INTRODUCTION
Oral squamous cell carcinoma (OSCC) is the sixth most common malignancy worldwide, with 599,000 new cases and 325,000 cancer deaths in 2012[1, 2]. OSCC, which accounts for more than 90% of all mouth malignancies and approximately 40% of head and neck tumours has, a high potential for local invasion and lymph node metastasis. Indeed, the estimated 5-year overall survival rate of less than 50% has not improved significantly over the last three decades [3]. OSCC often causes postsurgical dysfunctions in chewing, swallowing, and speech, as well as aesthetics loss [4]. Moreover, although early-stage OSCCs has a high treatment success rate approximately 70% of patients with progressive disease cannot be successfully treated due to relatively high local and regional recurrence rates [5]. Therefore, early detection and clarification of the detailed molecular mechanism of OSCC are urgently needed [6, 7].
The HMGB1 gene belongs to the high-mobility group protein family, and the protein product contains two 80-amino acid DNA-binding domains (A-box and B-box) and an negatively charged C-terminus [8]. The gene sequence is located on the long arm of chromosome 13; the transcriptional region spans approximately 6,000-bp, with a promoter region of at least 1,700-bp, and 5 exons of approximately 2,600-bp [8]. HMGB1 functions in the nucleus as a chromatin structural protein and extracellularly as a pro-inflammatory cytokine [9, 10]. Nuclear HMGB1 is a non-histone DNA-binding protein, which suggests that it facilitates the assembly of site-specific DNA targets by acting as a DNA chaperone [11]. In contrast, extracellular HMGB1 functions as a damage-associated molecular pattern that propagates infection- or injury-elicited inflammatory responses [12]. Moreover, evidences confirm that HMGB1 over-expression is closely related to tumour development through functions in proliferation, invasion and migration of cancer cells [13-16]. Therefore, HMGB1 may serve a biomarker of inflammation and/or a prognostic marker for OSCC progression [9, 17, 18].
Although the development of OSCC may take several decades years, early detection of this cancer is seldom achieved due to the lack of reliable markers [19]. Therefore, the disease runs a largely asymptomatic course until it becomes too advanced for successful treatment [20, 21]. Aberrations in some genes may be responsible for certain clinical features of OSCC [22]. For instance, differences in the level of HMGB1 expression have been demonstrated between precancerous and malignant lesions [23-25] yet little is known regarding the joint effects of HMGB1 gene variants and behavioural exposure to cancer-causing substances on predisposition to OSCC. Moreover, previous studies have reported the effect of HMGB1 gene polymorphisms on human cancer susceptibility, and the polymorphisms may efficiently predict the risk of cancers [26-30]. Thus, we hypothesized that four polymorphisms (rs1412125, rs2249825, rs1045411, and rs1360485; Table 1) in HMGB1 are associated with susceptibility to OSCC. The aim of this study was to examine the potential clinical relevance of four SNPs in the HMGB1 gene in patients with OSCC, as well as associations between HMGB1 polymorphisms, haplotypes and environmental risk factors.
Table 1: Variants, positions, function and of high mobility group box 1 (HMGB1) sequence variations
Chromosome position (Chr.13) |
||||
|---|---|---|---|---|
SNP 1 |
SNP 2 |
SNP 3 |
SNP 4 |
|
Genome position † |
31,041,595 |
31,037,903 |
31,033,232 |
31,031,884 |
Nucleotide change |
T>C |
C>G |
C>T |
T>C |
dbSNP (rs number) |
rs1412125 |
rs2249825 |
rs1045411 |
rs1360485 |
Molecular consequences ‡ |
2KB upstream |
intron 1 |
3’-UTR |
500B downstream |
Function prediction |
enhancer |
Transcriptional factor |
miRNA binding site |
- |
Heterozygous (%) ‖ |
44.4 |
28.0 |
33.5 |
36.3 |
† GRCh37.p13
‡ The Molecular consequences annotated by human HMGB1 mRNA (NM_002128.5).
‖ Specific heterozygosity frequencies using the East Asian population (1000 Genomes Project phase 3).
RESULTS
Study population
A total of 1,972 participants, including 772 OSCC cases and 1,200 controls were recruited to explore the effects of the HMGB1 gene on OSCC risk. Demographic characteristics including mean age, betel quid chewing, cigarette smoking and alcohol drinking are shown in Table 2. We discovered significant differences when grouping by betel quid chewing (P<0.001), cigarette smoking (P<0.001) and alcohol drinking (P<0.001) in the healthy controls and OSCC patients (Table 2).
Table 2: The distributions of demographical characteristics in 1,200 controls and 772 patients with oral squamous cell carcinoma (OSCC)
Variables |
Controls (N=1,200) |
Patients (N=772) |
P value † |
|---|---|---|---|
Age (yrs) |
Mean ± S.D. |
Mean ± S.D. |
|
53.91 ± 10.02 |
54.80 ± 11.04 |
P=0.063 |
|
Betel quid chewing |
n (%) |
n (%) |
|
No |
1,001 (83.4%) |
154 (19.9%) |
P<0.001 |
Yes |
199 (16.6%) |
618 (80.1%) |
|
Cigarette smoking |
|||
No |
564 (47.0%) |
86 (63.2%) |
P<0.001 |
Yes |
636 (53.0%) |
686 (88.9%) |
|
Alcohol drinking |
|||
No |
963 (80.3%) |
340 (44.0%) |
P<0.001 |
Yes |
237 (19.8%) |
432 (56.0%) |
|
† Mann–Whitney U test or Fisher’s exact test was used between controls and patients with OSCC.
HMGB1 associated polymorphisms and environmental risk factors
Table 3 summarizes the basic characteristics of the HMGB1 SNPs in the study population. In both the OSCC patients and healthy control subjects, genotypes T/T, C/C, C/C, and T/T exhibited the highest frequencies for rs1412125, rs2249825, rs1045411 and rs1360485, respectively. In these controls, the genotypic frequency of HMGB1 SNP rs1412125 met the Hardy-Weinberg equilibrium (P=0.282, χ2 value: 1.159). The frequencies of HMGB1 SNPs rs2249825, rs1045411 and rs1360485 were also in the Hardy-Weinberg equilibrium (P=0.678, χ2 value: 0.172; P=0.451, χ2 value: 0.569; and P=0.537, χ2 value: 0.382, respectively). According to the adjusted odds ratios (AORs) and 95% confidence intervals (CIs) in a multiple logistic regression model for HMGB1 polymorphism and OSCC, compared with their corresponding wild-type homozygotes (C/C), only rs1045411 C/T or C/T+T/T presented a significant (P<0.05) protective role after adjusting confounding factors: 0.754-fold (95% CI, 0.582-0.978) and 0.778-fold (95% CI, 0.609-0.995), respectively (Table 3).
Table 3: Adjusted odds ratio (AOR) and 95% confidence interval (CI) of oral squamous cell carcinoma (OSCC) associated with HMGB1 genotypic frequencies
Variables |
Controls (n=1,200) |
Patients (n=772) |
OR (95% CI) † |
AOR (95% CI) ‡ |
|---|---|---|---|---|
rs1412125 |
||||
TT |
649 (54.1%) |
438 (56.7%) |
1.00 |
1.00 |
TC |
457 (38.1%) |
274 (35.5%) |
0.888 (0.733-1.077) |
0.993 (0.772-1.277) |
CC |
94 (7.8%) |
60 (7.8%) |
0.946 (0.669-1.337) |
0.975 (0.622-1.527) |
TC+CC |
551 (45.9%) |
334 (43.3%) |
0.898 (0.749-1.078) |
0.990 (0.780-1.255) |
rs2249825 |
||||
CC |
852 (71.0%) |
573 (74.2%) |
1.00 |
1.00 |
CG |
316 (26.3%) |
183 (23.7%) |
0.861 (0.698-1.063) |
0.848 (0.644-1.116) |
GG |
32 (2.7%) |
16 (2.1%) |
0.743 (0.404-1.367) |
0.512 (0.243-1.077) |
CG+GG |
348 (29.0%) |
199 (25.8%) |
0.850 (0.693-1.043) |
0.807 (0.619-1.053) |
rs1045411 |
||||
CC |
723 (60.3%) |
507 (65.7%) |
1.00 |
1.00 |
CT |
411 (34.3%) |
226 (29.3%) |
0.784 (0.643-0.956) |
0.754 (0.582-0.978) |
TT |
66 (5.5%) |
39 (5.1%) |
0.843 (0.558-1.272) |
0.962 (0.565-1.636) |
CT+TT |
477 (39.8%) |
265 (34.3%) |
0.792 (0.656-0.956) |
0.778 (0.609-0.995) |
rs1360485 |
||||
TT |
682 (56.8%) |
452 (58.5%) |
1.00 |
1.00 |
TC |
440 (36.7%) |
273 (35.4%) |
0.936 (0.772-1.135) |
0.926 (0.719-1.919) |
CC |
78 (6.5%) |
47 (6.1%) |
0.909 (0.621-1.331) |
0.998 (0.609-1.636) |
TC+CC |
518 (43.2%) |
320 (41.5%) |
0.932 (0.776-1.119) |
0.936 (0.737-1.190) |
† The odds ratio (OR) with their 95% confidence intervals were estimated bylogistic regression models.
‡ The adjusted odds ratio (AOR) with their 95% confidence intervals were estimated by multiple logistic regression models after controlling for betel quid chewing, alcohol and tobacco consumption.
Functional effect of rs1045411 on HMGB1 transcript stability
All of the subjects were genotyped for the four SNPs, which were selected to cover (r2 ≥ 0.70) most of the SNPs located in a 12 kb region that includes the HMGB1 gene (8 kb), its promoter (2 kb) and the 3’-UTR (2 kb) (Figure 1A). The 3’-untranslated region (3’-UTR) of the HMGB1 gene covers 2.0 kb and might be among the region most sensitive to microRNA (miRNA) epigenetic regulation [31]. An miRNA, hsa-miR-505-5p [32, 33] (miRBase [34] ID: MI0003190, Figure 1B), shares binding site complementarily with rs1045411 in the 3’-UTR (Figures 1C and 1D). In addition, the OSCC-associated risk [C]-allele creates a slight kink in the HMGB1 mRNA structure compared to the [T]-allele, which results in a less negative free-energy state, and less stable hybridization [MFE (minimum free energy) change: 6.04%, from -19.3 kcal/mol to -18.2 kcal/mol] (Figure 1E). Furthermore, starBase [35] analysis revealed an 1.86-fold (P=2.39x10-10) increase in hsa-miRNA-505-5p expression in 420 OSCC patients and 43 subjects from the Pan-Cancer dataset [36] (Figure 1F). Accordingly, the location of the 3’-UTR SNP rs1045411 [C-allele] alter HMGB1 mRNA stability and increase susceptibility to OSCC (Figure 1E).
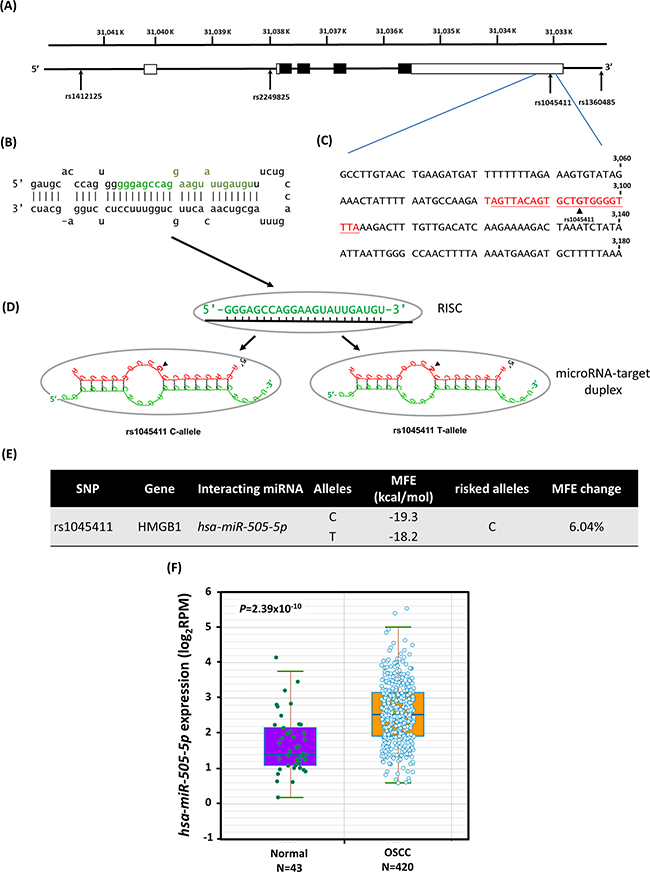
Figure 1: Binding site polymorphism for SNP rs1045411 [G/A] in the human HMGB1 3’-UTR mRNA with a microRNA hsa-miR-505-5p binding site, decreases OSCC susceptibility among the Taiwan OSCC population. (A) Exons of HMBG1 are shown as filled boxes along the chromosome (chr.13, reference genome GRCh37.p13). (B) The stem-loop portion of the miRNA-miRNA duplex structure of pre-miRNAs (hsa-miR-505; miRBase ID: MI0003190) was identified by miRNA target prediction using the MicroRNA.org resource. The hsa-miR-505-5p sequence is marked in green. (C) Sequence of the human HMBG1 3’-UTR region; the number shows the positions of the mRNA (NM_002128.5). The predicted hsa-miR-505-5p binding site of SNP rs1045411 is highlighted in color red. (D) Models of the miRNA-target duplex were determined using the RNAhybrid web tool on Bielefeld Bioinformatics Server. RISC, RNA-induced silencing complex; arrows indicate the location of rs1045411. (E) The SNP rs1045411 C-allele reduces the free binding energy (MFE, minimum free energy; change: 6.04%). (F) Boxplot chart showing the differential expressions of miRNA hsa-miR-505-5p in 420 OSCC patients and 43 normal controls, as taken from the Pan-Cancer dataset.
Synergistic effects of genetic variants and betel quid chewing behavior with smoking
The habit of betel quid chewing widespread in Taiwan, and chewers are often smokers. Thus, we analyzed the synergistic effects of cigarette smoking and betel chewing combined with the four SNPs in the OSCC patients. Among the 1,322 smokers in our study, the estimated AORs of the four selected HMGB1 polymorphisms were elevated in individuals who were both betel quid chewers and smokers (10.071-10.659-fold) compared to abstainers (Table 4). However, the risk of OSCC tended to decline among those who had quit betel quid chewing, suggesting that the four selected HMGB1 variants are associated with betel quid chewing and OSCC susceptibility in cigarette smokers (Table 4).
Table 4: Associations of the combined effect of HMGB1 gene polymorphisms and betel quid chewing with the susceptibility to oral squamous cell carcinoma (OSCC) among 1,322 smokers
Variable |
Controls |
Patients |
OR (95% CI) † |
AOR (95% CI) ‡ |
|---|---|---|---|---|
rs1412125 |
||||
TT genotype & non-betel quid chewing a |
231 (36.3%) |
59 (8.6%) |
1.00 |
1.00 |
TC or CC genotype or betel quid chewing b |
331 (52.0%) |
375 (54.7%) |
4.436 (3.215-6.120) |
3.604 (2.534-5.126) |
TC or CC genotype with betel quid chewing c |
74 (11.6%) |
252 (36.7%) |
13.333 (9.065-19.611) |
10.164 (6.688-15.446) |
rs2249825 |
||||
CC genotype & non-betel quid chewing a |
314 (49.4%) |
82 (12.0%) |
1.00 |
1.00 |
CG or GG genotype or betel quid chewing b |
269 (42.3%) |
446 (65.0%) |
6.349 (4.768-8.454) |
5.465 (3.982-7.501) |
CG or GG genotype with betel quid chewing c |
53 (8.3%) |
158 (23.0%) |
11.416 (7.692-16.942) |
10.026 (6.505-15.453) |
rs1045411 |
||||
CC genotype & non-betel quid chewing a |
280 (44.0%) |
71 (10.3%) |
1.00 |
1.00 |
CT or TT genotype or betel quid chewing b |
288 (45.3%) |
403 (58.7%) |
5.518 (4.083-7.458) |
4.877 (3.494-6.809) |
CT or TT genotype with betel quid chewing c |
68 (10.7%) |
212 (30.9%) |
12.295 (8.430-17.932) |
10.071 (6.670-15.208) |
rs1360485 |
||||
TT genotype & non-betel quid chewing a |
262 (41.2%) |
66 (9.6%) |
1.00 |
1.00 |
TC or CC genotype or betel quid chewing b |
301 (47.3%) |
371 (54.1%) |
4.893 (3.589-6.670) |
4.383 (3.111-6.176) |
TC or CC genotype with betel quid chewing c |
73 (11.5%) |
249 (36.3%) |
13.540 (9.303-19.709) |
10.659 (7.073-16.065) |
† The odds ratio (OR) with their 95% confidence intervals were estimated bylogistic regression models.
‡ The adjusted odds ratio (AOR) with their 95% confidence intervals were estimated by multiple logistic regression models after controlling for betel quid chewing, alcohol and tobacco consumption.
a. Individuals are homozygous for ancestral allele but without betel quid chewing.
b. Individuals with either at least one polymorphic allele or betel quid chewing.
c. Individuals with both at least one polymorphic allele and betel quid chewing.
Haplotypes analysis for the HMGB1 gene
HMGB1 polymorphisms were further characterized using linkage disequilibrium (LD) and haplotype analyses. LD was determined pairwise among all 4 SNPs and the haplotype structure of the HMGB1 gene was analyzed (D’ and r2) according to 1000 Genomes Project data [37] for the East Asian population (CHB+JPT, Figure 2). Haplotype blocks divided by the D’ confidence interval method with a D’ value of 95% CI 0.70-0.98 in adjacent SNPs was classified as the same haplotype block. One LD block was detected by Solid Spine [38], a haplotype phasing technique. Block1 (3 kb) consisted of two closely selective SNPs showing strong linkage, rs1045411 and rs1360485, in the 3’-UTR of HMGB1 (Figure 2).
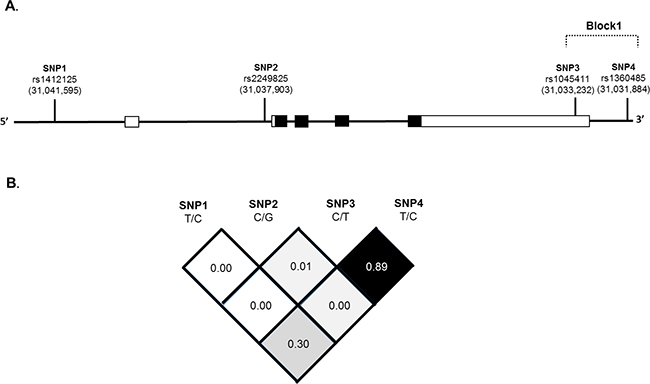
Figure 2: The HMGB1 gene, locations of the genotyped variants and their pairwise linkage disequilibrium patterns. Schematic presentation of the HMGB1 (gene ID: 3146), (A) indicating the locations of the analyzed variants (rs1412125, rs2249825, rs1045411 and rs1360485). SNPs within and around the HMGB1 gene were plotted against the chromosomal positions chr.13: 29,928,000 to 29,940,000 (HapMap version 3 release 27). Exons, introns and untranslated regions are shown by filled boxes, unfilled and thin lines from 5’- to 3’- end of this gene, respectively. (B) In the LD map SNPs-pairwise correlation coefficients, r2, in East Asian population (HCB+JPT) are shown in the square black when D’=1.0, while the square is white when D’=0. A “Confidence intervals” conventional gray scale is used to display LD were shown in black through gray (color intensity decreasing with decrease D’ value) generated using Haploview version 4.2.
Furthermore, a haplotype-based association study was performed to show association between HMGB1 haplotypes and OSCC risk (Table 5). Block1 of the 3’-UTR SNPs constituted virtually four haplotypes of approximately equal frequencies in the control subjects (75.0%, 22.5%, 2.3% and 0.1%). In contrast, one haplotype in the OSCC cases, “C-C”, was associated with increased susceptibility to OSCC (AOR=1.808; 95% CI, 1.257-2.601, P=0.001) compared with the most common “C-T” haplotype (Table 5).
Table 5: The estimated haplotype frequencies of four examined polymorphisms in HMGB1 gene and the corresponding risk for oral squamous cell carcinoma (OSCC)
Haplotype block |
Controls (n=2,400) |
Patients (n=1,544) |
OR (95% CI) |
P value |
|
|---|---|---|---|---|---|
rs1045411 C/T |
rs1360485 T/C |
||||
C |
T |
1,801 (75.0%) |
1,174 (76.0%) |
1.00 |
|
T |
C |
540 (22.5 %) |
301 (19.5%) |
0.855 (0.729-1.002) |
0.054 |
C |
C |
56 (2.3%) |
66 (4.3%) |
1.808 (1.257-2.601) |
0.001 |
T |
T |
3 (0.1%) |
3 (0.2%) |
1.534 (0.309-7.613) |
0.598 |
The effects of rs1412125 and rs2249825 on HMGB1 transcriptional regulation
As a preliminary assessment of the putative functional role of these SNPs, we investigated whether rs1412125 and rs2249825 are associated with differential expression of HMGB1, According to the Genome-Tissue Expression (GTEx) database [39], statistically significant down-regulation of HMGB1 mRNA expression with rs2249825 variant genotype (C/G+G/G) compared with the wild-type homozygous genotype (C/C, P=0.026, Figure 3A) is observed in whole blood. Moreover, OSCC-risk-associated environmental factors were further explored by examining functional annotation in the Encyclopedia of DNA elements (ENCODE) data [40] for sites in the 3,500-bp promoter and intron 1 sequences (Figure 3B and 3C). We determined that rs1412125 and rs2249825 are situated at a site for transcription factor binding, with histone modification patterns and DNA hyposensitivity characterized in the promoter or enhancers in several cell type (Figure 3C). The risk allele [T] of rs1412125 in located in the core of a CCAAT/Enhancer box [41] (Figure 3D), which is one of the most common elements in eukaryotic promoter. The effect of rs2249825 may be attributed to the suboptimal avian myeloblastosis viral oncogene homolog-like 2 (v-Myb) binding site [42] (Figure 3E), with the consensus motif of (C/T)AA(G/T)G surrounding intron 1 of the predicted transcriptional start site of human HMGB1 gene (Figure 3B), which enables modulating of initiation rates in response to transcriptional status. Accordingly, these promoter polymorphisms in the CCAAT/Enhancer box and v-Myb binding site might collaborate in the regulation of HMGB1 expression.
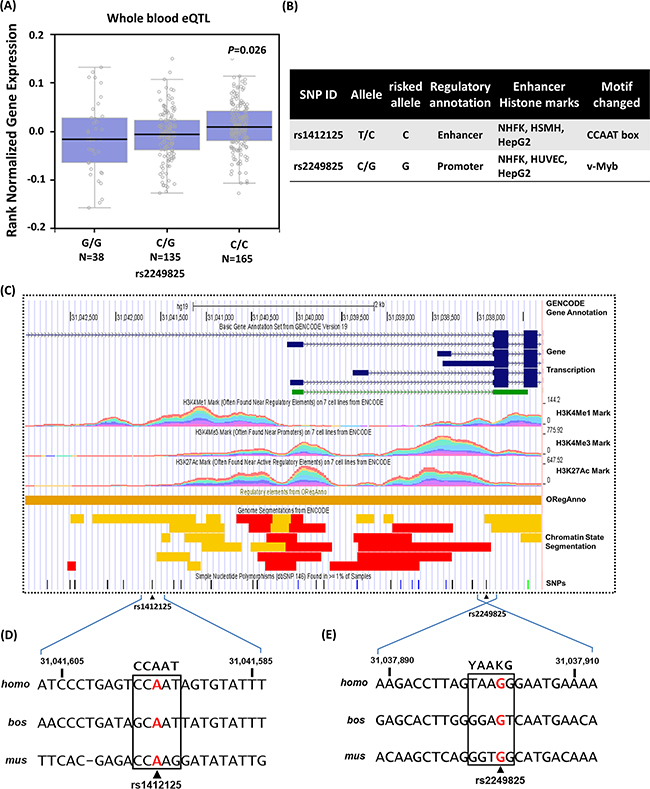
Figure 3: Binding sites polymorphism for SNPs rs1412125 [T/C] and rs2249825 [C/G] provide transcriptional control to regulate expression of the human HMGB1 gene (NM_002128.5). (A) Expression quantitative trait locus (eQTL) association between rs2249825 and HMGB1 expression in whole blood (GTEx data set). The numbers in parentheses indicate the number of cases. (B) Regulatory annotation of variants from ENCODE data showing evidence of enhancer elements coinciding with rs1412125 and rs2249825 in many different cell types. In addition, the CCAAT/Enhancer box and v-Myb motif are predicted to be affected. (C) Expanded view of ENCODE data for the HMGB1 promoter and intron 1, containing rs1412125 and rs2249825, in the GRCh37/hg19 assembly using the UCSC genome browser. In Gene Transcription track, shown color blue and green indicated coding and non-coding HMGB1 proteins, respectively. H3K4Me1, H3K4Me3, and H3K27Ac tracks show the genome-wide levels of enrichment of the mono-methylation of lysine 4, tri-methylation of lysine 4, and acetylation of lysine 27 of the H3 histone protein, as determined by the ChIP-seq assays. These levels are thought to be associated with promoter and enhancer regions. Chromatin State Segmentation track displays chromatin state segmentations by integrating ChIP-seq data using a Hidden Markov Model for H1-hESC (embryonic stem cells; color yellow), HepG2 (hepatocellular carcinoma cells; color orange), HUVEC (umbilical vein endothelial cells; color light blue), HMEC mammary epithelial cells, HSMM (skeletal muscle myoblasts; color green), NHEK (epidermal keratinocytes; color purple), and NHLF (lung fibroblasts; color red). The chromatin state regions predicted for promoters and enhancer are highlighted by red and yellow, respectively. ORegAnno track shows transcription factor binding sites. Enhancers are typically common near transcription start sites and may be associated with promoter regions. (D) Sequence of the human HMGB1 promoter region and rs1412125. Consensus residues of the CCAAT/Enhancer box core consensus sequence are indicated in bold fonts. (E) Reversed sequence of the human HMGB1 intron region and rs2249825. Consensus residues of the core v-Myb consensus sequence are indicated in bold font, where Y denotes C or T and K denotes G or T.
DISCUSSION
Cancer is the result of a series of genetic or epigenetic regulations, and environmental risk factors may contribute to the development and progression of tumors [21, 43]. Thus, in-depth understanding of the mechanisms underlying carcinogenesis is needed to characterize genetic alterations linked to OSCC development [44]. Such information may provide relevant strategies for prevention, for the surveillance of patients at risk, and for early detection to reduce morbidity and mortality from OSCC [45]. Several studies have suggested that genetic alternations harbored on chromosome 13q12 are involved in multiple tumor types [46-50] and this locus is highly susceptibility to genomic instability in OSCC [51]. Recent studies have reported that HMGB1 polymorphism is associated with susceptibility to several carcinomas including breast cancer [52], melanoma [53], gastric cancer [54] and colorectal cancer [55]. Loss of HMGB1 increase DNA damage due to decreases in DNA chaperone efficiency in response chemotherapy, cell stress, and environmental risk factors [56]. Our data revealed an increased risk of OSCC among patients with the HMGB1 polymorphism rs1045411 C/C compared with those with the C/T or C/T+T/T genotype (Table 3). We present additional evidence for a role for HMGB1 in OSCC by showing a significant interaction between betel quid chewing and at least one polymorphic allele of four HMGB1 SNPs in individuals, higher incidence of OSCC with smoking behavior (Table 4).
The present study confirms the association of the clinically examined rs1045411 and rs1360485 haplotype in the HMGB1 3’-UTR region with OSCC risk most likely because a putative miRNA-505 binding site (Figure 1). We confirmed that these two HMGB1 3’-UTR variants are synergistically involved in the etiopathogenesis of OSCC in a haplotype-specific fashion rather than as single genetic variants (Table 5). Therefore, an SNP haplotype corresponding to the RNA bulge region may effects the binding strength of specific miRNA/target duplexes, resulting in low minimum free energy and modulating mRNA stability (Figure 1). Although directly testing this hypothesis is beyond the scope of the current study, evidence suggests that mRNA stability is associated with susceptibility to OSCC, with HMGB1 downregulation being associated with an increased susceptibility to OSCC. Previous studies demonstrate that miRNA-505 acts as tumor suppressor in endometrial carcinoma [57]. Matamala et al. also reported that circulating miRNA-505 was significantly overexpressed in breast cancer patients [58]. Moreover, miRNA-505 suppresses proliferation and invasion in hepatoma cells by directly targeting HMGB1 has been previously reported [59]. Hence, the present analyses increased our understanding of naturally occurring HMGB1 variants, a lesion category that, while not infrequent, has been relatively neglected in terms of elucidation of the underlying pathogenic mechanisms. HMGB1 expression is considered as a novel and independent predictor of decreased disease-free survival of patients with prostate cancer [25, 60], and it may also be used as a novel and independent predictor of prognosis for OSCC patients.
Converging lines of evidence support that OSCC development is a complicated process regulated by both environmental and genetic factors. In the present study, we estimated the magnitude of the statistical interaction between two measurable exposures with respect to a given outcome with HMGB1 genetic variants and found that the use of betel quid with smoking conferred a higher susceptibility to OSCC (Table 4). Strikingly, polymorphisms altering DNA chaperone activity, which maintains nuclear homeostasis, may lead to synergistic effects in areca nut carcinogen-induced OSCC risk [61]. However, the effects of environmental risks on predisposition to OSCC may be underestimated because of the inability to quantitatively access the degree of betel quid chewing and DNA chaperone gene variations with smoking. Nevertheless, this study provides comprehensive evidence for the association of several genetic variation in the DNA-chaperon HMGB1 gene and multiple environmental exposures in OSCC development [62]. However, there are several limitations in our investigation. One of the limitations is that information on exposures of carcinogens is dichotomized into ‘‘ever-user’’ versus ‘‘never-user.’’ More detailed analysis based on amount on exposures of carcinogens should be performed. Furthermore, the molecular functional role in the HMGB1 3’UTR mRNA with a miRNA-505-binding site of OSCC requires further investigation.
To our knowledge, there are no previous studies of the effects of gene-environment interactions on DNA chaperon gene polymorphisms and betel-chewing habits in oral tumorigenesis. Nonetheless, the co-effects of genetic and environmental factors interact to facilitate OSCC development. Overall, we present a possible role for HMGB1 variants and provide deeper insights into naturally occurring haplotype-based variants. Furthermore, characterizing the molecular basis of polymorphisms in cancer provides insight into tumorigenesis. Accurate biomarkers for such types of variants are required for developing optimal therapeutic approaches that will eventually ameliorate the clinical phenotype in patients harboring the corresponding lesions.
MATERIALS AND METHODS
Description of the participants
This study recruited 772 male OSCC patients between 2008 and 2015 at Chung Shan Medical University Hospital, Taiwan. Clinically information including TNM staging, primary tumor size, lymph node involvement, and histologic grade was examined according to American Joint Committee on Cancer (AJCC) classification [63, 64]. Subjects with neither self-reported history of cancer of any site nor oral precancerous disease such as oral submucous fibrosis, leukoplakia, erythroplakia, or verrucous hyperplasia were selected to the control group from Taiwan Biobank. Before the study began, approval was obtained from the Institutional Review Board of Chung Shan Medical University Hospital, and informed written consent was obtained from each individual.
SNPs selection and genotyping
Genomic DNA was isolated from peripheral blood using the QIAamp DNA blood mini kit as described in detail previously [65, 66]. The final preparation was stored at -20°C, quantified by optical density measurement at 260 nm and used as the template for polymerase chain reaction (PCR). Genotyping of four HMGB1 SNPs (rs1412125, rs2249825, rs1045411, and rs1360485; Table 1) with minor allele frequencies >5% in the HapMap Chinese Han Beijing (CHB) population was performed using the TaqMan SNP genotyping assay (Applied Biosystems, Foster City, CA, USA) [67]. Allele frequencies were determined by the ABI SDS software.
Bioinformatics analysis
Several semi-automated bioinformatics tools were applied to assess whether SNPs or their linked genetic variants are associated with a putative function that might affect patient outcomes. HaploReg [68] v4 and the GTEx database [39] from the ENCODE project [69] were used to identify the regulatory potential of candidate functional variants. Particular sites of interest were examined including transcription factor (TF)-ChIP signals, DNase peaks, DNase footprints and predicted DNA sequence motifs for TFs. The GTEx data were used to identify correlations between SNPs and whole blood-specific gene expression levels. The publically available cBioPortal for Cancer Genomics [70] and UCSC Cancer Genomics Browser [71] for OSCC were utilized to analyse HMGB1 gene expression, enhancer modulation, molecular features, and clinical outcomes.
Characteristics of miRNA candidates
We predicted targets using the web-based tools RNAhybrid on BiBiServ2 (http://bibiserv.techfak.uni-bielefeld.de/rnahybrid). RNAhybrid [72] determines the most energetically favorable hybridization patterns using the MFE of two RNA fragments with different lengths, i.e. long (3’-UTR of HMGB1) and short (mature miRNA sequences). The parameters used in the analysis were as follows: number of hits per target, 3 nucleotides, maximum mismatch size, 1 nucleotide, overhangs 2 nucleotides. The MFE considered for each miRNA/target duplex was higher than -15 kcal/mole as assessed for a perfect match between the mature miRNA and its target.
Statistical analysis
The Mann–Whitney U test and Fisher’s exact test were used to compare differences in the distribution of age and demographic characteristics between the controls and OSCC patients. ORs with 95% CIs were estimated using logistic regression models. AORs with 95% CIs were used to assess association between genotype frequencies with OSCC risk and clinical factors. P values less than 0.05 were considered significant. The data were analysed with SPSS 12.0 statistical software (SPSS Inc., Chicago, IL, USA).
Author contributions
YFL and CWL conceived and designed the experiments; SFY, CMY and YEC performed the experiments; YFL and SFY analyzed the data; SFY and CYC contributed samples; YFL and CWL wrote the paper.
ACKNOWLEDGMENTS
The authors would like to thank the Ministry of Science and Technology, Taiwan, for a research grant under Contract No. NSC-102-2320-B-040-012-MY2, Taichung Taiwan.
CONFLICTS OF INTEREST
The authors declared no conflict of interest.
NOVELTY AND IMPACT STATEMENTS
The presence of the HMGB1 SNPs, combined with betel quid chewing showed profoundly synergistic effects on the risk of OSCC. The finding may help in development of optimal therapeutic approaches for OSCC patients.
REFERENCES
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65: 87-108.
2. Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010; 127: 2893-2917.
3. Kademani D, Bell RB, Bagheri S, Holmgren E, Dierks E, Potter B, Homer L. Prognostic factors in intraoral squamous cell carcinoma: the influence of histologic grade. J Oral Maxillofac Surg. 2005; 63: 1599-1605.
4. Tei K, Maekawa K, Kitada H, Ohiro Y, Yamazaki Y, Totsuka Y. Recovery from postsurgical swallowing dysfunction in patients with oral cancer. J Oral Maxillofac Surg. 2007; 65: 1077-1083.
5. Mydlarz WK, Hennessey PT, Califano JA. Advances and Perspectives in the Molecular Diagnosis of Head and Neck Cancer. Expert Opin Med Diagn. 2010; 4: 53-65.
6. Petti S. Oral cancer screening usefulness: between true and perceived effectiveness. Oral Dis. 2016; 22: 104-108.
7. Chien MH, Lin CW, Cheng CW, Wen YC, Yang SF. Matrix metalloproteinase-2 as a target for head and neck cancer therapy. Expert Opin Ther Targets. 2013; 17: 203-216.
8. Ferrari S, Finelli P, Rocchi M, Bianchi ME. The active gene that encodes human high mobility group 1 protein (HMG1) contains introns and maps to chromosome 13. Genomics. 1996; 35: 367-371.
9. Hsieh MJ, Hsieh YH, Lin CW, Chen MK, Yang SF, Chiou HL. Transcriptional regulation of Mcl-1 plays an important role of cellular protective effector of vincristine-triggered autophagy in oral cancer cells. Expert Opin Ther Targets. 2015; 19: 455-470.
10. Zhao L, An R, Yang Y, Yang X, Liu H, Yue L, Li X, Lin Y, Reiter RJ, Qu Y. Melatonin alleviates brain injury in mice subjected to cecal ligation and puncture via attenuating inflammation, apoptosis, and oxidative stress: the role of SIRT1 signaling. J Pineal Res. 2015; 59: 230-239.
11. Bonaldi T, Langst G, Strohner R, Becker PB, Bianchi ME. The DNA chaperone HMGB1 facilitates ACF/CHRAC-dependent nucleosome sliding. EMBO J. 2002; 21: 6865-6873.
12. Scaffidi P, Misteli T, Bianchi ME. Release of chromatin protein HMGB1 by necrotic cells triggers inflammation. Nature. 2002; 418: 191-195.
13. Chen M, Liu Y, Varley P, Chang Y, He XX, Huang H, Tang D, Lotze MT, Lin J, Tsung A. High-Mobility Group Box 1 Promotes Hepatocellular Carcinoma Progression through miR-21-Mediated Matrix Metalloproteinase Activity. Cancer Res. 2015; 75: 1645-1656.
14. Chung HW, Jang S, Kim H, Lim JB. Combined targeting of high-mobility group box-1 and interleukin-8 to control micrometastasis potential in gastric cancer. Int J Cancer. 2015; 137: 1598-1609.
15. Huber R, Meier B, Otsuka A, Fenini G, Satoh T, Gehrke S, Widmer D, Levesque MP, Mangana J, Kerl K, Gebhardt C, Fujii H, Nakashima C, et al. Tumour hypoxia promotes melanoma growth and metastasis via High Mobility Group Box-1 and M2-like macrophages. Sci Rep. 2016; 6: 29914.
16. Shi Z, Huang Q, Chen J, Yu P, Wang X, Qiu H, Chen Y, Dong Y. Correlation of HMGB1 expression to progression and poor prognosis of adenocarcinoma and squamous cell/adenosquamous carcinoma of gallbladder. Am J Transl Res. 2015; 7: 2015-2025.
17. Sasahira T, Kirita T, Oue N, Bhawal UK, Yamamoto K, Fujii K, Ohmori H, Luo Y, Yasui W, Bosserhoff AK, Kuniyasu H. High mobility group box-1-inducible melanoma inhibitory activity is associated with nodal metastasis and lymphangiogenesis in oral squamous cell carcinoma. Cancer Sci. 2008; 99: 1806-1812.
18. Wild CA, Brandau S, Lotfi R, Mattheis S, Gu X, Lang S, Bergmann C. HMGB1 is overexpressed in tumor cells and promotes activity of regulatory T cells in patients with head and neck cancer. Oral Oncol. 2012; 48: 409-416.
19. Markopoulos AK. Current aspects on oral squamous cell carcinoma. Open Dent J. 2012; 6: 126-130.
20. Feitelson MA. Parallel epigenetic and genetic changes in the pathogenesis of hepatitis virus-associated hepatocellular carcinoma. Cancer Lett. 2006; 239: 10-20.
21. Su SC, Hsieh MJ, Yang WE, Chung WH, Reiter RJ, Yang SF. Cancer metastasis: Mechanisms of inhibition by melatonin. J Pineal Res. 2017; 62. doi: 10.1111/jpi.12370.
22. Thorgeirsson SS, Grisham JW. Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet. 2002; 31: 339-346.
23. Bascones-Martinez A, Lopez-Duran M, Cano-Sanchez J, Sanchez-Verde L, Diez-Rodriguez A, Aguirre-Echebarria P, Alvarez-Fernandez E, Gonzalez-Moles MA, Bascones-Ilundain J, Muzio LL, Campo-Trapero J. Differences in the expression of five senescence markers in oral cancer, oral leukoplakia and control samples in humans. Oncol Lett. 2012; 3: 1319-1325.
24. Nguyen A, Bhavsar S, Riley E, Caponetti G, Agrawal D. Clinical Value of High Mobility Group Box 1 and the Receptor for Advanced Glycation End-products in Head and Neck Cancer: A Systematic Review. Int Arch Otorhinolaryngol. 2016; 20: 382-389.
25. Wu T, Zhang W, Yang G, Li H, Chen Q, Song R, Zhao L. HMGB1 overexpression as a prognostic factor for survival in cancer: a meta-analysis and systematic review. Oncotarget. 2016; 7: 50417-50427. doi: 10.18632/oncotarget.10413.
26. Wang B, Yeh CB, Lein MY, Su CM, Yang SF, Liu YF, Tang CH. Effects of HMGB1 Polymorphisms on the Susceptibility and Progression of Hepatocellular Carcinoma. Int J Med Sci. 2016; 13: 304-309.
27. Wang D, Qi X, Liu F, Yang C, Jiang W, Wei X, Li X, Mi J, Tian G. A multicenter matched case-control analysis on seven polymorphisms from HMGB1 and RAGE genes in predicting hepatocellular carcinoma risk. Oncotarget. 2017 Feb 8. doi: 10.18632/oncotarget.15202. [Epub ahead of print].
28. Wang JX, Yu HL, Bei SS, Cui ZH, Li ZW, Liu ZJ, Lv YF. Association of HMGB1 Gene Polymorphisms with Risk of Colorectal Cancer in a Chinese Population. Med Sci Monit. 2016; 22: 3419-3425.
29. Wu HH, Liu YF, Yang SF, Lin WL, Chen SC, Han CP, Wang HL, Lin LY, Wang PH. Association of single-nucleotide polymorphisms of high-mobility group box 1 with susceptibility and clinicopathological characteristics of uterine cervical neoplasia in Taiwanese women. Tumour Biol. 2016; 12: 15813-15823.
30. Yue L, Zhang Q, He L, Zhang M, Dong J, Zhao D, Ma H, Pan H, Zheng L. Genetic predisposition of six well-defined polymorphisms in HMGB1/RAGE pathway to breast cancer in a large Han Chinese population. J Cell Mol Med. 2016; 20: 1966-1973.
31. Bao G, Qu F, He L, Zhao H, Wang N, Ji G, He X. Prognostic Significance of Tag SNP rs1045411 in HMGB1 of the Aggressive Gastric Cancer in a Chinese Population. PLoS One. 2016; 11: e0154378.
32. Lu L, Qiu C, Li D, Bai G, Liang J, Yang Q. MicroRNA-505 suppresses proliferation and invasion in hepatoma cells by directly targeting high-mobility group box 1. Life Sci. 2016;
33. Yang Q, Jia C, Wang P, Xiong M, Cui J, Li L, Wang W, Wu Q, Chen Y, Zhang T. MicroRNA-505 identified from patients with essential hypertension impairs endothelial cell migration and tube formation. Int J Cardiol. 2014; 177: 925-934.
34. Griffiths-Jones S. miRBase: the microRNA sequence database. Methods Mol Biol. 2006; 342: 129-138.
35. Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014; 42: D92-97.
36. Cancer Genome Atlas Research N, Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, Ellrott K, Shmulevich I, Sander C, Stuart JM. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet. 2013; 45: 1113-1120.
37. Genomes Project C, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA. An integrated map of genetic variation from 1,092 human genomes. Nature. 2012; 491: 56-65.
38. Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005; 21: 263-265.
39. Consortium GT. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013; 45: 580-585.
40. Kellis M, Wold B, Snyder MP, Bernstein BE, Kundaje A, Marinov GK, Ward LD, Birney E, Crawford GE, Dekker J, Dunham I, Elnitski LL, Farnham PJ, et al. Defining functional DNA elements in the human genome. Proc Natl Acad Sci U S A. 2014; 111: 6131-6138.
41. van der Krieken SE, Popeijus HE, Mensink RP, Plat J. CCAAT/enhancer binding protein beta in relation to ER stress, inflammation, and metabolic disturbances. Biomed Res Int. 2015; 2015: 324815.
42. Calvisi DF, Simile MM, Ladu S, Frau M, Evert M, Tomasi ML, Demartis MI, Daino L, Seddaiu MA, Brozzetti S, Feo F, Pascale RM. Activation of v-Myb avian myeloblastosis viral oncogene homolog-like2 (MYBL2)-LIN9 complex contributes to human hepatocarcinogenesis and identifies a subset of hepatocellular carcinoma with mutant p53. Hepatology. 2011; 53: 1226-1236.
43. Su SC, Lin CW, Yang WE, Fan WL, Yang SF. The urokinase-type plasminogen activator (uPA) system as a biomarker and therapeutic target in human malignancies. Expert Opin Ther Targets. 2016; 20: 551-566.
44. Sadikovic B, Al-Romaih K, Squire JA, Zielenska M. Cause and consequences of genetic and epigenetic alterations in human cancer. Curr Genomics. 2008; 9: 394-408.
45. Sasahira T, Kirita T, Kuniyasu H. Update of molecular pathobiology in oral cancer: a review. Int J Clin Oncol. 2014; 19: 431-436.
46. Chattopadhyay I, Singh A, Phukan R, Purkayastha J, Kataki A, Mahanta J, Saxena S, Kapur S. Genome-wide analysis of chromosomal alterations in patients with esophageal squamous cell carcinoma exposed to tobacco and betel quid from high-risk area in India. Mutat Res. 2010; 696: 130-138.
47. Domanska K, Malander S, Staaf J, Karlsson A, Borg A, Jonsson G, Nilbert M. Genetic profiles distinguish different types of hereditary ovarian cancer. Oncol Rep. 2010; 24: 885-895.
48. Yamamoto S, Midorikawa Y, Morikawa T, Nishimura Y, Sakamoto H, Ishikawa S, Akagi K, Aburatani H. Identification of chromosomal aberrations of metastatic potential in colorectal carcinoma. Genes Chromosomes Cancer. 2010; 49: 487-496.
49. Gollin SM. Cytogenetic alterations and their molecular genetic correlates in head and neck squamous cell carcinoma: a next generation window to the biology of disease. Genes Chromosomes Cancer. 2014; 53: 972-990.
50. Nowacka-Zawisza M, Forma E, Walczak M, Rozanski W, Brys M, Krajewska WM. Loss of heterozygosity for chromosomal regions 15q14-21.1, 17q21.31, and 13q12.3-13.1 and its relevance for prostate cancer. Med Oncol. 2015; 32: 246.
51. Ekanayaka RP, Tilakaratne WM. Oral submucous fibrosis: review on mechanisms of malignant transformation. Oral Surg Oral Med Oral Pathol Oral Radiol. 2016.
52. Brezniceanu ML, Volp K, Bosser S, Solbach C, Lichter P, Joos S, Zornig M. HMGB1 inhibits cell death in yeast and mammalian cells and is abundantly expressed in human breast carcinoma. FASEB J. 2003; 17: 1295-1297.
53. Poser I, Golob M, Buettner R, Bosserhoff AK. Upregulation of HMG1 leads to melanoma inhibitory activity expression in malignant melanoma cells and contributes to their malignancy phenotype. Mol Cell Biol. 2003; 23: 2991-2998.
54. Akaike H, Kono K, Sugai H, Takahashi A, Mimura K, Kawaguchi Y, Fujii H. Expression of high mobility group box chromosomal protein-1 (HMGB-1) in gastric cancer. Anticancer Res. 2007; 27: 449-457.
55. Volp K, Brezniceanu ML, Bosser S, Brabletz T, Kirchner T, Gottel D, Joos S, Zornig M. Increased expression of high mobility group box 1 (HMGB1) is associated with an elevated level of the antiapoptotic c-IAP2 protein in human colon carcinomas. Gut. 2006; 55: 234-242.
56. Kang R, Zhang Q, Zeh HJ 3rd, Lotze MT, Tang D. HMGB1 in cancer: good, bad, or both? Clin Cancer Res. 2013; 19: 4046-4057.
57. Chen S, Sun KX, Liu BL, Zong ZH, Zhao Y. MicroRNA-505 functions as a tumor suppressor in endometrial cancer by targeting TGF-alpha. Mol Cancer. 2016; 15: 11.
58. Matamala N, Vargas MT, Gonzalez-Campora R, Minambres R, Arias JI, Menendez P, Andres-Leon E, Gomez-Lopez G, Yanowsky K, Calvete-Candenas J, Inglada-Perez L, Martinez-Delgado B, Benitez J. Tumor microRNA expression profiling identifies circulating microRNAs for early breast cancer detection. Clin Chem. 2015; 61: 1098-1106.
59. Lu L, Qiu C, Li D, Bai G, Liang J, Yang Q. MicroRNA-505 suppresses proliferation and invasion in hepatoma cells by directly targeting high-mobility group box 1. Life Sci. 2016; 157: 12-18.
60. Zhao CB, Bao JM, Lu YJ, Zhao T, Zhou XH, Zheng DY, Zhao SC. Co-expression of RAGE and HMGB1 is associated with cancer progression and poor patient outcome of prostate cancer. Am J Cancer Res. 2014; 4: 369-377.
61. Lin KH, Lin CY, Liu CC, Chou MY, Lin JK. Arecoline N-oxide: its mutagenicity and possible role as ultimate carcinogen in areca oral carcinogenesis. J Agric Food Chem. 2011; 59: 3420-3428.
62. Ho SM, Johnson A, Tarapore P, Janakiram V, Zhang X, Leung YK. Environmental epigenetics and its implication on disease risk and health outcomes. ILAR J. 2012; 53: 289-305.
63. Cheng HL, Liu YF, Su CW, Su SC, Chen MK, Yang SF, Lin CW. Functional genetic variant in the Kozak sequence of WW domain-containing oxidoreductase (WWOX) gene is associated with oral cancer risk. Oncotarget. 2016; 7: 69384-69396. doi: 10.18632/oncotarget.12082.
64. Chou CH, Chou YE, Chuang CY, Yang SF, Lin CW. Combined effect of genetic polymorphisms of AURKA and environmental factors on oral cancer development in Taiwan. PLoS One. 2017; 12: e0171583.
65. Ho HY, Lin CW, Chien MH, Reiter RJ, Su SC, Hsieh YH, Yang SF. Melatonin suppresses TPA-induced metastasis by downregulating matrix metalloproteinase-9 expression through JNK/SP-1 signaling in nasopharyngeal carcinoma. J Pineal Res. 2016; 61: 479-492.
66. Su SC, Hsieh MJ, Liu YF, Chou YE, Lin CW, Yang SF. ADAMTS14 Gene Polymorphism and Environmental Risk in the Development of Oral Cancer. PLoS One. 2016; 11: e0159585.
67. Zavras AI, Yoon AJ, Chen MK, Lin CW, Yang SF. Metallothionein-1 genotypes in the risk of oral squamous cell carcinoma. Ann Surg Oncol. 2011; 18: 1478-1483.
68. Ward LD, Kellis M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2016; 44: D877-881.
69. Pazin MJ. Using the ENCODE Resource for Functional Annotation of Genetic Variants. Cold Spring Harb Protoc. 2015; 2015: 522-536.
70. Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012; 2: 401-404.
71. Goldman M, Craft B, Swatloski T, Cline M, Morozova O, Diekhans M, Haussler D, Zhu J. The UCSC Cancer Genomics Browser: update 2015. Nucleic Acids Res. 2015; 43: D812-817.
72. Kruger J, Rehmsmeier M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res. 2006; 34: W451-454.

