INTRODUCTION
The lack of useful biomarkers is one of the most important clinical problems of bone and soft tissue sarcomas. Osteosarcoma, the most common primary bone malignant tumor arising in children and young adults [1, 2], is no exception; the detection of the primary tumor or tumor relapse has generally relied on imaging methods such as X-ray, computed tomography (CT), positron emission tomography (PET)-CT, magnetic resonance imaging (MRI), and scintigraphy. Along with the development of surgical techniques [3, 4] and multi-agent chemotherapy [5, 6], patient prognosis has gradually improved over the past 30 years. Current multi-chemotherapeutic regimens, including neoadjuvant and adjuvant doxorubicin, cisplatin, methotrexate, and/or ifosfamide have maintained 5-year overall survival rates at approximately 60–80% [5–9]. However, for patients who present with local recurrence or metastasis, outcomes are far worse, with survival rates below 30% within 5 years of diagnosis [9, 10]. Furthermore, the histological response to neoadjuvant chemotherapy is the most dependable and reproducible prognostic indicator of the probability of disease-free survival [5–9]. Thus, development of methodologies for real-time monitoring of drug response and early detection of recurrence or metastasis will further improve patient prognosis. Currently, less complex monitoring using patient blood has not been developed. Alkaline phosphatase (ALP), a known serum-based tumor marker of osteosarcoma, sometimes provides false positives since ALP is generally elevated in children and affected by organ damage. Thus, the development of highly sensitive, specific, and minimally invasive biomarkers that can be used to detect and monitor tumor burden of tumors is the most important challenge for osteosarcoma management.
Evidence of microRNA (miRNA) dysregulation in malignant tumors has emerged in recent years. miRNAs are small non-coding RNA molecules that modulate the expression of multiple target genes and play important roles in various physiological and pathological processes, such as development, differentiation, cell proliferation, apoptosis, organogenesis, and homeostasis [11, 12]. After the discovery of miRNA in 1993 [13], its importance in malignant disease was suggested in 2004 when miRNA genes were found to be specifically deleted in leukemia [14, 15]. Subsequent reports have demonstrated that miRNAs are dysregulated in many malignant tumors, and these miRNAs can initiate carcinogenesis or drive progression [15, 16]. Following the discovery of miRNA dysregulation in cancer, miRNA dysregulation in osteosarcoma cells and/or tissues was reported since 2009 [17–44].
Tumor cells have recently been demonstrated to secrete miRNAs into the circulation [45]. Despite the presence of RNase activity in human blood, data have demonstrated that serum miRNAs remain stable under protection by exosomes or argonaute 2 [46, 47]. Analysis of circulating miRNA levels in patient blood presents a novel approach for diagnostic cancer screening or monitoring. Lawrie et al. was the first to report that tumor-associated miRNA levels in patient serum were higher than those in healthy individuals, indicating that circulating miRNAs may be used as biomarkers to monitor cancer cells [48]. To date, differential expression of circulating miRNA has been reported in cancers of the breast, lung, stomach, liver, kidney, bladder, prostate, and ovaries, among others [49–51]. This evidence on the potential of circulating miRNAs in patient blood is transforming the field of clinical biomarkers. However, comprehensive profiling studies of circulating miRNAs in osteosarcoma are limited and a consistent diagnostic signature for circulating miRNAs is not available.
In this study, we performed global miRNA screening in patient serum, following a validation study of in vitro and in vivo analysis. In addition, we evaluated whether the specified miRNA could monitor osteosarcoma and its drug response during multimodal treatment including chemotherapy and surgery.
RESULTS
Global miRNA expression analysis of serum in osteosarcoma patients, age-matched controls, and healthy individuals
This study was designed as an initial screening phase and a subsequent validation phase. The initial global miRNA screening included the serum samples from 10 osteosarcoma patients, 10 age-matched non-osteosarcoma patients, 10 healthy volunteers, and the culture medium from seven osteosarcoma cell lines (Supplementary Figure 1). The mean age of the osteosarcoma patients was 23.4 ± 13.4 years and that of the benign tumor patients was 20.0 ± 12.4 years; no significant difference in age between the two groups was observed (P = 0.56). The healthy volunteers were adults with a mean age of 32.4 ± 10.2 years. In the combined pool of human miRNAs, 236 serum miRNAs were identified to be highly expressed (>1.5 fold-change) in the serum of osteosarcoma patients compared with benign tumor patients and healthy volunteers. Among these miRNAs, eight miRNAs overlapped with the secretory miRNAs detected in the culture medium of the seven osteosarcoma cell lines (SaOS2, U2OS, HOS, MNNG/HOS, 143B, MG63, and HuO9) (Figure 1, Supplementary Table 1).
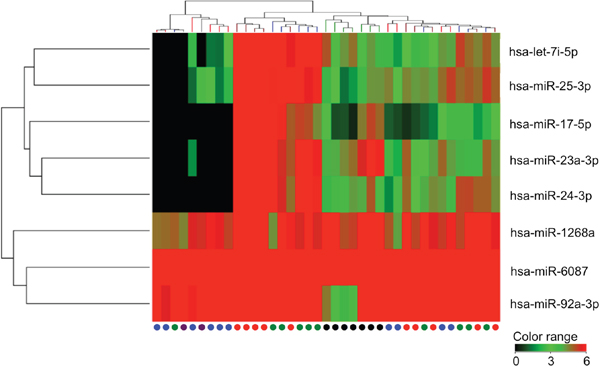
Figure 1: Hierarchical clustering of serum samples and culture media. Heatmap showing expression of candidate miRNAs that are upregulated in the serum of oseteosarcoma patients compared with controls and are expressed in the culture media of osteosarcoma cell lines. Serum from osteosarcoma patients at preoperative state and postoperative state are indicated with red and purple, respectively. Serum from patients with benign tumors and healthy individuals are indicated with green, and blue circles, respectively. Culture media from osteosarcoma cell lines are indicated with black circles.
Upregulated expression of miR-25a-3p and miR-17-5p in osteosarcoma cells and cell culture media
To determine whether these miRNAs acted in a secretory manner, we analyzed their expression levels in cultured human osteosarcoma cells and control normal cells, including human mesenchymal stem cells (MSCs) [52, 53] and HOB human osteoblast cells, as well as in culture media of these cells. qRT-PCR analyzes of RNA isolated from cell lines confirmed significant upregulation of miR-25-3p and miR-17-5p, but not other miRNAs (including miR-23a-3p), in both osteosarcoma cells and their culture media relative to normal control cells and their culture media (Figure 2A, 2B). Interestingly, the tendency of the expression levels of miR-25-3p and miR-17-5p between the cells and culture media were slightly different. Next, we determined whether miR-25-3p and miR-17-5p levels in the culture media were time course- and cell number-dependent. We observed that the expression levels of both miR-25-3p and miR-17-5p in the culture media from all seven cell lines increased with culture duration (24 and 48 h; P < 0.05) and with increasing numbers of tumor cells (P < 0.05) (Figure 2C, miR-17-5p; Figure 2D, miR-25-3p). These data suggest that miR-25-3p and miR-17-5p are secreted from human osteosarcoma cells.
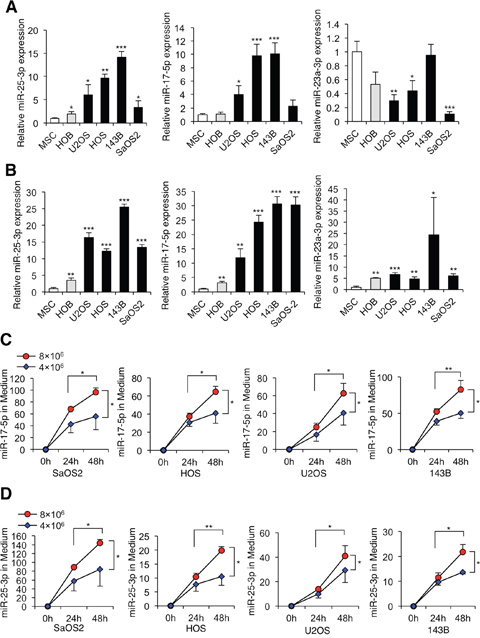
Figure 2: Secretory miRNAs from osteosarcoma cells. (A) The expression levels of miR-25-3p, miR-17-5p, and miR-23a3-p in human osteosarcoma cells (SaOS2, HOS, U2OS, and 143B), human mesenchymal stem cells, and human osteoblast cells. Data are presented as means ± standard deviations (SD; n = 3 per group). *, p < 0.05; **, p < 0.01; ***, p < 0.001; Student’s t test. (B) The expression of candidate miRNAs in the culture media of human osteosarcoma cells, human mesenchymal stem cells, and human osteoblast cells. Data are presented as means ± SD (n = 3 per group). *, p < 0.05; **, p < 0.01; ***, p < 0.001; Student’s t test. (C) The expression of miR-17-5p in the culture media of osteosarcoma cell lines (SaOS2, HOS, U2OS, and 143B). The expression levels of miR-17-5p in the media of all cell lines increased with cell counts and longer incubation time. Data are presented as mean ± SD (n = 3 per group). *, p < 0.05; **, p < 0.01; ***, p < 0.001; one-way ANOVA with Bonferroni’s multiple comparison. (D) The expression of miR-25-3p in the culture media of osteosarcoma cell lines. The expression levels of miR-25-3p in the media of all cell lines increased with cell counts and longer incubation time. Data are presented as mean ± SD (n = 3 per group). *, p < 0.05; **, p < 0.01; ***, p < 0.001; one-way ANOVA with Bonferroni’s multiple comparison.
miR-25-3p and miR-17-5p expression levels in exosomes derived from osteosarcoma cells
In order to analyze whether circulating miR-25-3p and miR-17-5p are embedded in tumor-derived exosomes, we purified exosomes from the culture medium of osteosarcoma cell lines (U2OS, HOS, 143B, and SaOS2) and MSCs. We then analyzed the expression levels of exosomal and cellular miR-25-3p by qRT-PCR and compared expression ratios. Our results demonstrated that miR-25-3p was expressed to a greater extent in the exosomes compared with donor cells; this tendency was not observed in MSCs (Supplementary Figure 2).
Expression profiles of serum miRNA levels during osteosarcoma development in tumor-bearing mice
To evaluate whether the serum expression levels of miR-25-3p and miR-17-5p could be used to monitor tumor dynamics in vivo, we examined the expression levels of these miRNAs in osteosarcoma-bearing mice. Serum miRNA expression levels were examined at different time points during tumor progression. Median serum levels in mice were plotted at day 0 before tumor cell inoculation and 3 weeks, 5 weeks, and 10 weeks after tumor cell inoculation. qRT-PCR analysis revealed that the expression levels of both miR-25-3p and miR-17-5p consistently increased with tumor development (Figure 3A, 3B). We also compared the expression levels of these miRNAs in pre- and post-operative states and identified that tumor resection decreased serum expression levels of miR-25-3p (Figure 3C). Among four mice with tumor resections, two developed lung metastasis 2 weeks after tumor resection. Interestingly, the expression levels of miR- 25-3p and miR-17-5p increased in the serum of the mice with lung metastasis (Figure 3D). These results suggest that these miRNAs reflect tumor burden in osteosarcoma-bearing mice.
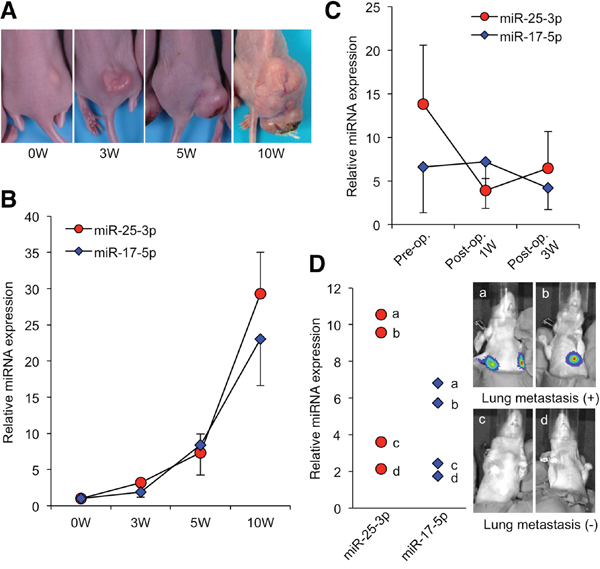
Figure 3: Serum miRNA levels during the development of osteosarcoma xenograft. (A) The macroscopic appearances of 143B-luc tumors in each group of mice 0, 3, 5, and 10 weeks after tumor inoculation. (B) The medians of the relative expression levels of miR-17-5p and miR-25-3p plotted at 0, 3, 5, and 10 weeks. Data are presented as mean ± SD (n = 4 per group). (C) The medians of the relative expression levels of miR-17-5p and miR-25-3p plotted before and after (1 and 3 weeks) tumor resection (n = 4 per group). (D) The serum miR-17-5p and miR-25-3p levels of each mouse 3 weeks after tumor resection. The expression levels of miR-25-3p and miR-17-5p were higher in the serum of the two mice with lung metastasis.
Evaluation of miR-25-3p and miR-17-5p expression levels in the serum of osteosarcoma patients
We next evaluated serum expression levels of miR-25-3p and miR-17-5p by qRT–PCR in a validation cohort of 14 osteosarcoma patients, 14 age-matched non-osteosarcoma patients, and eight healthy controls. Serum concentrations of these two miRNAs in osteosarcoma patients were statistically compared with those in non-osteosarcoma patients and healthy volunteers. Serum concentrations of miR-25-3p in osteosarcoma patients were significantly higher than in non-osteosarcoma patients (P = 0.004) and healthy volunteers (P = 0.004) (Figure 4A). Serum concentrations of miR-17-5p in osteosarcoma patients were also significantly higher than healthy volunteers (P = 0.021). However, we did not identify a statistical difference in miR-17-3p levels between osteosarcoma patients and non-osteosarcoma patients (P = 0.070) (Figure 4B). Then, we further analyzed serum miR-25-3p levels in patients with chondrosarcoma and Ewing sarcoma, which are second and third most common types of primary bone sarcoma after osteosarcoma, and observed elevated serum miR-25-3p levels in patients with osteosarcoma relative to those with other sarcomas (Supplementary Figure 3).
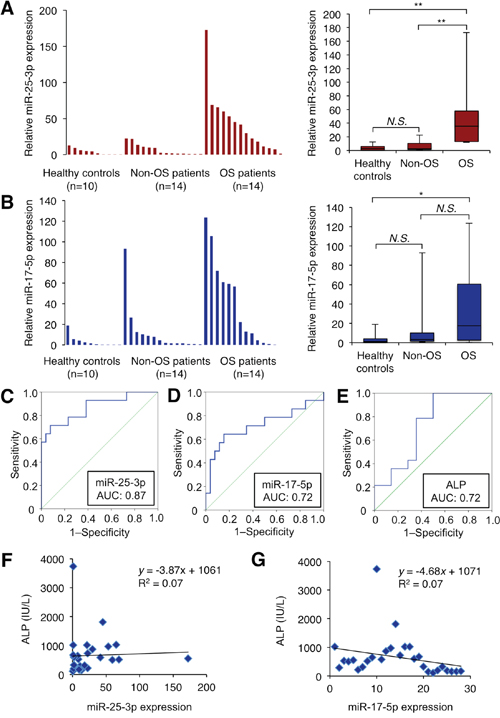
Figure 4: Serum miR-25-3p and miR-17-5p expression levels in the validation cohort of osteosarcoma patients. (A, B) Serum miR-25-3p concentrations in 14 consecutive osteosarcoma patients, 14 age-matched non-osteosarcoma patients, and 10 healthy individuals. Serum miR-25-3p (A) and miR-17-5p (B) expression was detectable in all samples (left). Differential expressions of serum miR-25-3p and miR-17-5p in osteosarcoma patients were compared with those of controls by a waterfall plot (right). Concentrations of miR-25-3p were significantly higher in serum from osteosarcoma patients than in controls (P = 0.004). Concentrations of miR-17-5p were significantly higher in serum from osteosarcoma patients than in healthy volunteers (P = 0.021) but there was no statistical significance (P = 0.070) compared to the age-matched non-osteosarcoma patients. *, P < 0.05; **, P < 0.01; one-way ANOVA with Bonferroni’s multiple comparison. (C, D) Receiver-operating characteristic (ROC) curve analysis of miR-25-3p and miR-17-5p for detecting osteosarcoma patients. ROC curve analysis showed the AUC of 0.868 (95% confidence interval, 0.743–0.993) for miR-25-3p (C) and 0.720 (95% confidence interval, 0.531–0.909) for miR-17-5p (D), respectively. The analysis revealed that the sensitivity and specificity of miR-25-3p was 71.4% and 92.3%, respectively, whereas the sensitivity and specificity of miR-17-5p was 64.3% and 84.6%, respectively. (E) The results of ROC curve analysis of ALP. ROC analysis revealed that the sensitivity and the specificity was 100% and 50%, respectively, with an AUC of 0.724 (95% confidence interval, 0.530–0.919). (F, G) Correlation between serum miR-25-3p and miR-17-5p concentrations and serum ALP levels in osteosarcoma patients. There was no significant correlation between serum miRNA concentrations and ALP levels (Pearson correlation scatter plot).
Next, we performed receiver-operating characteristic (ROC) curve analysis to assess the potential usefulness of serum miR-25-3p and miR-17-5p as noninvasive biomarkers for osteosarcoma. ROC analyses revealed that serum miR-25-3p levels were robust in discriminating patients with osteosarcoma from non-osteosarcoma patients and healthy volunteers, with an AUC value of 0.868 (95% confidence interval = 0.743 to 0.993) (Figure 4C). Using a cutoff value determined by the Youden Index [54], the sensitivity and specificity values for identifying a patient with osteosarcoma were 71.4% and 92.3%, respectively. For miR-17-5p, the AUC value from ROC analysis based on the expression levels was 0.720 (95% confidence interval = 0.743 to 0.993), and sensitivity and specificity values were 64.3% and 84.6%, respectively (Figure 4D).
We also compared the AUC based on the expression levels of these miRNAs with that of ALP, a conventional biomarker for osteosarcoma [55]. Interestingly, the AUC value of miR-25-3p levels was higher than that of ALP (0.724, 95% confidence interval = 0.530 to 0.919) (Figure 4E). These results indicate that serum levels of miR-25-3p may be a useful biomarker for differentiating patients with osteosarcoma from those with non-osteosarcoma and healthy individuals. Indeed, there was no significant correlation between serum miR-25-3p or miR-17-5p concentrations and serum ALP (Figure 4F, 4G).
Correlation between miR-25-3p and hematocytes in the patient blood
A recent report has cautioned that circulating miRNAs may reflect a blood-based phenomenon rather than a cancer-specific origin [56]. Accordingly, we analyzed the correlation between serum miR-25-3p and the hematocytes of peripheral blood in the validation cohort. No significant correlation between serum miR-25-3p concentrations and any type of hematocyte were observed (Figure 5A-5D). These results indicate that serum miR-25-3p may not be derived from hematocytes. Furthermore, serum miR-25-3p levels did not correlate with other data, including albumin (ALB), aspartate transaminase (AST), alanine transaminase (ALT), creatinine (Cr), and C-reactive protein (CRP) (Supplementary Figure 4).
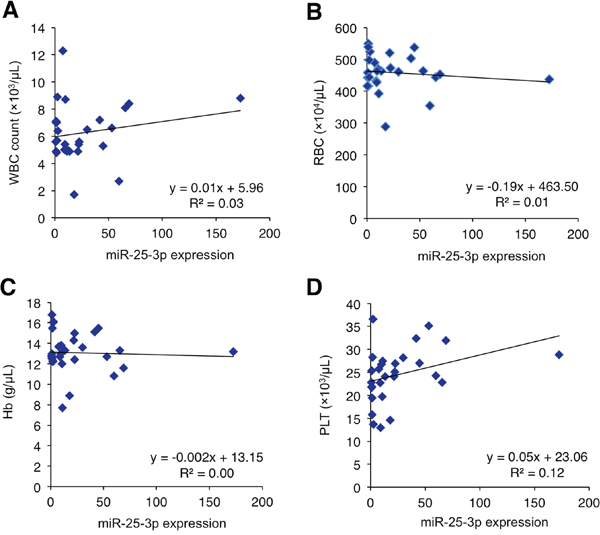
Figure 5: Correlation between serum miR-25-3p levels and the hematocytes of peripheral blood in the validation cohort of osteosarcoma patients. (A-D) There was no significant correlation between serum miR-25-3p levels and any type of peripheral hematocytes (Pearson correlation scatter plot).
Clinicopathological evaluation of serum miR-25-3p expression levels in osteosarcoma patients
In order to evaluate the clinical relevance of miR-25-3p in the serum of osteosarcoma patients, we analyzed its prognostic significance with regard to various clinicopathological factors of patients who could be followed up in the validation cohort (Figure 6 and Table 1). An analysis of disease-free survival was performed using the Kaplan–Meier approach and the log-rank test. Patients were divided into two groups according to their serum miR-25-3p expression level by ROC curve analysis (Supplementary Figure 5). The prognostic value of miR-25-3p expression levels at diagnosis revealed a significant difference in the survival between patients with higher and lower expression levels of serum miR-25-3p (P = 0.023; Figure 6). The 3-year metastasis-free survival rate in patients with lower serum miR-25-3p expression was 83.3%, whereas that of patients with higher miR-25-3p expression was 16.7%. According to univariate analysis, we found that metastasis at diagnosis, distant metastasis, and serum level of miR-25-3p all significantly impacted survival (P = 0.001, P = 0.004, and P = 0.023, respectively). In contrast, age, gender, tumor site, and response to chemotherapy did not significantly impact survival (P = 0.454, P = 0.775, P = 0.403, and P = 0.932, respectively).
Table 1: Clinicopathological correlation with metastasis-free survival based on the univariate analysis in the validation cohort of osteosarcoma patients
Variable |
Number of patients |
Metastasis free 3-year survival (%) |
P value |
|---|---|---|---|
Age (years) |
0.454 |
||
0–20 |
9 |
41.7 |
|
21+ |
3 |
66.7 |
|
Gender |
0.775 |
||
Male |
7 |
57.1 |
|
Female |
5 |
40.0 |
|
Site |
0.403 |
||
Femur |
8 |
33.3 |
|
Tibia |
2 |
50.0 |
|
Humerus |
2 |
100.0 |
|
Metastasis at diagnosis |
0.001 |
||
Present |
1 |
0.0 |
|
Absent |
11 |
53.0 |
|
Distant metastasis during follow-up |
0.004 |
||
Present |
7 |
0.0 |
|
Absent |
5 |
100.0 |
|
Response to neoadjuvant chemotherapy |
0.932 |
||
Good (necrosis > 90%) |
6 |
50.0 |
|
Poor (necrosis < 90%) |
6 |
50.0 |
|
miR-25-3p expression |
0.023 |
||
High |
6 |
16.7 |
|
Low |
6 |
83.3 |
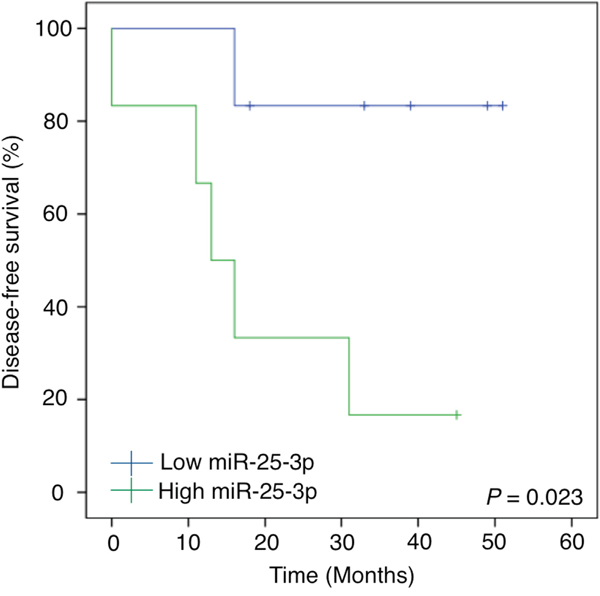
Figure 6: The prognostic value of serum miR-25-3p expression levels at diagnosis. Kaplan-Meier analysis and log-rank test of disease-free survival according to serum miR-25-3p levels. The 3-year metastasis-free survival rate for patients with lower serum miR-25-3p levels was 83.3%, whereas that for patients with higher miR-25-3p levels was 16.7% (P = 0.023; log-rank test).
Evaluation of serum miR-25-3p expression for monitoring tumor dynamics in osteosarcoma patients
To investigate whether serum miR-25-3p could monitor tumor dynamics in osteosarcoma patients, we evaluated serum miR-25-3p in several osteosarcoma patients according to the multi-modal treatment. Two cases (Case 1 and 2) could be monitored with their serum obtained at preoperative and postoperative status. Case 1 was a 17 year-old-male with osteosarcoma arising in the distal femur (Figure 7A). Both serum ALP and miR-25-3p expression decreased after tumor resection. Case 2 was an 8 year-old-female with osteosarcoma arising in the proximal tibia (Figure 7B). Similar to case 1, both serum ALP and miR-25-3p expression decreased after tumor resection. Four cases were monitored with their serum obtained at several points during treatment. Case 3 was a 24-year-old-male patient with osteosarcoma arising in the distal femur (Figure 7C). Neoadjuvant chemotherapy (high-dose methotrexate, doxorubicin, and cisplatin) did not reduce the SUVmax (11.22 to 20.81) and the subsequent 40% tumor necrosis rate in the resected specimens indicated a poor response. Serum ALP levels increased during chemotherapy (44%), whereas serum miR-25-3p levels decreased (54%), while both levels decreased after tumor resection. Regarding tumor necrosis, serum miR-25-3p levels, rather than serum ALP levels, appeared to correspond with the tumor burden. Case 4 was a 20-year-old male patient with a pathological fracture caused by osteosarcoma arising in the proximal femur (Figure 7D). Abnormal accumulation observed during thallium scintigraphy was reduced by neoadjuvant chemotherapy (high-dose methotrexate, doxorubicin, cisplatin, and ifosfamide), suggesting a favorable tumor response, and surviving tumor cells were not pathologically identified. Although his serum miR-25-3p levels decreased in response to neoadjuvant chemotherapy (70%), ALP levels did not change. Subsequently, serum miR-25-3p, rather than ALP, levels appeared to correlate with tumor necrosis. Case 5 was a 50-year-old male patient with osteosarcoma arising in the distal fibula (Figure 7E). Although decreased accumulation during thallium scintigraphy indicated a good tumor response to neoadjuvant chemotherapy (doxorubicin, cisplatin, and ifosfamide), the preoperative serum ALP levels plateaued. However, a slight decrease in serum miR-25-3p during chemotherapy (32%) was observed. Pathological evaluation of the resected specimens revealed 80% necrosis following chemotherapy, which correlated with a decrease in serum miR-25-3p, rather than ALP. Case 6 was a 24-year-old male patient with osteosarcoma arising in the calcaneous (Figure 7F). Neoadjuvant chemotherapy (high-dose methotrexate, doxorubicin, and cisplatin) reduced FDG-PET accumulation, indicating a good therapeutic response. Serum miR-25-3p levels also decreased during neoadjuvant chemotherapy (82%). However, ALP remained within the normal range at all time points and did not reflect drug response. Wide tumor resection did not reduce serum miR-25-3p levels; this finding corresponded with the >95% rate of tumor necrosis, suggesting that miR-25-3p might reflect viable cells and, therefore, drug response. Overall, data from these individualized cases suggest that serum miR-25-3p expression may be useful for tumor monitoring during multi-modal treatment.
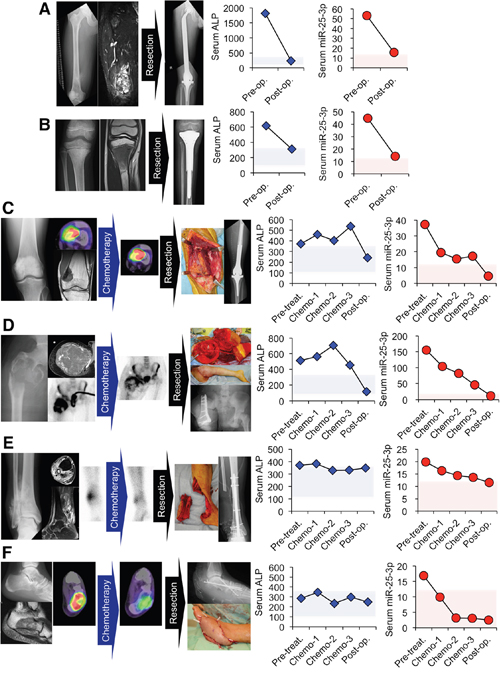
Figure 7: Tumor monitoring by serum miR-25-3p levels according to multi-modal treatment. (A, B) Two cases whose serum ALP and miR-25-3p could be evaluated before and after surgical resection. Both serum ALP and serum miR-25-3p in a 17 year-old-male (A) and an 8 year-old-female (B) decreased after surgical resection. (C-F) Four cases whose serum ALP and miR-25-3p could be evaluated during neoadjuvant chemotherapy. (C) A 24 year-old-male with osteosarcoma in the distal femur. Tumor necrosis by neoadjuvant chemotherapy was 40%. Serum ALP levels increased during chemotherapy, while serum miR-25-3p levels decreased; both levels were reduced after tumor resection. (D) A 20 year-old-male with osteosarcoma in the proximal femur. Surviving tumor cells were not identified in the pathological evaluation after neoadjuvant chemotherapy. ALP levels did not change, but serum miR-25-3p levels decreased during chemotherapy. (E) A 50 year-old-male with osteosarcoma arising in the distal fibula. While serum ALP levels before surgical resection plateaued, serum miR-25-3p slightly decreased during chemotherapy. The pathological evaluation of the resected specimens reported 80% necrosis. (F) A 24 year-old-male with osteosarcoma in the calcaneous. Neoadjuvant chemotherapy reduced FDG accumulation and serum miR-25-3p levels also decreased during chemotherapy. ALP was within the normal range at all time points. Serum miR-25-3p levels corresponded with tumor necrosis, which was revealed as more than 95%. Pale blue; normal range of ALP, pale red; miR-25-3p levels below cut-off point.
DISCUSSION
Clinical outcomes of osteosarcoma treatment have not significantly improved in over 20 years. To date, the most powerful predictors of outcome have remained the ability to detect metastatic disease at diagnosis and the histopathologic response of the tumor to neoadjuvant chemotherapy. Despite steady progress in the identification of genetic alterations in osteosarcoma, no individual molecular marker has been demonstrated to monitor drug response or have a better prognostic significance than the current clinical markers. Recent studies have identified that circulating mature miRNA in contrast to mRNA or snRNA is strikingly stable in the blood stream and cell culture medium [46, 47]. These circulating miRNAs were discovered in the peripheral blood of patients with malignant diseases and have provided novel insights into the biology of tumors and the effects of therapeutic interventions [46, 47].
To date, emerging evidence has begun to accumulate on miRNA dysregulation in osteosarcoma cells and/or tissues [26, 38]. Several studies have indicated that some of the miRNAs dysregulated in tumor tissue may exist in the blood samples of osteosarcoma patient [57–59]. Importantly, these studies have simply focused on the miRNA whose expression is dysregulated in osteosarcoma cells or tissues. However, there is no consensus whether or not miRNA dysregulation in tumor cells or tissues reflects those in the patient’s blood. Indeed, recent investigations have demonstrated that miRNA profiling between tumor tissue and corresponding serum is largely dissimilar [60–62]. Pigati et al. performed miRNA microarray analysis of cellular and extracellular RNAs of MCF7, a breast cancer cell line, and found that approximately 66% of released miRNAs closely reflected cellular miRNAs [61]. Similarly, Chan et al. demonstrated that, among 20 miRNAs differentially expressed in breast cancer tumors compared with adjacent normal tissues, only seven miRNAs were upregulated in both tumors and serum [60]. These data suggest the existence of a cellular selection mechanism for miRNA release and indicates that released miRNAs do not necessarily reflect the abundance of miRNA in the cell of origin. Therefore, a circulating miRNA signature that shows favorable sensitivity and specificity could be detected by global expression analysis of patient’s serum rather than the simple focus on dysregulated miRNAs in tumor cells or tissues. Indeed, this study, based on global miRNA screening, revealed that several oncogenic miRNAs in osteosarcoma cells and/or tissues such as miR-21 [63] and miR-214 [34, 64] were not detected as highly upregulated miRNAs in the patient serum, which is similar to other cancers [60–62]. Thus, this study is important as the first global miRNA screening of patient serum that detected circulating miRNA, with favorable sensitivity and specificity compared to the conventional marker.
In this study, global miRNA screening of patient serum, followed by in vitro analysis, detected miR-17-5p and miR-25-3p as secretory and circulating miRNAs in osteosarcoma patients. These miRNAs have also been reported as dysregulated miRNAs in osteosarcoma cells and/or tissues. The miR-17-92 cluster was among the first miRNAs to be validated as showing oncogenic potential [65, 66]. To date, the oncogenic function of miR-17-92 in osteosarcoma has been demonstrated by several researchers [24, 67–69]. On the other hand, upregulated miR-25 levels in osteosarcoma tissues compared with adjacent healthy tissues has been reported and its overexpression promoted cell proliferation and tumor growth [70]. In this study, we found that the sensitivity and specificity of serum miR-25-3p concentrations was superior to those of miR-17-5p. Notably, the relationship between the dysregulation of the miR-17-92 cluster and aging was reported [71], although the mechanistic connection between downregulation of the cluster’s members and aging has yet to be elucidated [71]. Hackl et al. investigated different cell and tissue types representing aging and identified the downregulation of miR-17, miR-19b, miR-20a, and miR-106a [72]. Considering these reports, serum miR-25-3p expression levels would be a suitable non-invasive biomarker compared with serum miR-17-5p concentrations in osteosarcoma patients.
Various reports have shown that miR-25 plays an oncogenic role in esophageal cancer [73], cholangiocarcinoma [74], gastric cancer [75], and lung cancer [76], whereas other reports have indicated that miR-25 may function as a tumor suppressor in colon cancer and anaplastic thyroid carcinoma [77, 78]. Thus, whether miR-25 acts as an oncogene or a tumor suppressor may depend on the cellular context. Recent reports have shown that circulating miR-25 has diagnostic and/or prognostic value in patients with lung and hepatocellular cancer [79–81]. Therefore, this is the first evidence of the value of circulating miR-25 expression in mesenchymal malignancies, which enlarges the utility of this miRNA for various malignancies. In this study, we identified that there was no significant correlation between serum miR-25-3p concentrations and any type of hematocyte, which is consistent with other reports [56], while the use of whole blood may lead to the isolation of miRNAs from various cell types including those within the blood cells. Although Pitchard et al. suggested caution in the interpretation of circulating miRNAs as they may reflect a blood cell-based phenomenon rather than a cancer-specific origin, they demonstrated no correlation between miR-25, blood cells, and hemolysis [56]. Furthermore, while recent studies have demonstrated age-related miRNAs such as miR-21, miR-126-3p, miR-151a-5p, miR-181a-5p, and miR-1248 [71, 72, 82, 83], there has been no correlation of miR-25 and aging. Indeed, recent reports have demonstrated that miR-25 is upregulated in osteo-differentiated MSCs compared with MSCs [84], suggesting the correlation between miR-25 expression and bone metabolism, which may be different from ALP-related miRNAs including miR-27, miR-133, and miR-206 [85]. In this study, serum miR-25-3p concentration was a novel prognostic factor for osteosarcoma patients, which correlated with clinical distant metastasis. However, there was no statistical significance between the serum expression of this marker at diagnosis and drug sensitivity, as well as the other clinicopathological elements, in the validation cohort (Supplementary Table 4), which might be due to the small number of patients. An additional external validation based on a larger cohort is needed as the next step, which may yield clinically important information whether the detected serum miRNA could be a non-invasive biomarker of osteosarcoma.
Recent reports have demonstrated that most of the circulating miRNAs are included in lipid or lipoprotein complexes, such as apoptotic bodies, microvesicles, or exosomes, and are, therefore, highly stable [86]. In this study, we demonstrated that both miR-25-3p and miR-17-5p were enriched in the exosomes derived from osteosarcoma cell lines, indicating that they might be stable in the patient serum. One of the limitations of this study is that it was impossible to analyze the exosomal expression levels of miR-25-3p and miR-17-5p in patient serum because of the limited amount of serum that had been preserved. Indeed, the extraction procedures of exosomes from patient serum in clinical practice require additional time and efforts, which might inhibit the standardization of this methodology. Therefore, the simple procedure, without extraction of exosomes, could be practical for clinical application. On the other hand, liquid biopsies based on other methods, such as measurements of exosomes or circulating tumor cells (CTCs), have been reported. Suetsugu et al. successfully imaged tumor-derived exosomes in tumor-bearing mouse models by observing GFP-tagged CD63, a general exosomal marker, and found that these elements were incorporated into tumor-associated cells and circulated in the blood of mice with metastatic breast cancer [87]. Yoshioka et al. identified CD147 as a exosomal marker in colorectal cancer and demonstrated a sensitive and rapid ExoScreen-based analytical technique. Therefore, our future research plans include the use of tumor-derived exosomes as a novel methods of liquid biopsy. In addition, various trials have suggested the utility of CTCs as a liquid biopsy target. Shigeyasu et al. developed a method to capture live CTCs using a green fluorescent protein (GFP)-expressing telomerase-specific adenovirus and successfully imaged epithelial and mesenchymal tumor cells in clinical blood samples from patients with colorectal cancer [88]. Indeed, the roles of CTCs in tumor development have been evident based on the molecular imaging technologies in various malignancies including prostate, lung, and breast cancers [89–95]. To date, osteosarcoma CTCs have been detected by several authors [96] but have not been applied for liquid biopsy, which represents a future challenge.
In this study, we compared serum miRNA levels before and after tumor resection in a subcutaneous xenograft osteosarcoma model. To date, various reports have described molecular mechanisms of sarcoma metastasis based on orthotopic sarcoma models [97–100]. Indeed, we initially used an orthotopic osteosarcoma model. However, the orthotopically xenografted mice died after tumor resection (amputation) following complications such as bleeding or infection. In contrast, all mice with subcutaneous xenografts remained alive after tumor resection. Therefore, subcutaneous tumor inoculation might be less harmful and appropriate if the study protocol includes tumor resection and subsequent observation. As no reports have described differences in miRNA expression between orthotopic and non-orthotopic models, secretory miRNA profiling of in vivo osteosarcoma models would be an interesting step toward elucidating the roles of secretory miRNA in tumor tissue–microenvironment interactions.
In conclusion, serum-based miRNA signatures associated with osteosarcoma were successfully documented and validated. miR-25-3p was secreted from the established osteosarcoma cell lines and detected in tumor-derived exosomes. Serum miR-25-3p levels were upregulated in osteosarcoma patients and showed favorable sensitivity and specificity, correlating with poor prognosis. Furthermore, these concentrations closely correlated with tumor burden in animal models as well as with the therapeutic status of osteosarcoma patients. The clinical development of this methodology as a noninvasive diagnostic or monitoring strategy may be promising for clinicians and patients with malignant diseases.
MATERIALS AND METHODS
Patients and samples
Blood samples were collected from osteosarcoma patients, age-matched non-osteosarcoma patients with other benign tumors including enchondroma, non-ossifying fibroma, solitary bone cyst, lipoma, and hemangioma, and healthy volunteers at the National Cancer Center Hospital and Okayama University Hospital. Patient clinical information is summarized in Supplementary Tables 2, 3. Post-operative samples from three patients were obtained at 1–2 weeks after tumor-wide resection. Post-chemotherapy samples were obtained at the end of the cource. Serum samples were stored at –80°C until further processing. All patients provided written, informed consent authorizing the collection and use of their samples for research purposes. The study protocol was approved by the Institutional Review Board of the National Cancer Center Hospital and Okayama University Hospital.
Cell lines and cell culture
The human osteosarcoma cell lines SaOS2, U2OS, HOS, and 143B were purchased from the American Type Culture Collection (ATCC). Human mesenchymal stem cells and human osteoblast cell line (HOB) were purchased from Takara Bio. The human osteosarcoma cell line 143B-luc was previously established in our laboratory [101]. We cultured SaOS2 in RPMI 1640 (Life Technologies). U2OS, HOS, 143B, and 143B-luc cells were cultured in DMEM (Life Technologies). All media were supplemented with 10% heat-inactivated FBS (Life Technologies), penicillin (100 U/mL), and streptomycin (100 mg/mL). Cells were maintained under 5% CO2 in a humidified incubator at 37°C.
Isolation of exosomes
Conditioned medium was collected from 48-h cell cultures and centrifuged at 2,000 x g for 10 min, followed by filtration through a 0.22-μm filter to remove cell debris. The conditioned medium was then used for exosome isolation. Exosomes were harvested by ultracentrifugation at 110,000 x g for 70 min at 4°C. Pellets were washed with phosphate-buffered saline (PBS), subject to ultracentrifugation, and resuspended in PBS.
RNA extraction
Total RNA was extracted from 800 μL of serum and culture media samples using miRNeasy RNA isolation Kits (Qiagen). Total RNA was isolated from tissue specimens, cultured cells, and exosomal pellets using the TRIzol reagent (Invitrogen), followed by homogenization by pipetting up and down several times. To allow for normalization of sample-to-sample variation, 25 fmol of synthetic C. elegans miRNA cel-miR-39 (5 μL of 5 nM miRNA mimic) (Qiagen) was added to the denatured samples. Total RNA in serum samples was then extracted according to the manufacturer’s protocol.
miRNA array
Global miRNA expression profiling was performed using a miRNA microarray manufactured by Agilent Technologies (Santa Clara). Labeling and hybridization of total RNA samples were performed according to the manufacturer’s protocol. Each microarray experiment used 2 ng of miRNA containing total RNA. The microarray results were extracted using Agilent Feature Extraction software (v10.7.3.1) and analyzed using GeneSpring 12.6.1 software (Agilent Technologies).
Quantitative reverse-transcription polymerase chain reaction
The expression levels of miRNAs in osteosarcoma cells were analyzed by TaqMan quantitative real-time PCR (TaqMan MicroRNA Assays; Applied Biosystems) using a 7300 Real-Time PCR system (Applied Biosystems) according to the manufacturer’s protocol. Relative expression was calculated using the 2–ΔΔCt method [102].
Animal experiments
The animal experiments in this study were performed in compliance with the guidelines of the Institute for Laboratory Animal Research at Okayama University. Athymic nude mice (CLEA Japan) were purchased at 4 weeks of age and given at least 1 week to adapt to their new environment prior to tumor transplantation. On day 0, the mice were anesthetized with 3% isoflurane, and the right leg was disinfected with 70% ethanol. A 100 μL volume of solution containing 143B-luc cells (1 × 106) was subcutaneously injected. The size of tumor growth was monitored, and whole blood was taken by cardiac puncture from mice at different times (3, 5, and 10 weeks after tumor transplantation) or after tumor resection. Blood samples were allowed to stand at room temperature for at least 1 h and for a maximum of 2 h. Mouse serum was separated from clots by centrifugation at 3500 rpm for 15 min at 4°C and stored at –80°C.
Statistical analysis
All statistical analyses were performed using the SPSS statistical software package (version 23; SPSS Inc.). Student’s t-test or one-way ANOVA, corrected for multiple comparisons as appropriate, were used to compare differences in cellular, secretory, and serum miRNA concentrations and miRNA ratios between the osteosarcoma, age-matched control, and healthy control groups. Receiver-operating characteristic (ROC) curves and area under the curve (AUC) were used to assess the feasibility of using serum miRNA concentrations as a diagnostic tool for detecting osteosarcoma. The Younden Index was used to determine the cutoff value for serum miRNA concentrations [103]. The Kaplan–Meier method and the log-rank test were used to compare the survival of patients. For all analyses, we considered a P value of 0.05 or less to be significant.
Author contributions
Conception and design: T. Fujiwara
Development of methodology: T. Ochiya
Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): T. Fujiwara, K. Uotani, A. Yoshida, T. Morita
Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): T. Fujiwara, A. Yoshida
Writing, review, and/or revision of the manuscript: T. Fujiwara
Administrative, technical, or material support (i.e., reporting or organizing data, constructing databases): Y. Nezu, A. Yoshida, T. Uehara, T. Omori, T. Komatsubara, K. Sugiu, K. Takeda, T. Kunisada, M. Kawamura, A. Kawai, T. Ozaki
Study supervision: T. Ochiya
ACKNOWLEDGMENTS
The authors are supported by the Grant-in-Aid of Japan Orthopaedics and Traumatology Research Foundation, Inc. No. 311, the NOVARTIS Foundation for the Promotion of Science, JSPS KAKENHI Grant Number 16K20054, and JSPS KAKENHI Grant Number 16H05449. The authors also acknowledge the National Cancer Center Biobank for providing the serum samples, all patients who agreed to this study project, and Dr. Shintaro Iwata from the Division of Orthopaedic Surgery, Chiba Cancer Center, for helpful discussions.
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
REFERENCES
1. Baldini N, Scotlandi K, Barbanti-Bròdano G, Manara MC, Maurici D, Bacci G, Bertoni F, Picci P, Sottili S, Campanacci M. Expression of P-glycoprotein in high-grade osteosarcomas in relation to clinical outcome. New England Journal of Medicine. 1995; 333:1380-1385.
2. Mirabello L, Troisi RJ, Savage SA. Osteosarcoma incidence and survival rates from 1973 to 2004: data from the Surveillance, Epidemiology, and End Results Program. Cancer. 2009; 115:1531-1543.
3. Gupta A, Meswania J, Pollock R, Cannon SR, Briggs TW, Taylor S, Blunn G. Non-invasive distal femoral expandable endoprosthesis for limb-salvage surgery in paediatric tumours. The Journal of bone and joint surgery British volume. 2006; 88:649-654.
4. Kawaguchi N, Ahmed AR, Matsumoto S, Manabe J, Matsushita Y. The concept of curative margin in surgery for bone and soft tissue sarcoma. Clinical orthopaedics and related research. 2004; 419:165-172.
5. Bielack SS, Kempf-Bielack B, Delling G, Exner GU, Flege S, Helmke K, Kotz R, Salzer-Kuntschik M, Werner M, Winkelmann W. Prognostic factors in high-grade osteosarcoma of the extremities or trunk: an analysis of 1,702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. Journal of clinical oncology. 2002; 20:776-790.
6. Ferrari S, Smeland S, Mercuri M, Bertoni F, Longhi A, Ruggieri P, Alvegard TA, Picci P, Capanna R, Bernini G, Muller C, Tienghi A, Wiebe T, et al. Neoadjuvant chemotherapy with high-dose Ifosfamide, high-dose methotrexate, cisplatin, and doxorubicin for patients with localized osteosarcoma of the extremity: a joint study by the Italian and Scandinavian Sarcoma Groups. Journal of clinical oncology. 2005; 23:8845-8852.
7. Bacci G, Longhi A, Versari M, Mercuri M, Briccoli A, Picci P. Prognostic factors for osteosarcoma of the extremity treated with neoadjuvant chemotherapy: 15-year experience in 789 patients treated at a single institution. Cancer. 2006; 106:1154-1161.
8. Iwamoto Y, Tanaka K, Isu K, Kawai A, Tatezaki SI, Ishii T, Kushida K, Beppu Y, Usui M, Tateishi A. Multiinstitutional phase II study of neoadjuvant chemotherapy for osteosarcoma (NECO study) in Japan: NECO-93J and NECO-95J. Journal of Orthopaedic Science. 2009; 14:397-404.
9. Smith MA, Seibel NL, Altekruse SF, Ries LA, Melbert DL, O'Leary M, Smith FO, Reaman GH. Outcomes for children and adolescents with cancer: challenges for the twenty-first century. Journal of clinical oncology. 2010; 28:2625-2634.
10. Bacci G, Rocca M, Salone M, Balladelli A, Ferrari S, Palmerini E, Forni C, Briccoli A. High grade osteosarcoma of the extremities with lung metastases at presentation: treatment with neoadjuvant chemotherapy and simultaneous resection of primary and metastatic lesions. Journal of surgical oncology. 2008; 98:415-420.
11. Ambros V. microRNAs: tiny regulators with great potential. Cell. 2001; 107:823-826.
12. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004; 116:281-297.
13. Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993; 75:843-854.
14. Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proceedings of the National Academy of Sciences. 2002; 99:15524-15529.
15. Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nature reviews Genetics. 2009; 10:704-714.
16. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nature reviews Cancer. 2006; 6:857-866.
17. He C, Xiong J, Xu X, Lu W, Liu L, Xiao D, Wang D. Functional elucidation of MiR-34 in osteosarcoma cells and primary tumor samples. Biochemical and biophysical research communications. 2009; 388:35-40.
18. Song B, Wang Y, Xi Y, Kudo K, Bruheim S, Botchkina GI, Gavin E, Wan Y, Formentini A, Kornmann M. Mechanism of chemoresistance mediated by miR-140 in human osteosarcoma and colon cancer cells. Oncogene. 2009; 28:4065-4074.
19. Creighton CJ, Fountain MD, Yu Z, Nagaraja AK, Zhu H, Khan M, Olokpa E, Zariff A, Gunaratne PH, Matzuk MM. Molecular profiling uncovers a p53-associated role for microRNA-31 in inhibiting the proliferation of serous ovarian carcinomas and other cancers. Cancer research. 2010; 70:1906-1915.
20. Zhang H, Cai X, Wang Y, Tang H, Tong D, Ji F. microRNA-143, down-regulated in osteosarcoma, promotes apoptosis and suppresses tumorigenicity by targeting Bcl-2. Oncology reports. 2010; 24:1363-1369.
21. Duan Z, Choy E, Harmon D, Liu X, Susa M, Mankin H, Hornicek F. MicroRNA-199a-3p is downregulated in human osteosarcoma and regulates cell proliferation and migration. Molecular Cancer Therapeutics. 2011; 10:1337-1345.
22. Gougelet A, Pissaloux D, Besse A, Perez J, Duc A, Dutour A, Blay JY, Alberti L. Micro-RNA profiles in osteosarcoma as a predictive tool for ifosfamide response. International Journal of Cancer. 2011; 129:680-690.
23. Maire G, Martin JW, Yoshimoto M, Chilton-MacNeill S, Zielenska M, Squire JA. Analysis of miRNA-gene expression-genomic profiles reveals complex mechanisms of microRNA deregulation in osteosarcoma. Cancer genetics. 2011; 204:138-146.
24. Huang G, Nishimoto K, Zhou Z, Hughes D, Kleinerman ES. miR-20a encoded by the miR-17–92 cluster increases the metastatic potential of osteosarcoma cells by regulating Fas expression. Cancer research. 2012; 72:908-916.
25. Jones KB, Salah Z, Del Mare S, Galasso M, Gaudio E, Nuovo GJ, Lovat F, LeBlanc K, Palatini J, Randall RL. miRNA Signatures Associate with Pathogenesis and Progression of Osteosarcoma. Cancer research. 2012; 72:1865-1877.
26. Kobayashi E, Hornicek FJ, Duan Z. MicroRNA Involvement in Osteosarcoma. Sarcoma. 2012; 2012:359739.
27. Zhu J, Feng Y, Ke Z, Yang Z, Zhou J, Huang X, Wang L. Down-regulation of miR-183 promotes migration and invasion of osteosarcoma by targeting Ezrin. The American journal of pathology. 2012; 180:2440-2451.
28. Chen L, Wang Q, Wang GD, Wang HS, Huang Y, Liu XM, Cai XH. miR-16 inhibits cell proliferation by targeting IGF1R and the Raf1-MEK1/2-ERK1/2 pathway in osteosarcoma. FEBS letters. 2013; 587:1366-1372.
29. Fujiwara T, Katsuda T, Hagiwara K, Kosaka N, Yoshioka Y, Takahashi Ru, Takeshita F, Kubota D, Kondo T, Ichikawa H. Clinical relevance and therapeutic significance of microRNA-133a expression profiles and functions in malignant osteosarcoma-initiating cells. Stem Cells. 2013; 32:959-73.
30. Kuijjer ML, Hogendoorn PC, Cleton-Jansen AM. Genome-wide analyses on high-grade osteosarcoma: making sense of a genomically most unstable tumor. International journal of cancer. 2013; 133:2512-2521.
31. Lauvrak SU, Munthe E, Kresse SH, Stratford EW, Namlos HM, Meza-Zepeda LA, Myklebost O. Functional characterisation of osteosarcoma cell lines and identification of mRNAs and miRNAs associated with aggressive cancer phenotypes. British journal of cancer. 2013; 109:2228-2236.
32. van der Deen M, Taipaleenmaki H, Zhang Y, Teplyuk NM, Gupta A, Cinghu S, Shogren K, Maran A, Yaszemski MJ, Ling L, Cool SM, Leong DT, Dierkes C, et al. MicroRNA-34c inversely couples the biological functions of the runt-related transcription factor RUNX2 and the tumor suppressor p53 in osteosarcoma. The Journal of biological chemistry. 2013; 288:21307-21319.
33. Wang Y, Zhao W, Fu Q. miR-335 suppresses migration and invasion by targeting ROCK1 in osteosarcoma cells. Molecular and cellular biochemistry. 2013; 384:105-111.
34. Wang Z, Cai H, Lin L, Tang M, Cai H. Upregulated expression of microRNA-214 is linked to tumor progression and adverse prognosis in pediatric osteosarcoma. Pediatric blood & cancer. 2013; 61:206-10.
35. Won KY, Kim YW, Kim HS, Lee SK, Jung WW, Park YK. MicroRNA-199b-5p is involved in the Notch signaling pathway in osteosarcoma. Human pathology. 2013; 44:1648-1655.
36. Zhao G, Cai C, Yang T, Qiu X, Liao B, Li W, Ji Z, Zhao J, Zhao H, Guo M. MicroRNA-221 Induces Cell Survival and Cisplatin Resistance through PI3K/Akt Pathway in Human Osteosarcoma. PloS one. 2013; 8:e53906.
37. Zhao H, Ma B, Wang Y, Han T, Zheng L, Sun C, Liu T, Zhang Y, Qiu X, Fan Q. miR-34a inhibits the metastasis of osteosarcoma cells by repressing the expression of CD44. Oncology reports. 2013; 29:1027-1036.
38. Zhou G, Shi X, Zhang J, Wu S, Zhao J. MicroRNAs in osteosarcoma: from biological players to clinical contributors, a review. The Journal of international medical research. 2013; 41:1-12.
39. Zhou X, Wei M, Wang W. MicroRNA-340 suppresses osteosarcoma tumor growth and metastasis by directly targeting ROCK1. Biochem Biophys Res Commun. 2013; 437:653-658.
40. Liu Z, Zhang G, Li J, Liu J, Lv P. The Tumor-Suppressive MicroRNA-135b Targets c-Myc in Osteoscarcoma. PloS one. 2014; 9:e102621.
41. Mu Y, Zhang H, Che L, Li K. Clinical significance of microRNA-183/Ezrin axis in judging the prognosis of patients with osteosarcoma. Med Oncol. 2014; 31:821.
42. Shen L, Wang P, Yang J, Li X. MicroRNA-217 regulates WASF3 expression and suppresses tumor growth and metastasis in osteosarcoma. PloS one. 2014; 9:e109138.
43. Wang Z, Cai H, Lin L, Tang M, Cai H. Upregulated expression of microRNA-214 is linked to tumor progression and adverse prognosis in pediatric osteosarcoma. Pediatric blood & cancer. 2014; 61:206-210.
44. Zhang K, Zhang Y, Ren K, Zhao G, Yan K, Ma B. MicroRNA-101 inhibits the metastasis of osteosarcoma cells by downregulation of EZH2 expression. Oncology reports. 2014; 32:2143-2149.
45. Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecologic oncology. 2008; 110:13-21.
46. Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nature reviews Cancer. 2011; 11:426-437.
47. Schwarzenbach H, Nishida N, Calin GA, Pantel K. Clinical relevance of circulating cell-free microRNAs in cancer. Nature reviews Clinical oncology. 2014; 11:145-156.
48. Lawrie CH, Gal S, Dunlop HM, Pushkaran B, Liggins AP, Pulford K, Banham AH, Pezzella F, Boultwood J, Wainscoat JS, Hatton CS, Harris AL. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br J Haematol. 2008; 141:672-675.
49. Agaoglu FY, Kovancilar M, Dizdar Y, Darendeliler E, Holdenrieder S, Dalay N, Gezer U. Investigation of miR-21, miR-141, and miR-221 in blood circulation of patients with prostate cancer. Tumor Biology. 2011; 32:583-588.
50. Cheng H, Zhang L, Cogdell DE, Zheng H, Schetter AJ, Nykter M, Harris CC, Chen K, Hamilton SR, Zhang W. Circulating plasma MiR-141 is a novel biomarker for metastatic colon cancer and predicts poor prognosis. PloS one. 2011; 6:e17745.
51. Heneghan HM, Miller N, Lowery AJ, Sweeney KJ, Newell J, Kerin MJ. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Annals of surgery. 2010; 251:499-505.
52. Mohseny AB, Szuhai K, Romeo S, Buddingh EP, Briaire-de Bruijn I, de Jong D, van Pel M, Cleton-Jansen AM, Hogendoorn PC. Osteosarcoma originates from mesenchymal stem cells in consequence of aneuploidization and genomic loss of Cdkn2. The Journal of pathology. 2009; 219:294-305.
53. Tataria M, Quarto N, Longaker MT, Sylvester KG. Absence of the p53 tumor suppressor gene promotes osteogenesis in mesenchymal stem cells. Journal of pediatric surgery. 2006; 41:624-632.
54. Youden W. Index for rating diagnostic tests. Cancer. 1950; 3:32-35.
55. Bacci G, Picci P, Orlandi M, Avella M, Manfrini M, Pignatti G, Dallari D, Manduchi R. Prognostic value of serum alkaline phosphatase in osteosarcoma. Tumori. 1987; 73:331-336.
56. Pritchard CC, Kroh E, Wood B, Arroyo JD, Dougherty KJ, Miyaji MM, Tait JF, Tewari M. Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer prevention research. 2012; 5:492-497.
57. Yuan J, Chen L, Chen X, Sun W, Zhou X. Identification of Serum MicroRNA-21 as a Biomarker for Chemosensitivity and Prognosis in Human Osteosarcoma. Journal of International Medical Research. 2012; 40:2090-2097.
58. Ouyang L, Liu P, Yang S, Ye S, Xu W, Liu X. A three-plasma miRNA signature serves as novel biomarkers for osteosarcoma. Med Oncol. 2013; 30:340.
59. Tian Q, Jia J, Ling S, Liu Y, Yang S, Shao Z. A causal role for circulating miR-34b in osteosarcoma. European journal of surgical oncology. 2013; 40: 67-72.
60. Chan M, Liaw CS, Ji SM, Tan HH, Wong CY, Thike AA, Tan PH, Ho GH, Lee AS. Identification of circulating microRNA signatures for breast cancer detection. Clinical cancer research. 2013; 19:4477-4487.
61. Pigati L, Yaddanapudi SC, Iyengar R, Kim DJ, Hearn SA, Danforth D, Hastings ML, Duelli DM. Selective release of microRNA species from normal and malignant mammary epithelial cells. PloS one. 2010; 5:e13515.
62. Wu Q, Lu Z, Li H, Lu J, Guo L, Ge Q. Next-generation sequencing of microRNAs for breast cancer detection. Journal of biomedicine & biotechnology. 2011; 2011:597145.
63. Ziyan W, Shuhua Y, Xiufang W, Xiaoyun L. MicroRNA-21 is involved in osteosarcoma cell invasion and migration. Medical Oncology. 2011; 28:1469-1474.
64. Xu Z, Wang T. miR-214 promotes the proliferation and invasion of osteosarcoma cells through direct suppression of LZTS1. Biochem Biophys Res Commun. 2014; 449:190-195.
65. He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, Powers S, Cordon-Cardo C, Lowe SW, Hannon GJ. A microRNA polycistron as a potential human oncogene. Nature. 2005; 435:828-833.
66. Uziel T, Karginov FV, Xie S, Parker JS, Wang YD, Gajjar A, He L, Ellison D, Gilbertson RJ, Hannon G, Roussel MF. The miR-17~92 cluster collaborates with the Sonic Hedgehog pathway in medulloblastoma. Proceedings of the National Academy of Sciences of the United States of America. 2009; 106:2812-2817.
67. Baumhoer D, Zillmer S, Unger K, Rosemann M, Atkinson MJ, Irmler M, Beckers J, Siggelkow H, von Luettichau I, Jundt G. MicroRNA profiling with correlation to gene expression revealed the oncogenic miR-17-92 cluster to be up-regulated in osteosarcoma. Cancer genetics. 2012; 205:212-219.
68. Gao Y, Luo LH, Li S, Yang C. miR-17 inhibitor suppressed osteosarcoma tumor growth and metastasis via increasing PTEN expression. Biochemical and biophysical research communications. 2014; 444:230-234.
69. Li X, Yang H, Tian Q, Liu Y, Weng Y. Upregulation of microRNA-17-92 cluster associates with tumor progression and prognosis in osteosarcoma. Neoplasma. 2014; 61:453-60.
70. Wang XH, Cai P, Wang MH, Wang Z. microRNA-25 promotes osteosarcoma cell proliferation by targeting the cell-cycle inhibitor p27. Molecular medicine reports. 2014; 10:855-859.
71. Grillari J, Hackl M, Grillari-Voglauer R. miR-17-92 cluster: ups and downs in cancer and aging. Biogerontology. 2010; 11:501-506.
72. Hackl M, Brunner S, Fortschegger K, Schreiner C, Micutkova L, Mück C, Laschober GT, Lepperdinger G, Sampson N, Berger P. miR-17, miR-19b, miR-20a, and miR-106a are down-regulated in human aging. Aging cell. 2010; 9:291-296.
73. Xu X, Chen Z, Zhao X, Wang J, Ding D, Wang Z, Tan F, Tan X, Zhou F, Sun J, Sun N, Gao Y, Shao K, et al. MicroRNA-25 promotes cell migration and invasion in esophageal squamous cell carcinoma. Biochem Biophys Res Commun. 2012; 421:640-645.
74. Razumilava N, Bronk SF, Smoot RL, Fingas CD, Werneburg NW, Roberts LR, Mott JL. miR-25 targets TNF-related apoptosis inducing ligand (TRAIL) death receptor-4 and promotes apoptosis resistance in cholangiocarcinoma. Hepatology. 2012; 55:465-475.
75. Zhao H, Wang Y, Yang L, Jiang R, Li W. MiR-25 promotes gastric cancer cells growth and motility by targeting RECK. Molecular and cellular biochemistry. 2014; 385:207-213.
76. Wu T, Chen W, Kong D, Li X, Lu H, Liu S, Wang J, Du L, Kong Q, Huang X, Lu Z. miR-25 targets the modulator of apoptosis 1 gene in lung cancer. Carcinogenesis. 2015; 36:925-935.
77. Esposito F, Tornincasa M, Pallante P, Federico A, Borbone E, Pierantoni GM, Fusco A. Down-regulation of the miR-25 and miR-30d contributes to the development of anaplastic thyroid carcinoma targeting the polycomb protein EZH2. The Journal of Clinical Endocrinology & Metabolism. 2012; 97:E710-E718.
78. Li Q, Zou C, Zou C, Han Z, Xiao H, Wei H, Wang W, Zhang L, Zhang X, Tang Q. MicroRNA-25 functions as a potential tumor suppressor in colon cancer by targeting Smad7. Cancer letters. 2013; 335:168-174.
79. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Q, Li X, Wang W, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008; 18:997-1006.
80. Chen X, Hu Z, Wang W, Ba Y, Ma L, Zhang C, Wang C, Ren Z, Zhao Y, Wu S, Zhuang R, Zhang Y, Hu H, et al. Identification of ten serum microRNAs from a genome-wide serum microRNA expression profile as novel noninvasive biomarkers for nonsmall cell lung cancer diagnosis. International journal of cancer. 2012; 130:1620-1628.
81. Li LM, Hu ZB, Zhou ZX, Chen X, Liu FY, Zhang JF, Shen HB, Zhang CY, Zen K. Serum microRNA profiles serve as novel biomarkers for HBV infection and diagnosis of HBV-positive hepatocarcinoma. Cancer research. 2010; 70:9798-9807.
82. Bonafè M, Olivieri F. Circulating microRNAs in aging. Oncotarget. 2015; 6:1340-1341. doi: 10.18632/oncotarget.3175.
83. Zhang Y, Mattison JA, Becker KG, Zonderman AB, Evans MK. Age-related changes in microRNA levels in serum. Aging (Albany NY). 2013; 5. doi: 10.18632/aging.100603.
84. Gao J, Yang T, Han J, Yan K, Qiu X, Zhou Y, Fan Q, Ma B. MicroRNA expression during osteogenic differentiation of human multipotent mesenchymal stromal cells from bone marrow. Journal of cellular biochemistry. 2011; 112:1844-1856.
85. Kapinas K, Delany AM. MicroRNA biogenesis and regulation of bone remodeling. Arthritis Res Ther. 2011; 13:220.
86. Kosaka N, Iguchi H, Ochiya T. Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010; 101:2087-2092.
87. Suetsugu A, Honma K, Saji S, Moriwaki H, Ochiya T, Hoffman RM. Imaging exosome transfer from breast cancer cells to stroma at metastatic sites in orthotopic nude-mouse models. Adv Drug Deliv Rev. 2013; 65:383-390.
88. Shigeyasu K, Tazawa H, Hashimoto Y, Mori Y, Nishizaki M, Kishimoto H, Nagasaka T, Kuroda S, Urata Y, Goel A. Fluorescence virus-guided capturing system of human colorectal circulating tumour cells for non-invasive companion diagnostics. Gut. 2014:gutjnl-2014-306957.
89. Glinskii AB, Smith BA, Jiang P, Li XM, Yang M, Hoffman RM, Glinsky GV. Viable circulating metastatic cells produced in orthotopic but not ectopic prostate cancer models. Cancer research. 2003; 63:4239-4243.
90. Berezovskaya O, Schimmer AD, Glinskii AB, Pinilla C, Hoffman RM, Reed JC, Glinsky GV. Increased expression of apoptosis inhibitor protein XIAP contributes to anoikis resistance of circulating human prostate cancer metastasis precursor cells. Cancer research. 2005; 65:2378-2386.
91. Glinsky GV, Glinskii AB, Berezovskaya O, Smith BA, Jiang P, Li XM, Yang M, Hoffman RM. Dual-color-coded imaging of viable circulating prostate carcinoma cells reveals genetic exchange between tumor cells in vivo, contributing to highly metastatic phenotypes. Cell Cycle. 2006; 5:191-197.
92. Kolostova K, Pinterova D, Hoffman RM, Bobek V. Circulating human prostate cancer cells from an orthotopic mouse model rapidly captured by immunomagnetic beads and imaged by GFP expression. Anticancer research. 2011; 31:1535-1539.
93. Bobek V, Hoffman RM, Kolostova K. Site-specific cytomorphology of disseminated PC-3 prostate cancer cells visualized in vivo with fluorescent proteins. Diagnostic cytopathology. 2013; 41:413-417.
94. Kolostova K, Zhang Y, Hoffman RM, Bobek V. In vitro culture and characterization of human lung cancer circulating tumor cells isolated by size exclusion from an orthotopic nude-mouse model expressing fluorescent protein. Journal of fluorescence. 2014; 24:1531-1536.
95. Kolostova K, Zhang Y, Hoffman RM, Bobek V. Morphological and gene-expression characterization of viable heterogeneous circulating tumor cells size-captured and cultured from triple-negative breast cancer mouse models. Int J Clin Exp Med. 2016; 9:7772-7779.
96. Chang L, Asatrian G, Dry SM, James AW. Circulating tumor cells in sarcomas: a brief review. Med Oncol. 2015; 32:430.
97. Tome Y, Tsuchiya H, Hayashi K, Yamauchi K, Sugimoto N, Kanaya F, Tomita K, Hoffman RM. In vivo gene transfer between interacting human osteosarcoma cell lines is associated with acquisition of enhanced metastatic potential. Journal of cellular biochemistry. 2009; 108:362-367.
98. Tome Y, Kimura H, Maehara H, Sugimoto N, Bouvet M, Tsuchiya H, Kanaya F, Hoffman RM. High lung-metastatic variant of human osteosarcoma cells, selected by passage of lung metastasis in nude mice, is associated with increased expression of αvβ3 integrin. Anticancer research. 2013; 33:3623-3627.
99. Miwa S, Hiroshima Y, Yano S, Zhang Y, Matsumoto Y, Uehara F, Yamamoto M, Kimura H, Hayashi K, Bouvet M, Tsuchiya H, Hoffman RM. Fluorescence-guided surgery improves outcome in an orthotopic osteosarcoma nude-mouse model. Journal of orthopaedic research. 2014; 32:1596-1601.
100. Miwa S, Zhang Y, Baek KE, Uehara F, Yano S, Yamamoto M, Hiroshima Y, Matsumoto Y, Kimura H, Hayashi K. Inhibition of spontaneous and experimental lung metastasis of soft-tissue sarcoma by tumor-targeting Salmonella typhimurium A1-R. Oncotarget. 2014; 5:12849-12861. doi: 10.18632/oncotarget.2561.
101. Osaki M, Takeshita F, Sugimoto Y, Kosaka N, Yamamoto Y, Yoshioka Y, Kobayashi E, Yamada T, Kawai A, Inoue T. MicroRNA-143 regulates human osteosarcoma metastasis by regulating matrix metalloprotease-13 expression. Molecular Therapy. 2011; 19:1123-1130.
102. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods. 2001; 25:402-408.
103. Akobeng AK. Understanding diagnostic tests 3: receiver operating characteristic curves. Acta paediatrica. 2007; 96:644-647.

