INTRODUCTION
The most widely type of solid tumor in men is prostate cancer (PCa). Prostate cancer is the secondary highest cause of malignancy mortality, behind lung neoplasms, in developed occidental countries. It is assessed that prostate cancer hits approximately 2.8 million men in USA and the incidence rate raises with advanced years and in many countries [1]. Prostate cancer occurrence and mortality are up to 20-fold higher in industrialized countries respect to developing ones, thus for this purpose diet and lifestyle have been proposed as factors causing this discrepancy. Nutrition and exercise modify serum factors that slow down the growth and induce apoptosis in androgen-dependent PCa cells, while high levels of body mass index, blood pressure and several metabolic factors were correlated with high risk of prostate cancer death.
Many studies establish relevant ethnic diversity in death rates: Caribbean hold the highest rates in the world (26.3%), following Sub-Saharan Africans (10%) and, at last, the Asians show the lowest (2.5%). Perhaps genetic and environmental components contribute to generate this ethnic variety. Moreover, it was found that the risk of developing prostate cancer among American men, in particular African Americans, is highest than the other populations in the United States [2].
New therapies with established benefits on survival rate have been produced in last years [3–10], but nevertheless the gain in terms of survival is minimal. Prostate cancer is a very heterogeneous tumor which can show up indolent or very aggressive, usually with metastasis to other organs and bone [11–14], thus generating morbidity and mortality [15]. Prostate cancer can be clinically diagnosed as local or advanced and different treatments including surveillance, radiotherapy [16–18], radical prostatectomy and androgen-deprivation treatment are expected.
First line therapeutic options for local prostate cancer include different approaches: surgical prostatectomy, neoadjuvant therapy and radiation therapy. At present, the only advantageous treatment in PCa is surgical prostatectomy, but it involves adverse effects. Therefore, several studies recommended neoadjuvant therapy to improve survival rate. The mentioned therapy, known as androgen deprivation therapy (ADT), leads a reduction of androgens in prostate cancer cells of patients with advanced asymptomatic disease. It involves the use of drugs such as luteinising hormone-releasing hormone (LHRH) agonists, LHRH antagonists, antiandrogens, maximum androgen blockade and intermittent androgen deprivation. Radiation therapy is the main local disease treatment after surgical approach, in particular the hormonal therapy and radiotherapy combination improved survival rate [19]. On the contrary, treatment of choice of advanced prostate cancer is founded on chemotherapy: docetaxel in combination with prednisone, abiraterone acetate (AA) with prednisone, enzalutamide and cabazitaxel. Nevertheless, advanced prostate cancer remains an incurable disease and new alternative treatment are required.
The main screening exams to diagnose prostate cancer include digital rectal exam (DRE), serum level of prostate-specific antigen (PSA) and transrectal ultrasound guided biopsy. A suspected DRE alone reveals PCa, regardless of psa level, in about 18% of all patients; moreover, a suspected digital rectal exam has a positive predictive value of 5–30% in patients who have a PSA level up to 2 ng/ml. At present, measurement of PSA is the most usual marker able to correlate with prostate cancer risk, aggressiveness and outcome in order to detect prostate cancer [20]. Anyway, it has been found that several patients develop this type of cancer in spite of low PSA levels though a higher PSA level shows the presence of PCa [21–22]. Contrariwise, several factors as well as benign prostatic hyperplasia, gland inflammation and some drugs could arise from high PSA levels [23–26]. PSA screening is usually associated with over-diagnosis although it has decreased the death rate of PCa [27]. An important prognostic indicator of prostate cancer used to performe the histopathologic grading is Gleason score. Gleason-based grading allows to classify tumors according to their relative degree of differentiation by assigning a score from 1 (most differentiated) to 5 (least differentiated). Since it has been elucidated that tumors with similar histological patterns may provide different clinical outcomes, it became clear that Gleason score cannot exactly predict the aggressiveness of the disease although it is a powerful prognostic indicator [31–33]. The diagnosis of prostate cancer essentially results from cytologic or histopathologic confirmation, in particular in patients with PSA 4–10 ng/ml (“grey zone”). Hence, sometimes over-diagnosis and over-therapy of patients undergoing surgical or pharmacological therapy without a real necessity occur. For this reason and for the molecular heterogeneity of prostate cancer [34], it is necessary to identify alternative prostate cancer biomarkers, such as miRNA, to expedite and improve cancer diagnosis, prognosis and response to treatment, that will open the way to personalized therapeutic strategy. The purpose of this review is to analyze the improvement made to our knowledge of the function of miRNA in PCa tumourigenesis and for discovering which miRNAs may be considered helpful serum biomarkers.
MiRNAs biogenesis and function
Small endogenous non-coding RNA (18–25 nt) are known as miRNAs. They can prevent protein expression through cleavage of specific target mRNAs or through inhibition of their translation [35]. Their biogenesis begins in the nucleus: a primary RNA (pri-miRNA), previously transcribed by RNA polymerase II (Pol II), is identified and processed by RNase III Drosha-DGCR8 complex to obtain a miRNA precursor (pre-miRNA), that is carried into the cytoplasm by Exportin-5 and Ran-GTP61. The RNase III Dicer, coupled to the transactivation-response RNA-binding protein (TRBP) enzyme, cleaves the pre-miRNA into a mature double-stranded miRNA of about 22 nucleotides. Afterwards, the functional strand of the mature miRNA is loaded with Argonaute (AGO) proteins into the RISC (RNA induced silencing complex): here miRNA drives RISC to bind the 3’ UTR of a mRNA target and resulting in either mRNA cleavage, translational repression or deadenylation. On the contrary, the not functional strand will be degraded. In mammals there are four AGO (AGO1-4) proteins that induce mRNA degradation by interaction with deadenylation factors and translational machinery. Besides the aforementioned canonical mechanisms of biogenesis of miRNAs other “non canonical” mechanisms are emerging where some passages are bypassed. Mirtrons are one of the first noncanonical miRNAs that during their production do not require Drosha [36–38]. A recent study identifies another group called “5′ capped pre-miRNAs” that contains a 7-methylguanosine as a cap that includes miR-320 and miR-484. In addition there is also a Dicer-independent maturation of which is part miR-451 that is directly embedded in AGO2 and subsequently cut by exoribonuclease PARN into a mature form of miR-451 [39–43]. With respect to the phase of nuclear export, a current study revealed that miR-320 needs exportin 1 instead of exportin-5. MiRNAs can act by binding either the 3’ UTR of the mRNA target through imperfect complementarity or via multiple sites by inhibition of the mRNA interaction with ribosomal complex and translational machinery. Furthermore, the imperfect complementarity with target ensures that miRNAs have multiple intracellular targets that lead to amplificate the biological effects. The identification of specific targets of miRNAs has been very difficult due to limited complementarity between miRNAs and mRNA transcripts. Functional characterization of miRNAs depends strongly on identification of their specific mRNA binding partners [44]. Several tools for target prediction have been realized to discover miRNA targets and most of them are conceived to concurrently decrease the false-positive incidence and raise the accuracy of their findings [45]. For example, TargetScan [46–49], one of the most used, allows to search miRNA families across several species by miRNA name, gene name or from more to less conserved. TargetScan is easy to use, but only considers stringent seeds, and ignores many potential targets. Therefore, computational prediction of miRNA target sites is an efficient tool to understand the molecular mechanisms of miRNA-mediated interactions and plays a key role in miRNA-based therapeutics in clinic. It is not unexpected that miRNAs have been shown to be involved in almost all key cellular processes, such as proliferation, differentiation, migration, apoptosis, stemness maintenance and metabolism, because each miRNA can modulate the expression of multiple mRNAs and, therefore, each of them may be targeted by several different miRNAs [50]. The deregulation of miRNAs expression in malignant diseases is due to the apoptotic evasion that results in tumorigenesis and drug-resistance, in particular during the tumor initiation and progression where act as oncogenes or as tumor suppressors and that are related to apoptosis. Therapies based on miRNA are used not only alone but in combination with anti-cancer therapies. This may reduce the toxicity and reduce the tumor drug resistance. For this reason, the studies are directed to new molecular therapies that block oncogenic miRNAs and stimulate the expression of those tumor suppressor [51–55]. In order to inhibit miRNA oncogenes, various strategies for miRNA-based therapeutics have been studied: a direct method involves oligonucleotides or virus-based constructs to inhibit the oncogenic miRNAs expression and to restore the expression loss of tumor suppressor miRNA, whereas an indirect method includes drugs that regulate miRNAs expression through their transcripts [56]. In the context of direct strategies, antisense oligonucleotides, miRNA sponges, miRNA-Masking Antisense oligonucleotides Technology (miRNA-mask) and small RNA inhibitors are used to block oncogenic miRNA. The action of the antisense oligonucleotides as competitive inhibitors is carried out through annealing to the target miRNA guide and inducing degradation or duplex formation. These oligonucleotides have been modified in their chemical structure to increase their stability, specificity and affinity binding. The most common changes include the insertion of 2′-O- methoxyethyl groups (2′-MOE), the 2′-O- methyl groups and locked nucleic acid (LNA) [57, 58]. Locked nucleic acid (LNA) is a modified RNA oligonucleotide, whose ribose moiety is structurally linked to an extra bridge connecting the 2′-O and the 4′-C atoms. This bridge “blocks” the ribose in the 3′-endo conformation, which is peculiar of the A-form duplexes. LNA oligonucleotides have a big ability to hybridize complementary single- or double-stranded DNA and complementary single-stranded RNA [59]. MiRNA sponges have been developed as new oligonucleotide constructs containing multiple complementary miRNA binding sites to the target miRNA. A repression of miRNA targets was observed when introduced into the cell and it seems to mimic an in vitro silencing of miRNAs. A bulge at the position normally cleaved by Argonaute 2 was introduced to increase the affinity of these decoy transcripts, thus allowing strong association of miRNA sponges with ribonucleoproteins of the corresponding miRNA [60]. A novel approach provides the use of the MiR-mask: 2′-O-methyl-modified antisense oligonucleotides with complementarity to the 3′ UTR target mRNA able to compete with endogenous miRNAs for own target. MiR-Mask protects the miRNA binding site in order to inhibit its target gene (mRNA) and, thereby, has the ability to repress oncogenic miRNA injurious functions at the target level [61]. MiRNA expression is regulated at transcriptional level by small molecule miRNA inhibitors. The chance to use synthetic miRNAs (called miRNA mimics) or viral constructs containing genes coding for miRNAs, such as the adeno associated viral vectors, is helpful to avoid the loss or decrease of a tumour suppressor miRNA [62–64]. Several investigations show that miRNAs could be considered excellent biomarkers for cancer diagnosis, prognosis and therapy. In cancer cells the over-expressed miRNAs that promote carcinogenesis by inhibition of tumor suppressor genes are considered oncogenic miRNAs (OncomiRs), while the down-expressed ones, that normally avoid cancer development by inhibition of proto-oncogenes expression, are tumor suppressor miRNAs [65].
Disease-specific miRNA expression profiles in different type of human cancers have been detected and identified by many reports [66–69]. The utility in diagnosis, staging, progression, prognosis and response to treatment makes miRNA expression profiling of human cancers still more important. However, the study of miRNAs is constantly changing because a better comprehension of the molecular mechanisms underlying their biogenesis and function will enable the development of new therapeutic miRNA-based techniques.
MiRNAs as useful biomarkers in prostate cancer diagnosis and prognosis
MiRNAs represent an interesting target for biomarker discovery. Every miRNA can interact with various cellular pathways, and therefore alterations in the expression of few miRNAs may imply a bad regulation of several cellular processes conforming to the cancer complexity.
The issue of early cancer detection represents a promising field for microRNA-based diagnostics due to the currently lack of early detection methods and screening tests in many types of cancer.
In last years, it has been demonstrated through clinical and experimental discoveries that microRNAs have different expression in normal and cancer cells according to specific tissue expression signature [70]; microRNAs can also influence all hallmarks of cancer either by promoting or suppressing tumour development and progression [71]. Next to their intracellular functions, microRNAs have been detected as circulating molecules in many bio-organic fluids (urine, blood, saliva etc). Current investigations suggest that microRNAs are not only passively circulating molecules as byproducts, but also can act as mediators of intercellular communication via exosome-mediated transport in an ‘hormone-like’ way, thus mimicking the endocrine pathway [72]. Relying on these features, microRNAs could be considered excellent biomarkers. The big advantage they have is related to their high chemical stability in fresh or even formalin-fixed tissues and body fluids, so their potential as diagnostic markers increases in comparison with long noncoding RNAs or longer messenger RNAs and makes miRNA levels well suitable for testing in patient samples [73].
Several approaches to study miRNA profiling have been carried out [74]. It might be useful to identify a miRNAs panel testable in serum or tissue in order to discriminate patients with local prostate cancer that will eventually progress and will likely undergone to aggressive treatment from those ones with more advanced disease that have higher chances to benefit from special therapies such as novel agents acting on androgen pathway.
MiRNA profiling on serum is really exciting because it can be carried out in a non-invasive way, and it allows to establish a diagnostic serum miRNA panel that would probably prevent prostatic biopsies in patients with high PSA levels, some of whom at last will not have PCa. To date, the attempt to define a clear miRNA profile in prostate cancer has proved inconclusive. Numerous conflicting outcomes regarding miRNAs expression in prostate cancer still raise. Several literature studies reported the opposite expression of some miRNA in this disease: some studies establish an overall down-expression of miRNAs in tumours, while other studies prove a general over-expression of miRs in cancer [75] (Figure 1).
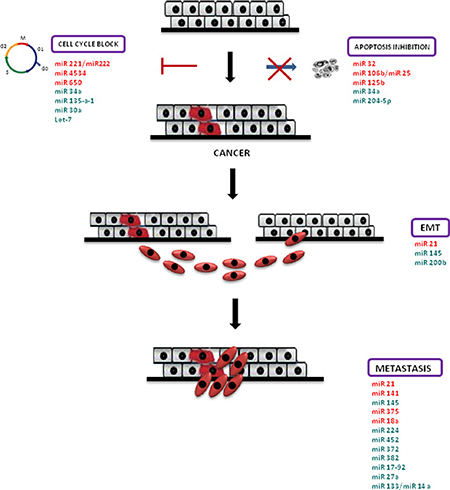
Figure 1: MiRNAs in PCa pathogenesis. Schematic representation of prostate cancer pathogenesis and miRNAs involved in this process. Cell cycle block and apoptosis resistance lead to cancer; afterwards, prostate cancer cells are subjected to epithelial mesenchymal transition (EMT), which confers increased migratory capacity and invasiveness to generate metastases. Over-expressed miRNAs are highlighted in red, down-expressed miRNAs are highlighted in green.
Oncogenic MiRNAs
One of the most common over-expressed oncomiR in cancer is miR-21. It controls the expression of many mRNA targets related to micro-vascular proliferation and tumor invasiveness. Its positive expression correlates with weak biochemical recurrence-free survival and has predictive value for biochemical recurrence risk in prostate cancer patients after radical prostatectomy [76]. Its expression is correlated with castration resistance and metastatic disease and enhances concurrently with clinical parameters (Gleason score, lymph node metastasis). Therefore, miR-21 is also helpful as biomarker to predict cancer progression [77]. Several literature studies showed that miR-221/miR-222 are increased in prostate cancer and that they could increase S-phase kinase associated protein 2 (Skp2), cyclin A and cyclin D1 via targeting p27Kip1 suppression, leading to cell cycle progression at G1- to -S phase, increased clonogenicity in vitro and enhanced tumorigenicity in vivo [78–80]. To date, the miR-141 function in PCa is not clear, but it has been revealed that it is over-expressed after castration, thus it’s likely its involvement in androgen mechanism, both in androgen-dependent and in metastatic castration-resistant prostate cancer. For this reason, the role of miR-141 in cancer progression seems interesting [81]. However, miRNAs expression in body fluids, in particular in serum, allows to obtain many interesting results concerning their role. In 2008, it was demonstrated for the first time by Mitchell et al. that cancer associated miRNAs can be easily detected and measured in biological fluids and that miR-141 levels were higher in serum of PCa patients [82].
In this context, miR-141 could be considered an important tool for prostate cancer diagnosis and prognosis. In fact, it has been elucidated that discrimination among low, intermediate and high risk patients with PCa can be carried out by miRNAs [77]. According to a good study concerning circulating miRNAs, it has been established that a decreased expression in non metastatic patients was also showed by miR-375 and therefore it appeared really important for early diagnosis [83, 84]. miR-18a is strongly over-expressed in prostate cancer tissues, thus acting as oncogenic miRNA. Its expression is really increased in peripheral blood of patients with prostate cancer compared with benign prostatic hyperplasia (BPH) patients and healthy individuals, as showed by Al-Kafaji G et al.
In addition, they have found that an higher miR-18a expression level correlates with PCa progression. These studies indicate that peripheral blood oncogenic miR-18a can function as a new potential non-invasive biomarker for PCa and can also make easy the distinction between PCa and BPH [85]. Another oncogenic miRNA overexpressed in PCa and positively correlated with poor overall and PSA recurrence free survival is miR-4534. It is hypermethylated in normal cells and tissues compared to those of PCa and exerts its oncogenic effects partly by arresting the tumor suppressor PTEN gene [86]. Its overexpression induces pro-cancerous characteristics in non-cancer cell line whereas its knockdown impairs cell proliferation, migration/invasion and induces apoptosis and then G0/G1 cell cycle block in PCa [87]. High level of miR-650 was discovered in PCa samples. Oncogenic activity of miR-650 in PCa is explicated through suppression of cellular stress response 1 (CSR1) expression. It was showed that miR-650 inhibition reversed the suppressed expression of CSR1, repressed colony formation and blocked cell cycle entry to the S phase in DU145 and PC3 cells [88].
MiR-32 is highly expressed in castration resistant prostate cancer (CRPC) samples compared to benign prostatic hyperplasia samples. A study demonstrated that miR-32 exerts oncogenic characteristics by targeting on both B-cell translocation gene 2 (BTG-2) and phosphoinositide-3-kinase interacting protein 1 (PIK3IP1), which controls the inhibition of PI3K, a well-known regulator of cell proliferation, migration and survival [89]. MiR-106b/miR-25 cluster is an oncomir cluster, identified by computational approach, that is found associated with prostate cancer progression by targeting CASP7 (caspase 7, apoptosis related cysteine peptidase) mRNA, which is down-expressed both in primary prostate cancer and metastatic lesions [90]. MiR-125b is an androgen receptor-induced miRNA. Its induction in prostate cancer cell line LNCaP inhibits apoptosis and enhances cell proliferation. It was demonstrated that miR-125b activates the p53 network by interrupting Mdm2 degradation via targeting p14ARF, which mediates the Mdm2 sequestration [91]. A summary of several oncogenic miRNAs found in prostate cancer is included in Table 1.
Table 1: Some of oncogenic MiRNAs involved in prostate cancer
Oncogenic miRNAs |
Function |
Reference |
|---|---|---|
miR-21 |
promotes tumor invasiveness and induces castration-resistance phenotype. |
|
miR-221/miR-222 |
enhance cell proliferation, invasion, cell survival, increase clonogenicity and enhance tumorigenicity in vivo. |
|
miR-141 |
is important in androgen-dependent and in metastatic castration-resistant. |
|
miR-375 |
is important for an early diagnosis. |
|
miR-18a |
promotes cancer progression. |
[85] |
miR-4534 |
induces pro-cancerous characteristics in non-cancer cell line. |
[87] |
miR-650 |
suppresses the cellular stress response 1 (CSR1) expression. |
[88] |
miR-32 |
inhibits apoptosis and enhances proliferation. |
[89] |
miR-106/miR-25 |
facilitate tumor progression. |
[90] |
miR-125b |
enhances cell proliferation and inhibits apoptosis. |
[91] |
Tumour suppressor MiRNAs
The involvement of miR-34 family as tumour suppressor in cancer has been reported by several investigations and many strategies to replace miR-34 have been developed in order to treat cancer [92, 93]. MiR-34a may exert its tumor suppressor role via targeting several signaling molecules involved in various stages of PCa progression. DNA damage and oncogenic stress strongly induced miR-34 expression in a p53-dependent manner. The distinctive features of p53 activity are promoted by miR-34 activation [94–95]: induction of cell cycle block and apoptosis through down-modulation of different proteins (CDK4, CDK6, cyclin D1, cyclin E2, E2F3 and BCL2) [96–98].
MiR-145 is a tumor suppressor and its expression is controlled by p53 pathway [99]. MiR-145 has been found down-expressed in breast cancer and colorectal cancer [100, 101]. Avgeris et al. found that higher risk for biochemical recurrence and significantly decreased disease-free survival were caused by miR-145 down-expression and that its decrease in prostate cancer was related to higher Gleason scores, advanced clinical stage, greater tumor diameter and higher PSA levels during follow-up [102]. Recently, it was discovered that miR-224 leads to migration and invasion of prostate cancer cells by targeting a miR-224 target gene, called apelin, whose inhibited expression allows the miRNA to inhibit cell migration and invasion. Moreover, miR-224 down-expression correlates significantly with advanced clinical stage and metastasis [103]. Kristensen et al. elucidated that proliferation, migration and invasion in prostate cancer cell lines PC3 and DU145 were inhibited by miR-224 and miR-452 through blocking cell cycle, cellular adhesion and motility [104]. Although the miR-200 family can be regulated at post-transcriptional levels, Zheng Zhang et al. found that ERG directly over-expresses the tumour suppressive miR-200b subfamily in human prostate tumours. In particular ERG can directly bind to miR-200b/a/429 promoter to facilitate its transcription in prostate cancer cells [105]. Tumour suppressor miR-382 represents an important new mechanism for understanding prostate cancer. It suppresses PCa cell proliferation, migration, invasion and metastasis through inhibiting COUP-TFII, including Snail and matrix metalloproteinase 2 (MMP2); in contrast, miR-382 suppression exhibited an opposite effect [106]. miR-372 has the ability to inhibit proliferation activity, migration, and invasion of DU145 cell line. One confirmed target of miR-372 is p65, whose knocked down expression induced a decrease in proliferation activity, migration and invasion. In both these processes CDK8, MMP-9, and prostate-specific antigen were involved [107]. miR-17-92a cluster miRNAs arises from a polycistronic transcription unit C13orf25 that generates six mature miRNAs: miR-17, miR-18a, miR-19a, miR-19b, miR-20a and miR-92a. These cluster miRNAs have a therapeutic benefit for both androgen-independent and -dependent PCa cells. Their expression are reduced in cancerous prostate tissues and also in aggressive prostate cancer cells. Restoration of expression of all members of miR-17-92a cluster showed depressed expression of cell cycle regulatory proteins cyclin D1 and SSH1 and inhibited EMT by reducing cell migration and mesenchymal markers expression and by increasing surface localization and expression of the E-Cadherin. Moreover, this cluster improves sensitivity of LNCaP androgen dependent prostate cancer cells to anti-androgen drug Casodex, AKT inhibitor MK-2206 2HCl and docetaxel [108]. miR-27a functions as tumour suppressor in PCa cell by repression of MAP2K4 which works as an oncogene in PCa cell lines [109]. Xu B et al. found that hsa-miR-135a-1 exerts its action as a potential tumour suppressor in metastatic PCa by targeting EGFR [110]. Human tumour suppressor miRNA-204-5p facilitates apoptosis by BCL2 in PCa cells. In miR-204-5p-transfected cells BCL2 mRNA and protein expression decreased and consequently cytochrome C delivery from mitochondria and caspase 3 and caspase 3/7 activation were induced [111]. Zhang et al. suggest that miR-30a may function as a novel tumour suppressor in castration-resistant prostate cancer (CRPC) tissues. Its anti-oncogenic activity may occur by the reduced expression of a distinct cell cycle protein, cyclin E2 (CCNE2) [112]. Let-7 gene encodes a highly conserved miRNA family, which are significantly down-expressed in localized prostate cancer. Let-7 has been shown to target oncogenes implicated in cell-cycle progression, cell proliferation, migration, differentiation and epithelial-to-mesenchymal transition (EMT) progression. Let-7 miRNA family exerts tumour suppressor characteristics via targeting multiple oncogenes including RAS, HMGA2, Ezh2, Lin28 and c-Myc [113]. Thus, loss of let-7 miRNAs correlates with prostate cancer progression [114]. It has been discovered that miR-133 and miR-146a suppress prostate cancer tumor progression via targeting EGFR, a tumour promoter for this disease. Thus, loss of miR-133 and miR-146a may be attributed to enhancement of EGFR signaling, leading to aggressive prostate cancer progression [115]. A summary of some tumor suppressor miRNAs in prostate cancer is included in Table 2.
Table 2: Some of suppressive miRNAs involved in prostate cancer
Tumour suppressor miRNAs |
Function |
Reference |
|---|---|---|
miR-34a |
induces cell-cycle arrest, cell senescence and apoptosis and inhibits cell proliferation and cell invasion. |
|
miR-145 |
inhibits invasion, migration and arrests cell cycle. |
|
miR-224 |
inhibits invasion and migration of PCa cells. |
[103] |
miR-452 |
regulates cell cycle, cellular adhesion and motility. |
[104] |
miR-200b |
inhibits PCa cell growth and invasion. |
[105] |
miR-382 |
inhibits PCa cell proliferation, migration, invasion and metastasis. |
[106] |
miR-372 |
inhibits proliferation, migration and invasion of DU145 cells. |
[107] |
miR-17-92a |
decreases cell cycle regulatory proteins and the expression of mesenchymal markers. |
[108] |
miR-27a |
suppresses MAP2K4 in PCa cell. |
[109] |
has-miR-135-a-1 |
inhibits cell growth, cell cycle progression, migration, invasion, and xenograft tumor formation. |
[110] |
miR-204-5p |
promotes apoptosis by targeting BCL2 in PCa cell. |
[111] |
miR-30a |
reduces expression of cell cycle protein, cyclin E2. |
[112] |
let-7 miRNAs |
regulate cell cycle, cell migration, cell proliferation and epithelial-to-mesenchymal transition progression. |
|
miR-133/miR-146a |
suppress tumor progression via targeting EGFR. |
[115] |
CONCLUSIONS
Prostate cancer is the secondary highest cause of malignancy mortality in male. It is estimated, in 2016, about two hundred thousand men will be diagnosed with PCa, and about twenty seven thousand will die of PCa. In this context it becomes extremely important to improve early diagnosis and treatment to reduce cancer-related mortality. In PCa there is a deregulation of several miRNAs that may act as oncogenes or tumour suppressors. The miRNA profiling studies demonstrate that mRNAs may act independently or in partnership with other transcription factors to regulate gene transcription, which ultimately leads to perturbed cellular processes in PCa. The knowledge of miRNAs biology, the study of their functions and molecular mechanisms in recent years are growing more and more. They are strongly involved in cancer development, in particular several miRNAs are implicated in cell transformation for the development of cancer cells, metastasis and treatment resistance. Many studies established that specific miRNAs are over-expressed or down-expressed concurrently in different types of human cancer and often related to specific cytogenetic abnormalities.
The most important strategies for miRNA-based therapeutics consist of a direct and an indirect method. Direct strategies include oligonucleotides or virus-based constructs in order to block the oncogenic miRNAs expression, while the indirect ones involve drugs to regulate miRNAs expression. Understanding the specific roles of some miRNAs and their involvement in prostate cancer development and progression could lead to improve the course of this cancer and could open new therapeutic options in prostate cancer treatment. Many discoveries have demonstrated that miRNAs are helpful tools to make diagnosis and prognosis of prostate cancer and can predict the clinical outcome of patients. To date, miRNAs can be evaluated using cheap technologies in serum of patients. Therefore, the development of easy kits to detect circulating miRNAs by nanotechnologic approach is necessary to provide the diffusion of these biomarkers in the national health system and could represent an important step forward in the field of molecular therapies [116].
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
GRANT SUPPORT
This work was supported by grant “Progetto ONCONET2.0 – Linea progettuale 14 per l’implementazione della prevenzione e diagnosi precoce del tumore alla prostata e testicolo – Regione Campania, Italy” (to G.F).
REFERENCES
1. Gronberg H. Prostate cancer epidemiology. Lancet. 2003; 361:859–64.
2. Caraglia M, Alaia C, Grimaldi A, Boccellino M, Quagliuolo L. MiRNA as prognostic and therapeutic targets in tumor of male urogenital tract. Mol Targets Strat Cancer Prev. 2016; 7:151–171.
3. Fong MK, Hare R, Jarkowski A. A new era for castrate resistant prostate cancer: a treatment review and update. J Oncol Pharm Pract. 2012; 18:343–354.
4. Rodrigues DN, Butler LM, Estelles DL, de Bono JS. Molecular pathology and prostate cancer therapeutics: from biology to bedside. J Pathol. 2013; 232:178–184.
5. Facchini G, Caraglia M, Morabito A, Marra M, Piccirillo MC, Bochicchio AM, Striano S, Marra L, Nasti G, Ferrari E, Leopardo D, Vitale G, Gentilini D, et al. Metronomic administration of zoledronic acid and taxotere combination in castration resistant prostate cancer patients. Phase I ZANTE Trial. Cancer Biol Ther. 2010; 10:543–8.
6. Marra M, Santini D, Tonini G, Meo G, Zappavigna S, Facchini G, Morabito A, Abbruzzese A, Cartenì G, Budillon A, Caraglia M. Molecular and preclinical models enhancing anti-tumour activity of zoledronic acid. Eur J Cancer. 2008; 6:79–85.
7. Caraglia M, Giuberti G, Marra M, Di Gennaro E, Facchini G, Caponigro F, Iaffaioli RV, Budillon A, Abruzzese A. Docetaxel induces P53-dependent apoptosis and synergizes with farnesyl transefrase inhibitor R115777 in human epithelial cancer cells. Front Biosci. 2005; 2566–2575.
8. Frasci G, Iaffaioli RV, Perone V, Facchini G, De Rosa G, Bianco AR. Prostatic cancer with cutaneous involvement treated with Goserelin. Curr Ther Res. 1991; 49:169–73.
9. Caffo O, Gernone A, Ortega C, Sava T, Cartenì G, Facchini G, Re GL, Amadio P, Bortolus R, Pagliarulo V, Prati V, Veccia A, Galligioni E. Central nervous system metastases from castration-resistant prostate cancer in the docetaxel era. J Neurooncol. 2012; 107:191–6.
10. Berretta M, Della Pepa C, Tralongo P, Fulvi A, Martellotta F, Lleshi A, Nasti G, Fisichella R, Romano C, De Divitiis C, Taibi R, Fiorica F, Di Francia R. Use of Complementary and Alternative Medicine (CAM) in cancer patients: An Italian multicenter survey. Oncotarget. 2016; 8:24401–24414. doi: 10.18632/oncotarget.14224.
11. Veccia A, Caffo O, De Giorgi U, Di Lorenzo G, Ortega C, Scognamiglio F, Aieta M, Facchini G, Mansueto G, Mattioli R, Procopio G, Zagonel V, D’Angelo A, et al. Clinical outcomes in octogenarians treated with docetaxel as first-line chemotherapy for castration-resistant prostate cancer. Future Oncology. 2016; 12:493–502.
12. Caffo O, De Giorgi U, Fratino L, Alesini D, Zagonel V, Facchini G, Gasparro D, Ortega C, Tucci M, Verderame F, Campadelli E, Lo Re G, Procopio G, et al. Clinical Outcomes of Castration-resistant Prostate Cancer Treatments Administered as Third or Fourth Line Following Failure of Docetaxel and Other Second-line Treatment: Results of an Italian Multicentre Study. Eur Urol. 2015; 68:147–153.
13. Della Pepa C, D’Aniello C, Rossetti S, Iovane G, Pisconti S, Fisichella R, Facchini G, Cavaliere C. “Real-life” effectiveness studies in mCRPC patients: systematic review. WCRJ. 2015; 2:e617.
14. Della Pepa C, Cavaliere C, Rossetti S, Di Napoli M, Cecere SC, Crispo A, De Sangro C, Rossi E, Turitto D, Germano D, Iovane G, Berretta M, D’Aniello C, et al. Predictive Comprehensive Geriatric Assessment in elderly prostate cancer patients: the prospective observational scoop trial results. Anticancer Drugs. 2017; 28:104–109.
15. Loberg RD, Logothetis CJ, Keller ET, Pienta KJ. Pathogenesis and treatment of prostate cancer bone metastases: targeting the lethal phenotype. J Clin Oncol. 2005; 23:8232–8241.
16. Di Franco R, Borzillo V, Ravo V, Ametrano G, Falivene S, Cammarota F, Rossetti S, D’Aniello C, Cavaliere C, Romano FJ, Malzone MG, Montanari M, Vanacore D, et al. Hypofractionated versus conventional radiotherapy in patients with low/intermediate-risk localized prostate cancer: review of rectal and urinary toxicity. WCRJ. 2016; 3: e751.
17. D’Aniello C, Cavaliere C, Fiorica F, Facchini G. Letter to the Editor - Unenhanced whole-body MRI versus PET-CT for the detection of prostate cancer metastases after primary treatment. Eur Rev Med Pharmacol Sci. 2016; 20:4626–4627.
18. Berretta M, Della Pepa C, Tralongo P, Fulvi A, Martellotta F, Lleshi A, Nasti G, Fisichella R, Romano C, De Divitiis C, Taibi R, Fiorica F, Di Francia R. Tumor infiltrating T lymphocytes expressing FoxP3, CCR7 or PD-1 predict the outcome of prostate cancer patients subjected to salvage radiotherapy after biochemical relapse. Cancer Biol Ther. 2016; 17:1213–1220.
19. Heidenreich A, Bellmunt J, Bolla M, Joniau S, Mason M, Matveev V, Mottet N, Schmid HP, van der Kwast T, Wiegel T, Zattoni F, European Association of Urology. EAU guidelines on prostate cancer. Part 1: screening, diagnosis, and treatment of clinically localised disease. Eur Urol. 2011; 59:61–71.
20. Loeb S, Catalona WJ. Prostate-specific antigen in clinical practice. Cancer Lett. 2007; 249:30–9.
21. Boccellino M, Alaia C, Misso G, Cossu AM, Facchini G, Piscitelli R, Quagliuolo L, Caraglia M. Gene interference strategies as a new tool for the treatment of prostate cancer. Endocrine. 2015; 49:588–605.
22. Facchini G, Caffo O, Ortega C, D'Aniello C, Di Napoli M, Cecere SC, Della Pepa C, Crispo A, Maines F, Ruatta F, Iovane G, Pisconti S, Montella M. Very Early PSA Response to Abiraterone in mCRPC Patients: a Novel Prognostic Factor Predicting Overall Survival. Front Pharmacol. 2016; 123.
23. Wang G, Chan ES, Kwan BC, Li PK, Yip SK, Szeto CC, Ng CF. Expression of microRNAs in the urine of patients with bladder cancer. Clin Genitourin Cancer. 2012; 10:106–13.
24. Lolli C, Caffo O, Scarpi E, Aieta M, Conteduca V, Maines F, Bianchi E, Massari F, Veccia A, Chiuri VE, Facchini G, De Giorgi U. Systemic Immune-Inflammation Index Predicts the Clinical Outcome in Patients With mCRPC Treated With Abiraterone. Front Pharmacol. 2016; 7:376.
25. Verzoni E, De Giorgi U, Derosa L, Caffo O, Boccardo F, Facchini G, Porcu L, De Vincenzo F, Zaniboni A, Chiuri VE, Fratino L, Santini D, Adamo V, et al. Predictors of long-term response to abiraterone in patients with metastastic castration-resistant prostate cancer: a retrospective cohort study. Oncotarget. 2016; 7:40085–40094. doi: 10.18632/oncotarget.9485.
26. Caffo O, Lo Re G, Sava T, Buti S, Sacco C, Basso U, Zustovich F, Lodde M, Perin A, Facchini G, Veccia A, Maines F, Barile C, et al. Intermittent docetaxel chemotherapy as first-line treatment for metastatic castration-resistant prostate cancer patients. Future Oncol. 2015; 11:965–973.
27. Schroder FH, Hugosson J, Roobol MJ, Tammela TL, Ciatto S, Nelen V, Kwiatkowski M, Lujan M, Lilja H, Zappa M, Denis LJ, Recker F, Berenguer A, et al. Screening and prostate-cancer mortality in a randomized European study. N Engl J Med. 2009; 360:1320–8.
28. Epstein JI. An update of the Gleason grading system. J Urol. 2010; 183:433–440.
29. Mellinger GT, Gleason D, Bailar J 3rd. The histology and prognosis of prostatic cancer. J Urol. 1967; 97:331–337.
30. Shen MM, Abate-Shen C. Molecular genetics of prostate cancer: new prospects for old challenges. Genes Dev. 2010; 24:1967–2000.
31. Squire JA, Park PC, Yoshimoto M, Alami J, Williams JL, Evans A, Joshua AM. Prostate cancer as a model system for genetic diversity in tumors. Adv Cancer Res. 2011; 112:183–216.
32. Schoenborn JR, Nelson P, Fang M. Genomic profiling defines subtypes of prostate cancer with the potential for therapeutic stratification. Clin Cancer Res. 2013; 19:4058–4066.
33. De Cobelli O, Buonerba C, Terracciano D, Danilo B, Lucarelli G, Bove P, Altieri V, Coman I, Perdonà S, Facchini G, Berretta M, Di Lorenzo G, Grieco P, et al. Urotensin II receptor on preoperative biopsy is associated with upstaging and upgrading in prostate cancer. Future Oncol. 2015; 11:3091–8.
34. Fraser M, Berlin A, Bristow RG, van der Kwast T. Genomic, pathological, and clinical heterogeneity as drivers of personalized medicine in prostate cancer. Urol Oncol. 2015; 33:85–94.
35. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004; 116:281–97.
36. Berezikov E, Chung WJ, Willis J, Cuppen E, Lai EC. Mammalian mirtron genes. Mol Cell. 2007; 28:328–336.
37. Okamura K, Hagen JW, Duan H, Tyler DM, Lai EC. The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell. 2007; 130:89–100.
38. Ruby JG, Jan CH, Bartel DP. Intronic microRNA precursors that bypass Drosha processing. Nature. 2007; 448:83–86.
39. Xie M, Li M, Vilborg A, Lee N, Shu MD, Yartseva V, Šestan N, Steitz JA. Mammalian 5′-capped microRNA precursors that generate a single microRNA. Cell. 2013; 155:1568–1580.
40. Cheloufi S, Dos Santos CO, Chong MM, Hannon GJ. A dicer-independent miRNA biogenesis pathway that requires Ago catalysis. Nature. 2010; 465:584–589.
41. Cifuentes D, Xue H, Taylor DW, Patnode H, Mishima Y, Cheloufi S, Ma E, Mane S, Hannon GJ, Lawson ND, Wolfe SA, Giraldez AJ. A novel miRNA processing pathway independent of Dicer requires Argonaute2 catalytic activity. Science. 2010; 328:1694–1698.
42. Yang JS, Maurin T, Robine N, Rasmussen KD, Jeffrey KL, Chandwani R, Papapetrou EP, Sadelain M, O’Carroll D, Lai EC. Conserved vertebrate mir-451 provides a platform for Dicerindependent, Ago2-mediated microRNA biogenesis. Proc Natl Acad Sci USA. 2010; 107:15163–15168.
43. Yoda M, Cifuentes D, Izumi N, Sakaguchi Y, Suzuki T, Giraldez AJ, Tomari Y. Poly(A)-specific ribonuclease mediates 3′-end trimming of Argonaute2-cleaved precursor microRNAs. Cell Rep. 2013; 5:715–726.
44. Watanabe Y, Tomita M, Kanai A. Computational methods for microRNA target prediction. Methods Enzymol. 2007; 427:65–86.
45. Saito T, Saetrom P. MicroRNAs – targeting and target prediction. N Biotechnol. 2010; 27:243–249.
46. Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell. 2007; 27:91–105.
47. Garcia DM, Baek D, Shin C, Bell GW, Grimson A, Bartel DP. Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat Struct Mol Biol. 2011; 18:1139–1146.
48. Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009; 19:92–105.
49. Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005; 120:15–20.
50. Jansson MD, Lund AH. MicroRNA and cancer. Mol Oncol. 2012; 6:590–610.
51. Hata A, Lieberman J. Dysregulation of microRNA biogenesis and gene silencing in cancer. Sci Signal. 2015;17;8:re3.
52. Wahid F, Shehzad A, Khan T, Kim YY. MicroRNAs: synthesis, mechanism, function, and recent clinical trials. Biochim Biophys Acta. 2010; 1803:1231–1243.
53. Lennox KA, Behlke MA. Chemical modification and design of anti-miRNA oligonucleotides. Gene Ther. 2011; 18:1111–1120.
54. Price C, Chen J. MicroRNAs in cancer biology and therapy: Current status and perspectives. Genes Dis. 2014; 1:53–63.
55. Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014; 20:460–469.
56. Garzon R, Marcucci G, Croce CM. Targeting microRNAs in cancer: rationale, strategies and challenges. Nat Rev Drug Discov. 2010; 9:775–789.
57. Hutzvanger G, Simard MJ, Mello CC, Zamore PD. Sequence specific inhibition of small RNA function. Plos Biol. 2004; 2:E98.
58. Weiler J, Hunziker J, Hall J. Anti-miRNA oligonucleotides (AMOs); amunition to target miRNAs implicated in human disease? Gene Therapy. 2006; 13:496–502.
59. Vester B, Wengel J. LNA (locked nucleic acid): high affinity targeting of complementary RNA and DNA. Biochemistry. 2004; 43:13233–13241.
60. Xiao J, Yang B, Lin H, Lu Y, Luo X, Wang Z. Novel approaches for gene-specific interference via manipulating actions of micro-RNAs: examination on the pacemaker channel genes HCN2 and HCN4. J Cell Physiol. 2007; 212:285–92.
61. Choi WY, Giraldez AJ, Schier AF. Target protectors reveal dampening and balancing of Nodal agonist and antagonist by miR-430. Science. 2007; 318:271–4.
62. Gumireddy K, Young DD, Xiong X, Hogenesch JB, Huang Q, Deiters A. Small-molecule inhibitors of microrna miR-21 function. Angew Chem Int Ed Engl. 2008; 47:7482–4.
63. Michelfelder S, Trepel M. Adeno-associated viral vectors and their redirection to cell-type specific receptors. Adv Genet. 2009; 67:29–60.
64. Aagaard L, Rossi JJ. RNAi therapeutics: Principles, prospects and challenges. Adv Drug Deliver Rev. 2007; 59:75–86.
65. Esquela-Kerscher A, Slack FJ. Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer. 2006; 6:259–269.
66. Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T, Shimotohno K. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene. 2006; 25 :2537–45.
67. He H, Jazdzewski K, Li W, Liyanarachchi S, Nagy R, Volinia S, Calin GA, Liu CG, Franssila K, Suster S, Kloos RT, Croce CM, de la Chapelle A. The role of microRNA genes in papillary thyroid carcinoma. PNAS USA. 2005; 102:19075–80.
68. Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T, Calin GA, Liu CG, Croce CM et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006; 9:189–98.
69. Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M, Prueitt RL, Yanaihara N, Lanza G, et al. 2006 A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA. 2006; 103:2257–61.
70. Franco R, Caraglia M, Facchini G, Abbruzzese A, Botti G. The role of tissue microarray in the era of target-based agents. Expert Rev Anticancer Ther. 2011; 11:859–69.
71. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006; 6:857–866.
72. Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA. MicroRNAs in body fluids–the mix of hormones and biomarkers. Nat Rev Clin Oncol. 2011; 8:467–477.
73. Blondal T, Jensby Nielsen S, Baker A, Andreasen D, Mouritzen P, Wrang Teilum M, Dahlsveen IK. Assessing sample and miRNA profile quality in serum and plasma or other biofluids. Methods. 2013; 59:S1-S6.
74. Pritchard CC, Cheng HH, Tewari M. MicroRNA profiling: approaches and considerations. Nat Rev Genet. 2012; 13:358–369.
75. Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435:834–838.
76. Li T, Li RS, Li YH, Zhong S, Chen YY, Zhang CM, Hu MM, Shen ZJ. miR-21 as an Independent Biochemical Recurrence Predictor and Potential Therapeutic Target for Prostate Cancer. J Urol. 2012; 187:1466–1472.
77. Cannistraci A, Di Pace AL, De Maria R, Bonci D. MicroRNA as new tools for prostate cancer risk. Assessment and therapeutic intervention: results from clinical data set and patients’ samples. Biomed Res Int. 2014; 146170. doi: 10.1155/2014/146170.
78. Mercatelli N, Coppola V, Bonci D, Miele F, Costantini A, Guadagnoli M, Bonanno E, Muto G, Frajese GV, De Maria R, Spagnoli LG, Farace MG, Ciafrè SA. The inhibition of the highly expressed miR-221 and miR-222 impairs the growth of prostate carcinoma xenografts in mice. PloS One. 2008; 3:e4029. doi:10.1371/journal.pone.0004029.
79. Le Sage C, Nagel R, Egan DA, Schrier M, Mesman E, Mangiola A, Anile C, Maira G, Mercatelli N, Ciafrè SA, Farace MG, Agami R. Regulation of the p27(Kip1) tumor suppressor by miR-221 and miR-222 promotes cancer cell proliferation. EMBO J. 2007; 26:3699–708.
80. Galardi S, Mercatelli N, Giorda E, Massalini S, Frajese GV, Ciafrè SA, Farace MG. miR-221 andmiR-222 expression affects the proliferation potential of human prostate carcinoma cell lines by targeting p27Kip1. J Biol Chem. 2007; 282:23716–24.
81. Zhang HL, Qin XJ, Cao DL, Zhu Y, Yao XD, Zhang SL, Dai B, Ye DW. An elevated serum miR-141 level in patients with bone-metastatic prostate cancer is correlated with more bone lesions. Asian J Androl. 2013; 15:231–5.
82. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A, Lin DW, Urban N, Drescher CW, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008; 105:10513–8.
83. Wen J, Li R, Wen X, Chou G, Lu J, Wang X, Jin Y. Dysregulation of cell cycle related genes and microRNAs distinguish the low- from high-risk of prostate cancer. Diagn Pathol. 2014; 9:156. doi: 10.1186/s13000-014-0156-1.
84. Bryant RJ, Pawlowski T, Catto JW, Marsden G, Vessella RL, Rhees B, Kuslich C, Visakorpi T, Hamdy FC. Changes in circulating microRNA levels associated with prostate cancer. Brit J Cancer. 2012; 106:768–774.
85. Al-Kafaji G, Al-Naieb ZT, Bakhiet M. Increased oncogenic microRNA-18a expression in the peripheral blood of patients with prostate cancer: A potential novel non-invasive biomarker. Oncol Lett. 2016; 11:1201–1206.
86. Rea D, Del Vecchio V, Palma G, Barbieri A, Falco M, Luciano A, De Biase D, Perdonà S, Facchini G, Arra C. Mouse Models in Prostate Cancer Translational Research: From Xenograft to PDX. Biomed Res Int. 2016.
87. Nip H, Dar AA, Saini S, Colden M, Varahram S, Chowdhary H, Yamamura S, Mitsui Y, Tanaka Y, Kato T, Hashimoto Y, Shiina M, Kulkarni P, et al. Oncogenic microRNA-4534 regulates PTEN pathway in prostate cancer. Oncotarget. 2016; 7:68371–68384. doi: 10.18632/oncotarget.12031.
88. Zuo ZH, Yu YP, Ding Y, Liu S, Martin A, Tseng G, Luo JH. Oncogenic Activity of miR-650 in Prostate Cancer Is Mediated by Suppression of CSR1 Expression. Am J Pathol. 2015; 185:1991–9.
89. Jalava SE, Urbanucci A, Latonen L, Waltering KK, Sahu B, Jänne OA, Seppälä J, Lähdesmäki H, TammelaTL, Visakorpi T. Androgenregulated miR-32 targets BTG2 and is overexpressed in castration-resistant prostate cancer. Oncogene. 2012; 31:4460–71.
90. Walter BA, Valera VA, Pinto PA, Merino MJ. Comprehensive microRNA profiling of prostate cancer. J Cancer. 2013; 4:350–7.
91. Amir S, Ma AH, Shi XB, Xue L, Kung HJ, Devere White RW. Oncomir miR-125b suppresses p14(ARF) to modulate p53-dependent and p53-independent apoptosis in prostate cancer. PloS One. 2013; 8:e61064.
92. Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE, Zhai Y, Giordano TJ, Qin ZS, Moore BB, MacDougald OA, Cho KR, Fearon ER. p53-mediated activation of miRNA34 candidate tumor suppressor genes. Curr Biol. 2007; 17:1298–1307.
93. He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y, Xue W, Zender L, Magnus J, Ridzon D, Jackson AL, Linsley PS, Chen C, et al. A microRNA component of the p53 tumour suppressor network. Nature. 2007; 447:1130–4.
94. Romano FJ, Rossetti S, Conteduca V, Schepisi G, Cavaliere C, Di Franco R, Lamantia E, Castaldo L, Nocerino F, Ametrano G, Cappuccio F, Malzone G, Montanari M, et al. Role of DNA repair machinery and p53 in the testicular germ cell cancer: a review. Oncotarget. 2016; 7:85641–85649. doi: 10.18632/oncotarget.13063.
95. Chieffi P, Franco R, Portella G. Molecular and cell biology of testicular germ cell tumors. Int. Rev Cell Mol Biol. 2009; 278: 277–308.
96. Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, Feldmann G, Yamakuchi M, Ferlito M, Lowenstein CJ, Arking DE, Beer MA, Maitra A, et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell. 2007; 26:745–52.
97. Raver-Shapira N, Marciano E, Meiri E, Spector Y, Rosenfeld N, Moskovits N, Bentwich Z, Oren M. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol Cell. 2007; 26:731–743.
98. Tazawa H, Tsuchiya N, Izumiya M, Nakagama H. Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. PNAS. 2007; 104:15472–15477.
99. Sachdeva M, Zhu SM, Wu FT, Wu HL, Walia V, Kumar S, Elble R, Watabe K, Mo YY. p53 represses c-Myc through induction of the tumor suppressor miR-145. PNAS USA. 2009; 106:3207–3212.
100. Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S, Magri E, Pedriali M, Fabbri M, Campiglio M, Menard S, Palazzo JP, Rosenberg A, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005; 65:7065–7070.
101. Slaby O, Svoboda M, Fabian P, Smerdova T, Knoflickova D, Bednarikova M, Nenutil R, Vyzula R. Altered expression of miR-21, miR- 31, miR-143 and miR-145 is related to clinicopathologic features of colorectal cancer. Oncology. 2007; 72:397–402.
102. Avgeris M, Stravodimos K, Fragoulis EG, Scorilas A. The loss of the tumour-suppressor miR-145 results in the shorter disease-free survival of prostate cancer patients. Brit J Cancer. 2013; 108:2573–2581.
103. Kim WT, Kim WJ. MicroRNAs in prostate cancer. Prostate Int. 2013; 1:3–9.
104. Karatas OF, Guzel E, Suer I, Ekici ID, Caskurlu T, Creighton CJ, Ittmann M, Ozen M. miR-1 and miR-133b Are Differentially Expressed in Patients with Recurrent Prostate Cancer. PLoS One. 2014; 9:e98675.
105. Zhang Z, Lanz RB, Xiao L, Wang L, Hartig SM, Ittmann MM, Feng Q, He B. The tumor suppressive miR-200b subfamily is an ERG target gene in human prostate tumors. Oncotarget. 2016; 7:37993–38003. doi: 10.18632/oncotarget.9366.
106. Zhang W, Liu J, Qiu J, Fu X, Tang Q, Yang F, Zhao Z, Wang H. MicroRNA-382 inhibits prostate cancer cell proliferation and metastasis through targeting COUP-TFII. Oncol Rep. 2016; 36:3707–3715.
107. Kong X, Qian X, Duan L, Liu H, Zhu Y, Qi J. microRNA-372 Suppresses Migration and Invasion by Targeting p65 in Human Prostate Cancer Cells. DNA Cell Biol. 2016; 35:828–835.
108. Ottman R, Levy J, Grizzle WE, Chakrabarti R. The other face of miR-17-92a cluster, exhibiting tumor suppressor effects in prostate cancer. Oncotarget. 2016; 7:73739–73753. doi:10.18632/oncotarget.12061.
109. Wan X, Huang W, Yang S, Zhang Y, Zhang P, Kong Z, Li T, Wu H, Jing F, Li Y. Androgen-induced miR-27A acted as a tumor suppressor by targeting MAP2K4 and mediated prostate cancer progression. Int J Biochem Cell Biol. 2016; 79:249–260.
110. Xu B, Tao T, Wang Y, Fang F, Huang Y, Chen S, Zhu W, Chen M. hsa-miR-135a-1 inhibits prostate cancer cell growth and migration by targeting EGFR. Tumor Biol. 2016; 37:14141–14151.
111. Lin YC, Lin JF, Tsai TF, Chou KY, Chen HE, Hwang TI. Tumor suppressor miRNA-204-5p promotes apoptosis by targeting BCL2 in prostate cancer cells. Asian J Surg. 2016; 9. pii:S1015-958430173–7.
112. Zhang L, Zhang XW, Liu CH, Lu K, Huang YQ, Wang YD, Xing L, Zhang LJ, Liu N, Jiang H, Sun C, Yang Y, Chen SQ, et al. miRNA-30a functions as a tumor suppressor by downregulating cyclin E2 expression in castration-resistant prostate cancer. Mol Med Rep. 2016; 14:2077–84.
113. Esposito F, Libertini S, Franco R, Abagnale A, Marra L, Portella G, Chieffi P. Aurora B expression in post-puberal testicular germ cell tumours. J Cell Physiol. 2009; 221:435–9.
114. Kong D, Heath E, Chen W, Cher ML, Powell I, Heilbrun L, Li Y, Ali S, Sethi S, Hassan O, Hwang C, Gupta N, Chitale D, et al. Loss of let-7 upregulates EZH2 in prostate cancer consistent with the acquisition of cancer stem cell signatures that are attenuated by BR-DIM. PloS One. 2012; 7:e33729.
115. Sherwood ER, Van Dongen JL, Wood CG, Liao S, Kozlowski JM, Lee C. Epidermal growth factor receptor activation in androgen-independent but not androgen-stimulated growth of human prostatic carcinoma cells. Br J Cancer. 1998; 77:855–61.
116. Vitale G, Dicitore A, Messina E, Sciammarella C, Faggiano A, Colao A. Epigenetics in Medullary Thyroid Cancer: From Pathogenesis to Targeted Therapy. Recent Pat Anticancer Drug Discov. 2016; 11:275–82.

