INTRODUCTION
Chromosomal translocations that lead to gene fusions are well-known cancer-causing genetic events that are actively used in clinical cancer diagnosis, and their gene products have been shown to be effective targets for directed therapy [1–3]. Prominent examples include BCR-ABL in chronic myelogenous leukemia [4] with the development of Gleevec as a paradigm for targeted therapy [5], frequent gene fusion TMPRSS2-ERG in prostate cancer [6], and the rapid targeting of ALK gene fusion products with crizotinib after the discovery of EML4-ALK in lung cancer [7, 8]. The success of these discoveries has led to the prevailing view that gene fusions and fusion products (RNA and protein) are generated due to chromosomal rearrangement at the DNA level, and thus are unique to cancer. However, others and we have shown that fusion transcripts can also be detected in normal human cell lines [9] and tissues [10–17]. They may be products of intergenic splicing instead of traditional chromosomal rearrangement [18–20]. In a recent study, we found a large number of fusion RNAs by analyzing nearly 300 RNA-Seq libraries, covering 30 different non-neoplastic human tissues and cells [17]. From the study, we identified 291 recurrent fusions, 51 in more than five tissue types. Among them, DUS4L-BCAP29, was found in six different tissues.
The DUS4L-BCAP29 fusion was previously discovered in gastric and prostate cancers [21, 22], and has been deposited in the Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer in the Cancer Genome Anatomy Project as a cancer-fusion. It was also reported to play a cancer-promoting role in gastric cancer [22], and proposed to be used as a cancer biomarker. However, our results showing its presence in multiple non-neoplastic tissues and cells, raise questions about its biomarker potential and challenge its relevance in tumorigenesis.
RESULTS
DUS4L-BCAP29 is widely expressed in non-neoplastic human tissues and cell lines
DUS4L-BCAP29 has been reported in both gastric and prostate cancers [21, 22]. However, recently, by analyzing RNA-Seq datasets, we detected the DUS4L-BCAP29 fusion transcript in multiple non-neoplastic tissues [17]. To confirm the finding that the fusion transcript exists in non-cancerous tissues, we designed primers flanking the fusion junction site, and used RT-PCR to detect the fusion in our collection of normal tissues. As shown in Figure 1A, the fusion RNA was indeed detected in multiple tissues, ranging from heart to testis (Figure 1A). Furthermore, it is expressed in diverse non-cancerous cell lines, including mammary gland (MCF10A), lung epithelial (Beas2B and 16HBE), and foreskin fibroblast (HFF) (Figure 1B). It is the same fusion form, involving the first seven exons of the DUS4L gene, and the last seven exons of BCAP29 as previously reported [22] (Supplementary Figure 1). The junction sequence is also identical to that reported in the gastric cancer study [22] (Figure 1C).
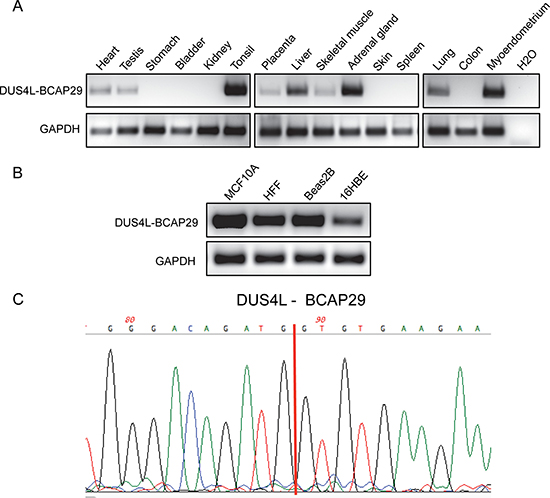
Figure 1: Detection of DUS4L-BCAP29 in non-cancer human tissues and cell lines. (A) Detection of the DUS4L-BCAP29 in several non-cancer human tissues by RT-PCR and followed by agarose electrophoresis. GAPDH was used as internal control. (B) Detection of the DUS4L-BCAP29 in several non-cancer cell lines by RT-PCR and followed by agarose electrophoresis. GAPDH was used as internal control. (C) Sanger sequencing validation of the RT-PCR product. Red line marks the fusion junction site.
DUS4L-BCAP29 is not significantly overexpressed in gastric or prostate cancer cells
To test whether the fusion RNA is expressed at a much higher level in cancer vs. non-cancer cell lines, we used qRT-PCR to quantify the difference of expression in several gastric and prostate non-cancer (GES-1 and RWPE-1), and cancer lines (SGC-7901, HGC-27, LNCaP, and PC3). In the gastric cells, contradictory to the previous report [22], DUS4L-BCAP29 is expressed at a comparable level in GES-1 as in SGC-7901, and even lower in HGC-27 (Figure 2A). In the prostate cells, the fusion is indeed expressed at lower levels in RWPE-1, than in LNCaP and PC3 cells (Figure 2A). We then compared the fusion RNA expression in clinical samples. No statistical difference was observed between the 21 gastric prostate cancer and normal matched pairs (Figure 2B). Similarly, no statistical difference was seen in 18 prostate cancer and 18 non-cancer prostate tissue samples (Figure 2B).
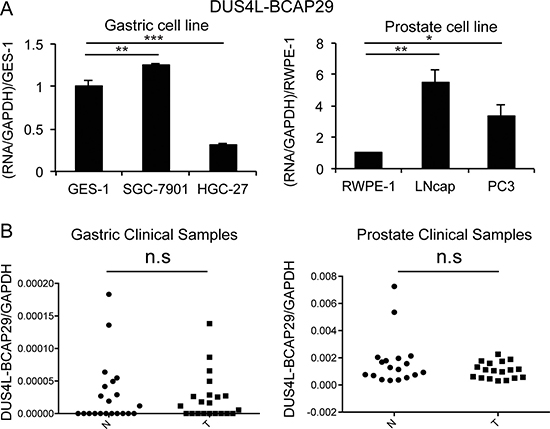
Figure 2: Quantification of DUS4L-BCAP29 expression in gastric and prostate tissues and cell lines. (A) qRT-PCR measuring DUS4L-BCAP29 expression in cell lines. Left panel is the comparison in gastric cell lines, GES-1 and SGC-7901, HGC-27 cell lines. The expression was normalized to GAPDH, and then normalized to that in GES-1. Right panel is the comparison in prostate cell lines, RWPE-1, LNCaP, and, PC3 cell lines. The expression level was normalized to GAPDH, and then normalized to that in RWPE-1. (B) qRT-PCR measuring DUS4L-BCAP29 expression in clinical samples. Left panel is the comparison in 21 gastric cancer and matched normals. Right panel is the comparison in 18 prostate cancer and 18 non-cancer prostate tissue samples. The expression level was normalized to GAPDH.
The fact that the fusion RNA is present in non-neoplastic tissues and cells, and expressed to a similar level raised the question of whether or not it is truly oncogenic, as previously suggested [22].
Loss-of-function system
Previously, a siRNA (siDUS4L-BCAP29) targeting the fusion was used to demonstrate its role in gastric cancer [22]. However, the siRNA targeting sequence lies in the common region of the fusion and wild type DUS4L (Supplementary Figures 1 and 2), thus difficult to justify its specificity on the fusion transcript as reported [22]. Indeed, we found that the siRNA silences both the fusion (Supplementary Figure 3A) and the DUS4L wild transcripts (Supplementary Figure 3B). To silence the fusion transcript specifically, we usually design siRNAs targeting the junction sequence. In the past, we were able to use this approach to achieve fusion-specific silencing, including SLC45A3-ELK4 [9, 23], CTBS-GNG5, and CTNNBIP1-CLSTN1 [17]. Unfortunately, we failed to develop such a siRNA for DUS4L-BCAP29. To determine whether the previously reported reduced cell proliferation was truly due to the fusion RNA silencing, we designed another siRNA targeting only the wild type DUS4L (Supplementary Figures 1 and 2). Since the fusion is expressed in non-neoplastic tissues and cells, we tested the loss-of-function system in GES-1 and RWPE-1 cells. As expected, siDUS4L-BCAP29, but not siDUS4L silenced the fusion transcripts in both cell lines (Supplementary Figure 3A), whereas both siRNAs silenced the wild type transcript to a similar extent (Supplementary Figure 3A and 3B). When we measured cell proliferation, we found a more dramatic effect in siDUS4L-transfected cells in both GES-1, and RWPE-1 (Figure 3A). Both cell lines also showed significantly reduced migration ability when transfected with either siRNA, and to a similar extent (Figure 3B and 3C). We thus suspect that the effect of this loss-of-function system is mainly due to the silencing of wild type DUS4L. In conjunction with the previous report [22], we could not detect wild type BCAP29. Therefore, the effect of the siRNAs on BCAP29 was not evaluated.
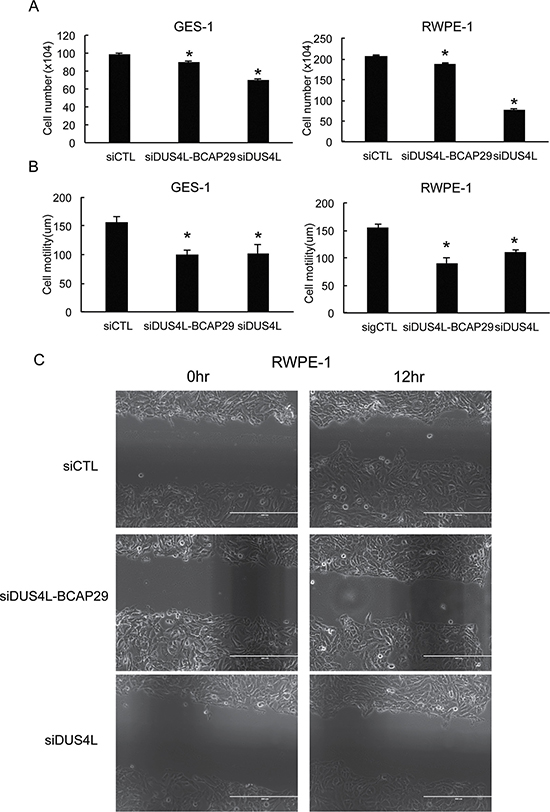
Figure 3: Loss-of-function system. (A) siRNAs silencing DUS4L-BCAP29 and wild type DUS4L decreased cell proliferation in GES-1 and RWPE-1 cell lines. The numbers of viable cells were counted three days after siRNA transfection. (B) siRNAs silencing DUS4L-BCAP29 and wild type DUS4L decreased cell motility in GES-1 and RWPE-1 cell lines. Wound healing assay was conducted after the cells achieved monolayer confluency. Distance of the wound was measured right after scratch and at a later time point. Differences of the distance were plotted. *p < 0.05 (C) Representative wound-healing assay pictures showing reduced motility in siDUS4L-BCAP29 and siDUS4L transfected RWPE-1 cells, compared with cells transfected with control siCTL. The distances of the gaps were measured and differences between two time points were compared.
We then performed microarray analyses on the GES-1 samples transfected with the control siRNA, or siDUS4L-BCAP29, or siDUS4L. Consistent with the above observation, we noticed that the most significantly enriched Gene Ontology (GO) term is negative regulators of apoptosis, and this term is shared by both siDUS4L-BCAP29, and siDUS4L (Supplementary Figure 5). Interestingly, a few GO terms unique to the siDUS4L-BCAP29 were enriched, suggesting that the fusion may have some unique functions.
Gain-of-function system
Previously, a DUS4L-BCAP29 expression vector was transfected into gastric cancer cells to demonstrate its effect on cancer cell proliferation [22]. We decided to test the effect of fusion overexpression on non-cancer cells. The full length of DUS4L-BCAP29 was cloned into a mammalian expression plasmid, pCDNA3.1, and a retroviral vector of pQXCIH-CMV. GES-1 and RWPE-1 cells were transfected, and stable cells were then selected by G418 (for pCDNA3.1), and hygromycin (for pQXCIH-CMV) respectively. In both systems, we observed enhanced proliferation rates in both cell lines (Figure 4A), and significantly increased cell motility, at least in GES-1 cells (Figure 4B). These results argue that the increased proliferation and motility effects of the fusion are not specific to cancer cells.
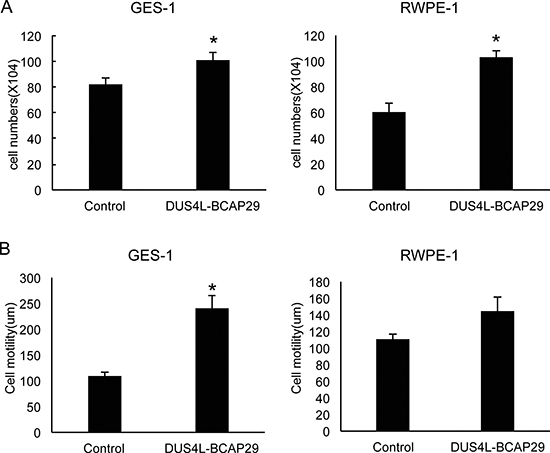
Figure 4: Gain-of-function system. Cells were transfected with either empty vector “control” or the plasmid expressing the fusion “DUS4L-BCAP29”. Stable cells expressing the fusion were selected with corresponding antibiotics. (A) Overexpression of DUS4L-BCAP29 promoted cell proliferation in GES-1 and RWPE-1 cell lines. (B) Overexpression of DUS4L-BCAP29 promoted cell motility in GES-1 and RWPE-1 cell lines. Wound healing assays were conducted on stable cells as described before. *p < 0.05
DUS4L-BCAP29 as a cis-SAGe fusion
DUS4L-BCAP29 is the type of chimeric transcript that combines the exons of adjacent genes, making them a candidate for cis-splicing between adjacent genes (cis-SAGe). This chimeric RNA also involves the second-to-last exon in DUS4L joining to the second exon in the BCAP29. As demonstrated in our previous study, this configuration is the most common type of cis-SAGe [9]. To prove its transcriptional read-through nature, we used a reverse primer annealing to the second exon of BCAP29 as the RT primer, and detected the fragment of the primary transcript between the last intron and the last exon of DUS4L (Figure 5), suggesting that the transcript runs from DUS4L to BCAP29. To avoid DNA contamination, DNaseI treatment and “no reverse transcriptase” control (Supplementary Figure 4) were used as we described before [9, 24, 25].
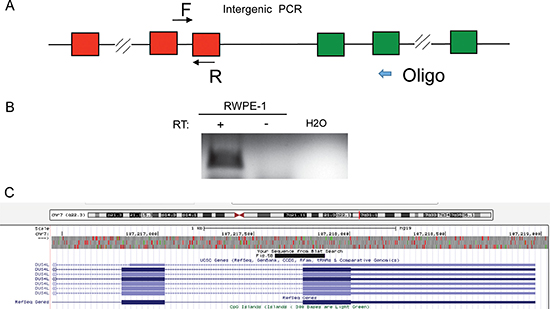
Figure 5: DUS4L-BCAP29 is a product of transcriptional read-through. (A) Scheme of the experiment. Reverse transcription was performed using an antisense primers annealing to the second exons of BCAP29 gene. A fragment between the last intron and last exon of DUS4L was then amplified using the cDNA from above. (B) In RWPE-1, the correct sized band was observed in “+RT” (with reverse transcriptase); “-RT”, no reverse transcriptase. The RNA was pretreated with DNaseI. (C) Sanger sequencing validated the PCR fragment sequence. The sequence matches the fragment as demonstrated in UCSC Genome browser.
DISCUSSION
Traditionally, gene fusions resulting from chromosomal rearrangement are considered ideal biomarkers in clinical diagnostics. For instance, PAX3-FOXO1 resulting from the t(2;13)(q35;q14) translocation [26] is detected in 55% of alveolar rhabdomyosarcoma (ARMS) [27], and it is used as a diagnostic aid in many pathology laboratories worldwide [28]. Fusion products can also be ideal therapeutic targets. Prominent examples include Gleevec targeting BCR-ABL in chronic myelogenous leukemia [5], and crizotinib targeting EML4-ALK in lung cancer [7].
However, recent research on intergenic splicing has demonstrated that trans-splicing and cis-SAGe are other mechanisms to generate chimeric RNAs [19], and that the chimeric RNAs can also exist in normal physiology [14–17]. For instance, PAX3-FOXO1 fusion RNA, was detected transiently during muscle differentiation, but no evidence of t(2;13) was detected in the normal cells [20, 29]. The assumption that all fusion (RNA)s are cancer-fusions resulted in an explosion in the deposition of fusion genes into the Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer in the Cancer Genome Anatomy Project [30]. The number of entries reached 10,256 in June 2016, compared to only a few hundred entries several years ago. This is largely due to the wide application of RNA-Sequencing of cancer samples. However, the vast majority of studies did not include a sufficient number of normal, and non-cancer control samples. Rushing to translate these fusions into cancer biomarkers will result in a disastrous amount of false positives [31]. Previously, we found 13 fusions that are listed in the Mitleman database list as of April 2015, are also present in multiple non-cancer tissues [17], and DUS4L-BCAP29 is one of them. Here, we provide further evidence to support that this fusion is not a cancer-specific event. In contrast, it is widely expressed in various tissues and cell types. Different from the traditional gene fusions produced via chromosomal rearrangement, it is a product of transcriptional read-through.
DUS4L encodes a dihydrouridine synthase 4 like protein. BCAP29 encodes the B-cell receptor associated protein 29. Based on the sequence, the fusion creates an in-frame chimeric protein, which contains the TIM_phosphate_binding superfamily domain from DUS4L and BAP31 superfamily domain from BCAP29. The exact function of the fusion in normal and cancer cells is not clear. The loss-of-function system, as reported before [2], is not sufficient to support the function of the fusion, as the siRNA effect may be purely due to the silencing of wild type DUS4L. Further support for the idea comes from microarray analyses Interestingly, where we found that the most enriched GO term shared by wild type DUS4L silencing and the fusion silencing is negative regulators of apoptosis. Interestingly, the fusion may have some novel functions different from those of the parental genes, as evidenced by some unique GO terms only seen in siDUS4L-BCAP29 group. We are actively investigating the fusion functional mechanisms and their implications in normal physiology.
MATERIALS AND METHODS
Cell culture
The cell line GES-1 was cultured in RPMI1640 media containing 10% fetal bovine serum (Gibco®, Invitrogen, Carlsbad, CA), 100 units/mL penicillin, and 100 μg/mL streptomycin, at 37°C in a 5% CO2 humidified incubator. RWPE-1 cells were grown as previously described [26].
RT-PCR and sanger sequencing
Total RNA was extracted with TRIzol reagent (Life Technologies, United States) and cDNA was generated by cDNA synthesis Kit (Bioline, United States), according to the manufacturer’s instructions. Following RT-PCR and gel electrophoresis, purified DNA bands were sent for Sanger sequencing by Eton Bioscience INC (NC, United States).
Real-time PCR
qRT-PCR was performed as described previously [9, 24, 32]. GAPDH was used as the endogenous control. The qPCR experiments were conducted with the ABI Step One Plus real time PCR system (Applied Biosystems, USA). The following primers are used for fusion and wild type RNAs. DUS4L-BCAP29-F: GCCAGTGC ACTATGATTCCA, DUS4L-BCAP29-R: GGAGGAATAA AAGGTAGGCAGAA; DUS4L-RT-F: TCCATGGAAG AACTGCTGAA, DUS4L-RT-R: AGAGTCCTCTTGCA ACCATCA. For the primer set to amplify fusion transcript, we designed the primers flanking the fusion junction. The forward primer anneals to the second-to-last exon of DUS4L, and the reverse primer anneals to the second exon of BCAP29. For primer sets specific for wild type parental genes, primer pairs annealing to only wild type but not fusion part were used.
siRNA and transfection
siRNA was purchased from Invitrogen and siRNA transfection was carried out using Lipofectamine RNAiMAX (Life Technologies, United States) following the manufacturer’s protocols. The siRNA sequence that was reported for silencing the DUS4L-BCAP29 fusion transcript is 5′-AUUAAUACAGAUAUGUUUC- 3′ [22]. The siRNA sequence used against the DUS4L wide type transcript is 5′-AGGCAAACAUUGACAAAUA - 3′.
DUS4L–BCAP29 expression constructs
Human DUS4L–BCAP29 sequence was amplified from GES-1 cell line by Reverse Transcription PCR, and cloned into mammalian expression vectors. The forward primer was 5′-CCGGAATTCCCACCATGAAGA GTGACTGCATGCAAACGACAATA- 3′ and the reverse primer was 5′-CGCTCGAGAAGCTTTCACAGTCTTT TCTTGTTGCCTCTTTCTAA- 3′. The cDNA was inserted into EcoRI/XhoI cloning sites of pCDNA3, or pQXCIH-CMV. Cells stably expressing the transgenes were selected with antibiotics, G418 (for pCDNA3.1), and hygromycin (for pQXCIH-CMV) respectively.
Cell proliferation and migration assays
Cells were plated at proper density and then transfected with siRNAs, or expression plasmids and controls. The number of total cells in each well was counted after three days of transfection. Cell migration was measured by a wound-healing assay. Briefly, a wound was created by scraping the cells using a 10 μl plastic pipette tip, and the medium was replaced with fresh medium. Images were captured immediately after making the scratch, and again 12 hours later. Cell migration was qualitatively assessed by the size of the wounds at the end of the experiment, as previously described [25].
Abbreviations
cis-SAGe: cis-Splicing of Adjacent Genes.
Authors’ contributions
Y.T., F.Q., and A.L. made unique, essential and extensive contributions to the manuscript, and are ordered by amount of time and effort contributed. H.L. conceived of the study, supervised the project and wrote the manuscript with contributions from Y.T., and F.Q.
ACKNOWLEDGMENTS
We thank Loryn Facemire for editing the manuscript. This work is supported by NIH grant CA190713.
CONFLICTS OF INTEREST
We declare that there is no conflicts of interest in relation to the work described.
REFERENCES
1. Rabbitts TH. Chromosomal translocations in human cancer. Nature. 1994; 372:143–149.
2. Rowley JD. The role of chromosome translocations in leukemogenesis. Semin Hematol. 1999; 36:59–72.
3. Heim S, Mitelman F. Molecular screening for new fusion genes in cancer. Nat Genet. 2008; 40:685–686.
4. Rowley JD. Letter: A new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature. 1973; 243:290–293.
5. Druker BJ, Talpaz M, Resta DJ, Peng B, Buchdunger E, Ford JM, Lydon NB, Kantarjian H, Capdeville R, Ohno-Jones S, Sawyers CL. Efficacy and safety of a specific inhibitor of the BCR-ABL tyrosine kinase in chronic myeloid leukemia. N Engl J Med. 2001; 344:1031–1037.
6. Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW, Varambally S, Cao X, Tchinda J, Kuefer R, Lee C, Montie JE, Shah RB, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science. 2005; 310:644–648.
7. Shaw AT, Yeap BY, Solomon BJ, Riely GJ, Gainor J, Engelman JA, Shapiro GI, Costa DB, Ou SH, Butaney M, Salgia R, Maki RG, Varella-Garcia M, et al. Effect of crizotinib on overall survival in patients with advanced non-small-cell lung cancer harbouring ALK gene rearrangement: a retrospective analysis. Lancet Oncol. 2011; 12:1004–1012.
8. Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S, Fujiwara S, Watanabe H, Kurashina K, Hatanaka H, Bando M, Ohno S, Ishikawa Y, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature. 2007; 448:561–566.
9. Qin F, Song Z, Babiceanu M, Song Y, Facemire L, Singh R, Adli M, Li H. Discovery of CTCF-sensitive Cis-spliced fusion RNAs between adjacent genes in human prostate cells. PLoS Genet. 2015; 11:e1005001.
10. Chase A, Ernst T, Fiebig A, Collins A, Grand F, Erben P, Reiter A, Schreiber S, Cross NC. TFG, a target of chromosome translocations in lymphoma and soft tissue tumors, fuses to GPR128 in healthy individuals. Haematologica. 2010; 95:20–26.
11. Ren G, Zhang Y, Mao X, Liu X, Mercer E, Marzec J, Ding D, Jiao Y, Qiu Q, Sun Y, Zhang B, Yeste-Velasco M, Chelala C, et al. Transcription-mediated chimeric RNAs in prostate cancer: time to revisit old hypothesis? OMICS. 2014; 18:615–624.
12. Yoshihara K, Wang Q, Torres-Garcia W, Zheng S, Vegesna R, Kim H, Verhaak RG. The landscape and therapeutic relevance of cancer-associated transcript fusions. Oncogene. 2015; 34:4845–4854.
13. Stransky N, Cerami E, Schalm S, Kim JL, Lengauer C. The landscape of kinase fusions in cancer. Nat Commun. 2014; 5:4846.
14. Magrangeas F, Pitiot G, Dubois S, Bragado-Nilsson E, Cherel M, Jobert S, Lebeau B, Boisteau O, Lethe B, Mallet J, Jacques Y, Minvielle S. Cotranscription and intergenic splicing of human galactose-1-phosphate uridylyltransferase and interleukin-11 receptor alpha-chain genes generate a fusion mRNA in normal cells. Implication for the production of multidomain proteins during evolution. J Biol Chem. 1998; 273:16005–16010.
15. Frenkel-Morgenstern M, Lacroix V, Ezkurdia I, Levin Y, Gabashvili A, Prilusky J, Del Pozo A, Tress M, Johnson R, Guigo R, Valencia A. Chimeras taking shape: potential functions of proteins encoded by chimeric RNA transcripts. Genome Res. 2012; 22:1231–1242.
16. Carrara M, Beccuti M, Cavallo F, Donatelli S, Lazzarato F, Cordero F, Calogero RA. State of art fusion-finder algorithms are suitable to detect transcription-induced chimeras in normal tissues? BMC Bioinformatics. 2013; 14:S2.
17. Babiceanu M, Qin F, Xie Z, Jia Y, Lopez K, Janus N, Facemire L, Kumar S, Pang Y, Qi Y, Lazar IM, Li H. Recurrent chimeric fusion RNAs in non-cancer tissues and cells. Nucleic Acids Res. 2016; 44:2859–72.
18. Finta C, Zaphiropoulos PG. Intergenic mRNA molecules resulting from trans-splicing. J Biol Chem. 2002; 277:5882–5890.
19. Jividen K, Li H. Chimeric RNAs generated by intergenic splicing in normal and cancer cells. Genes Chromosomes Cancer. 2014; 53:963–971.
20. Yuan H, Qin F, Movassagh M, Park H, Golden W, Xie Z, Zhang P, Sklar J, Li H. A chimeric RNA characteristic of rhabdomyosarcoma in normal myogenesis process. Cancer Discov. 2013; 3:1394–1403.
21. Nacu S, Yuan W, Kan Z, Bhatt D, Rivers CS, Stinson J, Peters BA, Modrusan Z, Jung K, Seshagiri S, Wu TD. Deep RNA sequencing analysis of readthrough gene fusions in human prostate adenocarcinoma and reference samples. BMC Med Genomics. 2011; 4:11.
22. Kim HP, Cho GA, Han SW, Shin JY, Jeong EG, Song SH, Lee WC, Lee KH, Bang D, Seo JS, Kim JI, Kim TY. Novel fusion transcripts in human gastric cancer revealed by transcriptome analysis. Oncogene. 2014; 33:5434–5441.
23. Zhang Y, Gong M, Yuan H, Park HG, Frierson HF, Li H. Chimeric transcript generated by cis-splicing of adjacent genes regulates prostate cancer cell proliferation. Cancer Discov. 2012; 2:598–607.
24. Qin F, Song Y, Zhang Y, Facemire L, Frierson H, Li H. Role of CTCF in Regulating SLC45A3-ELK4 Chimeric RNA. PLoS One. 2016; 11:e0150382.
25. Qin F, Song Z, Chang M, Song Y, Frierson H, Li H. Recurrent cis-SAGe chimeric RNA, D2HGDH-GAL3ST2, in prostate cancer. Cancer Lett. 2016; 380:39–46.
26. Downing JR, Khandekar A, Shurtleff SA, Head DR, Parham DM, Webber BL, Pappo AS, Hulshof MG, Conn WP, Shapiro DN. Multiplex RT-PCR assay for the differential diagnosis of alveolar rhabdomyosarcoma and Ewing’s sarcoma. Am J Pathol. 1995; 146:626–634.
27. Sorensen PH, Lynch JC, Qualman SJ, Tirabosco R, Lim JF, Maurer HM, Bridge JA, Crist WM, Triche TJ, Barr FG. PAX3-FKHR and PAX7-FKHR gene fusions are prognostic indicators in alveolar rhabdomyosarcoma: a report from the children’s oncology group. J Clin Oncol. 2002; 20:2672–2679.
28. Missiaglia E, Williamson D, Chisholm J, Wirapati P, Pierron G, Petel F, Concordet JP, Thway K, Oberlin O, Pritchard-Jones K, Delattre O, Delorenzi M, Shipley J. PAX3/FOXO1 fusion gene status is the key prognostic molecular marker in rhabdomyosarcoma and significantly improves current risk stratification. J Clin Oncol. 2012; 30:1670–1677.
29. Xie Z, Babiceanu M, Kumar S, Jia Y, Qin F, Barr FG, Li H. Fusion transcriptome profiling provides insights into alveolar rhabdomyosarcoma. Proc Natl Acad Sci USA. 2016; 113: 13126–13131.
30. Mitelman F JBaMFE. Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer 2015.
31. Jia Y, Xie Z, Li H. Intergenically Spliced Chimeric RNAs in Cancer. Trends Cancer. 2016; 2:475–484.
32. Qin FJ, Sun QW, Huang LM, Chen XS, Zhou DX. Rice SUVH histone methyltransferase genes display specific functions in chromatin modification and retrotransposon repression. Mol Plant. 2010; 3:773–782.

