular and cellular imaging. NMR Biomed. 2004; 17:484–99. doi: 10.1002/nbm.924.
3. Kircher MF, Gambhir SS, Grimm J. Noninvasive cell-tracking methods. Nat Rev Clin Oncol. 2011; 8:677–88. doi: 10.1038/nrclinonc.2011.141.
4. Addicott B, Willman M, Rodriguez J, Padgett K, Han D, Berman D, Hare JM, Kenyon NS. Mesenchymal stem cell labeling and in vitro MR characterization at 1.5 T of new SPIO contrast agent: Molday ION Rhodamine-B. Contrast Media Mol Imaging. 2011; 6:7–18. doi: 10.1002/cmmi.396.
5. Shen WB, Plachez C, Chan A, Yarnell D, Puche AC, Fishman PS, Yarowsky P. Human neural progenitor cells retain viability, phenotype, proliferation, and lineage differentiation when labeled with a novel iron oxide nanoparticle, Molday ION Rhodamine B. Int J Nanomedicine. 2013; 8:4593–600. doi: 10.2147/IJN.S53012.
6. Smirnov P, Gazeau F, Beloeil JC, Doan BT, Wilhelm C, Gillet B. Single-cell detection by gradient echo 9.4 T MRI: a parametric study. Contrast Media Mol Imaging. 2006; 1: 165–74. doi: 10.1002/cmmi.104.
7. Foster-Gareau P, Heyn C, Alejski A, Rutt BK. Imaging single mammalian cells with a 1.5 T clinical MRI scanner. Magn Reson Med. 2003; 49:968–71. doi: 10.1002/mrm.10417.
8. Danhier P, Magat J, Leveque P, De Preter G, Porporato PE, Bouzin C, Jordan BF, Demeur G, Haufroid V, Feron O, Sonveaux P, Gallez B. In vivo visualization and ex vivo quantification of murine breast cancer cells in the mouse brain using MRI cell tracking and electron paramagnetic resonance. NMR Biomed. 2015; 28:367–75. doi: 10.1002/nbm.3259.
9. Heyn C, Ronald JA, Ramadan SS, Snir JA, Barry AM, MacKenzie LT, Mikulis DJ, Palmieri D, Bronder JL, Steeg PS, Yoneda T, MacDonald IC, Chambers AF, et al. In vivo MRI of cancer cell fate at the single-cell level in a mouse model of breast cancer metastasis to the brain. Magn Reson Med. 2006; 56:1001–10. doi: 10.1002/mrm.21029.
10. Economopoulos V, Chen Y, McFadden C, Foster PJ. MRI detection of nonproliferative tumor cells in lymph node metastases using iron oxide particles in a mouse model of breast cancer. Transl Oncol. 2013; 6:347–54.
11. Magnitsky S, Roesch A, Herlyn M, Glickson JD. In vivo and ex vivo MR imaging of slowly cycling melanoma cells. Magn Reson Med. 2011; 66:1362–73. doi: 10.1002/mrm.22917.
12. Townson JL, Ramadan SS, Simedrea C, Rutt BK, MacDonald IC, Foster PJ, Chambers AF. Three-dimensional imaging and quantification of both solitary cells and metastases in whole mouse liver by magnetic resonance imaging. Cancer Res. 2009; 69:8326–31. doi: 10.1158/0008-5472.CAN-09-1496.
13. Aguirre-Ghiso JA. Models, mechanisms and clinical evidence for cancer dormancy. Nat Rev Cancer. 2007; 7:834–46. doi: 10.1038/nrc2256.
14. Danhier P, De Preter G, Magat J, Godechal Q, Porporato PE, Jordan BF, Feron O, Sonveaux P, Gallez B. Multimodal cell tracking of a spontaneous metastasis model: comparison between MRI, electron paramagnetic resonance and bioluminescence. Contrast Media Mol Imaging. 2014; 9:143–53. doi: 10.1002/cmmi.1553.
15. Wilhelm C, Gazeau F, Bacri JC. Magnetophoresis and ferromagnetic resonance of magnetically labeled cells. Eur Biophys J. 2002; 31:118–25. doi: 10.1007/s00249-001-0200-4.
16. Iannone A, Federico M, Tomasi A, Magin RL, Casasco A, Calligaro A, Vannini V. Detection and quantitation in rat tissues of the superparamagnetic magnetic resonance contrast agent dextran magnetite as demonstrated by electron spin resonance spectroscopy. Invest Radiol. 1992; 27:450–5.
17. Radermacher KA, Beghein N, Boutry S, Laurent S, Vander Elst L, Muller RN, Jordan BF, Gallez B. In vivo detection of inflammation using pegylated iron oxide particles targeted at E-selectin: a multimodal approach using MR imaging and EPR spectroscopy. Invest Radiol. 2009; 44:398–404.
18. Radermacher KA, Boutry S, Laurent S, Elst LV, Mahieu I, Bouzin C, Magat J, Gregoire V, Feron O, Muller RN, Jordan BF, Gallez B. Iron oxide particles covered with hexapeptides targeted at phosphatidylserine as MR biomarkers of tumor cell death. Contrast Media Mol Imaging. 2010; 5:258–67. doi: 10.1002/cmmi.382.
19. Radermacher KA, Magat J, Bouzin C, Laurent S, Dresselaers T, Himmelreich U, Boutry S, Mahieu I, Vander Elst L, Feron O, Muller RN, Jordan BF, Gallez B. Multimodal assessment of early tumor response to chemotherapy: comparison between diffusion-weighted MRI, 1H-MR spectroscopy of choline and USPIO particles targeted at cell death. NMR Biomed. 2012; 25:514–22. doi: 10.1002/nbm.1765.
20. Levy M, Luciani N, Alloyeau D, Elgrabli D, Deveaux V, Pechoux C, Chat S, Wang G, Vats N, Gendron F, Factor C, Lotersztajn S, Luciani A, et al. Long term in vivo biotransformation of iron oxide nanoparticles. Biomaterials. 2011; 32:3988–99. doi: 10.1016/j.biomaterials.2011.02.031.
21. Danhier P, Gallez B. Electron paramagnetic resonance: a powerful tool to support magnetic resonance imaging research. Contrast Media Mol Imaging. 2015; 10:266–81. doi: 10.1002/cmmi.1630.
22. Danhier P, De Preter G, Boutry S, Mahieu I, Leveque P, Magat J, Haufroid V, Sonveaux P, Bouzin C, Feron O, Muller RN, Jordan BF, Gallez B. Electron paramagnetic resonance as a sensitive tool to assess the iron oxide content in cells for MRI cell labeling studies. Contrast Media Mol Imaging. 2012; 7:302–7. doi: 10.1002/cmmi.497.
23. Boutry S, Forge D, Burtea C, Mahieu I, Murariu O, Laurent S, Vander Elst L, Muller RN. How to quantify iron in an aqueous or biological matrix: a technical note. Contrast Media Mol Imaging. 2009; 4:299–304. doi: 10.1002/cmmi.291.
24. de Vries IJ, Lesterhuis WJ, Barentsz JO, Verdijk P, van Krieken JH, Boerman OC, Oyen WJ, Bonenkamp JJ, Boezeman JB, Adema GJ, Bulte JW, Scheenen TW, Punt CJ, et al. Magnetic resonance tracking of dendritic cells in melanoma patients for monitoring of cellular therapy. Nat Biotechnol. 2005; 23:1407–13. doi: 10.1038/nbt1154.
25. Hong H, Yang Y, Zhang Y, Cai W. Non-invasive cell tracking in cancer and cancer therapy. Curr Top Med Chem. 2010; 10:1237–48.
26. Ottobrini L, Martelli C, Trabattoni DL, Clerici M, Lucignani G. In vivo imaging of immune cell trafficking in cancer. Eur J Nucl Med Mol Imaging. 2011; 38:949–68. doi: 10.1007/s00259-010-1687-7.
27. Arbab AS, Pandit SD, Anderson SA, Yocum GT, Bur M, Frenkel V, Khuu HM, Read EJ, Frank JA. Magnetic resonance imaging and confocal microscopy studies of magnetically labeled endothelial progenitor cells trafficking to sites of tumor angiogenesis. Stem Cells. 2006; 24:671–8. doi: 10.1634/stemcells.2005-0017.
28. Cho HR, Choi SH, Lee N, Hyeon T, Kim H, Moon WK. Macrophages homing to metastatic lymph nodes can be monitored with ultrasensitive ferromagnetic iron-oxide nanocubes and a 1.5T clinical MR scanner. PLoS One. 2012; 7:e29575. doi: 10.1371/journal.pone.0029575.
29. Cohen B, Dafni H, Meir G, Harmelin A, Neeman M. Ferritin as an endogenous MRI reporter for noninvasive imaging of gene expression in C6 glioma tumors. Neoplasia. 2005; 7:109–17. doi: 10.1593/neo.04436.
30. Cromer Berman SM, Kshitiz, Wang CJ, Orukari I, Levchenko A, Bulte JW, Walczak P. Cell motility of neural stem cells is reduced after SPIO-labeling, which is mitigated after exocytosis. Magn Reson Med. 2013; 69:255–62. doi: 10.1002/mrm.24216.
31. Schroeter M, Saleh A, Wiedermann D, Hoehn M, Jander S. Histochemical detection of ultrasmall superparamagnetic iron oxide (USPIO) contrast medium uptake in experimental brain ischemia. Magn Reson Med. 2004; 52:403–6. doi: 10.1002/mrm.20142.
32. Mukaide T, Hattori Y, Misawa N, Funahashi S, Jiang L, Hirayama T, Nagasawa H, Toyokuni S. Histological detection of catalytic ferrous iron with the selective turn-on fluorescent probe RhoNox-1 in a Fenton reaction-based rat renal carcinogenesis model. Free Radic Res. 2014; 48:990–5. doi: 10.3109/10715762.2014.898844.
33. Walczak P, Kedziorek DA, Gilad AA, Barnett BP, Bulte JW. Applicability and limitations of MR tracking of neural stem cells with asymmetric cell division and rapid turnover: the case of the shiverer dysmyelinated mouse brain. Magn Reson Med. 2007; 58:261–9. doi: 10.1002/mrm.21280.
34. Song HT, Jordan EK, Lewis BK, Liu W, Ganjei J, Klaunberg B, Despres D, Palmieri D, Frank JA. Rat model of metastatic breast cancer monitored by MRI at 3 tesla and bioluminescence imaging with histological correlation. J Transl Med. 2009; 7:88. doi: 10.1186/1479-5876-7-88.
35. Perera M, Ribot EJ, Percy DB, McFadden C, Simedrea C, Palmieri D, Chambers AF, Foster PJ. In Vivo Magnetic Resonance Imaging for Investigating the Development and Distribution of Experimental Brain Metastases due to Breast Cancer. Transl Oncol. 2012; 5:217–25.
36. Amsalem Y, Mardor Y, Feinberg MS, Landa N, Miller L, Daniels D, Ocherashvilli A, Holbova R, Yosef O, Barbash IM, Leor J. Iron-oxide labeling and outcome of transplanted mesenchymal stem cells in the infarcted myocardium. Circulation. 2007; 116:I38–45. doi: 10.1161/CIRCULATIONAHA.106.680231.
37. Terrovitis J, Stuber M, Youssef A, Preece S, Leppo M, Kizana E, Schar M, Gerstenblith G, Weiss RG, Marban E, Abraham MR. Magnetic resonance imaging overestimates ferumoxide-labeled stem cell survival after transplantation in the heart. Circulation. 2008; 117:1555–62. doi: 10.1161/CIRCULATIONAHA.107.732073.
38. Ma N, Cheng H, Lu M, Liu Q, Chen X, Yin G, Zhu H, Zhang L, Meng X, Tang Y, Zhao S. Magnetic resonance imaging with superparamagnetic iron oxide fails to track the long-term fate of mesenchymal stem cells transplanted into heart. Sci Rep. 2015; 5:9058. doi: 10.1038/srep09058.

